Abstract
Background
Tissue inhibitors of metalloproteases (TIMPs) are a multifunctional family of proteins that orchestrate extracellular matrix turnover, tissue remodelling and other cellular processes. In parasitic helminths, such as hookworms, TIMPs have been proposed to play key roles in the host-parasite interplay, including invasion of and establishment in the vertebrate animal hosts. Currently, knowledge of helminth TIMPs is limited to a small number of studies on canine hookworms, whereas no information is available on the occurrence of TIMPs in other parasitic helminths causing neglected diseases.
Methods
In the present study, we conducted a large-scale investigation of TIMP proteins of a range of neglected human parasites including the hookworm Necator americanus, the roundworm Ascaris suum, the liver flukes Clonorchis sinensis and Opisthorchis viverrini, as well as the schistosome blood flukes. This entailed mining available transcriptomic and/or genomic sequence datasets for the presence of homologues of known TIMPs, predicting secondary structures of defined protein sequences, systematic phylogenetic analyses and assessment of differential expression of genes encoding putative TIMPs in the developmental stages of A. suum, N. americanus and Schistosoma haematobium which infect the mammalian hosts.
Results
A total of 15 protein sequences with high homology to known eukaryotic TIMPs were predicted from the complement of sequence data available for parasitic helminths and subjected to in-depth bioinformatic analyses.
Conclusions
Supported by the availability of gene manipulation technologies such as RNA interference and/or transgenesis, this work provides a basis for future functional explorations of helminth TIMPs and, in particular, of their role/s in fundamental biological pathways linked to long-term establishment in the vertebrate hosts, with a view towards the development of novel approaches for the control of neglected helminthiases.
Keywords: Tissue inhibitors of metalloproteases, Parasitic helminths, Excretory/secretory products, Transcriptomics, Bioinformatics, Protein structure, Functional inferences
Background
Parasitic helminths cause devastating diseases in humans and animals worldwide [1,2]. Amongst these parasites, soil-transmitted helminths (STHs), including Ancylostoma duodenale and Necator americanus (hookworms), Ascaris sp. (roundworms) and Trichuris spp. (whipworms), are estimated to infect over one-sixth of all humans [1,2], while trematodes, including the blood flukes Schistosoma spp. and the carcinogenic liver flukes Clonorchis sinensis and Opisthorchis viverrini, affect >200 million people worldwide [3-5].
Traditionally, the control of helminth infections has relied on the repeated and frequent use of anthelmintics [6,7], which is likely to lead to the development of drug resistance against the compounds administered (cf. [8-10]). Indeed, some studies [11-15] have reported a reduction in efficacy of mebendazole and pyrantel in N. americanus and A. duodenale in areas of Mali, North-Western Australia and Zanzibar, which has been attributed to emerging anthelmintic resistance. Given the limited knowledge of the molecular mechanisms linked to the development of drug resistance in parasitic helminths [16], as well as the unavailability of effective vaccines, much attention is now directed towards the identification of novel targets for intervention [7,17]. A detailed understanding of the molecular biology of parasitic helminths, and in particular of the structure and function of key genes and gene products playing essential roles in host-parasite interactions, could provide a basis for the design of novel therapeutics.
Among several groups of helminth molecules involved in the host-parasite interplay, protease inhibitors have been the subject of intense investigations due to their roles in a range of fundamental molecular processes, including regulation of host proteases and modulation of the host’s immune response [18]. Amongst these molecules, the inhibitors of cysteine- and serine-proteases (= cystatins and serpins, respectively; MEROPS family I25 and I04, respectively) are known to participate in the cascades of molecular events leading to parasite development through the larval stages (cystatins) [19], as well as in the inhibition of host molecules responsible for the initiation of blood coagulation (serpins) [20,21]. In addition, both molecular groups have been proposed to play key roles in the evasion and modulation of the immune response of the vertebrate host [19,21,22]. In contrast to data on cystatins and serpins, knowledge of the biological roles of parasite-derived tissue inhibitors of metalloproteases (TIMPs) is limited. Eukaryote TIMPs are a multifunctional family of inhibitors of matrix metalloproteases (MMPs), including collagenases and gelatinases, which function as important regulators of extracellular matrix (ECM) turnover, tissue remodelling and cellular behaviour [23]. The N-terminal domain of TIMPs contains a netrin module (=‘NTR’; Prosite: PDOC50189) which, in addition to harbouring the functional site responsible for the primary metalloprotease inhibitory activity, is associated with a range of biological roles, including axon guidance, regulation of cell-cell interactions during embryogenesis, cell proliferation, angiogenesis and pro- and anti-apoptotic pathways [23,24]. The NTR domain of TIMPs is also found in other groups of proteins, such as the frizzled-related (PDOC50038) and the laminyn-type EGF-like (PDOC00961) proteins, in which it fulfills distinct biological roles (cf. [24]).
In the canine hookworm Ancylostoma caninum, TIMPs are abundant components of the excretory/secretory (ES) products of the adult worm [25-27]. Ac-TMP-1 and Ac-TMP-2, two abundantly expressed TIMPs recognized by sera from dogs vaccinated with irradiated third-stage larvae (L3s) of A. caninum, were proposed to play key roles in the host-parasite interplay [25,27]. To date, no data on the occurrence of TIMP homologues/orthologues in other parasitic helminths is available. Over the last decade, advances in next-generation sequencing (NGS) technologies and bioinformatics [28-31] have provided the infrastructure for large-scale analyses of the genomes and transcriptomes of a range of parasitic helminths of public health significance, including the nematodes N. americanus, Ascaris suum, Trichuris suis and Trichinella spiralis (gastrointestinal nematodes) [32-36] and the trematodes Schistosoma mansoni, S. japonicum, S. haematobium (blood flukes), C. sinensis and O. viverrini (liver flukes) [37-41]. These advances have resulted in an expansion of sequence data available in public databases (e.g., http://www.gasserlab.org/, http://www.genedb.org/, http://www.ncbi.nlm.nih.gov/, http://nematode.net/NN3_frontpage.cgi and http://www.sanger.ac.uk/research/projects/parasitegenomics/), which represent an invaluable resource for studies of TIMPs in parasitic helminths. In the present study, we (i) conducted the first large-scale investigation of TIMP proteins in a range of parasitic helminths of both human and veterinary health significance; (ii) inferred phylogenetic relationships between/among helminth TIMPs based on predictions of secondary structures of protein sequences; and (iii) investigated differences in the levels of transcription of genes encoding putative TIMPs in different developmental stages of A. suum (cf. [34]), N. americanus (cf. [36]) and S. haematobium (cf. [40]).
Methods
Sequence data, and identification and bioinformatic analyses of TIMPs
The sequence data obtained from public sequence databases (i.e. National Center for Biotechnology Information at http://www.ncbi.nlm.nih.gov/; ENSEMBL Genome Browser at http://www.ensembl.org/index.html; WormBase, at http://www.wormbase.org; GeneDB at http://www.genedb.org/; http://www.gasserlab.org) [32-34,39,40,42-45] and analysed herein included known TIMP amino acid sequences from Homo sapiens (GenBank accession numbers XP_010392.1, NP_003246.1, P35625.1 and Q99727.1), Mus musculus (accession numbers P12032.2, P25785.2, P39876.1 and Q9JHB3.1), Canis familiaris (AF112115.1), Gallus gallus (AAB69168.1), Oryctolagus cuniculus (AAB35920.1), Drosophila melanogaster (AAL39356.1), A. caninum (AF372651.1 and EU523698.1), A. duodenale (ABP88131.1) and Caenorhabditis elegans (NP_505113.1), as well as predicted peptides inferred from (i) the whole or draft genome sequences of S. mansoni, S. japonicum, S. haematobium (http://www.genedb.org), A. suum (http://www.wormbase.org), T. spiralis (http://www.ncbi.nlm.nih.gov/nuccore/316979833), Brugia malayi and Wuchereria bancrofti (human filarial nematodes) (http://www.sanger.ac.uk/; [46]), N. americanus (human hookworm; [36]), and (ii) the transcriptomes of T. suis (swine whipworm), Oesophagostomum dentatum (swine nodule worm) (http://www.gasserlab.org), Dictyocaulus filaria (sheep lungworm; [47]) and C. sinensis, O. viverrini (human liver flukes), Fasciola hepatica and F. gigantica (bovine and deer liver fluke, respectively) (http://www.gasserlab.org). The algorithms BLASTp [48] and InterProScan [49] were used to identify TIMP proteins in each of the genomic and transcriptomic datasets based on sequence homology (e-value cut-off: 10-5) with known TIMP proteins from eukaryotes [50]. In addition, the software pScan (http://www.psc.edu/general/software/packages/emboss/appgroups/pscan.html) was used to identify regular-expression based diagnostic patterns for TIMPs (Prosite: PS00288). Signal peptides were also predicted using the program SignalP 3.0, employing both the neural network and hidden Markov Models [51]. Putative ES TIMP proteins were identified based on the presence of a signal peptide and sequence homology to one or more known ES proteins listed in the Secreted Protein (http://spd.cbi.pku.edu.cn/; [52]) and the Signal Peptide (http://proline.bic.nus.edu.sg/spdb/index.html; [53]) databases.
