Abstract
Whole-genome enabled prediction of complex traits has received enormous attention in animal and plant breeding and is making inroads into human and even Drosophila genetics. The term “Bayesian alphabet” denotes a growing number of letters of the alphabet used to denote various Bayesian linear regressions that differ in the priors adopted, while sharing the same sampling model. We explore the role of the prior distribution in whole-genome regression models for dissecting complex traits in what is now a standard situation with genomic data where the number of unknown parameters (p) typically exceeds sample size (n). Members of the alphabet aim to confront this overparameterization in various manners, but it is shown here that the prior is always influential, unless n ≫ p. This happens because parameters are not likelihood identified, so Bayesian learning is imperfect. Since inferences are not devoid of the influence of the prior, claims about genetic architecture from these methods should be taken with caution. However, all such procedures may deliver reasonable predictions of complex traits, provided that some parameters (“tuning knobs”) are assessed via a properly conducted cross-validation. It is concluded that members of the alphabet have a room in whole-genome prediction of phenotypes, but have somewhat doubtful inferential value, at least when sample size is such that n ≪ p.
Keywords: Bayesian alphabet, whole-genome prediction, genomic selection, SNPs, marker-assisted selection, genetic architecture, quantitative traits
WHOLE-genome enabled prediction of complex traits has received much attention in animal and plant breeding (e.g., Meuwissen et al. 2001; Heffner et al. 2009; Lorenz et al. 2011; de los Campos et al. 2012a; Heslot et al. 2012) and is making inroads into human and even Drosophila genetics (e.g., de los Campos et al. 2010, 2012b; Makowsky et al. 2011; Ober et al. 2012; Vázquez et al. 2012). This approach is known as “genomic selection” in breeding of agricultural species. The term “Bayesian alphabet” was coined by Gianola et al. (2009) to refer to a (growing) number of letters of the alphabet used to denote various Bayesian linear regressions used in genomic selection that differ in the priors adopted while sharing the same sampling model: a Gaussian distribution with mean vector represented by a regression on p markers, typically SNPs, and a residual variance, . A recent review of some of these methods is in de los Campos et al. (2012a). In addition to prediction, this whole-genome approach lends itself to investigation of “genetic architecture,” often defined as the number of genes affecting a quantitative trait, the allelic effects on phenotypes, and the frequency distribution spectrum of alleles at these genes (e.g., Hill 2012). If epistasis and pleiotropy are brought into the picture, this definition of genetic architecture needs to be expanded significantly.
Most researchers in genomic selection are familiar with most letters of the alphabet, but we provide a brief review of its ontogeny. The alphabet started with Bayes A and B (Meuwissen et al. 2001), but there has been rapid expansion since, as illustrated by the Bayes Cπ and Dπ methods (Habier et al. 2011). Apart from between-letter variation, there is also variation within letters, such as fast EM-Bayes A (Sun et al. 2012), fast Bayes B (Meuwissen et al. 2009), and BRR (Bayesian ridge regression on markers), which is equivalent to G-BLUP (Van Raden 2008) but with variance parameters estimated Bayesianly; the equivalence between G-BLUP and ridge regression is given, for example, in de los Campos et al. (2009a,b). The letter D has several variants: Bayes D0, D1, D2, and D3 (Wellman and Bennewitz 2012).
Here, L is used to denote the Bayesian Lasso (Park and Casella 2008; de los Campos et al. 2009a,b) while L1 and L2 can be used to refer to variants due to Legarra et al. (2011). There is also the EL Bayesian Lasso of Mutshinda and Sillanpää (2010), with EL standing for “extended Lasso.” An almost empty hiatus spans from letters D to R (Erbe et al. 2012) with Bayes RS emerging even more recently (Brondum et al. 2012). Wang et al. (2013) presented Bayes TA, TB, and TCπ, extensions of the corresponding letters to threshold models. The upper bound of the alphabet seems to have been defined by Larry Schaeffer (personal communication, Interbull Meeting, Guelph, 2011) when he threatened attendants of this conference with Bayes Z-Δ, although full details have not been published yet. The preceding review may not be comprehensive, as there may be other members of the alphabet that are unknown to the author. It is tempting to conjecture that there may be issues with individual members of the alphabet, as this continued growth is suggestive of a state of lack of satisfaction with any given letter.
This article explores the role of the prior distribution in whole-genome regression models for predicting or dissecting complex traits. In particular, we address a standard situation encountered in genomic selection: with genomic data, the number of unknown parameters exceeds sample size. Section General Setting presents the regression model and reminds readers that, for the preceding situation, regression coefficients on marker genotypes are not identified in the likelihood function so that the data do not contain information for inference that is uncontaminated from the effects of the prior, except in a subspace. Bayesian methods for confronting the blatant overparameterization of genomic selection models are reviewed in this section, where it is shown that the prior is always influential in this setting. The section Bayesian Shrinkage discusses how ridge regression produces frequency-dependent shrinkage, while Bayes A, Bayes B, Bayes L, and Bayes R effect a type of shrinkage that is both allelic frequency and effect-size dependent. After establishing in the preceding sections, hopefully in a firm manner, that all members of the alphabet do not lead to inferences that are devoid of the influence of the prior, it is argued in the Discussion that all such methods may deliver reasonable predictions of complex traits, provided that some parameters (“tuning knobs”) are assessed properly. It is concluded that, while members of the alphabet cannot be construed as providing solid inferences about “genetic architecture,” they do have a room in whole-genome prediction of phenotypes.
General Setting
Let y be an n × 1 vector of target responses (e.g., phenotypes, preprocessed data). Using molecular markers, all members of the alphabet pose the same linear regression of phenotypes on marker codes, that is
| (1) |
where X is an n × p matrix of marker codes (e.g., −1, 0, 1 for aa, Aa, and AA genotypes, respectively); when additive action is assumed, β = {βj} is a vector of allelic substitution effects for each of p markers, and e is a vector of residuals typically assigned the normal distribution where is the residual variance, defined earlier.
In the standard additive model of quantitative genetics (e.g., Falconer and Mackay 1996), the βj are fixed parameters, while the elements xij of X are random variables; e.g., members of the jth column of X may be realizations from a Hardy–Weinberg distribution with corealizations in columns j and j′ reflecting some linkage disequilibrium distribution. The maximum-likelihood estimator of β treats X as a fixed matrix and satisfies the system of equations
where β(0) may not be a unique solution (Searle 1971). If n < p X′X is singular so the maximum-likelihood estimator is not unique, as there is an infinite number of solutions to the equations above. Letting (X′X)− be a generalized inverse of X′X, one solution is β(0) = (X′X)−X′y with expectation E(β(0)|β) = (X′X)−X′Xβ, producing a biased estimator of β, with at least p − n of its elements being equal to 0. On the other hand, E(y|β) = Xβ = g (the genetic signal captured by markers) is estimated uniquely because its estimator, Xβ(0) is unique, although this reproduces y exactly in the n < p situation. Fisher’s information content about β in the sample is and, because this matrix is singular, one cannot speak about information pertaining to individual marker effects in a strict sense. However, the information content about g = {gi} is , meaning that information about each genotypic value gi is proportional to that conveyed by a sample of size 1. Hence, in an n < p model, maximum-likelihood cannot be used either as an inferential or as a predictive machine. In the latter case, it does not generalize to new samples, because it copies both noise and signal contained in model training data.
In their proposals for employing whole-genome markers in a linear regression model, Meuwissen et al. (2001) were inspired by the fact that animal breeders had dealt with the n ≪ p problem successfully in the context of predicting random effects via best linear unbiased prediction (BLUP); see Henderson (1984) for a review, with a gentler treatment in Mrode (2005). BLUP assumes that marker effects are drawn from some distribution with known variance components; only knowledge of the covariance structure is needed and the form of the distribution is immaterial, although a linear model must hold. An alternative is provided by the Bayesian treatment but, here, the meaning of probability and the manner in which unknowns are inferred are different from their frequentist counterparts (Gianola and Fernando 1986; Robinson 1991). The distinctions between these two views are emphasized next, but the two approaches confront n ≪ p by bringing external information to the problem, as noted early in the game by Robertson (1955).
The BLUP approach to whole-genome prediction assumes that β has a null mean vector and some known covariance matrix Vβ; then E(y) = 0 and the best linear unbiased predictor of β is
Here, β is regarded as a random draw from the distribution indicated above so, on average Ey[BLUP(β)] = 0, meaning that BLUP(β) is unbiased with respect to the mean of the random effects distribution, E(β|Vβ). BLUP envisages a sampling scheme where one draws a different realization of marker effects in every repetition of the sampling, such that, over all repetitions, 0 is obtained, on average. BLUP estimates zero without bias! However, when one is interested about individual marker effects (or about the genetic values of a given individual), the inference to be made pertains to the specific item of interest, and not to the average of their distribution. If so, BLUP is biased with respect to specific marker effects (the classical fixed model of quantitative genetics) because
so that the bias is where Ip is an identity matrix of order p. The random effects treatment results in that BLUP(β) is unique whether n ≪ p or not, but it produces a biased estimator of marker effects; this bias never disappears when n ≪ p. On the other hand, if n → ∞ and p stays fixed, bias goes away, given that the model is true. A toy example of the bias of BLUP with respect to the true, fixed, substitution effects is shown in the Appendix (Bias of BLUP with respect to marker effects).
For the n ≪ p situation, Fan and Li (2001) discuss estimators that induce sparsity. However, to meet their so-called “oracle properties” (e.g., asymptotic unbiasedness), Fisher’s information matrix must be nonsingular for p0 nonzero parameters (p0 < n), these being the “true” effects of some markers on quantitative traits. BLUP and most members of the Bayesian alphabet do not produce a sparse model automatically; rather, they produce shrinkage of regression coefficients. Consider a sequence of P models of increasing dimensionality fitted to the same data, with p0 < p1 < p2 …< pP. The size of the “true” signal is dictated by the “true” effects p0 and the size of the models could be viewed as corresponding to the number of SNPs in platforms of increasing density applied to the same data set. As marker density increases while n and p0 remain fixed, estimates of marker effects must become necessarily smaller. How can “true” effects be learned properly if the model forces estimates to become smaller as p grows? Given the rhythm of technology, it is unlikely that we will reach the situation where n ≫ pP. At this point there is not much hope for learning marker effects in a manner that is free from making additional untestable assumptions. A minor complication: for the oracle properties to hold the true model must be “hit.” This is probably an unrealistic proposal when dealing with complex traits where many difficulties arise; for example, linkage disequilibrium creates ambiguity because many markers can act as proxies for others and complex forms of epistasis are bound to produce havoc in a naive linear model on additive effects.
