Abstract
We previously identified a set of 50 genes that were differentially transcribed in the hippocampal CA1 region of aged, learning-impaired rats compared to aged, superior learning animals during a Morris water maze paradigm. In the current study we expressed three of these genes (Pctk1, Tcf12 and Ccnd1), which had shown increased transcription in aged, learning impaired rats, in the hippocampus of young rats using viral gene transfer and tested for learning and memory deficits at age 7–14 months. Pctk1 injected animals displayed a modest deficit in acquiring latency in both the Morris Water Maze and the reverse Morris maze. In the radial arm water maze paradigm, Pctk1, Tcf12 and Ccnd1 expressing animals all showed significant deficits in spatial working memory compared to controls. Rats injected with Ccnd1 and Tcf12, but not Pctk1, also showed a significant deficit in spatial reference memory in the radial arm water maze. Electrophysiological experiments revealed no difference in LTP in Ccnd1 and Pctk1 animals. However, LTD induced by low frequency stimulation was observed in control and Ccnd1 animals, but not in Pctk1 treated animals. In addition, neither Ccnd1 nor Pctk1 expression produced any detectable neuropathology. In contrast Tcf12 expressing animals displayed significant neurodegeneration in both CA1 and dentate gyrus. Several Tcf12 animals also developed tumors that appeared to be glioblastomas, suggesting that aberrant Tcf12 expression in the hippocampus is tumorigenic. Thus, behavioral experiments suggested that overexpression of Pctk1 and Ccnd1 produce a deficit in learning and memory, but electrophysiological experiments do not point to a simple mechanism. In contrast, the learning and memory deficits in Tcf12 animals are likely due to neuropathology associated with Tcf12 gene expression.
INTRODUCTION
Aging is associated with cognitive declines in both humans and rodents. Humans, as well as rodents, display a large variability between individuals in age-related impairments in learning and memory. The underlying causes for this variability remain unknown. The formation of long term memory requires transcription and synthesis of new proteins (Hernandez and Abel 2008). We have hypothesized that differential changes in gene expression between individuals may be responsible for the variability in memory-related behaviors in aged rodents. We previously used the Morris water maze (MWM) to segregate aged learning-competent and aged learning-impaired animals, and used microarray analysis to investigate the genome-wide transcriptional changes that occurred in both CA1 and dentate gyrus of rats that showed age-related learning impairment (AI) in MWM compared to rats that showed less cognitive impairment (superior learners (SL). We identified two sets of genes that are differentially expressed in CA1 and dentate gyrus, respectively, in AI and SL rats (Burger, Lopez et al. 2007; Burger, Lopez et al. 2008). Several genes that we identified had already been validated for their roles in learning and memory processes by other studies, but most remained to be investigated. In this study, we focus on validating the roles of three genes identified by differential transcription in CA1, which may play a role in cognition. The three genes were PCTAIRE protein kinase 1 (Pctk1, Cdk16), G1/S-specific cyclin D1 (Ccnd1), and Transcription factor 12 (Tcf12, HEB, HTF4). The messenger RNA levels for all three genes were upregulated in the CA1 region in aged, learning-impaired rats, suggesting that their elevated expression might contribute to learning impairment (Burger, Lopez et al. 2007).
Pctk1 is a member of the serine/threonine protein kinase family with a conserved cyclin-dependent kinase-like kinase domain and variable N- and C-terminal domains (Meyerson, Enders et al. 1992; Okuda, Cleveland et al. 1992). Although originally identified as a cdc2-like kinase, Pctk1 is not involved in the regulation of cell cycle progression, nor does it require cyclins for its activity (Graeser, Gannon et al. 2002). The expression of Pctk1 is ubiquitous in mammalian tissue with particular abundance in pyramidal neurons in the hippocampus (Besset, Rhee et al. 1999).
Ccnd1 (cyclin D1) is a key regulator of cell cycle progression that functions as a mitogenic sensor and allosteric activator of cyclin dependent kinases 4 and 6 (CDK4/6) (Sherr and Roberts 1999; Kozar and Sicinski 2005). Ccnd1/CDK4/6 holoenzyme phosphorylates the retinoblastoma protein (Rb) and promotes progression through the G1-S phase of the cell cycle. Ccnd1 is important for normal development of retina, mammary gland and cerebellum as homozygous deletion of Ccnd1 results in impaired development of these tissues (Fantl, Stamp et al. 1995; Kim, Pomeroy et al. 2000; Kozar and Sicinski 2005). Interestingly, Ccnd1 is also expressed abundantly in adult brain neurons such as pyramidal cells in the CA1 region of the hippocampal formation and SGZ zone of dentate gyrus. These regions are known to be sites of neurogenesis in adult brain; alternatively, Ccnd1 may have functions independent of promoting cell cycle progression in brain.
Tcf12 is believed to be a transcription factor. It is a member of the basic helix-loop-helix family of proteins that recognizes the DNA E-box motif (Zhang, Babin et al. 1991; Hu, Olson et al. 1992). It has been shown to form homoligomers or heteroligomers with myogenin, E12 and ITF2, and interacts with PTF-1 and RUNX1T1 and the Tcf12 regulator, ID1. Tcf12 has been shown to have a role in neuron differentiation and is upregulated in some tumors (Zhuang, Cheng et al. 1996; Uittenbogaard and Chiaramello 2002; O'Neil, Shank et al. 2004). All three of the genes we chose to study (Tcf12, Ccnd1, Pctk1) are abundantly expressed in hippocampal neurons (Allen 2004).
To investigate the potential involvement of these genes in learning and memory, we used targeted overexpression in the hippocampus with recombinant Adeno-associated virus (rAAV)-mediated gene transfer, an approach that had previously worked successfully for determining gene function in the hippocampus as well as the substantia nigra (Kirik, Rosenblad et al. 2002; Rex, Gavin et al. 2010). Our results showed that rAAV-mediated gene expression in hippocampus caused deficits in spatial working memory (SWM) performance in a radial arm water maze (RAWM) task in animals injected with Ccnd1, Tcf12 or Pctk1. Overexpression of Ccnd1 or Tcf12, but not Pctk1, also resulted in impairments in spatial reference memory (SRM) in the same task. Overexpression of Pctk1 had a significant effect on LTD but not on LTP, while Ccnd1 did not significantly affect neuronal plasticity in hippocampus of aged animals. In addition, expression of Pctk1 (but not Ccnd1 or Tcf12) produced deficits in the MWM and reverse MWM. Finally, although neither Ccnd1 nor Pctk1 produced significant pathology or neurodegeneration in the hippocampus, the third gene, Tcf12, induced measurable neurodegeneration that was accompanied by the formation of tumors.
MATERIALS AND METHODS
rAAV vectors
The coding sequences for Pctk1, Tcf12 and Ccnd1 were cloned from a rat brain cDNA library (Biochain). The primers for Pctk1 cloning were TCCAAGCTTCCACCATGGATCGGATGAAGAAGATCAAACG and ATCAAGCTTTCAAGCGTAGTCTGGGACGTCGTATGGGTAGAACTCGGTATCCACCAC ACGG. Pctk1 was tagged with hemagglutinin (HA) at its C-terminal. Pctk1 tagged with HA retains its kinase activity. The primers for Ccnd1 cloning were CGTTGGCGGCCGCCACCATGGAACAACAGCTCCTGTGCTGCGAAG and GGATGGTCGACTCAGATGTCCACATCTCGGACGTCGG. Ccnd1 was not tagged. The Tcf12 primers were: TCCAAGCTTCCACCATGAATCCCCAGCAGCAGCGCAT and ATCAAGCTTTCAAGCGTAGTCTGGGACGTCGTATGGGTACAGATGACCCATAGGGTT GGTTGTCT. Tcf12 was HA-tagged at its C-terminal end.
Coding sequences for Pctk1 and Tcf12 were cloned into an AAV cloning vector plasmid called pTR2 MCS at the HindIII site; Ccnd1 sequences were cloned into the NotI and SalI sites. Constructs were verified by DNA sequencing and immunoblotting using anti-Pctk1, anti-Tcf12 and anti-Ccnd1 antibodies against HEK 293 cell extracts from cells transfected with the constructs (not shown). An empty pTR2 MCS was used as null control construct. The pTR2 MCS (tr2), HA-tagged Pctk1 and Tcf12 constructs were serotyped with AAV8 capsid and the Ccnd1 construct was serotyped with AAV5 capsid during virus production as previously described (Zolotukhin, Potter et al. 2002). The recombinant viruses were generated and purified as described (Hauswirth, Lewin et al. 2000; Zolotukhin, Potter et al. 2002). rAAV particles are expressed as vector genomes (v/g)/ml. Vector genomes were quantitated using the dot plot protocol, with a probe for the chicken β-actin promoter, as described by (Hauswirth, Lewin et al. 2000; Zolotukhin, Potter et al. 2002). The titers were 3.4 × 1012, 2.3 × 1012, 2.4 × 1013, and 8.8 × 1012 vector genome per ml for tr2 null, Pctk1, Tcf12, and Ccnd1 and vectors, respectively.
Subjects and surgery
Three month old male Sprague-Dawley rats were injected with rAAV vector bilaterally in the hippocampus. Specifically, each animal was injected at two positions on each side of the brain with a total of 3 µl of vector at each position. At each position, 1 µl of vector was injected at each depth for total of 3 depths. The coordinates for injection at the first position were AP −3.48 mm; ML ± 2.2 mm; DV −3.7mm, −3 mm, and −2.8 mm. The coordinates for the injection at the second position were AP −5.28 mm; ML ± 5 mm; DV −7.5mm, −6 mm, and −4 mm. All surgical procedures were performed by using aseptic techniques and isoflurane gas anesthesia. Procedures involving animal subjects have been reviewed and approved by the Institutional Animal Care and Use Committee, and were in accordance with guidelines established by the U.S. Public Health Service Policy on Humane Care and Use of Laboratory Animals. Animals were group housed (2 per cage), maintained on a 12:12 hr light schedule, and provided ad lib access to food and water. Behavioral tests and electrophysiological experiments were carried out as shown in the timeline in Fig. 1A. Additional 3 month old rats were injected with Ccnd1, Tr2 and Pctk1 vector and taken for electrophysiological measurements at 1 month post-injection.
