Abstract
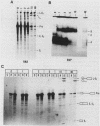
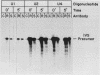
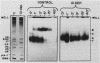
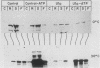
U1 small nuclear ribonucleoproteins (snRNPs) are required for in vitro splicing of pre-mRNA. Sequences within U1 RNA hybridize to, and thus recognize, 5' splice junctions. We have investigated the mechanism of association of U1 snRNPs with the spliceosome. U1-specific antibodies detected U1 association with precursor RNA early during assembly. Removal of the 5' terminal sequences of U1 RNA by oligo-directed cleavage or removal of U1 snRNPs by immunoprecipitation prior to the addition of precursor RNA depressed the association of all snRNPs with precursor RNA as detected by immunoprecipitation of splicing complexes by either Sm or U1-specific antibodies. Assembly of the spliceosome as monitored by gel electrophoresis was also depressed after cleavage of U1 RNA. The dependency of Sm precipitability of precursor RNA upon the presence of U1 snRNPs suggests that U1 snRNPs participate in the early recognition of substrate RNAs by U2 to U6 snRNPs. Although removal of the 5'-terminal sequences of U1 depressed U1 snRNP association with precursor RNA, it did not eliminate it, suggesting semistable association of U1 snRNPs with the assembling spliceosome in the absence of U1 RNA hybridization. This association was not dependent upon 5' splice junction sequences but was dependent upon 3' intronic sequences, indicating that U1 snRNPs interact with factors recognizing 3' intronic sequences. Mutual dependence of 5' and 3' recognition factors suggests significant snRNP-snRNP communication during early assembly.
Full text
PDF
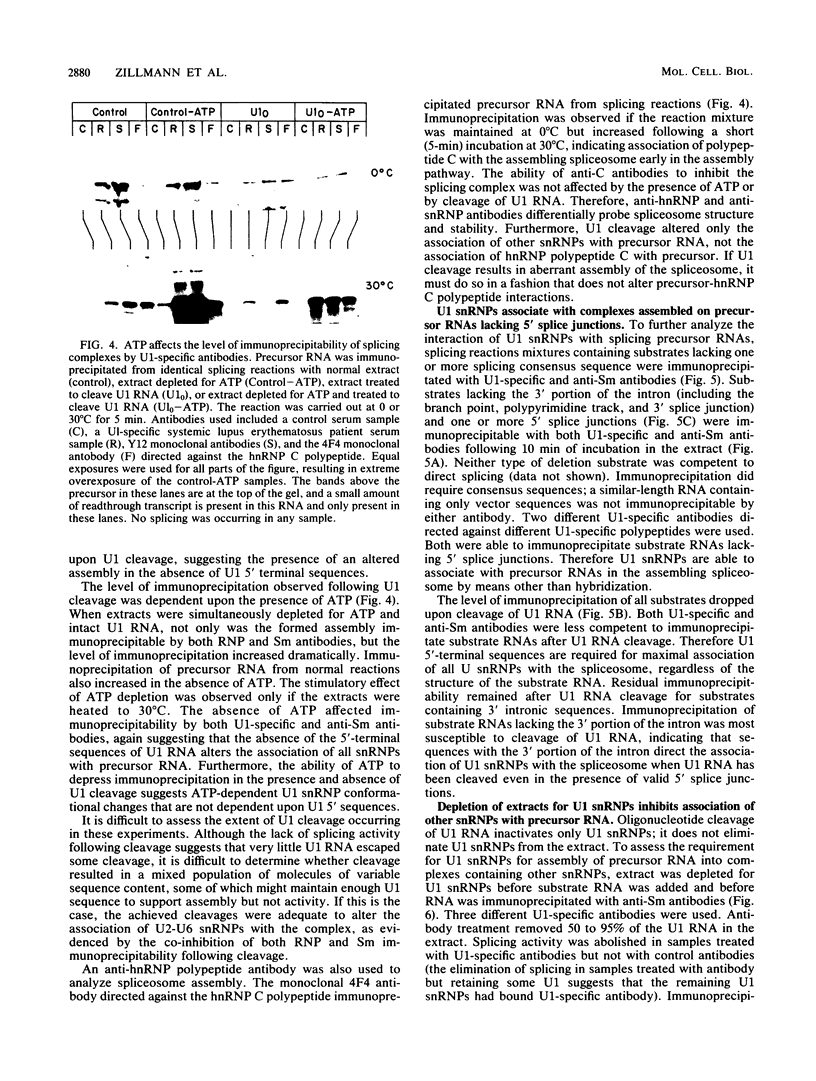




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Berget S. M., Robberson B. L. U1, U2, and U4/U6 small nuclear ribonucleoproteins are required for in vitro splicing but not polyadenylation. Cell. 1986 Aug 29;46(5):691–696. doi: 10.1016/0092-8674(86)90344-2. [DOI] [PubMed] [Google Scholar]
- Bindereif A., Green M. R. Ribonucleoprotein complex formation during pre-mRNA splicing in vitro. Mol Cell Biol. 1986 Jul;6(7):2582–2592. doi: 10.1128/mcb.6.7.2582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Black D. L., Chabot B., Steitz J. A. U2 as well as U1 small nuclear ribonucleoproteins are involved in premessenger RNA splicing. Cell. 1985 Oct;42(3):737–750. doi: 10.1016/0092-8674(85)90270-3. [DOI] [PubMed] [Google Scholar]
