Abstract
Porcine pleuropneumonia is a highly contagious respiratory disease that causes great economic losses worldwide. In this study, we aimed to explore the underlying relationship between infection and injury by investigation of the whole porcine genome expression profiles of swine lung tissues post-inoculated with experimentally Actinobacillus pleuropneumoniae. Expression profiling experiments of the control group and the treatment group were conducted using a commercially available Agilent Porcine Genechip including 43,603 probe sets. Microarray analysis was conducted on profiles of lung from challenged versus non-challenged swine. We found 11,929 transcripts, identified as differentially expressed at the p ≤0.01 level. There were 1188 genes annotated as swine genes in the GenBank Data Base. GO term analysis identified a total of 89 biological process categories, 82 cellular components and 182 molecular functions that were significantly affected, and at least 27 biological process categories that were related to the host immune response. Gene set enrichment analysis identified 13 pathways that were significantly associated with host response. Many proinflammatory-inflammatory cytokines were activated and involved in the regulation of the host defense response at the site of inflammation; while the cytokines involved in regulation of the host immune response were suppressed. All changes of genes and pathways of induced or repressed expression not only led to a decrease in antigenic peptides presented to T lymphocytes by APCs via the MHC and alleviated immune response injury induced by infection, but also stimulated stem cells to produce granulocytes (neutrophils, eosinophils, and basophils) and monocyte, and promote neutrophils and macrophages to phagocytose bacterial and foreign antigen at the site of inflammation. The defense function of swine infection with Actinobacillus pleuropneumoniae was improved, while its immune function was decreased.
Keywords: porcine pleuropneumonia, infection, injury, Actinobacillus pleuropneumoniae, agilent porcine genechip, microarray analyses, cytokine, host defense response
1. Introduction
Porcine pleuropneumonia (PP) is a highly contagious respiratory disease that causes great economic losses worldwide [1]. The disease, which occurs in swine of all ages, is highly infectious, often fatal, and characterized by necrotizing, hemorrhagic bronchopneumonia and serofibrinous pleuritis [1]. Actinobacillus pleuropneumoniae (APP) is the causative agent of PP and can spread quickly by air-borne particles and/or touching a contaminated surface, and often kills infected animals in the acute phase when extensive lung hemorrhage and necrosis occur. Swine that survive often develop pleurisy, the sequelaes of local necrosis of the pleura, or became healthy carriers of APP.
The porcing lung infected with APP has previously been reported to result in local production of proinflammatory proteins or to mRNA encoding the cytokines interleukin (IL)-1α, IL-1α, IL-6 and the chemokine IL-8 [2]. Likewise, bioactive protein and/or mRNA code IL10, IL12p35, TNF-α and INF-α have shown to be up-regulated after infection with APP in vivo or in vitro [2–4]. Using cDNA microarrays, Moser and co-workers found 307 anonymous transcripts in blood leukocytes from swine that were significantly affected by experimental infection with APP [5]. Hedegaard et al. investigated the molecular characterization of the early response in pigs to experimental infection with APP serotype 5B, using cDNA microarrays [6]. In this study, two-colour microarray analysis was conducted to identify genes being significantly differently expressed in non-inflamed lung tissue compared with inflamed lung tissue sampled from the same animal [6]. The samples of lung tissue were studied by manual hybridization to the pig array DIAS_PIG_27K2 that contains 5375 PCR products amplified from unique cDNA clones [6]. Hedegaard and co-workers found three subsets of genes consistently expressed at different levels depending upon the infection status, and a total of 357 genes differed significantly in their expression levels between infected and non-infected lung tissue from infected versus non-infected animals [6]. Mortensen et al. studied the local transcriptional response in different locations of lung from pigs experimentally infected with the respiratory pathogen APP 5B, using porcine cDNA microarrays (DJF Pig 55 K v1) representing approximately 20,000 porcine genes printed in duplicate [7]. Within the lung, Mortensen and co-workers found a clear division of induced genes as, in unaffected areas a large part of differently expressed genes were involved in systemic reaction to infections, while differently expressed genes in necrotic areas were mainly concerned with homeostasis regulation [7]. However, a limited number of genes relative to the whole Porcine Genome have been studied in previous documents by using cDNA microarrays [5–7]. Thus, transcriptional profiling of whole porcine genome in lung tissue sampled from inoculated versus non-inoculated swine would lead to greater knowledge of the host response dynamics to bacterial infection in the lung. This knowledge is important to obtain a more complete picture of the lung-specific host reactions in the pathogenesis of respiratory infection.
In the present study, the Agilent Whole Porcine Genome Oligo (4 × 44 K) Microarrays (one-color platform), which is a commercially available Agilent Porcine Genechip that included 43,603 probe sets, were used to detect the changes in gene expression of infected pigs’ lungs from non-inoculated animals. Ten transcripts (top six up-regulated and top four down-regulated in microarray data) were selected to verify the accuracy and reproducibility of the microarray data by real-time qRT-PCR.
2. Results
2.1. Clinical Symptoms and Necropsy Findings
The symptoms of lung lesions in the TG were typical after swine infected with APP. Swine showed hyperthermia (40.6–42.0 °C), dyspnea and anorexia after inoculation with APP 24–48 h. Two swine died with respiratory distress at post-inoculation 36–48 h. In the autopsy, the lungs were found to be severely damaged by acute, multifocal, fibrino-necrotizing and hemorrhagic pneumonia complicated by acute diffuse fibrinous pleuritis. The tracheobronchial lymphoid nodes were enlarged and congested.
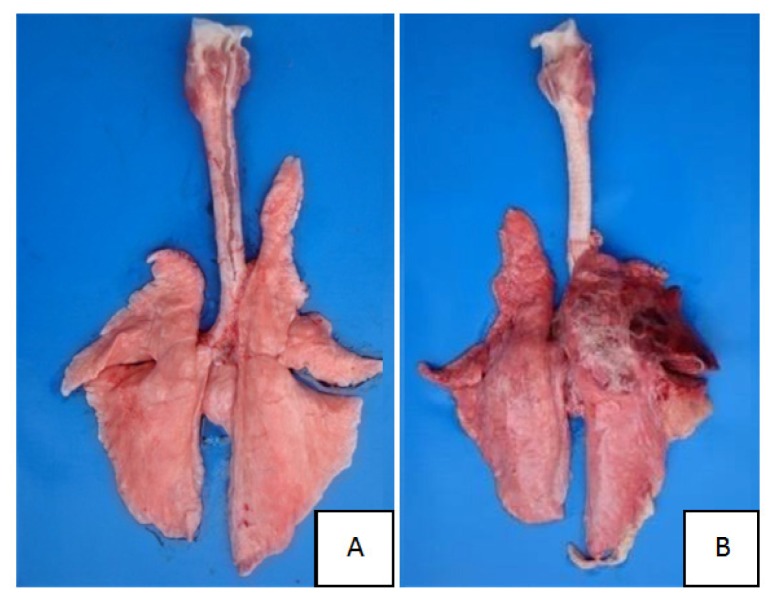
No lesions were observed in CG lung (Figures 1A and 2). Lung showed swelling, bleeding and fibrinous exudate sticking to the lung surface in TG (Figure 1B). The histopathologic changes were characterized by hemorrhage, lymphocyte infiltration, fibrinous exudation vascular thrombosis, necrotic focus and edema in TG (Figure 3).
Figure 1.
(A) Normal lung from healthy swine; (B) Damaged lung after APP infection (Lung in TG showing swelling, bleeding and fibrinous exudate sticking to the lung surface; while no lesion in CG).
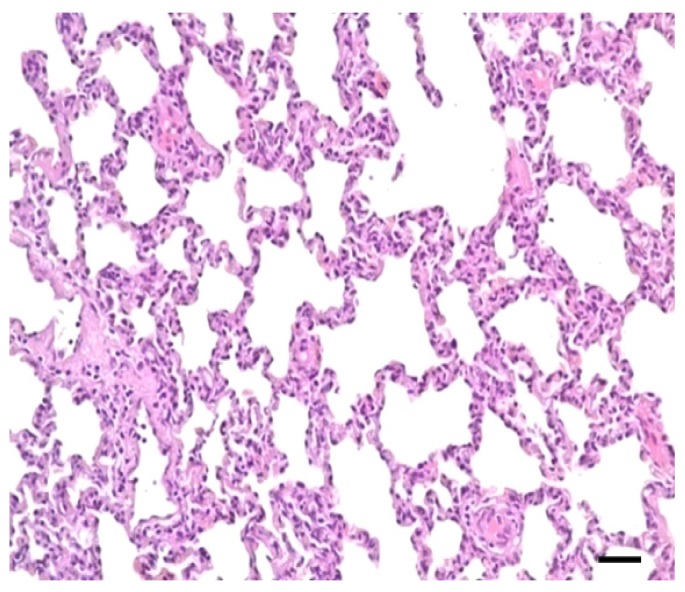
Figure 2.
No lesions were observed in CG lung tissue (scale bar = 50 μm, 200×).
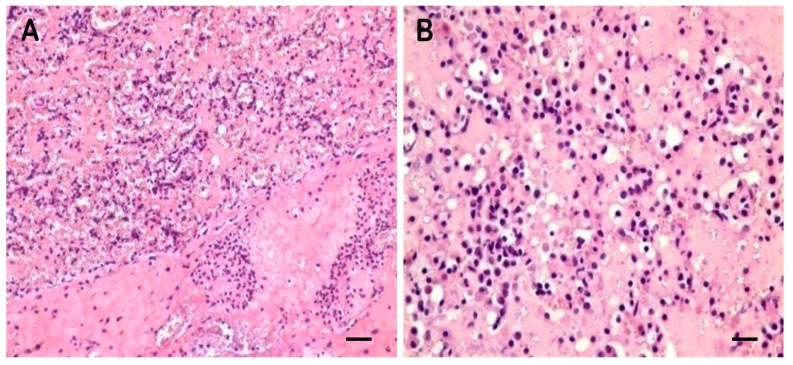
Figure 3.
Alveolar cavities were filled with pink serum and red blood cells (A) (scale bar = 50 μm, 200×). and filled with serum, lymphocyte infiltration in the alveolar wall (B) (scale bar = 25 μm, 400×) in the lung of TG.
2.2. Microarray Profiling
Expression profiling was conducted using a commercially available Agilent Porcine Genechip that included 43,603 probe sets. The transcriptome of the lung was determined. Expression was detected for 30,574 probes (70.12% of all probe sets) of the CG. A total of 31,957 probes (73.29% of all probe sets) were expressed in TG. When probe set intensities were normalized and filtered, there were still 26353 probes used to significantly identify DE genes. There were 11,929 genes identified as DE at the p ≤ 0.01 level by comparing the log2 (normalized signal) of the two groups using T-test analysis.
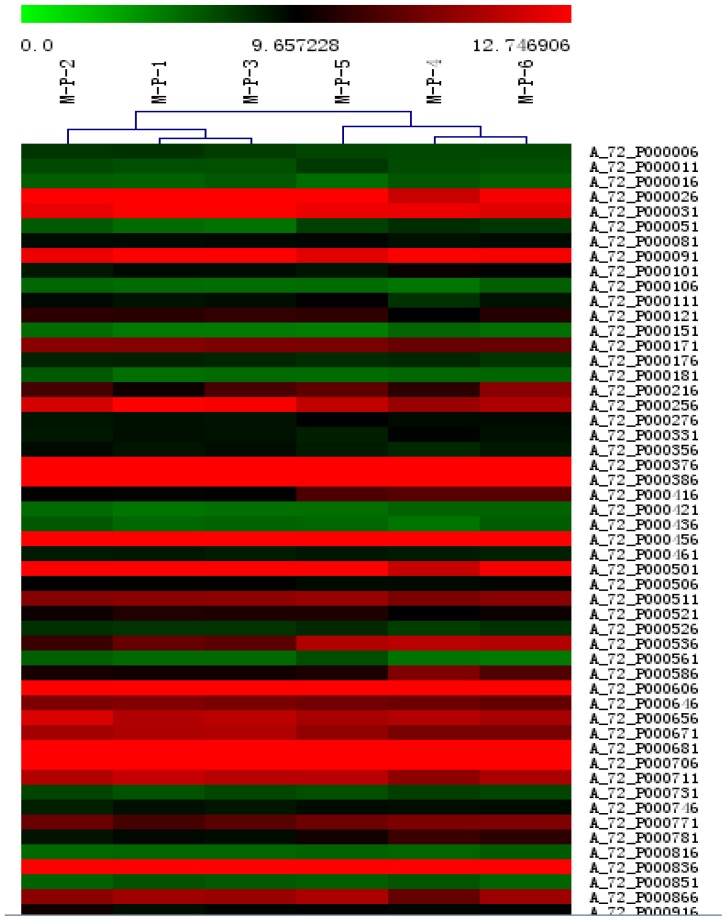
Hierarchical clustering was applied to the mean log-ratio of the replicated spots from the DE genes by the average linkage and using euclidean distance as the similarity metric (Figure 4). The expression profiles of samples were divided into two groups—one from the non-inoculated swine (M-P-1, M-P-2, M-P-3) and the other group from the inoculated swine (M-P-4, M-P-5, M-P-6).
Figure 4.
Hierarchical clustering analysis and clustering segmentation.
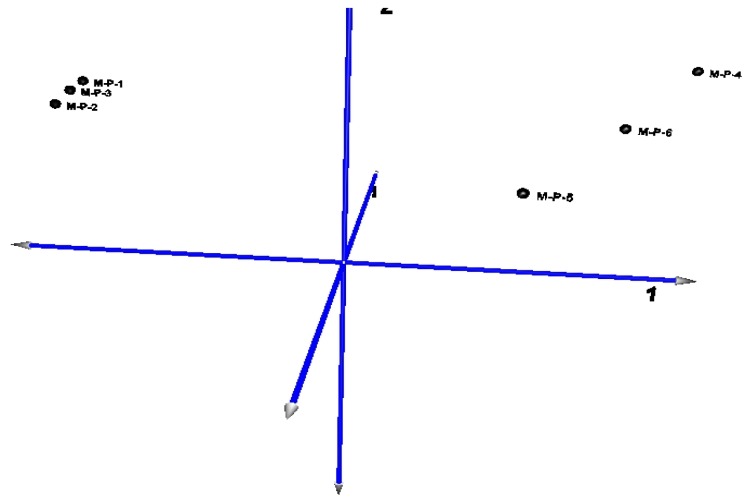
The principal component map to three-dimensional space, also found that the distance of CG (samples M-P-1, M-P-2, M-P-3) is short, and the gene expression pattern is more consistent. Also the distance of TG (samples M-P-4, M-P-5, M-P-6) is relatively discrete because of the differences in the degree of lesion, and the gene expression pattern is similar (Figure 5).
Figure 5.
Three-dimensional map of principal component analysis (PCA) for mapping samples obtained from clustering segmentation.
The six samples were set as variables, the principal component analysis (PCA) of the co-expressed differentially genes (CG vs TG) showed the contribution rate of the first principal component which reached 96.95%, the first three principal components of the total contribution rate reach 99.754% (Table 1).
Table 1.
Eigenvalues and contribution ratio of principal component analysis (PCA) for differential expression genes.
| Principal component | Eigenvalues | Contribution ratio |
|---|---|---|
| 1 | 50.17 | 96.95% |
| 2 | 1.339 | 2.59% |
| 3 | 0.111 | 0.21% |
| 4 | 0.088 | 0.17% |
| 5 | 0.027 | 0.05% |
| 6 | 0.013 | 0.02% |
2.3. DE Genes Profiling
Of the 11,929 DE genes, 1188 were annotated as swine genes in the GenBank Database (DB). GO and KEGG pathway analyses of the 1188 DE gene lists were conducted using DAVID. There were 89 biological process (BP) categories (Table 2), 82 cellular components (Table 3), and 182 molecular functions (Table 4) were significantly affected by infection with APP (p = 0). The BP of the test for over-representation of specific GO terms among the affected genes related to the immune responses were at least 27 (Table 5). Furthermore, a number of BP related to metabolism were also identified.
