Abstract
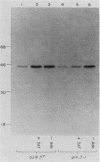
Production of glutamine synthetase in Saccharomyces cerevisiae is controlled by three regulatory systems. One system responds to glutamine levels and depends on the positively acting GLN3 product. This system mediates derepression of glutamine synthetase in response to pyrimidine limitation as well, but genetic evidence argues that this is an indirect effect of depletion of the glutamine pool. The second system is general amino acid control, which couples derepression of a variety of biosynthetic enzymes to starvation for many single amino acids. This system operates through the positive regulatory element GCN4. Expression of histidinol dehydrogenase, which is under general control, is not stimulated by glutamine limitation. A third system responds to purine limitation. No specific regulatory element has been identified, but depression of glutamine synthetase is observed during purine starvation in gln3 gcn4 double mutants. This demonstrates that a separate purine regulatory element must exist. Pulse-labeling and immunoprecipitation experiments indicate that all three systems control glutamine synthetase at the level of subunit synthesis.
Full text
PDF

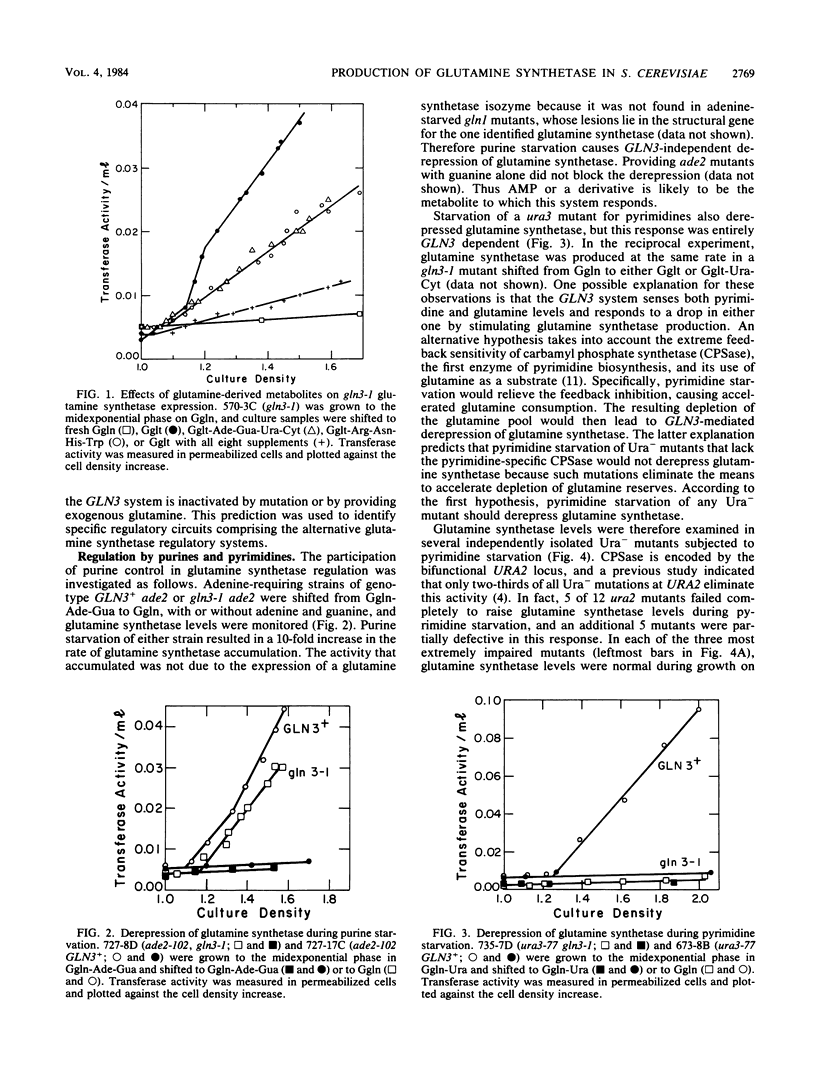

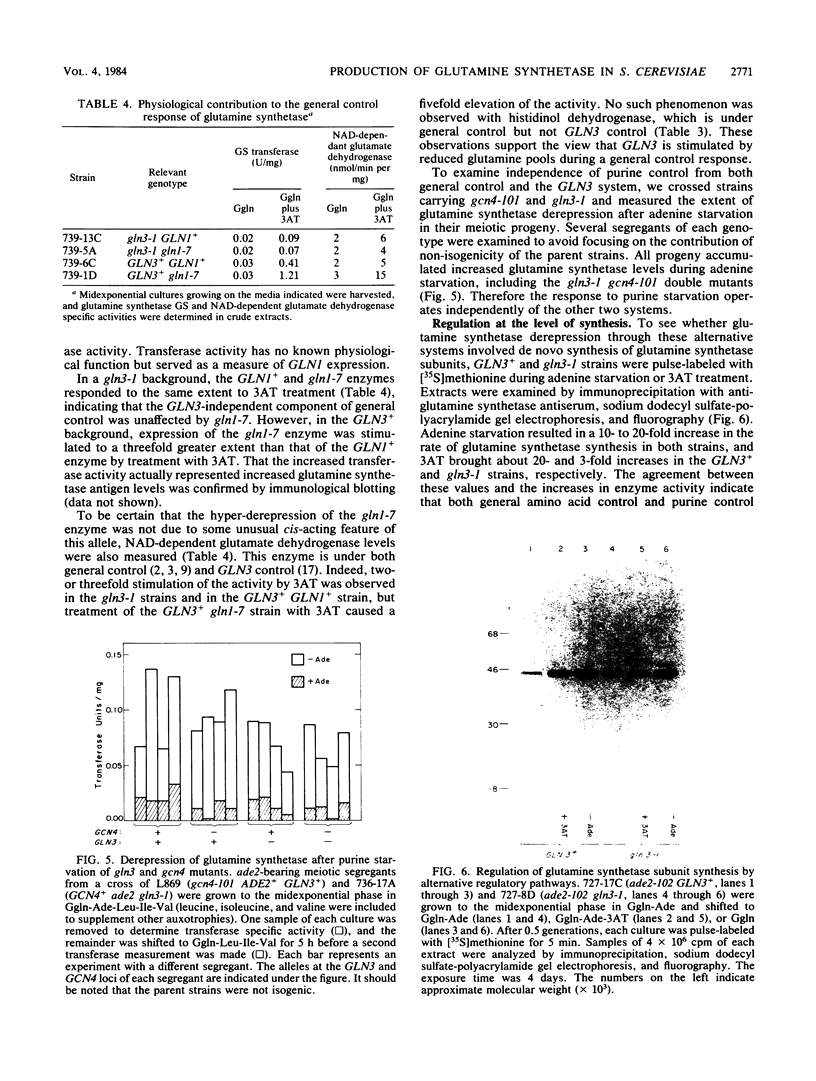


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Brandriss M. C., Magasanik B. Genetics and physiology of proline utilization in Saccharomyces cerevisiae: enzyme induction by proline. J Bacteriol. 1979 Nov;140(2):498–503. doi: 10.1128/jb.140.2.498-503.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cherest H., Surdin-Kerjan Y., Robichon-Szulmajster H. Methionine-mediated repression in Saccharomyces cerevisiae: a pleiotropic regulatory system involving methionyl transfer ribonucleic acid and the product of gene eth2. J Bacteriol. 