Abstract
Metabolomics or biochemical profiling is a fast emerging science; however, there are still many associated bottlenecks to overcome before measurements will be considered robust. Advances in MS resolution and sensitivity, ultra pressure LC MS, ESI, and isotopic approaches such as flux analysis and stable-isotope dilution, have made it easier to quantitate biochemicals. The digitization of mass spectrometers has simplified informatic aspects. However, issues of analytical variability, ion suppression and metabolite identification still plague metabolomics investigators. These hurdles need to be overcome for accurate metabolite quantitation not only for in vitro systems, but for complex matrices such as biofluids and tissues, before it is possible to routinely identify biomarkers that are associated with the early prediction and diagnosis of diseases. In this report, we describe a novel isotopic-labeling method that uses the creation of distinct biochemical signatures to eliminate current bottlenecks and enable accurate metabolic profiling.
Metabolite profiling & stable isotopes
Standardization of current practices in MS-based metabolic profiling is required to ensure laboratory-to-laboratory comparative analysis for the development of precise discriminatory diagnostic tests and therapeutic treatments. In order to accurately compare data and ratios of individual metabolite concentrations, several analytical challenges must be overcome. The ability to measure inherent biological variation is directly compromised by the introduction of variability through sample handling, extraction, ionization efficiency (including suppression), or variation of analytical parameters (including instrumentation conditions). Standardization techniques to remove introduced variability in the determination of biological information have historically been addressed either through the use of multiple internal standard compounds to normalized data [1] or through stable-isotope-assisted methods [2,3].
When employing multiple internal standards as a means to standardize, the choice of which standard to apply to a particular metabolite is difficult. The retention time and mass-to-charge ratio of a metabolite compound is dependent on its chemical properties, and the use of a single standard or standard normalization factor for multiple chemical species may not be practical or appropriate as the best standard is almost always an isotopomer of the compound itself. This can be prohibitively expensive when the number of compounds to be measured is significant. For this reason there are many techniques for the creation of assemblages of isotopic standards used in NMR and LC–MS data analysis [1,4–8].
Stable-isotope assisted methods employ either an in vivo or in vitro (chemical) incorporation of an isotopically enriched element (e.g., 13C or 15N), an amino acid or salt into a biological sample. The in vivo methods are the oldest and most frequent usage of stable isotopes in studies probing metabolism, to understand flux [9,10] and also for biomarker discovery [11,12]. In these studies, a stable isotope is incorporated into a specific molecule at a specific location in order to achieve a precision in understanding the metabolic fate of that molecule or its substituent. Investigators employ wholly labeled molecules where >99% of an atom is replaced with an isotopic equivalent, and universally labeled isotopes where the isotope is universally distributed within the target molecule at less than saturation levels. The isotopically labeled molecule, or ‘precursor’, is fed to an in vivo organism, in vitro cell system or in vitro cell-free system for either a brief or extended period of time, after which the fate of the isotope is determined either by the use of NMR, MS, chemical degradation or other detection technique. There are many studies in which more than one isotope is incorporated into a target molecule, and all of the isotopic fragments are examined for their differential fates. In all cases, these methods are targeted analyses; that is, they seek the incorporation of a specific labeled atom into other specific molecules [2]. In vivo tracer labeling or metabolic-flux analysis typically measures metabolic fluxes or the rate of turnover of molecules through a metabolic pathway [13,14]. Other commonly used in vivo stable-isotopic-labeling approaches include stable-isotope labeling with amino acids [15], and stable-isotope labeling by amino acids in cell culture (SILAC) [16]. In both of these, the experimental and control samples are differentially labeled so that the pooled sample may be analyzed as a single sample in order to get comparative information on the experimental relative to the control. These approaches are primarily used in proteomic studies. The benefits of these techniques are that they require fewer analyses because both the control and experimental samples are analyzed at once, and data quality is enhanced as there can be no sample-to-sample variance between the commonly prepped and analyzed experimental and control. One of the biggest problems in the interpretation of mass spectral data is the variability of ionization efficiency, commonly known as ‘ion suppression’. This is one of the major sources of sample-to-sample variance. Where an isotopically labeled standard is chemically identical to the analyte, both the standard and the analyte are measured in an identical environment and, therefore, share identical ionization efficiencies making the measurements more accurate. This reduction of the impact of suppression is common to many of the isotopic techniques discussed below.
Chemical or post vivo labeling techniques include isotope-coded affinity tag [17], isotope-coded protein labeling [18], or the utilization of isobaric mass tags for labeling, such as isobaric tags for relative and absolute quantitation [19] and tandem mass tag [20]. These techniques also run as a single, pooled sample composed of multiple differentially tagged samples, and also demonstrate improved data quality due to the reduction of sample-to-sample variance.
Stable-isotopic internal standards
Another use of stable-isotopically labeled compounds is as internal standards for their nonlabeled counterparts, referred to as stable-isotope dilution methodology [21]. In such an experiment, an isotopically enriched molecule is added to a sample or extract at a known concentration prior to an analysis, and the final measurement determines the exact concentration of the nonlabeled material by comparison. In this type of study, it is not uncommon for a researcher to add more than one isotopically-distinct standard if more than one molecule is to be quantified. In this situation, the same standard is introduced into all samples, but there is no information carried by the standard other than for purposes of relative quantitation; that is, the standard has no relation to the experiment at hand.
More recently, there have been a number of variants of the above whereby an organism is grown on an isotopically defined feedstock, such that the entire organism is heavily, if not entirely, composed of molecules consisting of only one or two isotopes [22,23]. Historically, such standards are carefully constructed to differ from any other analyte by a specific mass difference. For example, by labeling one cell population with a cell-culture component or components containing isotopically light atoms, for example 12C, and another cell population with isotopically heavy atoms, for example 13C. Once a stressor is applied to the experimental group – for example, drug or chemical treatment – equal amounts of both populations are combined and analyzed by LC–MS. Again, the composite sample is mass-spectroscopically analyzed for analyte peaks, with a reduction of sample-to-sample variance. The ratio of the first to second isotope (isotopic ratio) for each compound is determined. While the practice of isotopic-labeling standardization to introduce a specific stable-isotope-labeled compound for each metabolite has provided a means to effectively reduce variability introduced during analysis and sample processing, the fact that the peaks carry no inherent means of identification, artifactual confounding peaks and metabolite misidentification remain unsolved sources of error.