Table 2.
The significant gene ontology biological processes in pigs.
| Name | Description | Probe | Genes |
|---|---|---|---|
| [GO:0050896] | response to stimulus | 96 | 96 |
| [GO:0051179] | Localization | 92 | 92 |
| [GO:0006810] | Transport | 88 | 88 |
| [GO:0006807] | nitrogen compound metabolic process | 84 | 84 |
| [GO:0019222] | regulation of metabolic process | 67 | 67 |
| [GO:0002376] | immune system process | 56 | 56 |
| [GO:0055114] | oxidation reduction | 53 | 53 |
| [GO:0006955] | immune response | 53 | 53 |
| [GO:0032501] | multicellular organismal process | 52 | 52 |
| [GO:0006950] | response to stress | 51 | 51 |
| [GO:0009056] | catabolic process | 50 | 50 |
| [GO:0032502] | developmental process | 38 | 38 |
| [GO:0065008] | regulation of biological quality | 35 | 35 |
| [GO:0007275] | multicellular organismal development | 35 | 35 |
| [GO:0022610] | biological adhesion | 29 | 29 |
| [GO:0048518] | positive regulation of biological process | 28 | 28 |
| [GO:0016043] | cellular component organization | 27 | 27 |
| [GO:0009605] | response to external stimulus | 25 | 25 |
| [GO:0048856] | anatomical structure development | 22 | 22 |
| [GO:0008219] | cell death | 22 | 22 |
| [GO:0048519] | negative regulation of biological process | 21 | 21 |
| [GO:0033036] | macromolecule localization | 21 | 21 |
| [GO:0048523] | negative regulation of cellular process | 19 | 19 |
| [GO:0048522] | positive regulation of cellular process | 19 | 19 |
| [GO:0044281] | small molecule metabolic process | 18 | 18 |
| [GO:0048869] | cellular developmental process | 17 | 17 |
| [GO:0042221] | response to chemical stimulus | 17 | 17 |
| [GO:0006066] | alcohol metabolic process | 17 | 17 |
| [GO:0006996] | organelle organization | 16 | 16 |
| [GO:0051641] | cellular localization | 15 | 15 |
| [GO:0019882] | antigen processing and presentation | 15 | 15 |
| [GO:0008104] | protein localization | 15 | 15 |
| [GO:0048583] | regulation of response to stimulus | 13 | 13 |
| [GO:0042592] | homeostatic process | 12 | 12 |
| [GO:0032879] | regulation of localization | 12 | 12 |
| [GO:0007049] | cell cycle | 12 | 12 |
| [GO:0048584] | positive regulation of response to stimulus | 11 | 11 |
| [GO:0009893] | positive regulation of metabolic process | 11 | 11 |
| [GO:0002682] | regulation of immune system process | 11 | 11 |
| [GO:0051716] | cellular response to stimulus | 10 | 10 |
| [GO:0019725] | cellular homeostasis | 10 | 10 |
| [GO:0016192] | vesicle-mediated transport | 10 | 10 |
| [GO:0009653] | anatomical structure morphogenesis | 10 | 10 |
| [GO:0002252] | immune effector process | 10 | 10 |
| [GO:0051239] | regulation of multicellular organismal process | 9 | 9 |
| [GO:0048731] | system development | 9 | 9 |
| [GO:0065009] | regulation of molecular function | 8 | 8 |
| [GO:0051704] | multi-organism process | 8 | 8 |
| [GO:0051128] | regulation of cellular component organization | 8 | 8 |
| [GO:0050778] | positive regulation of immune response | 8 | 8 |
| [GO:0044085] | cellular component biogenesis | 8 | 8 |
| [GO:0007610] | behavior | 8 | 8 |
| [GO:0003008] | system process | 8 | 8 |
| [GO:0051301] | cell division | 7 | 7 |
| [GO:0030029] | actin filament-based process | 7 | 7 |
| [GO:0022607] | cellular component assembly | 7 | 7 |
| [GO:0009607] | response to biotic stimulus | 7 | 7 |
| [GO:0000003] | reproduction | 7 | 7 |
| [GO:0055085] | transmembrane transport | 6 | 6 |
| [GO:0051707] | response to other organism | 6 | 6 |
| [GO:0051129] | negative regulation of cellular component organization | 6 | 6 |
| [GO:0050878] | regulation of body fluid levels | 6 | 6 |
| [GO:0050793] | regulation of developmental process | 6 | 6 |
| [GO:0023052] | signaling | 6 | 6 |
| [GO:0022414] | reproductive process | 6 | 6 |
| [GO:0016044] | cellular membrane organization | 6 | 6 |
| [GO:0048646] | anatomical structure formation involved in morphogenesis | 5 | 5 |
| [GO:0040011] | locomotion | 5 | 5 |
| [GO:0019953] | sexual reproduction | 5 | 5 |
| [GO:0019637] | organophosphate metabolic process | 5 | 5 |
| [GO:0010817] | regulation of hormone levels | 5 | 5 |
| [GO:0009892] | negative regulation of metabolic process | 5 | 5 |
| [GO:0060348] | bone development | 4 | 4 |
| [GO:0044087] | regulation of cellular component biogenesis | 4 | 4 |
| [GO:0043933] | macromolecular complex subunit organization | 4 | 4 |
| [GO:0042330] | taxis | 4 | 4 |
| [GO:0040008] | regulation of growth | 4 | 4 |
| [GO:0022402] | cell cycle process | 4 | 4 |
| [GO:0070271] | protein complex biogenesis | 3 | 3 |
| [GO:0048609] | reproductive process in a multicellular organism | 3 | 3 |
| [GO:0046903] | secretion | 3 | 3 |
| [GO:0040012] | regulation of locomotion | 3 | 3 |
| [GO:0034621] | cellular macromolecular complex subunit organization | 3 | 3 |
| [GO:0019748] | secondary metabolic process | 3 | 3 |
| [GO:0010605] | negative regulation of macromolecule metabolic process | 3 | 3 |
| [GO:0009719] | response to endogenous stimulus | 3 | 3 |
| [GO:0009628] | response to abiotic stimulus | 3 | 3 |
| [GO:0007017] | microtubule-based process | 3 | 3 |
| [GO:0002520] | immune system development | 3 | 3 |
Table 3.
The significant gene ontology cellular components in pigs.
| Name | Description | Probe | Genes |
|---|---|---|---|
| [GO:0005886] | plasma membrane | 98 | 98 |
| [GO:0005634] | nucleus | 86 | 86 |
| [GO:0032991] | macromolecular complex | 86 | 86 |
| [GO:0044422] | organelle part | 84 | 84 |
| [GO:0043234] | protein complex | 62 | 62 |
| [GO:0043228] | non-membrane-bounded organelle | 46 | 46 |
| [GO:0044421] | extracellular region part | 45 | 45 |
| [GO:0044459] | plasma membrane part | 44 | 44 |
| [GO:0031090] | organelle membrane | 41 | 41 |
| [GO:0005739] | mitochondrion | 37 | 37 |
| [GO:0005783] | endoplasmic reticulum | 35 | 35 |
| [GO:0005794] | Golgi apparatus | 33 | 33 |
| [GO:0005615] | extracellular space | 29 | 29 |
| [GO:0012505] | endomembrane system | 25 | 25 |
| [GO:0044429] | mitochondrial part | 23 | 23 |
| [GO:0016023] | cytoplasmic membrane-bounded vesicle | 23 | 23 |
| [GO:0005856] | cytoskeleton | 23 | 23 |
| [GO:0031974] | membrane-enclosed lumen | 22 | 22 |
| [GO:0031975] | envelope | 21 | 21 |
| [GO:0005840] | ribosome | 19 | 19 |
| [GO:0044428] | nuclear part | 18 | 18 |
| [GO:0005578] | proteinaceous extracellular matrix | 17 | 17 |
| [GO:0044430] | cytoskeletal part | 15 | 15 |
| [GO:0071212] | subsynaptic reticulum | 15 | 15 |
| [GO:0005740] | mitochondrial envelope | 15 | 15 |
| [GO:0005829] | cytosol | 14 | 14 |
| [GO:0031966] | mitochondrial membrane | 14 | 14 |
| [GO:0019898] | extrinsic to membrane | 14 | 14 |
| [GO:0042611] | MHC protein complex | 13 | 13 |
| [GO:0048770] | pigment granule | 12 | 12 |
| [GO:0044431] | Golgi apparatus part | 12 | 12 |
| [GO:0005773] | vacuole | 11 | 11 |
| [GO:0005764] | lysosome | 10 | 10 |
| [GO:0044432] | endoplasmic reticulum part | 10 | 10 |
| [GO:0005792] | microsome | 9 | 9 |
| [GO:0009898] | internal side of plasma membrane | 9 | 9 |
| [GO:0005743] | mitochondrial inner membrane | 9 | 9 |
| [GO:0005887] | integral to plasma membrane | 8 | 8 |
| [GO:0005789] | endoplasmic reticulum membrane | 8 | 8 |
| [GO:0005768] | endosome | 8 | 8 |
| [GO:0005759] | mitochondrial matrix | 8 | 8 |
| [GO:0005654] | nucleoplasm | 8 | 8 |
| [GO:0015630] | microtubule cytoskeleton | 8 | 8 |
| [GO:0030054] | cell junction | 7 | 7 |
| [GO:0031300] | intrinsic to organelle membrane | 7 | 7 |
| [GO:0042613] | MHC class II protein complex | 7 | 7 |
| [GO:0044451] | nucleoplasm part | 7 | 7 |
| [GO:0031301] | integral to organelle membrane | 6 | 6 |
| [GO:0005635] | nuclear envelope | 6 | 6 |
| [GO:0042612] | MHC class I protein complex | 6 | 6 |
| [GO:0015629] | actin cytoskeleton | 6 | 6 |
| [GO:0016469] | proton-transporting two-sector ATPase complex | 6 | 6 |
| [GO:0031225] | anchored to membrane | 5 | 5 |
| [GO:0031968] | organelle outer membrane | 5 | 5 |
| [GO:0031965] | nuclear membrane | 5 | 5 |
| [GO:0043235] | receptor complex | 5 | 5 |
| [GO:0033279] | ribosomal subunit | 5 | 5 |
| [GO:0005819] | spindle | 4 | 4 |
| [GO:0030173] | integral to Golgi membrane | 4 | 4 |
| [GO:0048471] | perinuclear region of cytoplasm | 4 | 4 |
| [GO:0005911] | cell-cell junction | 4 | 4 |
| [GO:0043292] | contractile fiber | 4 | 4 |
| [GO:0000502] | proteasome complex | 4 | 4 |
| [GO:0030135] | coated vesicle | 4 | 4 |
| [GO:0016459] | myosin complex | 4 | 4 |
| [GO:0005874] | microtubule | 4 | 4 |
| [GO:0042995] | cell projection | 3 | 3 |
| [GO:0045259] | proton-transporting ATP synthase complex | 3 | 3 |
| [GO:0044420] | extracellular matrix part | 3 | 3 |
| [GO:0015935] | small ribosomal subunit | 3 | 3 |
| [GO:0005777] | peroxisome | 3 | 3 |
| [GO:0034702] | ion channel complex | 3 | 3 |
| [GO:0005901] | caveola | 3 | 3 |
| [GO:0045202] | synapse | 3 | 3 |
| [GO:0031227] | intrinsic to endoplasmic reticulum membrane | 3 | 3 |
| [GO:0016323] | basolateral plasma membrane | 3 | 3 |
| [GO:0005681] | spliceosomal complex | 3 | 3 |
| [GO:0030141] | secretory granule | 3 | 3 |
| [GO:0005667] | transcription factor complex | 3 | 3 |
| [GO:0033176] | proton-transporting V-type ATPase complex | 3 | 3 |
| [GO:0033177] | proton-transporting two-sector ATPase complex, proton-transporting domain | 3 | 3 |
| [GO:0005730] | nucleolus | 3 | 3 |
Table 4.
The significant gene ontology molecular functions in pigs.