Secondary structure predictions and homology modelling
Structure-based sequence alignments of TIMP proteins were computed and manually edited with SBAL [54] guided by secondary structure elements predicted using the PSIPRED software [55]. Individual structure-based alignments of amino acid sequences were subjected to analysis by Bayesian inference (BI) using the program MrBayes v.3.1.2 [56] and verified by Maximum Likelihood analysis using the program MEGA v.5 [57] and the Jones-Taylor-Thornton substitution model with uniform rates among sites (JTT + G + I). Each BI analysis was conducted for 1,000,000 generations (ngen = 1,000,000), with every 100-th tree being saved, using the following parameters: rates = gamma, aamodelpr = mixed, and the other parameters left at the default settings. Tree and branch lengths were measured employing the parameter ‘sumt burnin = 1000’; an unrooted, consensus tree was constructed, with ‘contype = halfcompat’ nodal support being determined using consensus posterior probabilities and displayed employing the software FigTree (http://tree.bio.ed.ac.uk/software/figtree/). For selected TIMPs, homologues with known three-dimensional structures were identified using the protein-fold recognition software pGenTHREADER [58] and selected as templates for comparative modelling using MODELLER [59]. Twenty independent models were generated, and the model with the lowest energy was selected, its geometry analysed using PROCHECK [60] and then inspected visually with PyMOL [61].
Assessment of levels of transcription of TIMP-encoding genes
The raw sequence reads derived from each of the non-normalized cDNA libraries from A. suum infective L3s (iL3s; from eggs), migrating L3s (from liver and lung), fourth-stage larvae (L4s, from the small intestine) and muscular and reproductive tissues from each adult male and female [34], N. americanus iL3s and adults (mixed males and females) [36], as well as S. haematobium eggs and adult male and female [40] were mapped to the longest contigs encoding individual putative TIMP proteins using the program SOAP2 [62]. Briefly, raw sequence reads were aligned to the non-redundant transcriptomic data, such that each raw sequence read was uniquely mapped (i.e. to a unique transcript). Reads that mapped to more than one transcript (designated ‘multi-reads’) were randomly assigned to a unique transcript, such that they were recorded only once. To provide a relative assessment of transcript abundance, the number of raw reads that mapped to each sequence was normalized for length (i.e. reads per kilobase per million reads, RPKM) [34,40,63].
Results and discussion
TIMP proteins of parasitic helminths
A total number of 15 protein sequences with high homology (e-value cut-off: 10-5) to known eukaryotic TIMPs were predicted from the complement of sequence data available for parasitic helminths (Table 1), thus representing a solid resource for future structural and functional investigations of this protein family in parasites. The sequence data in FASTA format analysed in the present article is available in Additional file 1. Of the datasets included here, the complement of protein coding genes available for N. americanus and A. suum encoded the largest number of predicted TIMP proteins (n = 8 and 3, respectively; cf. Table 1). Three N. americanus (i.e. NECAME_13168, NECAME_07191 and NECAME_08458; cf. Table 1) and all A. suum TIMPs (GS_21732, GS_04796 and GS_08199; cf. Table 1) were predicted to contain an N-terminal signal peptide, in accordance with previous observations of A. caninum Ac-TMP-1 and Ac-TMP-2 and a netrin-domain containing homologue from Ancylostoma ceylanicum (= excretory-secretory protein 2, AceES-2), respectively [25-27,64]. Despite the sequence similarities between Ac-TMP-1, Ac-TMP-2 and AceES-2, the latter did not display human MMP inhibitory activity in vitro, thus suggesting a different function of this protein in vivo[64]. However, it should be noted that the partial MMP-inhibitory activity of Ac-TMP-2 described by Zhan et al.[26] was based on a vast molar excess of recombinant TMP-2, well beyond the 1:1 inhibitor:enzyme molar ratio required for inhibition of mammalian MMPs by their TIMP counterparts [23]. Moreover, TIMPs seem to require the C-X-C motif at the N-terminus to allow insertion into the MMP active site cleft and subsequent inhibition of catalytic activity; recombinant Ac-TMP-2 was engineered to contain a long N-terminal extension donated by the plasmid vector, so it is premature to unequivocally assign MMP inhibitory activity to the hookworm TIMPs without further work. In A. ceylanicum, secretion of AceES-2 begins soon after infection of the experimental hamster host, and steadily increases in correspondence with the onset of blood-feeding activity [65]. Furthermore, a single oral dose of recombinant AceES-2 resulted in reduced anaemia following challenge infection of hamsters with A. ceylanicum[66], which led to speculations that this molecule may play a role in the pathogenesis of hookworm disease [66]. A role for hookworm TIMPs in molecular processes linked to the invasion of the mammalian hosts and/or the inhibition of hosts MMPs at the final site of attachment has also been hypothesized, based on the fact that Ac-TMP-2 could be isolated solely from extracts and ES products of A. caninum adults, despite the corresponding mRNA being detected from both L3s and adults of this parasite [26].
Table 1.
Number of tissue inhibitor of metalloproteases (TIMP) and netrin module (NTR)-containing protein sequences, respectively, identified in each sequence dataset and listed according to taxa
| TIMPs (no. with SP) | NTR-module containing proteins (no. with SP) | |
|---|---|---|
| Nematodes |
|
|
| Ascaris suum |
3* (3) |
2 (1) |
| Brugia malayi |
- |
1 (1) |
| Dictyocaulus filaria |
1 (1) |
1 (−) |
| Necator americanus |
8** (3) |
1 (1) |
| Oesophagostomum dentatum |
2 (2) |
1 (−) |
| Trichinella spiralis |
- |
1 (1) |
| Trichuris suis |
- |
2 (−) |
| Wuchereria bancrofti |
- |
2 (−) |
| Trematodes |
|
|
| Clonorchis sinensis |
- |
1 (−) |
| Fasciola gigantica |
- |
2 (−) |
| Fasciola hepatica |
- |
1 (−) |
| Opisthorchis viverrini |
- |
4 (1) |
| Schistosoma haematobium |
1*** (1) |
5 (−) |
| Schistosoma japonicum |
- |
1 (−) |
| Schistosoma mansoni |
- |
1 (−) |
| Total | 15 (10) | 26 (5) |
The number of proteins containing a predicted N-terminal signal peptide (SP) is also indicated.
*Of these, Ascaris suum GS_04796 was up-regulated in the reproductive tissue of the adult female, while GS_21732 was up-regulated in the muscular tissue of adult males.
**Necator americanus NECAME_13168 and NECAME_07191 were up-regulated in infective L3s, while NECAME_08457 and NECAME_08458 were up-regulated in adult N. americanus, ***Schistosoma haematobium A_01727 was up-regulated in the adult male.