Metabolomics & stable-isotope labeling
The diverse chemical nature of the metabolome makes the comprehensive chemical tagging methods unfeasible. Until recently, other than metabolic-flux analysis and simple targeted analyses, very few additional methods that employ stable isotopes were used for the full quantitation of biochemical metabolites. In 2004, Mashego et al. reported the use of stable-isotope dilution for the metabolome characterization of Saccharomyces cerevisiae, termed ‘Mass Isotopomer Ratio Analysis of U-13C-Labeled Extracts’ (MIRACLE), in which metabolites were quantitated relative to their U-13C-labeled equivalents [24]. Using the same isotope ratio-based quantitation approach, both Birkemeyer et al. [23] and Bennett et al. [25,26] describe the quantitation of microbial metabolites using uniformly labeled U-13C carbon sources. Others have also utilized this approach for specific classes of metabolites, including nitrogen-containing metabolites [27] and microbial folates [28]. Other unique metabolite labeling approaches have been described by Nagai et al., namely, an isotopic tracing method for the metabolic assessment of lipogenesis, lipolysis and glucose uptake in adipocytes using LC–MS through the absorption of 13C-labeled compounds [29], while Huang and Regnier differentially derivatized control and experimental samples with isotopically light and heavy (D6) forms of the silylation reagent N-methyl-N-(tert-butyldimethylsilyl)trifluoroacetamide, which were then mixed and analyzed using 2D GC–MS (GC × GC–MS) [30]. These methods are useful for the simultaneous analysis of multiple compounds, as metabolites are quantitated relative to their U-13C-labeled equivalents. They are targeted methods, either to specific classes of compounds or individual metabolites and their pathways; however, if a new pathway is triggered and metabolites synthesized in the unlabeled samples, then these compounds will not be identified by the mass ratio-based analysis of the type described here, since the U-13C equivalents will be absent.
Isotopic Ratio Outlier Analysis™ as a protocol for metabolic profiling: quantitation & identification
Metabolic profiling is rapidly gaining adoption as a methodology to measure metabolic phenotypes associated with disease and drug therapies, and MS is excellently suited for the analysis of labeling patterns of individual atoms (i.e., isotopomers) in a wide range of metabolites. The Isotopic Ratio Outlier Analysis™ (IROA) method is not a targeted analysis [101], it utilizes full metabolic labeling instead, such as SILAC in cell culture, but uses specifically labeled materials with random, universal enrichments centered on either 5 or 95% 13C. The reduction in purity of the label from either natural abundance, or the high purity 13C (generally >98%) creates an isotopic signature of such molecules that is dramatic and imparts many analytical advantages compared with standard isotopic labeling, as outlined in Table 1. With the creation of distinct chemical signatures in every biological molecule, it is possible to develop software algorithms to sort these biological signals. Using comprehensive (>98%) labeling, the base peak can usually be detected, but the M+1 peak that provides important diagnostic information is generally less useful for data interpretation because of its small size (e.g., the M+1 peak is 6% of the height of the base peak for a six-carbon molecule). If the percentage of 13C is increased to 5%, then the M+1 for a six-carbon molecule is significantly larger, namely 32%. With the IROA 13C-labeling approach, the number of carbons in a biological molecule can be determined by the distance between the two base peaks, 12C and 13C, and the relative height of the M+1 and M-1 provide confirmation of this fact, providing triply redundant information. Isotopically labeled compounds for the IROA are made for NextGen Metabolomics, Inc. (MI, USA) under patent protection and through special contract by Cambridge Isotope Laboratories (MA, USA).
Table 1.
Features of standard isotopic labeling using U-13C versus Isotopic Ratio Outlier Analysis™.
| Feature | U-13C (99 atom %) | IROA |
|---|---|---|
| Sample throughput | High to medium | High to medium |
| Removal of sample-to-sample variance | Yes – control and experimental samples are prepped and analyzed together, removing variability | Yes – control and experimental samples are prepped and analyzed together, removing variability |
| Removal of ion suppression | Yes – ionization is similar for isotopomers | Yes – ionization is similar for isotopomers |
| Reduction of the number of samples to be analyzed | Yes – control and experimental samples pooled and run as a single sample, reducing the number of samples to be analyzed by half | Yes – control and experimental samples pooled and run as a single sample, reducing the number of samples to be analyzed by half |
| Distinguishes artifacts and noise | No | Yes – allows classification of all peaks as compounds or artifacts |
| Metabolite coverage | Targeted | Global |
| Metabolite Identification | No | Number of carbon atoms and residual mass defined |
IROA: Isotopic Ratio Outlier Analysis™.
When considering the IROA experiment, the use of an isotope that exhibits minimal biological isotope effect is of importance. For instance, the use of deuterium isotopes with a mass twice that of hydrogen is known to cause a reduction in the kinetics of some enzyme mechanisms, but not in others [31]. On the other hand, the carbon 13C isotope effect is minimal, being an increase of only 8%, not 100% as in the case of deuterium. In the majority of biological systems, except in Crassulacean acid metabolism plants [32], no significant isotopic effects have been shown for 13C in most commonly used model systems. In an IROA experiment, described below, in order to accommodate for any minor 13C isotope effects, the 13C-labeled samples are always used as the control and directly compared with their 12C-labeled equivalents to identify any 13C effects. Furthermore, during the analysis of the IROA samples, it is important to select injection volumes to minimize detector saturation while allowing optimal compound identification.