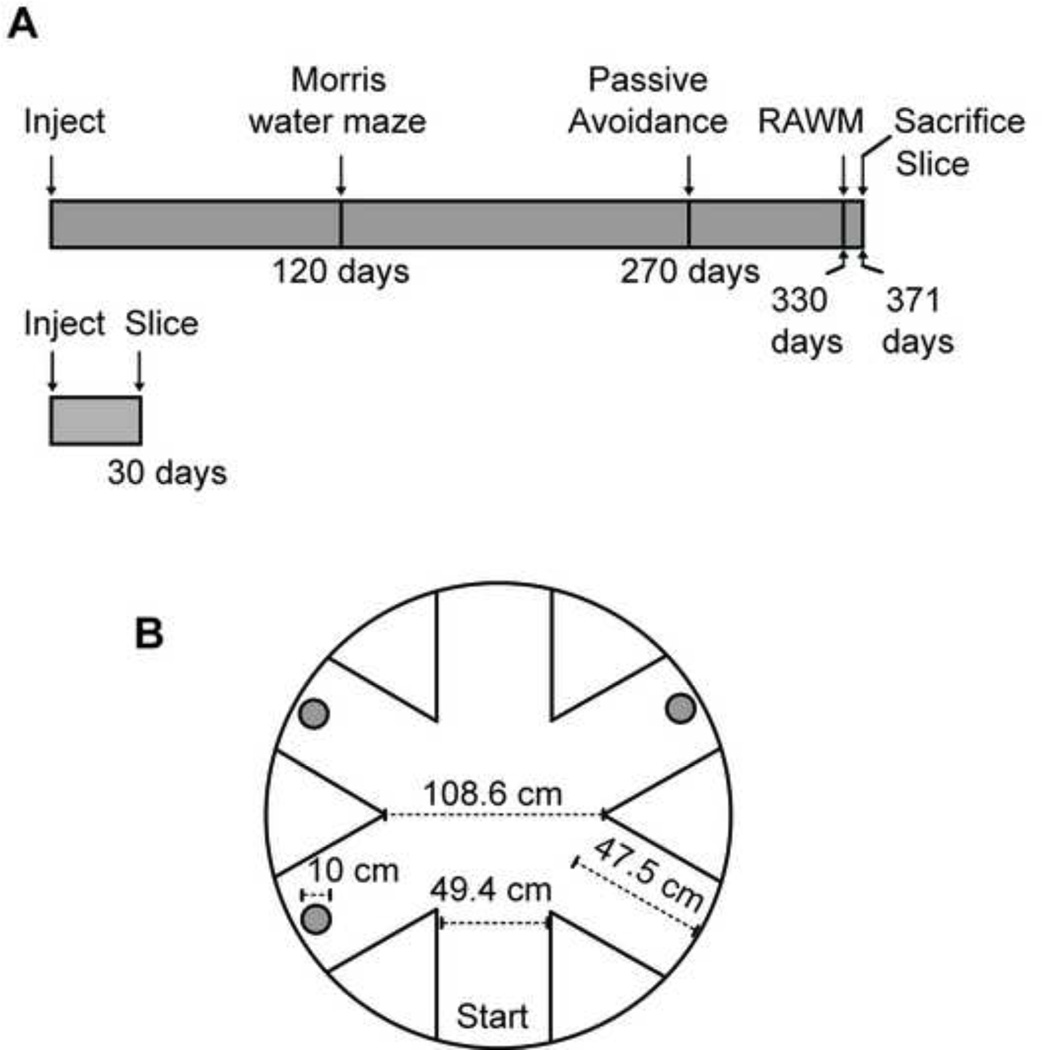
Fig. 1.
(A) Timeline for study. (B) Schematic and dimensions of 3 platform radial arm water maze used in the study.
Morris water maze
Four months after vector injections, animals received 8 days of 3 training sessions/day beginning in the morning of each day. Twenty-four hours after the last training session, animals received a probe trial in which the hidden platform was removed. During training, escape latencies were measured for each session. During training, animals were hand-guided to the platform if they did not find it by 90 s; these training intervals were given a 90 s value. Swimming distance was measured for the entire 24 trials for each animal. During the probe trial, percentage time in target vs non-target quadrants was determined over a 60-s interval. Paths swum by animals were recorded and analyzed using a computerized video monitoring system (EthoVision, Noldus Information Technology).
Passive avoidance behavior
Animals were trained in a two-chamber passive avoidance paradigm. They were placed in the lit compartment and allowed to enter the dark adjoining chamber. Animals that entered the second chamber received a mild foot shock (0.8 mA) for 1 sec. Rats were tested for latency to enter the dark chamber 24 hr later for up to 5 min. Statistical analyses used rank order nonparametric comparisons of latencies.
Radial arm water maze
Rats were tested in the radial-arm maze shown in Fig. 1B. The maze was constructed of aluminum, and was placed in a large pool of water in a black tank 2 m in diameter. No dye was added. It had black escape platforms (2 cm below the water surface) placed at the ends of three of the six arms. The distance from the water level to the top of the maze was approximately 20 cm. The temperature of water was 25°C. Each subject had different platform locations that were randomly determined, and that remained fixed throughout the experiment. There was no platform in the arm from which the rats was released. The testing room had salient extra-maze cues, including a door, solid black panels on a wall, black and white stripped panels on the opposite wall. In each trial, the rat was introduced into the start arm with its head pointing towards the tank wall. The rat had 90 s to locate a platform. If the allotted time expired, the subject was guided to the nearest available platform. Once a platform was found, the animal remained on it for 20 s, and was returned to its heated cage for 30 s. During the interval, the just-chosen platform was removed from the maze. The animal was then placed back into the start arm and allowed to locate another platform. A session consisted of this sequence of events repeated until all three platforms were located, resulting in a total of three trials per session. After the first day of training, rats were tested for 9 more days using the same procedure as day one. Behavioral testing took place between 9:30 am and 3:30 pm.
An arm entry is counted when the tip of a rat’s snout reaches a mark on the outside of the arm (~11 cm into the arm), or all four paws enter the arm. Working memory errors are the number of first and repeat entries into any arm from which a platform has been removed previously during the session and the number of entries into any arm with an existing platform that is not located by rats. Reference memory errors are the number of first entry into any arm that never contains a platform. In order to determine whether rats employed non-spatial strategies similar to thigmotaxis in MWM by simply going from one arm to an adjacent arm (chaining) during the RAWM task, a radial index was calculated. A radial index is defined as the mean of the angles formed by successively chosen arms (Roullet, Lassalle et al. 1993). When a rat swims out of an arm, it can enter the adjacent arm (60°), the second arm (120°), the third arm (180°), or it can reenter the same arm (0°). If a rat developed chaining behavior, its radial index would approach 60°. When it had no preference for a particular angle, the radial index would remain 90°.
Stereology
The unbiased stereological estimation of the total number of the cresyl violet stained neurons in hippocampus was performed by using the optical fractionator method, as described (Tran and Kelly 2003) with the MicroBrightfield Stereo Investigator System. Hippocampal subregions were outlined according to the atlas published in The Rat Brain (Paxinos, Watson et al. 1985). The estimate of the total number of neurons and coefficient of error due to the estimation was calculated according to the optical fractionator formula as described (Tran and Kelly 2003).
Immunoblotting and real-time PCR
Tissues were suspended in 300 µl of lysis buffer (50 mM Tris, pH 7.5, 0.15 M NaCl) containing protease mixture (0.1 mM PMSF, 0.5 µg/ml leupeptin, 0.7 µg/ml pepstatin A) (Roche) and homogenized for 10 s. Each aliquot was adjusted to a final concentration of 1% Nonidet P-40, 0.1% SDS, incubated on ice for 30 min, and centrifuged for 15 min at 4°C. Protein concentrations were determined by the Bradford protein assay. Fifty micrograms of each protein pool was separated on Bio-Rad precast 4–20% SDS/PAGE gradient gel, transferred to PVDF-LFP (Amersham) membranes, and immunoblotted. Mouse anti-Pctk1 (sc-53410, Santa Cruz Biotechnology) and rat anti-HA peroxidase high affinity (3F10, Roche) antibodies were used as recommended by the supplier.
For detection of rat Ccnd1 messenger RNA, rat hippocampal tissues were homogenized in TRIzol reagent according to the manufacturer’s manual (Invitrogen). After DNase I treatment, 500 ng of RNA was reverse-transcribed in 25 µl reaction using SuperScriptTM III first strand cDNA synthesis kit (Invitrogen). Real time PCR was performed on the MyiQ Single-Color Real-Time PCR Detection System (Bio-Rad) using IQ SYBR Green Supermix (Bio-Rad), 2 µl of cDNA, and Ccnd1 primers. All values obtained were normalized with respect to levels of β-actin mRNA. The primer pairs used for RT-PCR experiments were forward 5'-AGTTGCTGCAAATGGAACTG −3' and reverse 5'- TGGAGAGGAAGTGTTCGA G −3' for Ccnd1 gene, and forward 5'-CACTGCCGCATCCTCTTCCT-3' and reverse 5'-AACCGCTCATTGCCGATAGTG-3' for β-actin gene. All real-time PCR s were performed in duplicate using RNA from individual rats to give an average value for each animal. Serial dilutions of plasmid DNA containing Ccnd1 gene were used as standard templates. Relative quantification was performed using the ΔCt method.
Immunohistochemistry and pathology
Rat brain was harvested and fixed in 4% paraformaldehyde followed by freezing in optimal cutting temperature media. Frozen sections (40 um) were stained by immunohistochemistry and immunofluorescence using minor modifications of previously published methods (Campbell-Thompson, Dixon et al. 2009). Primary antibodies included anti-GFAP (Novus Biologicals, NB300141), anti-HA (sc-7932, Santa Cruz Biotechnology), Pctk1 (sc-53410, Santa Cruz Biotechnology) and Tcf12 (sc-23128, Santa Cruz Biotechnology). Secondary fluorescent antibodies were conjugated with Alexa Fluor (Invitrogen) fluorochromes (488 and 594); DAPI was used to counter stain nuclei (blue).
Electrophysiological recordings
The methods for hippocampal slice preparation have been published previously (Kumar and Foster 2004; Kumar and Foster 2007; Kumar, Thinschmidt et al. 2007; Bodhinathan, Kumar et al. 2010; Kumar 2010). Briefly, rats were anesthetized with isoflurane (Halocarbon Laboratories, River Edge, NJ) and swiftly decapitated. The brains were rapidly removed and the hippocampi were dissected. Hippocampal slices (~ 400 µm) were cut parallel to the alvear fibers using a tissue chopper. The slices were incubated in a holding chamber (room temperature) containing standard artificial cerebrospinal fluid (ACSF) (in mM): NaCl 124, KCl 2, KH2PO4 1.25, MgSO4 1.5, CaCl2 2.4, NaHCO3 26, and glucose 10. Thirty to sixty min before recording, 2–3 slices were transferred to a standard interface recording chamber (Harvard Apparatus, Boston, MA); the chamber was continuously perfused with standard oxygenated (95% O2, 5% CO2) ACSF at a flow rate of 2 ml/min. The pH and temperature were maintained at 7.4 and 30 ± 0.5°C, respectively. Humidified air (95% O2, 5% CO2) was continuously blown over the slices.
Extracellular synaptic field potentials from CA3-CA1 synaptic contacts were recorded with glass micropipettes (4–6 MΏ) filled with recording medium (ACSF). Two concentric bipolar stimulating electrodes (outer pole: stainless steel, 200 µm diameter; inner pole: Platinum/Iridium, 25 µm diameter, FHC, Bowdoinham, ME) were positioned approximately 1 mm from either side of the recording electrode localized in the middle of the stratum radiatum. A single diphasic stimulus pulse of 100 µsec was passed via stimulators (SD9 Stimulator, Grass Instrument Co, West Warwick, RI) to the Schaffer collateral commissural pathway, in order to evoke field potentials at 0.033 Hz. The signals were amplified, filtered between 1 Hz and 1 kHz, and stored for off-line analysis. Two cursors were placed around the initial descending phase of the excitatory post synaptic potential (EPSP) waveform, and the maximum slope (mV/ms) of the EPSP was determined by a computer algorithm that found the maximum change across all sets of 20 consecutively recorded points (20 kHz sampling rate) between the two cursors. For examination of synaptic plasticity, the stimulus current was adjusted to produce a response 50–60% of the maximal EPSP slope. Responses were collected for at least twenty minutes prior to pattern stimulation to insure a stable baseline before induction of synaptic plasticity. LTP was induced by employing 4 trains of 100 Hz (100 pulses, 10 ms apart, each train 10 sec apart). LTD was induced by using 1 Hz paired-pulse (1 Hz PP, with 50 ms inter-pulse interval, total 900 pulses) low frequency stimulation. Changes in transmission properties induced by patterned stimulation were calculated as the percent change from the averaged response collected during baseline. For examination of paired-pulse facilitation (PPF), a 50 ms inter-pulse interval was used. The PPF ratio was calculated by dividing the slope of the second synaptic response by the slope of the first response.