| Name | Description | Probe | Genes |
|---|---|---|---|
| [GO:0017076] | purine nucleotide binding | 91 | 91 |
| [GO:0003676] | nucleic acid binding | 88 | 88 |
| [GO:0032555] | purine ribonucleotide binding | 81 | 81 |
| [GO:0004872] | receptor activity | 75 | 75 |
| [GO:0004690] | cyclic nucleotide-dependent protein kinase activity | 68 | 68 |
| [GO:0004691] | cAMP-dependent protein kinase activity | 67 | 67 |
| [GO:0016491] | oxidoreductase activity | 67 | 67 |
| [GO:0008270] | zinc ion binding | 64 | 64 |
| [GO:0030554] | adenyl nucleotide binding | 63 | 63 |
| [GO:0005102] | receptor binding | 57 | 57 |
| [GO:0032559] | adenyl ribonucleotide binding | 53 | 53 |
| [GO:0005215] | transporter activity | 52 | 52 |
| [GO:0008233] | peptidase activity | 46 | 46 |
| [GO:0070011] | peptidase activity, acting on l-amino acid peptides | 43 | 43 |
| [GO:0003677] | DNA binding | 43 | 43 |
| [GO:0004888] | transmembrane receptor activity | 40 | 40 |
| [GO:0005509] | calcium ion binding | 39 | 39 |
| [GO:0022892] | substrate-specific transporter activity | 39 | 39 |
| [GO:0004687] | myosin light chain kinase activity | 38 | 38 |
| [GO:0030528] | transcription regulator activity | 37 | 37 |
| [GO:0022857] | transmembrane transporter activity | 36 | 36 |
| [GO:0022891] | substrate-specific transmembrane transporter activity | 35 | 35 |
| [GO:0030234] | enzyme regulator activity | 35 | 35 |
| [GO:0005506] | iron ion binding | 33 | 33 |
| [GO:0004175] | endopeptidase activity | 33 | 33 |
| [GO:0003700] | transcription factor activity | 32 | 32 |
| [GO:0015075] | ion transmembrane transporter activity | 30 | 30 |
| [GO:0019001] | guanyl nucleotide binding | 28 | 28 |
| [GO:0005198] | structural molecule activity | 28 | 28 |
| [GO:0004857] | enzyme inhibitor activity | 26 | 26 |
| [GO:0016788] | hydrolase activity, acting on ester bonds | 26 | 26 |
| [GO:0008324] | cation transmembrane transporter activity | 25 | 25 |
| [GO:0009055] | electron carrier activity | 24 | 24 |
| [GO:0004930] | G-protein coupled receptor activity | 24 | 24 |
| [GO:0003723] | RNA binding | 23 | 23 |
| [GO:0005126] | cytokine receptor binding | 22 | 22 |
| [GO:0016874] | ligase activity | 22 | 22 |
| [GO:0048037] | cofactor binding | 20 | 20 |
| [GO:0016817] | hydrolase activity, acting on acid anhydrides | 19 | 19 |
| [GO:0003735] | structural constituent of ribosome | 19 | 19 |
| [GO:0005125] | cytokine activity | 18 | 18 |
| [GO:0030414] | peptidase inhibitor activity | 18 | 18 |
| [GO:0050662] | coenzyme binding | 17 | 17 |
| [GO:0016757] | transferase activity, transferring glycosyl groups | 17 | 17 |
| [GO:0030246] | carbohydrate binding | 17 | 17 |
| [GO:0008092] | cytoskeletal protein binding | 16 | 16 |
| [GO:0022890] | inorganic cation transmembrane transporter activity | 16 | 16 |
| [GO:0016746] | transferase activity, transferring acyl groups | 16 | 16 |
| [GO:0016879] | ligase activity, forming carbon-nitrogen bonds | 16 | 16 |
| [GO:0000287] | magnesium ion binding | 15 | 15 |
| [GO:0008237] | metallopeptidase activity | 15 | 15 |
| [GO:0016614] | oxidoreductase activity, acting on CH–OH group of donors | 15 | 15 |
| [GO:0022804] | active transmembrane transporter activity | 15 | 15 |
| [GO:0019955] | cytokine binding | 14 | 14 |
| [GO:0017171] | serine hydrolase activity | 14 | 14 |
| [GO:0046906] | tetrapyrrole binding | 14 | 14 |
| [GO:0016616] | oxidoreductase activity, acting on the CH–OH group of donors, NAD or NADP as acceptor | 14 | 14 |
| [GO:0016747] | transferase activity, transferring acyl groups other than amino-acyl groups | 14 | 14 |
| [GO:0008528] | peptide receptor activity, G-protein coupled | 14 | 14 |
| [GO:0003779] | actin binding | 14 | 14 |
| [GO:0004252] | serine-type endopeptidase activity | 13 | 13 |
| [GO:0016705] | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen | 13 | 13 |
| [GO:0004497] | monooxygenase activity | 12 | 12 |
| [GO:0042578] | phosphoric ester hydrolase activity | 12 | 12 |
| [GO:0008234] | cysteine-type peptidase activity | 11 | 11 |
| [GO:0046873] | metal ion transmembrane transporter activity | 11 | 11 |
| [GO:0008289] | lipid binding | 11 | 11 |
| [GO:0016791] | phosphatase activity | 11 | 11 |
| [GO:0050660] | FAD binding | 10 | 10 |
| [GO:0015078] | hydrogen ion transmembrane transporter activity | 10 | 10 |
| [GO:0016758] | transferase activity, transferring hexosyl groups | 10 | 10 |
| [GO:0004867] | serine-type endopeptidase inhibitor activity | 10 | 10 |
| [GO:0004428] | inositol or phosphatidylinositol kinase activity | 10 | 10 |
| [GO:0016798] | hydrolase activity, acting on glycosyl bonds | 10 | 10 |
| [GO:0005516] | calmodulin binding | 9 | 9 |
| [GO:0016810] | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds | 9 | 9 |
| [GO:0042623] | ATPase activity, coupled | 9 | 9 |
| [GO:0001871] | pattern binding | 9 | 9 |
| [GO:0016776] | phosphotransferase activity, phosphate group as acceptor | 9 | 9 |
| [GO:0005179] | hormone activity | 9 | 9 |
| [GO:0070851] | growth factor receptor binding | 9 | 9 |
| [GO:0004197] | cysteine-type endopeptidase activity | 9 | 9 |
| [GO:0005057] | receptor signaling protein activity | 9 | 9 |
| [GO:0004950] | chemokine receptor activity | 9 | 9 |
| [GO:0004222] | metalloendopeptidase activity | 9 | 9 |
| [GO:0043565] | sequence-specific DNA binding | 9 | 9 |
| [GO:0005216] | ion channel activity | 8 | 8 |
| [GO:0016853] | isomerase activity | 8 | 8 |
| [GO:0000826] | inositol pyrophosphate synthase activity | 8 | 8 |
| [GO:0019842] | vitamin binding | 8 | 8 |
| [GO:0005539] | glycosaminoglycan binding | 8 | 8 |
| [GO:0005529] | sugar binding | 8 | 8 |
| [GO:0005066] | transmembrane receptor protein tyrosine kinase signaling protein activity | 8 | 8 |
| [GO:0020037] | heme binding | 8 | 8 |
| [GO:0004356] | glutamate-ammonia ligase activity | 8 | 8 |
| [GO:0005507] | copper ion binding | 7 | 7 |
| [GO:0016209] | antioxidant activity | 7 | 7 |
| [GO:0008238] | exopeptidase activity | 7 | 7 |
| [GO:0008009] | chemokine activity | 7 | 7 |
| [GO:0016860] | intramolecular oxidoreductase activity | 7 | 7 |
| [GO:0004721] | phosphoprotein phosphatase activity | 7 | 7 |
| [GO:0015291] | secondary active transmembrane transporter activity | 7 | 7 |
| [GO:0016563] | transcription activator activity | 6 | 6 |
| [GO:0005244] | voltage-gated ion channel activity | 6 | 6 |
| [GO:0008201] | heparin binding | 6 | 6 |
| [GO:0031420] | alkali metal ion binding | 6 | 6 |
| [GO:0046983] | protein dimerization activity | 6 | 6 |
| [GO:0015082] | di-, tri-valent inorganic cation transmembrane transporter activity | 6 | 6 |
| [GO:0004312] | fatty-acid synthase activity | 6 | 6 |
| [GO:0042802] | identical protein binding | 6 | 6 |
| [GO:0016684] | oxidoreductase activity, acting on peroxide as acceptor | 6 | 6 |
| [GO:0016829] | lyase activity | 5 | 5 |
| [GO:0008047] | enzyme activator activity | 5 | 5 |
| [GO:0003924] | GTPase activity | 5 | 5 |
| [GO:0004091] | carboxylesterase activity | 5 | 5 |
| [GO:0015399] | primary active transmembrane transporter activity | 5 | 5 |
| [GO:0005261] | cation channel activity | 5 | 5 |
| [GO:0019904] | protein domain specific binding | 5 | 5 |
| [GO:0004694] | eukaryotic translation initiation factor 2alpha kinase activity | 5 | 5 |
| [GO:0016627] | oxidoreductase activity, acting on the CH–CH group of donors | 5 | 5 |
| [GO:0031406] | carboxylic acid binding | 5 | 5 |
| [GO:0042803] | protein homodimerization activity | 5 | 5 |
| [GO:0004518] | nuclease activity | 5 | 5 |
| [GO:0019899] | enzyme binding | 5 | 5 |
| [GO:0003774] | motor activity | 5 | 5 |
| [GO:0008430] | selenium binding | 5 | 5 |
| [GO:0004725] | protein tyrosine phosphatase activity | 5 | 5 |
| [GO:0015293] | symporter activity | 5 | 5 |
| [GO:0004713] | protein tyrosine kinase activity | 5 | 5 |
| [GO:0046915] | transition metal ion transmembrane transporter activity | 4 | 4 |
| [GO:0008135] | translation factor activity, nucleic acid binding | 4 | 4 |
| [GO:0008134] | transcription factor binding | 4 | 4 |
| [GO:0010857] | calcium-dependent protein kinase activity | 4 | 4 |
| [GO:0050661] | NADP or NADPH binding | 4 | 4 |
| [GO:0060589] | nucleoside-triphosphatase regulator activity | 4 | 4 |
| [GO:0008373] | sialyltransferase activity | 4 | 4 |
| [GO:0004896] | cytokine receptor activity | 4 | 4 |
| [GO:0008083] | growth factor activity | 4 | 4 |
| [GO:0016709] | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, NADH or NADPH as one donor, and incorporation of one atom of oxygen | 4 | 4 |
| [GO:0004576] | oligosaccharyl transferase activity | 4 | 4 |
| [GO:0008509] | anion transmembrane transporter activity | 4 | 4 |
| [GO:0051540] | metal cluster binding | 4 | 4 |
| [GO:0004177] | aminopeptidase activity | 4 | 4 |
| [GO:0016765] | transferase activity, transferring alkyl or aryl (other than methyl) groups | 4 | 4 |
| [GO:0015929] | hexosaminidase activity | 4 | 4 |
| [GO:0016741] | transferase activity, transferring one-carbon groups | 4 | 4 |
| [GO:0004129] | cytochrome-c oxidase activity | 4 | 4 |
| [GO:0004521] | endoribonuclease activity | 4 | 4 |
| [GO:0051287] | NAD or NADH binding | 3 | 3 |
| [GO:0005385] | zinc ion transmembrane transporter activity | 3 | 3 |
| [GO:0004774] | succinate-CoA ligase activity | 3 | 3 |
| [GO:0004090] | carbonyl reductase (NADPH) activity | 3 | 3 |
| [GO:0005249] | voltage-gated potassium channel activity | 3 | 3 |
| [GO:0008757] | S-adenosylmethionine-dependent methyltransferase activity | 3 | 3 |
| [GO:0005160] | transforming growth factor beta receptor binding | 3 | 3 |
| [GO:0008235] | metalloexopeptidase activity | 3 | 3 |
| [GO:0005275] | amine transmembrane transporter activity | 3 | 3 |
| [GO:0019840] | isoprenoid binding | 3 | 3 |
| [GO:0019838] | growth factor binding | 3 | 3 |
| [GO:0005128] | erythropoietin receptor binding | 3 | 3 |
| [GO:0016801] | hydrolase activity, acting on ether bonds | 3 | 3 |
| [GO:0016701] | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen | 3 | 3 |
| [GO:0003712] | transcription cofactor activity | 3 | 3 |
| [GO:0015144] | carbohydrate transmembrane transporter activity | 3 | 3 |
| [GO:0003995] | acyl-CoA dehydrogenase activity | 3 | 3 |
| [GO:0004869] | cysteine-type endopeptidase inhibitor activity | 3 | 3 |
| [GO:0031402] | sodium ion binding | 3 | 3 |
| [GO:0019888] | protein phosphatase regulator activity | 3 | 3 |
| [GO:0016651] | oxidoreductase activity, acting on NADH or NADPH | 3 | 3 |
| [GO:0004180] | carboxypeptidase activity | 3 | 3 |
| [GO:0030695] | GTPase regulator activity | 3 | 3 |
| [GO:0010181] | FMN binding | 3 | 3 |
| [GO:0003743] | translation initiation factor activity | 3 | 3 |
| [GO:0016861] | intramolecular oxidoreductase activity, interconverting aldoses and ketoses | 3 | 3 |
| [GO:0004522] | pancreatic ribonuclease activity | 3 | 3 |
| [GO:0015294] | solute:cation symporter activity | 3 | 3 |
| [GO:0016790] | thiolester hydrolase activity | 3 | 3 |
| [GO:0008026] | ATP-dependent helicase activity | 3 | 3 |
| [GO:0030145] | manganese ion binding | 3 | 3 |
| [GO:0008417] | fucosyltransferase activity | 3 | 3 |
| [GO:0008194] | UDP-glycosyltransferase activity | 3 | 3 |
| [GO:0008199] | ferric iron binding | 3 | 3 |
Table 5.
Significant GO terms related to the immune responses caused by infection with APP.
| NO | GO term | Biological process | Number PermineJ | |
|---|---|---|---|---|
| 1 | GO:0050896 | response to stimulus | 96 | 0 |
| 2 | GO:0051179 | Localization | 92 | 0 |
| 3 | GO:0002376 | immune system process | 56 | 0 |
| 4 | GO:0006955 | immune response | 53 | 0 |
| 5 | GO:0055114 | oxidation reduction | 53 | 0 |
| 1 | GO:0050896 | response to stimulus | 96 | 0 |
| 6 | GO:0006950 | response to stress | 51 | 0 |
| 7 | GO:0022610 | biological adhesion | 29 | 0 |
| 8 | GO:0009605 | response to external stimulus | 25 | 0 |
| 9 | GO:0008219 | cell death | 22 | 0 |
| 10 | GO:0033036 | macromolecule localization | 21 | 0 |
| 11 | GO:0042221 | response to chemical stimulus | 17 | 0 |
| 12 | GO:0019882 | antigen processing and presentation | 15 | 0 |
| 13 | GO:0048583 | regulation of response to stimulus | 13 | 0 |
| 14 | GO:0032879 | regulation of localization | 12 | 0 |
| 15 | GO:0042592 | homeostatic process | 12 | 0 |
| 16 | GO:0048584 | positive regulation of response to stimulus | 11 | 0 |
| 17 | GO:0002682 | regulation of immune system process | 11 | 0 |
| 18 | GO:0051716 | cellular response to stimulus | 10 | 0 |
| 19 | GO:0002252 | immune effector process | 10 | 0 |
| 20 | GO:0050778 | positive regulation of immune response | 8 | 0 |
| 21 | GO:0009607 | response to biotic stimulus | 7 | 0 |
| 22 | GO:0023052 | signaling | 6 | 0 |
| 23 | GO:0040011 | locomotion | 5 | 0 |
| 24 | GO:0042330 | taxis | 4 | 0 |
| 25 | GO:0002520 | immune system development | 3 | 0 |
| 26 | GO:0009628 | response to abiotic stimulus | 3 | 0 |
| 27 | GO:0009719 | response to endogenous stimulus | 3 | 0 |
A total of 513 genes were analyzed using gene set enrichment analysis (GSEA). Three hundred and thirty (64.3%) database genes correlated with TG, while the other 183 (35.7%) genes correlated with CG. One hundred and thirty pathways remained for further analysis after size filtering (2 ≤ sizes ≤ 20). Altogether, 102 pathways (Table 6) were enriched and up-regulated in the TG and down-regulated in the CG. One pathway (SSC04664) was significantly enriched at a false discovery rate <25%. Eight pathways (i.e., SSC04664, SSC04930, SSC04914, SSC00140, SSC04621, SSC05221, SSC05218 and SSC03040) were significantly enriched at nominal p values of less than 1% and 5%.
Table 6.
