1 Introduction
In statistical literature, interval censoring usually represents a sampling scheme or an incomplete data structure. By interval censoring, we mean that a random variable of interest is known only to lie within an interval instead of being observed exactly. For applications in survival analysis, the random variable is the time to some event such as death, a disease recurrence or a distant metastasis. Many clinical trials and longitudinal studies may generate interval-censored data[1,2,3]. One common example occurs in medical or health studies that entail periodic follow-up. In this situation, an individual due for the pre-scheduled observations for a clinically observable change in disease or health status may miss some observations and return with a changed status. Accordingly, we only know that the true event time is greater than the last observation time at which the change has not occurred and less than or equal to the first observation time at which the change has been observed to occur, thus giving an interval which contains the real (but unobserved) time of occurrence of the change.
Another example of interval-censored data arises in the acquired immune deficiency syndrome (AIDS) trials[4] that, for example, are interested in times to AIDS for human immunodeficiency virus (HIV) infected subjects. In these cases, the determination of AIDS onset is usually based on blood testing, which can be performed obviously only periodically but not continuously. In consequence, only interval-censored data may be available for AIDS diagnosis times. A similar case is for studies on HIV infection times. If a patient is HIV positive at the beginning of a study, then the HIV infection time is usually determined by a retrospective analysis of his or her medical history. Therefore, we are only able to obtain an interval given by the last HIV negative test date and the first HIV positive test date for the HIV infection time.
An important special case of interval-censored data is the so called current status data[5,6]. This type of censoring means that each subject is observed only once for the status of the occurrence of the event of interest. In other words, we do not directly observe the survival endpoint but instead, we only know the observation time and whether or not the event of interest has occurred at the time. In consequence, the survival time is either left- or right-censored. One such example is the data arising from cross-sectional studies on survival events[7]. Another example is given by the tumourigenicity study and in this situation, the time to tumor onset is usually of interest, but not directly observable[8]. As a matter of fact, we only have the exact measurement of the observation time, which is often the death or sacrifice time of the subject. Note that for the first example, current status data occur due to the study design, while for the second case, they are often observed because of our inability of measuring the variable directly and exactly. Sometimes we also refer current status data to as case I interval-censored data and the general case as case II interval-censored data[9].
We now establish some notation for interval censoring. Let T denote the survival time of interest. When T is interval-censored we use I = (L, R] to denote the interval containing T. Using this notation, we see that current status data correspond to the situation where either L = 0 or R = ∞. Interval censoring also contains right censoring and left censoring as special cases and if R = ∞, we have a right-censored observation, while if L = 0 we obtain a left-censored observation.
To this point, the survival time has been defined as the time between a fixed starting time point, usually zero, and the event time. One can apply a more general framework that defines the survival time as the time between two related events whose occurrence times are random variables and both could suffer censoring. If there is right- or interval-censoring on both occurrence times, the resulting data are commonly referred to as doubly censored data[4,10]. An example of such complicated type of data is provided by the AIDS studies discussed above when the variable of interest is AIDS incubation time[4], the time from HIV infection to AIDS diagnosis, with both the HIV infection time and the AIDS diagnosis time being right- or interval-censored.
In applications, interval-censored data can be easily confused with grouped survival data. There is actually a fundamental difference between these two data structures although both usually appear in the form of intervals. The grouped survival data can be seen as a special case of interval-censored data and commonly mean that the intervals for any two subjects either are completely identical or have no overlapping. In contrast, the intervals for interval-censored data may overlap in any way. As a consequence of this structure difference, statistical methods for grouped survival data are much more straightforward than those for interval-censored data.
To illustrate the concepts described above, consider a set of well-known case II interval-censored data on breast cancer, which can be found in Finkelstein and Wolfe[11] and Sun[3] among others. The data consist of 94 early breast cancer patients treated at the Joint Center for Radiation Therapy in Boston between 1976 and 1980. For their treatments, the patients were given either radiation therapy alone (RT, 46 patients) or radiation therapy plus adjuvant chemotherapy (RCT, 48 patients). Each patient was supposed to have clinic visits every 4 to 6 months to be examined for cosmetic appearance such as breast retraction. However, actual visit times differ from patient to patient and the times between the visits also vary. As a consequence, only interval-censored data are observed for breast retraction times. Specifically, among the 94 patients, 38 of them did not experience breast retraction during the study, giving right-censored observations for the breast retraction times. For other patients, intervals such as (25, 37] were observed for their breast retraction times. Here the interval (25, 37] means that the patient had a clinic visit at month 25 and no breast retraction was detected at the visit, while at the next visit at month 37, breast retraction was found to be present already. There are 5 patients for whom the breast retraction was detected at their first clinical visits, giving the observed intervals with the left end points being zero or left-censored observations. One objective of the study was to compare the two treatments through their effects on breast retraction and more discussion on this will be given below.
The rest of this paper will follow the style of Sun[12] with the focus on medical applications, some basic issues and available methods for them, and the recent development in the literature for interval-censored data. The readers are referred to Sun[3] for more complete references. In the following, we will begin in Section 2 with describing a fundamental and important assumption behind most methodologies dealing with interval censoring: noninformative interval censoring. It basically states that the censoring mechanism does not contribute to the likelihood function. For the analysis of interval-censored data, we will first discuss nonparametric estimation of a survival function as well as a hazard function in Section 3. In Section 4, the methods for comparing several survival functions will be reviewed. Regression analysis of interval-censored data under various semiparametric models is then considered in Section 5. Section 6 briefly covers a few other topics including parametric approaches, interval-censored data with truncation, multivariate interval-censored data, competing risks interval-censored data and informative interval censoring. Finally, some concluding remarks are given in Section 7.
2 Noninformative Interval Censoring
In the case of right-censored failure time data, a common assumption is that the censoring time is independent of the survival time of interest marginally or conditionally given external covariates. It is clear that this assumption cannot be directly generalized to interval censoring since in this case with the notation defined above, the endpoints of the interval, L and R, together with the survival time T, have a natural relationship L < T ≤ R. Instead, for interval-censored data, the noninformative interval censoring assumption specified as
| (1) |
is usually used[3,13]. It essentially says that, except for the fact that T lies between l and r which are the realizations of L and R, the interval (L, R] (or equivalently its endpoints L and R) does not provide any extra information for T. In other words, the probabilistic behavior of T remains the same except that the original sample space T ≥ 0 is now reduced to l = L < T ≤ R = r.