Statistical analysis
Analyses of variance (ANOVAs) were used with treatment effects and interactions as independent varaibles. For electrophysiological recordings F tests of main effects, t-tests for planned contrasts, and Fisher’s PLSD or Bonferroni post hoc comparisons were employed. A p-value of P < 0.05, was considered statistically significant when testing changes in synaptic response induced by pattern stimulation, as well as any possible differences between baseline and PPF ratio. The data were log transformed when appropriate. Statistical analyses were performed using StatView 5.0 (SAS Institute) and Prism 5 (GraphPad).
RESULTS
Behavioral Experiments
Morris Water Maze
Three month old rats were injected bilaterally into their hippocampus with rAAV virus expressing Pctk1, Ccnd1 or Tcf12. The timeline for the study is shown in Fig. 1A. Four months after rats were injected with rAAV virus they were tested for their performance in a MWM task along with uninjected, age-matched animals. AAV viral gene transfer has previously been shown to produce no pathology or cell loss when control genes such as green fluorescent protein (GFP) have been expressed in rodent brain at the input doses used in this study (Gorbatyuk, Li et al. 2008). In addition, previous work has shown that expression of an irrelevant gene such as GFP in the hippocampus using rAAV mediated transduction had no effect on memory formation in rats (Rex, Gavin et al, 2010). To be certain that virus injection itself did not induce behavioral deficits, we compared uninjected animals with animals that had been injected with virus carrying a null gene, tr2. Tr2 was an empty cloning vector with multiple stop codons in all possible reading frames that was driven by the same CMV/β-actin hybrid promoter that was used to drive expression of Ccnd1, Pctk1 and Tcf12. Although it was expected to make mRNA at the same rate as the experimental constructs, no protein product could be made from tr2. Thus, tr2 injection controlled for effects from the surgery, injection of virus and production of a foreign RNA.
When the data were analyzed by two-way ANOVA, there was a significant main effect of treatment for latency between experimental groups (Table 1, Fs4,42 = 2.87, P = 0.0346), with no interaction between treatment and day of treatment (Fs28,294 =0.81, P = 0.7378). A two-way repeated measures ANOVA analysis showed no difference in performance between animals from the uninjected, age-matched group and those receiving the tr2 null vector (Fig. 2A and Table 1, for gene Fs1,112 = 0.8968, P = 0.358). Because there was no difference in performance between animals from the uninjected, age-matched group and those receiving the tr2 null vector, they were combined as a single control group to simplify subsequent comparison with each experimental group. See Supplementary Table 1 for additional comparisons between each experimental group (Pctk1, Ccnd1, Tcf12) and each control group (uninjected, tr2).
Table 1.
P Values for Treatment Comparisons*
| Control Treatment |
Experimental Treatment |
MWM Latency |
Reverse MWM Latency |
RAWM Reference Memory |
RAWM Working Memory |
|---|---|---|---|---|---|
| control** | pctk1 | 0.044 | 0.004 | 0.638 | 0.003 |
| control | ccnd1 | 0.054 | 0.219 | 0.009 | <0.0001 |
| control | tcf12 | 0.067 | 0.08 | 0.019 | 0.004 |
| unlnjected tr2 | pctkl ccndl tcf 12 | 0.035 | 0.046 | 0.001 | <0.0001 |
| uninjected | tr2 | 0.358 | 0.341 | 0.607 | 0.882 |
two-way repeated measures ANOVA P values for main effect of the four treatment groups (pctkl, ccndl, tcf 12, tr2) compared to the indicated control groups; shaded values: P<0.05
control group = uninjected + tr2 animals
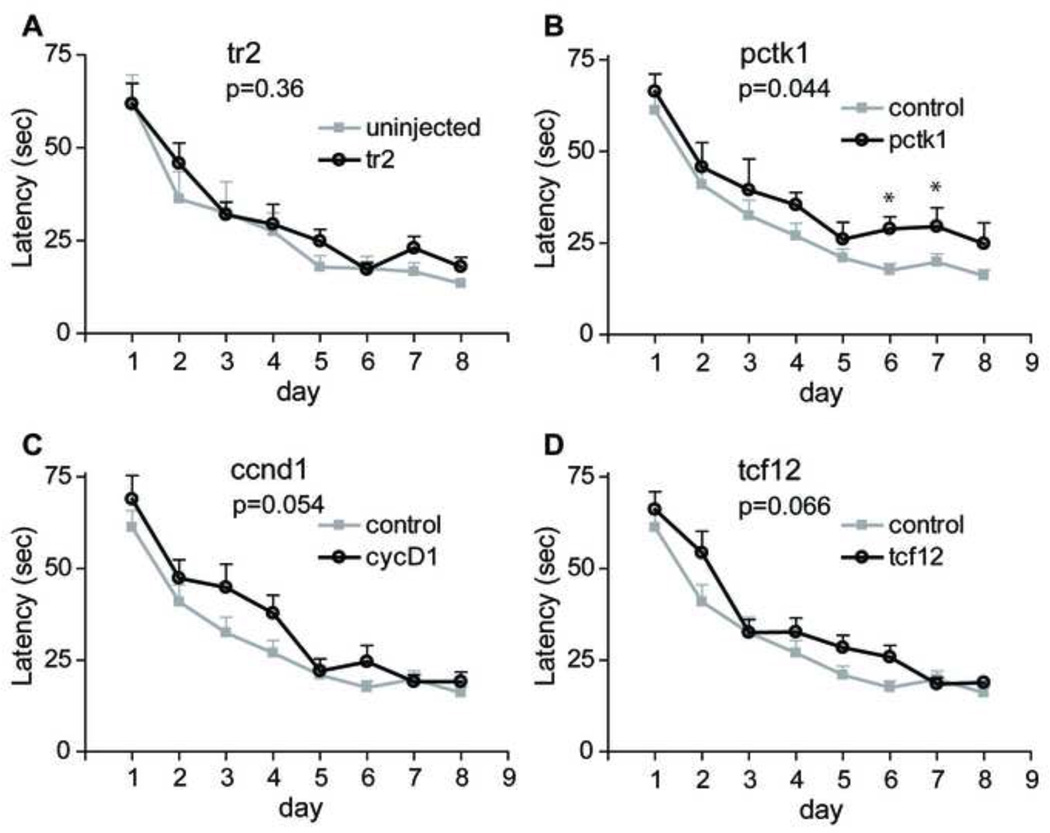
Fig. 2.
Morris water maze task performance. Each day block represents the average of three trials. Each data point represents the mean ± SEM of time to find the platform. (A) There was no significant difference between uninjected (n = 8) and tr2 vector-injected rats (n = 10). For ease of further analysis, these 2 groups were combined and designated as the control group. (B) A significant difference was found between control (n = 18) and Pctk1 (n = 10) animals in the acquisition phase of the MWM task. (C, D) Comparison of control and Ccnd1 (n = 9) or control and Tcf12 animals (n = 10) did not reach significance. Asterisks indicate a significant difference between control and treatment animals on a particular day by Bonferroni post hoc comparison (*, P < 0.05). See text and Table 1 for additional details.
Comparison of each of the experimental groups (Pctk1, Ccnd1, and Tcf12) with the control group in a two-way repeated measures ANOVA (Fig. 2B–D) suggested that there was a significant difference for latency between Pctk1 and controls (Fig. 2B, Fs1,175 = 4.494, P = 0.044), but not between Ccnd1 and controls (Fig. 2C, Fs1,168 = 4.14, P = 0.054) or between Tcf12 and controls (Fig. 2D, Fs1,175 = 3.709, P = 0.066). In addition, during the MWM probe trial, all treatment groups were able to recall the position of the platform. There was no difference in the time spent in the target quadrant among treatment groups (one-way ANOVA, F3,46 = 0.5317, P = 0.663, Supplemental Fig. 1A) or in the number of target crossings (F3,43 = 2.034, P = 0.123, Supplemental Fig. 1B). In addition, there was no difference in the swimming speed between groups (Supplemental Fig. 2).
Reverse Morris Water Maze
A subsequent reverse MWM test also showed a significant difference between all groups for the gene in a two-way ANOVA (Table 1, F4,42 = 2.65, P = 0.046) and a significant interaction between time and gene (F12,126 = 1.97, P = 0.032). As in the MWM above, two-way ANOVA comparison of the two control groups revealed no significant difference between animals that had been uninjected or injected with the null tr2 gene (Supplemental Fig. 3A and Table 1, Fs1,48 = 0.9635, P = 0.341), and these groups were pooled and used as the control group for further comparisons. Comparison of the control group with individual genes (Supplemental Fig. 3B–D and Table 1) in the reverse MWM showed a significant difference for Pctk1 (Fs1,78 = 9.966, P = 0.004), but Tcf12 (Fs1,78 = 5.28, P = 0.08) and Ccnd1 (Fs1,75 = 1.593, P = 0.219) arewere not significant. Additionally, during the probe trial, there was no significant difference between groups in target crossings (one-way ANOVA, F3,38 = 1.265, P = 0.300, Supplemental Fig. 1C) or time spent in the target quadrant (F3,38 = 1.51, P = 0.341, Supplemental Fig. 1D).
Finally, five months after the MWM task, the animals were evaluated for their performance in the hippocampus-dependent passive avoidance task. One day after training, animals from all groups spent a similar amount of time in the light chamber (data not shown).
Radial Arm Water Maze
Taken together, the data suggested that Pctk1 animals had a deficit in acquiring the platform in the MWM. The failure to detect a robust difference in MWM and passive avoidance tasks between control groups and the Ccnd1 and Tcf12 groups raised the possibility that the difference between these treatment groups might be small, and that it might only be revealed in a more sensitive behavioral performance test. We, therefore, tested the same rats in a radial arm water maze that contained six arms in a water tank with 3 hidden platforms in 3 arms other than the start arm (Fig. 1B). The platform was removed after being located by the rats in each session. To perform well in this task, the rats needed to remember the position of each hidden platform (reference memory) as well as whether they had visited a platform in previous sessions (working memory). Therefore, this testing scheme allowed us to measure the SWM as well as SRM performance by rats in a single task.
Reference memory errors
Analysis of reference memory errors of all groups in a two-way ANOVA revealed a significant effect of gene (F4,38 = 5.59, P = 0.001, Table 1) with no interaction between gene and session day (F32,304 = 1.26, P = 0.1613). Two-way (treatment and sessions) repeated measures ANOVA comparison of the two control groups (tr2 and uninjected) confirmed that there was no significant difference between the two controls (Fig. 3A and Table 1, Fs1,120 = 0.2760, P = 0.607) and these two groups were pooled to simplify further comparisons. (See Supplemental Table 1 for separate comparisons with each control group.)