One way of tackling the n ≪ p problem is by introducing restrictions on the size of the regression coefficients, i.e., shrinkage or “regularization.” In the machine-learning literature this is attained via ad hoc penalty functions that produce regularization (e.g., Bishop 2006; Hastie et al. 2009). Bayesian methods with proper priors produce regularization automatically, to an extent that depends on the prior adopted. The various members of the Bayesian alphabet effect shrinkage in different manners, an issue explored subsequently in this article. Let all unknown parameters of a model be represented by θ = (θ1, θ2), where θ1 and θ2 denote distinct parameters, e.g., marker effects and their apparent (the reason for these terms is made clear later) variances, respectively, in Bayes A. The posterior distribution of θ (assume, for simplicity, that the residual variance is known) is
| (2) |
| (3) |
Above, H is a set of more or less arbitrarily specified hyperparameters. Expression (3) results from the assumption that, given location effects θ1 (e.g., allelic substitution effects), the data are conditionally independent of θ2. Further,
with the expression on the right side resulting because, given θ2, location effects θ1 do not depend on H (e.g., Sorensen and Gianola 2002). Note that p(θ1|θ2) is a conditional prior distribution, while the marginal prior distribution actually assigned to θ1 is
| (4) |
Likewise, p(θ1|y, H) and p(g(θ1)|y, H) denote the marginal posterior distributions of θ1 and g(θ1), the latter being any function of θ1. For example, if θ1 is the vector β, one may be interested in the posterior distribution of Xβ, the marked signal. The results of a Bayesian analysis should not be interpreted from a frequentist perspective, as the meaning of probability is different in the two camps (Bernardo and Smith 1994; O’Hagan 1994; Sorensen and Gianola 2002). For example, BLUP is an unbiased predictor in conceptual repeated sampling, but corresponds to the posterior mean of marker effects in a Bayesian Gaussian model with known covariance structure. In the latter, the data are fixed; in the BLUP setting, the data vary at random.
An important issue is the influence of priors on inference. Theory on Bayesian asymptotics dictates that, as sample size grows, the influence of the prior vanishes gradually. In the limit, the posterior distribution becomes normal, centered at the maximum-likelihood estimator and with covariance matrix given by the inverse of Fisher’s information measure, so the prior matters little in large samples (Bernardo and Smith 1994). However, this result holds for parameters that are likelihood identifiable, i.e., when their maximum-likelihood estimator exists, but it must be kept in mind that markers are not QTL, so the marker-based model is arguably wrong. In an n ≪ p setting, true Bayesian learning can take place for at most n parameters or functions thereof, since p − n parameters are unidentified. Gelfand and Sahu (1999) show that one can learn about at most n linearly independent functions of marker effects, such as . Carlin and Louis (1996) and Sorensen and Gianola (2002) give an example where the marginal posterior distributions of unidentified parameters exist if these are assigned proper priors; however, the priors will always matter and their influence will never vanish asymptotically. In the n ≪ p setting, inferences about marker effects (often referred to as learning genetic architecture, e.g., inferring effects of some QTL) are always influenced by the priors adopted, apart from the fact that the model is wrong, as argued above. This means that stories that can be made from posterior distributions will depend on stories that are made a priori. For example, Lehermeier et al. (2013) demonstrated the influence of priors on predictive ability from various Bayesian models (Bayes A, B, L) with simulated and empirical data. Also, Gianola et al. (2009) showed that the priors in Bayes A and B drive inferences on variances of marker-specific effects.
A formal verification that individual marker effects are not identified from a Bayesian perspective using a definition by Dawid (1979) is presented in the Appendix (Marker effects are not identified from a Bayesian perspective in the n < p setting); this holds for any model, linear or nonlinear. A proof that is specific to the linear regression model on p markers with sample size n is given in the Appendix as well (Inferences in a linear model with unidentified parameters); there, it is shown that what is learned about β is a function of what is learned about Xβ. In other words, Bayesian learning occurs for n items but then this knowledge is “distributed” into p pieces via the relationship between β and Xβ induced by the prior.
In summary, proper Bayesian learning from data in a linear regression model with n < p takes place only for linear combinations of marker effects that are identified in the likelihood, that is, estimable. Any other marker effects or linear combinations thereof are redundant in the sampling model, but their posterior distributions exist and the posterior mean will differ from the prior mean. It follows that mechanistic conjectures about genetic architecture in the n < p situation are, to a large extent, driven by prior assumptions and not by data. This observation has been corroborated empirically (e.g., Heslot et al. 2012; Ober et al. 2012; Lehermeier et al. 2013): Bayesian models differing in their prior produce different inferences about individual marker effects, but most often deliver similar predictive abilities if tuned properly. Not surprisingly, the posterior distributions of (the signal to be predicted for datum i) from varying models are more similar to each other than the corresponding priors, as this function is likelihood identifiable. In short, extant theory says that given that a model is “true” (oracle principle 1, Fan and Li 2001), the posterior mean of an identifiable parameter or of a likelihood-identified combination of parameters will converge to its true value, including any “true zero,” as sample size goes to infinity (oracle principle 2). This works for n > p or for some estimators where sparsity is built in automatically, but the model must be true; Fan and Li (2001) describe several such estimators.
A situation in which proper Bayesian learning can take place is presented in the Appendix (An example of proper Bayesian learning).
Bayesian Shrinkage
Given that learning about genetic architecture without contamination from effects of the prior does not take place whenever n ≪ p, a question is what the various different members of the alphabet do. We examine ridge regression (BLUP), Bayes A, Bayes B, Bayes L, and Bayes R and also give a warning about a commonly used description of a specific prior; these procedures are prototypical so there is no need to consider other letters of the alphabet. All these methods have been reported in the genomic selection literature. Since the marginal posterior distribution of marker effects (with the exception of that of BLUP under normality) cannot be arrived at analytically, the methods are appraised from a heuristic perspective.
BLUP (ridge regression)
The vector of marker effects β is assigned the normal prior . The structure of the problem is well known and the mixed model equations leading to BLUP satisfy
| (5) |
where is the variance ratio, and β(0) = (X′X)− X′y is as before. One can write
| (6) |
so (unique) is a matrix weighted average of solution β(0) (not unique) and of the prior mean 0, where the weights are X′X and λ, respectively. For fixed p, as n increases, the rank of X′X will increase, eventually exceeding p and, in the limit, the posterior distribution will be centered at the unique maximum-likelihood estimator (by consistency, this will converge to the “true” value of β, given the model).
Representation (5) suggests that the same amount of shrinkage is effected to all p markers (because the same variance ratio λ is added to every diagonal element of X′X), but this is not the case. This is clear from Equation 6 where, for each marker effect, the contributions from the data vary over markers; this is more transparent from inspection of the solutions in scalar form. For marker 1, as an example, the estimator of substitution effect is
where
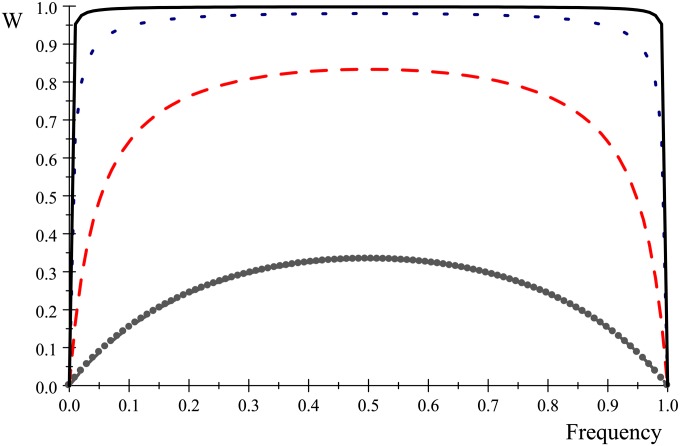
Then, the BLUP of the allele substitution effect can be viewed, heuristically, as a weighted average of a “data driven” estimate and of the mean of the prior distribution (0) where the respective weights are and respectively. This suggests less shrinkage toward zero for markers (j, say) having larger values of . Now, if for any marker genotypes are coded as −1, 0, 1 for aa, Aa, and AA, respectively, it follows (assuming Hardy–Weinberg proportions and centered marker codes) that so where pj is the frequency of the A-type allele at that locus. Hence, at fixed sample size n, BLUP effects less shrinkage toward zero of markers that have intermediate allelic frequencies, simply because 2pj(1 − pj) is maximum at . To illustrate, we use this Hardy–Weiberg approximation and plot (Figure 1) the weight “assigned to the data” for marker j
against allelic frequency at and 0.001, respectively. As depicted in Figure 1, the extent of shrinkage is frequency and sample size dependent, with some differential shrinkage (bottom two curves) taking place at large values of , that is, at small sample sizes, but with little or no differential shrinkage otherwise, unless alleles are rare. Then, the often-made statement that BLUP or ridge regression perform an homogeneous shrinkage of marker effects is not correct. In short, shrinkage is frequency and sample size dependent but effect-size independent.
Figure 1.
Approximate weight (W) assigned to the data (the weight assigned to prior information is 1 − W) as a function of allelic frequency at a marker locus. From top to bottom the lines give the trajectory for λ/n values of 0.001 (solid line), 0.01 (short dashes), 0.1 (long dashes), and 1 (dots).
