Abstract
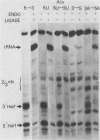
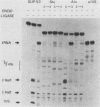
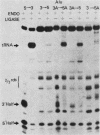
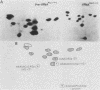
The Saccharomyces cerevisiae amber suppressor tRNA gene SUP53 (a tRNALeu3 allele) was used to investigate the role of intron structure and sequence on precursor tRNA splicing in vivo and in vitro. This gene encodes a pre-tRNA which contains a 32-base intervening sequence. Two types of SUP53 intron mutants were constructed: ones with an internal deletion of the natural SUP53 intron and ones with a novel intron. These mutant genes were transcribed in vitro, and the end-processed transcripts were analyzed for their ability to serve as substrates for the partially purified S. cerevisiae tRNA endonuclease and ligase. The in vitro phenotype of these mutant RNAs was correlated with the in vivo suppressor tRNA function of these SUP53 alleles after integration of the genes into the yeast genome. Analysis of these mutant pre-tRNAs, which exhibited no perturbation of the mature domain, clearly showed that intron structure and sequence can have profound effects on pre-tRNA splicing. All of the mutant RNAs, which were inefficiently spliced or unspliced, evidenced cleavage only at the 5' splice junction. Base changes in the intron proximal to the 3' splice junction could partially rescue the splicing defect. The implications of these data for tRNA endonuclease-substrate interactions are discussed.
Full text
PDF
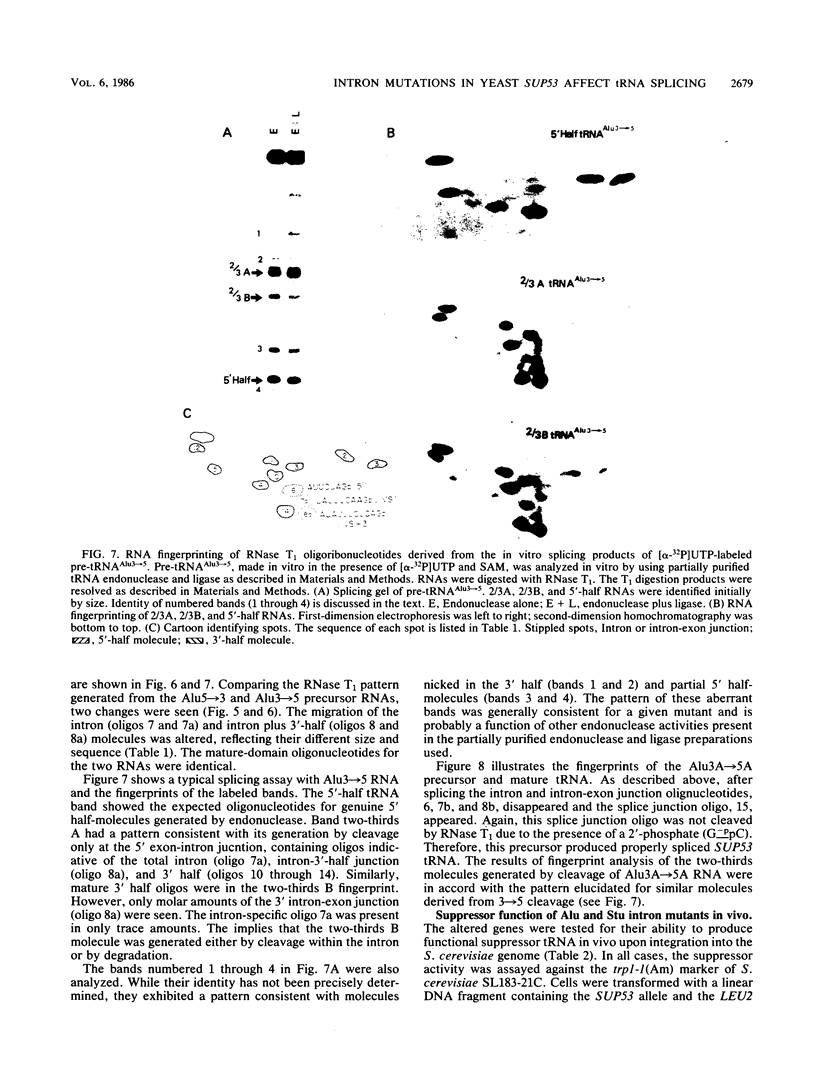

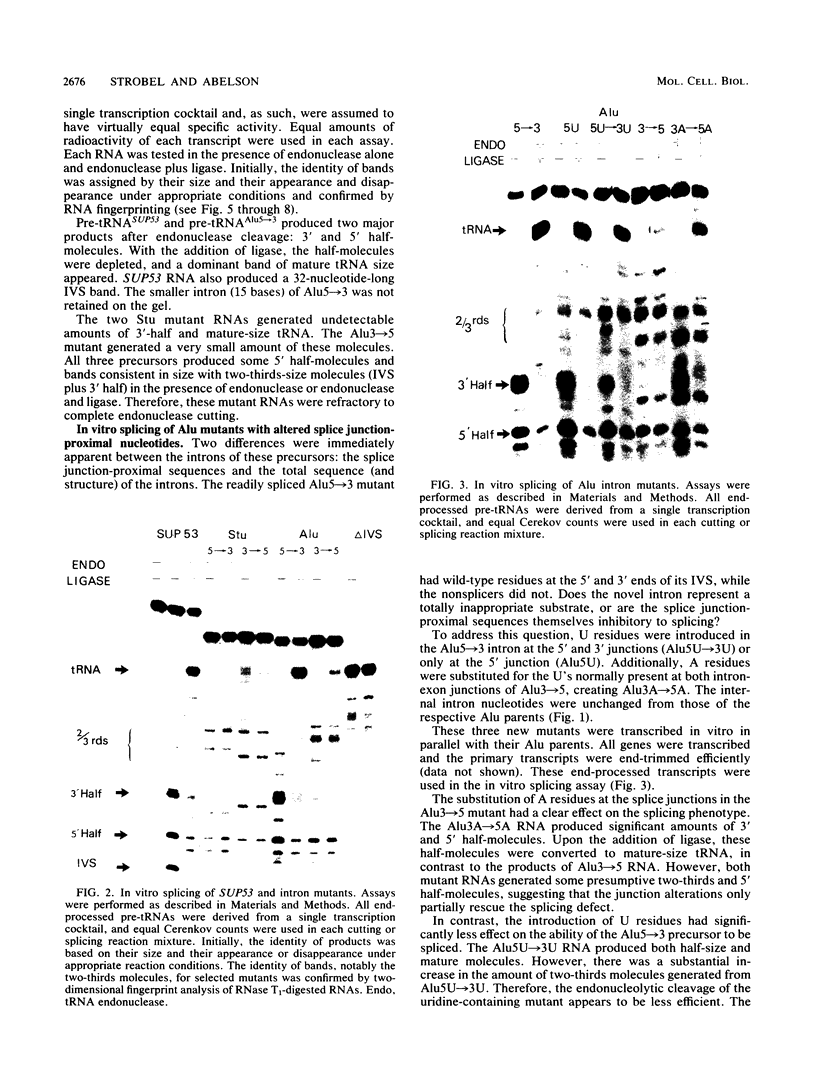




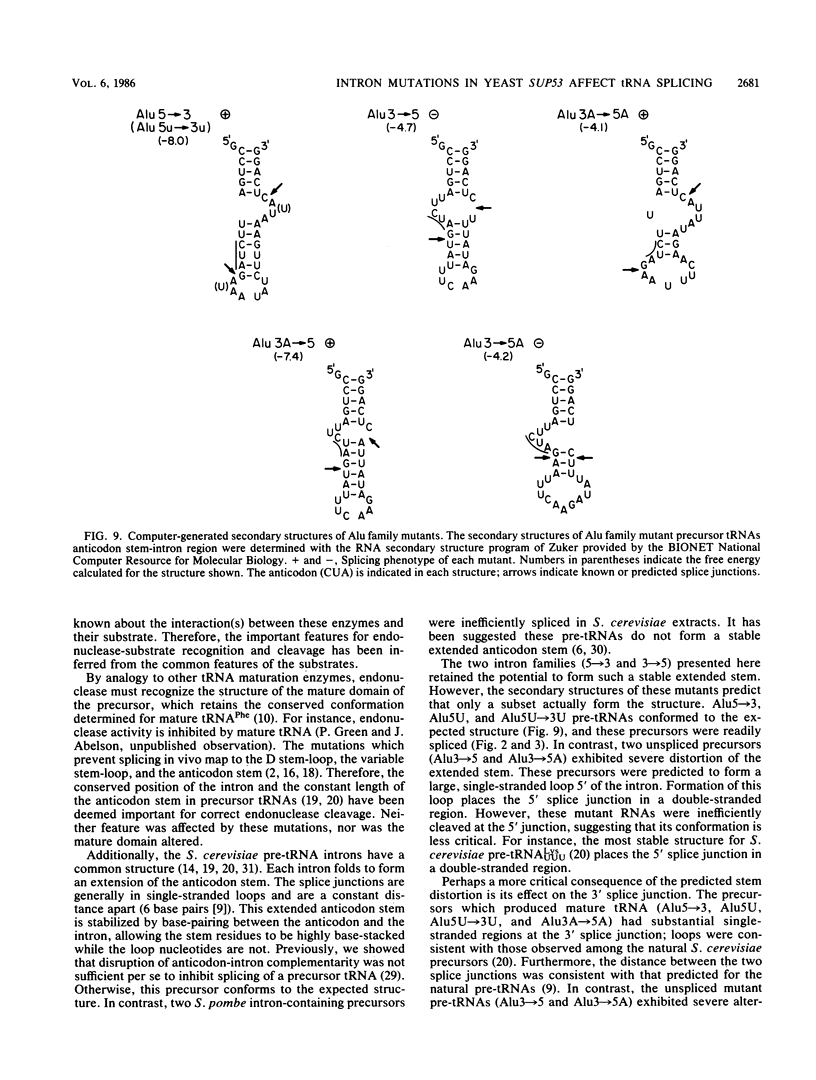


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Andreadis A., Hsu Y. P., Kohlhaw G. B., Schimmel P. Nucleotide sequence of yeast LEU2 shows 5'-noncoding region has sequences cognate to leucine. Cell. 1982 Dec;31(2 Pt 1):319–325. doi: 10.1016/0092-8674(82)90125-8. [DOI] [PubMed] [Google Scholar]