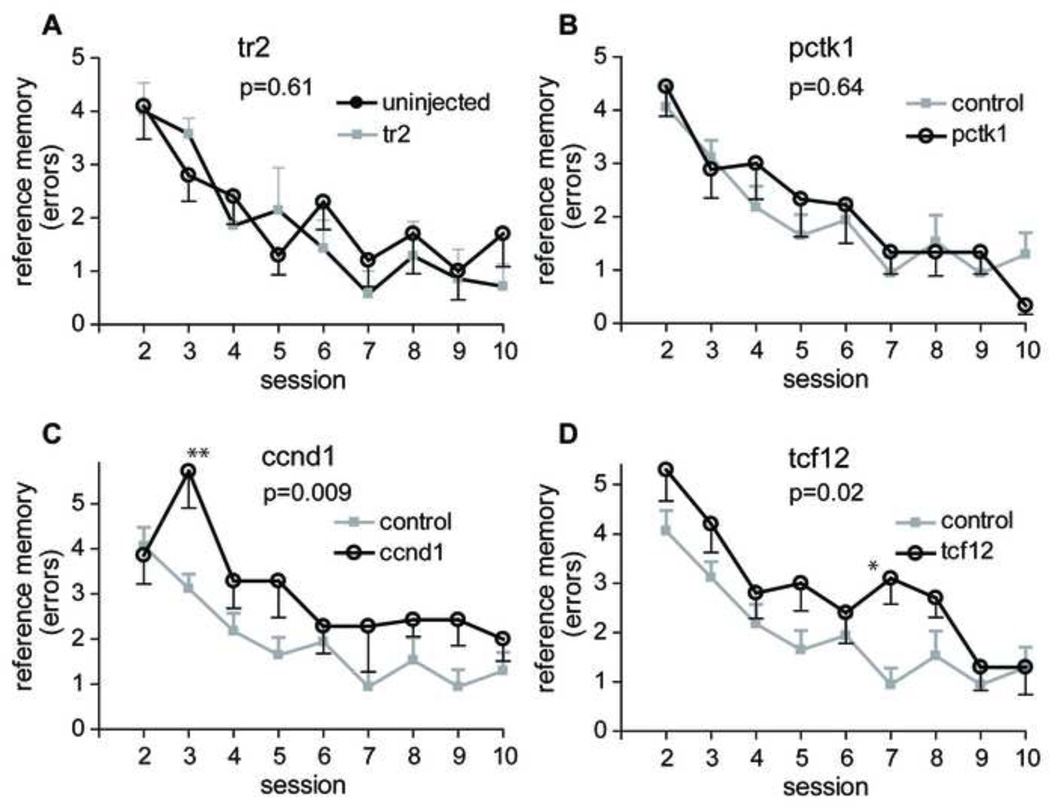
Fig. 3.
Spatial reference memory performance in radial arm water maze. Each data point represents the mean ± SEM of spatial reference memory errors committed in each session. (A) There was no difference between tr2 vector (n = 10) injected and uninjected animals (n = 7). These 2 groups were combined as a control group. (B) No significant difference was found between Pctk1 (n = 10) and control animals (n = 17). (C, D) A significant difference was found between control animals and the Ccnd1 (n = 7, panel C) or Tcf12 (n = 10, panel D) treatment groups. Asterisks indicate a significant difference between control and treatment animals on a particular day by Bonferroni post hoc comparison (*, P < 0.05; **, P < 0.01). See text and Table 1 for additional details.
Direct comparison between each gene and the control group (Fig. 3B–D, Table 1) revealed that there was a significant difference in the number of reference memory errors between control and Ccnd1 rats (Fs1,176 = 8.117, P = 0.009) or control and Tcf12 rats (Fs1,200 = 6.323, P = 0.019), but surprisingly, not between control and Pctk1 rats (Fs1,192 = 0.2276, P = 0.638). For each of the genes there was no significant interaction between gene and session day.
Working memory correct errors
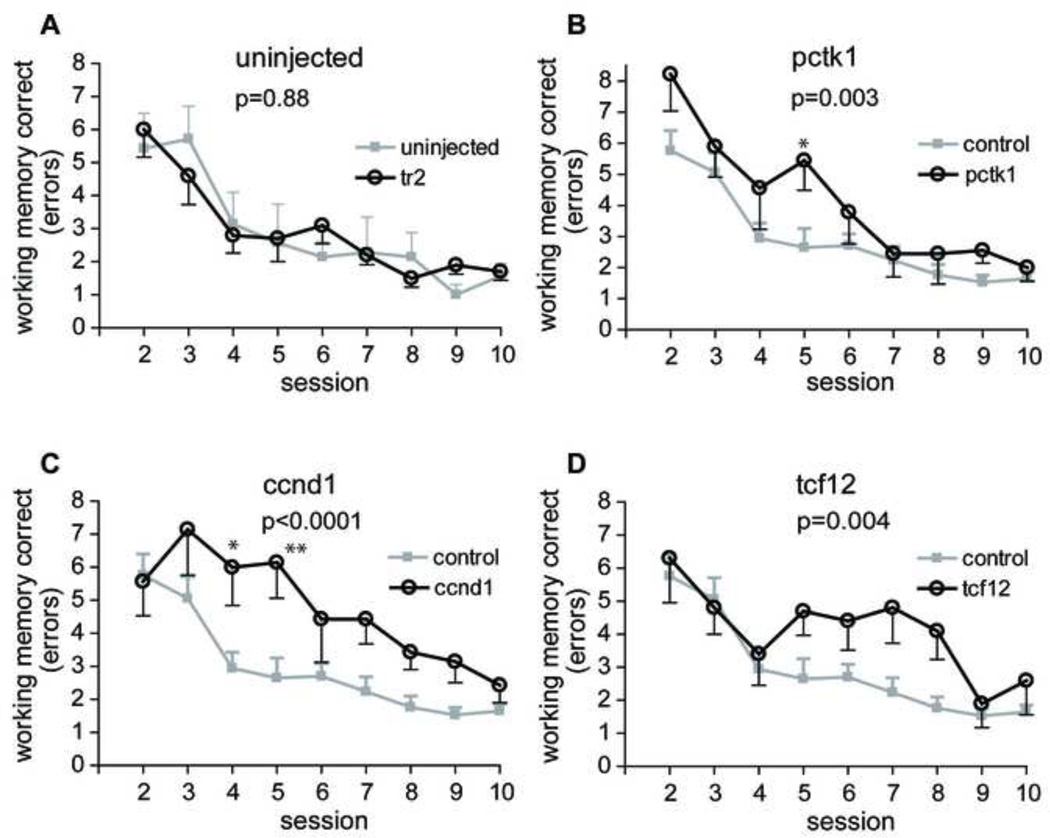
Analysis of working memory correct errors of all groups in a two-way ANOVA revealed a significant effect of gene (Table 1, F4,38 = 8.33, P < 0.0001) with no interaction between gene and session day (F32,304 = 0.93, P = 0.5838). Two-way (treatment and sessions) repeated measures ANOVA comparison of the two control groups (tr2 and uninjected) confirmed that there was no significant difference between the two controls (Fig. 4A, Fs1,120 = 0.0227, P = 0.882) and these two groups again were pooled for further comparisons. Comparison of each experimental group with the control revealed a significant difference between the control group and each of the experimental groups, Pctk1 (Fig. 4B, Fs1,192 = 10.77; P = 0.003), Ccnd1 (Fig. 4C, Fs1,176 = 25.42; P < 0.0001), and Tcf12 (Fig. 4D, Fs1,200 = 10.10; P = 0.004). (See Supplemental Table 1 for separate comparisons with each control group.)
Fig. 4.
Spatial working memory performance in radial arm water maze. Each data point represents the mean ± SEM of spatial working memory correct errors committed in each session. (A) There was no difference between tr2 vector (n = 10) injected and uninjected animals (n = 7). These 2 groups were combined as control group. (B, C, D) Significant differences were found between control animals (n = 17) and Pctk1 (n = 9, panel B), or Ccnd1 (n = 7, panel C), or Tcf12 (n = 10, panel D). Asterisks indicate a significant difference between control and treatment animals on a particular day by Bonferroni post hoc comparison (*, P < 0.05; **, P < 0.01). See text and Table 1 for additional details.
All groups significantly reduced the number of errors with number of sessions in all memory categories (Fs8,312 = 20.36, and 14.10 for reference memory and working memory errors, respectively; Ps for session < 0.0001 for both reference and working memory correct). We concluded that all three genes had an effect on working memory but only Ccnd1 and Tcf12 had an effect on reference memory.
To determine if the rats were solving the maze by simply moving from one arm to the next adjacent arm (chaining), we calculated the radial index scores over sessions for each treatment (Supplemental Fig. 4). The radial index was 90° or greater for both control groups and for Pctk1 and Ccnd1, suggesting that chaining was not occurring. There was no significant main effect of treatment (Fs2,248 = 1.061, P = 0.358). There also was no significant main effect of sessions (Fs8,248 = 0.5596, P = 0.810), nor was there a significant main effect of treatment × sessions interaction (Fs16,248 = 0.9821, P = 0.476).
Taken together, the RAWM data suggested that all three genes produced a learning deficit when overexpressed in hippocampus. Ccnd1 and Tcf12 produced robust deficits in both working and reference memory; in contrast, Pctk1 affected only working memory.
Gene Expression and Pathology
One week after RAWM, rats were sacrificed to confirm gene expression in the injected tissue, for electrophysiology assays and for pathological examination. Gene delivery of Pctk1 resulted in expression of the HA tagged Pctk1 and a significant increase in the total level of Pctk1 protein in rat hippocampus as judged by immunoblotting of tissues with HA and Pctk1 antibodies (Supplemental Fig. 5). The level of expression was constant over a broad region of the hippocampus and was similar between animals (Supplemental Fig. 6A). Interestingly, we could not detect endogenous Pctk1 expression under current experimental conditions, probably due to the low endogenous protein levels (Supplemental Fig. 5, Fig. 6A). Attempts to detect Ccnd1 protein in rat hippocampus after gene delivery were not successful due to high backgrounds (not shown). However, real-time PCR using primers specific for Ccnd1 mRNA, showed a seven-fold increase in the levels of Ccnd1 transcript in the hippocampus of Ccnd1-injected animals compared to controls (Supplemental Fig. 6C). Immunoblots of horse fibroblast cells transduced with Ccnd1 vector using anti-Ccnd1 antibody confirmed that gene transfer results in Ccnd1 protein expression (Supplemental Fig. 6B, C).