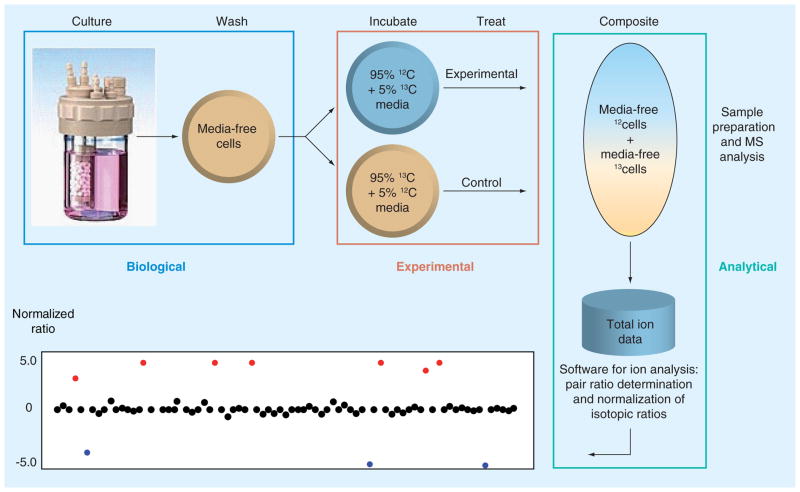
In the IROA experiment (Figure 1), as in other in vivo experiments, an isotopically-defined media is biotransformed by experimental systems to induce identifiable isotopic-labeling patterns (signatures) in all of their biological compounds. In brief, a homogenous cell population is divided into two populations: a ‘control’ population and an ‘experimental’ population, which will be subsequently treated with vehicle or a stressor, respectively. All of the biological compounds in the control sample are labeled with ‘13C media’ signatures and all of the biological compounds in the experimental sample are labeled with ‘12C media’ signatures. The key element is that both the 12C and 13C media are chemically identical but isotopically different. Unlike the previously discussed techniques, all of the carbon sources in the 12C media are randomly and universally composed of approximately 95% 12C with 5% 13C. Conversely, all of the carbon sources in the 13C media are composed of 95% 13C with 5% 12C, again, and importantly, the isotopes are universally and randomly incorporated into all carbon positions.
Figure 1. The Isotopic Ratio Outlier Analysis™ method for determining the biological response to drugs, toxins or other stressors.
Isotopic Ratio Outlier Analysis isotope ratios of 12C/13C 95/5% and 5/95% allow for control and experimental samples to be run simultaneously, eliminating sample-to-sample variability.
If both the control and the experimental cell samples are grown in the IROA isotopically-defined media for a sufficient time to replace their original natural-abundance carbon, then all of their contents will demonstrate distinctive isotopic patterns (orange box, Figure 1). Once labeled, the experimental sample is treated with a stressing regimen and the control sample is treated with only the vehicle for the same period of time. The stressing agent may be chemical, genetic, environmental, or any element or combination of elements that induce physiological alteration. At the conclusion of the experiment, the experimental sample is mixed with a control sample and the pooled sample prepped and analyzed in a single analysis. This has the effect of, as noted above, reducing sample-to-sample variance and increasing data quality.
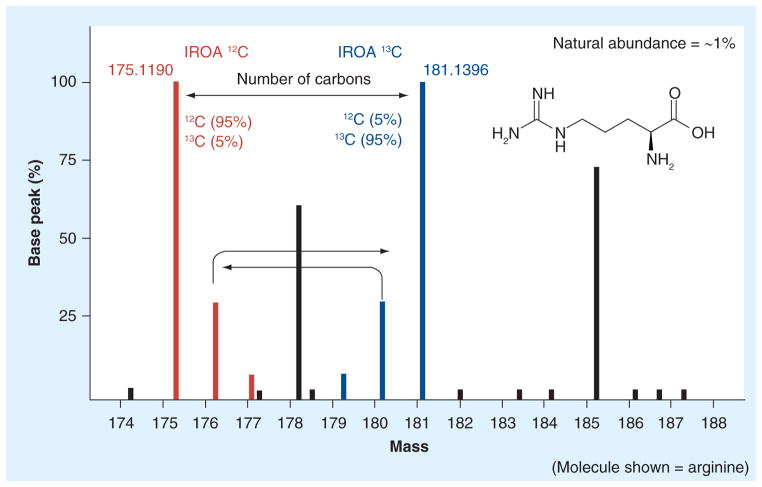
In natural abundance, biochemicals or metabolites, the M+1 secondary peak is quite small. In the IROA experiment, all biologically derived molecules may be easily distinguished from artifacts (which are present at only natural abundance), and each peak carries a ready identifier of its origin, a diagnostically enhanced M+1 for the 12C-derived sample and an identical diagnostically enhanced M-1 for the 13C-derived sample. The compounds associated with the IROA 13C control and IROA 12C experimental samples are both differentiatable (see the IROA peaks, Figure 2) and carry a tremendous amount of additional information; namely, the height of the M-1 relative to the 13C-base peak is a clear step function indicating the number of carbons in the molecule; unlike the 12C-base peaks, only the noncarbon components contribute to the M+1 and M+2 of the 13C-base peak, and may be examined without interference from carbon; and the exact mass of the 13C-base peak may be used to calculate the probable formula of the molecule. When armed with the number of carbons in the molecule, the ability to closely examine the remaining elements is generally quite discriminatory, and not only affords correct molecular formulae with a great deal of assurance, but allows the prediction of the characteristics of the 12C-based isotopomer of the same compound. Once labeled with the 12C and 13C media, all of the biological components in the control sample and experiment sample will carry unique signatures, and this means that the control sample can be directly embedded into and prepared simultaneously with an experimental sample and analyzed as a composite (green box, Figure 1). As noted previously, this not only reduces the number of samples to be analyzed by half, but also removes the sample-to-sample variance (or noise) that will normally be present in most metabolomics experiments, with not only no loss of information, but actually improved information content. IROA also fully removes the problem of ion suppression because a compound’s ability to ionize is a function of its structure and is not significantly altered by its isotopic distribution, similar to the technique noted above.