In the existence of covariates denoted by Z, one can relax the assumption (1) to
| (2) |
All methods discussed below will be based on the assumption (1) or (2) unless specified otherwise. In practice, one question of interest is the conditions under which the assumption (1) or (2) holds and for this, several authors described the constant-sum condition[13,14,15]. Some discussion will be given in Section 6 for situations where the assumption (1) or (2) does not hold.
Instead of the assumption (1) or (2), another approach to define the noninformative interval censoring is through or using the stochastic process that yields interval censoring[16]. For example, one can assume that there exists a sequence of observation times or an observation process and this process is independent of the survival time T[9]. Of course, in practice, one could estimate and/or examine the observation process to check the validity of the assumption [16]. Since this alternative approach requires more advanced mathematical tools, it will not be discussed in depth here.
3 Nonparametric Estimation
In medical and health studies, estimation of the cumulative distribution function (CDF) or the survival function of a survival variable is often the first task in statistical analysis. Let F(t) = P(T ≤ t) be the CDF of T and S(t) = 1 − F(t), the survival function. It is apparent that estimating one is equivalent to estimating the other and therefore, one only needs to discuss one of them. Here we will focus on estimation of S(t) because of its popularity in survival analysis.
By using the notation defined above, interval-censored data can usually be represented by , where Ii = (Li, Ri] is the interval known or observed to contain the unobserved T associated with the ith subject and n denotes the sample size. Let denote the unique ordered elements of , i.e., 0 = t0 < t1 < … < tm < tm+1 =∞, αij the indicator of the event (tj−1, tj] ⊆ Ii, and pj = S(tj−1) − S(tj). Then under assumption (1), the likelihood function for p = (p1, …, pm+1)′ is proportional to
and the problem of finding the nonparametric maximum likelihood estimator (NPMLE) of S becomes that of maximizing LS(p) under the constraint that and pj ≥ 0 (j = 1, …, m + 1)[9,17,18,19]. Obviously, the likelihood function LS depends on S only through the values . Thus the NPMLE of S, which we shall denote by Ŝ can be uniquely determined only over the observed intervals (tj−1, tj] and the behavior of S within these intervals will be unknown. Conventionally, however, Ŝ(t) is often taken to be a right continuous step function. That is, Ŝ(t) = Ŝ(tj−1) when tj−1 ≤ t < tj.
Several methods have been proposed for maximizing LS(p) with respect to p. Here we will briefly describe three algorithms commonly used for case II interval-censored data. The first and simplest one is the self-consistency algorithm given in Turnbull[19] and was developed based on the equation
where I(·) is the indicator function and Ti is the unobserved survival time associated with subject i. It can be easily seen that the algorithm is essentially an application of the EM algorithm[20]. Computationally, for obtaining Ŝ, we may iterate the equation
until convergence. This approach is easy to implement (available in SPLUS and R, see Section 7 for some details) but has a slow convergence rate.
The second approach that one can apply to maximize LS(p) is the iterative convex minorant (ICM) algorithm introduced by Groeneboom and Wellner[9] and improved by Jongbloed[21], which converges faster than the self-consistency algorithm. In fact, it can be seen as an optimized version of the well known pool-adjacent-violator algorithm[22]. Due to the complexity of this algorithm we will not describe the computational details here but refer readers to [3] and [9]. The third commonly used algorithm is the EM-ICM algorithm presented in Wellner and Zhan[23]. As suggested by the name, it is a hybrid (and the fastest) algorithm which combines the first two approaches. All the above algorithms are iterative and in fact, there is no closed form for the NPMLE of S.
It should be noted that the algorithms discussed above may yield multiple solutions[17]. In other words, the NPMLE may not be unique. One sufficient condition for the uniqueness of the NPMLE is that the log likelihood is strictly concave. Furthermore, it is possible that the solutions derived from these algorithms may not be the NPMLE. To check this, one may apply the so-called Kuhn-Tucker conditions[17], which gives the sufficient and necessary condition for a solution to be the NPMLE. An alternative is to use the fact that an estimate Ŝ is the NPMLE if and only if [24].
For current status data, however, a closed form of the NPMLE can be derived. Remember that in this case, each subject is observed only once and the only available information is if the survival event has occurred at the observation time point. Let the tj’s denote the ordered observation times as above and Qj the set of subjects who are observed at tj, j = 1, …, m. Define dj = Σi∈Qj 1(Ti ≤ tj) and let nj denote the number of elements in Qj. Then the NPMLE of S can be shown[3] to be equal to the isotonic regression of {d1/n1, …, dm/nm} with weights {n1, …, nm}. Using the max-min formula for the isotonic regression[25], we have
Other methods for computing the NPMLE that are not discussed in details here include Vandal et al.[26], who connected graph theoretic representation and the NPMLE, and Sen and Banerjee[27], who used a pseudo-likelihood approach based on Poisson processes.
The discussion above shows a clear difference between right censoring and interval censoring as the NPMLE of a survival function based on right-censored data is given by the closed-form Kaplan-Meier estimate. More importantly, the NPMLE for the two situations have quite different asymptotic behavior. Before discussing this asymptotic difference, it is worth to remark that interval censoring induces many challenging problems in large sample studies. One is that unlike with right-censored data, the use of the counting process technique is quite difficult in the case of interval-censored data and as a consequence, the martingale theory cannot be applied. For investigating asymptotic properties of various procedures, instead, one has to rely on complicated empirical process arguments.
It has been shown that for interval-censored data, Ŝ is strongly consistent[9,28]. However, Ŝ(t) converges only in the rate of n1/3 and furthermore its limiting distribution is nonnormal[9,29]. This clearly differs from the usual n1/2-convergence rate and the limiting normal distribution for the Kaplan-Meier estimate. Heuristically, this is because interval censoring in general provides much less information than right censoring. On the other hand, as with right-censored data, the linear functionals of Ŝ still have asymptotic normal distributions with the n1/2 convergence rate[29,30].