- Baldi M. I., Mattoccia E., Tocchini-Valentini G. P. Role of RNA structure in splicing: excision of the intervening sequence in yeast tRNA3leu is dependent on the formation of a D stem. Cell. 1983 Nov;35(1):109–115. doi: 10.1016/0092-8674(83)90213-1. [DOI] [PubMed] [Google Scholar]
- Colby D., Leboy P. S., Guthrie C. Yeast tRNA precursor mutated at a splice junction is correctly processed in vivo. Proc Natl Acad Sci U S A. 1981 Jan;78(1):415–419. doi: 10.1073/pnas.78.1.415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Engelke D. R., Gegenheimer P., Abelson J. Nucleolytic processing of a tRNAArg-tRNAAsp dimeric precursor by a homologous component from Saccharomyces cerevisiae. J Biol Chem. 1985 Jan 25;260(2):1271–1279. [PubMed] [Google Scholar]
- Fischhoff D. A., Waterston R. H., Olson M. V. The yeast cloning vector YEp13 contains a tRNALeu3 gene that can mutate to an amber suppressor. Gene. 1984 Mar;27(3):239–251. doi: 10.1016/0378-1119(84)90069-6. [DOI] [PubMed] [Google Scholar]
- Gamulin V., Mao J., Appel B., Sumner-Smith M., Yamao F., Söll D. Six Schizosaccharomyces pombe tRNA genes including a gene for a tRNALys with an intervening sequence which cannot base-pair with the anticodon. Nucleic Acids Res. 1983 Dec 20;11(24):8537–8546. doi: 10.1093/nar/11.24.8537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Greer C. L., Peebles C. L., Gegenheimer P., Abelson J. Mechanism of action of a yeast RNA ligase in tRNA splicing. Cell. 1983 Feb;32(2):537–546. doi: 10.1016/0092-8674(83)90473-7. [DOI] [PubMed] [Google Scholar]
- Johnson J. D., Ogden R., Johnson P., Abelson J., Dembeck P., Itakura K. Transcription and processing of a yeast tRNA gene containing a modified intervening sequence. Proc Natl Acad Sci U S A. 1980 May;77(5):2564–2568. doi: 10.1073/pnas.77.5.2564. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koski R. A., Allison D. S., Worthington M., Hall B. D. An in vitro RNA polymerase III system from S. cerevisiae: effects of deletions and point mutations upon SUP4 gene transcription. Nucleic Acids Res. 1982 Dec 20;10(24):8127–8143. doi: 10.1093/nar/10.24.8127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koski R. A., Clarkson S. G., Kurjan J., Hall B. D., Smith M. Mutations of the yeast SUP4 tRNATyr locus: transcription of the mutant genes in vitro. Cell. 1980 Nov;22(2 Pt 2):415–425. doi: 10.1016/0092-8674(80)90352-9. [DOI] [PubMed] [Google Scholar]
- Kurjan J., Hall B. D., Gillam S., Smith M. Mutations at the yeast SUP4 tRNATyr locus: DNA sequence changes in mutants lacking suppressor activity. Cell. 1980 Jul;20(3):701–709. doi: 10.1016/0092-8674(80)90316-5. [DOI] [PubMed] [Google Scholar]
- Lee M. C., Knapp G. Transfer RNA splicing in Saccharomyces cerevisiae. Secondary and tertiary structures of the substrates. J Biol Chem. 1985 Mar 10;260(5):3108–3115. [PubMed] [Google Scholar]
- Liebman S. W., Srodulski Z., Reed C. R., Stewart J. W., Sherman F., Brennan G. Yeast amber suppressors corresponding to tRNA3Leu genes. J Mol Biol. 1984 Sep 15;178(2):209–226. doi: 10.1016/0022-2836(84)90140-2. [DOI] [PubMed] [Google Scholar]
- Mattoccia E., Baldi M. I., Pande G., Ogden R., Tocchini-Valentini G. P. Mutation in the a block of the yeast tRNAleu3 gene that allows transcription but abolishes splicing and 5'-end maturation. Cell. 1983 Jan;32(1):67–76. doi: 10.1016/0092-8674(83)90497-x. [DOI] [PubMed] [Google Scholar]
- Newman A. J., Ogden R. C., Abelson J. tRNA gene transcription in yeast: effects of specified base substitutions in the intragenic promoter. Cell. 1983 Nov;35(1):117–125. doi: 10.1016/0092-8674(83)90214-3. [DOI] [PubMed] [Google Scholar]
- Nishikura K., Kurjan J., Hall B. D., De Robertis E. M. Genetic analysis of the processing of a spliced tRNA. EMBO J. 1982;1(2):263–268. doi: 10.1002/j.1460-2075.1982.tb01157.