Figure 2. The Isotopic Ratio Outlier Analysis™ peaks.
In the case of arginine, the 12C M+ located at 175.1190 and its 13C mate at 181.1396 clearly indicate a six-carbon molecule. The corresponding M+1 and M-1 peaks are a mass difference of 1.00335 amu (the mass difference between a 13C and 12C isotope). Natural abundance peaks (in black) from exogenous sources do not have a 13C counterpart and are not considered in the analysis. IROA 12C peaks (red) and IROA 13C peaks (blue) are from biologically derived compounds.
IROA: Isotopic Ratio Outlier Analysis™.
Another advantage of the IROA patterning is that the M+1 and M-1 peak height, relative to the base peak of each molecule, generally indicates the number of carbons in the molecule. Since the height of the peaks of an isotopically defined compound may be effectively calculated (this is technically a polynomial expansion in which the dominance of carbon makes the remaining terms less important) by the binomial expansion of the expression: (12C% + 13C%)N where N equals the number of carbons and 12C% and 13C% equals the relative isotopic abundances [33]. This is not sufficiently accurate when all atoms are present at their natural abundance, but at 5% the M+1 is diagnostic. For the 12C (5% 13C) peaks, if there are: seven carbons the M+1 will be approximately 36.8%; six carbons the M+1 will be approximately 31.6%; five carbons the M+1 will be approximately 26.3%; four carbons the M+1 will be approximately 21.1%; three carbons the M+1 will be approximately 15.8% and so on. This calculation indicates exactly where the 13C-paired mate will be located. Together with accurate mass, the identification of metabolite entities in the sample is made possible.
The distinctive patterns are important in the interpretation of the resulting composite spectra and because it is possible to discriminate 12C-derived molecules, 13C-derived molecules, artifacts and derivatives of exogenously applied compounds, and the origin of every peak in the composite (see the IROA peaks, Figure 2).
The IROA peaks are all mathematically calculable and each set (12C and 13C) of carbon isotopomers will reliably and accurately account for the other set providing a redundant QC check point. As such, it is possible to interpret the analytical results of the composite sample to an even greater extent through the creation of IROA software algorithms that achieve data reduction of complex raw data, to concise, high-value information, as follows:
Characterize all peaks according to source (artifact, experimental [12C], control [13C], or standard);
Remove all artifacts;
Align and pair all remaining peaks across all scans;
Normalize and identify all pairs;
Determine the relative 12C/13C ratios of analytes in each sample;
Determine the statistical variance of the sample ratios.
Any experimental analyte compound that has a ratio that is a significant deviation (two or more standard deviations) from the average ratio will indicate a point where the biochemistry was altered. For instance, if the average ratio for all of the analytes is 1 (1:1 12C/13C ratio), but some analytes have ratios of 10 (10:1) or 0.1 (1:10) then the analytes that are outliers to the general population are those most strongly affected by the stressor.
A software package named ClusterFinder™ has been written in Java (JRE 1.7). To achieve step 1, ClusterFinder does a scan-by-scan analysis of the complete dataset and identifies all IROA peaks based on their extended isotopic envelopes. The removal of all non-IROA peaks follows logically upon the conclusion of step 1, and results in a significantly simplified dataset that may then be aligned based on mass across adjacent scans. The mirrored symmetry of the 12C and 13C halves of each IROA cluster allow the isotopic peaks to be correctly associated with their appropriate 12C- or 13C-base peak. As the isotopic dilution distributes molecules over a number of masses, the sum of the base peaks and their associated isotopic peaks must be used to calculate their respective areas accurately. Once the summed areas are available their ratio is simply calculated. On the whole, because the isotopic patterns carry so much information, the tasks the software performs are well defined but not overly difficult. The software has two modes of use; where both the 12C and 13C media are utilized, the software can find all IROA peaks regardless of their source. This represents a fully unbiased analysis as compounds of biological origin present or absent in either samples (control or experimental) will be identified by the software and artifacts will be simultaneously identified and removed from the dataset. The software will also find all IROA-labeled compounds where the IROA peak is present in one sample but not the other. This allows for targeted analysis whereby the control may contain 13C IROA peaks that may be used to identify their associated natural abundance peaks. This mode of analysis has been called the ‘phenotypic mode’ (see section titled ‘The use of IROA phenotyping application for the metabolic profiling of tissues or biofluids’ later in this article).
Normalization of IROA datasets
Various normalization techniques to account for matrix and analytical variances in transcriptomics, proteomics and metabolomics have been explored extensively. Methods that account for dilution or cell number have included total ion count (TIC), a standard ‘housekeeping’ compound or other physical characteristics of the sample (e.g., DNA, protein and osmality).
In metabolomics studies there have not yet been any molecules identified as good candidates for ‘housekeeping’ compounds. Creatinine has been proposed as such a ‘housekeeping’ compound in urine, but it has been discounted as it is, in itself, quite variable [34]. The physical measurements of the sample are often destructive and performed on like aliquots and not the samples to be analyzed, which lowers its precision. In a recent publication, Warrack et al. demonstrated the benefits of normalization with respect to the TIC of a spectrum, utilizing only the total intensity of components that are common to all samples after baseline correction – the ‘MS total useful signal’ (MSTUS) approach [35]. They noted the superiority of this approach in avoiding the influence of matrix- and solvent-related intensity measurements. This approach is similar to the common practice used in 1H NMR-based metabonomics analyses wherein each spectrum is normalized to the total integrated proton signal, after removal of regions corresponding to xenobiotics, internal standards and artifacts [36].