Of the eight genes encoding putative TIMPs in N. americanus, transcription of NECAME_13168 and NECAME_07191 was significantly up-regulated in iL3s (cf. Table 1; [36]), thus supporting a role for these proteins in the infection process of the human host. Conversely, NECAME_08457 and NECAME_08458 displayed high transcription levels in adult N. americanus (cf. Table 1; [36]), which likely reflects a diversification of function of members of this protein family in different developmental stages of this parasite. In the future, studies of differential transcription of genes encoding TIMPs in both genders and different tissues of N. americanus may help elucidate the roles that these molecules play in the fundamental molecular biology of the adult nematode. In A. suum, transcription of GS_04796 was significantly up-regulated in the adult female reproductive tissue of this nematode, whereas GS_21732 was up-regulated in the male muscle (cf. Table 1; cf. [34]). The putative TIMP proteins encoded by GS_04796 and GS_21732 share ~40% similarity with C. elegans CRI-2 (WBGene00019478; http://www.wormbase.org), the expression of which has been localized to the body wall musculature and to the vulval, anal and pharyngeal muscles of the adult nematode (cf. http://www.wormbase.org). In C. elegans, cri-2 is known to function in the cascade of molecular events linked to the regulation of the innate immune response to lipopolysaccharide (LPS) [67]. In a previous study, inhibition by small interfering RNAs (siRNAs) of the M. musculus ortholog of C. elegans cri-2 in a mouse macrophage cell line stimulated with Escherichia coli LPS resulted in decreased production of interleukin-6 (IL-6) [67]. This cytokine, in vivo, is associated with a wide range of biological activities, which include the generation of acute-phase reactions in response to infections by pathogens [68]. The putative role/s that parasite homologs of C. elegans cri-2 play in the modulation of innate immunity in vertebrate hosts remain/s unknown. However, recent evidence that recombinant Ac-TMP-1 promotes the development of a regulatory immune response by modifying the functions of bone marrow-derived dendritic cells and subsequent development of regulatory T cells [69], supports a key role for this TIMP in establishing an anti-inflammatory environment.
In flatworms, the S. haematobium gene A_01727 encoded the only trematode TIMP protein that could be identified using computational methods. Analysis of transcriptional regulation of S. haematobium A_01727 in different developmental stages revealed that this molecule is up-regulated in the adult male of this parasitic trematode (Table 1; cf. [40]). The transcript encoding mouse TIMP-1 is up-regulated in male gonads during testis morphogenesis, while expression of the corresponding protein was restricted to the cords of foetal testes [70]. In addition, the human and mouse genes encoding TIMP-2 are known to include the differential display clone 8 (DDC8) gene, whose transcription is enhanced during spermatogenesis [71]. These observations, together with earlier findings of increased expression of TIMP-1 in human foetal Sertoli cells [72,73] and testicular expression of TIMP-2 in rats [74], led to the hypothesis that these molecules may play specific roles during testis organogenesis and development [70], as well as in the migration of germ cells through the seminiferous epithelium [71]. Therefore, it is tempting to speculate a role for S. haematobium A_01727 in biological processes linked to the reproductive activity of the adult male fluke; however, this hypothesis requires rigorous testing. In the future, genetic manipulation of N. americanus, A. suum and S. haematobium by RNA interference (RNAi) and/or transgenesis [75-78], may help elucidate the function/s of putative helminth TIMPs in the reproductive biology of these organisms, as well as in other fundamental molecular processes, for instance those linked to host invasion and modulation of the host’s innate immune response.
Genomic sequence data with identity to S. haematobium A_01727 were detected in both S. mansoni (Smp_087690; e-value 3e-110) and S. japonicum (Sjp_0053050.1; e-value 6.3e-64). However, the sequence overlap between the amino acid sequence predicted from S. haematobium A_01727 and the corresponding homologues from S. mansoni and S. japonicum was limited to the NTR N-terminal module (cf. Figure 1), which would make any inference of the presence of TIMP-encoding genes in the genome sequences of the latter two species highly speculative. While it is possible that fragmentation of the Open Reading Frames (ORFs) of TIMP-encoding genes in the current assemblies of the S. mansoni and S. japonicum genomes might have occurred, the absence of homologues of eukaryote TIMPs in other species whose whole-genome sequences are currently available (e.g. B. malayi and T. spiralis) may reflect the substantial variations, both in sequence and in length, among members of this protein family in helminths [23]. Indeed, a search of the characteristic features of the N-terminal NTR module of eukaryote TIMPs using the PScan software revealed the presence of members of the netrin protein family in all parasitic helminths analysed herein (n = 26; range 1–5; cf. Table 1). This finding is in accordance with current knowledge that the genomes of helminths encode single-domain TIMP proteins that are homologous to the N-terminal domain of vertebrate TIMPs, while lacking the corresponding C-terminal region [79]. In eukaryotes, the N-terminal NTR domain of TIMPs is known to be responsible for their metalloprotease inhibitory activity [24,80,81], whereas the C-terminal domain provides binding sites for the metalloproteases [80,82,83] or for binding TIMPs to the cell surface and/or the extracellular matrix [24,81,84]. When separated from the corresponding C-terminus, the N-terminal domain of TIMPs retains its metalloprotease inhibitory activity [24,81-84]. While, based on this knowledge, single-domain helminth TIMPs may be hypothesized to exert similar metalloprotease inhibitory activities as their vertebrate counterparts, the amino acid residues present at position 2 of some mature helminth molecules (e.g. lysine, arginine and glutamine; cf. Figure 1) are atypical for vertebrate TIMPs and suggest that these proteins may perform functions that are unrelated to the inhibition of metalloprotease activity (see [23,85]). Comparative structural analyses of the amino acid sequences of TIMP proteins, as well as the N-terminal NTR module are essential to assist in-depth investigations of the function/s of this family of helminth proteins.
Figure 1.
Amino acid sequence alignment of tissue inhibitors of metalloproteases (TIMPs) based on predictions of their secondary structures.Homo sapiens TIMP-1 (GenBank accession number XP_010392.1), TIMP-2 (NP_003246.1), TIMP-3 (P35625.2), TIMP-4 (Q99727.1), Canis familiaris TIMP-2 (AF112115.1), Gallus gallus TIMP-2 (AAB69168.1), Oryctolagus cuniculus TIMP-2 (AAB35920.1), Mus musculus TIMP-1 (P12032.2), TIMP-2 (P25785.2), TIMP-3 (P39876.1), TIMP-4 (Q9JHB3.1), Drosophila melanogaster TIMP (AAL39356.1), Caenorhabditis elegans CRI-2 (K07C11.5), Ancylostoma caninum TMP-1 (AF372651.1), TMP-2 (EU523696.1), Ancylostoma duodenale TIMP-1 (ABP88131.1), Necator americanus (NECAME_13168, NECAME_07191, NECAME_01063, NECAME_05356, NECAME_05357, NECAME_14664, NECAME_08457 and NECAME_08458), Dictyocaulus filaria (1495356.2; http://www.gasserlab.org), Oesophagostomum dentatum (E59TEJM01BU99S and E59TEJM02GRTKW; http://www.gasserlab.org), Ascaris suum (GS_21732, GS_04796, GS_08199; http://www.wormbase.org), Schistosoma haematobium A_01727, Schistosoma mansoni Smp_087690 and Schistosoma japonicum Sjp_0053050 (http://www.genedb.org). Ancylostoma ceylanicum AceES-2 (GenBank Q6R7N7) is also included.
Structural analyses of eukaryote TIMPs
Structurally, the four human TIMPs are well characterized (cf. http://www.rcsb.org). These proteins consist of two domains, an N-terminal domain (N-TIMP) adopting the NTR fold, and a C-terminal domain (C-TIMP). Tertiary structures of full-length TIMP-1, TIMP-2, as well as N-TIMP-1, N-TIMP-2 and N-TIMP-3 have been determined, some in complex with their target MMPs (for an overview, see Table 2). Both N-TIMP and C-TIMP are internally stabilised by three intra-domain disulphide bridges and their structural elements are not intertwined, suggesting that the two moieties are indeed individual folding units, i.e. domains. This notion is further supported by the observation that N-TIMPs can be obtained as folded entities in vitro that display MMP inhibitory activity [79,86-88]. The shape of full-length TIMPs appears wedge-like, and the extreme N-terminus is responsible for the inhibitory action of MMPs by interaction with the protease active site cleft. In some instances, additional interactions have been observed between C-TIMP and peripheral areas of the protease that are distant to the catalytic site. However, in the case of the TIMP-2/MMP-2 complex, the interaction of C-TIMP-2 and the hemopexin domain of MMP-2 significantly enhances the affinity of the inhibitor [89,90].
Table 2.