- Black D. L., Steitz J. A. Pre-mRNA splicing in vitro requires intact U4/U6 small nuclear ribonucleoprotein. Cell. 1986 Aug 29;46(5):697–704. doi: 10.1016/0092-8674(86)90345-4. [DOI] [PubMed] [Google Scholar]
- Brody E., Abelson J. The "spliceosome": yeast pre-messenger RNA associates with a 40S complex in a splicing-dependent reaction. Science. 1985 May 24;228(4702):963–967. doi: 10.1126/science.3890181. [DOI] [PubMed] [Google Scholar]
- Chabot B., Black D. L., LeMaster D. M., Steitz J. A. The 3' splice site of pre-messenger RNA is recognized by a small nuclear ribonucleoprotein. Science. 1985 Dec 20;230(4732):1344–1349. doi: 10.1126/science.2933810. [DOI] [PubMed] [Google Scholar]
- Chabot B., Steitz J. A. Multiple interactions between the splicing substrate and small nuclear ribonucleoproteins in spliceosomes. Mol Cell Biol. 1987 Jan;7(1):281–293. doi: 10.1128/mcb.7.1.281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choi Y. D., Grabowski P. J., Sharp P. A., Dreyfuss G. Heterogeneous nuclear ribonucleoproteins: role in RNA splicing. Science. 1986 Mar 28;231(4745):1534–1539. doi: 10.1126/science.3952495. [DOI] [PubMed] [Google Scholar]
- Frendewey D., Keller W. Stepwise assembly of a pre-mRNA splicing complex requires U-snRNPs and specific intron sequences. Cell. 1985 Aug;42(1):355–367. doi: 10.1016/s0092-8674(85)80131-8. [DOI] [PubMed] [Google Scholar]
- Gerke V., Steitz J. A. A protein associated with small nuclear ribonucleoprotein particles recognizes the 3' splice site of premessenger RNA. Cell. 1986 Dec 26;47(6):973–984. doi: 10.1016/0092-8674(86)90812-3. [DOI] [PubMed] [Google Scholar]
- Grabowski P. J., Seiler S. R., Sharp P. A. A multicomponent complex is involved in the splicing of messenger RNA precursors. Cell. 1985 Aug;42(1):345–353. doi: 10.1016/s0092-8674(85)80130-6. [DOI] [PubMed] [Google Scholar]
- Grabowski P. J., Sharp P. A. Affinity chromatography of splicing complexes: U2, U5, and U4 + U6 small nuclear ribonucleoprotein particles in the spliceosome. Science. 1986 Sep 19;233(4770):1294–1299. doi: 10.1126/science.3638792. [DOI] [PubMed] [Google Scholar]
- Konarska M. M., Sharp P. A. Electrophoretic separation of complexes involved in the splicing of precursors to mRNAs. Cell. 1986 Sep 12;46(6):845–855. doi: 10.1016/0092-8674(86)90066-8. [DOI] [PubMed] [Google Scholar]
- Krainer A. R., Maniatis T. Multiple factors including the small nuclear ribonucleoproteins U1 and U2 are necessary for pre-mRNA splicing in vitro. Cell. 1985 Oct;42(3):725–736. doi: 10.1016/0092-8674(85)90269-7. [DOI] [PubMed] [Google Scholar]
