Abstract
Human retinoblastoma binding protein 9 (RBBP9) is an interacting partner of the retinoblastoma susceptibility protein (Rb). RBBP9 is a tumor-associated protein required for pancreatic neoplasia, affects cell cycle control, and is involved in the TGF-β signalling pathway. Sequence analysis suggests that RBBP9 belongs to the α/β hydrolase superfamily of enzymes. The serine hydrolase activity of RBBP9 is required for development of pancreatic carcinomas in part by inhibiting TGF-β antiproliferative signaling through suppressing Smad2/3 phosphorylation. The crystal structure of human RBBP9 confirms the α/β hydrolase fold, with a six-stranded parallel β-sheet flanked by α helixes. The structure of RBBP9 resembles that of the YdeN protein from Bacillus subtilis, which is suggested to have carboxylesterase activity. RBBP9 contains a Ser75-His165-Asp138 catalytic triad, situated in a prominent pocket on the surface of the protein. The side chains of the LxCxE sequence motif that is important for interaction with Rb is mostly buried in the structure. Structure-function studies of RBBP9 suggest possible routes for novel cancer drug discovery programs.
Keywords: α/β hydrolase, pancreatic cancer, protein structure, structural genomics, RBBP9
1. Function of RBBP9
Retinoblastoma binding protein 9 (RBBP9) was first shown to be overexpressed in several transformed rat liver epithelial cell lines, resulting in the loss of TGF-β1 growth inhibition [1]. The retinoblastoma (Rb) protein is one of the most extensively studied tumor-suppressor genes [2-4]. RBBP9 contains the Rb binding motif LxCxE and has been identified to bind the Rb protein by yeast two-hybrid and co-immunoprecipitation studies [1]. Overexpression of RBBP9 overcomes TGF-β1 induced growth arrest [1]. In addition to its Rb-binding function, RBBP9 may play a role in cellular responses to chronic low dose radiation [5], and is a candidate regulator of aging in hematopoietic stem cells [6].
Recently, the Northeast Structural Genomics Consortium (NESG, www.nesg.org) determined a high-resolution 1.72 Å crystal structure of human RBBP9 [7], as a part of biomedical theme focusing on networks of interacting proteins involved in cancer biology [8]. RBBP9 is a member of Pfam family [9] DUF1234 (PF06821), a proposed α/β hydrolase family of unknown function. The crystal structure of human RBBP9 (PDB ID: 2QS9) has an α/β hydrolase fold, and reveals a serine hydrolase active site, with a Ser-His-Asp catalytic triad [7]. Recent biochemical studies with pancreatic cancer cells and site-directed mutagenesis of the putative nucleophile Ser75 to Ala have further confirmed that the protein does indeed function as a hydrolase and that Ser75 is essential for this activity [10]. In fact, the RBBP9 serine hydrolase activity is elevated in 40% of pancreatic tumor biopsies [10]. It is required to suppress Smad2/3 phosphorylation and overcome the TGF-β-mediated anti-proliferative response. Accordingly, it has been proposed that RBBP9 may function to maintain the Smad signaling pathway in the “off state”.
RBBP9 serine hydrolase activity has been detected in a range of carcinoma cell lines (lung, breast, colon, and ovary) [10], implicating RBBP9 in a wide range of cancers. RBBP9-mediated suppression of TGF-β signaling is also required for E-cadherin expression and the formation of adherens junctions and may contribute to the epithelial phenotype of pancreatic tumor cells [10]. The serine hydrolase activity may have broad-ranging implications for epithelial neoplasia.
RBBP9 may become a potentially viable cancer drug target [11]. Although the substrates remain unknown, oxime esters have been identified as selective, covalent inhibitors of RBBP9’s serine hydrolase activity [12], and the bioactive natural alkaloid emetine has been identified as a reversible inhibitor of RBBP9 [13]. A structure-based drug design approach can be developed for RBBP9 [14,15], utilizing the atomic coordinates, expression constructs, and sample preparation protocols that are freely available through the NIH Protein Structure Initiative (PSI) Knowledge Base (http://kb.psi-structuralgenomics.org/) and PSI Materials Repository (http://psimr.asu.edu/) for NESG target HR2978.
2. Protein-protein interactions of RBBP9
In addition to binding Rb, RBBP9 has been found in complexes with p107 (RBL1) and p130 (RBL2) [1], the two other Rb family members (Fig. 1) [16,17]. p107 and p130 are more similar to each other in sequence than to Rb. The Rb family negatively modulates the transition from the G1 phase to the S phase, by inactivation of transcription factors, such as those of the E2F family that promote cell entrance into the S phase [16]. The Rb family also regulates a wide spectrum of complex biological phenomena, such as differentiation, embryonic development and apoptosis [3,16,17]. RBBP9 can compete with and displace E2F1 from E2F1-Rb complexes [1].