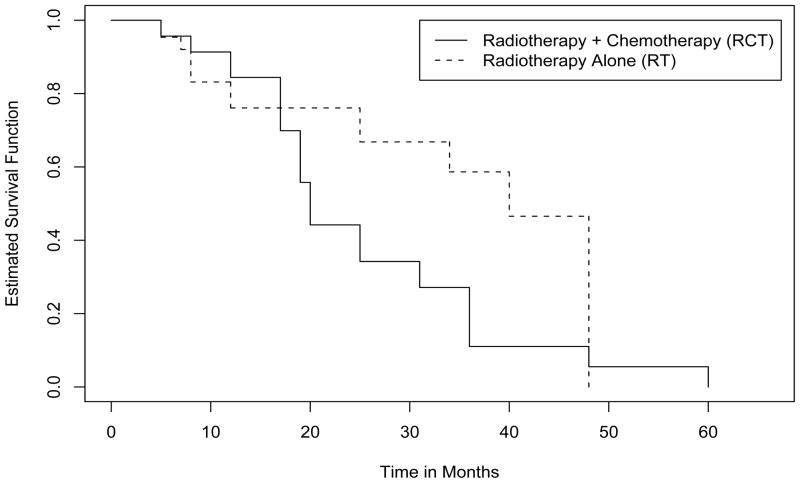
As an illustration, we apply the algorithms described above to the breast cancer data discussed in Section 1. Note that here the survival time of interest is the time to breast retraction for which only case II interval-censored data are available. For the analysis, we applied the algorithms to the data from the two treatment groups separately and the obtained NPMLEs are presented in Figure 1. Note that here the three algorithms discussed above yielded the same estimates for both groups. Figure 1 suggests that although there is no difference at the beginning, the patients in the RCT group seem to develop breast retraction more quickly than those in the RT group overall. Some formal comparison results will be given below.
Figure 1.
NPMLE of survival functions of time to breast retraction.
To conclude this section, we briefly discuss estimation of the hazard function based on interval-censored data. For this, as with right-censored data, the estimated hazard function may provide some insight information about the shape of a survival function although it may not be of primary interest in many applications. A simple and direct estimate of the hazard function is clearly the empirical estimate but one may not want to use it due to its roughness and the interpretation issue. In most cases, therefore, certain smoothing techniques are needed for deriving more descriptive estimators. Among others, Kooperberg and Stone[31] and Rosenberg[32] described some spline-based estimators and Bebchuk and Betensky[33] and Betensky et al.[34] gave a multiple imputation approach and a local likelihood approach, respectively.
4 Comparison of Survival Functions
Comparison of different treatments or survival functions is another primary objective in most medical or clinical studies. To formalize the problem, suppose there are K treatment arms in a clinical study and let S(k)(t) denote the survival function of the kth arm with k = 1, …, K. Then the problem becomes testing the null hypothesis
It is clear that if some relationship among or regression models for S(k)’s are known or can be assumed, one should make use of them to derive, say, parametric or score tests for H0. On the other hand or in general, one probably prefers some nonparametric or distribution-free test procedures.
With right censoring, many nonparametric test procedures have been developed and most of them can be classified into two categories: rank-based tests and survival-based tests. The fundamental difference between them is that the former relies on the differences between the estimated hazard functions while the latter bases the comparison on the differences between the estimated survival functions. Among them, the log-rank test is perhaps the most widely used method. A few of these rank-based or survival-based test procedures have been generalized to the case of interval-censored data and will be described below. However, we caution that most of these generalized methods require strong assumptions about the censoring mechanism or distribution, or can only be applied to the two-sample situation.
Here we review three procedures that are direct generalizations of the corresponding methods for right-censored data. First we discuss a rank-based approach[35] that is a direct generalization of the log-rank test. Consider a survival study consisting of n independent subjects and giving rise to interval-censored data. Let the Ti’s, Ii’s, tj’s and αij’s be defined as in Section 3 and Ŝ0 denote the NPMLE of the S(k)’s under H0. For subject i, define δi = 0 if the observation on Ti is right-censored and 1 otherwise and ρij = I(δi = 0, Li ≥ tj), which is equal to 1 if Ti is right-censored and subject i is still at risk at tj−.
Also for each j = 1, …, m, define
and
They can be regarded as the estimates of the total observed failure and risk numbers, respectively, at time tj under H0. Similarly one can define djk and njk, the estimates of the observed failure and risk numbers, respectively, from subjects in arm k, as dj and nj with the summation over i replaced by the summation over only subjects in arm k, j = 1, …, m, k = 1, …, K.
To test = (H0, following the log-rank test, one can define a test statistic as U0 = U1, …, UK)′ with
It can be easily shown that if right-censored data are available, the statistic U reduces to the log-rank test statistic. For estimation of the covariance matrix of U, Zhao and Sun[35] give a multiple imputation approach which will not be discussed here. A major advantage of this generalized log-rank procedure is its simplicity in both its implementation and interpretation. Of course, to apply it, one has to determine Ŝ0. Recently Kim et al. [36] suggested a modification to the procedure that does not require Ŝ0 by assuming that Ti follows the uniform distribution over the observed interval.
Instead of the generalizations of the rank-based tests for right-censored data such as the one described above, there exist a few generalizations of the survival-based tests. For simplicity, assume that K = 2 and let Ŝ(1) and Ŝ(2) denote the NPMLE of S(1) and S(2) separately. Then to test H0, a natural statistic is given by
| (3) |
where τ denotes the longest follow-up time and W is a weight function. This statistic measures the integrated weighted survival difference and by taking W(t) = 1, it gives the difference of the estimated sample means. Among others, Petroni and Wolfe[37] and Fang et al.[38] discussed the statistics given in (3) for discrete and continuous survival times, respectively. Especially, the latter showed that the standardized statistic has asymptotically a normal distribution with mean zero under H0 if the observed data do not include exact failure times. Note that as an alternative to the integrated weighted differences, for testing H0 based on the survival difference, one could base the comparison on the supremum or absolute difference among the estimated survival functions[39].
The third method[40] that we will briefly discuss is a generalization of the method given in Peto and Peto[41] and it bases testing H0 on the statistic
Here zi is a vector of group indicators associated with subject i and ξ is a known function over (0,1). A limitation of the statistic given above is that it requires that the observed data are purely interval-censored or no exact failure time is observed. To relax this, Zhao et al.[42] generalized it to situations where one may observe both exact failure times and interval-censored failure times.