The IROA technology is designed to identify and remove noise and artifacts, which has the advantage in that an artificial cut-off for noise removal or baseline correction is not required, so that any low abundance metabolites are not artificially removed and can be considered the data-set. As such, IROA metabolomics utilizing TIC normalization follows the MSTUS approach; namely, normalizing only the components that are common to all signals. Since artifacts and non-biological compounds may be identified as they carry no IROA signatures, their removal further strengthens the MSTUS normalization approach.
Application of the IROA approach in S. cerevisiae
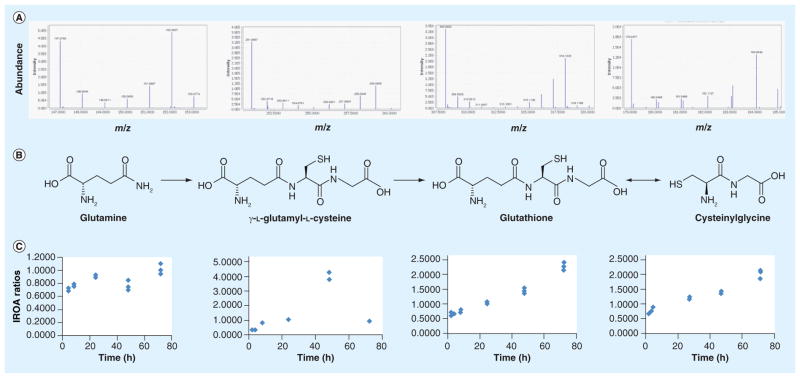
To demonstrate the application of IROA in the biological system of S. cerevisiae S288C, the bio-synthetic pathways surrounding glutathione in an aerobic culture were measured over a 72-h time course (Figure 3). The fermentation process of S. cerevisiae was measured using isotopically defined media (whereby all carbon sources were labeled with either 5% 13C/95% 12C or 95% 13C/5% 12C) and analyzed using LC–MS. The dataset produced was analyzed by the IROA ClusterFinder software (not shown). Figure 3 shows the representative IROA peaks for the compounds glutamine, γ-l-glutamyl-l-cysteine and glutathione and the normalized concentrations of these biochemical metabolites measured in culture samples collected at 4, 8, 24, 48 and 72 h. The glutathione pathway metabolites, some of the most important biological metabolites controlling many aspects of physiological response, are shown to be increasing over the 72-h time course, which is most likely a result of glucose levels being depleted over the 72-h time period (not shown) [37].
Figure 3. The biosynthetic pathways surrounding glutathione in Saccharomyces cerevisiae during a 72-h time course.
(A) IROA peaks, (B) Biological compounds and (C) Isotopic Ratio Outlier Analysis ratios.
IROA: Isotopic Ratio Outlier Analysis™.
The use of IROA phenotyping application for the metabolic profiling of tissues or biofluids
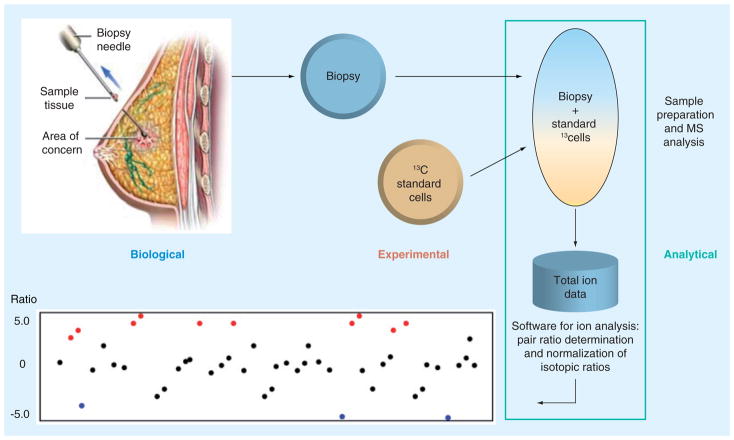
The clinical usefulness of metabolic profiling is just beginning to take shape [38,39]. In the basic IROA methodology described above (Figure 1), both the control sample and experimental samples are fully isotopically labeled so both may be easily found with absolute assurance of their identity and any artifacts may be removed. This leads to very clean, high-resolution datasets that clearly define the biological response of a biological system. Where it is not possible to isotopically label the biological sample, the IROA phenotyping application is applied and the sample is collected at natural abundance and mixed with a fully predefined ‘standard’ that has been isotopically labeled using IROA 13C media (Figure 4). An ideal standard would be one that represented the entire metabolome of the fluid or tissue under study. While this could be generated in a number of ways, including the use of IROA grown cells lines (the IROA standard) to achieve accurate quantification, this aspect of the application will not be considered here.
Figure 4. The Isotopic Ratio Outlier Analysis™ phenotyping application.
The material to be phenotyped is mixed with 13C (Isotopic Ratio Outlier Analysis) cells and/or standard compounds that allow one to find and pair all peaks. The deviation from the standard is diagnostic of the sample’s biochemical phenotype.
Using an IROA-labeled standard, all of the peaks may be easily identified according to the presence of their characteristic M-1 peak. The natural abundance peaks corresponding to each standard peak may be readily identified because, although they do not carry any isotopic labeling, their exact mass and position are established relative to the standard. If the pooled standard is well characterized and the compounds that are present in it have already been identified, then it can be used as a point of comparison for all of its contained compounds. Artifacts will have no match in the standard and, thus, need not be considered in a final dataset. Whereas, in a basic IROA dataset the ratio of the peak areas represents the relative deviation of the metabolic pool sizes brought about by the experimental condition, in a phenotyping experiment the overall pattern of deviations from the standard will define phenotype by difference from the standard. It is possible to consider the phenotyping experiment a complex targeted analysis relative to the unbiased analysis of the full IROA experiment.