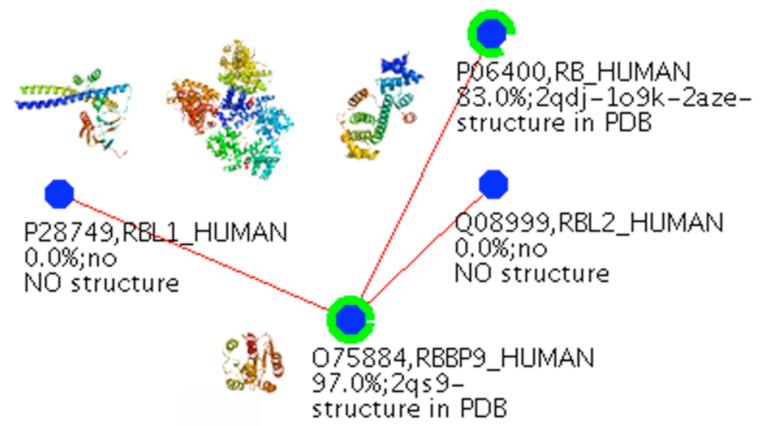
Figure 1.
RBBP9 interacts with the Rb family members, including Rb, RBL1 (p107 in the text), and RBL2 (p130 in the text). The percentage of structural coverage of these proteins are indicated by green rings around each node, and images of the corresponding domain structures are shown on the interaction network.
p107 binds a variety of cellular proteins to affect the expression of many target genes during cell cycle progression [17]. p107 can act as a Smad cofactor [18]. It forms complex with Smad3, a direct mediator of transcriptional regulation by the TGF-β receptor, and with the transcription factors E2F4/5 and DP1 [18]. This p107-E2F4/5-DP1-Smad3 complex pre-exists in the cytoplasm. In response to the TGF-β receptor, this complex moves into the nucleus, associates with Smad4, and represses c-myc expression [18].
RBBP9 is primarily restricted to the extranuclear compartment [10]. It is possible that RBBP9 regulates the formation of the p107-E2F4/5-DP1-Smad3 complex, by interacting with p107 [1]. The fact that RBBP9 can compete with and displace E2F1 from E2F1-Rb complexes in vitro [1] suggests that RBBP9 might compete with and displace E2F4/5 from the E2F4/5-p107 complex in the extranuclear compartment.
3. Structure of RBBP9
Amino-acid sequence analysis suggests that RBBP9 belongs to the DUF1234 superfamily and may have the α/β hydrolase fold. Members of the DUF1234 superfamily include acyl transferases, chlorophyllase (chlase), lipase, thioesterase, serine carboxypeptidase and others. The crystal structure of human RBBP9 confirms that it has a classic α/β hydrolase superfamily fold [7], consisting of a central six-stranded parallel β-sheet with topology order 213456 surrounded by seven α-helixes (Figs. 2A, 2B). According to ESTHER classification [19], the RBBP9 fold is more similar to the lipase 2 family, except for the insertion of an additional αD helix (aa. 106-111) after the β4 strand (Fig. 2A). The lipase 2 family currently includes 86 members, most of which are from prokaryotes or nematodes. The closest sequence homolog with three-dimensional structural information is that of the YdeN protein from Bacillus subtilis [20], which shares 26% sequence identity with RBBP9. YdeN was also determined as part of a Protein Structure Initiative (PSI) structural genomics project [20]. RBBP9 is more similar to prokaryotic proteins structurally than to human α/β hydrolases – the closest human proteins are valacyclovirase (PDB ID: 2OCL) [21] and monoglyceride lipase (PDB ID: 3HJU) [22], sharing around 15% sequence identity.