Bayes A
Bayes A (Meuwissen et al. 2001) consists of a three-stage hierarchical model. The first stage is the normal regression (1); the second stage assigns a normal conditional prior to each of the marker effects, all possessing a null mean but with a variance that is specific to each marker; the third stage assigns the same scaled inverted chi-square distribution with known scale and degrees of freedom (ν) parameters to each of the marker variances. The mechanistic argument for the Bayes A prior was that markers may contribute differentially to genetic variance (they do, to an extent depending on their effects, allelic frequencies, and linkage disequilibrium relationships with causal variants), so it seemed a good idea to “estimate” such variances. There are two difficulties: the first one is that the marginal prior for the markers effects, resulting from deconditioning the second stage over the third stage as done in (4), is the same for all markers. Second, there is scant Bayesian learning for marker-specific variances. This was pointed out by Gianola et al. (2009), who showed that all markers have the same prior distribution: a process with null mean and variance . Given that this prior is homoscedastic over markers, why is it that it behaves differently from ridge regression BLUP, where all individual marker effects are assigned the prior
In Bayes A, the marginal posterior distribution of marker effects cannot be arrived at in closed form, but insight can be obtained from inspection of the joint mode of the posterior distribution of β, assuming that the residual variance is known; recall that and ν are known hyperparameters in Bayes A. The hierarchical model is then
where is the ith row of X. Conditionally on , and ν, the joint posterior density is
| (7) |
Using results presented in the Appendix (Mode of the conditional posterior distribution in Bayes A), an iterative scheme for locating a mode of (7) is given by
| (8) |
with successive updating; here,
is a diagonal matrix. If this converges, it will do so to one of perhaps many stationary points, as it is known that t-regression models may produce multimodal log-posterior surfaces, especially if ν is small (McLachlan and Krishnan 1997). Hence, iteration (8) may lead to a point receiving little posterior plausibility.
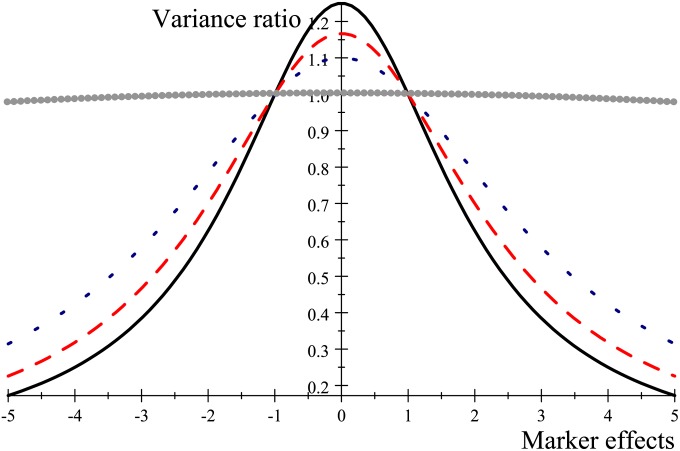
The role of in (8) parallels that of the inverse of the genetic variance–covariance matrix (times ) in standard BLUP (Henderson 1984), so that the larger is, the stronger the shrinkage toward 0 (mean of the prior distribution). However, while the variance ratio is constant in ridge regression BLUP, here it varies over markers, as it takes the form . As ν → ∞ (the t-distribution approaches a normal one), , resembling λ of BLUP. On the other hand, if the t prior has a finite number of degrees of freedom, markers whose effects are closer to 0 are shrunk more strongly than those with larger absolute values, simply because λj is larger for the former. To illustrate, let so that the “variance ratio” is . Figure 2 illustrates the impact of the marker effect on the “variance ratio” for ν = 4, 6, 10, and 1000. It is seen that λj becomes smaller (less shrinkage toward zero) as the absolute value of the effect of the marker increases; also, shrinkage increases as the degrees of freedom of the distribution increase, at any given marker effect. Eventually, when ν → ∞ (so that the prior is normal) the variance ratio takes the same value for all markers (thick line in Figure 2, almost horizontal, corresponding to ν = 1000). For markers with effects near zero, the t-distribution shrinks effects more strongly than the normal process, but it does not severely penalize markers having strong effects on the phenotype. Hence, in Bayes A shrinkage is marker-effect specific, with this specificity becoming milder as the value of ν increases. Note that (7) also induces frequency-specific shrinkage, due to the Bayesian compromise between the prior and X′X, as in the case of BLUP. Hence, apart from the effects of sample size, there are two sources of shrinkage in Bayes A, contrary to a single one in BLUP. This seems to confer Bayes A more flexibility than BLUP, but this is not necessarily good because the extra parameters ν and (this one playing the role of ) are influential, and may affect “inference” of marker effects adversely (Lehermeier et al. 2013). Naturally, these parameters can be assigned priors and inferred from the resulting Bayesian model, but this was not suggested by Meuwissen et al. (2001).
Figure 2.
Impact of marker effect and of the degrees of freedom (d.f.) parameter on the extent of shrinkage toward the prior distribution in Bayes A: the larger the value in the y-axis, the stronger the shrinkage toward 0. d.f. = 4, solid line; d.f. = 6, long-dashed line; d.f. = 10, short-dashed line; d.f. = 1000, gray circles, almost horizontal.
Bayes B
A formulation of Bayes B as a mixture at the level of effects, but not of their variances, as in Meuwissen et al. (2001), is in Gianola et al. (2009) and Habier et al. (2011). The hierarchical model is
The prior is a mixture of a “0-state” (a point mass at 0) with a t-distribution, the mixing probabilities being π and 1 − π, respectively, where π is assumed known and specified arbitrarily. Recall (in informal notation) that
where is a scaled-inverted chi-square distribution assigned as prior to the variance of the jth marker effect, . Meuwissen et al. (2001) formulated the mixture at the level of these variances, arguing as follows: “the distribution of genetic variances across loci is that there are many loci with no genetic variance (not segregating) and a few with genetic variance.” Gianola et al. (2009) were critical of this formulation, both from statistical and genetic points of view.
The hierarchical prior is deceiving because, in fact, Bayes B ends up assigning the same marginal prior to every marker. This follows from consideration of the mean and variance of a mixture, e.g., Gianola et al. (2006). The mean of a mixture is the weighted average of the means of the components (the weights being the mixing probabilities π and 1 − π) and the variance is the weighted average of the component variances, plus a term that can be interpreted as “variance” among component means. One has
where is the mean of the t-distribution, and
Above, is the variance of the t-distribution. It follows that Bayes B assigns, a priori, the same mean and variance to all marker effects and that it uses a prior that is even more precise than the prior in Bayes A (the prior variance is reduced by a fraction π in Bayes B, relative to that of Bayes A). This makes effective Bayesian learning even more difficult in Bayes B than in Bayes A, as it takes more information from the data to “neutralize” the prior of Bayes B than that of Bayes A. At any rate, none of these two regression models allows for proper learning about marker effects or genetic architecture in the n ≪ p setting, as argued earlier in this article.
As for Bayes A, no closed forms for the marginal posterior distributions of marker effects exist for Bayes B. The posterior expectation of β is
| (9) |
This indicates that shrinkage toward 0 is stronger than in Bayes A since posterior means are smaller in Bayes B by a fraction π. Coupled with the arbitrary assignment of a value to π, the implication is that the the prior is even more influential in Bayes B than in Bayes A. This could have been expected intuitively, but the point has not been made before, at least in this manner.
Wimmer et al. (2012) noted that methods such as Bayes B have yielded better predictive abilities than BLUP in many simulation studies reported in the literature, but that this has not been observed with real data (e.g., Ober et al. 2012). Wimmer et al. (2012) investigated predictive abilities of these two methods in maize and in Arabidopsis. The target populations differed in effective population size and in extent of linkage disequilibrium. Despite expected differences in genetic architecture among populations and traits, predictive abilities delivered by BLUP and Bayes B did not differ significantly for their target traits. Further, they found via simulation (personal communication) that Bayes B was effective for learning genetic architecture in the n ≪ p setting only when the number of true nonzero marker effects (s) is such that s ≪ n, given the true model. Otherwise, the error of estimation of marker effects was as poor as that of BLUP, the latter found to be more robust over a wide range of situations (this is ironic, because BLUP or G-BLUP were not tailored for learning genetic architecture). In short, they confirmed that, provided one “hits” the true model (thus fulfilling oracle property 1 of Fan and Li 2001), effective learning of “true genetic architecture” is possible only if the model is very sparse relative to sample size. The condition s ≪ n would lead to oracle property 2, as anticipated by standard asymptotic Bayesian theory under regularity conditions.
BayesSSVS was proposed by Verbyla et al. (2009), but it is not discussed here because it is similar to Bayes B. Bayes Cπ of Habier et al. (2011) provides a more sensible formulation of the mixture, but it is similar in spirit and shares the same limitations of Bayes B, since parameter identification is not attained for most of the unknowns. An interesting example of consequences of overparameterization in Bayes Cπ is provided by Duchemin et al. (2012); these authors noted that as values of π went up in the sampling process, realizations of marker effect variances went down. Hints about genetic architecture from Bayes B or Bayes Cπ or from other members of the alphabet should be taken very cautiously, at least when n ≪ p.
Bayes L
Lasso regression (Tibshirani 1996) inspired the Bayesian Lasso (Bayes L here) of Park and Casella (2008), a method with followers such as Vázquez et al. (2010) and Crossa et al. (2010) and with an implementation available in the software R described by Pérez et al. (2010). The linear regression model is given in (1), but the prior assigned to marker effects is a Laplace (double exponential, DE) distribution. All marker effects are assumed to be independently and identically distributed as DE with the prior density being
| (10) |
Here, E(β|λ) = 0 and for all markers; as λ increases the variance of the DE distribution decreases and the density becomes sharper. This prior assigns the same variance or prior uncertainty to all marker effects, but it possesses thicker tails than the normal prior. A comparative discussion of the DE prior is in de los Campos et al. (2012a). Even though Bayes L bears a parallel with the Lasso, it does not “kill” or remove markers from the model, contrary to what happens in variable selection approaches. Bayes L poses a leptokurtic prior, so it is expected to shrink effects more strongly toward zero than the Gaussian prior, as opposed to inducing sparsity in the strict sense of the Lasso.