If current status data are available, it is apparent that one can apply the procedures discussed above to test H0. Of course, one can also apply some specially developed methods that are able to take into account the special structure of the data[6,43]. Here we will discuss one of such methods that is different from those discussed above for general interval-censored data. Consider a study that yields current status data {(Ci, Ni(Ci) = I(Ti ≤ Ci), zi) ; i = 1, …, n} using the notation introduced above, where Ci denotes the observation time on subject i. Assume that the Ci’s may follow different distributions for subjects in different arms and their hazard functions are given by the proportional hazards model (4) described in the next section. More comments on this are given below. To test H0, Sun[44] suggested to use the statistic
where z̄ denotes the sample mean of the zi’s, β̂ the maximum likelihood estimate (MLE) of the regression parameters in model (4), and
with Ñi(t) =I(t ≥ Ci) and . Note that a major advantage of this statistic is that it allows different distributions for observation times in different treatment arms, while most of the procedures for general interval-censored data do not as mentioned above. We remark that the assumed model (4) for the Ci’s is not too restrictive as it is commonly used and more importantly can be easily checked because of the availability of the complete data here.
We will conclude this section by applying the first two test procedures described above to the breast cancer data discussed in Section 1. Let S(1)(t) and S(2)(t) denote the underlying survival functions of the times to breast retraction for the patients in the RT and RCT groups, respectively. Then the comparison of the two treatments is equivalent to testing the hypothesis: S(1)(t) = S(2)(t) for all t. The application of the first procedure, the generalized log-rank test, gave U1 = −9.43 with the estimated standard error of 3.42 based on the multiple imputation approach. This corresponds to a p-value of 0.006 and suggests that as shown in Figure 1, the patients in the RT group had a significantly lower breast retraction rate than those in the RCT group. To apply the second, survival-based procedure given in (3), we assumed that the breast retraction could occur only at six-month time points and took W(t) = 1. The test statistic in (3) then yielded the value 8.97 with the estimated standard error being 2.36. This again indicates that the use of adjuvant chemotherapy significantly increased the breast retraction risk.
5 Regression Analysis
Regression analysis is usually performed if one is interested in quantifying the effect of some covariates on the survival time of interest or predicting survival probabilities for new individuals. Of course, the first step for regression analysis is to specify an appropriate regression model. In this section, we shall discuss a few commonly used semiparametric or parametric models and some corresponding inference procedures for interval-censored data. Unlike most methods developed for right-censored data, estimating regression parameters under interval censoring usually involves estimation of both the parametric and the nonparametric parts. In other words, for interval-censored data, one has to deal with estimation of some unknown baseline functions in order to estimate regression parameters.
The proportional hazards model[45] has been the most commonly used semiparametric regression model for survival analysis for about the past three decades. It postulates
| (4) |
for the hazard function of the survival time T of interest given covariates Z = z, where λ0(t) denotes the unknown baseline hazard function (the hazard function for subjects with Z = 0) and β the vector of unknown regression parameters. For situations of current status data, Huang[46] provided a complete study of this model and an ICM-type algorithm for estimation of unknown parameters. A Newton-Raphson algorithm is also available for the estimation[3]. In the case of general or case II interval-censored data, Finkelstein[1] in her seminal paper proposed to apply the Newton-Raphson algorithm to determine the MLE of β and the baseline cumulative hazard function together. The approach actually simplifies the situation to a finite-dimensional parametric estimation problem. Alternatively, one can apply the marginal likelihood approach and the stochastic approximation algorithm given in Satten[47], the Markov Chain Monte Carlo EM algorithm developed by Goggins et al.[48], or the multiple imputation-based method presented in Pan[49]. It should be noted that all of these methods require a great deal of computational efforts. The readers are referred to Huang and Wellner[50] for a rigorous study of the efficiency and asymptotic properties of the MLE based on case II interval-censored data. A few generalizations of model (4) have been proposed for interval-censored data[3] and among them, Kim and Jhun[51] recently studied the situation where there exists a group of cured subjects or the subjects who may never experience the survival event of interest.
Another commonly used semiparametric model in survival analysis is the proportional odds model. This model can be expressed as
| (5) |
with respect to the CDF F(t|z) of the survival time T given Z = z, where h(t) is an unknown monotone-increasing function, also referred to as the baseline log odds. For inference about this model based on current status data, both Huang[52] and Rossini and Tsiatis[53] considered the maximum likelihood approach but used different approximations for the baseline log odds function. In the case of case II interval-censored data, one may apply the sieve estimation procedures discussed in Huang and Rossini[54] and Shen[55]. The former employed a piecewise linear function, while the latter used a monotone spline to approximate the baseline log odds function. An alternative to these methods is that given by Rabinowitz et al.[56], which does not require estimation of the baseline log odds function but assumed that subjects are still under follow-up even after the failure event of interest has occurred. It is apparent that this continuous follow-up may not be realistic in practice. Huang and Wellner[50] provided some discussion on the asymptotic properties of the MLE for model (5).
As models (4) and (5), the accelerated failure time model is also widely used in survival analysis. It assumes that the survival time T and the covariate Z have the following relationship
| (6) |
where ε is an error term whose distribution is usually unspecified. Suppose that general case II interval-censored data are available and for inference about model (6), among others, Rabinowitz et al.[57] and Betensky et al.[58] developed some procedures. The former employed a class of score statistics, while the latter used the estimating equation idea by treating observation times from the same subject as if they were from different subjects. Both methods require estimation of the distribution of the error term ε. In contrast, Li and Pu[59] gave an inference procedure that does not require estimation of the distribution of ε. The approach employs a rank-based estimating equation and may not be efficient. More recently, Tian and Cai[60] and Xue et al.[61] also investigated the inference problem regarding model (6).
In addition to the models discussed above, another attractive semiparametric regression model is the additive hazards model given by
| (7) |
with respect to the hazard function of T given Z = z. It specifies that the effects of the covariates are additive rather than multiplicative as in model (4). To estimate the regression parameter β based on current status data, Lin et al.[62] proposed a simple estimating equation approach that does not require estimation of the baseline hazard function λ0(t). Martinussen and Scheike[63] studies the same problem and provided an approach that can be more efficient than that given in Lin et al.[62]. However, the latter involves estimation of the baseline hazard function and can be much more complicated. For inference about model (7) based on case II interval-censored data, both Huang and Wellner[50] and Zeng et al.[64] investigated the maximum likelihood approach. Furthermore, Chen and Sun[65] and Zhu et al.[66] recently gave a multiple imputation-based procedure and a transformation approach, respectively.
