Abstract
Expression of the c-myc gene can be controlled by transcriptional or posttranscriptional mechanisms (or both), depending on the cell type and the growth conditions. An important mechanism of posttranscriptional regulation is modulation of cytoplasmic c-myc mRNA stability; normal human and murine c-myc mRNAs have cytoplasmic half-lives of 30 min or less. To elucidate the c-myc sequences which impart this unusually high rate of cytoplasmic transcript turnover, we have constructed various deletion and hybrid c-myc genes and analyzed the cytoplasmic stability of the mRNAs produced from them in stably transfected murine fibroblasts. The results indicate that sequences contained within the 5' and 3' ends of the c-myc transcript can affect cytoplasmic stability. Specifically, the 3' untranslated sequences of c-myc exon 3 are required for, but do not ensure, a high rate of transcript turnover in the cytoplasm. Exon 2 coding sequences do not seem to be involved, and exon 1 sequences at the 5' end of the transcript have only a small effect on cytoplasmic transcript stability. The sequences that are primarily responsible for the short c-myc RNA half-life were localized to a region of 140 bases in the 3' untranslated region.
Full text
PDF



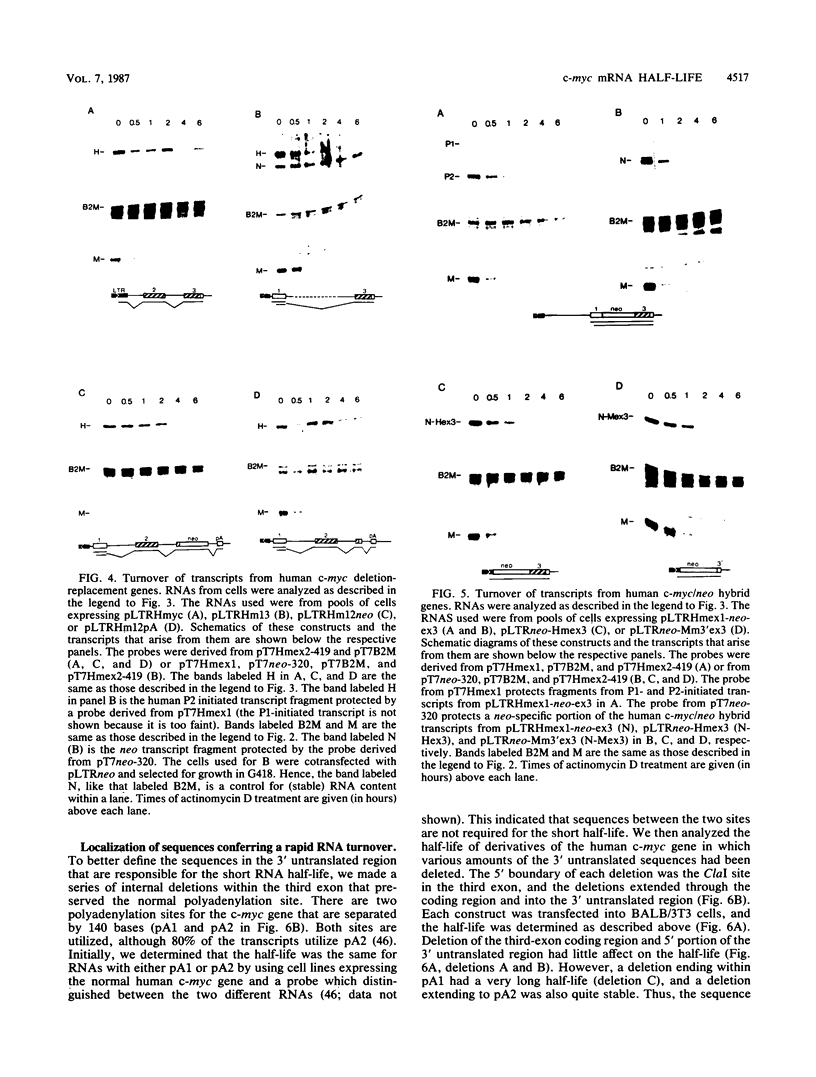




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Battey J., Moulding C., Taub R., Murphy W., Stewart T., Potter H., Lenoir G., Leder P. The human c-myc oncogene: structural consequences of translocation into the IgH locus in Burkitt lymphoma. Cell. 1983 Oct;34(3):779–787. doi: 10.1016/0092-8674(83)90534-2. [DOI] [PubMed] [Google Scholar]
- Bentley D. L., Groudine M. A block to elongation is largely responsible for decreased transcription of c-myc in differentiated HL60 cells. Nature. 1986 Jun 12;321(6071):702–706. doi: 10.1038/321702a0. [DOI] [PubMed] [Google Scholar]
- Bernard O., Cory S., Gerondakis S., Webb E., Adams J. M. Sequence of the murine and human cellular myc oncogenes and two modes of myc transcription resulting from chromosome translocation in B lymphoid tumours. EMBO J. 1983;2(12):2375–2383. doi: 10.1002/j.1460-2075.1983.tb01749.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blanchard J. M., Piechaczyk M., Dani C., Chambard J. C., Franchi A., Pouyssegur J., Jeanteur P. c-myc gene is transcribed at high rate in G0-arrested fibroblasts and is post-transcriptionally regulated in response to growth factors. Nature. 1985 Oct 3;317(6036):443–445. doi: 10.1038/317443a0. [DOI] [PubMed] [Google Scholar]
- Campisi J., Gray H. E., Pardee A. B., Dean M., Sonenshein G. E. Cell-cycle control of c-myc but not c-ras expression is lost following chemical transformation. Cell. 1984 Feb;36(2):241–247. doi: 10.1016/0092-8674(84)90217-4. [DOI] [PubMed] [Google Scholar]