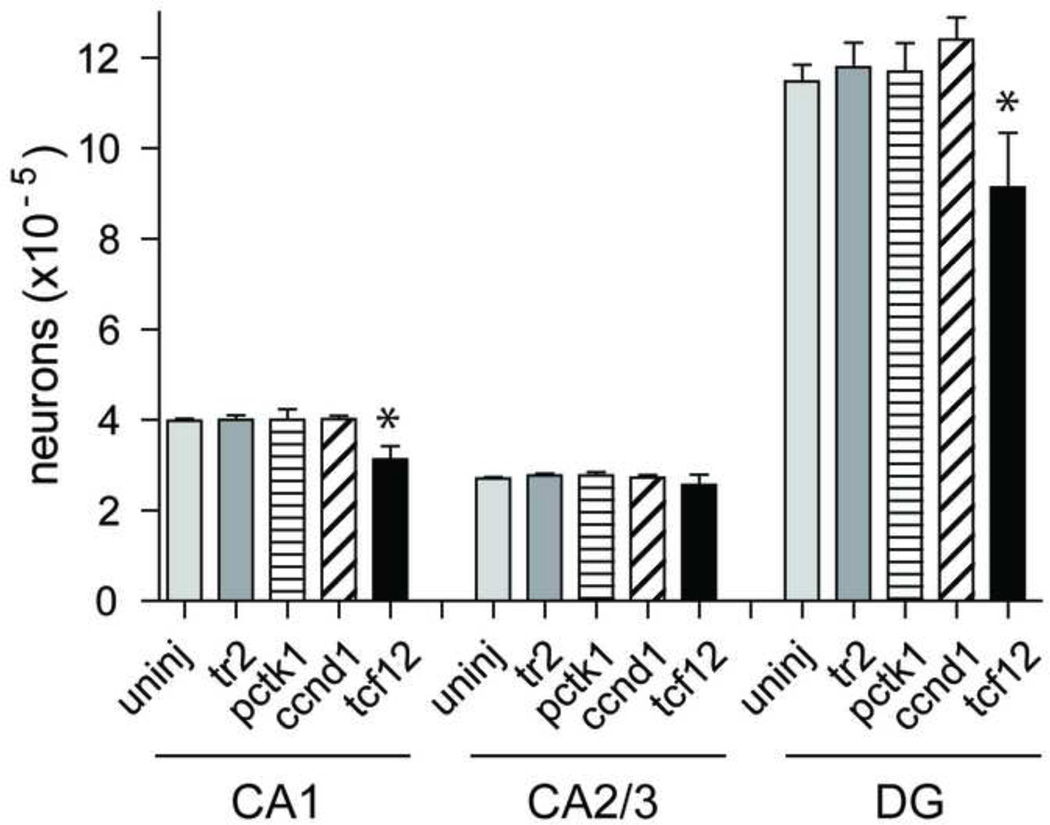
Fig. 6.
Unbiased stereological estimate of the total number of cresyl violet stained (mean ± SEM) pyramidal neurons in CA1 and CA2/3, and granule cells in dentate gyrus (DG) of uninjected rats (n=5), and tr2 (n = 4), Pctk1 (n = 5), Ccnd1 (n = 4) and Tcf12 (n = 4) injected rats. Each bar represents the mean ± SEM for each group. In a one way ANOVA analysis, Tcf12 was significantly different from either control in CA1 and DG (*, P < 0.05)
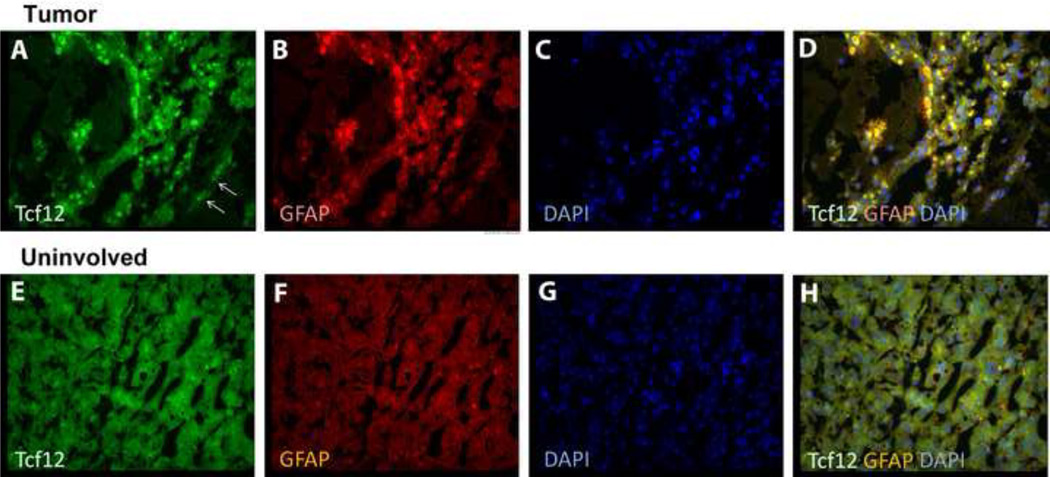
When we examined tissues at one year post injection, both the Ccnd1 and Pctk1 expressing vectors showed no overt pathology at one year post injection (not shown). In contrast, three of 10 rats that had been injected with Tcf12 expressing virus developed tumors in the area of injection. All 3 tumors were located between hippocampus and thalamus. Although the injection site was hippocampus, there were cystic epithelial features in one rat. All three tumors showed varying degree of vascularization/hemorrhage, with cells primarily neuronal and with no to very infrequent mitotic bodies observed. Immunohistological staining of the tumors with Tcf12 antibody revealed that they were expressing Tcf12. Two of the rat brain tumors were positive for glial fibrillary acidic protein (GFAP) indicating glial cell-origin and therefore glioma or astrocytoma are the likely tumor type (Fig. 5). In contrast to nearby non-tumor tissue, Tcf12 expression appeared to localize to nuclei in tumor tissue (which is its normal location) and resulted in the accumulation of autofluorescent material. One tumor was not positive for GFAP (not shown), yet was positive for Tcf12 indicating some heterogeneity, most likely at the level of transduced cells and subsequent tumorgenesis.
Fig. 5.
Elevated Tcf12 expression was associated with tumors formed in Tcf12 vector-injected rat brains. Tumor tissue (A–D) and nearby non-tumor (uninvolved) tissue (E–H) was stained with Tcf12 antibody (A, E, green), GFAP antibody (B, F, red), DAPI (C, G, blue) or merged (D, H). Increased Tcf12 expression appeared to be localized inside nuclei (panels A, C, D compared to E, F, H) and was associated with activation of astrocytes as shown by strong GFAP staining (panels B and D). In contrast, there was no detectable Tcf12 expression that colocalized with GFAP staining in normal brain tissues (panels E-H) and Tcf12 staining did localize to nuclei (panels E, G, H).
The occurrence of tumors raised the question of whether gene expression of Tcf12 or the other two genes, Ccnd1 or Pctk1, might be exerting their effect by reducing the number of viable cells in the injected hippocampus. To determine this we used non-biased methods to count the number of neuronal cells in the CA1, CA2/3 and dentate gyrus regions of uninjected, tr2 injected and experimental animals (Fig. 6). There was no significant difference in the number of surviving neurons when Pctk1and Ccnd1 injected animals were compared to uninjected and tr2 animals (one-way ANOVA: P = 0.999 for CA1; P = 0.743 for CA2/3; P = 0.719 for DG). In contrast, animals expressing Tcf12 had lost approximately 20–25% of their neurons in CA1 and dentate gyrus (Fig. 6) and were significantly different from the tr2 and uninjected controls (oneway ANOVA: Fs3,18= 10.89, P = 0.001 for CA1 and F3,18 = 4.024, P= 0.024 for DG).
We concluded that the behavioral deficits seen for Tcf12 injected animals could at least in part be the result of the pathology (tumors and loss of cells) that resulted from Tcf12 gene expression. In contrast, the absence of pathology in Ccnd1 and Pctk1 animals suggested that behavioral deficits in these animals were the result of a functional defect due to gene expression. These animals, therefore, were chosen for further electrophysiological analyses.
Synaptic Transmission and Plasticity
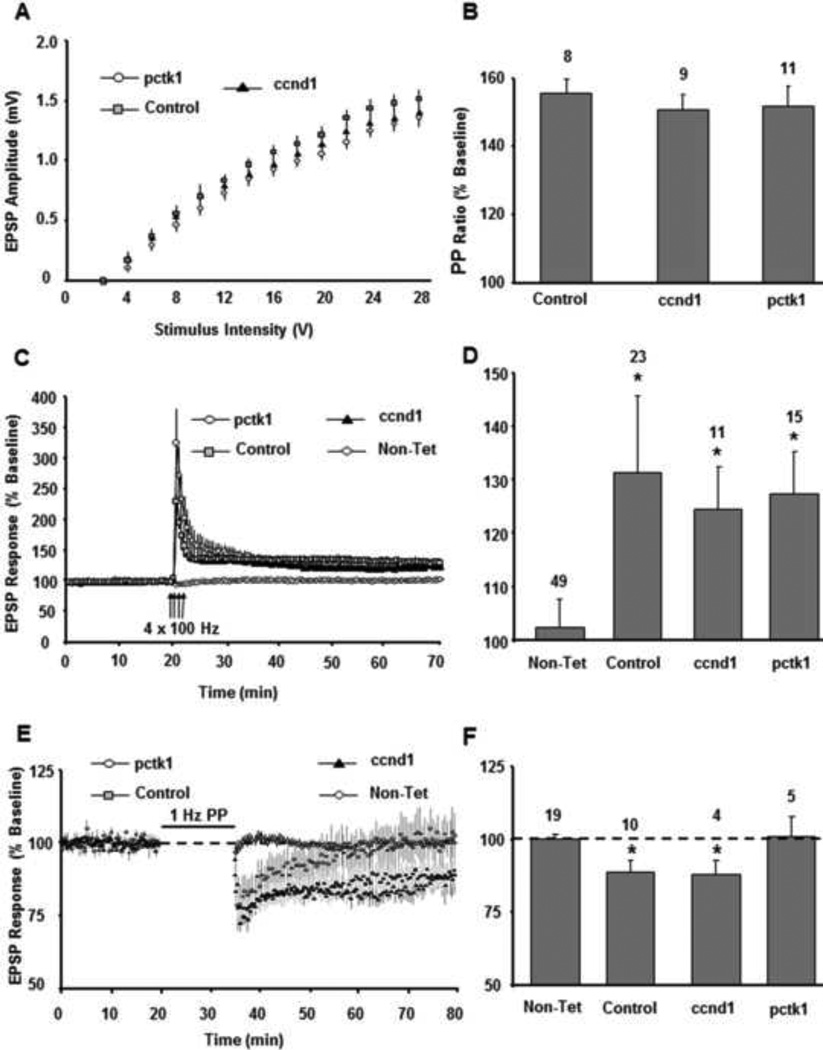
To evaluate the influence of enhanced expression of Ccnd1 and Pctk1 genes on synaptic function, 3 month old rats were injected with null, Pctk1 or Ccnd1 vectors. One month after injection, when gene expression reaches a maximum plateau (Reimsnider, Manfredsson et al. 2007), the input-output synaptic response and level of LTP and LTD were examined in slices near the injection site. Examination of input-output curves of the synaptic response indicated no group difference in baseline synaptic strength (Fig. 7A). Furthermore, paired-pulse stimulation resulted in a similar level of PPF for controls (155.9 ± 34.2, n = 8), Ccnd1 (150.9 ± 4.5, n = 9), and Pctk1 (152.2 ± 5.8, n = 11) treated animals (Fig. 7B), suggesting that expression of Ccnd1 or Pctk1did not influence short-term plasticity mechanisms. Pattern stimulation to induce LTP resulted in an increase in the synaptic response compared to the control path (P < 0.05) in each group and the level of LTP was similar across the three groups: control (131.4 ± 3.5, n = 23, P < 0.0001), Ccnd1 (124.5 ± 6.1, n = 11, P < 0.0002), and Pctk1 (127.3 ± 7.6, n = 15, P < 0.0001) (Fig 7C & D). Similar to young animals, middle aged animals, (13–15 months, 12–13 months post injection) showed no group differences in LTP induced by theta burst pattern stimulation; the magnitude of LTP between control and experimental animals were not different (Supplementary Fig. 7).
Fig. 7.
