SUMMARY
Splicing of pre-mRNAs in eukaryotes is catalyzed by the spliceosome a large RNA–protein metalloenzyme. The catalytic center of the spliceosome involves a structure comprised of the U2 and U6 snRNAs and includes a metal bound by U6 snRNA. The precise architecture of the splicesome active site however, including the question of whether it includes protein components, remains unresolved. A wealth of evidence places the protein PRP8 at the heart of the spliceosome through assembly and catalysis. Here we provide evidence that the RNase H domain of PRP8 undergoes a conformational switch between the two steps of splicing rationalizing yeast prp8 alleles promoting either the first or second step. We also show that this switch unmasks a metal-binding site involved in the second step. Together these data establish that PRP8 is a metalloprotein that promotes exon ligation within the spliceosome.
The spliceosome is a large RNP particle consisting of the U1, U2, and U4–U5–U6 snRNPs (small nuclear ribonucleoprotein particles) each containing a unique snRNA and associated proteins bound to the pre-mRNA substrate1–3. Spliceosome assembly is a complex process that includes formation of a U2–U6 snRNA structure that has been shown to be essential for catalysis of the transesterifications4. Although pre-mRNA splicing is therefore believed to be intrinsically RNA catalyzed, there is considerable evidence suggesting an intimate interaction between spliceosomal proteins and the active site of the spliceosome. The highly conserved U5 snRNP protein PRP8 has been shown in a crosslinking study to directly contact the 5′ splice site and U6 snRNA during spliceosome assembly5–7. Mutant prp8 alleles in yeast also strongly suggest interactions of this factor with both substrate and snRNA catalytic structures in the spliceosome8–12. Therefore, the catalytic center of the spliceosome may include protein as well as RNA components13.
A large number of mutant prp8 alleles related to spliceosome activation and possibly catalysis have been characterized in yeast, which suggest a central role for this factor in both processes including a proposed conformational change between the two steps of splicing8,12. The clustering of these alleles, along with crosslinking data, suggested the presence of a core domain within PRP8 that could be part of or interact with the spliceosomal catalytic machinery12,13. In order to test these hypotheses, we examined the role of this domain of PRP8 within the spliceosome using a combination of X-ray structure determination and characterization of mutant prp8 alleles in yeast and yeast splicing extracts.
RESULTS
PRP8 rearrangement unmasks a conserved metal binding site
We and others recently solved the X-ray structures of the human and yeast domain IV of PRP814–16 (hereafter referred to as PRP8 RH). This domain is C-terminal to reverse transcriptase-derived and endonuclease-like domains and N-terminal to a Jab-MPN domain within PRP817 (Supplementary Fig. 1a,b). Here, we report our detailed structural analysis of PRP8 RH and a panel of domain mutants highlighting a conformational change that unmasks a Mg2+ binding site that promotes splicing (Fig. 1, Table 1, Supplementary Table 1,2). Our previously described crystals of human PRP8 RH (amino acids 1769–1990) included two monomers in the asymmetric unit. A salient feature was the presence of an N-terminal RNase H domain that included a seventeen amino acid insertion (amino acids 1787–1803) between two adjacent β-strands of the fold. Within the crystal, in what we describe as the closed conformation, this insertion forms a two-stranded antiparallel β-hairpin while in the second, open conformation this hairpin is disrupted to form a displaced loop (Supplementary Note 1). The translation of this loop back ~45° pulls amino acids 1782–1784 to extend the β1 strand of the RNase H fold (Fig. 1a, Supplementary Movie 1). This movement is related to the disruption and rearrangement of elements of the α1 helix and part of the α2 helix (amino acids 1823– 1839). These residues now partially fill the space formerly occupied by the base of the hairpin.
Figure 1.
Conformational switch in the PRP8 RH domain unmasks a Mg2+ binding site. (a) X-ray structure of the PRP8 RH domain. Superposition of the closed (yellow) and open (cyan) conformations observed in the asymmetric unit. (b) 2Fo-Fc maps contoured at 1.0σ showing octahedral coordination of Mg2+ (purple) bound in the open conformation of wild-type and R1865A PRP8 RH. (c) Detail of Mg2+ ion (purple) coordination by Asp1781 and inner-sphere waters (red) in the open conformation of PRP8 RH. (d) Superposition of the X-ray structure of the PRP8 RH domain closed (yellow) and open (cyan) conformations detailing the displacement of Thr1783 to allow Mg2+ coordination. The metal ion bound in the open conformation is not shown for clarity.
Table 1.