- Caput D., Beutler B., Hartog K., Thayer R., Brown-Shimer S., Cerami A. Identification of a common nucleotide sequence in the 3'-untranslated region of mRNA molecules specifying inflammatory mediators. Proc Natl Acad Sci U S A. 1986 Mar;83(6):1670–1674. doi: 10.1073/pnas.83.6.1670. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chung J., Sinn E., Reed R. R., Leder P. Trans-acting elements modulate expression of the human c-myc gene in Burkitt lymphoma cells. Proc Natl Acad Sci U S A. 1986 Oct;83(20):7918–7922. doi: 10.1073/pnas.83.20.7918. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cole M. D. The myc oncogene: its role in transformation and differentiation. Annu Rev Genet. 1986;20:361–384. doi: 10.1146/annurev.ge.20.120186.002045. [DOI] [PubMed] [Google Scholar]
- Corcoran L. M., Cory S., Adams J. M. Transposition of the immunoglobulin heavy chain enhancer to the myc oncogene in a murine plasmacytoma. Cell. 1985 Jan;40(1):71–79. doi: 10.1016/0092-8674(85)90310-1. [DOI] [PubMed] [Google Scholar]
- Cory S. Activation of cellular oncogenes in hemopoietic cells by chromosome translocation. Adv Cancer Res. 1986;47:189–234. doi: 10.1016/s0065-230x(08)60200-6. [DOI] [PubMed] [Google Scholar]
- Dani C., Blanchard J. M., Piechaczyk M., El Sabouty S., Marty L., Jeanteur P. Extreme instability of myc mRNA in normal and transformed human cells. Proc Natl Acad Sci U S A. 1984 Nov;81(22):7046–7050. doi: 10.1073/pnas.81.22.7046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dani C., Mechti N., Piechaczyk M., Lebleu B., Jeanteur P., Blanchard J. M. Increased rate of degradation of c-myc mRNA in interferon-treated Daudi cells. Proc Natl Acad Sci U S A. 1985 Aug;82(15):4896–4899. doi: 10.1073/pnas.82.15.4896. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dean M., Levine R. A., Campisi J. c-myc regulation during retinoic acid-induced differentiation of F9 cells is posttranscriptional and associated with growth arrest. Mol Cell Biol. 1986 Feb;6(2):518–524. doi: 10.1128/mcb.6.2.518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dean M., Levine R. A., Ran W., Kindy M. S., Sonenshein G. E., Campisi J. Regulation of c-myc transcription and mRNA abundance by serum growth factors and cell contact. J Biol Chem. 1986 Jul 15;261(20):9161–9166. [PubMed] [Google Scholar]
- Dony C., Kessel M., Gruss P. Post-transcriptional control of myc and p53 expression during differentiation of the embryonal carcinoma cell line F9. Nature. 1985 Oct 17;317(6038):636–639. doi: 10.1038/317636a0. [DOI] [PubMed] [Google Scholar]
- Eick D., Bornkamm G. W. Transcriptional arrest within the first exon is a fast control mechanism in c-myc gene expression. Nucleic Acids Res. 1986 Nov 11;14(21):8331–8346. doi: 10.1093/nar/14.21.8331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eick D., Piechaczyk M., Henglein B., Blanchard J. M., Traub B., Kofler E., Wiest S., Lenoir G. M., Bornkamm G. W. Aberrant c-myc RNAs of Burkitt's lymphoma cells have longer half-lives. EMBO J. 1985 Dec 30;4(13B):3717–3725. doi: 10.1002/j.1460-2075.1985.tb04140.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gorski K., Roch J. M., Prentki P., Krisch H. M. The stability of bacteriophage T4 gene 32 mRNA: a 5' leader sequence that can stabilize mRNA transcripts. Cell. 1985 Dec;43(2 Pt 1):461–469. doi: 10.1016/0092-8674(85)90176-x. [DOI] [PubMed] [Google Scholar]