Conclusion & future perspective
Isotopic techniques have proven valuable in improving the quality of data for many experimental systems. In general, where a standard can be embedded into an experimental sample, the measurements of the experimental analytes are greatly improved. In metabolomics, there have been many uses of isotopic standards for understanding flux and biomarker discovery. The use of high purity isotopic labels has been a common technique in many other omics sciences and has just recently been used in metabolomics [24–30,40]. The IROA technique has advanced this by using specific isotopic balances to create definable patterns in metabolites. This makes the mass-spectral signals complex but increases the information content that can be extracted from these signals. It is clear that metabolomics has just begun to explore the potential of using isotopic labels to further its effectiveness.
Executive summary.
Metabolite profiling & stable isotopes
Standardization techniques through the use of multiple internal standard compounds and stable-isotope-assisted methods have been historically employed to overcome analytical challenges in metabolite profiling.
These analytical challenges include ion suppression, sample-to-sample variability, noise and artifactual confounding peaks and metabolite identification.
Several approaches go a long way in rectifying most of these issues, but they all employ targeted methods and fall short of solving all analytical problems.
The use of an internal standard for each metabolite measured is optimal; however, for broad metabolic profiling when using multiple internal standards to normalize, the choice of which standard to apply to a particular metabolite can be problematic.
In vivo stable-isotope-assisted methods include flux analysis, stable-isotope labeling with amino acids and stable-isotope labeling by amino acids in cell culture. Stable-isotope labeling with amino acids, stable-isotope labeling by amino acids in cell culture and various chemical labeling techniques improve data quality by removing ion suppression and sample-to-sample variability, and also provide the benefit of requiring fewer analyses (as the control and experimental samples are analyzed simultaneously).
Stable-isotopic internal standards
Rather than the use of natural abundance isotopic standards, stable isotopes can also be applied as internal standards through the use of stable-isotope dilution methodology.
While this method also utilizes a composite sample for analysis resulting in the removal of ion suppression and sample-to-sample variability, the problem of confounding artifactual peaks and metabolite misidentification remains.
Metabolomics & stable-isotope labeling
Stable-isotope dilution labeling was initially applied to proteomics and was later introduced to metabolomics through the use of ‘Mass Isotopomer Ratio Analysis of U-13C-Labeled Extracts’, an isotope ratio-based quantitation approach.
While many researchers have applied this protocol to quantitate specific classes of metabolites, the protocol does not eliminate issues of metabolite identification and confounding artifactual peaks.
Isotopic Ratio Outlier Analysis™ as a protocol for metabolic profiling, quantitation & identification
Similar to other stable isotope approaches, the same cell pool can be used to create both control and experimental samples using Isotopic Ratio Outlier Analysis™ (IROA) technology, so that variability is removed, fewer samples are required and ion suppression is reduced.
Artifacts and noise are removed using the IROA protocol. Since all material from either biological system (experimental or control) has a unique isotopic signature, all unsigned molecules may be discarded or considered separately. This means all artifactual signals can be removed from the dataset (usually the largest portion of the dataset), which is not possible in current metabolomics datasets where the high number of ‘observations’ increases the false discovery rate. IROA ClusterFinder™ software algorithms sort through the dataset, removing irrelevant noise and artifacts, allowing for a very dramatic reduction in data size.
When employing IROA, because samples are isotopically labeled, the number of carbons for each biochemical compound can be calculated from its mass spectra and the compound readily identified. The IROA technology uses a central point of reference (biochemical) embedded in every sample, thereby enabling measurements of samples with quantitative exactness.
Normalization of IROA datasets
Since the IROA technology is designed to identify artifacts and nonbiological peaks, normalization can be achieved using ‘MS total useful signal’ normalization using only the components that are common to all signals, following the removal of xenobiotics, noise and artifacts.
Application of the IROA approach in Saccharomyces cerevisiae
To illustrate the IROA methodology, the metabolite profiles of the biosynthetic compounds surrounding glutathione during the fermentation process of Saccharomyces cerevisiae were measured using isotopically defined media (whereby all carbon sources were labeled with either 5%13C/95%12C or 95%13C/5%12C) and LC–MS analysis.
The use of IROA phenotyping application for the metabolic profiling of tissues or biofluids
Where it is not possible to isotopically label experimental samples, such as biopsies or biological fluids, a ‘phenotypic’ approach to the IROA technique can be applied. Here, a complex internal standard that represents the metabolome of the fluid or tissue under study is isotopically labeled using IROA 13C media and then spiked into the experimental sample prior to analysis.
While this is a targeted approach, all labeled metabolites will have a characteristic M-1 peak and the experimental natural abundance equivalents can be identified relative to the labeled standards and artifactual peaks can be removed as they will have no match.
Conclusion & future perspective
The IROA technology has analytical advantages over existing isotopic metabolite profiling methods. However, the application of these methods for metabolomics is still in its infancy.
Key Terms
- Metabolic profiling
Quantitative analysis of metabolites, generally performed by either NMR or MS, to measure a change of the metabolome in response to a stressor
- Stable-isotopic labeling
A technique that incorporates a stable (nonradioactive) isotope into a molecule creating a higher molecular weight compound
- Isotopic Ratio Outlier Analysis™
MS method of metabolic profiling that utilizes stable isotopes to create distinct molecular signatures in biological compounds
Footnotes
For reprint orders, please contact reprints@future-science.com
Financial & competing interests disclosure
The authors are principals in NextGen Metabolomics, Inc., a Michigan business entity that holds an exclusive license to the Isotopic Ratio Outlier Analysis™ technology. The authors have no other relevant affiliations or financial involvement with any organization or entity with a financial interest in or financial conflict with the subject matter or materials discussed in the manuscript apart from those disclosed.
No writing assistance was utilized in the production of this manuscript.