Gene sets enriched in phenotype treatment group (TG).
| NO. | Pathway | Size | ES | NES | NOM p-val | FDR q-val | FWER p-val | Rank at max | Leading edge |
|---|---|---|---|---|---|---|---|---|---|
| 1 | ssc04664: Fc epsilon RI signaling pathway | 13 | −0.74 | −1.83 | 0.001 | 0.084 | 0.064 | 89 | Tags = 77%, list = 17%, signal = 91% |
| 2 | ssc04930: Type II diabetes mellitus | 5 | −0.82 | −1.62 | 0.017 | 0.715 | 0.685 | 42 | Tags = 60%, list = 8%, signal = 65% |
| 3 | ssc04914: Progesterone-mediated oocyte maturation | 12 | −0.66 | −1.61 | 0.012 | 0.508 | 0.701 | 56 | Tags = 42%, list = 11%, signal = 46% |
| 4 | ssc00140: Steroid hormone biosynthesis | 4 | −0.85 | −1.58 | 0.012 | 0.529 | 0.814 | 44 | Tags = 75%, list = 9%, signal = 81% |
| 5 | ssc04621: NOD-like receptor signaling pathway | 13 | −0.62 | −1.56 | 0.019 | 0.49 | 0.861 | 49 | tags = 46%, list = 10%, signal = 50% |
| 6 | ssc05221: Acute myeloid leukemia | 11 | −0.63 | −1.54 | 0.034 | 0.478 | 0.901 | 133 | tags = 73%, list = 26%, signal = 96% |
| 7 | ssc05218: Melanoma | 10 | −0.64 | −1.51 | 0.049 | 0.562 | 0.955 | 100 | tags = 50%, list = 19%, signal = 61% |
| 8 | ssc03040: Spliceosome | 9 | −0.67 | −1.5 | 0.039 | 0.524 | 0.961 | 101 | tags = 44%, list = 20%, signal = 54% |
| 9 | ssc04640: Hematopoietic cell lineage | 19 | −0.53 | −1.48 | 0.055 | 0.55 | 0.982 | 120 | tags = 58%, list = 23%, signal = 73% |
| 10 | ssc04210: Apoptosis | 10 | −0.61 | −1.48 | 0.066 | 0.504 | 0.984 | 158 | tags = 70%, list = 31%, signal = 99% |
| 11 | Ssc05214: Glioma | 11 | −0.61 | −1.47 | 0.052 | 0.467 | 0.985 | 100 | tags = 45%, list = 19%, signal = 55% |
| 12 | ssc04012: ErbB signaling pathway | 13 | −0.56 | −1.45 | 0.076 | 0.539 | 0.999 | 102 | tags = 38%, list = 20%, signal = 47% |
| 13 | ssc05020: Prion diseases | 8 | −0.64 | −1.44 | 0.07 | 0.533 | 1 | 65 | tags = 50%, list = 13%, signal = 56% |
| 14 | ssc04666: Fc gamma R-mediated phagocytosis | 12 | −0.56 | −1.4 | 0.098 | 0.641 | 1 | 135 | tags = 67%, list = 26%, signal = 88% |
| 15 | ssc04650: Natural killer cell mediated cytotoxicity | 19 | −0.52 | −1.4 | 0.087 | 0.605 | 1 | 89 | tags = 42%, list = 17%, signal = 49% |
| 16 | ssc00650: Butanoate metabolism | 5 | −0.72 | −1.39 | 0.088 | 0.581 | 1 | 6 | tags = 20%, list = 1%, signal = 20% |
| 17 | ssc00410: Beta-Alanine metabolism | 5 | −0.71 | −1.38 | 0.076 | 0.603 | 1 | 6 | tags = 20%, list = 1%, signal = 20% |
| 18 | ssc04660: T cell receptor signaling pathway | 18 | −0.49 | −1.35 | 0.112 | 0.689 | 1 | 122 | tags = 56%, list = 24%, signal = 70% |
| 19 | ssc04370: VEGF signaling pathway | 8 | −0.61 | −1.35 | 0.152 | 0.664 | 1 | 122 | tags = 63%, list = 24%, signal = 81% |
| 20 | ssc05219: Bladder cancer | 10 | −0.56 | −1.34 | 0.132 | 0.648 | 1 | 147 | tags = 60%, list = 29%, signal = 82% |
| 21 | ssc04920: Adipocytokine signaling pathway | 12 | −0.53 | −1.34 | 0.146 | 0.634 | 1 | 78 | tags = 42%, list = 15%, signal = 48% |
| 22 | ssc04114: Oocyte meiosis | 12 | −0.54 | −1.34 | 0.132 | 0.608 | 1 | 56 | tags = 25%, list = 11%, signal = 27% |
| 23 | ssc00750: Vitamin B6 metabolism | 2 | −0.86 | −1.33 | 0.09 | 0.6 | 1 | 72 | tags = 100%, list = 14%, signal = 116% |
| 24 | ssc00260: Glycine, serine and threonine metabolism | 4 | −0.71 | −1.32 | 0.131 | 0.597 | 1 | 128 | tags = 75%, list = 25%, signal = 99% |
| 25 | ssc04960: Aldosterone-regulated sodium reabsorption | 5 | −0.69 | −1.32 | 0.139 | 0.579 | 1 | 106 | tags = 60%, list = 21%, signal = 75% |
| 26 | ssc00910: Nitrogen metabolism | 3 | −0.76 | −1.29 | 0.164 | 0.646 | 1 | 91 | tags = 67%, list = 18%, signal = 81% |
| 27 | ssc05220: Chronic myeloid leukemia | 14 | −0.5 | −1.27 | 0.206 | 0.69 | 1 | 172 | tags = 57%, list = 34%, signal = 84% |
| 28 | ssc04115: P53 signaling pathway | 12 | −0.51 | −1.27 | 0.197 | 0.687 | 1 | 156 | tags = 75%, list = 30%, signal = 105% |
| 29 | ssc04630: Jak-STAT signaling pathway | 19 | −0.44 | −1.22 | 0.234 | 0.813 | 1 | 105 | tags = 42%, list = 20%, signal = 51% |
| 30 | ssc00640: Propanoate metabolism | 9 | −0.53 | −1.22 | 0.23 | 0.798 | 1 | 61 | tags = 22%, list = 12%, signal = 25% |
| 31 | ssc05213: Endometrial cancer | 10 | −0.51 | −1.21 | 0.248 | 0.787 | 1 | 133 | tags = 50%, list = 26%, signal = 66% |
| 32 | ssc00591: Linoleic acid metabolism | 4 | −0.66 | −1.19 | 0.27 | 0.829 | 1 | 44 | tags = 75%, list = 9%, signal = 81% |
| 33 | ssc05215: Prostate cancer | 17 | −0.44 | −1.19 | 0.243 | 0.811 | 1 | 115 | tags = 35%, list = 22%, signal = 44% |
| 34 | ssc00280: Valine, leucine and isoleucine degradation | 11 | −0.49 | −1.18 | 0.269 | 0.826 | 1 | 157 | tags = 45%, list = 31%, signal = 64% |
| 35 | ssc00620: Pyruvate metabolism | 7 | −0.54 | −1.17 | 0.285 | 0.818 | 1 | 239 | tags = 100%, list = 47%, signal = 185% |
| 36 | ssc00010: Glycolysis/Gluconeogenesis | 12 | −0.47 | −1.17 | 0.26 | 0.796 | 1 | 243 | tags = 83%, list = 47%, signal = 155% |
| 37 | ssc04150: MTOR signaling pathway | 7 | −0.56 | −1.17 | 0.291 | 0.78 | 1 | 42 | tags = 29%, list = 8%, signal = 31% |
| 38 | ssc00250: Alanine, aspartate and glutamate metabolism | 5 | −0.6 | −1.16 | 0.302 | 0.79 | 1 | 6 | tags = 20%, list = 1%, signal = 20% |
| 39 | ssc00511: Other glycan degradation | 4 | −0.63 | −1.16 | 0.312 | 0.772 | 1 | 195 | tags = 100%, list = 38%, signal = 160% |
| 40 | ssc05212: Pancreatic cancer | 13 | −0.45 | −1.16 | 0.304 | 0.754 | 1 | 172 | tags = 54%, list = 34%, signal = 79% |
| 41 | ssc00604: Glycosphingolipid biosynthesis | 4 | −0.63 | −1.15 | 0.299 | 0.755 | 1 | 195 | tags = 100%, list = 38%, signal = 160% |
| 42 | ssc00052: Galactose metabolism | 3 | −0.66 | −1.12 | 0.338 | 0.835 | 1 | 52 | tags = 33%, list = 10%, signal = 37% |
| 43 | ssc00520: Amino sugar and nucleotide sugar metabolism | 6 | −0.54 | −1.11 | 0.353 | 0.831 | 1 | 116 | tags = 50%, list = 23%, signal = 64% |
| 44 | ssc04070: Phosphatidylinositol signaling system | 6 | −0.55 | −1.11 | 0.358 | 0.813 | 1 | 100 | tags = 67%, list = 19%, signal = 82% |
| 45 | ssc04662: B cell receptor signaling pathway | 12 | −0.45 | −1.11 | 0.336 | 0.798 | 1 | 120 | tags = 58%, list = 23%, signal = 74% |
| 46 | ssc00500: Starch and sucrose metabolism | 4 | −0.6 | −1.1 | 0.37 | 0.809 | 1 | 57 | tags = 50%, list = 11%, signal = 56% |
| 47 | ssc05014: Amyotrophic lateral sclerosis (ALS) | 5 | −0.57 | −1.09 | 0.404 | 0.812 | 1 | 49 | tags = 40%, list = 10%, signal = 44% |
| 48 | ssc00310: Lysine degradation | 4 | −0.59 | −1.09 | 0.381 | 0.795 | 1 | 149 | tags = 75%, list = 29%, signal = 105% |
| 49 | ssc04144: Endocytosis | 18 | −0.41 | −1.08 | 0.384 | 0.814 | 1 | 26 | tags = 17%, list = 5%, signal = 17% |
| 50 | ssc00533: Keratan sulfate biosynthesis | 2 | −0.69 | −1.07 | 0.389 | 0.801 | 1 | 162 | tags = 100%, list = 32%, signal = 146% |
| 51 | ssc04623: Cytosolic DNA-sensing pathway | 8 | −0.47 | −1.07 | 0.4 | 0.791 | 1 | 30 | tags = 25%, list = 6%, signal = 26% |
| 52 | ssc00340: Histidine metabolism | 5 | −0.55 | −1.06 | 0.397 | 0.806 | 1 | 149 | tags = 60%, list = 29%, signal = 84% |
| 53 | ssc00980: Metabolism of xenobiotics by cytochrome P450 | 9 | −0.45 | −1.04 | 0.442 | 0.835 | 1 | 44 | tags = 33%, list = 9%, signal = 36% |
| 54 | ssc00270: Cysteine and methionine metabolism | 5 | −0.53 | −1.04 | 0.478 | 0.824 | 1 | 245 | tags = 100%, list = 48%, signal = 190% |
| 55 | ssc04330: Notch signaling pathway | 6 | −0.5 | −1.03 | 0.473 | 0.833 | 1 | 4 | tags = 17%, list = 1%, signal = 17% |
| 56 | ssc00330: Arginine and proline metabolism | 10 | −0.43 | −1.03 | 0.447 | 0.824 | 1 | 174 | tags = 50%, list = 34%, signal = 74% |
| 57 | ssc05223: Non-small cell lung cancer | 7 | −0.47 | −1.01 | 0.487 | 0.844 | 1 | 89 | tags = 43%, list = 17%, signal = 51% |
| 58 | ssc04720: Long-term potentiation | 8 | −0.46 | −1.01 | 0.48 | 0.84 | 1 | 89 | tags = 25%, list = 17%, signal = 30% |
| 59 | ssc04110: Cell cycle | 19 | −0.36 | −1 | 0.472 | 0.832 | 1 | 220 | tags = 58%, list = 43%, signal = 98% |
| 60 | ssc04730: Long-term depression | 13 | −0.39 | −1 | 0.464 | 0.822 | 1 | 34 | tags = 15%, list = 7%, signal = 16% |
| 61 | ssc05211: Renal cell carcinoma | 13 | −0.4 | −1 | 0.467 | 0.812 | 1 | 193 | tags = 62%, list = 38%, signal = 96% |
| 62 | ssc04912: GnRH signaling pathway | 13 | −0.38 | −0.98 | 0.507 | 0.842 | 1 | 122 | tags = 31%, list = 24%, signal = 39% |
| 63 | ssc00983: Drug metabolism | 5 | −0.5 | −0.97 | 0.515 | 0.838 | 1 | 44 | tags = 60%, list = 9%, signal = 65% |
| 64 | ssc00450: Selenoamino acid metabolism | 3 | −0.58 | −0.97 | 0.547 | 0.84 | 1 | 176 | tags = 67%, list = 34%, signal = 101% |
| 65 | ssc00071: Fatty acid metabolism | 8 | −0.43 | −0.95 | 0.556 | 0.856 | 1 | 244 | tags = 75%, list = 48%, signal = 141% |
| 66 | ssc00020: Citrate cycle (TCA cycle) | 9 | −0.41 | −0.95 | 0.533 | 0.845 | 1 | 309 | tags = 100%, list = 60%, signal = 247% |
| 67 | ssc03018: RNA degradation | 2 | −0.6 | −0.93 | 0.583 | 0.866 | 1 | 206 | tags = 100%, list = 40%, signal = 166% |
| 68 | ssc00603: Glycosphingolipid biosynthesis | 6 | −0.44 | −0.93 | 0.58 | 0.866 | 1 | 195 | tags = 83%, list = 38%, signal = 133% |
| 69 | ssc00053: Ascorbate and aldarate metabolism | 3 | −0.54 | −0.92 | 0.599 | 0.86 | 1 | 239 | tags = 100%, list = 47%, signal = 186% |
| 70 | ssc04020: Calcium signaling pathway | 20 | −0.33 | −0.92 | 0.578 | 0.85 | 1 | 29 | tags = 10%, list = 6%, signal = 10% |
| 71 | ssc00510: N-Glycan biosynthesis | 8 | −0.41 | −0.91 | 0.607 | 0.863 | 1 | 306 | tags = 100%, list = 60%, signal = 244% |
| 72 | ssc00903: Limonene and pinene degradation | 3 | −0.54 | −0.91 | 0.631 | 0.851 | 1 | 239 | tags = 100%, list = 47%, signal = 186% |
| 73 | ssc03320: PPAR signaling pathway | 13 | −0.36 | −0.91 | 0.598 | 0.843 | 1 | 78 | tags = 31%, list = 15%, signal = 35% |
| 74 | ssc04142: Lysosome | 16 | −0.34 | −0.89 | 0.623 | 0.867 | 1 | 195 | tags = 75%, list = 38%, signal = 117% |
| 75 | ssc05210: Colorectal cancer | 13 | −0.35 | −0.89 | 0.599 | 0.857 | 1 | 42 | tags = 15%, list = 8%, signal = 16% |
| 76 | ssc04622: RIG-I-like receptor signaling pathway | 12 | −0.35 | −0.87 | 0.626 | 0.867 | 1 | 49 | tags = 25%, list = 10%, signal = 27% |
| 77 | ssc04320: Dorso-ventral axis formation | 2 | −0.57 | −0.87 | 0.674 | 0.86 | 1 | 34 | tags = 50%, list = 7%, signal = 53% |
| 78 | ssc04130: SNARE interactions in vesicular transport | 2 | −0.57 | −0.87 | 0.676 | 0.857 | 1 | 123 | tags = 50%, list = 24%, signal = 66% |
| 79 | ssc00230: Purine metabolism | 6 | −0.42 | −0.85 | 0.68 | 0.869 | 1 | 21 | tags = 17%, list = 4%, signal = 17% |
| 80 | ssc00380: Tryptophan metabolism | 4 | −0.46 | −0.85 | 0.699 | 0.865 | 1 | 239 | tags = 75%, list = 47%, signal = 139% |
| 81 | ssc04910:Insulin signaling pathway | 16 | −0.32 | −0.84 | 0.688 | 0.859 | 1 | 104 | tags = 25%, list = 20%, signal = 30% |
| 82 | ssc00190: Oxidative phosphorylation | 17 | −0.31 | −0.84 | 0.666 | 0.85 | 1 | 317 | tags = 94%, list = 62%, signal = 238% |
| 83 | ssc00051: Fructose and mannose metabolism | 4 | −0.46 | −0.84 | 0.697 | 0.848 | 1 | 69 | tags = 25%, list = 13%, signal = 29% |
| 84 | ssc04614: Renin-angiotensin system | 3 | −0.5 | −0.84 | 0.715 | 0.839 | 1 | 143 | tags = 33%, list = 28%, signal = 46% |
| 85 | ssc00860: Porphyrin and chlorophyll metabolism | 3 | −0.49 | −0.83 | 0.709 | 0.836 | 1 | 238 | tags = 67%, list = 46%, signal = 124% |
| 86 | ssc00350: Tyrosine metabolism | 5 | −0.43 | −0.83 | 0.692 | 0.833 | 1 | 263 | tags = 80%, list = 51%, signal = 163% |
| 87 | ssc00561: Glycerolipid metabolism | 6 | −0.38 | −0.79 | 0.733 | 0.87 | 1 | 239 | tags = 83%, list = 47%, signal = 154% |
| 88 | ssc00030: Pentose phosphate pathway | 5 | −0.41 | −0.79 | 0.744 | 0.868 | 1 | 57 | tags = 20%, list = 11%, signal = 22% |
| 89 | ssc04540: Gap junction | 11 | −0.32 | −0.77 | 0.764 | 0.884 | 1 | 134 | tags = 27%, list = 26%, signal = 36% |
| 90 | ssc04916: Melanogenesis | 14 | −0.29 | −0.75 | 0.787 | 0.892 | 1 | 145 | tags = 29%, list = 28%, signal = 39% |
| 91 | ssc00562: Inositol phosphate metabolism | 7 | −0.35 | −0.75 | 0.775 | 0.884 | 1 | 100 | tags = 43%, list = 19%, signal = 53% |
| 92 | ssc03050 Proteasome: | 4 | −0.4 | −0.74 | 0.814 | 0.887 | 1 | 97 | tags = 25%, list = 19%, signal = 31% |
| 93 | ssc00600: Sphingolipid metabolism | 5 | −0.36 | −0.71 | 0.821 | 0.919 | 1 | 107 | tags = 40%, list = 21%, signal = 50% |
| 94 | ssc00630: Glyoxylate and dicarboxylate metabolism | 2 | −0.46 | −0.7 | 0.907 | 0.916 | 1 | 142 | tags = 50%, list = 28%, signal = 69% |
| 95 | ssc04512: ECM-receptor interaction | 11 | −0.29 | −0.69 | 0.853 | 0.916 | 1 | 8 | tags = 9%, list = 2%, signal = 9% |
| 96 | ssc00531: Glycosaminoglycan degradation | 4 | −0.35 | −0.65 | 0.899 | 0.946 | 1 | 240 | tags = 75%, list = 47%, signal = 140% |
| 97 | ssc05216: Thyroid cancer | 8 | −0.29 | −0.63 | 0.904 | 0.948 | 1 | 133 | tags = 38%, list = 26%, signal = 50% |
| 98 | ssc04520: Adherens junction | 10 | −0.26 | −0.61 | 0.902 | 0.957 | 1 | 170 | tags = 40%, list = 33%, signal = 59% |
| 99 | ssc00564: Glycerophospholipid metabolism | 5 | −0.3 | −0.58 | 0.94 | 0.969 | 1 | 219 | tags = 60%, list = 43%, signal = 104% |
| 100 | ssc00360: Phenylalanine metabolism | 5 | −0.26 | −0.51 | 0.972 | 0.992 | 1 | 219 | tags = 60%, list = 43%, signal = 104% |
| 101 | ssc04270: Vascular smooth muscle contraction | 14 | −0.18 | −0.46 | 0.988 | 0.998 | 1 | 34 | tags = 7%, list = 7%, signal = 7% |
| 102 | ssc04350: TGF-beta signaling pathway | 14 | −0.18 | −0.45 | 0.982 | 0.99 | 1 | 172 | tags = 36%, list = 34%, signal = 52% |
Twenty-eight pathways (Table 7) were down-regulated in the TG but upregulated in the CG. Six pathways (i.e., SSC05320, SSC04940, SSC05330, SSC04530, SSC04260 and SSC05412) were significant at a false discovery rate of <25%. Five pathways (i.e., SSC05320, SSC04940, SSC05330, SSC04530 and SSC04260) were significantly enriched at a nominal p value of less than 1% and 5%.
