Abstract
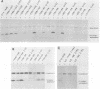
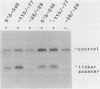
To examine the internal organization of the promoter of the Xenopus laevis rRNA gene, we constructed a series of linker-scanning mutants that traverse the rDNA initiation region. The mutant genes, which have 3 to 11 clustered base substitutions set within an otherwise unaltered rDNA promoter sequence, were injected into Xenopus oocyte nuclei, and their transcriptional capacity was assessed by S1 nuclease analysis of the resultant RNA. The data demonstrate that there are two essential promoter domains, the distal boundaries of which coincide with the promoter boundaries established previously by analysis of 5' and 3' deletion mutants. The upstream promoter domain is relatively small and extends from residues ca. -140 to -128. The downstream domain is considerably larger, encompassing residues ca. -36 to +10, and exactly corresponds in both size and position to the mammalian minimal promoter region. The Xenopus rDNA sequence between these two essential domains has a much smaller effect on the level of transcriptional initiation. In light of the fact that a large portion of this intervening region consists of a segment (residues -114 to -72) that is duplicated many times in the upstream spacer to form an rDNA enhancer sequence, it is noteworthy that a "-115/-77 linker scanner," in which virtually this entire segment is replaced by a polylinker sequence, has full promoter activity in the injected Xenopus borealis oocytes. Analysis of a parallel series of spacing change linker-scanning mutants revealed the unexpected result that the relative positions of the upstream and downstream promoter domains are very critical: all spacing alterations of more than 2 base pairs within this 100-base-pair region virtually abolish promoter activity. We conclude that the factors that bind to these two distant promoter domains must interact in a very precise stereospecific manner.
Full text
PDF



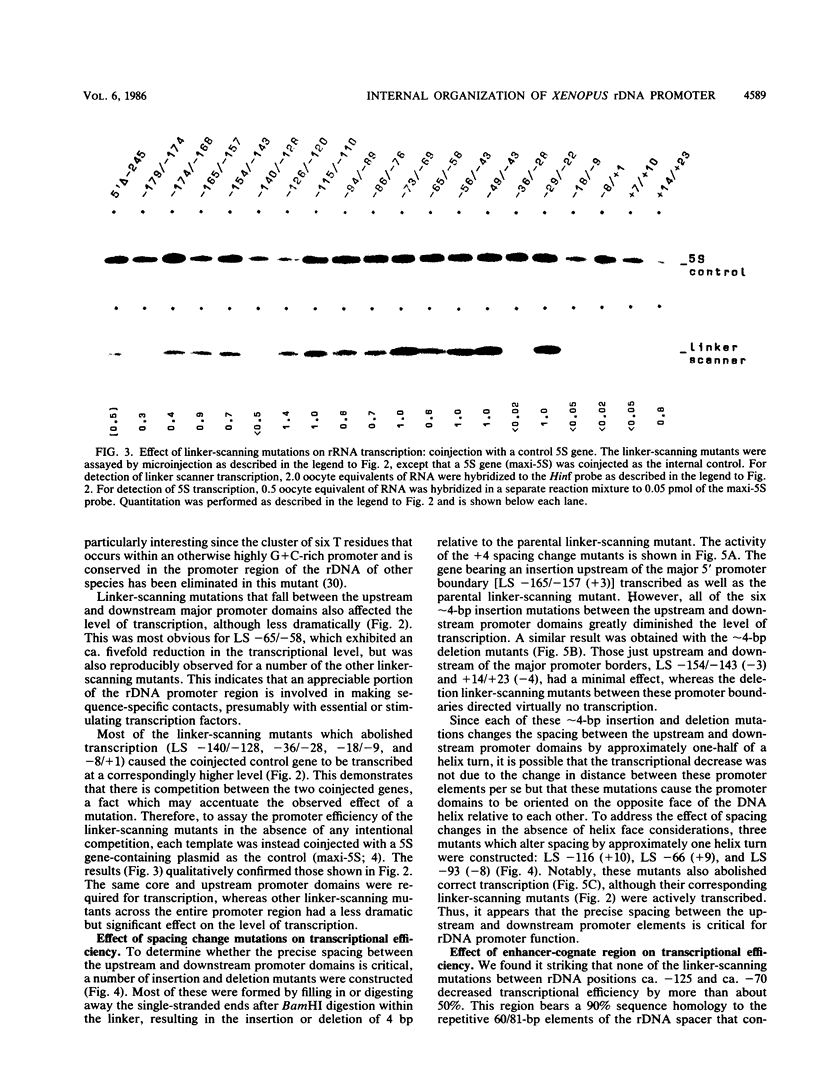




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bakken A., Morgan G., Sollner-Webb B., Roan J., Busby S., Reeder R. H. Mapping of transcription initiation and termination signals on Xenopus laevis ribosomal DNA. Proc Natl Acad Sci U S A. 1982 Jan;79(1):56–60. doi: 10.1073/pnas.79.1.56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bateman E., Iida C. T., Kownin P., Paule M. R. Footprinting of ribosomal RNA genes by transcription initiation factor and RNA polymerase I. Proc Natl Acad Sci U S A. 1985 Dec;82(23):8004–8008. doi: 10.1073/pnas.82.23.8004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benoist C., O'Hare K., Breathnach R., Chambon P. The ovalbumin gene-sequence of putative control regions. Nucleic Acids Res. 1980 Jan 11;8(1):127–142. doi: 10.1093/nar/8.1.127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bogenhagen D. F., Brown D. D. Nucleotide sequences in Xenopus 5S DNA required for transcription termination. Cell. 1981 Apr;24(1):261–270. doi: 10.1016/0092-8674(81)90522-5. [DOI] [PubMed] [Google Scholar]
- Breathnach R., Chambon P. Organization and expression of eucaryotic split genes coding for proteins. Annu Rev Biochem. 1981;50:349–383. doi: 10.1146/annurev.bi.50.070181.002025. [DOI] [PubMed] [Google Scholar]
