Abstract
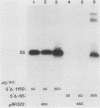
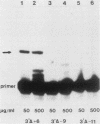
The DNA sequences involved in promoting transcription of the Xenopus laevis rRNA genes were determined by microinjecting a series of deletion mutants into oocyte nuclei. A very small promoter region is sufficient to direct efficient transcription when templates are microinjected at high rDNA concentration, since 5'delta- 9 and 3'delta +6 templates are fully active. However, as the concentration of injected template is decreased, an increasing requirement for upstream domains, extending to nucleotide approximately -170, is observed. The major downstream border of the required region does not change. This apparently expanding 5' promoter border results from the fact that, as the rDNA concentration is decreased, transcription from templates lacking the upstream promoter domain falls off much more sharply than does transcription from a complete promoter. In fact, the deleted promoters are virtually inactive below a threshold rDNA concentration. It is indeed the rDNA concentration that is important, for coinjected vector DNA does not increase the level of transcription obtained from low concentrations of the 5' deletions. From these data we conclude that polymerase I transcription factors can recognize and initiate transcription from a small core promoter domain, but that sequences extending upstream to nucleotide approximately -170 increase the efficiency of initiation. A model is presented that could account for these results.
Full text
PDF

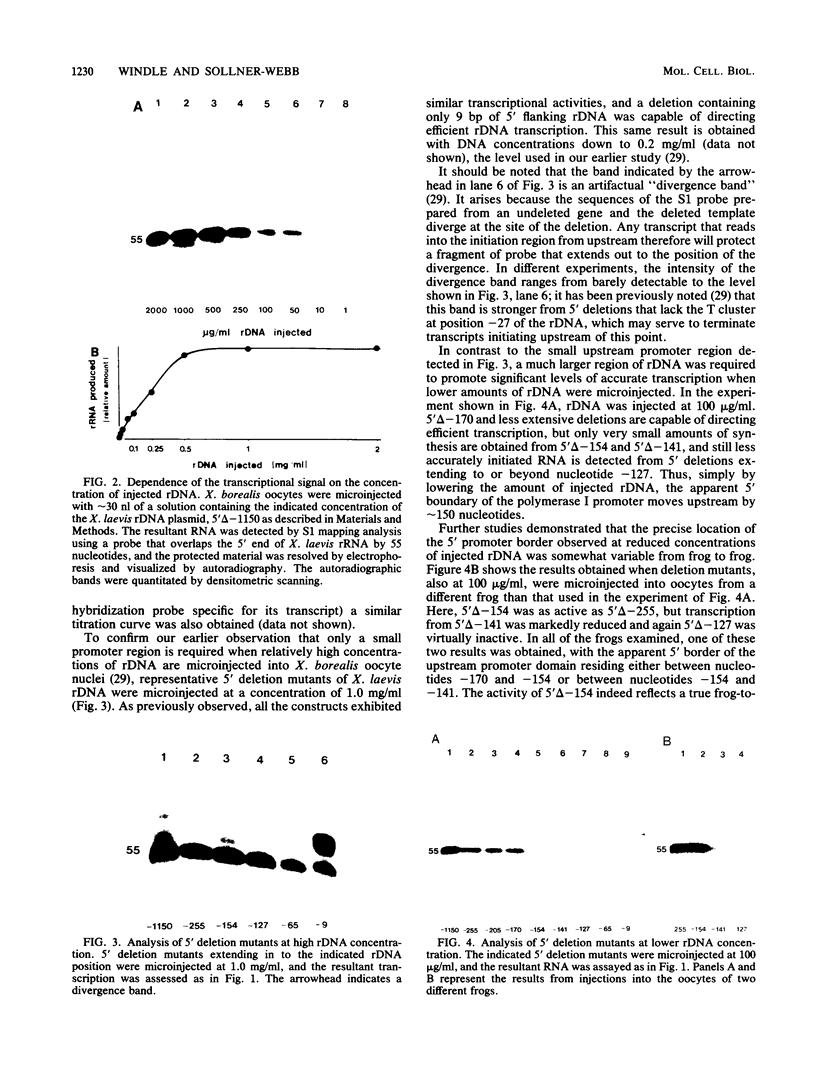




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bakken A., Morgan G., Sollner-Webb B., Roan J., Busby S., Reeder R. H. Mapping of transcription initiation and termination signals on Xenopus laevis ribosomal DNA. Proc Natl Acad Sci U S A. 1982 Jan;79(1):56–60. doi: 10.1073/pnas.79.1.56. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boseley P., Moss T., Mächler M., Portmann R., Birnstiel M. Sequence organization of the spacer DNA in a ribosomal gene unit of Xenopus laevis. Cell. 1979 May;17(1):19–31. doi: 10.1016/0092-8674(79)90291-5. [DOI] [PubMed] [Google Scholar]
- Botchan P., Reeder R. H., Dawid I. B. Restriction analysis of the nontranscribed spacers of Xenopus laevis ribosomal DNA. Cell. 1977 Jul;11(3):599–607. doi: 10.1016/0092-8674(77)90077-0. [DOI] [PubMed] [Google Scholar]
- Brand A. H., Breeden L., Abraham J., Sternglanz R., Nasmyth K. Characterization of a "silencer" in yeast: a DNA sequence with properties opposite to those of a transcriptional enhancer. Cell. 1985 May;41(1):41–48. doi: 10.1016/0092-8674(85)90059-5. [DOI] [PubMed] [Google Scholar]