- Graves R. A., Pandey N. B., Chodchoy N., Marzluff W. F. Translation is required for regulation of histone mRNA degradation. Cell. 1987 Feb 27;48(4):615–626. doi: 10.1016/0092-8674(87)90240-6. [DOI] [PubMed] [Google Scholar]
- Greenberg M. E., Ziff E. B. Stimulation of 3T3 cells induces transcription of the c-fos proto-oncogene. Nature. 1984 Oct 4;311(5985):433–438. doi: 10.1038/311433a0. [DOI] [PubMed] [Google Scholar]
- Grosso L. E., Pitot H. C. Transcriptional regulation of c-myc during chemically induced differentiation of HL-60 cultures. Cancer Res. 1985 Feb;45(2):847–850. [PubMed] [Google Scholar]
- Hann S. R., Thompson C. B., Eisenman R. N. c-myc oncogene protein synthesis is independent of the cell cycle in human and avian cells. 1985 Mar 28-Apr 3Nature. 314(6009):366–369. doi: 10.1038/314366a0. [DOI] [PubMed] [Google Scholar]
- Hayday A. C., Gillies S. D., Saito H., Wood C., Wiman K., Hayward W. S., Tonegawa S. Activation of a translocated human c-myc gene by an enhancer in the immunoglobulin heavy-chain locus. 1984 Jan 26-Feb 1Nature. 307(5949):334–340. doi: 10.1038/307334a0. [DOI] [PubMed] [Google Scholar]
- Jonak G. J., Knight E., Jr Selective reduction of c-myc mRNA in Daudi cells by human beta interferon. Proc Natl Acad Sci U S A. 1984 Mar;81(6):1747–1750. doi: 10.1073/pnas.81.6.1747. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keath E. J., Kelekar A., Cole M. D. Transcriptional activation of the translocated c-myc oncogene in mouse plasmacytomas: similar RNA levels in tumor and proliferating normal cells. Cell. 1984 Jun;37(2):521–528. doi: 10.1016/0092-8674(84)90382-9. [DOI] [PubMed] [Google Scholar]
- Kelekar A., Cole M. D. Tumorigenicity of fibroblast lines expressing the adenovirus E1a, cellular p53, or normal c-myc genes. Mol Cell Biol. 1986 Jan;6(1):7–14. doi: 10.1128/mcb.6.1.7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kelly K., Cochran B. H., Stiles C. D., Leder P. Cell-specific regulation of the c-myc gene by lymphocyte mitogens and platelet-derived growth factor. Cell. 1983 Dec;35(3 Pt 2):603–610. doi: 10.1016/0092-8674(83)90092-2. [DOI] [PubMed] [Google Scholar]
- Knight E., Jr, Anton E. D., Fahey D., Friedland B. K., Jonak G. J. Interferon regulates c-myc gene expression in Daudi cells at the post-transcriptional level. Proc Natl Acad Sci U S A. 1985 Feb;82(4):1151–1154. doi: 10.1073/pnas.82.4.1151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lachman H. M., Skoultchi A. I. Expression of c-myc changes during differentiation of mouse erythroleukaemia cells. Nature. 1984 Aug 16;310(5978):592–594. doi: 10.1038/310592a0. [DOI] [PubMed] [Google Scholar]
- Leder P., Battey J., Lenoir G., Moulding C., Murphy W., Potter H., Stewart T., Taub R. Translocations among antibody genes in human cancer. Science. 1983 Nov 18;222(4625):765–771. doi: 10.1126/science.6356357. [DOI] [PubMed] [Google Scholar]