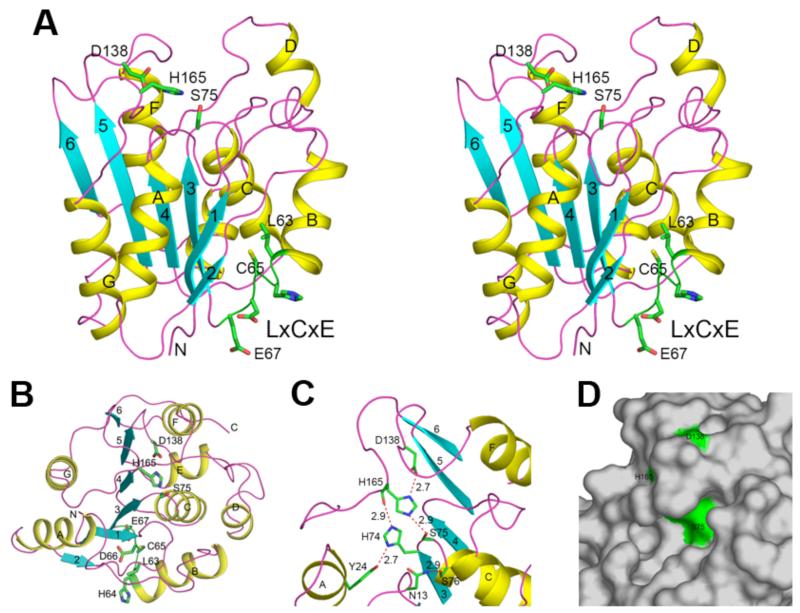
Figure 2.
Structure of human RBBP9. (A). Ribbon representation in stereo of the RBBP9 structure. The secondary structure is depicted as yellow (α-helixes), cyan (β-strands) and magenta (loops). The catalytic triad and the LxCxE residues are shown in green and labeled. (B). The structure of RBBP9, viewed down the active site and the central β-sheet. The viewing direction is from the top of panel A. (C). The active site of RBBP9. The catalytic triad and several additional residues are shown. Hydrogen-bonding interactions are indicated with the dashed lines in red, with the hydrogen-bonding distances indicated. (D). Molecular surface of RBBP9 is shown in gray, and the catalytic triad Ser-His-Asp is shown in green. The catalytic nucleophile Ser75 is located in a prominent pocket in the surface.
The active site of RBBP9 is located at the top of the β-sheet, with the catalytic triad Ser75-His165-Asp138 (Fig. 2B). As in other α/β hydrolases, Ser75 is located in a nucleophile elbow [23], a tight turn connecting strand β3 and helix αC (Fig. 2C), and assumes a strained main-chain conformation. The side chains of these three residues are hydrogen-bonded to each other in the structure (Fig. 2C) and Ser75 is located in a prominent groove on the surface of the protein (Fig. 2D). Typical Gly-flanked environment of the nucleophilic Ser75 (Gly-X1-Ser-X2-Gly) enables the tight turn at the tip of the elbow to bring Ser75 close to His165 (2.9 Å). Interestingly, in YdeN the first Gly of this sequence motif is replaced with an Ala, which was made possible by a slight opening of the elbow [20]. The X1 position is occupied by a highly conserved His74, forming a hydrogen bond with Tyr24 (2.7 Å) and the main chain carbonyl of His165 (2.9 Å), which also helps to stabilize the nucleophile elbow (Fig. 2C). Tyr24 is also conserved among eukaryotes, but is replaced by Phe in prokaryotes. In RBBP9, there is additional stabilization of the nucleophile elbow through Ser76 (at the X2 position) hydrogen-bonding to Asn13 (2.9 Å). This hydrogen bond is absent in YdeN and other prokaryotic homologs (Ser76 replaced by Leu and Asn13 by His).
Another important feature of α/β hydrolases is the presence of hydrogen-bond donors arranged in an oxyanion hole close to the active site. Based on close proximity to Ser75 and superposition with the carboxylesterase from Pseudomonas fluorescens [24], the main-chain amides of Asn13 and Ser76 may form the oxyanion hole in RBBP9 (Fig. 2C). These amino acids correspond to Leu23 and Gln115 in carboxylesterase and Tyr11 and Leu72 in YdeN [20,24].
A structural difference between lipase 2 family and RBBP9 is that the active site of the lipases is usually buried under the C-terminus α-helix (flap or lid). The flap conceals the active site of the lipases and has to be rearranged to make it accessible. In RBBP9 and YdeN, the much smaller αD helix could play a similar role. In the RBBP9 structure, the αD helix is located relatively far form the active site, but the presence of two long flanking linkers could allow the αD helix to close the active site (Fig. 2B). In comparison, both human valacyclovirase and monoglyceride lipase have very well formed flaps.