Effect of expression of Ccnd1 and Pctk1 on hippocampal synaptic function. A) Input– output curves of the baseline EPSP response vs. stimulus intensity for control (filled square, n = 45 slices), Ccnd1 (filled triangle, n = 23 slices), and Pctk1 (filled circle, n = 25 slices) treated rats. Each point represents the mean ± the SEM. B) The bar diagram shows the mean PPF ratio for control, Ccnd1, and Pctk1 treated rats. C) Time course of changes in the field EPSP obtained from hippocampal slices 20 min before and 50 min after stimulation to induce LTP for the control (gray square, Ccnd1 (filled triangle), and Pctk1 (open circle) treated slices. A non-tetanized control path (Non-Tet, open diamond) was used to insure that changes in EPSP were specific to pattern stimulation and not due to change in slice health. D) Bar diagram showing the average magnitude of LTP during the last 5 min of recording for control non-tetanized path, control, Ccnd1, and Pctk1 treated animals. The number above each bar indicates number of animals recorded for each group at each point. E) Time course of the field EPSP measurements obtained from hippocampal slices 20 min before and 45 min after stimulation to induce LTD for the control, Ccnd1, and Pctk1 treated slices. F) Bar diagram showing the average magnitude of LTD during the last 5 min of recording for control not-tetanized path, controls, Ccnd1 and Pctk1 treated animals. Asterisk represents significant difference between tetanized and not-tetanized control paths. For each panel, error bars equal SEM.
In contrast, when we analyzed LTD-inducing stimulation in middle aged animals (13–15 months, 12–13 months post injection), we found decreased synaptic responses of controls (88.9 ± 3.9, n = 10, P = < 0.007) and animals injected with Ccnd1 (87.9 ± 5.2, n = 4, P < 0.03), when examined 45 min following induction (Fig 7E & F). In contrast, the synaptic response of Pctk1 (101.4 ± 6.8, n = 5) treated animals was not different from the not-tetanized control path (100.3 ± 1.7, n = 19). The results suggest that enhanced expression of Pctk1 prevents induction of LTD, while Ccnd1 over expression has no influence on this form of synaptic plasticity.
DISCUSSION
The genes studied in this report were originally identified by microarray studies; they showed elevated expression in aged (24 months) learning impaired rats compared to aged learning superior animals. That study identified a limited number of genes in CA1 (~50), many of which had not previously been associated with learning and memory deficits, and none of which were elevated in the nearby dentate gyrus region (Burger, Lopez et al. 2007; Burger, Lopez et al. 2008). We reasoned that an increase in expression of these genes might also produce a learning deficit at younger ages (7–14 months). To see if we could validate these genes, we used a vector transfer protocol that has previously been successful in studies of learning and memory and Parkinson Disease (Klugmann, Wymond Symes et al. 2005; Gorbatyuk, Li et al. 2008). AAV vectors quantitatively transduce the neurons near the site of injection and then slowly ramp up expression of the transferred gene until expression plateaus at 1 month post-injection. Expression then remains constant for the lifetime of the animal (Peel and Klein 2000; Reimsnider, Manfredsson et al. 2007). Expression levels are typically 2–4 times endogenous levels (Gorbatyuk, Li et al. 2008; Gorbatyuk, Li et al. 2010), which is two to three times higher than the elevations detected in aged animals by microarray in our previous study (Burger, Lopez et al. 2007). Thus, this approach allowed us to independently evaluate the effect of overexpressing each gene on learning and memory.
All three of the genes chosen for this study displayed significant deficits in learning and memory in some but not all of the behavioral tests that were tried, and each of the three genes revealed a unique behavioral profile (Table 1). Pctk1 showed a significant difference in latency to learn the position of the platform in the conventional MWM and reverse MWM, and also showed a defect in working memory in the RAWM (Table 1). Pctk1 did not, however, display any significant difference in RAWM reference memory. The effect of Pctk1 is likely due to hippocampal dysfunction rather than a sensorimotor deficit. In general, sensorimotor deficits result in thigmotaxis and increased latencies, primarily during the early acquisition trials (Devan and White 1999; Dolleman-van der Weel et al., 2009). In contrast, deficits in Pctk1 animals emerged during later trials, suggesting impaired hippocampal-dependent spatial learning (Morris et al., 1990). In contrast to Pctk1, both Ccnd1 and Tcf12 both showed robust defects in RAWM reference and working memory, but did not reach significance when tested in the conventional MWM. Finally, Pctk1 and Ccnd1 expressing animals were tested for LTP and LTD. The results suggested that Pctk1 overexpression may result in reduced LTD, but no other significant differences for Pctk1 or Ccnd1 expressing groups compared to control groups was found.
The deficits in MWM, SWM and SRM are not likely the results of sensorimotor deficit because all rats had similar swimming speed (Supplemental Fig. 2) and learned the task as indicated by the decreasing number of errors in both working and reference memory during sessions. Further, the radial index scores for all groups in the RAWM were well above 60°, indicating a lack of chaining behavior (Roullet, Lassalle et al. 1993) and supporting the notion that behavioral deficits were the results of learning and memory impairments (Supplemental Fig. 4).
We note that although there was a clear effect of Pctk1 on latency in the MWM, there was no significant difference in the subsequent probe tests for any group, suggesting that all groups successfully learned the platform position (Supplemental Fig. 1). Additionally, none of the genes showed an effect in the passive avoidance test (not shown). In contrast, many previous experiments with aged animals, including our own (Burger, Lopez et al. 2007; Burger, Lopez et al. 2008) have shown an effect on memory acquisition in the probe trials and in passive avoidance tests. The most likely explanation is that learning deficits in aged animals are the result of multiple gene expression changes. Our previous microarray studies identified at least 135 genes whose expression pattern was significantly modified in aged rodents in CA1 or DG. In contrast, the study described here changed the expression of only one gene at a time, and only at an earlier age, when all other genes were likely to be expressed at normal levels, and possibly compensated for the engineered defect. In spite of this limitation, the effect of Ccnd1 and Pctk1 on memory in some of the behavioral tests was quite robust.
Tcf12
Tcf12 is a basic helix loop helix (bHLB) transcription factor that binds to E box sequences (Zhang, Babin et al. 1991; Hu, Olson et al. 1992). It forms homo and heteroduplex complexes with other bHLB factors (Tcf3, E12, myogenin, ITF2, RUNX1T1, TAL1) and can act both as a repressor and activator of transcription depending on the cellular context. A family of proteins (Id1-Id3) retain the protein interaction loop but not the DNA binding domain and act as inhibitors of the Tcf proteins. Tcf12 has been extensively characterized as a key factor controlling B and T cell progenitor expansion and differentiation as well as myogenesis (Quong, Romanow et al. 2002; Parker, Perry et al. 2006). In T cells it is involved in TCRα and TCRβ rearrangement. Tcf12 knockout mice die shortly after birth (Uittenbogaard and Chiaramello 2002). Relatively little is known about Tcf12 in neuronal tissues. Its pattern of expression suggests that it may be involved in neuronal stem cell and progenitor cell proliferation or may be necessary to maintain an undifferentiated state (Uittenbogaard and Chiaramello 2002). In knockout mice heterozygous for Tcf12, Tal1 induced T cell acute lymphoblastic leukemia was accelerated, suggesting that Tcf12 normally inhibited genes that lead to leukemia induction (O'Neil, Shank et al. 2004). In contrast, our results suggest that its expression may lead to neurodegeneration and tumor formation in the adult rodent brain. AAV has been used to deliver genes in many animal models, and has been used in several clinical trials that address neurodegenerative diseases (Manfredsson and Mandel 2010). To date, there has been no evidence of vector related toxicity in the brain. Tcf12 is the first example of an AAV delivered gene product that induced brain tumors.
Clearly, the loss of neurons in CA1 and dentate gyrus and possibly tumor formation are the most likely mechanisms for the learning deficits seen in the RAWM with Tcf12 injected animals. However, since we did not determine the time course of neurodegeneration, we cannot rule out a direct effect on learning and memory. In PC12 cells, overexpression of Tcf12 has been shown to repress the expression of the low-affinity neurotrophin receptor p75 gene (Uittenbogaard and Chiaramello 2002), which could be responsible for the neurodegeneration observed in CA1 and dentate gyrus.
Pctk1 and Ccnd1
Gene transfer of Pctk1 and Ccnd1 into rat hippocampus did not cause neurodegeneration, as indicated by the unaltered numbers of hippocampal neurons between treatment groups (Fig. 6). The gross morphology of pyramidal neurons in CA1, CA2/3 subregions and granule cells in dentate gyrus was also unaltered in Pctk1 and Ccnd1 rats compared to control animals (unpublished observation). These results suggest that the memory deficits that we observed in Pctk1 and Ccnd1 rats were due to altered gene expression and an apparent functional role for Pctk1 and Ccnd1 in learning and memory. Our results validate our previous microarray study that showed elevated transcript levels of Pctk1 and Ccnd1 in the hippocampus of aged, learning-impaired rats. The fact that increased levels of these genes also induce a learning deficit in young and middle-aged animals suggests that they are likely to play a role in learning and memory at all ages.
Pctk1
Pctk1 is abundantly expressed in pyramidal neurons (Besset, Rhee et al. 1999) and has been shown to be a negative regulator of N-ethylmaleimide-sensitive fusion (NSF) protein (Liu, Cheng et al. 2006). NSF is required for the disassembly and recycling of SNARE proteins during the interval between exocytosis and endocytosis, an event important for maintaining normal neurotransmitter release in presynaptic termini (Schweizer, Dresbach et al. 1998; Littleton, Barnard et al. 2001; Parnas, Rashkovan et al. 2006). NSF is also important for synaptic plasticity, which is believed to be the cellular mechanism for the formation of memory. NSF binds to the glutamate receptor subunit 2 (GluR2) of the postsynaptic α-amino-3-hydroxy-5-methyl-4-isoxazolepropionic acid receptor (AMPAR). This interaction promotes trafficking and stabilization of AMPAR at the postsynaptic membrane (Luscher, Xia et al. 1999; Lee, Liu et al. 2002; Hanley 2007). The NSF-GluR2 interaction is thought to be essential for the expression of LTD, a form of synaptic plasticity in the hippocampus (Lei and McBain 2004; Steinberg, Huganir et al. 2004). Pctk1 binds to NSF at the D2 domain, and phosphorylates NSF at serine 569 (Liu, Cheng et al. 2006). This inhibits oligomerization of NSF, which is required for normal NSF function. Expression of a kinase-dead mutant of Pctk1 or NSF-S569A in PC12 cells significantly increases high K(+)-stimulated growth hormone release, whereas expression of wild type Pctk1 had the opposite effect. These data suggest that Pctk1 may be involved in the formation of memory by modulating the activity of NSF at presynaptic or postsynaptic termini. Our result that Pctk1 inhibits induction of LTD is consistent with the Pctk1 interaction with NSF.
Pctk1 also interacts with the coat protein II (COPII) complex (Palmer, Konkel et al. 2005), which mediates the export of secretory cargo from the endoplasmic reticulum. Pctk1 interacts specifically with Sec23Ap, and inhibition of Pctk1 kinase activity produces a defect in cargo transport in the endoplasmic reticulum.