Table 7.
Gene sets enriched in phenotype control group (CG).
| No. | Pathway | Size | ES | NES | NOM p-val | FDR q-val | FWER p-val | Rank at Max | Leading edge |
|---|---|---|---|---|---|---|---|---|---|
| 1 | ssc05320: Autoimmune thyroid disease | 15 | 0.64 | 1.97 | 0.003 | 0.034 | 0.058 | 139 | tags = 87%, list = 27%, signal = 115% |
| 2 | ssc04940: Type I diabetes mellitus | 16 | 0.59 | 1.86 | 0.003 | 0.053 | 0.165 | 139 | tags = 88%, list = 27%, signal = 116% |
| 3 | ssc05330: Allograft rejection | 17 | 0.54 | 1.72 | 0.015 | 0.116 | 0.458 | 139 | tags = 82%, list = 27%, signal = 109% |
| 4 | Ssc04530: Tight junction | 17 | 0.52 | 1.67 | 0.013 | 0.141 | 0.628 | 2 | tags = 12%, list = 0%, signal = 11% |
| 5 | ssc04260: Cardiac muscle contraction | 13 | 0.55 | 1.65 | 0.03 | 0.132 | 0.683 | 107 | tags = 46%, list = 21%, signal = 57% |
| 6 | ssc05412: Arrhythmogenic right ventricular cardiomyopathy | 12 | 0.56 | 1.57 | 0.051 | 0.196 | 0.853 | 155 | tags = 75%, list = 30%, signal = 105% |
| 7 | ssc02010: ABC transporters | 2 | 0.83 | 1.34 | 0.136 | 0.6 | 0.998 | 89 | tags = 100%, list = 17%, signal = 121% |
| 8 | ssc05340: Primary immunodeficiency | 7 | 0.54 | 1.3 | 0.179 | 0.619 | 0.999 | 61 | tags = 43%, list = 12%, signal = 48% |
| 9 | ssc05217: Basal cell carcinoma | 5 | 0.56 | 1.25 | 0.241 | 0.668 | 1 | 114 | tags = 60%, list = 22%, signal = 76% |
| 10 | ssc04740: Olfactory transduction | 3 | 0.67 | 1.22 | 0.259 | 0.664 | 1 | 172 | Tags = 100%, list = 34%, signal = 150% |
| 11 | ssc04120: Ubiquitin mediated proteolysis | 8 | 0.47 | 1.18 | 0.271 | 0.683 | 1 | 32 | Tags = 25%, list = 6%, signal = 26% |
| 12 | ssc03010: Ribosome | 17 | 0.37 | 1.17 | 0.257 | 0.657 | 1 | 328 | Tags = 100%, list = 64%, signal = 268% |
| 13 | ssc01040: Biosynthesis of unsaturated fatty acids | 2 | 0.73 | 1.15 | 0.336 | 0.634 | 1 | 11 | Tags = 50%, list = 2%, signal = 51% |
| 14 | ssc05012: Parkinson’s disease | 17 | 0.35 | 1.13 | 0.302 | 0.633 | 1 | 337 | Tags = 100%, list = 66%, signal = 282% |
| 15 | ssc05332: Graft-versus-host disease | 15 | 0.36 | 1.1 | 0.367 | 0.646 | 1 | 139 | Tags = 80%, list = 27%, signal = 107% |
| 16 | ssc00590: Arachidonic acid metabolism | 11 | 0.39 | 1.08 | 0.352 | 0.634 | 1 | 55 | Tags = 36%, list = 11%, signal = 40% |
| 17 | ssc04360: Axon guidance | 12 | 0.36 | 1.05 | 0.393 | 0.653 | 1 | 56 | Tags = 33%, list = 11%, signal = 37% |
| 18 | ssc04310: Wnt signaling pathway | 14 | 0.35 | 1.03 | 0.462 | 0.66 | 1 | 132 | Tags = 43%, list = 26%, signal = 56% |
| 19 | ssc04340: Hedgehog signaling pathway | 3 | 0.55 | 1 | 0.479 | 0.671 | 1 | 114 | Tags = 67%, list = 22%, signal = 85% |
| 20 | ssc00982: Drug metabolism | 12 | 0.35 | 0.99 | 0.455 | 0.651 | 1 | 40 | Tags = 25%, list = 8%, signal = 26% |
| 21 | ssc04080: Neuroactive ligand-receptor interaction | 17 | 0.29 | 0.94 | 0.525 | 0.706 | 1 | 43 | tags = 29%, list = 8%, signal = 31% |
| 22 | ssc00830: Retinol metabolism | 7 | 0.39 | 0.94 | 0.524 | 0.682 | 1 | 16 | Tags = 29%, list = 3%, signal = 29% |
| 23 | ssc00565: Ether lipid metabolism | 4 | 0.43 | 0.84 | 0.686 | 0.811 | 1 | 295 | Tags = 100%, list = 58%, signal = 233% |
| 24 | ssc05222: Small cell lung cancer | 14 | 0.26 | 0.79 | 0.718 | 0.848 | 1 | 51 | Tags = 21%, list = 10%, signal = 23% |
| 25 | ssc00480: Glutathione metabolism | 6 | 0.34 | 0.78 | 0.746 | 0.832 | 1 | 339 | Tags = 100%, list = 66%, signal = 291% |
| 26 | ssc05310: Asthma | 11 | 0.27 | 0.77 | 0.741 | 0.806 | 1 | 139 | Tags = 73%, list = 27%, signal = 98% |
| 27 | ssc00563: Glycosylphosphatidylinositol (GPI)-anchor biosynthesis | 2 | 0.49 | 0.77 | 0.769 | 0.776 | 1 | 263 | Tags = 100%, list = 51%, signal = 204% |
| 28 | ssc00601: Glycosphingolipid biosynthesis | 4 | 0.33 | 0.68 | 0.846 | 0.857 | 1 | 164 | Tags = 75%, list = 32%, signal = 109% |
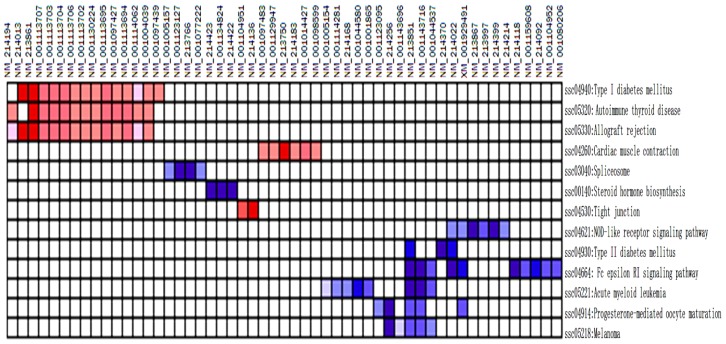
Further analysis revealed that several immune response genes were induced by leading edge analysis for the 13 significant pathways (Figure 6). The genes included those encoding CCL2, GM-CSF, HLA-B associated transcript 1, IGF-1, IL-6, IL-8, IL-18, TNF, Hsp70s, Hsp70.2, Fc fragment of IgE, MAP2K1, PIK3R5, MAPK 14, STAT3 and STAT5B, among others. Many genes related with metabolism as well as ribosomal protein genes were also induced in the inflamed lung. These genes included adiponectin, the Saccharomyces pombe cell division cycle 25 homolog C, cytochrome P450 (3A29, 3A39 and 3A46), FYN oncogene related to SRC, FGR and YES (FYN), the phosphatase and tensin homolog PTEN, PDK, Snrpa, Syk, SPI1, and v-Ha-ras, c-Myc, avian, among others.
Figure 6.
The heat map shows the clustered genes in the leading edge subsets. In the heat map, expression values are represented as colors, where the range of colors (red, pink, light blue, dark blue) represents the range of expression values (high, moderate, low, lowest) in the CG. This pattern is reversed in the TG.
The repressed genes comprised those encoding members of the MHC, including (SLA- 2, SLA-3, SLA-6, SLA-8, SLA-DRA, SLA-DQA1, SLA-DRB1, SLA-DMB, SLA-DQA, SLA-DMA, SLA-DQB1), CD40 molecule, CD40, IL-12B, IL-2, myosin, MYH, MYH2, CACNB4, CACNA2D, CPE, and FXYD2, among others.
Genes were frequently induced in the TG included p101, MAP2K1, H-RAS, TNF, MAPK14 and IGF1, while the genes such as CD40, IL12B, IL-2, SLA-2, SLA-3, SLA-6, SLA-8, SLA-DRA, LA-DQA1, SLA-DRB1, SLA-DMB, SLA-DQA, SLA-DMA and SLA-DQB1 were frequently suppressed.
2.4. Verification of Gene Expression Pattern from Microarray Data Using Real-Time QRT-PCR
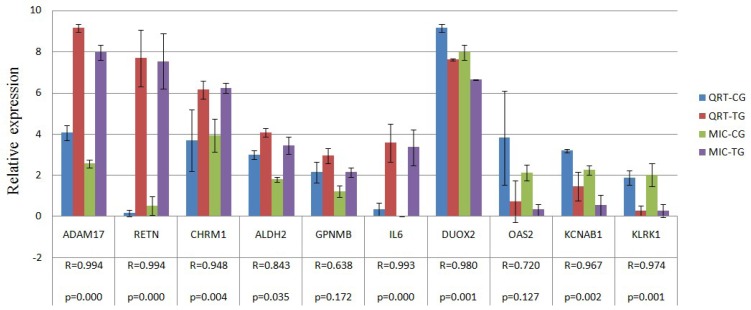
Ten genes (i.e., RETN, ADAM17, GPNMB, CHRM1, ALDH2, IL6, KLRK1, DUOX2, OAS2 and KCNAB1) were selected to confirm expression patterns using real-time qRT-PCR. The results indicate that the expression patterns of all the genes were consistent with the microarray data (r = 0.905 ± 0.125, Figure 7).
Figure 7.
Validation of the microarray data by the real-time qRT-PCR analyses of ten representative genes. The x-axis represents the genes and the y-axis shows their relative expression levels (−ΔCt) values for quantitative real-time RT-PCR; Log (Sample signal, 10) for microarray. Three biological replicates were conducted for both assays. R represents the Pearson correlation coefficient. The significance of differences for gene expression between the CG and the TG was calculated using a two-tailed T-test.
3. Discussion
In the present study, we revealed 11,929 DE genes using Agilent Whole Porcine Genome Oligo Microarrays (one-color platform) that contain 43,603 probes. There were 1188 genes annotated as swine genes in the GenBank Data Base (DB). GO term analysis identified that a total of 89 BP categories, 82 cellular components and 182 molecular functions were significantly affected and at least 27 BP categories were related to the host immune response.
The NOD-like receptor signaling pathway, Fc epsilon RI signaling pathway, acute myeloid leukemia, melanoma, progesterone-mediated oocyte maturation, spliceosome, type II diabetes mellitus and steroid hormone biosynthesis etc. were significantly enriched in inflamed lung. Five pathways as type I diabetes mellitus, autoimmune thyroid disease, allograft rejection, tight junction and cardiac muscle contraction were significantly enriched in non-inflamed lung. The NOD-like receptor signaling pathway is one of the most important pathways associated with microbial recognition and host defense [8,9]. The innate immune system comprises several classes of pattern recognition receptors, including Toll-like receptors (TLRs), NOD-like receptors (NLRs) and RIG-1-like receptors. Two NLRs, NOD1 and NOD2, sense the cytosolic presence of the peptidoglycan fragments, meso-DAP and muramyl dipeptide, respectively, and drive the activation of MAPK and the transcription factor NF-kappaB (NF-κB). A different set of NLRs induces caspase-1 activation through the assembly of large protein complexes named inflammasomes [9]. Inflammasomes are critical for generating mature proinflammatory cytokines in concert with Toll-like receptor signaling pathways [10]. Nod proteins fight off bacterial infections by stimulating proinflammatory signaling and cytokine networks and by inducing antimicrobial effectors, such as nitric oxide and antimicrobial peptides [11]. Fc epsilon RI-mediated signaling pathways in mast cells are initiated by the interaction of antigen with IgE bound to the extracellular domain of the alpha chain of the Fc epsilon RI [12–16]. The activation pathways are regulated by mast cells that release histamines and proteoglycans (especially heparin), lipid mediators such as leukotrienes (LTC4, LTD4 and LTE4) and prostaglandins (especially PDG2), and cytokines such as TNF-alpha, IL-4 and IL-5. These mediators and cytokines contribute to inflammatory response [14].
The pathways activated in infected lung tissues also include the acute myeloid leukemia pathway characterized by uncontrolled proliferation of clonal neoplastic cells and accumulation in the bone marrow of blasts with an impaired differentiation program [17–22], progesterone-mediated oocyte maturation pathway involved in endocrine system either insulin/IGF-1 or the steroid hormone progesterone regulation [23–25], steroid hormone biosynthesis pathway involved in lipid metabolism [26–29], type II diabetes mellitus involved in endocrine and metabolic diseases [30–33], spliceosome pathway involved in genetic information processing and transcription [34–36], and melanoma pathway involved in cancer arising from the malignant transformation of melanocytes [37–40].
