Abstract
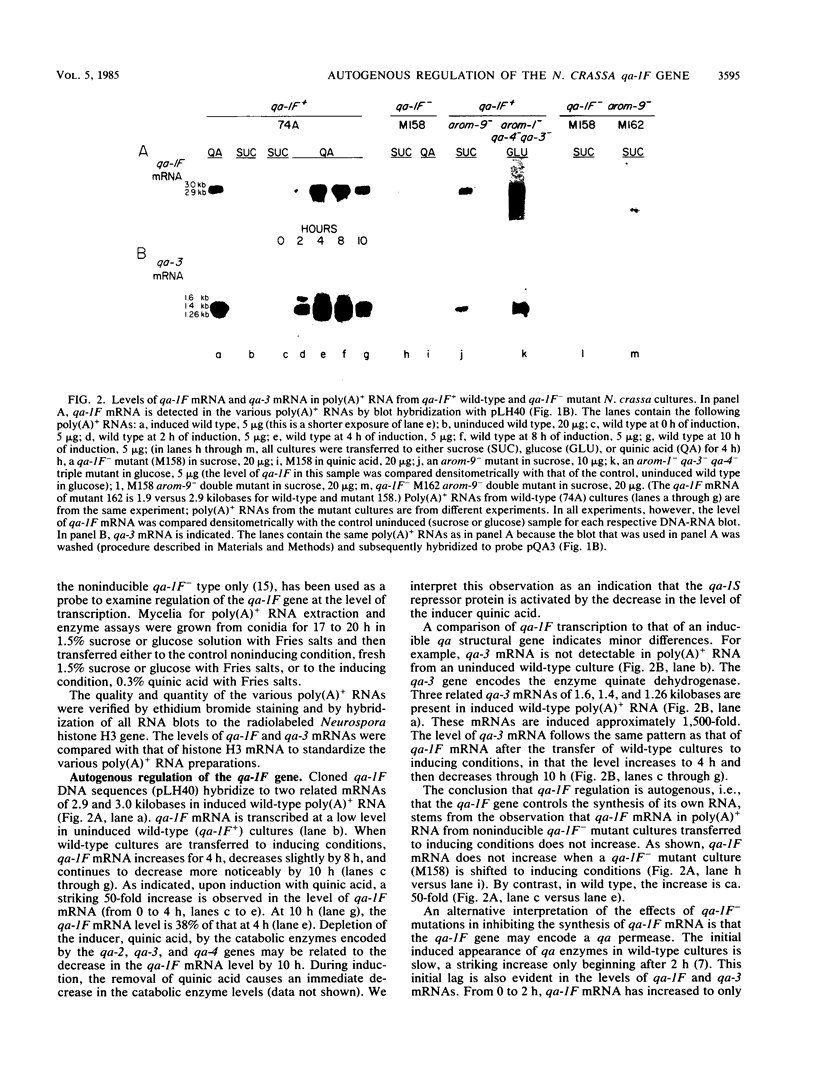
In Neurospora crassa, the qa-1F regulatory gene positively controls transcription of all genes in the quinic acid (qa) gene cluster. qa-1F is transcribed at a low, uninduced level but is subject to strong (50-fold), autogenous regulation as well as to control by the negative regulatory gene, qa-1S, and the inducer quinic acid. Cloned qa-1F DNA sequences hybridize to two related mRNAs of 2.9 and 3.0 kilobases. When wild-type (qa-1F+) cultures are transferred to inducing conditions, qa-1F mRNA increases for 4 h, remains somewhat level, and decreases after 8 to 10 h. That this control is autogenous, i.e., that the qa-1F gene controls the synthesis of its own mRNA, is indicated by the presence of approximately the same low level of qa-1F mRNA in poly(A)+ RNA from noninducible qa-1F- mutant cultures under inducing conditions as that observed in uninduced wild-type cultures. The qa-1S gene also regulates the transcription of qa-1F, since a qa-1S- mutant, whether in noninducing or inducing conditions, contains a level of qa-1F mRNA that corresponds to the low level observed in uninduced wild-type cultures. These results corroborate the hypothesis (M. E. Case and N. H. Giles, Proc. Natl. Acad. Sci. USA 72:553-557, 1975; V. B. Patel, M. Schweizer, C. C. Dykstra, S. R. Kushner, and N. H. Giles, Proc. Natl. Acad. Sci. USA 78:5783-5787, 1981; L. Huiet, Proc. Natl. Acad. Sci. USA 81:1174-1178, 1984) that the qa-1F gene encodes an activator protein and acts positively in controlling transcription of itself and the other qa genes.