- Corden J., Wasylyk B., Buchwalder A., Sassone-Corsi P., Kedinger C., Chambon P. Promoter sequences of eukaryotic protein-coding genes. Science. 1980 Sep 19;209(4463):1406–1414. doi: 10.1126/science.6251548. [DOI] [PubMed] [Google Scholar]
- Dunaway M., Reeder R. H. DNase I footprinting shows three protected regions in the promoter of the rRNA genes of Xenopus laevis. Mol Cell Biol. 1985 Feb;5(2):313–319. doi: 10.1128/mcb.5.2.313. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Efstratiadis A., Posakony J. W., Maniatis T., Lawn R. M., O'Connell C., Spritz R. A., DeRiel J. K., Forget B. G., Weissman S. M., Slightom J. L. The structure and evolution of the human beta-globin gene family. Cell. 1980 Oct;21(3):653–668. doi: 10.1016/0092-8674(80)90429-8. [DOI] [PubMed] [Google Scholar]
- Grummt I. Nucleotide sequence requirements for specific initiation of transcription by RNA polymerase I. Proc Natl Acad Sci U S A. 1982 Nov;79(22):6908–6911. doi: 10.1073/pnas.79.22.6908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grummt I., Roth E., Paule M. R. Ribosomal RNA transcription in vitro is species specific. Nature. 1982 Mar 11;296(5853):173–174. doi: 10.1038/296173a0. [DOI] [PubMed] [Google Scholar]
- Haltiner M. M., Smale S. T., Tjian R. Two distinct promoter elements in the human rRNA gene identified by linker scanning mutagenesis. Mol Cell Biol. 1986 Jan;6(1):227–235. doi: 10.1128/mcb.6.1.227. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hochschild A., Ptashne M. Cooperative binding of lambda repressors to sites separated by integral turns of the DNA helix. Cell. 1986 Mar 14;44(5):681–687. doi: 10.1016/0092-8674(86)90833-0. [DOI] [PubMed] [Google Scholar]
- Iida C. T., Kownin P., Paule M. R. Ribosomal RNA transcription: proteins and DNA sequences involved in preinitiation complex formation. Proc Natl Acad Sci U S A. 1985 Mar;82(6):1668–1672. doi: 10.1073/pnas.82.6.1668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kohorn B. D., Rae P. M. Localization of DNA sequences promoting RNA polymerase I activity in Drosophila. Proc Natl Acad Sci U S A. 1983 Jun;80(11):3265–3268. doi: 10.1073/pnas.80.11.3265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kovacs B. J., Butterworth P. H. The effect of changing the distance between the TATA-box and cap site by up to three base pairs on the selection of the transcriptional start site of a cloned eukaryotic gene in vitro and in vivo. Nucleic Acids Res. 1986 Mar 25;14(6):2429–2442. doi: 10.1093/nar/14.6.2429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Learned R. M., Smale S. T., Haltiner M. M., Tjian R. Regulation of human ribosomal RNA transcription. Proc Natl Acad Sci U S A. 1983 Jun;80(12):3558–3562. doi: 10.1073/pnas.80.12.3558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- McKnight S. L. Functional relationships between transcriptional control signals of the thymidine kinase gene of herpes simplex virus. Cell. 1982 Dec;31(2 Pt 1):355–365. doi: 10.1016/0092-8674(82)90129-5. [DOI] [PubMed] [Google Scholar]
- McKnight S. L., Kingsbury R. Transcriptional control signals of a eukaryotic protein-coding gene. Science. 1982 Jul 23;217(4557):316–324. doi: 10.1126/science.6283634. [DOI] [PubMed] [Google Scholar]
- Miller K. G., Tower J., Sollner-Webb B. A complex control region of the mouse rRNA gene directs accurate initiation by RNA polymerase I. Mol Cell Biol. 1985 Mar;5(3):554–562. doi: 10.1128/mcb.5.3.554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moss T. A transcriptional function for the repetitive ribosomal spacer in Xenopus laevis. Nature. 1983 Mar 17;302(5905):223–228. doi: 10.1038/302223a0. [DOI] [PubMed] [Google Scholar]