The NOD-like receptor signaling pathway, Fc epsilon RI signaling pathway, acute myeloid leukemia pathway, progesterone-mediated oocyte maturation pathway were strongly linked to the MAPK signaling pathway, which are involved in environmental information processing and signal transduction [41–43]; the acute myeloid leukemia pathway progesterone-mediated oocyte maturation pathway, melanoma pathway were all strongly linked to the cell cycle pathway, which plays an important role in the regulation of cell growth and death [44–46]; and the NOD-like receptor signaling pathway, acute myeloid leukemia pathway, type II diabetes mellitus pathway, melanoma pathway were all directly or indirectly linked to the apoptosis pathway, which plays an important role in the regulation of apoptosis (programmed cell death) [47–49]. Hence, the pathways linked to the cell function regulation, including the MAPK signaling pathway, apoptosis and Cell cycle pathway, were also affected directly or indirectly by the process of the body’s resistance to infection.
Many cytokines as shown by leading edge analysis were activated at the site of inflammation, including IL-6, IL-8, IL-18, TNF, GM-CSF, CCL2, p101 protein, HLA-B associated transcript 1, Fc fragment of IgE, MAPK14, MAP2K1, IGF-1, STAT3 and STAT5B, etc. IL-6 is responsible for stimulating acute phase protein synthesis, as well as the production of neutrophils in the bone marrow. IL-8 is synthesized by macrophages, endothelial cells and epithelial cells as host defenses against severe infection [50,51]. It serves as a chemical signal that attracts neutrophils to the site of any inflammation. Significant increases in IL-8 and IL6- mRNA after infection with APP have previously been observed in lung lavage as well as lung tissue using northern blotting and in situ hybridization [52,53]. IL-18 plays multiple roles in chronic inflammation and in a number of infections and enhances both Th-1- and Th-2-mediated immune response [54]. IL-18 is able to induce IFN gamma, GM-CSF, TNF-α and IL-1 in immunocompetent cells to activate killing by lymphocytes and to up-regulate the expression of certain chemokine receptors. GM-CSF stimulates stem cells to produce granulocytes (neutrophils, eosinophils, and basophils) and monocytes. Monocytes exit the circulation and migrate into tissues, whereupon they mature into macrophages. Thus, these cells play a part in the immune-inflammatory cascade, by which activation of a small number of macrophages can rapidly lead to an increase in their number, a process crucial for fighting infection. CCL2 recruits monocytes, memory T cells and dendritic cells to the sites of tissue injury, infection, and inflammation [55,56]. TNF-α can promote inflammatory response by inducing the production of other proinflammatory cytokines at the vicinity of the infection [57], and increase the expression of endothelial surface HLA-B by activation of the nuclear transcription factor NF-κB [58,59]. P101 protein is a single regulatory subunit of the phosphoinositide 3-kinase gamma (PI3Kα), which plays a crucial role in inflammatory and allergic processes [60,61], including neutrophil chemotaxis, mast cell degranulation, and cardiac function [62,63].
Genes involved in a variety of cellular function, including proliferation, differentiation, growth arrest or apoptosis of normal cells were affected including those encoding HLA-B associated transcript 1 [64,65], Fc fragment of IgE [66,67], MAPK14 [68], MAP2K1 [69], H-ras [70,71], IGF-1 [72], STAT3 and STAT5B [73]. Activations of all these genes can stimulate stem cells to produce granulocytes (neutrophils, eosinophils, and basophils) and monocytes, and also induce neutrophils and macrophages to phagocytose bacterial and foreign antigens.
Immunomodulatory cytokines were significantly suppressed at the site of inflammation. In this study, genes encoding IL2, IL12B, CD40, members of the MHC (SLA-2, SLA-3, SLA-6, SLA-8, SLA-DRB1, SLA-DMB, SLA-DQA, SLA-DMA and SLA-DQB1), as well as SLA-DRA and SLA-DQA1 in a previous study [74], were significantly down-regulated at the site of inflammation.
IL-2 is a type of cytokine immune system signaling molecule which is a leukocytotrophic hormone made in response to microbial infection that can identify the difference between self and non-self [75,76]. When environmental substances (molecules or microbes) gain access to the body, these substances (termed antigens) are recognized as foreign by antigen receptors that are expressed on the surface of lymphocytes. MHC can present antigenic peptides to T lymphocytes, which are responsible for a specific immune response that can destroy the pathogen producing those antigens [77]. CD40 is a co-stimulatory protein found on antigen presenting cells (APC) and is essential in mediating a broad variety of immune and inflammatory responses including T cell-dependent immunoglobulin class switching, memory B cell development, and germinal center formation [78]. The macrophage can express more CD40 and TNF receptors on its surface, which can increase the level of activation culminating in the induction of potent microbicidal substances in the macrophage; these include reactive oxygen species and nitric oxide, leading to the destruction of the ingested microbe [79–82]. IL-12 is an essential inducer of Th1 cell development, and has an important role in sustaining a sufficient number of memory/effector Th1 cells to mediate long-term protection against an intracellular pathogen [83]. The suppression of these immunomodulatory cytokines leads to a decrease in antigenic peptides presented to T lymphocytes by APC via the MHC, as well as to alleviate immune response injury induced by infection at the site of inflammation.
Many genes encoding metabolism as well as ribosomal proteins were affected at the site of inflammation. Genes related to metabolism and ribosomal proteins synthesis were induced in the inflamed lung, including adiponectin, cell division cycle 25 homolog C, cytochrome P450 (CYP 3A29, CYP 3A39 and CYP 3A46), FYN, PTEN, PDK, Snrpa, Syk and SPI1, among others. The repressed genes comprised those encoding MYH1, MYH2, tropomyosin (alpha, beta), troponin I type 3 (cardiac), CACNB4, CACNA2D1, CPE and FXYD2; while the CYP2E1 and the CYP3A29 were known to be down-regulated during inflammation in another study [84].
SOCS3 and CISH, both found to be up-regulated in the present study, are members of the suppressor of cytokine signaling (SOCS) family of proteins whose members regulate protein turnover by targeting proteins for degradation [42]. Expression of the members of the SOCS family is induced by cytokines such as IL-6 and IL-10, both found to be up-regulated in this study; both function as negative feed- back regulators of cytokine signaling [85,86]. The statistically significant increase in mRNA coding for the anti-inflammatory cytokine IL-10, found in inflamed areas of the lung, is probably due to the function of IL-10 in counteracting the host mediated tissue damage caused by proinflammatory and chemotactic cytokines [87]. The lower expression levels observed for genes encoding ribosomal proteins could be due to a general downregulation of ribosomal biogenesis in the necrotic areas of the lung. Previous studies have shown that 41 out of 54 genes encoding ribosomal proteins were down-regulated in Pseudomonas aeruginosa after treatment with H2O2 induced oxidative stress [88].
As described above, we found that: (1) A total of 89 biological process categories, 82 cellular components and 182 molecular functions were significantly affected, and more than 27 biological process were involved in the host immune response; (2) At the site of inflammation, 13 pathways associated with host responses were affected significantly; many proinflammatory-inflammatory cytokines were activated and several immunomodulatory cytokines were suppressed at the gene expression level reflecting the complex machinery at work during an infection; (3) Many genes which were involved in a variety of cellular functions-proliferation, differentiation, growth arrest or apoptosis of normal cells that activated, could stimulate stem cells to produce granulocyte (neutrophil, eosinophil, and basophil) and monocyte. All changes of the genes and pathways which induced or repressed expression, not only led to decrease in antigenic peptides presented to T lymphocytes by APC via the MHC and alleviated immune response injury induced by infection, but also stimulated stem cells to produce granulocyte (neutrophil, eosinophil, and basophil ) and monocyte, and promote neutrophil and macrophages to phagocytose bacterial and foreign antigen at the site of inflammation. Additional work including more animals and time points is clearly needed to further delineate the host response to APP infection and will contribute to a more detailed description of the dynamics of host responses in general.
4. Experimental Section
4.1. Animals, Bacterial Inoculation and Samples
All animal procedures were performed according to protocols approved by the Biological Studies Animal Care and Use Committee of Sichuan Province, China. Twenty 12-week-old male castrated Danish Landrace/Yorkshire/Duroc crossbred swine from a healthy herd free from APP were divided equally into a control group (CG) and the treatment group (TG). APP serotype I (Strain provided by the Animal Biotechnology Center, Laboratory of Animal Disease and Human Health, Sichuan Agricultural University) was cultivated overnight at 37 °C in air on trypticase soy broth (TSB) (Hangwei, Hangzhou, China). Bacterial counts of the suspensions were performed at the same time as the start of the inoculation. The inoculation was performed by holding the pigs (1–10) from the TG in an upright sitting position and spraying 0.25mL diluent containing (3.5–4) × 107 CFU/mL APP per kilogram weight into the nostrils during inspiration. Swine from the CG (swines 11–20) were inoculated with physiological saline (0.9% wt/vol NaCl) by the same means. In the TG, lung tissue was collected from three swine (swines 1, 2 and 3) after abattage and used for total RNA extraction and pathological analysis. Another three swine (swines 11, 12 and 13) from the CG were sacrificed 48 h post-inoculation and their lung tissues were collected. The remaining swine were used for other trials.
4.2. Microarray Hybridizations and Data Analysis
Total RNA was extracted from tissues using Trizol reagent (Invitrogen, Carlsbad, CA, USA). RNA was purified and DNase treated using the RNeasy QIAGEN RNeasy® Mini Kit. cDNA was synthesized from 2 μg of total-RNA using the direct cDNA Labeling System. Aminoallyl-cRNA was synthesized from cDNA using the Superscript Indirect cDNA Labeling System. The cRNA was purified and DNase treated using RNeasy QIAGEN RNeasy® Mini Kit. RNA integrity was confirmed with a bioanalyzer (model 2100; Agilent Technologies, Palo Alto, CA, USA) according to the manufacturer’s protocol. Labeling and hybridization of the cRNA was performed with Agilent Whole Porcine Genome Oligo (4 × 44 K) Microarrays (one-color platform) at the National Engineering Center for Biochip at Shanghai, according to the manufacturer’s protocols. The slides were scanned and analyzed using the histogram method with default settings in an Agilent G2565AA and Agilent G2565BA Microarray Scanner System with SureScan Technology. The array data were submitted to GEO [89].
Comparisons between the CG and TG were carried out using three biological replicates for each group. CG samples and TG samples were used for microarray analysis. The six Microarray data were normalized using the quantile normalization method [90] with WebarrayDB (http://www.webarraydb.org/webarray/) [91] and were filtered and assessed by the MIDAW online analysis program (http://www.webarraydb.org/webarray/) [92] using the method of weighted K-nearest neighbor [93]. T-tests and hierarchical cluster analyses of the significantly differentially expressed (DE) genes (clustering method: complete linkage; similarity measure: Pearson product momentum correlation; ordering function: average value) for microarray data were carried out by a MultiExperiment Viewer (MeV) software package (Version 4.5, Dana-Farber Cancer Institute, Boston, MA, USA, 2009) [94].
Tests for statistical significance (p < 0.05), overrepresentation of Gene ontology (GO) terms [74,95], and pathway in Kyoto Encyclopedia of Genes and Genomes (KEGG) DB [10,96] (http://www.genome.jp/kegg/) both induced and repressed genes were conducted using the ErmineJ [97] and the Database for Annotation, Visualization and Integrated Discovery (DAVID) Online platform (http://david.abcc.ncifcrf.gov/) with a threshold of a minimum three genes annotated at each node. The leading edge analysis for the pathway of differential expression in microarray data with a threshold of a minimum of two genes and maximum of 20 genes annotated at each node was conducted using the GSEA V2.06 package [98,99]. More detailed descriptions of the microarray experiments are available at the NCBI Gene Expression Omnibus [100–102].
4.3. Real-Time QRT-PCR
In order to confirm the reliability the expression profile in the microarray analyses, the expression level 10 gene (six up-regulated and four down-regulated) were performed by real-time qRT-PCR. Sequences for primers were obtained from Genbank and NCBI. Primers were designed using Primer 5 and synthesized at Invitrogen (Shanghai, China) (Table 1). Extracted RNA was converted into cDNA by reverse transcription of 1 μL total RNA using SYBR® PrimeScriptTM RT-PCR Kit (TaKaRa, Japan) according to the manufacturer’s protocol and then cDNA was stored at −20 °C until use. Quantitative PCR was performed in a 25 μL reaction volume (2 μL cDNA, 12.5 μL of SYBR® Premix Ex TaqTM (2×) TaKaRa, Japan), 0.5 μL of 10 μM upstream and downstream primers respectively, and added ddH2O to 25 μL) on the BIO-RAD IQ5 System (BIO-RAD, Hercules, CA, USA). Real-time PCR conditions were as follows: 30 s at 95.0 °C, 40 cycles of denaturation at 95 °C for 5 s followed by 30 s annealing and elongation at 51.2–60 °C (Table 8). Efficiency of primer pairs is reported in Table 1. Melting curves were obtained at the end of each run to confirm a single PCR product. All samples were run in triplicate. Non-template controls were included in each run to exclude contamination and nonspecific amplification. Expression levels of samples were normalised by using a normalisation factor calculated by the program geNorm. This normalisation factor was calculated based on RT-qPCR results for three selected reference genes, ACTB, TOP2B and TBP.
Table 8.
Information on the primers used for qRT-PCR.
| Confirmation objects | Gene symbol | Primer sequence (5′→3′) | Amplicon length (bp) | Ta (°C) | GenBank No. |
|---|---|---|---|---|---|
| Reference gene | ACTB | TCTGGCACCACACCTTCT | 114 | 60 | DQ178122 |
| TGATCTGGGTCATCTTCTCAC | |||||
|
| |||||
| TBP | GATGGACGTTCGGTTTAGG | 124 | 60 | DQ178129 | |
| AGCAGCACAGTACGAGCAA | |||||
|
| |||||
| TOP2B | AACTGGATGATGCTAATGATGCT | 137 | 60 | AF222921 | |
| TGGAAAAACTCCGTATCTGTCTC | |||||
|
| |||||
| Up gene | RETN | AGTGCGCTGGCATAGACTGG | 197 | 60 | NM_213783 |
| CATCCTCTTCTCAAGGTTTATTTCC | |||||
|
| |||||
| ADAM17 | TTGAGGAAGGGGAAGCC | 158 | 56 | NM_001099926 | |
| ACGGAGCCCACGATGTT | |||||
|
| |||||
| GPNMB | GAGACCCAGCCTTCCTT | 130 | 51.2 | NM_001098584 | |
| TTGCTTTCTATCGCTTTGTA | |||||
|
| |||||
| CHRM1 | CGCTGGTCAAGGAGAAGAA | 185 | 56 | NM_214034 | |
| GCACATGGGGTTGATGGT | |||||
|
| |||||
| ALDH2 | AAACTGCTCTGCGGTGGA | 181 | 56 | NM_001044611 | |
| CGTACTTGGAATTGTTGGCTC | |||||
|
| |||||
| IL6 | GTCGAGGCTGTGCAGATTAG | 101 | 56 | NM_214399 | |
| GCATTTGTGGTGGGGTTAG | |||||
|
| |||||
| Down gene | KLRK1 | TGATGTGATAAACCGTGGTG | 107 | 56 | NM_213813 |
| TGGATCGGGCAAGGAAA | |||||
|
| |||||
| DUOX2 | CCCTTCTTCAACTCCCTG | 158 | 51.2 | NM_213999 | |
| CAAAAGTTCTCATAGTGGTGC | |||||
|
| |||||
| OAS2 | GACACGGCTGAAGGTTT | 291 | 51.2 | NM_001031796 | |
| TGGCACGTCCCAAGACT | |||||
|
| |||||
| KCNAB1 | AAGGGAGAAAACAGCAAAAC | 176 | 56 | NM_001105294 | |
| AACCTGAATGGCACCGA | |||||
This allowed quantification of the target gene in one sample relative to that in another (the calibrator) using the “2−ΔΔCt method” of calculating fold changes in gene expression [103]. Correlation analysis between qRT-PCR and microarray was conducted.