The four semiparametric models described above are all specific models in terms of the functional form of the effects of covariates. Sometimes one may prefer a model that gives more flexibility. One such model is the linear transformation model that specifies the relationship between the failure time T and the covariate Z as
| (8) |
where h :
 →
→
 (
(
 denotes the real line and
denotes the real line and
 the positive half real line) is an unknown strictly increasing function and the distribution of ε is assumed to be known. Model (8) gives different models depending on the specification of the distribution of ε and especially, it includes the proportional hazards model (4) and the proportional odds model (5) as special cases. Among others, Sun and Sun[67], Younes and Lachin[68] and Zhang et al.[69] considered the inference about model (8) when one only observes interval-censored data.
the positive half real line) is an unknown strictly increasing function and the distribution of ε is assumed to be known. Model (8) gives different models depending on the specification of the distribution of ε and especially, it includes the proportional hazards model (4) and the proportional odds model (5) as special cases. Among others, Sun and Sun[67], Younes and Lachin[68] and Zhang et al.[69] considered the inference about model (8) when one only observes interval-censored data.
Several other models or generalizations of the models discussed above are also available for regression analysis of interval-censored failure time data. For example, one may apply the partial linear model given by[70]
a generalization of the accelerated failure time model (6). Here both Z1 and Z2 are covariates which may or may not overlap, g is an unknown smooth function, and ε follows a prespecified distribution. Shiboski[71] presented some generalized additive models and Zhang and Davidian[72] recently gave a group of smooth semiparametric regression models.
In addition to the semiparametric models discussed above, one particular family of parametric models, piecewise exponential models, is worth mentioning. To introduce the models, consider model (4) or (7) and instead of assuming that the baseline hazard function λ0(t) in (4) or (7) is an arbitrary nonnegative function, the piecewise exponential model assumes that λ0(t) is constant over pre-specified intervals. Specifically, let 0 = s0 < s1 < … < sK < sK+1 = ∞ be a partition of the positive real line and assume that λ0(t) = λk if sk−1 < t ≤ sk, k = 1, …, K + 1, where {λ1, …, λK+1} are a group of nonnegative unknown parameters. Obviously the model defined above can be seen as an approximation to model (4) or (7). Although the piecewise exponential model may be less flexible (parametric) than the semiparametric model, it is simple and one can readily apply the maximum likelihood approach for parameter estimation. Of course, for the use of the piecewise exponential model, one needs to specify the partition, which may not be straightforward sometimes.
To perform regression analysis on the breast cancer data discussed in Section 1, define Z = 0 for subjects in the RT group and Z = 1 otherwise and assume that the time to breast retraction follows the proportional hazards model (4). Then the parameter β represents the group or treatment difference in terms of the hazard rate. The application of the maximum likelihood estimation approach discussed in Sun[3] gave the estimate of β as 0.800 with the estimated standard error equal to 0.290. The Wald test for testing β = 0 then yielded the p-value of 0.006. As an alternative, by assuming the piecewise exponential proportional hazards model with the partition points at 15.5, 26.5 and 46.5 months (these partition points are chosen such that each interval contains roughly the same number of observed time points), we obtained the maximum likelihood estimate of β as 0.829 with the estimated standard error of 0.301. This gives the same p-value as above for testing no treatment effect. The results here give similar conclusions to those obtained in the previous sections and indicate that the patients in the RCT group had the 2.2 times higher hazard to develop breast retraction than those in the RC group. We remark that in reality, in addition to the analysis described above, one may also want to perform some model diagnosis or checking[3], or to fit other models to the data.
6 Miscellaneous Topics
In the previous sections, we only discussed several basic issues in the analysis of interval-censored failure time data and there exist a number of other issues that were not touched. In this section, we will briefly discuss some of them including multivariate interval-censored data, doubly censored data, competing risks analysis of interval-censored data, and informatively interval-censored data as well as interval-censored data with truncation and parametric procedures.
Multivariate interval-censored data arise if a survival study involves several related survival variables of interest and each of them suffers interval censoring. It is apparent that in this case, one needs different inference procedures than those discussed above and one key and important feature of these different procedures is that they need to take into account the correlation among the survival variables. In addition to the basic issues discussed before, a new and unique issue for multivariate data is to make inference about the association between the survival variables. For this and in general, one of the tools that are commonly used for the analysis of multivariate interval-censored data but not univariate data is the copula model, which provides a very flexible way to model the joint survival function. For the analysis of multivariate interval-censored data, in addition to the methods described in Sun[3], some of the more recently developed approaches include those given in Chen et al.[73], Wang et al.[74] and Zhang et al.[75]. The first considered regression analysis of general multivariate interval-censored data by using model (5), while the last two presented efficient estimation procedures for fitting bivariate current status data to models (4) and (5), respectively. Also Cook et al.[76] applied a multi-state model for bivariate interval-censored data and Komàrek and Lesaffre[77] gave a Bayesian approach for inference about model (6).
One of the early work on the analysis of doubly censored failure time data is given by the seminal paper De Gruttola and Lagakos[4]. In the paper, they proposed a self-consistency algorithm for estimating the distribution of the survival variable of interest. Following their work, many authors considered various issues related to the analysis of doubly censored data and in addition to the review article Sun[78], Sun[3] devoted one chapter for the analysis of doubly censored data. More recently, Komàrek and Lesaffre[79] discussed the Bayesian approach for fitting the doubly censored data to model (6) and Kim[80] considered the use of the frailty model approach for the analysis.
Competing risks analysis is needed when the failure on an individual may be one of several distinct failure types. For example, death of a cancer patient may be classified as disease-related or non-disease-related. In the case of current status data, Groeneboom et al.[81,82] recently studied the asymptotic properties of the NPMLE of the sub-distribution functions.
As mentioned above, all methods discussed so far require the assumption (1) or (2). That is, the interval censoring involved is noninformative about the survival variable or event of interest. Several inference procedures have been developed in the literature for situations where the censoring may be informative[3]. For this, a common way is to jointly model the survival variable and the variables representing interval censoring by, for example, using the latent variable approach[83,84]. Park et al.[85] gave a different approach assuming the presence of a mixture of independent and dependent censoring.