1971 Jun;106(3):758–772. doi: 10.1128/jb.106.3.758-772.1971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Delforge J., Messenguy F., Wiame J. M. The regulation of arginine biosynthesis in Saccharomyces cerevisiae. The specificity of argR- mutations and the general control of amino-acid biosynthesis. Eur J Biochem. 1975 Sep 1;57(1):231–239. doi: 10.1111/j.1432-1033.1975.tb02295.x. [DOI] [PubMed] [Google Scholar]
- Denis-Duphil M., Kaplan J. G. Fine structure of the URA2 locus in Saccharomyces cerevisiae. II. Meiotic and mitotic mapping studies. Mol Gen Genet. 1976 Jun 15;145(3):259–271. doi: 10.1007/BF00325822. [DOI] [PubMed] [Google Scholar]
- Donahue T. F., Daves R. S., Lucchini G., Fink G. R. A short nucleotide sequence required for regulation of HIS4 by the general control system of yeast. Cell. 1983 Jan;32(1):89–98. doi: 10.1016/0092-8674(83)90499-3. [DOI] [PubMed] [Google Scholar]
- Guarente L., Lalonde B., Gifford P., Alani E. Distinctly regulated tandem upstream activation sites mediate catabolite repression of the CYC1 gene of S. cerevisiae. Cell. 1984 Feb;36(2):503–511. doi: 10.1016/0092-8674(84)90243-5. [DOI] [PubMed] [Google Scholar]
- Guarente L., Mason T. Heme regulates transcription of the CYC1 gene of S. cerevisiae via an upstream activation site. Cell. 1983 Apr;32(4):1279–1286. doi: 10.1016/0092-8674(83)90309-4. [DOI] [PubMed] [Google Scholar]
- Guarente L., Yocum R. R., Gifford P. A GAL10-CYC1 hybrid yeast promoter identifies the GAL4 regulatory region as an upstream site. Proc Natl Acad Sci U S A. 1982 Dec;79(23):7410–7414. doi: 10.1073/pnas.79.23.7410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- LOWRY O. H., ROSEBROUGH N. J., FARR A. L., RANDALL R. J. Protein measurement with the Folin phenol reagent. J Biol Chem. 1951 Nov;193(1):265–275. [PubMed] [Google Scholar]
- Lacroute F., Piérard A., Grenson M., Wiame J. M. The biosynthesis of carbamoyl phosphate in Saccharomyces cerevisiae. J Gen Microbiol. 1965 Jul;40(1):127–142. doi: 10.1099/00221287-40-1-127. [DOI] [PubMed] [Google Scholar]