Three-dimensional structures of tissue inhibitors of metalloproteases (TIMPs) and their complexes available in the Protein Data Bank (PDB; http://www.rcsb.org/pdb/home/home.do) as of Nov 2012
| Protein(s) | PDB accession code |
|---|---|
| N-TIMP-1 |
1d2b |
| MMP1:TIMP-1 |
2j0t |
| MMP3:TIMP-1 |
1uea |
| MMP3:N-TIMP-1 |
1oo9 |
| MMP10:TIMP-1 |
3v96 |
| MMP14:TIMP-1 |
3ma2 |
| TIMP-2 |
1br9 |
| N-TIMP-2 |
2tmp |
| pro-MMP2:TIMP-2 |
1gxd |
| MMP13:TIMP-2 |
2e2d |
| MMP14:TIMP-2 |
1bqq |
| MMP14:TIMP-2 |
1buv |
| TACE:N-TIMP-3 | 3cki |
The main interactions of TIMPs with their target proteases are formed by a continuous peptide at the N-terminal end (Cys1-Pro5 in human TIMP-1) and in a loop connecting two adjacent β-strands (Met66-Cys70 in human TIMP-1). The two regions are covalently linked by a disulphide bond (Cys1-Cys70 in human TIMP-1), and are located in the netrin module (N-TIMP) of the protein which adopts the fold of a five-stranded α-barrel with Greek key topology (OB-fold) flanked by two α-helices. The N-terminus of N-TIMP inserts into the active site of the target protease and the α-amino and the carbonyl group of Cys-1 (human TIMP-1) coordinate the active site zinc ion of the protease by displacing a water molecule otherwise bound to the metal [23]. Residue 2 (Ser, Thr) projects into the specificity (S1′) pocket of the protease. Residues 3–5 interact with the protease residues in the primed subsites, which normally harbour substrate residues C-terminal of the scissile bond. Similarly, residues 66–70 of TIMP-1 occupy the non-primed subsites of the protease that otherwise interact with the residues N-terminal to the scissile bond.
As apparent from the structure-based amino acid sequence alignment (Figure 1), TIMPs from parasitic helminths are characterised by higher sequence variation than their mammalian homologues, in accordance with the results of previous analyses of invertebrate TIMPs [23]. With respect to structure-function relationships, however, the most important feature grafted onto the netrin fold seems to be the conformation neighbouring Cys-1. In vertebrate TIMPs, 2 is either a serine or threonine that projects into the protease specificity pocket. It is important to note that neither Ac-TMP-1 nor Ac-TMP-2 have been convincingly shown (via 1:1 inhibitor:enzyme molar ratios) to possess MMP inhibitory activity. Moreover, AceES-2 produced with a flush N-terminus was screened for MMP activity at 15:1 and 115:1 molar ratios and did not display inhibitory activity (cf. [64]). The amino acid sequence alignment in Figure 1 highlights the general motif of TIMPs, C-X-C, in this region. It shows for the helminth TIMP with published inhibitory activity, Ac-TMP-2, that in addition to serine and threonine, lysine is a tolerated residue at position 2 for inhibition. Notably, AceES-2 and Ad-TIMP-1 from A. duodenale lack the second cysteine residue as well as a suitable residue at position 2 (Ser/Thr/Lys) able to protrude into the S1′ pocket of the protease for inhibition (cf. Figure 1). On this basis, one would predict Ad-TIMP-1 to not have any MMP-inhibitory activity. Thus, helminth TIMPs that show conservation at position 2 are likely to display inhibitory activities against human MMPs. The S. haematobium protein encoded by A_01727 possesses two residues (Arg-Ser) between the two N-terminal cysteine residues, which makes the prediction of functional effects difficult in the absence of experimental structures.
Helminth TIMPs for which complete amino acid sequence data is available, with the exception of Ad-TIMP-1, show conservation of the crucial structural elements of the NTR module, such as the two N-terminal cysteine residues and their covalent binding partners, as well as residues relevant for maintaining the OB-fold. The areas of largest variation are three surface-exposed loop areas, namely residues 28–41, 56–59 and 66–70 (Hs-TIMP-2 numbering; see Figure 1). Notably, there is high conservation of a basic residue (Arg20 in Hs-TIMP-1) in vertebrate and helminth TIMPs, which is an exposed residue on the surface distal to the protease interaction site (Figure 2). To our knowledge, a physiologically important function for this residue is yet to be described. Its location (at the surface of the protein) suggests a protein-protein or protein-matrix interaction; however, basic residues at this position have not been reported to be involved in extra-cellular matrix binding [91]. While S. haematobium A_01727 shares the lowest amino acid sequence identity with the other eukaryote TIMPs (cf. Figure 1), the structure-based sequence alignment, together with the accordingly predicted 3D structure, indicate that it may be a functional member of the TIMP family of proteins. This conclusion is based on the presence of all conserved cysteine residues required for intramolecular disulphide bonds of a netrin-like fold, as well as conservation of the serine residue (Ser3) expected to protrude into the catalytic site of an MMP.
Figure 2.
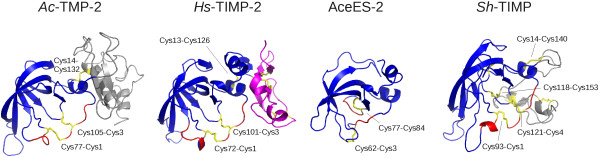
Structural comparison of four netrin domain-containing proteins. The netrin domains of Ac-TMP-2 (homology model based on Hs-TIMP-2), Hs-TIMP-2 (PDB accession code 1br9), AceES-2 (PDB accession code 3nsw) and Sh-TIMP (A_01727; homology model based on Hs-TIMP-2) are coloured blue, cysteine side chain residues are rendered as yellow sticks. Red highlighted areas indicate regions of interactions with MMPs; these regions are inferred for Ac-TMP-2, AceES-2 and Sh-TIMP based on the alignment in Figure 1. The parasite proteins Ac-TMP-2 and Sh-TIMP and human Hs-TIMP-2 share the same intra-domain disulphide bonding pattern. In contrast, AceES-2 possesses a different pattern with two intra-molecular disulphide bonds. The disulphide bond engaging the N-terminal cysteine (Cys3-Cys62) is reminiscent of that found in Ac-TMP-2, Sh-TIMP and Hs-TIMP-2. The other disulphide bond (Cys77-Cys84) is unique to AceES-2. The C-terminal domain of Hs-TIMP-2 is rendered magenta. The C-terminal domains of Ac-TMP-2 and Sh-TIMP are shown in grey for illustration only and the three-dimensional structures of these domains are neither based on computational nor experimental evidence. Comparative modelling was performed using MODELLER [59] based on the structure-based sequence alignment shown in Figure 1.
Phylogenetic analysis
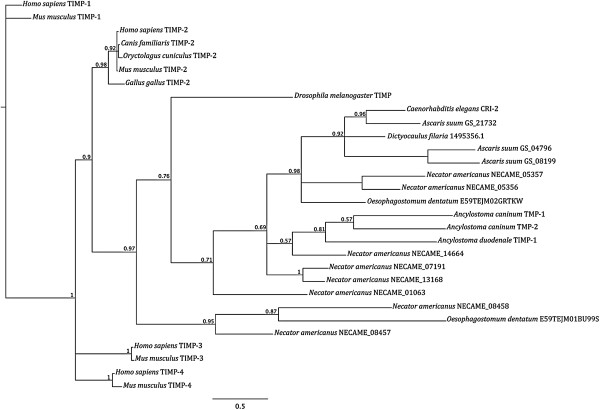
The phylogenetic analysis of eukaryote TIMPs allowed us to study the relationships between helminth TIMPs and their vertebrate counterparts (Figure 3). The analysis identified one main clade comprising TIMPs from invertebrates, including free-living and parasitic helminths (nodal support: 0.90), to the exclusion of clades formed by homologues from vertebrates (cf. Figure 3). Within the invertebrate clade, a sub-clade representing TIMPs from nematodes clustered to the exclusion of the TIMP protein from D. melanogaster (nodal support: 0.76; cf. Figure 3), supporting the existence of a monophyletic group of TIMPs for parasitic nematodes. Following the inclusion of S. haematobium A_01727 in the phylogenetic analysis, the monophyly of the nematode TIMP clade with respect to the vertebrate homologues was maintained (Additional file 2). No distinct separation between TIMPs from hookworms and those from other free-living and parasitic nematodes was observed, thus supporting the hypothesis that nematode TIMPs may be characterised by specific functional properties, distinct from those of their vertebrate homologues. Whether nematode TIMPs have originated following loss of the C-terminal domain from a vertebrate ancestor or from a distinct gene line (cf. [23]) remains to be explored.
Figure 3.