It is well-known that truncation may occur in survival studies and especially may occur together with interval censoring[19]. Although several procedures have been developed for one-sample estimation problem, there does not exist much literature on the topic. The same is true for the investigation of the use of parametric models and inference procedures for the analysis of interval-censored data. One major reason for this is that in most situations, there does not exist much prior information about the variable under study and thus one may prefer nonparametric or semiparametric approaches rather than parametric approaches.
7 Concluding Remarks
It is definitely important to implement the available inference procedures numerically for practitioners. Unfortunately and surprisingly, there is no commercially available statistical software yet that provides an extensive coverage for interval-censored data. This is perhaps due to the complexity of both the algorithms and the theory behind it. One can, however, find some simple functions in SPLUS, R and SAS that can be applied to interval-censored data. For example, in SPLUS, the function kaplanMeier can be used to compute the Turnbull estimator[19]. In R, the package Icens contains some routines that can perform statistical analysis when interval-censored data are present. A recent tutorial paper[86] has some details and examples of how to use R for interval-censored data. In SAS, the procedure LIFEREG allows one to fit the parametric accelerated failure time model to interval-censored data.
Methodologically, there are still many open questions in the analysis of interval-censored data. Examples include but are not limited to model checking techniques and joint modeling of longitudinal and interval-censored data. Some of the methods discussed in the previous sections also need proper theoretical justification. The major difficulty is that there lacks basic tools as simple and elegant as the partial likelihood and the martingale theory for right-censored data. The work by Groeneboom and Wellner[9] and Huang and Wellner[50] are perhaps the most comprehensive studies for interval censoring, which mainly rely on complicated empirical processes and the optimization theory and are difficult to generalize.
Acknowledgments
The authors wish to thank Professor Per Kragh Andersen and a reviewer for their helpful comments.
Contributor Information
Zhigang Zhang, Email: zhangz@mskcc.org, Department of Epidemiology and Biostatistics, Memorial Sloan-Kettering Cancer Center, 307 East 63rd Street, New York, NY 10065, U.S.A.
Jianguo Sun, Email: sunj@missouri.edu, Department of Statistics, University of Missouri, 146 Middlebush Hall, Columbia, Missouri 65211, U.S.A.
References
- 1.Finkelstein DM. A proportional hazards model for interval-censored failure time data. Biometrics. 1986;42:845–854. [PubMed] [Google Scholar]
- 2.Kalbfleisch JD, Prentice RL. The statistical analysis of failure time data. New York: Wiley; 2002. [Google Scholar]
- 3.Sun J. The statistical analysis of interval-censored failure time data. New York: Springer; 2006. [Google Scholar]
- 4.De Gruttola VG, Lagakos SW. Analysis of doubly-censored survival data, with application to AIDS. Biometrics. 1989;45:1–11. [PubMed] [Google Scholar]
- 5.Jewell NP, van der Laan MJ. Generalizations of current status data with applications. Lifetime Data Analysis. 1995;1:101–110. doi: 10.1007/BF00985261. [DOI] [PubMed] [Google Scholar]
- 6.Sun J, Kalbfleisch JD. The analysis of current status data on point processes. Journal of the American Statistical Association. 1993;88:1449–1454. doi: 10.1080/01621459.1993.10476336. [DOI] [PubMed] [Google Scholar]
- 7.Keiding N. Age-specific incidence and prevalence: A statistical perspective (with discussion) Journal of the Royal Statistical Society: Series A. 1991;154:371–412. [Google Scholar]
- 8.Dinse GE, Lagakos SW. Regression analysis of tumor prevalence data. Applied Statistics. 1983;32:236–248. [Google Scholar]
- 9.Groeneboom P, Wellner JA. DMV Seminar, Band 19. New York: Birkhauser; 1992. Information bounds and nonparametric maximum likelihood estimation. [Google Scholar]
- 10.Sun J. Statistical analysis of doubly interval-censored failure time data. In: Balakrishnan N, Rao CR, editors. Handbook of statistics: survival analysis. 2002. [Google Scholar]
- 11.Finkelstein DM, Wolfe RA. A semiparametric model for regression analysis of interval-censored failure time data. Biometrics. 1985;41:933–945. [PubMed] [Google Scholar]
- 12.Sun J. Encyclopedia of biostatistics. 2. John Wiley & Sons Ltd; 2005. Interval censoring; pp. 2603–2609. [Google Scholar]
- 13.Oller R, Gómez G, Calle ML. Interval censoring: model characterizations for the validity of the simplified likelihood. The Canadian Journal of Statistics. 2004;32:315–326. [Google Scholar]
- 14.Betensky RA. On nonidentifiability and noninformative censoring for current status data. Biometrika. 2000;87:218–221. [Google Scholar]
- 15.Oller R, Gómez G, Calle ML. Interval censoring: identifiability and the constant-sum property. Biometrika. 2007;94:61–70. [Google Scholar]
- 16.Lawless JF, Babineau D. Models for interval censoring and simulation-based inference for lifetime distributions. Biometrika. 2006;93:671–686. [Google Scholar]
- 17.Gentleman R, Geyer CJ. Maximum likelihood for interval censored data: Consistency and computation. Biometrika. 1994;81:618–623. [Google Scholar]
- 18.Li L, Watkins T, Yu Q. An EM algorithm for estimating survival functions with interval-censored data. Scandinavian Journal of Statistics. 1997;24:531–542. [Google Scholar]
- 19.Turnbull BW. The empirical distribution with arbitrarily grouped censored and truncated data. Journal of the Royal Statistical Society: Series B. 1976;38:290–295. [Google Scholar]
- 20.Dempster AP, Laird NM, Rubin DB. Maximum likelihood from incomplete data via the EM algorithm. Journal of the Royal Statistical Society: Series B. 1977;39:1–38. [Google Scholar]
- 21.Jongbloed G. The iterative convex minorant algorithm for nonparametric estimation. Journal of Computational and Graphical Statistics. 1998;7:310–321. [Google Scholar]