Bayes L shrinks strongly:
To appraise how Bayes L shrinks marker effects, we examine the mode(s) of the joint posterior distribution of β using the DE prior (10), assuming that λ and the residual variance are known. As in Tibshirani (1996), write with this representation , where . Using this, the log-posterior (apart from an additive constant) is
| (11) |
If, as in Tibshirani (1996), it is ignored that is a random matrix (because it is a function of |βj|) this takes the form of a standard BLUP representation, so the mode of the conditional posterior distribution of β satisfies
| (12) |
Contrary to BLUP–ridge regression where shrinkage factors are marker effect independent, these factors take the form in Bayes L, implying that markers with tiny effects are shrunk more strongly toward zero, as a larger number is added to the diagonal elements of the coefficient matrix leading to solution . Note, however, that (12) is not an explicit system, so it would make sense to iterate; details on an iterative scheme are in the Appendix (Mode of the conditional posterior distribution in Bayes L).
The preceding implies that Bayes L produces a more “effectively sparse” model. This can be seen from inspection of an “effective number of parameters” measure (e.g., Tibshirani 1996; Ruppert et al. 2003) given by
and
If X is orthonormalized, so that X′X = I (with dispersion parameters scaled accordingly)
| (13) |
and
| (14) |
This enables us to see that, in ridge regression, every degree of freedom (contributor to model complexity) represented by a column of the orthonormalized marker matrix is attenuated by the same factor, . On the other hand, in Bayes L markers having tiny effects are effectively, but not physically, wiped out of the model. Also, markers with strong effects receive a heavier weight in this overall measure of complexity.
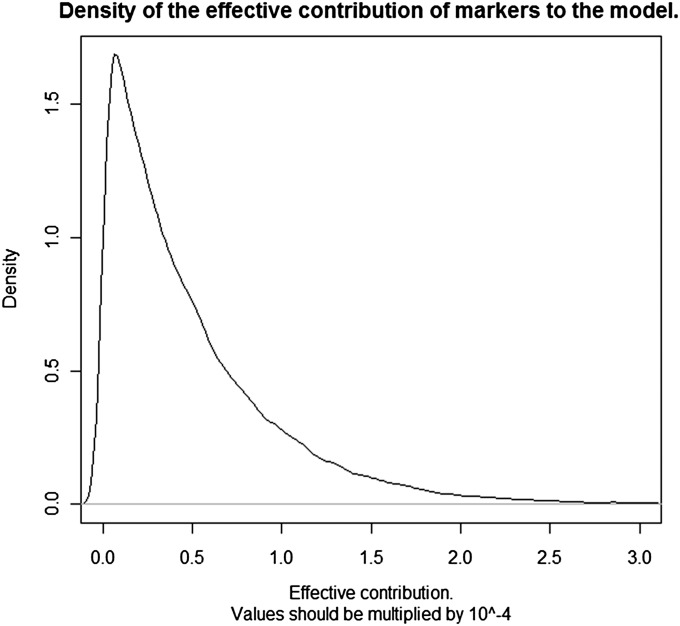
We simulated P = 100, 000 marker effects from DE distributions with mean 0 and variances 10−16, 10−8, or 10−4; setting , the preceding three values can be interpreted as the contribution of an individual marker to variance relative to residual variability. When a marker effect had a large variance (10−4), the entire battery of markers, assuming a priori independence of effects, represented of the total variance; on the other hand, when markers were assigned a variance of 10−16, markers accounted for about of the total variability. Since the variance of the DE distribution is the settings led to λ values of , , and , respectively; larger values of λ produce stronger shrinkage toward 0. The shrinkage factor is for marker j in Bayes L vs. in ridge regression–BLUP. The contribution of a marker to the model was assessed as follows: from (13), in ridge regression each marker contributes the same amount, to model complexity, whereas in Bayes L the corresponding metric is , as given in (14). For ridge regression, the effective number of parameters was approximately 10−11, 10−3, and 10, for , and 10−4, respectively. For Bayes L, the corresponding effective number of parameters was 4.96 × 10−12, 4.98 × 10−4, and 4.98, respectively. Clearly, Bayes L produced a model that was more sparse than ridge regression–BLUP. Each of the markers made a tiny contribution to model complexity; for instance, when the variance of the double exponential of marker effects was 10−16, the relative contributions to the model of individual markers ranged from 0 to 10−16; when the variance was 10−8, these ranged from 0 to 10−8, while the range was 0 − 10−4 for . A plot of the density of the effective contributions to the model of each of the 100, 000 markers is in Figure 3, for the case ; >95% of the markers contributed <2 × 10−4 effective degrees of freedom to the model. Hence, when a marker contributes to variance in a tiny manner, shrinkage of their individual effects toward 0 is very strong. Then, if a marker effect conveys the meaning of a fraction equal to 10−8, say, of some physical parameter, what can this tell us about the state of nature (i.e., genetic architecture) in the absence of effective Bayesian learning, as argued earlier in the article? Probably not much unless n ≫ p and the model fitted is the “true” one, the latter requiring the extraordinarily strong assumption that a complex trait is well represented by a (multiple) linear regression.
Figure 3.
Density (over 100,000 markers) of the “effective degrees of freedom” contributed by a marker to the model for a double exponential prior distribution with variance 10−4.
Bayes L With Gamma prior for λ2:
The DE density (10) is indexed by a single positive parameter λ, and if this is treated as unknown, the marginal prior density of a marker effect is
where p(λ) is the prior density of λ. Clearly E(β) = EλE(β|λ) = 0, but the prior variance of β will depend on the distribution assigned to λ. Typically, a Γ(r, δ) prior is placed on λ2 with the density being
| (15) |
and and . Since λ is positive, p(β|λ) = p(β|λ2), so that
| (16) |
Using in the above expression an approximation given in the Appendix (Approximation of an integral in Bayes L), Equation 16 gives
| (17) |
where ∝approx.means “approximately proportional to.” If only the first term of the approximation is used, after normalization one gets
| (18) |
and this is a DE density with parameter If both the first and second terms of the approximation are employed, one gets
| (19) |
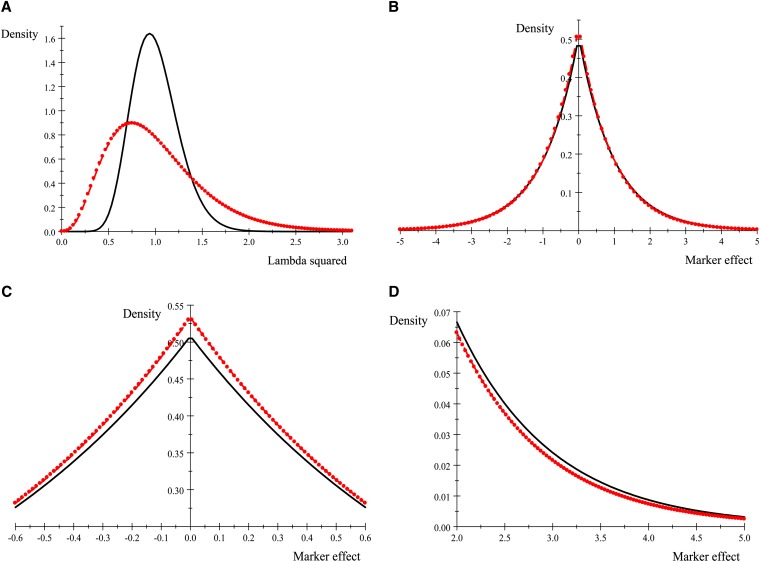
Next, we examine the shape of the unnormalized density (19) for two different Γ(r, δ) prior distributions of λ2. Setting r = δ gives Gamma distributions with expected value 1 and variance ; use of r = δ = 4 and r = δ = 16 produces prior distributions with variances and , respectively, and the corresponding densities are shown in Figure 4, top left. Taking into account that the prior distributions of marker effects have null means, the variance of approximation (19) to the marginal prior of β was evaluated by numerical integration between −9 and 9 as
yielding 1.73 (δ = 4) and 1.93(δ = 16). This produces a seemingly paradoxical situation, where the more uncertain prior for λ2 (δ = 4) gives a marginal prior for the marker effect that is more precise (as measured by the variance) than that for δ = 16. The densities, shown in Figure 4, top right, seem indistinguishable. However, if the plot is zoomed in the middle and right tails of the distribution (left and right bottom, respectively) the prior with δ = 16 turns out to be less sharp and with thicker tails, thus explaining its larger variance. Also, the prior probability that a marker has an effect ranging from −0.3 to 0.3 is 0.274 for δ = 4 (more variable prior for λ2) and 0.263 for δ = 16; the probabilities that a marker has an effect from 2 to 7 are 0.058 (δ = 4) and 0.065 (δ = 16), respectively.
Figure 4.
(A) Gamma prior density of lambda square for r = δ = 4 (dot–dash) and r = δ = 16 (solid). (B) Marginal prior densities of marker effects for r = δ = 4 (dot–dash) and r = δ = 16 (solid). (C) and (D) focus on the middle and right tails of the densities displayed in B, respectively.
Bayes L with uniform prior on λ:
In an attempt to make the prior in a Bayesian analysis less aggressive, one may naively think that Bayes’s “principle of insufficient reason” (the uniform prior) may render the analysis objective. Let the uniform prior on λ be λ|L, U ∼ Uniform(L, U), where L and U are the lower and upper bounds, respectively, of the prior distribution. Mixing the DE distribution with parameter λ over this prior gives as marginal density
As before, we employ a Taylor series to approximate exp(−λ|βj|), but now around the expectation of the uniform distribution, giving
Then
| (20) |
If the constant and the linear terms of the expansion are retained this produces
Since λ is positive, one can set L = 0 and yielding
A plot of punif,1(βj|L = 0, U) is shown in Figure 5. As U increases, the prior distribution of the marker effect gets increasingly concentrated near 0, reaching a point mass in the limit. This implies that the regression model becomes effectively very simple if U is assigned large values, as most regression coefficients take values close to 0. In theory, this should produce underfitting and out of sample predictions that do not generalize well. It is thus intriguing why Legarra et al. (2011) obtained reasonable predictive accuracies when placing a uniform prior on λ, with L = 0 and U = 106. This theoretical excursion suggests that a big warning should be inserted in documentation of software implementing DE regression models with a flat prior on the regularization parameter λ.
Figure 5.
Prior density of a marker effect when a uniform (0, U) prior is adopted for λ at varying values of upper bound U of the uniform distribution: 1 (solid line), 5 (dashes), 10 (dots–dashes).