The phylogenetic relationships of tissue inhibitor of metalloproteases (TIMPs) based on Bayesian Inference. The posterior probability supporting each clade is indicated. The corresponding phylogenetic reconstructions obtained by Maximum Likelihood (JTT + G + I) analyses of TIMP proteins are available from the primary author (CC) upon request. Homo sapiens TIMP-1 (GenBank accession number XP_010392.1), TIMP-2 (NP_003246.1), TIMP-3 (P35625.2), TIMP-4 (Q99727.1), Gallus gallus TIMP-2 (AAB69168.1), Canis familiaris TIMP-2 (AF112115.1), Oryctolagus cuniculus TIMP-2 (AAB35920.1), Drosophila melanogaster TIMP (AAL39356.1), Mus musculus TIMP-1 (P12032.2), TIMP-2 (P25785.2), TIMP-3 (P39876.1), TIMP-4 (Q9JHB3.1), Caenorhabditis elegans CRI-2 (K07C11.5), Ancylostoma caninum TMP-1 (AF372651.1), TMP-2 (EU523696.1), Ancylostoma duodenale TIMP-1 (ABP88131.1), Necator americanus (NECAME_13168, NECAME_07191, NECAME_01063, NECAME_05356, NECAME_05357, NECAME_14664, NECAME_08457 and NECAME_08458), Dictyocaulus filaria (1495356.2; http://www.gasserlab.org), Oesophagostomum dentatum (E59TEJM01BU99S and E59TEJM02GRTKW; http://www.gasserlab.org) and Ascaris suum (GS_21732, GS_04796, GS_08199; http://www.wormbase.org).
Conclusion
The current availability of ‘-omics’ technologies, applied to in-depth investigations of pathogens causing neglected diseases [31,92-94], are becoming pivotal for a better understanding of the structure and function of TIMP proteins in different species and developmental stages of parasitic helminths. For instance, data from in-depth comparative structural analyses between helminth TIMPs and their vertebrate counterparts, will be crucial in future studies aimed at assessing the suitability of parasite TIMPs as novel targets for intervention. Supported by the availability of the whole-genome sequences of, for instance, schistosomes and A. suum[34,37,38,40] and by current efforts to expand genomic sequencing to other neglected parasites (e.g. hookworms; [95]), the application of gene manipulation technologies such as RNAi and/or transgenesis [94,96,97], will allow the function of helminth TIMP proteins in fundamental biological pathways to be elucidated. Perhaps the most important question that is yet to be addressed in any depth is the function of helminth TIMPs. Are they inhibitors of metalloproteases? Is their primary purpose to suppress inflammation, and if so, how do they do it? We hope that the molecular information provided herein on parasitic helminth TIMPs will provide a framework on which to build intensive research activities around this intriguing family of proteins and their roles in host-parasite interactions.
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
CC, AH and AL conceived and designed the experiments; CC, AH, DP and SN analysed the data; CC, AH, MM and AL wrote the manuscript. All authors read and approved the final version of the manuscript.
Supplementary Material
TIMP amino acid sequences inferred from genomic and/or transcriptomic sequence data analysed in the present study.
Phylogenetic relationships. The phylogenetic relationships of tissue inhibitor of metalloproteases (TIMPs) based on Bayesian Inference. The posterior probability supporting each clade is indicated. The corresponding phylogenetic reconstructions obtained by Maximum Likelihood (JTT + G + I) analyses of TIMP proteins are available from the primary author (CC) upon request. Homo sapiens TIMP-1 (GenBank accession number XP_010392.1), TIMP-2 (NP_003246.1), TIMP-3 (P35625.2), TIMP-4 (Q99727.1), Gallus gallus TIMP-2 (AAB69168.1), Canis familiaris TIMP-2 (AF112115.1), Oryctolagus cuniculus TIMP-2 (AAB35920.1), Drosophila melanogaster TIMP (AAL39356.1), Mus musculus TIMP-1 (P12032.2), TIMP-2 (P25785.2), TIMP-3 (P39876.1), TIMP-4 (Q9JHB3.1), Caenorhabditis elegans CRI-2 (K07C11.5), Ancylostoma caninum TMP-1 (AF372651.1), TMP-2 (EU523696.1), Ancylostoma duodenale TIMP-1 (ABP88131.1), Necator americanus (NECAME_13168, NECAME_07191, NECAME_01063, NECAME_05356, NECAME_05357, NECAME_14664, NECAME_08457 and NECAME_08458Contig150, Contig565, Contig7; http://www.gasserlab.org), Dictyocaulus filaria (1495356.2; http://www.gasserlab.org), Oesophagostomum dentatum (E59TEJM01BU99S and E59TEJM02GRTKW; http://www.gasserlab.org) Ascaris suum (GS_21732, GS_04796, GS_08199; http://www.wormbase.org) and Schistosoma haematobium (A_01727; http://www.genedb.org).
Contributor Information
Cinzia Cantacessi, Email: cinzia.cantacessi@jcu.edu.au.
Andreas Hofmann, Email: a.hofmann@griffith.edu.au.
Darren Pickering, Email: darren.pickering@jcu.edu.au.
Severine Navarro, Email: severine.navarro@jcu.edu.au.
Makedonka Mitreva, Email: mmitreva@genome.wustl.edu.
Alex Loukas, Email: alex.loukas@jcu.edu.au.
Acknowledgements
This work was supported by a program grant from the National Health and Medical Research Council of Australia (NHMRC). AL and CC are supported by a NHMRC principal research fellowship and early-career research fellowship, respectively. The authors would like to thank Dr Alex Strongin (Sanford-Burnham Medical Research Institute, La Jolla, CA, USA) for helpful discussions on TIMP/MMP interactions.
References
- Hotez PJ, Brindley PJ, Bethony JM, King CH, Pearce EJ. et al. Helminth infections: the great neglected tropical diseases. J Clin Invest. 2008;118:1311–1321. doi: 10.1172/JCI34261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lustigman S, Prichard RK, Gazzinelli A, Grant WN, Boatin BA. et al. A research agenda for helminth diseases of humans: the problem of helminthiasis. PLoS Negl Trop Dis. 2012;6:e1582. doi: 10.1371/journal.pntd.0001582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rollinson D. A wake up call for urinary schistosomiasis: reconciling research effort with public health importance. Parasitology. 2009;136:1593–1610. doi: 10.1017/S0031182009990552. [DOI] [PubMed] [Google Scholar]
- Shin HR, Oh JK, Masuyer E, Curado MP, Bouvard V. et al. Epidemiology of cholangiocarcinoma: an update focusing on risk factors. Cancer Sci. 2010;101:579–585. doi: 10.1111/j.1349-7006.2009.01458.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fried B, Reddy A, Mayer D. Helminths in human carcinogenesis. Cancer Lett. 2011;305:239–249. doi: 10.1016/j.canlet.2010.07.008. [DOI] [PubMed] [Google Scholar]
- Keiser J, Utzinger J. The drugs we have and the drugs we need against major helminth infections. Adv Parasitol. 2010;73:197–230. doi: 10.1016/S0065-308X(10)73008-6. [DOI] [PubMed] [Google Scholar]
- Caffrey CR, Secor WE. Schistosomiasis: from drug deployment to drug development. Curr Opin Infect Dis. 2011;24:410–417. doi: 10.1097/QCO.0b013e328349156f. [DOI] [PubMed] [Google Scholar]
- Wolstenholme AJ, Fairweather I, Prichard R, von Samson-Himmelstjerna G, Sangster NC. Drug resistance in veterinary helminths. Trends Parasitol. 2004;20:469–476. doi: 10.1016/j.pt.2004.07.010. [DOI] [PubMed] [Google Scholar]
- Beech RN, Skuce P, Bartley DJ, Martin RJ, Prichard RK. et al. Anthelmintic resistance: markers for resistance, or susceptibility? Parasitology. 2011;138:160–174. doi: 10.1017/S0031182010001198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoste H, Torres-Acosta JF. Non chemical control of helminths in ruminants: adapting solutions for changing worms in a changing world. Vet Parasitol. 2011;180:144–154. doi: 10.1016/j.vetpar.2011.05.035. [DOI] [PubMed] [Google Scholar]