- 22.Robertson T, Wright FT, Dykstra R. Order restrict statistical inference. New York: John Wiley; 1988. [Google Scholar]
- 23.Wellner JA, Zhan Y. A hybrid algorithm for computation of the nonparametric maximum likelihood estimator from censored data. Journal of the American Statistical Association. 1997;92:945–959. [Google Scholar]
- 24.Böhning D, Schlattmann P, Dietz E. Interval censored data: A note on the non-parametric maximum likelihood estimator of the distribution function. Biometrika. 1996;83:462–466. [Google Scholar]
- 25.Barlow RE, Bartholomew DJ, Bremner JM, Brunk HD. Statistical inference under order restrictions. New York: Wiley; 1972. [Google Scholar]
- 26.Vandal AC, Gentleman R, Liu X. Constrained estimation and likelihood intervals for censored data. The Canadian Journal of Statistics. 2005;33:71–84. [Google Scholar]
- 27.Sen B, Banerjee M. A pseudolikelihood method for analyzing interval censored data. Biometrika. 2007;94:71–86. [Google Scholar]
- 28.Yu Q, Li L, Wong G. On consistency of self-consistent estimator of survival functions with interval-censored data. Scandinavian Journal of Statistics. 2000;27:35–44. [Google Scholar]
- 29.Geskus R, Groeneboom P. Asymptotically optimal estimation of smooth functionals for interval censoring, case 2. The Annals of Statistics. 1999;27:627–674. [Google Scholar]
- 30.Huang J, Wellner JA. Asymptotic normality of the NPMLE of linear functionals for interval censored data, case I. Statistica Neerlandica. 1995;49:153–163. [Google Scholar]
- 31.Kooperberg C, Stone CJ. Logspline density estimation for censored data. Journal of Computational and Graphical Statistics. 1992;1:301–328. [Google Scholar]
- 32.Rosenberg PS. Hazard function estimation using B-splines. Biometrics. 1995;51:874–887. [PubMed] [Google Scholar]
- 33.Bebchuk JD, Betensky RA. Multiple imputation for simple estimation of the hazard function based on interval censored data. Statistics in Medicine. 2000;19:405–419. doi: 10.1002/(sici)1097-0258(20000215)19:3<405::aid-sim325>3.0.co;2-2. [DOI] [PubMed] [Google Scholar]
- 34.Betensky RA, Lindsey JC, Ryan LM, Wand MP. Local EM estimation of the hazard function for interval-censored data. Biometrics. 1999;55:238–245. doi: 10.1111/j.0006-341x.1999.00238.x. [DOI] [PubMed] [Google Scholar]
- 35.Zhao Q, Sun J. Generalized log-rank test for mixed interval-censored failure time data. Statistics in Medicine. 2004;23:1621–1629. doi: 10.1002/sim.1746. [DOI] [PubMed] [Google Scholar]
- 36.Kim J, Kang DR, Nam CM. Logrank-type tests for comparing survival curves with interval-censored data. Computational Statistics & Data Analysis. 2006;50:3165–3178. [Google Scholar]
- 37.Petroni GR, Wolfe RA. A two-sample test for stochastic ordering with interval-censored data. Biometrics. 1994;50:77–87. [PubMed] [Google Scholar]
- 38.Fang H, Sun J, Lee M-LT. Nonparametric survival comparison for interval-censored continuous Data. Statistica Sinica. 2002;12:1073–1083. [Google Scholar]
- 39.Yuen K, Shi J, Zhu L. A k-sample test with interval censored data. Biometrika. 2006;93:315–328. [Google Scholar]
- 40.Sun J, Zhao Q, Zhao X. Generalized log rank tests for interval-censored failure time data. Scandinavian Journal of Statistics. 2005;32:49–57. [Google Scholar]
- 41.Peto R, Peto J. Asymptotically efficient rank invariant test procedures. Journal of the Royal Statistical Society: Series A. 1972;135:185–207. [Google Scholar]
- 42.Zhao X, Zhao Q, Sun J, Kim JS. Generalized log-rank tests for partly interval-censored failure time data. Biometrical Journal. 2009 doi: 10.1002/bimj.200710419. to appear. [DOI] [PubMed] [Google Scholar]
- 43.Andersen PK, Ronn BB. A nonparametric test for comparing two samples where all observations are either left- or right-censored. Biometrics. 1995;51:323–329. [PubMed] [Google Scholar]
- 44.Sun J. A nonparametric test for current status data with unequal censoring. Journal of the Royal Statistical Society: Series B. 1999;61:243–250. [Google Scholar]
- 45.Cox DR. Regression models and life-tables (with discussion) Journal of the Royal Statistical Society: Series B. 1972;34:187–220. [Google Scholar]
- 46.Huang J. Efficient estimation for the proportional hazards model with interval censoring. The Annals of Statistics. 1996;24:540–568. [Google Scholar]
- 47.Satten GA. Rank-based inference in the proportional hazards model for interval censored data. Biometrika. 1996;83:355–370. [Google Scholar]
- 48.Goggins WB, Finkelstein DM, Schoenfeld DA, Zaslavsky AM. A Markov chain Monte Carlo EM algorithm for analyzing interval censored data under the Cox proportional hazards model. Biometrics. 1998;54:1498–1507. [PubMed] [Google Scholar]
- 49.Pan W. A multiple imputation approach to Cox regression with interval-censored data. Biometrics. 2000;56:199–203. doi: 10.1111/j.0006-341x.2000.00199.x. [DOI] [PubMed] [Google Scholar]
- 50.Huang J, Wellner JA. Interval censored survival data: a review of recent progress. In: Lin D, Fleming T, editors. Proceedings of the first Seattle symposium in biostatistics: survival analysis. New York: Springer-Verlag; 1997. [Google Scholar]
- 51.Kim Y, Jhun M. Cure rate model with interval censored data. Statistics in Medicine. 2008;27:3–14. doi: 10.1002/sim.2918. [DOI] [PubMed] [Google Scholar]
- 52.Huang J. Maximum likelihood estimation for proportional odds regression model with current status data. Analysis of censored cata. 1995;27:129–146. IMS Lecture Notes - Monograph Series. [Google Scholar]
- 53.Rossini A, Tsiatis AA. A semiparametric proportional odds regression model for the analysis of current status data. Journal of the American Statistical Association. 1996;91:713–721. [Google Scholar]
- 54.Huang J, Rossini AJ. Sieve estimation for the proportional odds failure-time regression model with interval censoring. Journal of the American Statistical Association. 1997;92:960–967. [Google Scholar]
- 55.Shen X. Proportional odds regression and sieve maximum likelihood estimation. Biometrika. 1998;85:165–177. [Google Scholar]