5. Conclusions
We have generated reliable mRNA transcriptomes of swine lung tissues from APP-infected and negative control pigs. We have identified a set of differentially expressed (DE) genes in our current case-control study, and a functional enrichment analysis indicated that these DE genes mainly related to “host immune response” and “host response”. In addition, we also found that, in the APP-infected lung tissues, many proinflammatory-inflammatory cytokines were activated and involved in the regulation of the host defense response at the site of inflammation, while the cytokines involved in regulation of the host immune response were suppressed. The current study provides data that can be used in future studies to decipher the molecular mechanism of the systematic influences from porcine pleuropneumonia. Our findings will also help promote the further development of therapy for porcine pleuropneumonia.
Acknowledgments
We thank the farmer that provided swine. This work was supported financially by the Natural Science Foundation of Science and Technology Department of Sichuan Province.
Conflict of Interest
The authors declare no conflict of interest. Our experiments involving the use of swine, and the use of pigs and all experimental procedures involving animals were approved by Sichuan Agricultural University Animal Care and Use Committee.
References
- 1.Taylor D.J. Actinobacillus Pleuropneumoniae. In: Straw B.E., d’Allaire S., Mengeling W.L., Taylor D.J., editors. Diseases of Swine. Vol. 26. Iowa State University Press; Ames, IA, USA: 1999. pp. 343–354. [Google Scholar]
- 2.Baarsch M.J., Foss D.L., Murtaugh M.P. Pathophysiologic correlates of acute porcine pleuropneumonia. Am. J. Vet. Res. 2000;61:684–690. doi: 10.2460/ajvr.2000.61.684. [DOI] [PubMed] [Google Scholar]
- 3.Cho W.S., Chae C. Expression of nitric oxide synthase 2 and tumor necrosis factor alpha in swine naturally infected with Actinobacillus. pleuropneumoniae. Vet. Pathol. 2002;39:27–32. doi: 10.1354/vp.39-1-27. [DOI] [PubMed] [Google Scholar]
- 4.Cho W.S., Jung K., Kim J., Ha Y., Chae C. Expression of mRNA encoding interleukin (IL)-10, IL-12p35 and IL-12p40 in lungs from swines experimentally infected with Actinobacillus pleuropneumoniae. Vet. Res. Commun. 2005;29:111–122. doi: 10.1023/b:verc.0000047488.05304.3e. [DOI] [PubMed] [Google Scholar]
- 5.Moser R.J., Reverter A., Kerr C.A., Beh K.J., Lehnert S.A. A mixedmodel approach for the analysis of cDNA microarray gene expression data from extreme-performing swines after infection with Actinobacillus pleuropneumoniae. J. Anim. Sci. 2004;82:1261–1271. doi: 10.2527/2004.8251261x. [DOI] [PubMed] [Google Scholar]
- 6.Hedegaard J., Skovgaard K., Mortensen S., Sorensen P., Jensen T., Hornshoj H., Bendixen C., Heegaard P.M.H. Molecular characterisation of the early response in pigs to experimental infection with Actinobacillus pleuropneumoniae using cDNA microarrays. Acta Vet. Scand. 2007;49:11. doi: 10.1186/1751-0147-49-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Mortensen S., Skovgaard K., Hedegaard J., Bendixen C., Heegaard P.M. Transcriptional profiling at different sites in lungs of pigs during acute bacterial respiratory infection. Innate Immun. 2011;17:41–53. doi: 10.1177/1753425909349760. [DOI] [PubMed] [Google Scholar]
- 8.Shaw M.H., Reimer T., Kim Y.G., Nunez G. NOD-like receptors (NLRs): Bona fide intracellular microbial sensors. Curr. Opin. Immunol. 2008;20:377–382. doi: 10.1016/j.coi.2008.06.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Kanneganti T.D., Lamkanfi M., Nunez G. Intracellular NOD-like receptors in host defense and disease. Immunity. 2007;27:549–559. doi: 10.1016/j.immuni.2007.10.002. [DOI] [PubMed] [Google Scholar]
- 10.Aoki-Kinoshita K.F., Kanehisa M. Gene annotation and pathway mapping in KEGG. Methods Mol. Biol. 2007;396:71–91. doi: 10.1007/978-1-59745-515-2_6. [DOI] [PubMed] [Google Scholar]
- 11.Tattoli I., Travassos L.H., Carneiro L.A., Magalhaes J.G., Girardin S.E. The Nodosome: Nod1 and Nod2 control bacterial infections and inflammation. Semin. Immunopathol. 2007;29:289–301. doi: 10.1007/s00281-007-0083-2. [DOI] [PubMed] [Google Scholar]
- 12.Kawakami T., Galli S.J. Regulation of mast-cell and basophil function and survival by IgE. Nat. Rev. Immunol. 2002;2:773–786. doi: 10.1038/nri914. [DOI] [PubMed] [Google Scholar]
- 13.Siraganian R.P. Mast cell signal transduction from the high-affinity IgE receptor. Curr. Opin. Immunol. 2003;15:639–646. doi: 10.1016/j.coi.2003.09.010. [DOI] [PubMed] [Google Scholar]
- 14.Turner H., Kinet J.P. Signalling through the high-affinity IgE receptor Fc epsilonRI. Nature. 1999;402:B24–B30. doi: 10.1038/35037021. [DOI] [PubMed] [Google Scholar]
- 15.Gilfillan A.M., Tkaczyk C. Integrated signalling pathways for mast-cell activation. Nat. Rev. Immunol. 2006;6:218–230. doi: 10.1038/nri1782. [DOI] [PubMed] [Google Scholar]
- 16.Nadler M.J., Matthews S.A., Turner H., Kinet J.P. Signal transduction by the high-affinity immunoglobulin E receptor Fc epsilon RI: Coupling form to function. Adv. Immunol. 2000;76:325–355. doi: 10.1016/s0065-2776(01)76022-1. [DOI] [PubMed] [Google Scholar]
- 17.Steffen B., Muller-Tidow C., Schwable J., Berdel W.E., Serve H. The molecular pathogenesis of acute myeloid leukemia. Crit. Rev. Oncol. Hematol. 2005;56:195–221. doi: 10.1016/j.critrevonc.2004.10.012. [DOI] [PubMed] [Google Scholar]
- 18.Tenen D.G. Disruption of differentiation in human cancer: AML shows the way. Nat. Rev. Cancer. 2003;3:89–101. doi: 10.1038/nrc989. [DOI] [PubMed] [Google Scholar]
- 19.Martelli A.M., Nyakern M., Tabellini G., Bortul R., Tazzari P.L., Evangelisti C., Cocco L. Phosphoinositide 3-kinase/Akt signaling pathway and its therapeutical implications for human acute myeloid leukemia. Leukemia. 2006;20:911–928. doi: 10.1038/sj.leu.2404245. [DOI] [PubMed] [Google Scholar]
- 20.Lennartsson J., Jelacic T., Linnekin D., Shivakrupa R. Normal and oncogenic forms of the receptor tyrosine kinase kit. Stem Cells. 2005;23:16–43. doi: 10.1634/stemcells.2004-0117. [DOI] [PubMed] [Google Scholar]
- 21.Lorsbach R.B., Downing J.R. The role of the AML1 transcription factor in leukemogenesis. Int. J. Hematol. 2001;74:258–265. doi: 10.1007/BF02982058. [DOI] [PubMed] [Google Scholar]
- 22.Mizuki M., Schwäble J., Steur C., Choudhary C., Agrawal S., Sargin B., Steffen B., Matsumura I., Kanakura Y., Böhmer F.D. Suppression of myeloid transcription factors and induction of STAT response genes by AML-specific Flt3 mutations. Blood. 2003;101:3164–73. doi: 10.1182/blood-2002-06-1677. [DOI] [PubMed] [Google Scholar]
- 23.Nebreda A.R., Ferby I. Regulation of the meiotic cell cycle in oocytes. Curr. Opin. Cell. Biol. 2000;12:666–675. doi: 10.1016/s0955-0674(00)00150-2. [DOI] [PubMed] [Google Scholar]
- 24.Andersen C.B., Sakaue H., Nedachi T., Kovacina K.S., Clayberger C., Conti M., Roth R.A. Protein kinase B/Akt is essential for the insulin- but not progesterone-stimulated resumption of meiosis in Xenopus oocytes. Biochem. J. 2003;369:227–238. doi: 10.1042/BJ20021243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Abrieu A., Doree M., Fisher D. The interplay between cyclin-B-Cdc2 kinase (MPF) and MAP kinase during maturation of oocytes. J. Cell. Sci. 2001;114:257–267. doi: 10.1242/jcs.114.2.257. [DOI] [PubMed] [Google Scholar]
- 26.Ghayee H.K., Auchus R.J. Basic concepts and recent developments in human steroid hormone biosynthesis. Rev. Endocr. Metab. Disord. 2007;8:289–300. doi: 10.1007/s11154-007-9052-2. [DOI] [PubMed] [Google Scholar]
- 27.Holmes M.C., Seckl J.R. The role of 11beta-hydroxysteroid dehydrogenases in the brain. Mol. Cell. Endocrinol. 2006;248:9–14. doi: 10.1016/j.mce.2005.12.002. [DOI] [PubMed] [Google Scholar]
- 28.Thomas J.L., Duax W.L., Addlagatta A., Brandt S., Fuller R.R., Norris W. Structure/function relationships responsible for coenzyme specificity and the isomerase activity of human type 13 beta-hydroxysteroid dehydrogenase/isomerase. J. Biol. Chem. 2003;278:35483–35490. doi: 10.1074/jbc.M304752200. [DOI] [PubMed] [Google Scholar]
- 29.Tomlinson J.W., Stewart P.M. Cortisol metabolism and the role of 11beta-hydroxysteroid dehydrogenase. Best Pract. Res. Clin. Endocrinol. Metab. 2001;15:61–78. doi: 10.1053/beem.2000.0119. [DOI] [PubMed] [Google Scholar]
- 30.Gual P., le Marchand-Brustel Y., Tanti J.F. Positive and negative regulation of insulin signaling through IRS-1 phosphorylation. Biochimie. 2005;87:99–109. doi: 10.1016/j.biochi.2004.10.019. [DOI] [PubMed] [Google Scholar]
- 31.Kaneto H., Matsuoka T.-a., Nakatani Y., Kawamori D., Miyatsuka T., Matsuhisa M., Yamasaki Y. Oxidative stress, ER stress, and the JNK pathway in type 2 diabetes. J. Mol. Med. 2005;83:429–439. doi: 10.1007/s00109-005-0640-x. [DOI] [PubMed] [Google Scholar]
- 32.Sakai K., Matsumoto K., Nishikawa T., Suefuji M., Nakamaru K., Hirashima Y., Kawashima J., Shirotani T., Ichinose K., Brownlee M. Mitochondrial reactive oxygen species reduce insulin secretion by pancreatic beta-cells. Biochem. Biophys. Res. Commun. 2003;300:216–222. doi: 10.1016/s0006-291x(02)02832-2. [DOI] [PubMed] [Google Scholar]
- 33.Robertson R.P. Chronic oxidative stress as a central mechanism for glucose toxicity in pancreatic islet beta cells in diabetes. J. Biol. Chem. 2004;279:42351–42354. doi: 10.1074/jbc.R400019200. [DOI] [PubMed] [Google Scholar]
- 34.Wahl M.C., Will C.L., Luhrmann R. The spliceosome: Design principles of a dynamic RNP machine. Cell. 2009;136:701–718. doi: 10.1016/j.cell.2009.02.009. [DOI] [PubMed] [Google Scholar]
- 35.Chen Y.-I.G., Moore R.E., Helen Y.G., Young M.K., Lee T.D., Stevens S.W. Proteomic analysis of in vivo-assembled pre-mRNA splicing complexes expands the catalog of participating factors. Nucleic Acids Res. 2007;35:3928–3944. doi: 10.1093/nar/gkm347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ritchie D.B., Schellenberg M.J., Macmillan A.M. Spliceosome structure: Piece by piece. Biochim. Biophys. Acta. 2009;1789:624–633. doi: 10.1016/j.bbagrm.2009.08.010. [DOI] [PubMed] [Google Scholar]
- 37.Levy C., Khaled M., Fisher D.E. MITF: Master regulator of melanocyte development and melanoma oncogene. Trends Mol. Med. 2006;12:406–414. doi: 10.1016/j.molmed.2006.07.008. [DOI] [PubMed] [Google Scholar]
- 38.Tsao H., Goel V., Wu H., Yang G., Haluska F.G. Genetic interaction between NRAS and BRAF mutations and PTEN/MMAC1 inactivation in melanoma. J. Invest. Dermatol. 2004;122:337–341. doi: 10.1046/j.0022-202X.2004.22243.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Gruss C., Herlyn M. Role of cadherins and matrixins in melanoma. Curr. Opin. Oncol. 2001;13:117–123. doi: 10.1097/00001622-200103000-00006. [DOI] [PubMed] [Google Scholar]
- 40.Lázár-Molnár E., Hegyesi H., Tóth S., Falus A. Autocrine and paracrine regulation by cytokines and growth factors in melanoma. Cytokine. 2000;12:547–554. doi: 10.1006/cyto.1999.0614. [DOI] [PubMed] [Google Scholar]
- 41.Tanoue T., Nishida E. Docking interactions in the mitogen-activated protein kinase cascades. Pharmacol. Ther. 2002;93:193–202. doi: 10.1016/s0163-7258(02)00188-2. [DOI] [PubMed] [Google Scholar]
- 42.Yang S.H., Sharrocks A.D., Whitmarsh A.J. Transcriptional regulation by the MAP kinase signaling cascades. Gene. 2003;320:3–21. doi: 10.1016/s0378-1119(03)00816-3. [DOI] [PubMed] [Google Scholar]
- 43.Zhu X., Lee H.G., Raina A.K., Perry G., Smith M.A. The role of mitogen-activated protein kinase pathways in Alzheimer’s disease. Neurosignals. 2002;11:270–281. doi: 10.1159/000067426. [DOI] [PubMed] [Google Scholar]
- 44.Smith M.L., Chen I.T., Zhan Q., Bae I., Chen C.Y., Gilmer T.M., Kastan M.B., O’Connor P.M., Fornace A.J., Jr Interaction of the p53-regulated protein Gadd45 with proliferating cell nuclear antigen. Science. 1994;266:1376–1380. doi: 10.1126/science.7973727. [DOI] [PubMed] [Google Scholar]
- 45.Diehl J.A., Cheng M., Roussel M.F., Sherr C.J. Glycogen synthase kinase-3beta regulates cyclin D1 proteolysis and subcellular localization. Genes Dev. 1998;12:3499–3511. doi: 10.1101/gad.12.22.3499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Klein E.A., Assoian R.K. Transcriptional regulation of the cyclin D1 gene at a glance. J. Cell. Sci. 2008;121:3853–3857. doi: 10.1242/jcs.039131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Sprick M.R., Walczak H. The interplay between the Bcl-2 family and death receptor-mediated apoptosis. Biochim. Biophys. Acta. 2004;1644:125–132. doi: 10.1016/j.bbamcr.2003.11.002. [DOI] [PubMed] [Google Scholar]
- 48.Steelman L.S., Pohnert S.C., Shelton J.G., Franklin R.A., Bertrand F.E., McCubrey J.A. JAK/STAT, Raf/MEK/ERK, PI3K/Akt and BCR-ABL in cell cycle progression and leukemogenesis. Leukemia. 2004;18:189–218. doi: 10.1038/sj.leu.2403241. [DOI] [PubMed] [Google Scholar]
- 49.Martinon F., Holler N., Richard C., Tschopp J. Activation of a pro-apoptotic amplification loop through inhibition of NF-κB-dependent survival signals by caspase-mediated inactivation of RIP. FEBS Lett. 2000;468:134–136. doi: 10.1016/s0014-5793(00)01212-6. [DOI] [PubMed] [Google Scholar]