Pctk1 can be phosphorylated by protein kinase A (PKA). Phosphorylation of S153 inhibits Pctk kinase activity and kinase active forms of Pctk stimulate neurite outgrowth in Neuro-2A cells (Graeser, Gannon et al. 2002). Pctk1 can also be phosphorylated by cyclin dependent kinase 5 (Cdk5)/p35 complex. Like Cdk5, Pctk1 does not bind a cyclin; instead both Cdk5 and Pctk1 bind the same regulatory protein, p35. Whereas PKA phosphorylation of Pctk1 inhibits its kinase activity, phosphorylation of Pctk1 by Cdk5/p35 activates Pctk1 kinase activity (Graeser, Gannon et al. 2002). Pctk1 kinase activity in Cdk5 null mice is significantly reduced in brain and muscle.
The fact that Pctk1 rats showed a deficit in only SWM and not SRM lends support to the notion that SWM and SRM are encoded by different information processing mechanisms in hippocampus. However, it is not clear whether one of the interactions described above between Pctk1 and other neuronal complexes is responsible for the behavioral effect on learning and memory, or if some unknown interaction is essential. The present demonstration of impaired SWM but intact SRM in Pctk1 rats shows some parallels with the behavioral phenotypes of transgenic mice with functional loss of N-methyl-D-aspartate (NMDA) glutamate receptors in dentate gyrus or CA3 subregions (Niewoehner, Single et al. 2007).
Ccnd1
Ccnd1 rats displayed both SRM and SWM deficits in RAWM, but unlike Pctk1 injected animals, Ccnd1 rats did not show a significant deficit in the MWM. It is unclear whether the deficit in RAWM performance by Ccnd1 animals was due to an increase in memory load required in this paradigm or the difference in age of the animals during the two behavioral tests, 14 months old vs. 7 months old for RAWM and MW, respectively.
Ccnd1 is a key regulator of the transition from G1 to S phase in all cells (Weinberg 1995). It activates cdk4/6, which in turn phosphorylates Rb and releases E2f transcription factors. These then activate a variety of enzyme pathways essential for DNA synthesis. Despite its presence in adult dentate gyrus neurons, Ccnd1 does not appear essential for neurogenesis during development or in adults, because its ablation had little or no effect on the growth of neurons (Fantl, Stamp et al. 1995; Sicinski, Donaher et al. 1995; Kowalczyk, Filipkowski et al. 2004). However, overexpression of Ccnd1/cdk4 during development promoted the expansion of neural progenitors and inhibited neuron maturation leading to larger cortical and subventricular cell layers (Simpson, Moon et al. 2007; Lange, Huttner et al. 2009). Conversely, reduction of Cdnd1 had the opposite effect. Thus, the increase in Ccnd1 expression in mature adult brains might be expected to expand the pool of neuroprogenitors and delay their differentiation to mature neurons. It is worth noting that Ccnd1 expresses exclusively in neurons in adult brains and that it appears to be sequestered in cytoplasm in terminally differentiated neurons. In differentiated progenitor cells, nuclear localization of ectopic Ccnd1 induced apoptosis, and the DNA-damaging compound camptothecin caused nuclear accumulation of endogenous Ccnd1, accompanied by Rb phosphorylation (Sumrejkanchanakij, Tamamori-Adachi et al. 2003).
Ccnd1 may also influence learning and memory performance through its effect on several nuclear receptors/transcription factors (Ewen and Lamb 2004). Ccnd1 is a ligand-independent co-repressor of thyroid hormone receptor β1 (TRβ1) (Lin, Zhao et al. 2002; Petre-Draviam, Williams et al. 2005). Transgenic mice carrying a mutant TRβ1 that reduces its ligand binding activity (and, therefore, its activity) display hyperactivity and learning deficits, suggesting that the normal function of TRβ1 may be important for cognition (McDonald, Wong et al. 1998). Repression of TRβ1 by overexpression of Ccnd1 may lead to similar learning deficits.
Ccnd1 also modulates the activities of sex hormone receptors such as androgen receptors (ARs) and estrogen receptors (ERs). Ccnd1 selectively inhibits ligand-dependent AR functions in several cell types (Knudsen, Cavenee et al. 1999; Petre, Wetherill et al. 2002; Petre-Draviam, Williams et al. 2005). The roles of ARs in learning and memory have been controversial. Some studies have shown that adrogens and selective AR modulators enhanced learning and memory, whereas other studies showed that androgens impaired learning and memory (Naghdi, Nafisy et al. 2001; Acevedo, Tittle et al. 2008). Finally, Ccnd1 binds to and activates the estrogen receptor α (ERα) both in the presence and absence of estrogen (Neuman, Ladha et al. 1997; Zwijsen, Wientjens et al. 1997). Estrogen enhances the spatial memory performance of both male and female rats (Foy, Baudry et al. 2010; Mukai, Kimoto et al. 2010). This would suggest that the behavioral learning deficits induced by Ccnd1 expression in this study are independent of ERα.
To our knowledge, the results from the current study provide the first validation of the involvement of Pctk1 and Ccnd1 proteins in learning and memory processes. Further studies are needed to tease out the precise mechanisms by which these two proteins affect cognition. Additionally, 3 out of 3 genes chosen for this study all showed a defect in learning and memory. This validates the original studies of Burger et al (Burger, Lopez et al. 2007; Burger, Lopez et al. 2008) and suggests that other genes identified in those studies should probably be examined as well.
Supplementary Material
Acknowledgments
The authors were supported by NIH grants NS36302 to NM and AG014979, AG037984, AG036800 and the Evelyn F. McKnight Brain Research Foundation to TCF. NM was also supported by the Edward R. Koger American Cancer Society endowment fund. NM is an inventor of AAV patents related to recombinant AAV technology and owns equity in a gene therapy company that is commercializing AAV for gene therapy applications.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
REFERENCES
- Acevedo SF, Tittle S, et al. Transgenic expression of androgen receptors improves spatial memory retention in both sham-irradiated and 137Cs gamma-irradiated female mice. Radiat Res. 2008;170(5):572–578. doi: 10.1667/RR1435.1. [DOI] [PubMed] [Google Scholar]
- Allen Allen Brain Atlas (Internet) 2004 Retrieved 12/12/2010, 2010, from http://www.brain-map.org.
- Besset V, Rhee K, et al. The cellular distribution and kinase activity of the Cdk family member Pctaire1 in the adult mouse brain and testis suggest functions in differentiation. Cell Growth Differ. 1999;10(3):173–181. [PubMed] [Google Scholar]
- Bodhinathan K, Kumar A, et al. Intracellular redox state alters NMDA receptor response during aging through Ca2+/calmodulin-dependent protein kinase II. J Neurosci. 2010;30(5):1914–1924. doi: 10.1523/JNEUROSCI.5485-09.2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burger C, Lopez MC, et al. Genome-wide analysis of aging and learning-related genes in the hippocampal dentate gyrus. Neurobiol Learn Mem. 2008;89(4):379–396. doi: 10.1016/j.nlm.2007.11.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burger C, Lopez MC, et al. Changes in transcription within the CA1 field of the hippocampus are associated with age-related spatial learning impairments. Neurobiol Learn Mem. 2007;87(1):21–41. doi: 10.1016/j.nlm.2006.05.003. [DOI] [PubMed] [Google Scholar]
- Campbell-Thompson M, Dixon LR, et al. Pancreatic adenocarcinoma patients with localised chronic severe pancreatitis show an increased number of single beta cells, without alterations in fractional insulin area. Diabetologia. 2009;52(2):262–270. doi: 10.1007/s00125-008-1200-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ewen ME, Lamb J. The activities of cyclin D1 that drive tumorigenesis. Trends Mol Med. 2004;10(4):158–162. doi: 10.1016/j.molmed.2004.02.005. [DOI] [PubMed] [Google Scholar]
- Fantl V, Stamp G, et al. Mice lacking cyclin D1 are small and show defects in eye and mammary gland development. Genes Dev. 1995;9(19):2364–2372. doi: 10.1101/gad.9.19.2364. [DOI] [PubMed] [Google Scholar]
- Foy MR, Baudry M, et al. Regulation of hippocampal synaptic plasticity by estrogen and progesterone. Vitam Horm. 2010;82:219–239. doi: 10.1016/S0083-6729(10)82012-6. [DOI] [PubMed] [Google Scholar]
- Gorbatyuk OS, Li S, et al. alpha-Synuclein expression in rat substantia nigra suppresses phospholipase D2 toxicity and nigral neurodegeneration. Mol Ther. 2010;18(10):1758–1768. doi: 10.1038/mt.2010.137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gorbatyuk OS, Li S, et al. The phosphorylation state of Ser-129 in human alpha-synuclein determines neurodegeneration in a rat model of Parkinson disease. Proc Natl Acad Sci U S A. 2008;105(2):763–768. doi: 10.1073/pnas.0711053105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Graeser R, Gannon J, et al. Regulation of the CDK-related protein kinase PCTAIRE-1 and its possible role in neurite outgrowth in Neuro-2A cells. J Cell Sci. 2002;115(Pt 17):3479–3490. doi: 10.1242/jcs.115.17.3479. [DOI] [PubMed] [Google Scholar]
- Hanley JG. NSF binds calcium to regulate its interaction with AMPA receptor subunit GluR2. J Neurochem. 2007;101(6):1644–1650. doi: 10.1111/j.1471-4159.2007.04455.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hauswirth WW, Lewin AS, et al. Production and purification of recombinant adeno-associated virus. Methods Enzymol. 2000;316:743–761. doi: 10.1016/s0076-6879(00)16760-6. [DOI] [PubMed] [Google Scholar]
- Hernandez PJ, Abel T. The role of protein synthesis in memory consolidation: progress amid decades of debate. Neurobiol Learn Mem. 2008;89(3):293–311. doi: 10.1016/j.nlm.2007.09.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hu JS, Olson EN, et al. HEB, a helix-loop-helix protein related to E2A and ITF2 that can modulate the DNA-binding ability of myogenic regulatory factors. Mol Cell Biol. 1992;12(3):1031–1042. doi: 10.1128/mcb.12.3.1031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim HA, Pomeroy SL, et al. A developmentally regulated switch directs regenerative growth of Schwann cells through cyclin D1. Neuron. 2000;26(2):405–416. doi: 10.1016/s0896-6273(00)81173-3. [DOI] [PubMed] [Google Scholar]
- Kirik D, Rosenblad C, et al. Parkinson-like neurodegeneration induced by targeted overexpression of alpha-synuclein in the nigrostriatal system. J Neurosci. 2002;22(7):2780–2791. doi: 10.1523/JNEUROSCI.22-07-02780.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klugmann M, Wymond Symes C, et al. AAV-mediated hippocampal expression of short and long Homer 1 proteins differentially affect cognition and seizure activity in adult rats. Mol Cell Neurosci. 2005;28(2):347–360. doi: 10.1016/j.mcn.2004.10.002. [DOI] [PubMed] [Google Scholar]