- Moss T. Transcription of cloned Xenopus laevis ribosomal DNA microinjected into Xenopus oocytes, and the identification of an RNA polymerase I promoter. Cell. 1982 Oct;30(3):835–842. doi: 10.1016/0092-8674(82)90288-4. [DOI] [PubMed] [Google Scholar]
- Reeder R. H. Enhancers and ribosomal gene spacers. Cell. 1984 Sep;38(2):349–351. doi: 10.1016/0092-8674(84)90489-6. [DOI] [PubMed] [Google Scholar]
- Reeder R. H., Roan J. G., Dunaway M. Spacer regulation of Xenopus ribosomal gene transcription: competition in oocytes. Cell. 1983 Dec;35(2 Pt 1):449–456. doi: 10.1016/0092-8674(83)90178-2. [DOI] [PubMed] [Google Scholar]
- Smale S. T., Tjian R. Transcription of herpes simplex virus tk sequences under the control of wild-type and mutant human RNA polymerase I promoters. Mol Cell Biol. 1985 Feb;5(2):352–362. doi: 10.1128/mcb.5.2.352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sollner-Webb B., McKnight S. L. Accurate transcription of cloned Xenopus rRNA genes by RNA polymerase I: demonstration by S1 nuclease mapping. Nucleic Acids Res. 1982 Jun 11;10(11):3391–3405. doi: 10.1093/nar/10.11.3391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sollner-Webb B., Tower J. Transcription of cloned eukaryotic ribosomal RNA genes. Annu Rev Biochem. 1986;55:801–830. doi: 10.1146/annurev.bi.55.070186.004101. [DOI] [PubMed] [Google Scholar]
- Sollner-Webb B., Wilkinson J. A., Roan J., Reeder R. H. Nested control regions promote Xenopus ribosomal RNA synthesis by RNA polymerase I. Cell. 1983 Nov;35(1):199–206. doi: 10.1016/0092-8674(83)90222-2. [DOI] [PubMed] [Google Scholar]
- Sommerville J. RNA polymerase I promoters and transcription factors. Nature. 1984 Jul 19;310(5974):189–190. doi: 10.1038/310189a0. [DOI] [PubMed] [Google Scholar]
- Takahashi K., Vigneron M., Matthes H., Wildeman A., Zenke M., Chambon P. Requirement of stereospecific alignments for initiation from the simian virus 40 early promoter. Nature. 1986 Jan 9;319(6049):121–126. doi: 10.1038/319121a0. [DOI] [PubMed] [Google Scholar]
- Tower J., Culotta V. C., Sollner-Webb B. Factors and nucleotide sequences that direct ribosomal DNA transcription and their relationship to the stable transcription complex. Mol Cell Biol. 1986 Oct;6(10):3451–3462. doi: 10.1128/mcb.6.10.3451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vieira J., Messing J. The pUC plasmids, an M13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene. 1982 Oct;19(3):259–268. doi: 10.1016/0378-1119(82)90015-4. [DOI] [PubMed] [Google Scholar]
- Wilkinson J. K., Sollner-Webb B. Transcription of Xenopus ribosomal RNA genes by RNA polymerase I in vitro. J Biol Chem. 1982 Dec 10;257(23):14375–14383. [PubMed] [Google Scholar]
- Windle J., Sollner-Webb B. Upstream domains of the Xenopus laevis rDNA promoter are revealed in microinjected oocytes. Mol Cell Biol. 1986 Apr;6(4):1228–1234. doi: 10.1128/mcb.6.4.1228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamamoto O., Takakusa N., Mishima Y., Kominami R., Muramatsu M. Determination of the promoter region of mouse ribosomal RNA gene by an in vitro transcription system. Proc Natl Acad Sci U S A. 1984 Jan;81(2):299–303. doi: 10.1073/pnas.81.2.299. [DOI] [PMC free article] [PubMed] [Google Scholar]