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ogden R. C., Lee M. C., Knapp G. Transfer RNA splicing in Saccharomyces cerevisiae: defining the substrates. Nucleic Acids Res. 1984 Dec 21;12(24):9367–9382. doi: 10.1093/nar/12.24.9367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pearson D., Willis I., Hottinger H., Bell J., Kumar A., Leupold U., Söll D. Mutations preventing expression of sup3 tRNASer nonsense suppressors of Schizosaccharomyces pombe. Mol Cell Biol. 1985 Apr;5(4):808–815. doi: 10.1128/mcb.5.4.808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peebles C. L., Gegenheimer P., Abelson J. Precise excision of intervening sequences from precursor tRNAs by a membrane-associated yeast endonuclease. Cell. 1983 Feb;32(2):525–536. doi: 10.1016/0092-8674(83)90472-5. [DOI] [PubMed] [Google Scholar]
- Peebles C. L., Ogden R. C., Knapp G., Abelson J. Splicing of yeast tRNA precursors: a two-stage reaction. Cell. 1979 Sep;18(1):27–35. doi: 10.1016/0092-8674(79)90350-7. [DOI] [PubMed] [Google Scholar]
- Phizicky E. M., Schwartz R. C., Abelson J. Saccharomyces cerevisiae tRNA ligase. Purification of the protein and isolation of the structural gene. J Biol Chem. 1986 Feb 25;261(6):2978–2986. [PubMed] [Google Scholar]
- Quigley G. J., Teeter M. M., Rich A. Structural analysis of spermine and magnesium ion binding to yeast phenylalanine transfer RNA. Proc Natl Acad Sci U S A. 1978 Jan;75(1):64–68. doi: 10.1073/pnas.75.1.64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raymond G. J., Johnson J. D. The role of non-coding DNA sequences in transcription and processing of a yeast tRNA. Nucleic Acids Res. 1983 Sep 10;11(17):5969–5988. doi: 10.1093/nar/11.17.5969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothstein R. J. One-step gene disruption in yeast. Methods Enzymol. 1983;101:202–211. doi: 10.1016/0076-6879(83)01015-0. [DOI] [PubMed] [Google Scholar]
- Shaw K. J., Olson M. V. Effects of altered 5'-flanking sequences on the in vivo expression of a Saccharomyces cerevisiae tRNATyr gene. Mol Cell Biol. 1984 Apr;4(4):657–665. doi: 10.1128/mcb.4.4.657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Southern E. M. An improved method for transferring nucleotides from electrophoresis strips to thin layers of ion-exchange cellulose. Anal Biochem. 1974 Nov;62(1):317–318. doi: 10.1016/0003-2697(74)90395-9. [DOI] [PubMed] [Google Scholar]