- Lacroute F. Regulation of pyrimidine biosynthesis in Saccharomyces cerevisiae. J Bacteriol. 1968 Mar;95(3):824–832. doi: 10.1128/jb.95.3.824-832.1968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Legrain C., Vissers S., Dubois E., Legrain M., Wiame J. M. Regulation of glutamine synthetase from Saccharomyces cerevisiae by repression, inactivation and proteolysis. Eur J Biochem. 1982 Apr;123(3):611–616. doi: 10.1111/j.1432-1033.1982.tb06576.x. [DOI] [PubMed] [Google Scholar]
- Lucchini G., Hinnebusch A. G., Chen C., Fink G. R. Positive regulatory interactions of the HIS4 gene of Saccharomyces cerevisiae. Mol Cell Biol. 1984 Jul;4(7):1326–1333. doi: 10.1128/mcb.4.7.1326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Messenguy F., Colin D., ten Have J. P. Regulation of compartmentation of amino acid pools in Saccharomyces cerevisiae and its effects on metabolic control. Eur J Biochem. 1980 Jul;108(2):439–447. doi: 10.1111/j.1432-1033.1980.tb04740.x. [DOI] [PubMed] [Google Scholar]
- Mitchell A. P., Magasanik B. Biochemical and physiological aspects of glutamine synthetase inactivation in Saccharomyces cerevisiae. J Biol Chem. 1984 Oct 10;259(19):12054–12062. [PubMed] [Google Scholar]
- Mitchell A. P., Magasanik B. Purification and properties of glutamine synthetase from Saccharomyces cerevisiae. J Biol Chem. 1983 Jan 10;258(1):119–124. [PubMed] [Google Scholar]
- Mitchell A. P., Magasanik B. Regulation of glutamine-repressible gene products by the GLN3 function in Saccharomyces cerevisiae. Mol Cell Biol. 1984 Dec;4(12):2758–2766. doi: 10.1128/mcb.4.12.2758. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolfner M., Yep D., Messenguy F., Fink G. R. Integration of amino acid biosynthesis into the cell cycle of Saccharomyces cerevisiae. J Mol Biol. 1975 Aug 5;96(2):273–290. doi: 10.1016/0022-2836(75)90348-4. [DOI] [PubMed] [Google Scholar]