Full text
PDF



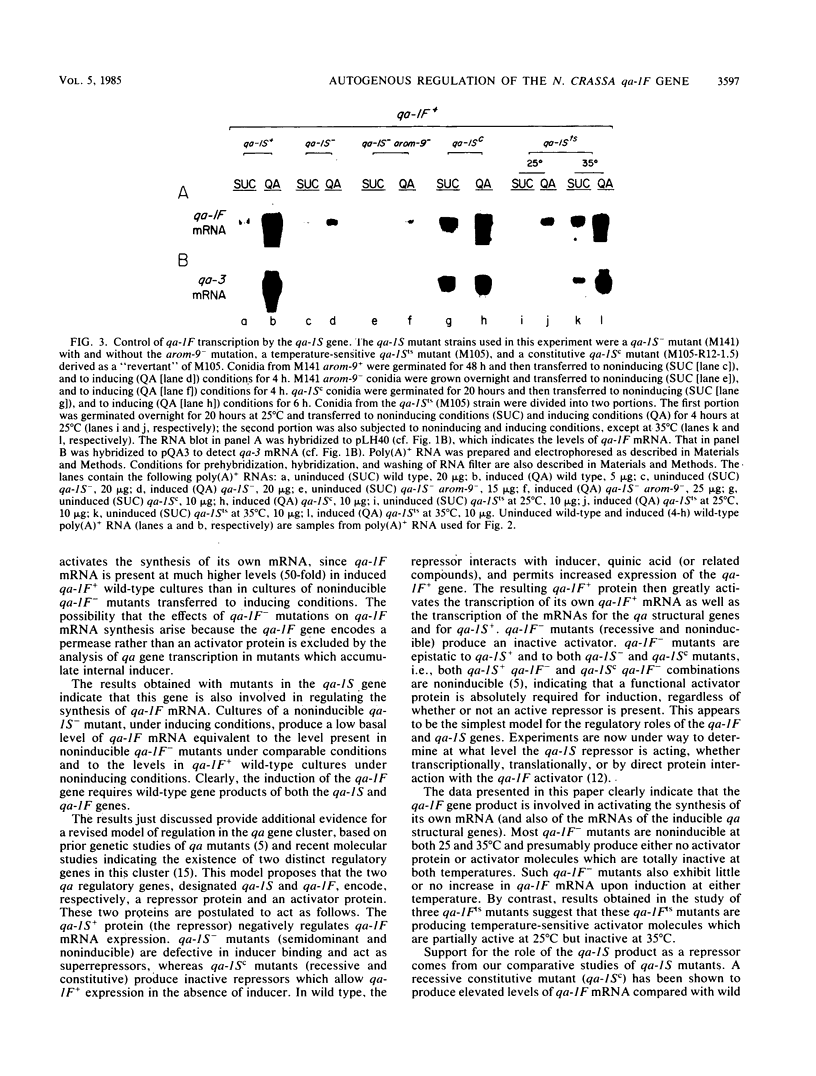


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alton N. K., Buxton F., Patel V., Giles N. H., Vapnek D. 5'-Untranslated sequences of two structural genes in the qa gene cluster of Neurospora crassa. Proc Natl Acad Sci U S A. 1982 Mar;79(6):1955–1959. doi: 10.1073/pnas.79.6.1955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alwine J. C., Kemp D. J., Stark G. R. Method for detection of specific RNAs in agarose gels by transfer to diazobenzyloxymethyl-paper and hybridization with DNA probes. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5350–5354. doi: 10.1073/pnas.74.12.5350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aviv H., Leder P. Purification of biologically active globin messenger RNA by chromatography on oligothymidylic acid-cellulose. Proc Natl Acad Sci U S A. 1972 Jun;69(6):1408–1412. doi: 10.1073/pnas.69.6.1408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bailey J. M., Davidson N. Methylmercury as a reversible denaturing agent for agarose gel electrophoresis. Anal Biochem. 1976 Jan;70(1):75–85. doi: 10.1016/s0003-2697(76)80049-8. [DOI] [PubMed] [Google Scholar]
- Case M. E., Giles N. H., Doy C. H. Genetical and biochemical evidence for further interrelationships between the polyaromatic synthetic and the quinate-shikimate catabolic pathways in Neurospora crassa. Genetics. 1972 Jul;71(3):337–348. doi: 10.1093/genetics/71.3.337. [DOI] [PubMed] [Google Scholar]
- Case M. E., Giles N. H. Genetic evidence on the organization and action of the qa-1 gene product: a protein regulating the induction of three enzymes in quinate catabolism in Neurospora crassa. Proc Natl Acad Sci U S A. 1975 Feb;72(2):553–557. doi: 10.1073/pnas.72.2.553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chaleff R. S. The inducible quinate-shikimate catabolic pathway in Neurospora crassa: genetic organization. J Gen Microbiol. 1974 Apr;81(2):337–355. doi: 10.1099/00221287-81-2-337. [DOI] [PubMed] [Google Scholar]
- DOUGLAS H. C., HAWTHORNE D. C. ENZYMATIC EXPRESSION AND GENETIC LINKAGE OF GENES CONTROLLING GALACTOSE UTILIZATION IN SACCHAROMYCES. Genetics. 1964 May;49:837–844. doi: 10.1093/genetics/49.5.837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feinberg A. P., Vogelstein B. A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal Biochem. 1983 Jul 1;132(1):6–13. doi: 10.1016/0003-2697(83)90418-9. [DOI] [PubMed] [Google Scholar]