References
Papers of special note have been highlighted as:
▪ of interest
- 1.Sysi-Aho M, Katajamaa M, Yetukuri L, Orešič M. Normalization method for metabolomics data using optimal selection of multiple internal standards. BMC Bioinformatics. 2007;8(93):1–17. doi: 10.1186/1471-2105-8-93. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Hellerstein MK. New stable isotope-mass spectrometric techniques for measuring fluxes through intact metabolic pathways in mammalian systems: introduction of moving pictures into functional genomics and biochemical phenotyping. Metab Eng. 2004;6(1):85–100. doi: 10.1016/j.ymben.2003.10.005. [DOI] [PubMed] [Google Scholar]
- 3.Hiller K, Metallo C, Stephanopoulos G. Elucidation of cellular metabolism via metabolomics and stable-isotope assisted metabolomics. Curr Pharm Biotechnol. 2011;12(7):105–1086. doi: 10.2174/138920111795909096. [DOI] [PubMed] [Google Scholar]
- 4.Kwok WH, Leung DKK, Leung GNW, et al. Unusual observations during steroid analysis. Rapid Commun Mass Spectrom. 2008;22:682–686. doi: 10.1002/rcm.3420. [DOI] [PubMed] [Google Scholar]
- 5.Mandal R, Guo AC, Chaudhary KK, et al. Multi-platform characterization of the human cerebrospinal fluid metabolome: a comprehensive and quantitative update. Genome Med. 2012;4:38. doi: 10.1186/gm337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Zhang Y, Tobias HJ, Brenna JT. Steroid isotopic standards for gas chromatography–combustion isotope ratio mass spectrometry (GCC–IRMS) Steroids. 2009;74(3):369–378. doi: 10.1016/j.steroids.2008.10.001. [DOI] [PubMed] [Google Scholar]
- 7.Stokvis E, Rosing H, Beijnen JH. Stable isotopically labeled internal standards in quantitative bioanalysis using liquid chromatography/mass spectrometry: necessity or not? Rapid Commun Mass Spectrom. 2005;19(3):401–407. doi: 10.1002/rcm.1790. [DOI] [PubMed] [Google Scholar]
- 8.Lewis IA, Karsten RH, Norton ME, Tonelli M, Westler WM, Markley JL. NMR methods for measuring carbon-13 isotopic enrichment of metabolites in complex solutions. Anal Chem. 2010;82(11):4558–4563. doi: 10.1021/ac100565b. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Hellerstein MK, Neese RA, Linfoot P, Christiansen M, Turner S, Letscher A. Hepatic gluconeogenic fluxes and glycogen turnover during fasting in humans. Clin Invest. 1997;100:1305–1319. doi: 10.1172/JCI119644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Schwender J, Ohlrogge JB. Probing in vivo metabolism by stable isotope labeling of storage lipiids and proteins in developing Brassica napus embryos. Plant Physiol. 2002;130(1):347–361. doi: 10.1104/pp.004275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Boros LG, Brackett DJ, Harrigan GG. Metabolic biomarker and kinase drug target discovery in cancer using stable isotope-based dynamic metabolic profiling (SIDMAP) Curr Cancer Drug Targets. 2003;3:447–455. doi: 10.2174/1568009033481769. [DOI] [PubMed] [Google Scholar]
- 12.Hood BL, Lucas DA, Kim G, et al. Quantitative analysis of the low molecular weight serum proteome using 18O stable isotope labeling in a lung tumor xenograft mouse model. Am Soc Mass Spectrom. 2005;16:1221–1230. doi: 10.1016/j.jasms.2005.02.005. [DOI] [PubMed] [Google Scholar]
- 13.Sonko BJ, Schmitt TC, Guo L, et al. Assessment of usnic acid toxicity in rat primary hepatocytes using 13C isotopomer distribution analysis of lactate, glutamate and glucose. Food Chem Toxicol. 2011;49(11):2968–2974. doi: 10.1016/j.fct.2011.07.047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Schwender J. Metabolic flux analysis as a tool in metabolic engineering of plants. Curr Opin Biotechnol. 2008;19(2):131–137. doi: 10.1016/j.copbio.2008.02.006. [DOI] [PubMed] [Google Scholar]
- 15.Beynon RJ. Stable isotope labeling with amino acids (SILAA) as an aid to protein identification in peptide mass fingerprinting. In: Conn PM, editor. Proteomics Methods and Reviews. Humana Press; NJ, USA: 2003. pp. 129–143. [Google Scholar]
- 16▪.Ong SE, Blagoev B, Kratchmarova I, et al. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol Cell Proteomics. 2002;1:376–386. doi: 10.1074/mcp.m200025-mcp200. Description of the stable-isotope labeling with amino acids methodology as a quantitative proteomic approach. [DOI] [PubMed] [Google Scholar]
- 17.Gygi SP, Rist B, Gerber SA, Turecek F, Gelb MH, Aebersold R. Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat Biotechnol. 1999;17(10):994–999. doi: 10.1038/13690. [DOI] [PubMed] [Google Scholar]
- 18.Schmidt A, Kellermann J, Lottspeich F. A novel strategy for quantitative proteomics using isotope-coded protein labels. Proteomics. 2005;5:4–15. doi: 10.1002/pmic.200400873. [DOI] [PubMed] [Google Scholar]
- 19.Ross PL, Huang YN, Marchese JN, et al. Multiplexed protein quantitation in Saccharomyces cerevisiae using amine-reactive isobaric tagging reagents. Mol Cell Proteomics. 2004;3(12):1154–1169. doi: 10.1074/mcp.M400129-MCP200. [DOI] [PubMed] [Google Scholar]
- 20.Thompson A, Schäfer J, Kuhn K, et al. Tandem mass tags: a novel quantification strategy for comparative analysis of complex protein mixtures by MS/MS. Anal Chem. 2003;75(8):1895–1904. doi: 10.1021/ac0262560. [DOI] [PubMed] [Google Scholar]