- Brown D. D., Gurdon J. B. Cloned single repeating units of 5S DNA direct accurate transcription of 5S RNA when injected into Xenopus oocytes. Proc Natl Acad Sci U S A. 1978 Jun;75(6):2849–2853. doi: 10.1073/pnas.75.6.2849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Das G. C., Niyogi S. K., Salzman N. P. SV40 promoters and their regulation. Prog Nucleic Acid Res Mol Biol. 1985;32:217–236. doi: 10.1016/s0079-6603(08)60349-9. [DOI] [PubMed] [Google Scholar]
- Feldman J. B., Hicks J. B., Broach J. R. Identification of sites required for repression of a silent mating type locus in yeast. J Mol Biol. 1984 Oct 5;178(4):815–834. doi: 10.1016/0022-2836(84)90313-9. [DOI] [PubMed] [Google Scholar]
- Gargiulo G., Razvi F., Worcel A. Assembly of transcriptionally active chromatin in Xenopus oocytes requires specific DNA binding factors. Cell. 1984 Sep;38(2):511–521. doi: 10.1016/0092-8674(84)90506-3. [DOI] [PubMed] [Google Scholar]
- Grummt I. Nucleotide sequence requirements for specific initiation of transcription by RNA polymerase I. Proc Natl Acad Sci U S A. 1982 Nov;79(22):6908–6911. doi: 10.1073/pnas.79.22.6908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iida C. T., Kownin P., Paule M. R. Ribosomal RNA transcription: proteins and DNA sequences involved in preinitiation complex formation. Proc Natl Acad Sci U S A. 1985 Mar;82(6):1668–1672. doi: 10.1073/pnas.82.6.1668. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones P. B., Galeazzi D. R., Fisher J. M., Whitlock J. P., Jr Control of cytochrome P1-450 gene expression by dioxin. Science. 1985 Mar 22;227(4693):1499–1502. doi: 10.1126/science.3856321. [DOI] [PubMed] [Google Scholar]
- Kmiec E. B., Worcel A. The positive transcription factor of the 5S RNA gene induces a 5S DNA-specific gyration in Xenopus oocyte extracts. Cell. 1985 Jul;41(3):945–953. doi: 10.1016/s0092-8674(85)80075-1. [DOI] [PubMed] [Google Scholar]
- Kohorn B. D., Rae P. M. A component of Drosophila RNA polymerase I promoter lies within the rRNA transcription unit. Nature. 1983 Jul 14;304(5922):179–181. doi: 10.1038/304179a0. [DOI] [PubMed] [Google Scholar]
- Kohorn B. D., Rae P. M. Localization of DNA sequences promoting RNA polymerase I activity in Drosophila. Proc Natl Acad Sci U S A. 1983 Jun;80(11):3265–3268. doi: 10.1073/pnas.80.11.3265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Labhart P., Reeder R. H. Enhancer-like properties of the 60/81 bp elements in the ribosomal gene spacer of Xenopus laevis. Cell. 1984 May;37(1):285–289. doi: 10.1016/0092-8674(84)90324-6. [DOI] [PubMed] [Google Scholar]