- De Clercq D, Sacko M, Behnke J, Gilbert F, Dorny P. et al. Failure of mebendazole in treatment of human hookworm infections in the southern region of Mali. Am J Trop Med Hyg. 1997;57:25–30. doi: 10.4269/ajtmh.1997.57.25. [DOI] [PubMed] [Google Scholar]
- Reynoldson JA, Behnke JM, Pallant LJ, Macnish MG, Gilbert F. et al. Failure of pyrantel in treatment of human hookworm infections (Ancylostoma duodenale) in the Kimberley region of north west Australia. Acta Trop. 1997;68:301–312. doi: 10.1016/s0001-706x(97)00106-x. [DOI] [PubMed] [Google Scholar]
- Sacko M, De Clercq D, Behnke JM, Gilbert FS, Dorny P. et al. Comparison of the efficacy of mebendazole, albendazole and pyrantel in treatment of human hookworm infections in the southern region of Mali, West Africa. Trans R Soc Trop Med Hyg. 1999;93:195–203. doi: 10.1016/s0035-9203(99)90306-1. [DOI] [PubMed] [Google Scholar]
- Albonico M, Engels D, Savioli L. Monitoring drug efficacy and early detection of drug resistance in human soil-trasmitted nematodes: a pressing public health agenda for helminth control. Int J Parasitol. 2004;34:1205–1210. doi: 10.1016/j.ijpara.2004.08.001. [DOI] [PubMed] [Google Scholar]
- Kotze AC, Kopp SR. The potential impact of density dependent fecundity on the use of the faecal egg count reduction test for detecting drug resistance in human hookworms. PLoS Negl Trop Dis. 2008;2:e297. doi: 10.1371/journal.pntd.0000297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gilleard JS, Beech RN. Population genetics of anthelmintic resistance in parasitic nematodes. Parasitology. 2007;134:1133–1147. doi: 10.1017/S0031182007000066. [DOI] [PubMed] [Google Scholar]
- Kotze AC. Target-based and whole-worm screening approaches to anthelmitic discovery. Vet Parasitol. 2012;186:118–123. doi: 10.1016/j.vetpar.2011.11.052. [DOI] [PubMed] [Google Scholar]
- Knox DP. Proteinase inhibitors and helminth parasite infection. Parasite Immunol. 2007;29:57–71. doi: 10.1111/j.1365-3024.2006.00913.x. [DOI] [PubMed] [Google Scholar]
- Klotz C, Ziegler T, Danilowicz-Luebert E, Hartmann S. Cystatins of parasitic organisms. Adv Exp Med Biol. 2011;712:208–221. doi: 10.1007/978-1-4419-8414-2_13. [DOI] [PubMed] [Google Scholar]
- Cappello M, Vlasuk GP, Bergum PW, Huang S, Hotez PJ. Ancylostoma caninum anticoagulant peptide: a hookworm-derived inhibitor of human anticoagulant factor Xa. Proc Natl Acad Sci USA. 1995;92:6152–6156. doi: 10.1073/pnas.92.13.6152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Molehin AJ, Gobert GN, McManus DP. Serine protease inhibitors of parasitic helminths. Parasitology. 2012;139:681–695. doi: 10.1017/S0031182011002435. [DOI] [PubMed] [Google Scholar]
- Hartmann S, Lucius R. Modulation of host immune responses by nematode cystatins. Int J Parasitol. 2003;33:1291–1302. doi: 10.1016/s0020-7519(03)00163-2. [DOI] [PubMed] [Google Scholar]
- Brew K, Nagase H. The tissue inhibitors of metalloproteinases (TIMPs): an ancient family with structural and functional diversity. Biochim Biophys Acta. 2010;1803:55–71. doi: 10.1016/j.bbamcr.2010.01.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bányai L, Patthy L. The NTR module: domains of netrins, secreted frizzled related proteins, and type I procollagen C-proteinase enhancer protein are homologous with tissue inhibitors of metalloproteases. Protein Sci. 1999;8:1636–1642. doi: 10.1110/ps.8.8.1636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhan B, Badamchian M, Meihua B, Ashcom J, Feng J. et al. Molecular cloning and purification of Ac-TMP, a developmentally regulated putative tissue inhibitor of metalloprotease released in relative abundance by adult Ancylostoma hookworm. Am J Trop Med Hyg. 2002;66:238–244. doi: 10.4269/ajtmh.2002.66.238. [DOI] [PubMed] [Google Scholar]
- Zhan B, Gupta R, Wong SPY, Bier S, Jiang D. et al. Molecular cloning and characterization of Ac-TMP-2, a tissue inhibitor of metalloproteinase secreted by adult Ancylostoma caninum. Mol Biochem Parasitol. 2008;162:142–148. doi: 10.1016/j.molbiopara.2008.08.008. [DOI] [PubMed] [Google Scholar]
- Mulvenna J, Hamilton B, Nagaraj SH, Smyth D, Loukas A. et al. Proteomics analysis of the excretory/secretory component of the blood-feeding stage of the hookworm, Ancylostoma caninum. Mol Cell Proteomics. 2009;8:109–121. doi: 10.1074/mcp.M800206-MCP200. [DOI] [PubMed] [Google Scholar]
- Margulies M, Egholm M, Altman WE, Attiya S, Bader JS. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature. 2005;437:376–380. doi: 10.1038/nature03959. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bentley DR, Balasubramanian S, Swerdlow HP, Smith GP, Milton J. et al. Accurate whole human genome sequencing using reversible terminator chemistry. Nature. 2008;456:53–59. doi: 10.1038/nature07517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harris TD, Buzby PR, Babcock H, Beer E, Bowers J. et al. Single-molecule DNA sequencing of a viral genome. Science. 2008;320:106–109. doi: 10.1126/science.1150427. [DOI] [PubMed] [Google Scholar]
- Cantacessi C, Campbell BE, Gasser RB. Key strongylid nematodes of animals – impact of next-generation transcriptomics on systems biology and biotechnology. Biotechnol Adv. 2012;30:469–488. doi: 10.1016/j.biotechadv.2011.08.016. [DOI] [PubMed] [Google Scholar]
- Cantacessi C, Mitreva M, Jex AR, Young ND, Campbell BE. et al. Massively parallel sequencing and analysis of the Necator americanus transcriptome. PLoS Negl Trop Dis. 2010;4:e684. doi: 10.1371/journal.pntd.0000684. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cantacessi C, Young ND, Nesjum P, Jex AR, Campbell BE. et al. The transcriptome of Trichuris suis – first molecular insights into a parasite with curative properties for key immune diseases of humans. PLoS One. 2011;6:e23590. doi: 10.1371/journal.pone.0023590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jex AR, Liu S, Li B, Young ND, Hall RS. et al. Ascaris suum draft genome. Nature. 2011;479:529–533. doi: 10.1038/nature10553. [DOI] [PubMed] [Google Scholar]
- Mitreva M, Jasmer JP, Zarlenga DS, Wang Z, Abubucker S. et al. The draft genome of the parasitic nematode Trichinella spiralis. Nat Genet. 2011;43:228–235. doi: 10.1038/ng.769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tang Y, Gao X, Rosa B, Abubucker S, Hallsworth-Pepin K, Genome of the human hookworm Necator americanus . 2013. submitted for publication. [DOI] [PMC free article] [PubMed]
- Berriman M, Haas BJ, LoVerde PT, Wilson RA, Dillon GP. et al. The genome of the blood fluke Schistosoma mansoni. Nature. 2009;460:352–358. doi: 10.1038/nature08160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- The Schistosoma japonicum Genome Sequencing and Functional Analysis Consortium. The Schistosoma japonicum genome reveals features of host-parasite interplay. Nature. 2009;460:345–351. doi: 10.1038/nature08140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young ND, Campbell BE, Hall RS, Jex AR, Cantacessi C. et al. Unlocking the transcriptomes of two carcinogenic parasites, Clonorchis sinensis and Opisthorchis viverrini. PLoS Negl Trop Dis. 2010;4:e719. doi: 10.1371/journal.pntd.0000719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young ND, Jex AR, Li B, Liu S, Yang L. et al. Whole genome sequence of Schistosoma haematobium. Nat Genet. 2012;44:221–225. doi: 10.1038/ng.1065. [DOI] [PubMed] [Google Scholar]