- Knudsen KE, Cavenee WK, et al. D-type cyclins complex with the androgen receptor and inhibit its transcriptional transactivation ability. Cancer Res. 1999;59(10):2297–2301. [PubMed] [Google Scholar]
- Kowalczyk A, Filipkowski RK, et al. The critical role of cyclin D2 in adult neurogenesis. J Cell Biol. 2004;167(2):209–213. doi: 10.1083/jcb.200404181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kozar K, Sicinski P. Cell cycle progression without cyclin D-CDK4 and cyclin D-CDK6 complexes. Cell Cycle. 2005;4(3):388–391. doi: 10.4161/cc.4.3.1551. [DOI] [PubMed] [Google Scholar]
- Kumar A. Carbachol-induced long-term synaptic depression is enhanced during senescence at hippocampal CA3-CA1 synapses. J Neurophysiol. 2010;104(2):607–616. doi: 10.1152/jn.00278.2010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kumar A, Foster TC. Enhanced long-term potentiation during aging is masked by processes involving intracellular calcium stores. J Neurophysiol. 2004;91(6):2437–2444. doi: 10.1152/jn.01148.2003. [DOI] [PubMed] [Google Scholar]
- Kumar A, Foster TC. Shift in induction mechanisms underlies an age-dependent increase in DHPG-induced synaptic depression at CA3 CA1 synapses. J Neurophysiol. 2007;98(5):2729–2736. doi: 10.1152/jn.00514.2007. [DOI] [PubMed] [Google Scholar]
- Kumar A, Thinschmidt JS, et al. Aging effects on the limits and stability of long-term synaptic potentiation and depression in rat hippocampal area CA1. J Neurophysiol. 2007;98(2):594–601. doi: 10.1152/jn.00249.2007. [DOI] [PubMed] [Google Scholar]
- Lange C, Huttner WB, et al. Cdk4/cyclinD1 overexpression in neural stem cells shortens G1, delays neurogenesis, and promotes the generation and expansion of basal progenitors. Cell Stem Cell. 2009;5(3):320–331. doi: 10.1016/j.stem.2009.05.026. [DOI] [PubMed] [Google Scholar]
- Lee SH, Liu L, et al. Clathrin adaptor AP2 and NSF interact with overlapping sites of GluR2 and play distinct roles in AMPA receptor trafficking and hippocampal LTD. Neuron. 2002;36(4):661–674. doi: 10.1016/s0896-6273(02)01024-3. [DOI] [PubMed] [Google Scholar]
- Lei S, McBain CJ. Two Loci of expression for long-term depression at hippocampal mossy fiber-interneuron synapses. J Neurosci. 2004;24(9):2112–2121. doi: 10.1523/JNEUROSCI.4645-03.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin HM, Zhao L, et al. Cyclin D1 Is a Ligand-independent Co-repressor for Thyroid Hormone Receptors. J Biol Chem. 2002;277(32):28733–28741. doi: 10.1074/jbc.M203380200. [DOI] [PubMed] [Google Scholar]
- Littleton JT, Barnard RJ, et al. SNARE-complex disassembly by NSF follows synaptic-vesicle fusion. Proc Natl Acad Sci U S A. 2001;98(21):12233–12238. doi: 10.1073/pnas.221450198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y, Cheng K, et al. Pctaire1 phosphorylates N-ethylmaleimide-sensitive fusion protein: implications in the regulation of its hexamerization and exocytosis. J Biol Chem. 2006;281(15):9852–9858. doi: 10.1074/jbc.M513496200. [DOI] [PubMed] [Google Scholar]
- Luscher C, Xia H, et al. Role of AMPA receptor cycling in synaptic transmission and plasticity. Neuron. 1999;24(3):649–658. doi: 10.1016/s0896-6273(00)81119-8. [DOI] [PubMed] [Google Scholar]
- Manfredsson FP, Mandel RJ. Development of gene therapy for neurological disorders. Discov Med. 2010;9(46):204–211. [PubMed] [Google Scholar]
- McDonald MP, Wong R, et al. Hyperactivity and learning deficits in transgenic mice bearing a human mutant thyroid hormone beta1 receptor gene. Learn Mem. 1998;5(4–5):289–301. [PMC free article] [PubMed] [Google Scholar]
- Meyerson M, Enders GH, et al. A family of human cdc2-related protein kinases. Embo J. 1992;11(8):2909–2917. doi: 10.1002/j.1460-2075.1992.tb05360.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mukai H, Kimoto T, et al. Modulation of synaptic plasticity by brain estrogen in the hippocampus. Biochim Biophys Acta. 2010;1800(10):1030–1044. doi: 10.1016/j.bbagen.2009.11.002. [DOI] [PubMed] [Google Scholar]
- Naghdi N, Nafisy N, et al. The effects of intrahippocampal testosterone and flutamide on spatial localization in the Morris water maze. Brain Res. 2001;897(1–2):44–51. doi: 10.1016/s0006-8993(00)03261-3. [DOI] [PubMed] [Google Scholar]
- Neuman E, Ladha MH, et al. Cyclin D1 stimulation of estrogen receptor transcriptional activity independent of cdk4. Mol Cell Biol. 1997;17(9):5338–5347. doi: 10.1128/mcb.17.9.5338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Niewoehner B, Single FN, et al. Impaired spatial working memory but spared spatial reference memory following functional loss of NMDA receptors in the dentate gyrus. Eur J Neurosci. 2007;25(3):837–846. doi: 10.1111/j.1460-9568.2007.05312.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Neil J, Shank J, et al. TAL1/SCL induces leukemia by inhibiting the transcriptional activity of E47/HEB. Cancer Cell. 2004;5(6):587–596. doi: 10.1016/j.ccr.2004.05.023. [DOI] [PubMed] [Google Scholar]
- Okuda T, Cleveland JL, et al. PCTAIRE-1 and PCTAIRE-3, two members of a novel cdc2/CDC28-related protein kinase gene family. Oncogene. 1992;7(11):2249–2258. [PubMed] [Google Scholar]
- Palmer KJ, Konkel JE, et al. PCTAIRE protein kinases interact directly with the COPII complex and modulate secretory cargo transport. J Cell Sci. 2005;118(Pt 17):3839–3847. doi: 10.1242/jcs.02496. [DOI] [PubMed] [Google Scholar]
- Parker MH, Perry RL, et al. MyoD synergizes with the E-protein HEB beta to induce myogenic differentiation. Mol Cell Biol. 2006;26(15):5771–5783. doi: 10.1128/MCB.02404-05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parnas I, Rashkovan G, et al. Role of NSF in neurotransmitter release: a peptide microinjection study at the crayfish neuromuscular junction. J Neurophysiol. 2006;96(3):1053–1060. doi: 10.1152/jn.01313.2005. [DOI] [PubMed] [Google Scholar]
- Paxinos G, Watson C, et al. Bregma, lambda and the interaural midpoint in stereotaxic surgery with rats of different sex, strain and weight. J Neurosci Methods. 1985;13(2):139–143. doi: 10.1016/0165-0270(85)90026-3. [DOI] [PubMed] [Google Scholar]
- Peel AL, Klein RL. Adeno-associated virus vectors: activity and applications in the CNS. J Neurosci Methods. 2000;98(2):95–104. doi: 10.1016/s0165-0270(00)00183-7. [DOI] [PubMed] [Google Scholar]
- Petre-Draviam CE, Williams EB, et al. A central domain of cyclin D1 mediates nuclear receptor corepressor activity. Oncogene. 2005;24(3):431–444. doi: 10.1038/sj.onc.1208200. [DOI] [PubMed] [Google Scholar]
- Petre CE, Wetherill YB, et al. Cyclin D1: mechanism and consequence of androgen receptor co-repressor activity. J Biol Chem. 2002;277(3):2207–2215. doi: 10.1074/jbc.M106399200. [DOI] [PubMed] [Google Scholar]
- Quong MW, Romanow WJ, et al. E protein function in lymphocyte development. Annu Rev Immunol. 2002;20:301–322. doi: 10.1146/annurev.immunol.20.092501.162048. [DOI] [PubMed] [Google Scholar]
- Reimsnider S, Manfredsson FP, et al. Time course of transgene expression after intrastriatal pseudotyped rAAV2/1, rAAV2/2, rAAV2/5, and rAAV2/8 transduction in the rat. Mol Ther. 2007;15(8):1504–1511. doi: 10.1038/sj.mt.6300227. [DOI] [PubMed] [Google Scholar]
- Rex CS, Gavin CF, et al. Myosin IIb regulates actin dynamics during synaptic plasticity and memory formation. Neuron. 2010;67(4):603–617. doi: 10.1016/j.neuron.2010.07.016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roullet P, Lassalle JM, et al. A study of behavioral and sensorial bases of radial maze learning in mice. Behav Neural Biol. 1993;59(3):173–179. doi: 10.1016/0163-1047(93)90926-9. [DOI] [PubMed] [Google Scholar]
- Schweizer FE, Dresbach T, et al. Regulation of neurotransmitter release kinetics by NSF. Science. 1998;279(5354):1203–1206. doi: 10.1126/science.279.5354.1203. [DOI] [PubMed] [Google Scholar]
- Sherr CJ, Roberts JM. CDK inhibitors: positive and negative regulators of G1-phase progression. Genes Dev. 1999;13(12):1501–1512. doi: 10.1101/gad.13.12.1501. [DOI] [PubMed] [Google Scholar]
- Sicinski P, Donaher JL, et al. Cyclin D1 provides a link between development and oncogenesis in the retina and breast. Cell. 1995;82(4):621–630. doi: 10.1016/0092-8674(95)90034-9. [DOI] [PubMed] [Google Scholar]
- Simpson PJ, Moon C, et al. Progressive and inhibitory cell cycle proteins act simultaneously to regulate neurotrophin-mediated proliferation and maturation of neuronal precursors. Cell Cycle. 2007;6(9):1077–1089. doi: 10.4161/cc.6.9.4132. [DOI] [PubMed] [Google Scholar]
- Steinberg JP, Huganir RL, et al. N-ethylmaleimide-sensitive factor is required for the synaptic incorporation and removal of AMPA receptors during cerebellar long-term depression. Proc Natl Acad Sci U S A. 2004;101(52):18212–18216. doi: 10.1073/pnas.0408278102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sumrejkanchanakij P, Tamamori-Adachi M, et al. Role of cyclin D1 cytoplasmic sequestration in the survival of postmitotic neurons. Oncogene. 2003;22(54):8723–8730. doi: 10.1038/sj.onc.1206870. [DOI] [PubMed] [Google Scholar]
- Tran TD, Kelly SJ. Critical periods for ethanol-induced cell loss in the hippocampal formation. Neurotoxicol Teratol. 2003;25(5):519–528. doi: 10.1016/s0892-0362(03)00074-6. [DOI] [PubMed] [Google Scholar]
- Uittenbogaard M, Chiaramello A. Expression of the bHLH transcription factor Tcf12 (ME1) gene is linked to the expansion of precursor cell populations during neurogenesis. Brain Res Gene Expr Patterns. 2002;1(2):115–121. doi: 10.1016/s1567-133x(01)00022-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weinberg RA. The retinoblastoma protein and cell cycle control. Cell. 1995;81(3):323–330. doi: 10.1016/0092-8674(95)90385-2. [DOI] [PubMed] [Google Scholar]
- Zhang Y, Babin J, et al. HTF4: a new human helix-loop-helix protein. Nucleic Acids Res. 1991;19(16):4555. doi: 10.1093/nar/19.16.4555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhuang Y, Cheng P, et al. B-lymphocyte development is regulated by the combined dosage of three basic helix-loop-helix genes, E2A, E2-2, and HEB. Mol Cell Biol. 1996;16(6):2898–2905. doi: 10.1128/mcb.16.6.2898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zolotukhin S, Potter M, et al. Production and purification of serotype 1, 2, and 5 recombinant adeno-associated viral vectors. Methods. 2002;28(2):158–167. doi: 10.1016/s1046-2023(02)00220-7. [DOI] [PubMed] [Google Scholar]
- Zwijsen RM, Wientjens E, et al. CDK-independent activation of estrogen receptor by cyclin D1. Cell. 1997;88(3):405–415. doi: 10.1016/s0092-8674(00)81879-6. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.