- Gaertner F. H., Cole K. W. A cluster-gene: evidence for one gene, one polypeptide, five enzymes. Biochem Biophys Res Commun. 1977 Mar 21;75(2):259–264. doi: 10.1016/0006-291x(77)91037-3. [DOI] [PubMed] [Google Scholar]
- Giles N. H., Case M. E., Baum J., Geever R., Huiet L., Patel V., Tyler B. Gene organization and regulation in the qa (quinic acid) gene cluster of Neurospora crassa. Microbiol Rev. 1985 Sep;49(3):338–358. doi: 10.1128/mr.49.3.338-358.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giles N. H., Case M. E., Partridge C. W., Ahmed S. I. A gene cluster in Nuerospora crassa coding for an aggregate of five aromatic synthetic enzymes. Proc Natl Acad Sci U S A. 1967 Oct;58(4):1453–1460. doi: 10.1073/pnas.58.4.1453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guarente L., Yocum R. R., Gifford P. A GAL10-CYC1 hybrid yeast promoter identifies the GAL4 regulatory region as an upstream site. Proc Natl Acad Sci U S A. 1982 Dec;79(23):7410–7414. doi: 10.1073/pnas.79.23.7410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huiet L. Molecular analysis of the Neurospora qa-1 regulatory region indicates that two interacting genes control qa gene expression. Proc Natl Acad Sci U S A. 1984 Feb;81(4):1174–1178. doi: 10.1073/pnas.81.4.1174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnston S. A., Hopper J. E. Isolation of the yeast regulatory gene GAL4 and analysis of its dosage effects on the galactose/melibiose regulon. Proc Natl Acad Sci U S A. 1982 Nov;79(22):6971–6975. doi: 10.1073/pnas.79.22.6971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krumlauf R., Marzluf G. A. Genome organization and characterization of the repetitive and inverted repeat DNA sequences in Neurospora crassa. J Biol Chem. 1980 Feb 10;255(3):1138–1145. [PubMed] [Google Scholar]
- Laughon A., Gesteland R. F. Isolation and preliminary characterization of the GAL4 gene, a positive regulator of transcription in yeast. Proc Natl Acad Sci U S A. 1982 Nov;79(22):6827–6831. doi: 10.1073/pnas.79.22.6827. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Loo M. Neurospora crassa temperature-sensitive mutant apparently defective in protein synthesis. J Bacteriol. 1975 Jan;121(1):286–295. doi: 10.1128/jb.121.1.286-295.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsumoto K., Adachi Y., Toh-e A., Oshima Y. Function of positive regulatory gene gal4 in the synthesis of galactose pathway enzymes in Saccharomyces cerevisiae: evidence that the GAL81 region codes for part of the gal4 protein. J Bacteriol. 1980 Feb;141(2):508–527. doi: 10.1128/jb.141.2.508-527.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nogi Y., Shimada H., Matsuzaki Y., Hashimoto H., Fukasawa T. Regulation of expression of the galactose gene cluster in Saccharomyces cerevisiae. II. The isolation and dosage effect of the regulatory gene GAL80. Mol Gen Genet. 1984;195(1-2):29–34. doi: 10.1007/BF00332719. [DOI] [PubMed] [Google Scholar]