- Learned R. M., Smale S. T., Haltiner M. M., Tjian R. Regulation of human ribosomal RNA transcription. Proc Natl Acad Sci U S A. 1983 Jun;80(12):3558–3562. doi: 10.1073/pnas.80.12.3558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller K. G., Tower J., Sollner-Webb B. A complex control region of the mouse rRNA gene directs accurate initiation by RNA polymerase I. Mol Cell Biol. 1985 Mar;5(3):554–562. doi: 10.1128/mcb.5.3.554. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moss T. A transcriptional function for the repetitive ribosomal spacer in Xenopus laevis. Nature. 1983 Mar 17;302(5905):223–228. doi: 10.1038/302223a0. [DOI] [PubMed] [Google Scholar]
- Moss T. Transcription of cloned Xenopus laevis ribosomal DNA microinjected into Xenopus oocytes, and the identification of an RNA polymerase I promoter. Cell. 1982 Oct;30(3):835–842. doi: 10.1016/0092-8674(82)90288-4. [DOI] [PubMed] [Google Scholar]
- Probst E., Kressmann A., Birnstiel M. L. Expression of sea urchin histone genes in the oocyte of Xenopus laevis. J Mol Biol. 1979 Dec 15;135(3):709–732. doi: 10.1016/0022-2836(79)90173-6. [DOI] [PubMed] [Google Scholar]
- Reeder R. H., Roan J. G., Dunaway M. Spacer regulation of Xenopus ribosomal gene transcription: competition in oocytes. Cell. 1983 Dec;35(2 Pt 1):449–456. doi: 10.1016/0092-8674(83)90178-2. [DOI] [PubMed] [Google Scholar]
- Sanger F., Coulson A. R. The use of thin acrylamide gels for DNA sequencing. FEBS Lett. 1978 Mar 1;87(1):107–110. doi: 10.1016/0014-5793(78)80145-8. [DOI] [PubMed] [Google Scholar]
- Smale S. T., Tjian R. Transcription of herpes simplex virus tk sequences under the control of wild-type and mutant human RNA polymerase I promoters. Mol Cell Biol. 1985 Feb;5(2):352–362. doi: 10.1128/mcb.5.2.352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sollner-Webb B., McKnight S. L. Accurate transcription of cloned Xenopus rRNA genes by RNA polymerase I: demonstration by S1 nuclease mapping. Nucleic Acids Res. 1982 Jun 11;10(11):3391–3405. doi: 10.1093/nar/10.11.3391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sollner-Webb B., Reeder R. H. The nucleotide sequence of the initiation and termination sites for ribosomal RNA transcription in X. laevis. Cell. 1979 Oct;18(2):485–499. doi: 10.1016/0092-8674(79)90066-7. [DOI] [PubMed] [Google Scholar]
- Sollner-Webb B., Wilkinson J. A., Roan J., Reeder R. H. Nested control regions promote Xenopus ribosomal RNA synthesis by RNA polymerase I. Cell. 1983 Nov;35(1):199–206. doi: 10.1016/0092-8674(83)90222-2. [DOI] [PubMed] [Google Scholar]
- Trendelenburg M. F., Gurdon J. B. Transcription of cloned Xenopus ribosomal genes visualised after injection into oocyte nuclei. Nature. 1978 Nov 16;276(5685):292–294. doi: 10.1038/276292a0. [DOI] [PubMed] [Google Scholar]
- Yamamoto O., Takakusa N., Mishima Y., Kominami R., Muramatsu M. Determination of the promoter region of mouse ribosomal RNA gene by an in vitro transcription system. Proc Natl Acad Sci U S A. 1984 Jan;81(2):299–303. doi: 10.1073/pnas.81.2.299. [DOI] [PMC free article] [PubMed] [Google Scholar]