RBBP9 contains the retinoblastoma (Rb) binding motif LxCxE in its sequence [25], and mutation of the Leu residue in this motif to Gln blocks the binding to Rb [1]. This LxCxE motif is not present in YdeN and other α/β hydrolases. The Leu residue is located at the end of helix αB, and the other residues are in the loop connecting helix αB and strand β3, near the bottom of the central β-sheet and far from the active site of RBBP9 (Fig. 2A). Somewhat surprisingly, the side chains of both the Leu and the Cys residues are buried in the hydrophobic core of the structure and are essentially not accessible to solvent. Only the side chain of the Glu residue is exposed (Fig. 2A). This suggests that a conformational change is needed for this motif to directly mediate interactions with Rb, although such a change may disrupt the hydrophobic core of RBBP9. Alternatively, this binding may involve a different surface area of RBBP9, with the LxCxE motif playing an indirect role in the recognition process.
4. Conclusions
Structural and functional studies have identified RBBP9 as an enzyme, in addition to its interactions with Rb. RBBP9 is a serine hydrolase, belonging to the α/β hydrolase superfamily of enzymes. The catalytic triad, Ser75-His165-Asp138, is situated in a prominent pocket on the surface of the protein. This serine hydrolase activity has been detected in a variety of tumor cell lines. Especially, RBBP9 catalytic activity was found to be elevated in pancreatic tumors, suggesting that RBBP9 may be a possible anti-cancer target.
Many questions remain about the exact mechanistic role of RBBP9. The native substrates of RBBP9 are not yet identified, and the consequences of substrate hydrolysis are unknown. Do RBBP9 Rb-binding and serine hydrolase activity act independently or cooperatively? How does substrate processing lead to regulation of Smad2/3 signaling? Further studies are needed to provide answers to these important questions.
Acknowledgements
We thank Prof. F. Liu for helpful comments on the manuscript. This research was supported by a grant from the Protein Structure Initiative of the National Institutes of Health (U54 GM074958). Data on the expression, purification, and characterization of RBBP9 (NESG ID HR2978) are available online at http://spine.nesg.org/target.cgi?id=HR2978.
References
- 1.Woitach JT, Zhang M, Niu C-H, Thorgeirsson SS. A retinoblastoma-binding protein that affects cell-cycle control and confers transforming ability. Nat. Genetics. 1998;19:371–374. doi: 10.1038/1258. [DOI] [PubMed] [Google Scholar]
- 2.Weinberg RA. Tumore suppressor genes. Science. 1991;254:1138–1146. doi: 10.1126/science.1659741. [DOI] [PubMed] [Google Scholar]
- 3.Burkhart DL, Sage J. Cellular mechanisms of tumour suppression by the retinoblastoma gene. Nat. Rev. Cancer. 2008;8:671–682. doi: 10.1038/nrc2399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Morris EJ, Dyson NJ. Retinoblastoma protein partners. Adv. Cancer Res. 2001;82:1–54. doi: 10.1016/s0065-230x(01)82001-7. [DOI] [PubMed] [Google Scholar]
- 5.Cassie S, Koturbash I, Hudson D, Baker M, Ilnytskyy Y, Rodriguez-Juarez R, Weber E, Kovalchuk O. Novel retinoblastoma binding protein RBBP9 modulates sex-specific radiation responses in vivo. Carcinogenesis. 2006;27:465–474. doi: 10.1093/carcin/bgi261. [DOI] [PubMed] [Google Scholar]
- 6.Geiger H, Rennebeck G, van Zant G. Regulation of hematopoietic stem cell aging in vivo by a distinct genetic element. Proc. Natl. Acad. Sci. USA. 2005;102:5102–5107. doi: 10.1073/pnas.0408654102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Vorobiev SM, Su M, Seetharaman J, Huang YJ, Chen CX, Maglaqui M, Janjua H, Proudfoot M, Yakunin A, Xiao R, Acton TB, Montelione GT, Tong L. Crystal structure of human retinoblastoma binding protein 9 (RBBP9) Proteins. 2009;74:526–529. doi: 10.1002/prot.22278. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Huang YJ, Hang D, Lu LJ, Tong L, Gerstein MB, Montelione GT. Targeting the human cancer pathway protein interaction network by structural genomics. Mol. Cell. Proteomics. 