- Patel V. B., Schweizer M., Dykstra C. C., Kushner S. R., Giles N. H. Genetic organization and transcriptional regulation in the qa gene cluster of Neurospora crassa. Proc Natl Acad Sci U S A. 1981 Sep;78(9):5783–5787. doi: 10.1073/pnas.78.9.5783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perlman D., Hopper J. E. Constitutive synthesis of the GAL4 protein, a galactose pathway regulator in Saccharomyces cerevisiae. Cell. 1979 Jan;16(1):89–95. doi: 10.1016/0092-8674(79)90190-9. [DOI] [PubMed] [Google Scholar]
- Reinert W. R., Patel V. B., Giles N. H. Genetic regulation of the qa gene cluster of Neurospora crassa: induction of qa messenger ribonucleic acid and dependency on qa-1 function. Mol Cell Biol. 1981 Sep;1(9):829–835. doi: 10.1128/mcb.1.9.829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rigby P. W., Dieckmann M., Rhodes C., Berg P. Labeling deoxyribonucleic acid to high specific activity in vitro by nick translation with DNA polymerase I. J Mol Biol. 1977 Jun 15;113(1):237–251. doi: 10.1016/0022-2836(77)90052-3. [DOI] [PubMed] [Google Scholar]
- Rines H. W., Case M. E., Giles N. H. Mutants in the arom gene cluster of Neurospora crassa specific for biosynthetic dehydroquinase. Genetics. 1969 Apr;61(4):789–800. doi: 10.1093/genetics/61.4.789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tyler B. M., Cowman A. F., Adams J. M., Harris A. W. Generation of long mRNA for membrane immunoglobulin gamma 2a chains by differential splicing. Nature. 1981 Oct 1;293(5831):406–408. doi: 10.1038/293406a0. [DOI] [PubMed] [Google Scholar]
- Tyler B. M., Geever R. F., Case M. E., Giles N. H. Cis-acting and trans-acting regulatory mutations define two types of promoters controlled by the qa-1F gene of Neurospora. Cell. 1984 Feb;36(2):493–502. doi: 10.1016/0092-8674(84)90242-3. [DOI] [PubMed] [Google Scholar]
- Valone J. A., Jr, Case M. E., Giles N. H. Constitutive mutants in a regulatory gene exerting positive control of quinic acid catabolism in Neurospora crassa. Proc Natl Acad Sci U S A. 1971 Jul;68(7):1555–1559. doi: 10.1073/pnas.68.7.1555. [DOI] [PMC free article] [PubMed] [Google Scholar]