On parameterizations of Bayes L:
How any Bayesian or “classical” model is parameterized depends on mechanistic (e.g., interpretation with respect to some theory) or computing considerations, but alternative parameterizations must be equivalent in terms of the inference attained. For example, a parameterization of the classical infinitesimal model (e.g., Hill 2012) in terms of additive genetic and environmental variances (VA, VE) must be equivalent to parameterization (V − VE, VE), where V is the phenotypic variance, or to parameterization (Vh2, (1 − h2)V), where h2 is heritability. The second and third parameterizations do not imply causally that the genetic variance depends on the environmental variance or that the environmental variance depends on heritability. In likelihood-based inference there is invariance of parameters under transformation. However, care must be exercised in Bayesian analysis because parameters are random, so any rotation of coordinates (some transformations involve nonlinear rotations) require intervention of the Jacobian of the transformation. One can go back and forth between parameterizations, provided that probability volumes are preserved properly. For instance, if one assigns independent priors to h2 and V in a (Vh2, (1 − h2)V) parameterization, those used in a VA, VE parameterization should be probabilistically consistent with the preceding, such that samples from the joint posterior of h2 and V produce the same distribution as that obtained by sampling from the joint posterior of VA, VE. Further, conditioning and deconditioning may be necessary due to computing issues, e.g., the Gibbs sampler works with conditional distributions, but the algorithm automates the deconditioning. It is precisely in this context that Legarra et al. (2011) misinterpreted the parameterization of Bayes L in Park and Casella (2008), de los Campos et al. (2009b), Weigel et al. (2009), and Vázquez et al. (2010) who, instead of working directly with prior (10) adopted a conditional prior discussed further below. All these authors have applied this parameterization successfully using data from animals and plants.
For reasons related to the behavior of Markov chain Monte Carlo algorithms for Bayes L, Park and Casella (2008) introduced a conditional DE distribution, with density This distribution has mean and variance this, of course, is not the variance of βj. Legarra et al. (2011) incorrectly wrote and made the statement: “we do expect the distribution of SNP effects not to be related to unobservable, unaccounted (residual) effects that can, for example, vary from site to site for the same individuals.“ It is fairly obvious that cannot be Var(βj) since
with the term dropping because it is null. Hence, Var(β|λ) depends on the prior adopted for . If is assigned a scaled inverted chi-square distribution on νe degrees of freedom and with scale , with density as in (29),
| (21) |
Therefore, the variance of the prior distribution of marker effects does not depend on but, rather, on λ2 and on the parameters of the prior distribution of . There is the additional complication that (21) does not take into account uncertainty associated with λ, and this is examined next.
Since λ must be positive, conditioning on λ is equivalent to conditioning on λ2, so that , and
Hence, unconditionally, use of (21) in produces
If a Γ(r, δ) prior is placed on λ2, with density (13)
Changing variables to gives
The integral is the expected value of a random variable (θ) following an inverted Gamma distribution with parameters r and δ, which is (r > 1), so
| (22) |
As argued in Gianola et al. (2009), the connection between the variance of the prior distribution of marker effects and additive genetic variance is subtle and elusive. If were to be viewed as the variance of an additive effect in some infinitesimal model, how are its different components interpreted? If the standard infinitesimal model is parameterized in terms of (VE, h2) one can write
In (22) is the counterpart of VE, since this is the expected value of the prior distribution assigned to the residual variance, . Then, plays the role of ; since is the prior expectation of , it would turn out that would be the counterpart of .
The statements made in Legarra et al. (2011) are misleading due to an incorrect interpretation of the parameterization of Bayes L proposed by Park and Casella (2008), used to address a multimodality problem that seems to arise in nonhierarchical implementations of Bayes L in the sense of Kärkkäinen and Sillanpäa (2012). These authors reported that hierarchical and nonhierarchical versions of the Bayesian Lasso led to different posterior inferences, but could not find clear reasons for this discrepancy. It might be related to lack of convergence of the Markov chain Monte Carlo scheme in the nonhierarchical parameterization or perhaps to some impropriety. Additional basic research is needed to explain this paradox, but Kärkkäinen and Sillanpäa (2012) recommended the hierarchical implementation, possibly because of easier computation.
Bayes R
Erbe et al. (2012) presented this method as follows. Bayes R starts the hierarchical model with (1) and poses a mixture of four zero-mean normal distributions as a conditional prior for a specific SNP effect:
| (23) |
Here, if the SNP effect is generated from the first component of the mixture (with probability π1) it will be 0 with complete certainty; if drawn from the second component it will have a normal distribution with null mean and variance and so on. In Bayes R, is the assumed genetic variance, r2 is the assumed reliability, and σ2 is the variance of the target trait. Presumably, the assumption about r2 is either model derived or based on prior cross-validation information, which is good Bayesian behavior, normatively. Makowsky et al. (2011) gave evidence that what one assumes about genetic variance from inference in training data is not recovered in cross-validation.
The mean of the mixture is obviously 0. Since the four components of the mixture have null means, the variance, given π = (π1, π2, π3, π4,), is
Further,
Erbe et al. (2012) used a Dirichlet distribution with parameter vector α = (α1,α2,α3,α4)′ as prior for the elements of π, so that
| (24) |
In particular, Erbe et al. (2012) took α1 = α2 = α3 = α4 = 1, producing a uniform distribution on π. It follows that all SNPs have the same marginal prior distribution, with null mean, and variance
This suggests that a simple ridge regression–BLUP obtained by solving
may deliver predictive abilities that are similar to those of Bayes R, except that it would differ with respect to Bayes R on how marker effects are shrunk.
Insight on how shrinkage takes place in Bayes R is gained by inspecting the joint posterior density of all marker effects, given r2, σ2, and π. Here
| (25) |
where , and (these values can be modified a piacere). Taking derivatives of the log-posterior with respect to β gives (apart from an additive constant)
| (26) |
where {.} denotes a p × 1 vector. Above, (i = 2, 3, 4) is the density of βj under the normal distribution corresponding to component i of the mixture, with
Employing the preceding expression in Equation 26 yields
| (27) |
Setting this to zero and rearranging leads to iteration
where is a p × p diagonal matrix with typical element
| (28) |
where
This is interpretable as the probability that a value βj in the course of iteration comes from the ith component of the mixture, as the value of βj changes iteratively. Note that Ωjj,β is a weighted average of the shrinkage factors corresponding to those that would be employed if the variance parameter of the ith component of the mixture were to be used in ridge regression–BLUP. If is taken as constant over the three “slab” components, Bayes R reduces to BLUP. On the other hand, when varies over components, the ratio will be larger for components having the smallest variance. Observe that π1 does not play a role in this posterior mode interpretation of how Bayes R effects shrinkage.
In summary, Bayes R assigns the same prior distribution to all markers in the battery of SNPs, one with null mean and variance (for a mixture of K components)
where the α′s are the parameters of the prior distribution of the mixing probabilities π. Bayes R takes .
The superior performance of Bayes R over other methods found by Erbe et al. (2012) probably results from using prior empirical knowledge about r2, the assumed reliability. Bayes R has been extended to Bayes RS (Brondum et al. 2012). This is a minor variant of Bayes R in which the mixture (23) is expanded by a factor S, so that there are now S mixtures of four normal distributions each. The letter S denotes a number of chromosome segments constructed in some manner that reflects prior knowledge that some such segments contribute more variance than others. Using the arguments outlined above, it is easy to see that Bayes RS leads to a shrinkage that, instead of being component specific, is now region-component specific.
An incorrect prior often used in the Bayesian alphabet
The following statement is found at high frequency in the genomic selection literature: “The prior distribution of the residual variance is , meaning that the degrees of freedom of the prior is −2 and that the scale parameter is null.” Examples are Meuwissen et al. (2001) and Jia and Jannink (2012), respectively. Note that Bayes’s theorem returns with null posterior density or probability any parameter value that is assigned 0 density or mass a priori. If the prior density (or probability) of parameter θ is such that p(θ|hyperparameters) = 0, it must be that
as well. The prior is absurd for two reasons. First, a scaled inverted chi-square distribution exists only if both νe and are >0. To see the second reason, we write the prior density explicitly, that is,
| (29) |
so for and any “legal” value of νe, . Then, it must be that as well. Hence, a scaled inverted chi-square with a null scale parameter is not a probability model at all, as it does not assign appreciable density to any value of the unknown residual variance. It does not convey uncertainty whatsoever: any value of the residual variance is assigned a density of zero, prior and posterior to observing data.
A possible reason for this mistake is as follows: Sorensen and Gianola (2002), as many other Bayesians often do, write the prior as being proportional to the kernel of the scaled inverted chi-square density, that is, as
and note that this kernel reduces to a uniform distribution by taking νe = −2 and , yielding . However, it takes more than a kernel to make a density, as multiplication of 1 times the integration constant produces zero.
Discussion
The main message from this article is that it is not clear how one can learn about genetic architecture from data in n ≪ p situations. This is because individual marker effects are not estimable from the likelihood, apart from the fact that it is unlikely that a multiple linear regression provides a sensible description of biological complexity. On the other hand, it is feasible to learn about the signal Xβ because there is information about this unknown vector in the data, although equivalent to that conveyed by a sample of size 1. Unfortunately, the Bayesian alphabet (Gianola et al. 2009) continues to grow under the incorrect perception that different specifications stemming from various choices of prior inform about genetic architecture; Erbe et al. (2012) and Brondum et al. (2012) provide good examples of this. It is difficult to defend such claims unless n ≫ p and provided that the model is “true” and effectively sparse (Wimmer et al. 2012). Otherwise, the prior always matters whenever n ≪ p and different priors lead to different claims about the state of nature, merely because their shrinkage behavior in finite samples varies. All members of the alphabet produce unique point and interval Bayesian estimates of marker effects, but the driver is the prior and not the data.