- Levine R. A., McCormack J. E., Buckler A., Sonenshein G. E. Transcriptional and posttranscriptional control of c-myc gene expression in WEHI 231 cells. Mol Cell Biol. 1986 Nov;6(11):4112–4116. doi: 10.1128/mcb.6.11.4112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Linial M., Gunderson N., Groudine M. Enhanced transcription of c-myc in bursal lymphoma cells requires continuous protein synthesis. Science. 1985 Dec 6;230(4730):1126–1132. doi: 10.1126/science.2999973. [DOI] [PubMed] [Google Scholar]
- Lipp M., Schilling R., Wiest S., Laux G., Bornkamm G. W. Target sequences for cis-acting regulation within the dual promoter of the human c-myc gene. Mol Cell Biol. 1987 Apr;7(4):1393–1400. doi: 10.1128/mcb.7.4.1393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mechti N., Piechaczyk M., Blanchard J. M., Marty L., Bonnieu A., Jeanteur P., Lebleu B. Transcriptional and post-transcriptional regulation of c-myc expression during the differentiation of murine erythroleukemia Friend cells. Nucleic Acids Res. 1986 Dec 22;14(24):9653–9666. doi: 10.1093/nar/14.24.9653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Persson H., Gray H. E., Godeau F. Growth-dependent synthesis of c-myc-encoded proteins: early stimulation by serum factors in synchronized mouse 3T3 cells. Mol Cell Biol. 1985 Nov;5(11):2903–2912. doi: 10.1128/mcb.5.11.2903. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piechaczyk M., Yang J. Q., Blanchard J. M., Jeanteur P., Marcu K. B. Posttranscriptional mechanisms are responsible for accumulation of truncated c-myc RNAs in murine plasma cell tumors. Cell. 1985 Sep;42(2):589–597. doi: 10.1016/0092-8674(85)90116-3. [DOI] [PubMed] [Google Scholar]
- Prehn J., Mercola M., Calame K. Translocation affects normal c-myc promoter usage and activates fifteen cryptic c-myc transcription starts in plasmacytoma M603. Nucleic Acids Res. 1984 Dec 11;12(23):8987–9007. doi: 10.1093/nar/12.23.8987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rabbitts P. H., Forster A., Stinson M. A., Rabbitts T. H. Truncation of exon 1 from the c-myc gene results in prolonged c-myc mRNa stability. EMBO J. 1985 Dec 30;4(13B):3727–3733. doi: 10.1002/j.1460-2075.1985.tb04141.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reitsma P. H., Rothberg P. G., Astrin S. M., Trial J., Bar-Shavit Z., Hall A., Teitelbaum S. L., Kahn A. J. Regulation of myc gene expression in HL-60 leukaemia cells by a vitamin D metabolite. Nature. 1983 Dec 1;306(5942):492–494. doi: 10.1038/306492a0. [DOI] [PubMed] [Google Scholar]
- Remmers E. F., Yang J. Q., Marcu K. B. A negative transcriptional control element located upstream of the murine c-myc gene. EMBO J. 1986 May;5(5):899–904. doi: 10.1002/j.1460-2075.1986.tb04301.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ross J., Kobs G. H4 histone messenger RNA decay in cell-free extracts initiates at or near the 3' terminus and proceeds 3' to 5'. J Mol Biol. 1986 Apr 20;188(4):579–593. doi: 10.1016/s0022-2836(86)80008-0. [DOI] [PubMed] [Google Scholar]