Data collection and refinement statistics for PRP8 RH domain structures
| wt 300mM MgCl2 | R1865A 300mM MgCl2 | |
|---|---|---|
| Data collection | ||
| Space group | P212121 | P212121 |
| Cell dimensions | ||
| a, b, c (Å) | 76.03, 78.02, 93.81 | 76.20, 78.04, 94.06 |
| Resolution (Å) | 1.4 | 1.15 |
| R sym | 0.069 (0.525) | 0.038 (0.482) |
| I/σI | 18.1 (2.3) | 26.7 (2.5) |
| Completeness (%) | 99.5 (99.5) | 98.9 (96.6) |
| Redundancy | 4.9 | 4.3 |
| Refinement | ||
| Resolution (Å) | 60-1.40 | 60-1.15 |
| No. reflections | 109740 | 186162 |
| Rwork/ Rfree | 0.136/0.177 | 0.140/0.160 |
| No. atoms | ||
| Protein | 3757 | 3884 |
| Ligand/ion | 19 | 26 |
| Water | 709 | 735 |
| B-factors | ||
| Protein | 21.5 | 23.4 |
| Ligand/ion | 28.2 | 27.4 |
| Water | 34.6 | 35.9 |
| R.m.s deviations | ||
| Bond lengths (Å) | 0.011 | 0.012 |
| Bond angles (°) | 1.44 | 1.51 |
Each dataset was collected from a single crystal
Value in parentheses corresponds to the highest resolution shell (10% of reflections)
The presence of an RNase H fold within a domain of PRP8 associated with the individual catalytic steps of splicing is provocative since RNase H is a metalloenzyme promoting chemistry on an RNA substrate. Catalysis of RNA cleavage by RNase H–like enzymes involves a two metal mechanism in which divalent magnesium ions, bound at adjacent sites separated by ~4 Å, promote hydrolysis by activation of a water nucleophile combined with transition state stabilization18. Thus, the presence of the RNase H domain, in a portion of PRP8 associated with the spliceosomal catalytic core, raised the possibility that a protein-bound metal could participate in catalysis of splicing. Inspection of the PRP8 RH structure14–16 showed that one of two canonical RNase H metal-binding sites is present with coordinating side chains—two aspartates (Asp1781 and Asp1782)— spatially conserved with respect to Mg2+ coordinating residues within the RNase H fold; this site is completely conserved in all PRP8 orthologues (Supplementary Fig. 1c). In our initial studies, despite the presence of MgCl2 during crystallization, we did not observe metal ion coordination at this site a result we attribute to the high ionic strength (2.5 M NaCl) of the crystallization conditions14. As well, the side chain of the conserved arginine residue Arg1865 positioned over this site possibly plays a shielding role. In the work reported here, we crystallized and solved the 1.4 Å structure of PRP8 RH and the 1.15 Å structure of the R1865A mutant under low ionic strength but in the presence of divalent magnesium. Again, we observed both closed and open conformations of PRP8 RH in the asymmetric units of the crystals. However, under these conditions we noted coordination of a single magnesium ion at the conserved metal binding site within the open, but not the closed, conformation of both wild-type and R1865A PRP8 confirming this observation by anomalous scattering from Co2+ soaked crystals (Fig. 1b, Supplementary Fig. 2a, Table 1). The electron density suggests that the Mg2+ ion is present at partial occupancy in the wild-type structure but is consistent with full occupancy in the R1865A structure. The overall features of the R1865A crystals and resulting structure are remarkably similar to wild-type. A higher metal occupancy is consistent with a shielding role of the Arg side chain in the wild-type protein which we propose could be relieved by rearrangement associated with RNA binding within the spliceosome.
Coordination of metal ion to the canonical RNase H site of PRP8 includes inner sphere contact with the side chain of Asp1781 and outer sphere coordination via five ordered waters to the carboxylate of Asp1782, carbonyls of Asp1782 and Leu1891, the amide carbonyl of Gln1894, and the hydroxyl of Thr1864 (Fig. 1c). One consequence of the structural rearrangement involved in the closed to open transition is that Asp1782 moves closer to Asp1781 and Thr1783 is displaced ~4 Å upwards (Fig. 1d, Supplementary Movie 1). In the closed conformation it is this positioning of Thr1783 that essentially blocks metal binding at this site; the substantial local reorganization of the PRP8 RH structure is essential to allow Mg2+ coordination. These striking observations led us to investigate the possible roles of both the conformational change and conformation-dependent Mg2+ coordination by PRP8 in the spliceosome.
prp8 suppressor alleles favor closed or open conformations
Query and Konarska have suggested an elegant model whereby PRP8 is involved in the equilibrium between two distinct spliceosomal conformations associated with the first and second transesterification steps19. Two sets of prp8 alleles, designated as first- or second-step, are proposed to act by shifting this equilibrium to favor one step over the other. Thus a first step allele suppresses a defect in the first but enhances a defect in the second step of splicing while a second step allele suppresses a defect in the second but enhances a defect in the first step of splicing. Intriguingly, the largest proportion of first or second step suppressor alleles associated with the PRP8 RH domain map to the two-stranded β-hairpin or rearranged loop14–16 (Supplementary Fig. 1d). This suggests that the disposition of this structure is critical to the function of this domain within the spliceosome. Because of this, we endeavored to determine whether first or second step phenotypes associated with specific alleles could be rationalized by reference to either of the two conformations observed crystallographically.