Mixtures of Gaussian distributions are widely used in nonparametric density estimation (Wasserman 2010) because most distributions can be approximated well. Mixtures can capture vagaries from cryptic distributions but at the expense of parsimony, thus posing the risk of copying noise, as opposed to signal, especially if the mixture model has too many parameters. McLachlan and Peel (2000) give a warning: estimation of the parameters of a mixture (Ψ) on the basis of data are meaningful only if Ψ is likelihood identifiable. In Bayes RS (apart from nuisance effects and the residual variance) the number of unknown parameters is 2p + 4S. Here, 2p comes from the fact that each marker is assigned a distinct variance; the 4S comes from the fact that there are S segments each having four segment-specific mixing probabilities πs(s = 1, 2, …, S). Unfortunately, n ≪≪< 2p + 4S, and this creates a huge identification deficit relative to the information content in a sample of size n. In a Bayesian context, there is the additional issue (occurring even when n > p) called label switching, leading Celeux et al. (2000) to write: “Although somewhat presumptuous, we consider that almost the entirety of Markov chain Monte Carlo samplers for mixture models has failed to converge!” In view of these pitfalls, one wonders what meaningful mechanistic sense can be extracted from these richly parameterized specifications intended to inform about genetic architecture.
Although their inferential outcomes may be misleading, one should not dismiss the potential value of Bayes B, C, R, RS, or of any of the mixture models proposed so far as prediction machines. Predictive distributions stemming from the various members of the alphabet may be analytically distinct from each other, but such differences are seldom revealed in cross-validation (e.g., Heslot et al. 2012); an exception is Lehermeier et al. (2013). Below we review how the alphabet can be interpreted from a predictive perspective.
A pioneer of Bayesian predictive inference (Geisser 1993) wrote:
Clearly hypothesis testing and estimation as stressed in almost all statistics books involve parameters…this presumes the truth of the model and imparts an inappropriate existential meaning to an index or parameter…inferring about observables is more pertinent since they can occur and be validated to a degree that is not possible for parameters.
Bayesian methods play an important role in machine learning (e.g., Bishop 2006; Barber 2012; Dehmer and Basak 2012; Rogers and Girolami 2012). A reason is that Bayes’s theorem provides a predictive distribution automatically, something that has not been appreciated in full yet in the whole-genome prediction literature.
The problem of prediction can be cast as one of making statements about future data yf, given past data y. A model M (e.g., Bayes L) with parameter vector θ is fitted (trained) to y, leading to the posterior distribution p(θ|y, H, M), where H denotes hyperparameters. If yf is treated as an unknown, the prior becomes p(θ, yf|H, M) = p(yf|θ, M)p(θ|H, M) so that
Since past observations do not depend on future observations, given the parameters, p(y|θ, yf, M) = p(y|θ, M), so that
| (30) |
This is the predictive distribution, where parameters θ do not necessarily play an “existential role” in the sense of Geisser (1993); rather, they are tools enabling one to go from past to future observations. Note that
meaning that the predictive distribution weights an infinite number of predictions made at a specific values of θ, with the averaging distributions being p(θ|y, H, M); this posterior conveys the plausibility assigned to a specific value of θ, posterior to the observed data y. For example, for ridge regression–BLUP, the posterior distribution of β is where is the solution to Equation 5. It follows that the posterior distribution of the signal is This implies that the predictive distribution of a future vector of data yf = Xfβ + ef would also be normal
Here, the strong assumption that the stochastic process generating current and future data are the same is made; typically, it is assumed that but this may not be realistic. While the different priors of the alphabet lead to different predictive distributions, it is to be expected that at least the point predictions will be fairly similar. This is because Xβ is identified in the likelihood, so some Bayesian learning about the signal will take place, especially when y is a vector of preprocessed means (e.g., means of daughter yield deviations for a battery of dairy cattle bulls with a large number of progeny records). In the latter case, the various members of the alphabet are expected to differ minimally in predicting ability.
The predictive distribution can be used to check whether observed data are consistent with what a model would lead one to expect. Sorensen and Waagepetersen (2003) used this idea to examine goodness of fit of model for litter size in pigs. However, the predictive approach outlined above does not take uncertainty about the model into account, and this may understate variability seriously. Bayesians address this via model averaging, where the predictive distribution is averaged over models, that is,
This integral represents both the situation where the number of models is finite and countable or infinite. In the first case the integral is a sum and the measure μ(M|y) is the posterior probability of model M. In the second case the number of possible models may be huge, e.g., in variable selection approaches for linear models aiming to include or exclude p markers, there are 2p possible specifications. If p is very large, the number of models is practically infinite, so the measure μ(M|y) is the posterior density assigned to a specific model.
Although p(yf|y) provides a more sensible assessment of predictive uncertainty, in practice one proceeds by constructing cross-validation distributions, with respect to one or several competing models. Each prediction generates an error, and this error will have a cross-validation distribution. The relevance of cross-validation is another important contribution of Meuwissen et al. (2001) to whole-genome prediction. Here, hyperparameters of genomic selection models (e.g., π in Bayes Cπ) can be viewed as “tuning knobs” and evaluated over a grid. Unfortunately, the reality is that Manhattan plots tend to overwhelm cross-validation graphs in genome-wide association studies.
Also, differences in predictive ability are often masked by the variation conveyed by a properly constructed cross-validation distribution (e.g., González-Camacho et al. 2012). On the other hand, the various Bayesian predictive machines resulting from different priors may possess differential robustness in finite samples. For instance, some priors may be less sensitive with respect to differences in true genetic architecture (Wimmer et al. 2012).
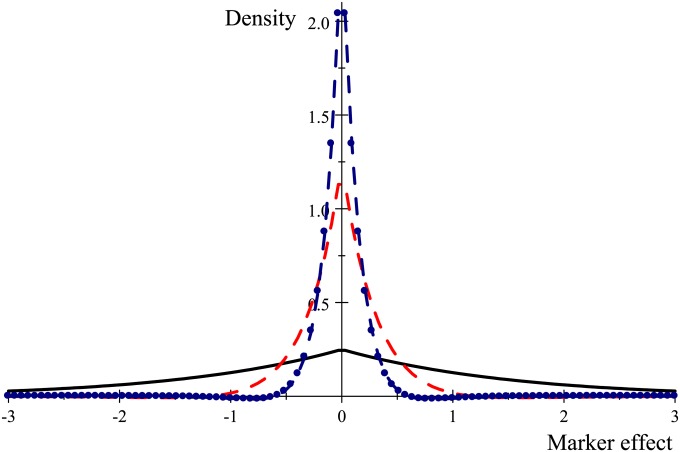
Given that the data do not contain information about individual marker effects, variation in inference is an artifact caused by the various priors. This leads to the question: How much does one prior differ from another one? Information on this can be obtained by use of some notion of statistical distance between distributions, such as the Kullback–Leibler (KL) metric. For example, Gianola et al. (2009) used KL for debunking the notion that marker-specific-effect variances in Bayes A tell us something about genetic variability of chromosomal regions. Recently, Lehermeier et al. (2013) used a metric that is easier to interpret than KL, the Hellinger distance or HD (e.g., Roos and Held 2011). They found that Bayesian learning in Bayes A and Bayes B was more limited than with Bayes L or Bayesian ridge regression. In our context, the HD between prior assigned to a marker effect in ridge regression and prior of Bayes A is
HD takes values between 0 and 1, with 1 corresponding to the situation where, say, any realization from is assigned 0 density under , and vice versa. Similar expressions hold for HD(N, DE), where DE(β|0, λ) is the zero-mean double-exponential distribution with parameter λ that is used in Bayes L and for HD(t, DE). To compare these three priors, we took , , and , so that the three priors had the same variance; for the t-distribution we assigned ν = 6, to produce sufficiently thick tails. With these assignments and so that, using numerical integration between −10 and 10, HD(N, t) = 0.0690. Further, HD(N, DE) = 0.122, and
This shows, at least when variances are matched, that these three priors are not too different from each other, so differences in inference would stem from difference in the type and extent of shrinkage effected. However, if priors are not matched, these distances would be expected to increase. Since ridge regression–BLUP, Bayes A, and Bayes L postulate the same sampling model, whenever n ≪ p differences in posterior inferences between these three members of the Bayesian alphabet must be due to the fact that the priors are very different and influential.
To conclude, whole-genome prediction can be useful for providing locally valid predictions of complex traits. However, the additive regression models employed therein should not be taken at face value from an inferential perspective unless an additive model with many 0 coefficients turns out to hold as approximately true (oracle property 1 met), and n ≫> p0, where p0 is the number of nonzero coefficients (oracle property 2 met). If these two conditions are (ever) fulfilled, it may be that the genetic architecture of the very elusive additive QTL (on whose existence the statistical abstraction of marker-assisted inference is based) will be unraveled by statistical means.
The question of the extent to which an additive genetic model is a good representer of complexity is another issue yet to be sorted out. The Bayesian alphabet may expand further on this matter, e.g., Bayes A may grow into Bayes AAA if additive × additive × additive epistasis is included in a model. Additional expansions of the Bayesian alphabet to accommodate epistatic interactions will further exacerbate the inferential problems, because of a vast increase in number of regression coefficients. It is far from obvious how genetic architecture of complex traits can be learned via highly dimensional statistical models.
Acknowledgments
A big note of thanks goes to Christos Dadousis, Christina Lehermeier, Valentin Wimmer, and Chris-Carolin Schön (Technische Universitat Munchen, TUM, Germany) and William G. Hill (University of Edinburgh) for providing a thorough external review of the manuscript. Eduardo Manfredi (Institut National de la Recherche Agronomique, Toulouse, France) is acknowledged for pointing out the article of Duchemin et al. (2012), who detected overparameterization problems of Bayes Cπ. Heather Adams, Juan Manuel González Camacho, Gota Morota, and Francisco Peñagaricano, all from Wisconsin, and Brad Carlin (Minnesota) are thanked for their comments on an earlier draft of this article. The author is indebted to Chiara Sabatti, the Associate Editor handling the review, and to two anonymous reviewers for their constructive criticism leading to a more succinct manuscript, albeit a much less humorous one than the original submission. Work was partially supported by the Wisconsin Agriculture Experiment Station.