- 21.Ciccimaro E, Blair IA. Stable isotope dilution LC–MS for quantitative biomarker analysis. Bioanalysis. 2010;2(2):311–341. doi: 10.4155/bio.09.185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Beynon RJ, Pratt JM. Metabolic labeling of proteins for proteomics. Mol Cell Proteomics. 2005;4:857–872. doi: 10.1074/mcp.R400010-MCP200. [DOI] [PubMed] [Google Scholar]
- 23.Birkemeyer C, Luedemann A, Wagner C, Erban A, Kopka J. Metabolome analysis: the potential of in vivo labeling with stable isotopes for metabolite profiling. Trends in Biotech. 2005;23(1):28–33. doi: 10.1016/j.tibtech.2004.12.001. [DOI] [PubMed] [Google Scholar]
- 24▪.Mashego MR, Wu L, Van Dam JC, et al. MIRACLE: Mass Isotopomer Ratio Analysis of U-13C-Labeled Extracts. A new method for accurate quantification of changes in concentrations of intracellular metabolites. Biotechnol Bioeng. 2004;85(6):620–628. doi: 10.1002/bit.10907. Quantitation of metabolite concentrations utilizing internal standards of U-13C-labeled metabolite extracts. [DOI] [PubMed] [Google Scholar]
- 25.Bennett BD, Yuan J, Kimball EH, Rabinowitz JD. Absolute quantitation of intracellual metabolite concentrations by an isotope ratio-based approach. Nat Protoc. 2008;3(8):1299–1311. doi: 10.1038/nprot.2008.107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Bennett BD, Kimball EH, Gao M, Osterhout R, Van Dien SJ, Rabinowitz JD. Absolute metabolite concentrations and implied enzyme active site occupancy in Escherichia coli. Nat Chem Biol. 2009;5(8):593–599. doi: 10.1038/nchembio.186. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Lu W, Kimball E, Rabinowitz JD. A high-performance liquid chromatography-tandem mass spectrometry method for quantitation of nitrogen-containing intracellular metabolites. J Am Soc Mass Spectrom. 2006;17(1):37–50. doi: 10.1016/j.jasms.2005.09.001. [DOI] [PubMed] [Google Scholar]
- 28.Lu W, Kwon YK, Rabinowitz JD. Isotope ratio-based profiling of microbial folates. J Am Soc Mass Spectrom. 2007;18(5):898–909. doi: 10.1016/j.jasms.2007.01.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Nagai H, Goto T, Takahashi N. A method for the simultaneous determination of 3T3-L1 adipocyte metabolites by liquid chromatography/mass spectrometry using [13C]-stable isotopes. Biosci Biotechnol Biochem. 2011;75(8):1485–1489. doi: 10.1271/bbb.110192. [DOI] [PubMed] [Google Scholar]
- 30.Huang X, Regnier FE. Differential metabolomics using stable isotope labeling and two-dimensional gas chromatography with time-of-flight mass spectrometry. Anal Chem. 2008;80 (1):107–114. doi: 10.1021/ac071263f. [DOI] [PubMed] [Google Scholar]
- 31.Cook PF, Yoon M-Y, Hara S, McClure GD. Product dependence of deuterium isotope effects in enzyme-catalyzed reactions. Biochemistry. 1993;32:1795–1802. doi: 10.1021/bi00058a013. [DOI] [PubMed] [Google Scholar]
- 32.Bender MM, Rouhani I, Vines HM, Black CC., Jr 13C/12C ratio changes in Crassulacean acid metabolism plants. Plant Physiol. 1973;52:427–430. doi: 10.1104/pp.52.5.427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Claesen J, Dittwald P, Burzykowski T, Valenborg D. An efficient method to calculate the aggregated isotopic distribution and exact center-masses. Am Soc Mass Spectrom. 2012;23:753–763. doi: 10.1007/s13361-011-0326-2. [DOI] [PubMed] [Google Scholar]
- 34.Miller RC, Brindle E, Holman DJ. Comparison of specific gravity and creatinine for normalizing urinary reproductive hormone concentrations. Clin Chem. 2004;50(5):924–932. doi: 10.1373/clinchem.2004.032292. [DOI] [PubMed] [Google Scholar]
- 35▪.Warrack BM, Hnatyshyn S, Ott KH, et al. Normalization strategies for metabonomic analysis of urine samples. J Chromatogr B Analyt Technol Biomed Life Sci. 2009;877:547–552. doi: 10.1016/j.jchromb.2009.01.007. Comparison and recommendations for normalization strategies in metabonomics relative to the total amount of all endogenous species. [DOI] [PubMed] [Google Scholar]
- 36.Craig A, Cloarec O, Holmes E, Nicholson JK, Lindon JC. Scaling and normalization effects in NMR spectroscopic metabonomic data sets. Anal Chem. 2006;78:2262–2267. doi: 10.1021/ac0519312. [DOI] [PubMed] [Google Scholar]
- 37.Liu Y, Song XD, Liu W, Zhang TY, Zuo J. Glucose deprivation induces mitochondrial dysfunction and oxidative stress in PC12 cell line. J Cell Mol Med. 2003;7(1):49–56. doi: 10.1111/j.1582-4934.2003.tb00202.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Serkova NJ, Standiford TJ, Stringer KA. The emerging field of quantitative blood metabolomics for biomarker discovery in critical illnesses. Am J Respir Crit Care Med. 2011;184(6):647–655. doi: 10.1164/rccm.201103-0474CI. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.McKillop AM, Flatt PR. Emerging applications of metabolomic and genomic profiling in diabetic clinical medicine. Diabetes Care. 2011;34(12):2624–2630. doi: 10.2337/dc11-0837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Huege J, Goetze J, Dethloff F, Junker B, Kopka J. Quantification of Stable Isotope Label in Metabolites Via Mass Spectrometry. Humana Press Inc; NJ, USA: 2012. [DOI] [PubMed] [Google Scholar]
Patent
- 101.Metabolic Analyses Inc. US7820963. 2010