- Strobel M. C., Abelson J. Effect of intron mutations on processing and function of Saccharomyces cerevisiae SUP53 tRNA in vitro and in vivo. Mol Cell Biol. 1986 Jul;6(7):2663–2673. doi: 10.1128/mcb.6.7.2663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sumner-Smith M., Hottinger H., Willis I., Koch T. L., Arentzen R., Söll D. The sup8 tRNALeu gene of Schizosaccharomyces pombe has an unusual intervening sequence and reduced pairing in the anticodon stem. Mol Gen Genet. 1984;197(3):447–452. doi: 10.1007/BF00329941. [DOI] [PubMed] [Google Scholar]
- Swerdlow H., Guthrie C. Structure of intron-containing tRNA precursors. Analysis of solution conformation using chemical and enzymatic probes. J Biol Chem. 1984 Apr 25;259(8):5197–5207. [PubMed] [Google Scholar]
- Volckaert G., Fiers W. A micromethod for base analysis of 32P-labeled oligoribonulcleotides. Anal Biochem. 1977 Nov;83(1):222–227. doi: 10.1016/0003-2697(77)90530-9. [DOI] [PubMed] [Google Scholar]
- Volckaert G., Fiers W. Micro thin-layer techniques for rapid sequence analysis of 32P-labeled RNA: double digestion and pancreatic ribonuclease analyses. Anal Biochem. 1977 Nov;83(1):228–239. doi: 10.1016/0003-2697(77)90531-0. [DOI] [PubMed] [Google Scholar]
- Volckaert G., Jou W. M., Fiers W. Analysis of 32P-labeled bacteriophage MS2 RNA by a mini-fingerprinting procedure. Anal Biochem. 1976 May 7;72:433–446. doi: 10.1016/0003-2697(76)90551-0. [DOI] [PubMed] [Google Scholar]
- Willis I., Hottinger H., Pearson D., Chisholm V., Leupold U., Söll D. Mutations affecting excision of the intron from a eukaryotic dimeric tRNA precursor. EMBO J. 1984 Jul;3(7):1573–1580. doi: 10.1002/j.1460-2075.1984.tb02013.x. [DOI] [PMC free article] [PubMed] [Google Scholar]