- Wang X, Chen W, Huang Y, Sun J, Men J. et al. The draft genome of the carcinogenic human liver fluke Clonorchis sinensis. Genome Biol. 2011;12:R107. doi: 10.1186/gb-2011-12-10-r107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cantacessi C, Jex AR, Hall RS, Young ND, Campbell BE. et al. A practical, bioinformatic workflow system for large data sets generated by next-generation sequencing. Nucleic Acids Res. 2010;38:e171. doi: 10.1093/nar/gkq667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young ND, Hall RS, Jex AR, Cantacessi C, Gasser RB. Elucidating the transcriptome of Fasciola hepatica – a key to fundamental and biotechnological discoveries for a neglected parasite. Biotechnol Adv. 2010;28:222–231. doi: 10.1016/j.biotechadv.2009.12.003. [DOI] [PubMed] [Google Scholar]
- Young ND, Jex AR, Cantacessi C, Hall RS, Campbell BE. et al. A portrait of the transcriptome of the neglected trematode, Fasciola gigantica – biological and biotechnological implications. PLoS Negl Trop Dis. 2011;5:e1004. doi: 10.1371/journal.pntd.0001004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Flicek P, Ridwan Amode M, Barrell D, Beal K, Brent S. et al. Ensembl 2012. Nucleic Acids Res. 2012;40:D84–D90. doi: 10.1093/nar/gkr991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ghedin E, Wang S, Spiro D, Caler E, Zhao Q. et al. Draft genome of the filarial nematode parasite Brugia malayi. Science. 2007;317:1756–1760. doi: 10.1126/science.1145406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mangiola S, Young ND, Korhonen P, Mondal A, Scheerlinck JP, Getting the most out of parasitic helminth transcriptomes using HelmDB: implications for biology and biotechnology. Biotechnol Adv. 2013. in press. [DOI] [PubMed]
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic Local Alignment Search Tool. J Mol Biol. 1990;215:403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- Hunter S, Jones P, Mitchell A, Apweiler R, Attwood TK. et al. InterPro in 2011: new developments in the family and domain prediction database. Nucleic Acids Res. 2012;40:D306–D312. doi: 10.1093/nar/gkr948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Docherty AJ, Lyons A, Smith BJ, Wright EM, Stephens PE. et al. Sequence of human tissue inhibitor of metalloproteinases and its identity to erythroid-potentiating activity. Nature. 1985;318:66–69. doi: 10.1038/318066a0. [DOI] [PubMed] [Google Scholar]
- Bendtsen JD, Nielsen H, von Heijne G, Brunak S. Improved prediction of signal peptides: SignalP 3.0. J Mol Biol. 2004;340:783–795. doi: 10.1016/j.jmb.2004.05.028. [DOI] [PubMed] [Google Scholar]
- Chen Y, Zhang Y, Yin Y, Gao G, Li S. et al. SPD—a web based secreted protein database. Nucleic Acids Res. 2005;33:D169–D173. doi: 10.1093/nar/gki093. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choo KH, Tan TW, Ranganathan S. SPdb – a signal peptide database. BMC Bioinformatics. 2005;6:249. doi: 10.1186/1471-2105-6-249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang CK, Broder U, Weeratunga SK, Gasser RB, Loukas A. et al. SBAL: a practical tool to generate and edit structure-based amino acid sequence alignments. Bioinformatics. 2012;28:1026–1027. doi: 10.1093/bioinformatics/bts035. [DOI] [PubMed] [Google Scholar]
- McGuffin LJ, Bryson K, Jones DT. The PSIPRED protein structure prediction server. Bioinformatics. 2000;16:404–405. doi: 10.1093/bioinformatics/16.4.404. [DOI] [PubMed] [Google Scholar]
- Ronquist F, Huelsenbeck J. MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics. 2003;19:1572–1574. doi: 10.1093/bioinformatics/btg180. [DOI] [PubMed] [Google Scholar]
- Tamura K, Peterson D, Peterson N, Stecher G, Nei M. et al. MEGA5: Molecular Evolutionary Genetics Analysis using maximum likelihood, evolutionary distances, and maximum parsimony methods. Mol Biol Evol. 2011;28:2731–2739. doi: 10.1093/molbev/msr121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lobley A, Sadowski MI, Jones DT. pGenTHREADER and pDomTHREADER: new methods for improved protein fold recognition and superfamily discrimination. Bioinformatics. 2009;25:1761–1767. doi: 10.1093/bioinformatics/btp302. [DOI] [PubMed] [Google Scholar]
- Sali A, Blundell T. Comparative protein modelling by satisfaction of spatial restraints. J Mol Biol. 1993;234:779–815. doi: 10.1006/jmbi.1993.1626. [DOI] [PubMed] [Google Scholar]
- Laskowski R, MacArthur M, Moss D, Thornton J. PROCHECK: A program to check the stereochemical quality of protein structures. J Appl Cryst. 1993;26:283–291. [Google Scholar]
- DeLano W. The PyMOL Molecular Graphics System. 2002. http://www.pymol.org/
- Li R, Yu C, Li Y, Lam TW, Yiu SM. et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics. 2009;25:1966–1967. doi: 10.1093/bioinformatics/btp336. [DOI] [PubMed] [Google Scholar]
- Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods. 2008;6:S22–S32. doi: 10.1038/nmeth.1226. [DOI] [PubMed] [Google Scholar]
- Kucera K, Harrison LM, Cappello M, Modis Y. Ancylostoma ceylanicum excretory-secretory protein 2 adopts a netrin-like fold and defines a novel family of nematode proteins. J Mol Biol. 2011;408:9–17. doi: 10.1016/j.jmb.2011.02.033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bungiro RD Jr, Cappello M. Detection of excretory/secretory coproantigens in experimental hookworm infection. Am J Trop Med Hyg. 2005;73:915–920. [PubMed] [Google Scholar]
- Bungiro RD Jr, Solis CV, Harrison LM, Cappello M. Purification and molecular cloning of and immunization with Ancylostoma ceylanicum excretory-secretory protein 2, an immunoreactive protein produced by adult hookworms. Infect Immun. 2004;72:2203–2213. doi: 10.1128/IAI.72.4.2203-2213.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alper S, Laws R, Lackford B, Boyd WA, Dunlap P. et al. Identification of innate immunity genes and pathways using a comparative genomics approach. Proc Natl Acad Sci USA. 2008;105:7016–7021. doi: 10.1073/pnas.0802405105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ding C, Cicuttini F, Li J, Jones G. Targeting IL-6 in the treatment of inflammatory and autoimmune diseases. Expert Opin Investig Drugs. 2009;18:1457–1466. doi: 10.1517/13543780903203789. [DOI] [PubMed] [Google Scholar]
- Cuellar C, Wu W, Mendez S. The hookworm tissue inhibitor of metalloproteases (Ac-TMP-1) modifies dendritic cell function and induces generation of CD4 and CD8 suppressor T cells. PLoS Negl Trop Dis. 2009;3:e439. doi: 10.1371/journal.pntd.0000439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guyot R, Magre S, Leduque P, Le Magueresse-Battistoni B. Differential expression of tissue inhibitor of metalloproteinases type 1 (TIMP-1) during mouse gonad development. Dev Dyn. 2003;227:357–366. doi: 10.1002/dvdy.10321. [DOI] [PubMed] [Google Scholar]
- Jaworski DM, Beem-Miller M, Lluri G, Barrantes-Reynolds R. Potential regulatory relationship between the nested gene DDC8 and its host gene tissue inhibitor of metalloproteinase-2. Physiol Genomics. 2007;28:168–178. doi: 10.1152/physiolgenomics.00160.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharpe RM. In: The physiology of reproduction. Knobil E, Neill JD, editor. New York: Raven Press; 1994. Regulation of spermatogenesis; pp. 1363–1434. [Google Scholar]
- Robinson LL, Sznajder NA, Riley SC, Anderson RA. Matrix metalloproteinases and tissue inhibitors of metalloproteinases in human fetal testis and ovary. Mol Hum Reprod. 2001;7:641–648. doi: 10.1093/molehr/7.7.641. [DOI] [PubMed] [Google Scholar]