- Krämer A., Keller W., Appel B., Lührmann R. The 5' terminus of the RNA moiety of U1 small nuclear ribonucleoprotein particles is required for the splicing of messenger RNA precursors. Cell. 1984 Aug;38(1):299–307. doi: 10.1016/0092-8674(84)90551-8. [DOI] [PubMed] [Google Scholar]
- Mount S. M., Pettersson I., Hinterberger M., Karmas A., Steitz J. A. The U1 small nuclear RNA-protein complex selectively binds a 5' splice site in vitro. Cell. 1983 Jun;33(2):509–518. doi: 10.1016/0092-8674(83)90432-4. [DOI] [PubMed] [Google Scholar]
- Noble J. C., Prives C., Manley J. L. In vitro splicing of simian virus 40 early pre mRNA. Nucleic Acids Res. 1986 Feb 11;14(3):1219–1235. doi: 10.1093/nar/14.3.1219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Padgett R. A., Konarska M. M., Grabowski P. J., Hardy S. F., Sharp P. A. Lariat RNA's as intermediates and products in the splicing of messenger RNA precursors. Science. 1984 Aug 31;225(4665):898–903. doi: 10.1126/science.6206566. [DOI] [PubMed] [Google Scholar]
- Perkins K. K., Furneaux H. M., Hurwitz J. RNA splicing products formed with isolated fractions from HeLa cells are associated with fast-sedimenting complexes. Proc Natl Acad Sci U S A. 1986 Feb;83(4):887–891. doi: 10.1073/pnas.83.4.887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pikielny C. W., Rosbash M. Specific small nuclear RNAs are associated with yeast spliceosomes. Cell. 1986 Jun 20;45(6):869–877. doi: 10.1016/0092-8674(86)90561-1. [DOI] [PubMed] [Google Scholar]
- Ruskin B., Green M. R. Specific and stable intron-factor interactions are established early during in vitro pre-mRNA splicing. Cell. 1985 Nov;43(1):131–142. doi: 10.1016/0092-8674(85)90018-2. [DOI] [PubMed] [Google Scholar]
- Sierakowska H., Szer W., Furdon P. J., Kole R. Antibodies to hnRNP core proteins inhibit in vitro splicing of human beta-globin pre-mRNA. Nucleic Acids Res. 1986 Jul 11;14(13):5241–5254. doi: 10.1093/nar/14.13.5241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tazi J., Alibert C., Temsamani J., Reveillaud I., Cathala G., Brunel C., Jeanteur P. A protein that specifically recognizes the 3' splice site of mammalian pre-mRNA introns is associated with a small nuclear ribonucleoprotein. Cell. 1986 Dec 5;47(5):755–766. doi: 10.1016/0092-8674(86)90518-0. [DOI] [PubMed] [Google Scholar]
- Zhuang Y., Weiner A. M. A compensatory base change in U1 snRNA suppresses a 5' splice site mutation. Cell. 1986 Sep 12;46(6):827–835. doi: 10.1016/0092-8674(86)90064-4. [DOI] [PubMed] [Google Scholar]