- Saito H., Hayday A. C., Wiman K., Hayward W. S., Tonegawa S. Activation of the c-myc gene by translocation: a model for translational control. Proc Natl Acad Sci U S A. 1983 Dec;80(24):7476–7480. doi: 10.1073/pnas.80.24.7476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shaw G., Kamen R. A conserved AU sequence from the 3' untranslated region of GM-CSF mRNA mediates selective mRNA degradation. Cell. 1986 Aug 29;46(5):659–667. doi: 10.1016/0092-8674(86)90341-7. [DOI] [PubMed] [Google Scholar]
- Stanton L. W., Watt R., Marcu K. B. Translocation, breakage and truncated transcripts of c-myc oncogene in murine plasmacytomas. Nature. 1983 Jun 2;303(5916):401–406. doi: 10.1038/303401a0. [DOI] [PubMed] [Google Scholar]
- Swartwout S. G., Preisler H., Guan W. D., Kinniburgh A. J. Relatively stable population of c-myc RNA that lacks long poly(A). Mol Cell Biol. 1987 Jun;7(6):2052–2058. doi: 10.1128/mcb.7.6.2052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taub R., Kelly K., Battey J., Latt S., Lenoir G. M., Tantravahi U., Tu Z., Leder P. A novel alteration in the structure of an activated c-myc gene in a variant t(2;8) Burkitt lymphoma. Cell. 1984 Jun;37(2):511–520. doi: 10.1016/0092-8674(84)90381-7. [DOI] [PubMed] [Google Scholar]
- Thompson C. B., Challoner P. B., Neiman P. E., Groudine M. Levels of c-myc oncogene mRNA are invariant throughout the cell cycle. 1985 Mar 28-Apr 3Nature. 314(6009):363–366. doi: 10.1038/314363a0. [DOI] [PubMed] [Google Scholar]
- Thorens B., Mermod J. J., Vassalli P. Phagocytosis and inflammatory stimuli induce GM-CSF mRNA in macrophages through posttranscriptional regulation. Cell. 1987 Feb 27;48(4):671–679. doi: 10.1016/0092-8674(87)90245-5. [DOI] [PubMed] [Google Scholar]
- Westin E. H., Wong-Staal F., Gelmann E. P., Dalla-Favera R., Papas T. S., Lautenberger J. A., Eva A., Reddy E. P., Tronick S. R., Aaronson S. A. Expression of cellular homologues of retroviral onc genes in human hematopoietic cells. Proc Natl Acad Sci U S A. 1982 Apr;79(8):2490–2494. doi: 10.1073/pnas.79.8.2490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wigler M., Pellicer A., Silverstein S., Axel R., Urlaub G., Chasin L. DNA-mediated transfer of the adenine phosphoribosyltransferase locus into mammalian cells. Proc Natl Acad Sci U S A. 1979 Mar;76(3):1373–1376. doi: 10.1073/pnas.76.3.1373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang J. Q., Remmers E. F., Marcu K. B. The first exon of the c-myc proto-oncogene contains a novel positive control element. EMBO J. 1986 Dec 20;5(13):3553–3562. doi: 10.1002/j.1460-2075.1986.tb04682.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zinn K., DiMaio D., Maniatis T. Identification of two distinct regulatory regions adjacent to the human beta-interferon gene. Cell. 1983 Oct;34(3):865–879. doi: 10.1016/0092-8674(83)90544-5. [DOI] [PubMed] [Google Scholar]