- Grima J, Calcagno K, Cheng CY. Purification, cDNA cloning, and developmental changes in the steady-state mRNA level of rat testicular tissue inhibitor of metalloproteases-2 (TIMP-2) J Androl. 1996;17:263–275. [PubMed] [Google Scholar]
- Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature. 1998;391:806–811. doi: 10.1038/35888. [DOI] [PubMed] [Google Scholar]
- Islam MK, Miyoshi T, Yamada M, Tsuij N. Pyrophosphatase of the roundworm Ascaris suum plays an essential role in the worm’s molting and development. Infect Immun. 2005;73:1995–2004. doi: 10.1128/IAI.73.4.1995-2004.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rinaldi G, Okatcha TI, Popratiloff A, Ayuk MA, Suttiprapa S. et al. Genetic manipulation of Schistosoma haematobium, the neglected schistosome. PLoS Negl Trop Dis. 2011;5:e1348. doi: 10.1371/journal.pntd.0001348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rinaldi G, Eckert SE, Tsai IJ, Suttiprapa S, Kines KJ. et al. Germline transgenesis and insertional mutagenesis in Schistosoma mansoni mediated by murine leukemia virus. PLoS Pathog. 2012;8:e1002810. doi: 10.1371/journal.ppat.1002820. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brew K, Dinakarpandian D, Nagase H. Tissue inhibitors of metalloproteinases: evolution, structure and function. Biochim Biophys Acta, Prot Struct Mol Enzymol. 2000;1477:267–283. doi: 10.1016/s0167-4838(99)00279-4. [DOI] [PubMed] [Google Scholar]
- Murphy G, Houbrechts A, Cockett MI, Williamson RA, O’Shea M. et al. The N-terminal domain of tissue inhibitor of metalloproteinases retains metalloproteinase inhibitory activity. Biochemistry. 1991;30:8097–8102. doi: 10.1021/bi00247a001. [DOI] [PubMed] [Google Scholar]
- Langton KP, Barker MD, McKie N. Localization of the functional domains of human tissue inhibitor of metalloproteinases-3 and the effects of a Sorsby’s fundus dystrophy mutation. J Biol Chem. 1998;273:16778–16781. doi: 10.1074/jbc.273.27.16778. [DOI] [PubMed] [Google Scholar]
- De Clerck YA, Yean TD, Lee Y, Tomich JM, Langley KE. Characterization of the functional domain of tissue inhibitor of metalloproteinase-2 (TIMP-2) Biochem J. 1993;289:65–69. doi: 10.1042/bj2890065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Willenbrock F, Crabbe T, Slocombe PM, Sutton CW, Docherty AJP. et al. The activity of the tissue inhibitors of metalloproteinases is regulated by C-terminal domain interactions: a kinetic analysis of the inhibition of gelatinase A. Biochemistry. 1993;32:4330–4337. doi: 10.1021/bi00067a023. [DOI] [PubMed] [Google Scholar]
- Ko YC, Langley KE, Mendiaz EA, Parker VP, Taylor SM. et al. The C-terminal domain of tissue inhibitor of metalloproteinase-2 is required for cell binding but not for antimetalloproteinase activity. Biochem Biophys Res Commun. 1997;236:100–105. doi: 10.1006/bbrc.1997.6859. [DOI] [PubMed] [Google Scholar]
- Meng Q, Malinovskii V, Huang W, Hu YJ, Chung L. et al. Residue 2 of TIMP-1 is a major determinant of affinity and specificity for matrix metalloproteinases but effects of substitutions do not correlate with those of the corresponding P1′ residue of substrate. J Biol Chem. 1999;274:10184–10189. doi: 10.1074/jbc.274.15.10184. [DOI] [PubMed] [Google Scholar]
- Huang W, Suzuki K, Nagase H, Arumugam S, Van Doren SR. et al. Folding and characterisation of the amino-terminal domain of human tissue inhibitor of metalloproteinases-1 (TIMP-1) expressed at high yield in E. coli. FEBS Lett. 1996;384:155–161. doi: 10.1016/0014-5793(96)00304-3. [DOI] [PubMed] [Google Scholar]
- Amour A, Slocombe PM, Webster A, Butler M, Knight CG. et al. TNF-alpha converting enzyme (TACE) is inhibited by TIMP-3. FEBS Lett. 1998;435:39–44. doi: 10.1016/s0014-5793(98)01031-x. [DOI] [PubMed] [Google Scholar]
- Kashiwagi M, Tortorella M, Nagase H, Brew K. TIMP-3 is a potent inhibitor of aggrecanase 1 (ADAM-TS4) and aggrecanase 2 (ADAM-TS5) J Biol Chem. 2001;276:12501–12504. doi: 10.1074/jbc.C000848200. [DOI] [PubMed] [Google Scholar]
- Olson MW, Gervasi DC, Mobahsery S, Fridman R. Kinetic analysis of the binding of human matrix metalloproteinse-2 and −9 to tissue inhibitor of matrix metalloproteinase (TIMP-1 and TIMP-2) J Biol Chem. 1997;272:29975–29983. doi: 10.1074/jbc.272.47.29975. [DOI] [PubMed] [Google Scholar]
- Hutton M, Willenbrock F, Brocklehurst K, Murphy G. Kinetic analysis of the mechanism of interaction of full-length TIMP-2 and gelatinase A – evidence for the existence of a low affinity intermediate. Biochemistry. 1998;37:10094–10098. doi: 10.1021/bi980616p. [DOI] [PubMed] [Google Scholar]
- Lee MH, Atkinson S, Murphy G. Identification of the Extracellular Matrix (ECM) Binding Motifs of Tissue Inhibitor of Metalloproteinases (TIMP)-3 and Effective Transfer to TIMP-1. J Biol Chem. 2007;282:6887–6898. doi: 10.1074/jbc.M610490200. [DOI] [PubMed] [Google Scholar]
- Webster JP, Oliviera G, Rollinson D, Gower CM. Schistosome genomes: a wealth of information. Trends Parasitol. 2010;26:103–106. doi: 10.1016/j.pt.2009.12.006. [DOI] [PubMed] [Google Scholar]
- Loukas A, Gaze S, Mulvenna JP, Gasser RB, Brindley PJ. et al. Vaccinomics for the major blood feeding helminths of humans. OMICS. 2011;15:567–577. doi: 10.1089/omi.2010.0150. [DOI] [PubMed] [Google Scholar]
- Hagen J, Lee EF, Fairlie WD, Kalinna BH. Functional genomics approaches in parasitic helminths. Parasite Immunol. 2012;34:163–182. doi: 10.1111/j.1365-3024.2011.01306.x. [DOI] [PubMed] [Google Scholar]
- Kumar S, Koutsovoulos G, Kaur G, Blaxter M. Toward 959 nematode genomes. Worm. 2012;1:1–9. doi: 10.4161/worm.19046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kalinna BH, Brindley PJ. Manipulating the manipulators: advances in parasitic helminth transgenesis and RNAi. Trends Parasitol. 2007;23:197–204. doi: 10.1016/j.pt.2007.03.007. [DOI] [PubMed] [Google Scholar]
- Selkirk ME, Huang SC, Knox DP, Britton C. The development of RNA interference (RNAi) in gastrointestinal nematodes. Parasitology. 2012;139:605–612. doi: 10.1017/S0031182011002332. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
TIMP amino acid sequences inferred from genomic and/or transcriptomic sequence data analysed in the present study.
Phylogenetic relationships. The phylogenetic relationships of tissue inhibitor of metalloproteases (TIMPs) based on Bayesian Inference. The posterior probability supporting each clade is indicated. The corresponding phylogenetic reconstructions obtained by Maximum Likelihood (JTT + G + I) analyses of TIMP proteins are available from the primary author (CC) upon request. Homo sapiens TIMP-1 (GenBank accession number XP_010392.1), TIMP-2 (NP_003246.1), TIMP-3 (P35625.2), TIMP-4 (Q99727.1), Gallus gallus TIMP-2 (AAB69168.1), Canis familiaris TIMP-2 (AF112115.1), Oryctolagus cuniculus TIMP-2 (AAB35920.1), Drosophila melanogaster TIMP (AAL39356.1), Mus musculus TIMP-1 (P12032.2), TIMP-2 (P25785.2), TIMP-3 (P39876.1), TIMP-4 (Q9JHB3.1), Caenorhabditis elegans CRI-2 (K07C11.5), Ancylostoma caninum TMP-1 (AF372651.1), TMP-2 (EU523696.1), Ancylostoma duodenale TIMP-1 (ABP88131.1), Necator americanus (NECAME_13168, NECAME_07191, NECAME_01063, NECAME_05356, NECAME_05357, NECAME_14664, NECAME_08457 and NECAME_08458Contig150, Contig565, Contig7; http://www.gasserlab.org), Dictyocaulus filaria (1495356.2; http://www.gasserlab.org), Oesophagostomum dentatum (E59TEJM01BU99S and E59TEJM02GRTKW; http://www.gasserlab.org) Ascaris suum (GS_21732, GS_04796, GS_08199; http://www.wormbase.org) and Schistosoma haematobium (A_01727; http://www.genedb.org).