Appendix
Bias of BLUP with Respect to Marker Effects
As a toy example, let n = 3 and p = 4. The model includes an intercept plus the effect of 3 markers, and the incidence matrix is
The first column contains the dummy variable for the intercept, and the remaining columns are the genotype codes for the markers at each of three loci. The first observation (row 1 of X) pertains to an individual that is Aa (coded as 0), bb(coded as −1), CC (coded as1), and so on. This matrix has rank 3, and a generalized inverse of X′X is
If the true values of the intercept and of the marker effects are denoted as a, b, c, and d, respectively, the expected value of the maximum-likelihood estimator of the four parameters is
with the expected value of the effect of the third marker being 0 instead of d because of the rank deficiency. Now, we use BLUP with and variance ratio ( is the variance of marker effects) and calculate it (Henderson 1984) as
For this example,
where k = λ3 + 9λ2 + 20λ + 2. Then
After tedious algebra, one arrives at
where
Conditionally on β, all marker effects are estimated with a bias that involves all other markers (and the intercept as well). Since inferences on genetic architecture are primarily based on point estimates (it should be noted that the biased estimator is more precise), it is quite clear that such inferences are not “clean.”
Marker Effects Are Not Identified from a Bayesian Perspective in the n < p Setting
Let a Bayesian linear model consist of location parameters θA and θB (this partition has a different meaning from the one given above), with likelihood p(y|θA, θB). If the conditional posterior density of θB is such that
then θB is not identifiable, meaning that observation of data does not increase knowledge about θB beyond what is conveyed by the conditional prior p(θB|θA) (Dawid 1979; Gelfand and Sahu 1999). For the model in (1), in the n < p situation matrix Xn×p has rank n, and one can reorganize its columns into
where X1 is n × n with rank n, and X2 is n × (p − n), with the vector of marker effects β partitioned accordingly. Changing variables as
produces the inverse transformations β2 = θB and ; because the transformation is linear, the Jacobian does not involve the parameters. Using the new parameterization model (1) can now be written as
implying that the data contain information about θA but not about θB (the latter can represent any marker effect, by construction). Then, irrespective of the joint prior distribution assigned to θA and θB, the posterior is
so
verifying that the p − n marker effects are not likelihood identified. As pointed out by Gelfand and Sahu (1999), this does not mean that there is not Bayesian learning about θB. It means, however, that data “speak” about θA and that what can be said about θB depends on what has been spoken about θA, with the pipe lining of knowledge done through the prior distribution. This can be seen more clearly by writing the posterior of θB as
This representation enables one to see that marginal inferences about individual marker effects are the weighted average of an infinite number of inferences made from the conditional prior p(θB|θA), where the averaging distribution is the posterior of the signal p(θA|y). If θB is any marker effect, say βj, the preceding becomes
In conclusion, for any letter of the alphabet and for any prior distribution adopted, any inference made about genetic architecture always depends on the form of p(βj|X1β1) or, more generally, of p(θB|θA), and these densities depend on the prior adopted, but not on the data. Proper Bayesian learning takes place for X1β1 only.
Inferences in a Linear Model with Unidentified Parameters
In the context of model (1), the likelihood function (assuming known ) is
For the n < p situation, and with β(0) being a solution to the normal equations corresponding to generalized inverse (X′X)−, the (singular) likelihood is expressible as
Letting r = rank(X) and using results from linear model theory (if n ≪ p, then r ≤ n), it follows that
where Q1 and Q2 are partitions of a p × p matrix of rank-preserving elementary operators (Searle 1966); α1 = Lβ is an r × 1 vector of likelihood-identified estimable functions and α2 = Hβ is a (p − r) × 1 vector of pseudoparameters; K1 = XQ1 and K2 = XQ2 are incidence matrices, with K2 = 0 (α2 is a pseudoparameter, because it is effectively wiped out of the model). The genetic signal is given by K1α1 but we include α2 as well, to see what Bayesian inference does for something on which the data lack information.
If β is assigned the normal prior N(β|0, Vβ),
| (A.1) |
The model is now y = K1α1 + K2α2 + e, and the likelihood under the new parameterization becomes
| (A.2) |
| (A.3) |
Expression (A.3) indicates that at most, r parameters are likelihood identified, but (A.2) is retained to illustrate what the prior does. It is well known (e.g., Gianola and Fernando 1986; Sorensen and Gianola 2002) that combining (A.1) with (A.2) leads to the posterior distribution
| (A.4) |
where
since K2 = 0, and where
The p-dimensional distribution (A.4) is nonsingular, but it is based on a likelihood that is defined in r dimensions only! Note that the posterior mean satisfies
| (A.5) |
The coefficient matrix in (A.5) is the counterpart of X′X and it is proportional to the negative matrix of second derivatives) of the log-posterior with respect to α1 and α2. This shows that proper Bayesian learning takes place only for α1, as the information about α2 and the co-information about α1 and α2 come from the prior only. Further, note the relationship
| (A.6) |
indicating that what is learned about α2 is solely a function of what is learned about α1. This is verified by inserting relationship (A.6) in Equations A.5 above, giving
Using properties of inverses of partitioned matrices, so that
| (A.7) |
The preceding confirms that the data inform about α1 but not about α2; what is learned about the latter from phenotypes is done indirectly, through α1. Such an “indirect” inference parallels the concept of “prediction of breeding values of individuals without phenotypes” (Henderson 1977). In the molecular markers setting, n linear combinations of markers are learned from the data, but p − n remain at the mercy of the prior. In other words, one does not clearly know what marker effects are being learned from the data, unless the model is parameterized deliberately. This is clearly shown in the first section of the Appendix.
An example of Proper Bayesian Learning
To illustrate a case of proper Bayesian learning where a connection with genomic BLUP arises, consider inferring the signal g = Xβ. This is likelihood identified (estimable) because E(y|Xβ) = g, and the likelihood is
with a maximum at . The information matrix is
meaning that, for each individual signal, the information content is proportional to what is conveyed by a sample of size n = 1 (if the response variates are means of preprocessed data, the information content will be higher). For the prior N(β|0, Vβ), the resulting prior for the signal is g|Vβ ∼ N(0, XVβX′) and standard results for Bayesian inference give as posterior distribution, where
and
A special case is when so that for being the variance ratio, and Using well-established results known from prediction of random variables dating back to Henderson (1977) but rediscovered recently (e.g., Janss et al. 2012) one can easily find the posterior distribution of β from that of g, and vice versa . Here, take α1 = Xβ so that and which is known as genomic BLUP. Any marker effect can be learned indirectly from using standard BLUP theory as
Mode of the Conditional Posterior Distribution in Bayes A
Taking logs of (7) yields
| (A.8) |
The gradient vector is
| (A.9) |
where and
Setting to zero, to satisfy the first-order condition, leads to
This system is not explicit in β (because marker effects appear nonlinearly in Wβ) but a functional iteration can be developed to locate stationary points.
Mode of the Conditional Posterior Distribution in Bayes L
As a side note, consider what happens if it is not ignored that is a random matrix, contrary to what was done by Tibshirani (1996) in a modal representation of Bayes L. Recalling that and that ,
Differentiating (11) with respect to β
where sβ is a vector containing the signs of the elements of β. Here, the first-order condition would lead to the iteration
Approximation of an Integral in Bayes L
The integral in (16) can be approximated using a second-order expansion around such that (ignoring the subscript in βj)
Use of this in (16) produces
Note that is the kernel of a distribution, so that
and
Footnotes
Communicating editor: C. Sabatti
Literature Cited
- Barber, D., 2012 Bayesian Reasoning and Machine Learning Cambridge University Press, Cambridge, UK. [Google Scholar]
- Bernardo, J. M., and A. F. M. Smith, 1994 Bayesian Theory Wiley, Chichester, UK. [Google Scholar]
- Bishop, C. M., 2006 Pattern Recognition and Machine Learning Springer, New York. [Google Scholar]
- Brondum R. F., Su G., Lund M. S., Bowman P. J., Goddard M. E., et al. , 2012. Genome specific priors for genomic prediction. BMC Genomics 10.1186/1471-2164-13-543 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carlin, B. P., and T. A. Louis, 1996 Bayes and Empirical Bayes Methods for Data Analysis Chapman & Hall, London. [Google Scholar]
- Celeux G., Hurn M., Robert C., 2000. Computational and inferential difficulties with mixture posterior distributions. J. Am. Stat. Assoc. 95: 957–979 [Google Scholar]
- Crossa J., de los Campos G., Pérez P., Gianola D., Burgueño J., et al. , 2010. Prediction of genetic value of quantitative traits in plant breeding using pedigree and molecular markers. Genetics 186: 713–724 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dawid A. P., 1979. Conditional independence in statistical theory (with discussion). J. R. Stat. Soc. B 41: 1–31 [Google Scholar]
- Dehmer, M., and S. C. Basak, 2012 Statistical and Machine Learning Approaches for Network Analysis Wiley, Hoboken, NJ. [Google Scholar]
- de los Campos G., Gianola D., Rosa G. J. M., 2009a Reproducing kernel Hilbert spaces regression: a general framework for genetic evaluation. J. Anim. Sci. 87: 1883–1887 [DOI] [PubMed] [Google Scholar]
- de los Campos G., Naya H., Gianola D., Crossa J., Legarra A., et al. , 2009b Predicting quantitative traits with regression models for dense molecular markers and pedigrees. Genetics 182: 375–385 [DOI] [PMC free article] [PubMed] [Google Scholar]