- 50.Wolff B., Burns A.R., Middleton J., Rot A. Endothelial cell “memory” of inflammatory stimulation: Human venular endothelial cells store interleukin 8 in Weibel-Palade bodies. J. Exp. Med. 1998;188:1757–1762. doi: 10.1084/jem.188.9.1757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Utgaard J.O., Jahnsen F.L., Bakka A., Brandtzaeg P., Haraldsen G. Rapid secretion of prestored interleukin 8 from Weibel-Palade bodies of microvascular endothelial cells. J. Exp. Med. 1998;188:1751–1756. doi: 10.1084/jem.188.9.1751. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Baarsch M.J., Scamurra R.W., Burger K., Foss D.L., Maheswaran S.K., Murtaugh M.P. Inflammatory cytokine expression in swine experimentally infected with Actinobacillus pleuropneumoniae. Infect. Immun. 1995;63:3587–3594. doi: 10.1128/iai.63.9.3587-3594.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Myers M.J., Baarsch M.J., Murtaugh M.P. Effects of pentoxifylline on inflammatory cytokine expression and acute pleuropneumonia in swine. Immunobiology. 2002;205:17–34. doi: 10.1078/0171-2985-00108. [DOI] [PubMed] [Google Scholar]
- 54.Nakanishi K., Yoshimoto T., Tsutsui H., Okamura H. Interleukin-18 regulates both Th1 and Th2 responses. Annu. Rev. Immunol. 2001;19:423–474. doi: 10.1146/annurev.immunol.19.1.423. [DOI] [PubMed] [Google Scholar]
- 55.Carr M.W., Roth S.J., Luther E., Rose S.S., Springer T.A. Monocyte chemoattractant protein 1 acts as a T-lymphocyte chemoattractant. Proc. Natl. Acad. Sci. USA. 1994;91:3652–3656. doi: 10.1073/pnas.91.9.3652. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Xu L.L., Warren M.K., Rose W.L., Gong W., Wang J.M. Human recombinant monocyte chemotactic protein and other C–C chemokines bind and induce directional migration of dendritic cells in vitro. J. Leukoc. Biol. 1996;60:365–371. doi: 10.1002/jlb.60.3.365. [DOI] [PubMed] [Google Scholar]
- 57.Toews G.B. Cytokines and the lung. Eur. Respir. J. 2001;34:S3–S17. doi: 10.1183/09031936.01.00266001. [DOI] [PubMed] [Google Scholar]
- 58.Krönke M., Schütze S., Scheurich P., Pfizenmaier K. TNF signal transduction and TNF-responsive genes. Immunol. Ser. 1992;56:189–216. [PubMed] [Google Scholar]
- 59.Schütze S., Potthoff K., Machleidt T., Berkovic D., Wiegmann K., Kronke M. TNF activates NF-kappa B by phosphatidylcholine-specific phospholipase C-induced “acidic” sphingomyelin breakdown. Cell. 1992;71:765–776. doi: 10.1016/0092-8674(92)90553-o. [DOI] [PubMed] [Google Scholar]
- 60.Brock C., Schaefer M., Reusch H.P., Czupalla C., Michalke M., Spicher K., Schultz G., Nürnberg B. Roles of G beta gamma in membrane recruitment and activation of p110 gamma/p101 phosphoinositide 3-kinase gamma. J. Cell. Biol. 2003;160:89–99. doi: 10.1083/jcb.200210115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Voigt P., Brock C., Nürnberg B., Schaefer M. Assigning functional domains within the p101 regulatory subunit of phosphoinositide 3-kinase γ. J. Biol. Chem. 2005;280:5121–5127. doi: 10.1074/jbc.M413104200. [DOI] [PubMed] [Google Scholar]
- 62.Wymann M.P., Zvelebil M., Laffargue M. Phosphoinositide 3-kinase signalling—Which way to target? Trends Pharmacol. Sci. 2003;24:366–376. doi: 10.1016/S0165-6147(03)00163-9. [DOI] [PubMed] [Google Scholar]
- 63.Crackower M.A., Oudit G.Y., Kozieradzki I., Sarao R., Sun H., Sasaki T., Hirsch E., Suzuki A., Shioi T., Irie-Sasaki J., et al. Regulation of myocardial contractibility and cell size by distinct PI3K-PTEN signaling pathways. Cell. 2002;110:737–749. doi: 10.1016/s0092-8674(02)00969-8. [DOI] [PubMed] [Google Scholar]
- 64.Peelman L.J., Chardon P., Nunes M., Renard C., Geffrotin C., Vaiman M., Zeveren A.V., Coppieters W., Weghe A.V., Bouquet Y., et al. The BAT1 gene in the MHC encodes an evolutionarily conserved putative nuclear RNA helicase of the DEAD family. Genomics. 1995;26:210–218. doi: 10.1016/0888-7543(95)80203-x. [DOI] [PubMed] [Google Scholar]
- 65.Spies T., Blanck G., Bresnahan M., Sands J., Strominger J.L. A new cluster of genes within the human major histocompatibility complex. Science. 1989;243:214–217. doi: 10.1126/science.2911734. [DOI] [PubMed] [Google Scholar]
- 66.Von Bubnoff D., Novak N., Kraft S., Bieber T. The central role of FcɛRI in allergy. Clin. Exp. Dermatol. 2003;28:184–187. doi: 10.1046/j.1365-2230.2003.01209.x. [DOI] [PubMed] [Google Scholar]
- 67.Gounni A., Lamkhioued B., Ochiai K., Tanaka Y., Delaporte E., Capron A., Kinet J.P., Capron M. High-affinity IgE receptor on eosinophils is involved in defence against parasites. Nature. 1994;367:183–186. doi: 10.1038/367183a0. [DOI] [PubMed] [Google Scholar]
- 68.Gesslein B., Hakansson G., Carpio R., Gustafsson L., Perez M.T., Malmsjo M. Mitogen-activated protein kinases in the porcine retinal arteries and neuroretina following retinal ischemia-reperfusion. Mol. Vis. 2010;16:392–407. [PMC free article] [PubMed] [Google Scholar]
- 69.Kubota Y., O’Grady P., Saito H., Takekawa M. Oncogenic Ras abrogates MEK SUMO ylation that suppresses the ERK pathway and cell transformation. Nat. Cell. Biol. 2011;13:282–291. doi: 10.1038/ncb2169. [DOI] [PubMed] [Google Scholar]
- 70.Reuther G.W., Der C.J. The Ras branch of small GTPases: Ras family members don’t fall far from the tree. Curr. Opin. Cell. Biol. 2000;12:157–165. doi: 10.1016/s0955-0674(99)00071-x. [DOI] [PubMed] [Google Scholar]
- 71.Malumbres M., Pellicer A. RAS pathways to cell cycle control and cell transformation. Front. Biosci. 1998;3:887–912. doi: 10.2741/a331. [DOI] [PubMed] [Google Scholar]
- 72.Yilmaz A., Davis M.E., Simmen R.C. Reproductive performance of bulls divergently selected on the basis of blood serum insulin-like growth factor I concentration. J. Anim. Sci. 1999;77:835–839. doi: 10.2527/1999.774835x. [DOI] [PubMed] [Google Scholar]
- 73.Kisseleva T., Bhattacharya S., Braunstein J., Schindler C.W. Signaling through the JAK/STAT pathway, recent advances and future challenges. Gene. 2002;285:1–24. doi: 10.1016/s0378-1119(02)00398-0. [DOI] [PubMed] [Google Scholar]
- 74.Mootha V.K., Lindgren C.M., Eriksson K.-F., Subramanian A., Sihag S., Lehar J., Puigserver P., Carlsson E., Ridderstråle M., Laurila E. PGC-1alpha-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat. Genet. 2003;34:267–273. doi: 10.1038/ng1180. [DOI] [PubMed] [Google Scholar]
- 75.Cantrell D.A., Smith K.A. The interleukin-2 T-cell system: A new cell growth model. Science. 1984;224:1312–1316. doi: 10.1126/science.6427923. [DOI] [PubMed] [Google Scholar]
- 76.Smith K.A. Interleukin-2: Inception, impact, and implications. Science. 1988;240:1169–1176. doi: 10.1126/science.3131876. [DOI] [PubMed] [Google Scholar]
- 77.Steinman R.M., Inaba K., Turley S., Pierre P., Mellman I. Antigen capture, processing, and presentation by dendritic cells: recent cell biological studies. Human immunol. 1999;60:562–567. doi: 10.1016/s0198-8859(99)00030-0. [DOI] [PubMed] [Google Scholar]
- 78.Entrez Gene: CD40 CD40 molecule, TNF receptor superfamily member 5. [(Accessed on 14 May 2013)]. Available online: http://www.ncbi.nlm.nih.gov/gene/958.
- 79.Tojo T., Aoki T., Kobayashi N., Ohishi T., Watanabe T., Yamamoto T., Inoue J. TRAF5, a novel tumor necrosis factor receptor-associated factor family protein, mediates CD40 signaling. Proc. Natl. Acad. Sci. USA. 1996;93:9437–9442. doi: 10.1073/pnas.93.18.9437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Cheng G., Schoenberger S.P. CD40 signaling and autoimmunity. Curr. Dir. Autoimmun. 2002;5:51–61. doi: 10.1159/000060547. [DOI] [PubMed] [Google Scholar]
- 81.Dallman C., Johnson P.W., Packham G. Differential regulation of cell survival by CD40. Apoptosis. 2003;8:45–53. doi: 10.1023/a:1021696902187. [DOI] [PubMed] [Google Scholar]
- 82.O’Sullivan B., Thomas R. Recent advances on the role of CD40 and dendritic cells in immunity and tolerance. Curr. Opin. Hematol. 2004;10:272–278. doi: 10.1097/00062752-200307000-00004. [DOI] [PubMed] [Google Scholar]
- 83.Oppmann B., Lesley R., Blom B., Timans J.C., Xu Y.M., Hunte B., Vega F., Yu N., Wang J., Singh K., et al. Novel p19 protein engages IL-12p40 to form a cytokine, IL-23, with biological activities similar as well as distinct from IL-12. Immunity. 2000;13:715–725. doi: 10.1016/s1074-7613(00)00070-4. [DOI] [PubMed] [Google Scholar]
- 84.Jover R., Bort R., Gomez-Lechon M.J., Castell J.V. Down-regulation of human CYP3A4 by the inflammatory signal interleukin 6: Molecular mechanism and transcription factors involved. FASEB. J. 2002;16:1799–1801. doi: 10.1096/fj.02-0195fje. [DOI] [PubMed] [Google Scholar]
- 85.Kile B.T., Schulman B.A., Alexander W.S., Nicola N.A., Martin H.M.E., Hiton D.J. The SOCS box: A tale of destruction and degradation. Trends Biochem. Sci. 2002;27:235–241. doi: 10.1016/s0968-0004(02)02085-6. [DOI] [PubMed] [Google Scholar]
- 86.Krebs D.L., Hilton D.J. SOCS proteins: Negative regulators of cytokine signaling. Stem Cells. 2001;19:378–387. doi: 10.1634/stemcells.19-5-378. [DOI] [PubMed] [Google Scholar]
- 87.Morrison D.F., Foss D.L., Murtaugh M.P. Interleukin-10 gene therapy-mediated amelioration of bacterial pneumonia. Infect. Immun. 2000;68:4752–4758. doi: 10.1128/iai.68.8.4752-4758.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Palma M., DeLuca D., Worgall S., Quadri L.E.N. Transcriptome analysis of the response of Pseudomonas aeruginosa to hydrogen peroxide. J. Bacteriol. 2004;186:248–252. doi: 10.1128/JB.186.1.248-252.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Series GSE42317. [(Accessed on 15 November 2012)]. Available online: http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE42317.
- 90.Hu J., He X. Enhanced quantile normalization of microarray data to reduce loss of information in gene expression profiles. Biometrics. 2007;63:50–59. doi: 10.1111/j.1541-0420.2006.00670.x. [DOI] [PubMed] [Google Scholar]
- 91.Xia X., McClelland M., Wang Y. WebArray: An online platform for microarray data analysis. BMC Bioinf. 2005;6:306. doi: 10.1186/1471-2105-6-306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Romualdi C., Vitulo N., Favero M.D., Lanfranchi G. MIDAW: A web tool for statistical analysis of microarray data. Nucleic Acids Res. 2005;33:W644–W649. doi: 10.1093/nar/gki497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Troyanskaya O., Cantor M., Sherlock G., Brown P., Hastie T., Tibshiran R., Bostein D., Altman R.B. Missing value estimation methods for DNA microarrays. Bioinformatics. 2001;17:520–525. doi: 10.1093/bioinformatics/17.6.520. [DOI] [PubMed] [Google Scholar]
- 94.Saeed A., Sharov V., White J., Li J., Liang W., Bhagabati N., Braisted J., Klapa M., Currier T., Thiagarajan M. TM4: A free, open-source system for microarray data management and analysis. Biotechniques. 2003;34:374–378. doi: 10.2144/03342mt01. [DOI] [PubMed] [Google Scholar]
- 95.Bild A., Febbo P.G. Application of a priori established gene sets to discover biologically important differential expression in microarray data. Proc. Natl. Acad. Sci. USA. 2005;102:15278–15279. doi: 10.1073/pnas.0507477102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96.Dennis G., Sherman B.T., Hosack D.A., Yang J., Gao W., Lane H.C., Lempicki R.A. DAVID: Database for annotation, visualization, and integrated discovery. Genome Biol. 2003;4:1–11. [PubMed] [Google Scholar]
- 97.Lee H.K., Braynen W., Keshav K., Pavlidis P. ErmineJ: Tool for functional analysis of gene expression data sets. BMC Bioinf. 2005;6:269. doi: 10.1186/1471-2105-6-269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Subramanian A., Tamayo P., Mootha V.K., Mukherjee S., Ebert B.L., Gilette M.A., Paulovich A., Pomeroy S.L., Golub T.R., Lander E.S. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA. 2005;102:15545–15550. doi: 10.1073/pnas.0506580102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Dopazo J. Functional interpretation of microarray experiments. OMICS. 2006;10:398–410. doi: 10.1089/omi.2006.10.398. [DOI] [PubMed] [Google Scholar]
- 100.Davis S., Meltzer P.S. GEOquery: A bridge between the Gene Expression Omnibus (GEO) and BioConductor. Bioinformatics. 2007;23:1846–1847. doi: 10.1093/bioinformatics/btm254. [DOI] [PubMed] [Google Scholar]
- 101.Barrett T., Suzek T.O., Troup D.B., Wilhite S.E., Ngau W.-C., Ledoux P., Rudnev D., Lash A.E., Fujibuchi W., Edgar R. NCBI GEO: Mining millionsof expression profiles-database and tools. Nucleic Acids Res. 2005;33:562–566. doi: 10.1093/nar/gki022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Edgar R., Domrachev M., Lash A.E. Gene expression omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 2002;30:207–210. doi: 10.1093/nar/30.1.207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 103.Livak K.J., Schmittgen T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001;25:402–408. doi: 10.1006/meth.2001.1262. [DOI] [PubMed] [Google Scholar]