- 56.Rabinowitz D, Betensky RA, Tsiatis AA. Using conditional logistic regression to fit proportional odds models to interval censored data. Biometrika. 2000;56:511–518. doi: 10.1111/j.0006-341x.2000.00511.x. [DOI] [PubMed] [Google Scholar]
- 57.Rabinowitz D, Tsiatis AA, Aragon J. Regression with interval-censored data. Biometrika. 1995;82:501–513. [Google Scholar]
- 58.Betensky RA, Rabinowitz D, Tsiatis AA. Computationally simple accelerated failure time regression for interval censored data. Biometrika. 2001;88:703–711. [Google Scholar]
- 59.Li L, Pu Z. Rank estimation of log-linear regression with interval-censored data. Lifetime Data Analysis. 2003;9:57–70. doi: 10.1023/a:1021882122257. [DOI] [PubMed] [Google Scholar]
- 60.Tian L, Cai T. On the accelerated failure time model for current status and interval censored data. Biometrika. 2006;93:329–342. [Google Scholar]
- 61.Xue H, Lam KF, Cowling BJ, de Wolf F. Semi-parametric accelerated failure time regression analysis with application to interval-censored HIV/AIDS data. Statistics in Medicine. 2006;25:3850–3863. doi: 10.1002/sim.2486. [DOI] [PubMed] [Google Scholar]
- 62.Lin DY, Oakes D, Ying Z. Additive hazards regression with current status data. Biometrika. 1998;85:289–298. [Google Scholar]
- 63.Martinussen T, Scheike TH. Efficient estimation in additive hazards regression with current status data. Biometrika. 2002;89:649–658. [Google Scholar]
- 64.Zeng D, Cai J, Shen Y. Semiparametric additive risks model for interval-censored data. Statistica Sinica. 2006;16:287–302. [Google Scholar]
- 65.Chen L, Sun J. A multiple imputation approach to the analysis of current status data with the additive hazards model. Communications in Statistics: Theory and Methods. 2009 to appear. [Google Scholar]
- 66.Zhu L, Tong X, Sun J. A transformation approach for the analysis of interval-censored failure time data. Lifetime Data Analysis. 2008;14:167–178. doi: 10.1007/s10985-007-9075-8. [DOI] [PubMed] [Google Scholar]
- 67.Sun J, Sun L. Semiparametric linear transformation models for current status data. The Canadian Journal of Statistics. 2005;33:85–96. [Google Scholar]
- 68.Younes N, Lachin J. Linked-based models for survival data with interval and continuous time censoring. Biometrics. 1997;53:1199–1211. [Google Scholar]
- 69.Zhang Z, Sun L, Zhao X, Sun J. Regression analysis of interval censored failure time data with linear transformation models. The Canadian Journal of Statistics. 2005;33:61–70. [Google Scholar]
- 70.Xue H, Lam KF, Li G. Sieve maximum likelihood estimation for semiparametric regression models with current status data. Journal of the American Statistical Association. 2004;99:346–356. [Google Scholar]
- 71.Shiboski SC. Generalized additive models for current status data. Lifetime Data Analysis. 1998;4:29–50. doi: 10.1023/a:1009652024999. [DOI] [PubMed] [Google Scholar]
- 72.Zhang M, Davidian M. “Smooth” semiparametric regression analysis for arbitrarily censored time-to-event data. Biometrics. 2008;64:567–576. doi: 10.1111/j.1541-0420.2007.00928.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Chen M, Tong X, Sun J. The proportional odds model for multivariate interval-censored failure time data. Statistics in Medicine. 2007;26:5147–5161. doi: 10.1002/sim.2907. [DOI] [PubMed] [Google Scholar]
- 74.Wang L, Sun J, Tong X. Efficient estimation for the proportional hazards model with bivariate current status data. Lifetime Data Analysis. 2008;14:134–153. doi: 10.1007/s10985-007-9058-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Zhang B, Tong X, Sun J. Efficient estimation for the proportional odds model with bivariate current status data. Far East Journal of Theoretical Statistics. 2009 to appear. [Google Scholar]
- 76.Cook RJ, Zeng L, Lee K-A. A Multistate Model for Bivariate Interval-Censored Failure Time Data. Biometrics. 2009 to appear. [Google Scholar]
- 77.Komàrek A, Lesaffre E. Bayesian accelerated failure time model for correlated interval-censored data with a normal mixture as error distribution. Statistica Sinica. 2007;17:549–569. [Google Scholar]
- 78.Sun J. Statistical analysis of doubly interval-censored failure time data. Advances in survival analysis, handbook of biostatistics. 2004;23:105–122. [Google Scholar]
- 79.Komàrek A, Lesaffre E. Bayesian accelerated failure time model with multivariate doubly interval-censored data and flexible distributional assumptions. Journal of the American Statistical Association. 2008;103:523–533. [Google Scholar]
- 80.Kim Y. Regression analysis of doubly censored failure time data with frailty. Biometrics. 2006;62:458–464. doi: 10.1111/j.1541-0420.2005.00487.x. [DOI] [PubMed] [Google Scholar]
- 81.Groeneboom P, Maathuis MH, Wellner JA. Current status data with competing risks: Consistency and rates of convergence of the MLE. The Annals of Statistics. 2008;36:1031–1063. doi: 10.1214/009053607000000983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Groeneboom P, Maathuis MH, Wellner JA. Current status data with competing risks: Limiting distribution of the MLE. The Annals of Statistics. 2008;36:1064–1089. doi: 10.1214/009053607000000983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Zhang Z, Sun J, Sun L. Statistical analysis of current status data with informative observation times. Statistics in Medicine. 2005;24:1399–1407. doi: 10.1002/sim.2001. [DOI] [PubMed] [Google Scholar]
- 84.Zhang Z, Sun L, Sun J, Finkelstein DM. Regression analysis of failure time data with informative interval censoring. Statistics in Medicine. 2007;26:2533–2546. doi: 10.1002/sim.2721. [DOI] [PubMed] [Google Scholar]
- 85.Park Y, Tian L, Wei LJ. One- and two-sample nonparametric inference procedures in the presence of a mixture of independent and dependent censoring. Biostatistics. 2006;7:252–267. doi: 10.1093/biostatistics/kxj005. [DOI] [PubMed] [Google Scholar]
- 86.Gómez G, Oller R, Calle ML, Langohr K. Tutorial on methods for interval-censored data and their implementation in R. Statistical Modelling. 2009 to appear. [Google Scholar]