- de los Campos G., Gianola D., Allison D. B., 2010. Predicting genetic predisposition in humans: the promise of whole-genome markers. Nat. Rev. Genet. 11: 880–886 [DOI] [PubMed] [Google Scholar]
- de los Campos G., Hickey J. M., Pong-Wong R., Daetwyler H. D., Calus M. P. L., 2012a Whole genome regression and prediction methods applied to plant an animal breeding. Genetics 193: 327-345 [DOI] [PMC free article] [PubMed] [Google Scholar]
- de los Campos G., Klimentidis Y. C., Vaźquez A. I., Allison D. B., 2012b Prediction of expected years of life using whole-genome markers. PLoS ONE 7: 1–7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duchemin S. I., Colombani C., Legarra A., Baloche G., Larroque H., et al. , 2012. Genomic selection in the French Lacaune dairy sheep breed. J. Dairy Sci. 95: 2723–2733 [DOI] [PubMed] [Google Scholar]
- Erbe M., Hayes B. J., Matukumali L. K., Goswami S., Bowman P. J., et al. , 2012. Improving accuracy of genomic predictions within and between dairy cattle breeds with imputed high-density single nucleotide polymorphism panels. J. Dairy Sci. 95: 4114–4129 [DOI] [PubMed] [Google Scholar]
- Falconer, D. S., and T. F. C. Mackay, 1996 Introduction to Quantitative Genetics, Ed. 4. Longmans Green, Harlow, UK. [Google Scholar]
- Fan J., Li R., 2001. Variable selection via nonconcave penalized likelihood and its oracle properties. J. Am. Stat. Assoc. 96: 1348–1360 [Google Scholar]
- Geisser, S., 1993 Predictive Inference: An Introduction Chapman & Hall, New York. [Google Scholar]
- Gelfand A. E., Sahu S. K., 1999. Identifiability, improper priors, and Gibbs sampling for generalized linear models. J. Am. Stat. Assoc. 94: 247–253 [Google Scholar]
- Gianola D., Fernando R. L., 1986. Bayesian methods in animal breeding theory. J. Anim. Sci. 63: 217–244 [Google Scholar]
- Gianola D., Heringstad B., Ødegård J., 2006. On the quantitative genetics of mixture characters. Genetics 173: 2247–2255 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gianola D., de los Campos G., Hill W. G., Manfredi E., Fernando R. L., 2009. Additive genetic variability and the Bayesian alphabet. Genetics 183: 347–363 [DOI] [PMC free article] [PubMed] [Google Scholar]
- González-Camacho J. M., de los Campos G., Pérez P., Gianola D., Cairns J. E., et al. , 2012. Genome-enabled prediction of genetic values using radial basis function neural networks. Theor. Appl. Genet. 125: 759–771 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Habier, D., R. L. Fernando, K. Kizilkaya, and D. J. Garrick, 2011 Extension of the Bayesian alphabet for genomic selection. BMC Bioinformatics. Available at: http://www.biomedcentral.com/1471–2105/12/186 [DOI] [PMC free article] [PubMed]
- Hastie, T., R. Tibshirani, and J. Friedman, 2009 The Elements of Statistical Learning, Ed. 2. Springer, New York. [Google Scholar]
- Heffner E. L., Sorrells M. E., Jannink J. L., 2009. Genomic selection for crop improvement. Crop Sci. 49: 1–12 [Google Scholar]
- Henderson C. R., 1977. Best linear unbiased prediction of breeding values not in the model for records. J. Dairy Sci. 60: 783–787 [Google Scholar]
- Henderson, C. R., 1984 Applications of Linear Models in Animal Breeding University of Guelph, Ontario, Canada. [Google Scholar]
- Heslot N., Sorrells M. E., Jannink J. L., Yang H. P., 2012. Genomic selection in plant breeding: a comparison of models. Crop Sci. 52: 146–160 [Google Scholar]
- Hill W. G., 2012. Quantitative genetics in the genomics era. Curr. Genomics 13: 196–206 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Janss L., de los Campos G., Sheehan N., Sorensen D., 2012. Inferences from genomic models in stratified populations. Genetics 92: 693–704 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jia Y., Jannink J.-L., 2012. Multiple trait genomic selection methods increase genetic value prediction accuracy. Genetics 192: 1513–1522 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kärkkäinen H. P., Sillanpää M. K., 2012. Back to basis for Bayesian model building in genomic selection. Genetics 191: 969–987 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Legarra A., Robert-Granié C., Croiseau P., Guillaume F., Fritz S., 2011. Improved Lasso for genomic selection. Genet. Res. 93: 77–87 [DOI] [PubMed] [Google Scholar]
- Lehermeier C., Wimmer V., Albrecht T., Auinger H., Gianola D., et al. , 2013. Sensitivity to prior specification in Bayesian genome-based prediction models. Stat. Appl. Genet. Mol. Biol. DOI: 10.1515/sagmb-2012-0042 [DOI] [PubMed] [Google Scholar]
- Lorenz A. J., Chao S., Asoro F. G., Heffner E. L., Hayashi T., et al. , 2011. Genomic selection in plant breeding: knowledge and prospects. Adv. Agron. 110: 77–123 [Google Scholar]
- Makowsky R., Pajewski N. M., Klimentidis Y. C., Vázquez A. I., Duarte C. W., et al. , 2011. Beyond missing heritability: prediction of complex traits. PLoS Genet. 7(4): 10.1371/journal.pgen.100205 [DOI] [PMC free article] [PubMed] [Google Scholar]
- McLachlan, G., and T. Krishnan, 1997 The EM Algorithm and Extensions Wiley, New York. [Google Scholar]
- McLachlan, G., and D. Peel, 2000 Finite Mixture Models Wiley, New York. [Google Scholar]
- Meuwissen T. H. E., Hayes B. J., Goddard M. E., 2001. Prediction of total genetic value using genome-wide dense marker maps. Genetics 157: 1819–1829 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meuwissen T. H. E., Solberg T. R., Shepherd R., Woolliams J. A., 2009. A fast algorithm for BayesB type of prediction of genome-wide estimates of genetic value. Genet. Sel. Evol. 41: (2) 1–10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mrode, R., 2005 Linear Models for the Prediction of Animal Breeding Values Ed. 2. CABI, Wallingford, UK. [Google Scholar]
- Mutshinda C.M., and M. J. Sillanpää, 2010. Extended Bayesian LASSO for multiple quantitative trait loci mapping and unobserved phenotype prediction. Genetics 86: 1067–1075 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ober U., Ayroles J. F., Stone E. A., Richards S., Zhu D., et al. , 2012. Using whole-genome sequence data to predict quantitative trait phenotypes in Drosophila melanogaster. PLoS Genet. 8(5): e1002685 10.1371/journal.pgen.1002685 [DOI] [PMC free article] [PubMed] [Google Scholar]
- O’Hagan, A., 1994 The Advanced Theory of Statistics: Vol. 2B. Bayesian Inference Arnold, Cambridge, UK. [Google Scholar]
- Park T., Casella G., 2008. The Bayesian Lasso. J. Am. Stat. Assoc. 103: 681–686 [Google Scholar]
- Pérez P., de los Campos G., Crossa J., Gianola D., 2010. Genomic-enabled prediction based on molecular markers and pedigree using the Bayesian Linear Regression Package in R. Plant Genome 3: 106–116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robertson A., 1955. Prediction equations in quantitative genetics. Biometrics 11: 95–98 [Google Scholar]
- Robinson G. K., 1991. That BLUP is a good thing: the estimation of random effects. Stat. Sci. 6: 15–32 [Google Scholar]
- Rogers, S., and M. Girolami, 2012 A First Course in Machine Learning CRC Press, Boca Raton, FL. [Google Scholar]
- Roos M., Held L., 2011. Sensitivity analysis in Bayesian generalized linear mixed models for binary data. Bayesian Anal. 6: 259–278 [Google Scholar]
- Ruppert, D., M. P. Wand, and R. J. Carroll, 2003 Semipara metric Regression Cambridge University Press, New York. [Google Scholar]
- Searle, S. R., 1971 Linear models Wiley, New York. [Google Scholar]
- Searle, S. R., 1966 Matrix Algebra for the Statistical Sciences Wiley, New York. [Google Scholar]
- Sillanpäa, M., 2012 Bayesian Lasso-Related Methods for Genomic Prediction Methods for Genomic Predictions and QTL Analysis Using SNP Data, p. 20. Eucarpia, Programme, Information, Abstracts, T4, Hohenheim University, Stuttgart, Germany. [Google Scholar]
- Sorensen, D., and D. Gianola, 2002 Likelihood, Bayesian, and MCMC Methods in Quantitative Genetics Springer, New York. [Google Scholar]
- Sorensen D., Waagepetersen R., 2003. Normal linear models with genetically structured residual variance heterogeneity: a case study. Genet. Res. 82: 207–222 [DOI] [PubMed] [Google Scholar]
- Sun X., Qu L., Garrick D. J., Dekkers J. C. M., Fernando R. L., 2012. A fast EM algorithm for Bayes A-like prediction of genomic breeding values. PLoS ONE 7(11): 1–9. e49157 10.1371/journal.pone.0049157 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tibshirani R., 1996. Regression shrinkage and selection via the Lasso. J. R. Stat. Soc. Ser. A Stat. Soc. 58: 267–288 [Google Scholar]
- Van Raden P. M., 2008. Efficient methods to compute genomic predictions. J. Dairy Sci. 91: 4414–4423 [DOI] [PubMed] [Google Scholar]
- Vázquez A. I., Rosa G. J. M., Weigel K. A., de los Campos G., Gianola D., et al. , 2010. Predictive ability of subsets of single nucleotide polymorphisms with and without parent average in US Holsteins. J. Dairy Sci. 93: 5942–5949 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vázquez A. I., de los Campos G., Klimentidis Y. C., Rosa G. J. M., Gianola D., et al. , 2012. A comprehensive genetic approach for improving prediction of skin cancer risk in humans. Genetics 192: 1493-1502 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Verbyla K. L., Bowman P. J., Hayes B. J., Goddard M. E., 2009. Sensitivity of genomic selection to using different prior distributions. BMC Proc. 03/2010; 4 (Suppl) 1S5. 1–4 ( 10.1186/1753-6561-4-S1-S5). [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang C.-L., Ding X.-D., Wang J.-Y., Liu J.-F., Fu W.-X., et al. , 2013. Bayesian methods for estimating GEBVs of threshold traits. Heredity 110: 213–219 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wasserman, L., 2010 All of Nonparametric Statistics Springer, New York. [Google Scholar]
- Weigel K. A., de los Campos G., González-Recio O., Naya H., Wu X. L., et al. , 2009. Predictive ability of direct genomic values for lifetime net merit of Holstein sires using selected subsets of single nucleotide polymorphism markers. J. Dairy Sci. 92: 5248–5257 [DOI] [PubMed] [Google Scholar]
- Wellmann R., Bennewitz J., 2012. Bayesian models with dominance effects for genomic evaluation of quantitative traits. Genet. Res. 94: 21–37 [DOI] [PubMed] [Google Scholar]
- Wimmer, V., T. Albrecht, C. Lehermeier, H.-J. Auinger, Y. Wang et al., 2012 Eucarpia: Programme, Information, Abstracts T7, p. 30. Hohenheim University, Stuttgart, Germany. [Google Scholar]