2008 doi: 10.1074/mcp.M700550-MCP200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Finn RD, Mistry J, Tate J, Coggill P, Heger A, Pollington JE, Gavin OL, Gunesekaran P, Ceric G, Forslund K, Holm L, Sonnhammer EL, Eddy S, Bateman A. The Pfam protein families database. Nucl. Acid Res. 2010;38:D211–D222. doi: 10.1093/nar/gkp985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Shields DJ, Niessen S, Murphy EA, Mielgo A, Desgrosellier JS, Lau SK, Barnes LA, Lesperance J, Bouvet M, Tarin D, Cravatt BF, Cheresh DA. RBBP9: a tumor-associated serine hydrolase activity required for pancreatic neoplasia. Proc. Natl. Acad. Sci. USA. 2010;107:2189–2194. doi: 10.1073/pnas.0911646107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Bogyo M. WFinding enzymes that are actively involved in cancer. Proc. Natl. Acad. Sci. USA. 2010;107:2379–2380. doi: 10.1073/pnas.0914955107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Bachovchin DA, Wolfe MR, Masuda K, Brown SJ, Spicer TP, Fernandez-Vega V, Chase P, Hodder PS, Rosen H, Cravatt BF. Oxime esters as selective, covalent inhibitors of the serine hydrolase retinoblastoma-binding protein 9 (RBBP9) Bioorg. Med. Chem. Lett. 2010;20:2254–2258. doi: 10.1016/j.bmcl.2010.02.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Bachovchin DA, Brown SJ, Rosen H, Cravatt BF. Identification of selective inhibitors of uncharacterized enzymes by high-throughput screening with fluorescent activity-based probes. Nat. Biotech. 2009;27:387–394. doi: 10.1038/nbt.1531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Weigelt J, McBroom-Cerajewski LD, Schapira M, Zhao Y, Arrowsmith CH. Structural genomics and drug discovery: all in the family. Curr. Opin. Chem. Biol. 2008;12:32–39. doi: 10.1016/j.cbpa.2008.01.045. [DOI] [PubMed] [Google Scholar]
- 15.van Montfort RL, Workman P. Structure-based design of molecular cancer therapeutics. Trends Biotechnol. 2009;27:315–328. doi: 10.1016/j.tibtech.2009.02.003. [DOI] [PubMed] [Google Scholar]
- 16.Paggi MG, Giordano A. Who is the boss in the retinoblastoma family? The point of view of Rb2/p130, the little brother. Cancer Res. 2001;61:4651–4654. [PubMed] [Google Scholar]
- 17.Wirt SE, Sage J. p107 in the public eye: an Rb understudy and more. Cell Div. 2010;5 doi: 10.1186/1747-1028-5-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Chen CR, Kang Y, Siegel PM, Massague J. E2F4/5 and p107 as Smad cofactors linking the TGFbeta receptor to c-myc repression. Cell. 2002;110:19–32. doi: 10.1016/s0092-8674(02)00801-2. [DOI] [PubMed] [Google Scholar]
- 19.Carr PD, Ollis DL. α/β hydrolase fold: an update. Prot. Pept. Lett. 2009;16:1137–1148. doi: 10.2174/092986609789071298. [DOI] [PubMed] [Google Scholar]
- 20.Janda I, Devedjiev Y, Cooper D, Chruszcz M, Derewenda U, Gabrys A, Minor W, Joachimiak A, Derewenda ZS. Harvesting the high-handing fruit: the structure of the YdeN gene product from Bacillus subtilis at 1.8 Å resolution. Acta Cryst. 2004;D60:1101–1107. doi: 10.1107/S0907444904007188. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lai L, Xu Z, Zhou J, Lee KD, Amido GL. Molecular basis of prodrug activation by human valacyclovirase, an alpha-amino acid ester hydrolase. J. Biol. Chem. 2008;283:9318–9327. doi: 10.1074/jbc.M709530200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Labar G, Bauvois C, Borel F, Ferrer JL, Wouters J, Lambert DM. Crystal structure of the human monoacylglycero lipase, a key actor in endocannabinoid signaling. Chembiochem. 2010;11:218–227. doi: 10.1002/cbic.200900621. [DOI] [PubMed] [Google Scholar]
- 23.Ollis DL, Cheah E, Cygler M, Dijkstra B, Frolow F, Franken SM, Harel M, Remington SJ, Silman I, Schrag J, Sussman JL, Verschueren KHG, Goldman A. The α/β hydrolase fold. Prot. Eng. 1992;5:197–211. doi: 10.1093/protein/5.3.197. [DOI] [PubMed] [Google Scholar]
- 24.Kim KK, Song HK, Shin DH, Hwang KY, Choe S, Yoo OJ, Suh SW. Crystal structure of carboxylesterase from Pseudomonas fluorescens, an alpha/beta hydrolase with broad substrate specificity. Structure. 1997;5:1571–1584. doi: 10.1016/s0969-2126(97)00306-7. [DOI] [PubMed] [Google Scholar]
- 25.Singh M, Krajewski M, Mikolajka A, Holak TA. Molecular determinants for the complex formation between the retinoblastoma protein and LXCXE sequences. J. Biol. Chem. 2005;280:37868–37876. doi: 10.1074/jbc.M504877200. [DOI] [PubMed] [Google Scholar]