We first generated mutant yeast strains to characterize the effect on splicing of a series of mutations within the PRP8 RH β-hairpin comparing these to both wild-type and known strong first (E1960K) and second (V1870N; Supplementary Fig. 1d, Supplementary Table 2) step alleles. We assayed growth using a copper-resistance reporter system in which the branch site adenosine of the ACT1-CUP1 intron was replaced with guanosine (BSG), a change that inhibits the second step of splicing20. Three mutant alleles (E1960K, V1860D, T1872E) were less resistant to copper consistent with a first step phenotype while three (V1870N, T1861P, V1862Y) were more resistant to copper consistent with a second step phenotype (Fig. 2a). We directly assayed steady-state pre-mRNA splicing efficiencies of the reporter gene in these mutant PRP8 backgrounds comparing splicing of the BSG mutant ACT1-CUP1 intron with an RNA containing the branch site cytidine mutation (BSC), known to inhibit both steps of splicing21. Consistent with the results of the growth assay, E1960K, V1860D, and T1872E acted as first step alleles enhancing the first step of splicing in the presence of BSC at the expense of the second step and worsening the effect of the BSG mutation. Conversely, V1870N, T1861P, and V1862Y acted as second step alleles enhancing the second step in splicing of the BSG introns (Fig. 2b, Supplementary Note 2).
Figure 2.
Characterization of PRP8 RH domain alleles. (a) Spot assays showing BSG ACT1-CUP1 reporter-dependent growth of yeast containing wild-type and mutant PRP8 alleles in the presence of the indicated concentration of Cu2+. (b) Denaturing PAGE analysis of RT-primer extension with 32P-labeled primer to examine steady-state splicing efficiencies in PRP8 mutant yeast strains. Primer extension of ACT1-CUP1 RNA with wild-type, BSC, or BSG sequences combined with PRP8 alleles. The products corresponding to the mRNA, pre-mRNA, and intron-lariat intermediate in the gel are indicated (top). Quantification of the first and second step efficiency is shown (bottom). BSG: branch site guanosine; BSC: branch site cytidine.
We next crystallized and solved the X-ray structures of four human PRP8 RH mutants corresponding to the yeast alleles mapping to the β-hairpin under conditions such that the asymmetric unit featured both the closed and open conformations (Fig. 3a,b, Supplementary Fig. 3a,b, Supplementary Table 1). In all cases, a bound metal ion was observed at the canonical RNase H site in the open conformation but was occluded in the closed conformation.
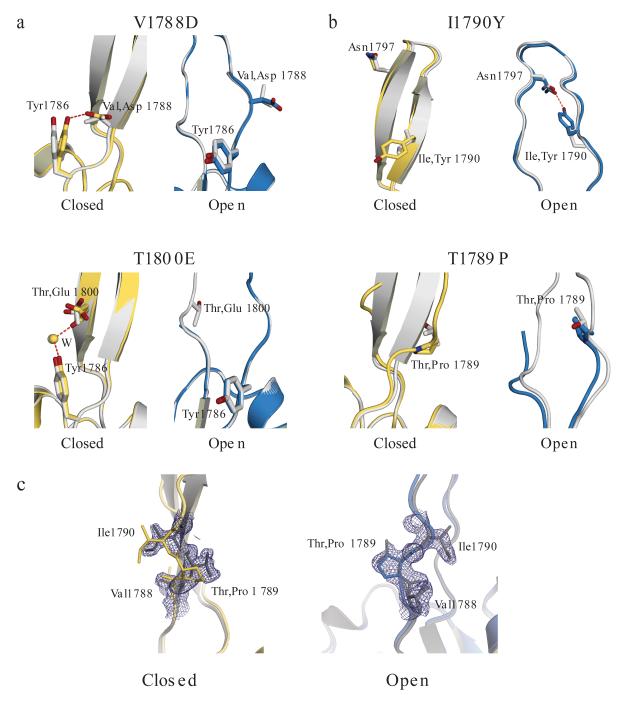
Figure 3.
PRP8 mutant alleles favor distinct conformations within the PRP8 RH domain. Shown are superpositions of X-ray structures of wild-type (gray) with mutant closed (yellow) or mutant open (cyan) conformers. (a) (top) Detail showing formation of an additional hydrogen-bonding interaction to Tyr1786 in the closed conformation of the V1788D structure; (bottom) detail showing formation of a water (W) mediated additional hydrogen-bonding interaction to Tyr1786 in the closed conformation of the T1800E structure. (b) (top) Detail showing formation of a hydrogen bond across the loop with Asn1797 in the open conformation of the I1790Y structure; (bottom) detail showing disruption of the β-hairpin of the closed conformation in the T1789P structure. (c) X-ray structural analysis from a crystallographic mixing experiment with T1789P-R1865A and R1865A proteins. Shown are models for structures of the R1865A (gray) and T1789P-R1865A protein for the closed (yellow, left panel) and open (cyan, right panel) conformation. A 2Fo-Fc map contoured at 1σ calculated from the dataset of the T1789P-R1865A + R1865A mixture crystal is shown in dark blue.
We solved the structure of human PRP8 RH mutants representing the V1788D PRP8 (yeast V1860D) and T1800E (yeast T1872E) first-step alleles; these revealed that both mutations introduce an additional hydrogen-bonding interaction within the closed conformation. Replacement of the Val1788 side chain with an aspartate results in formation of a hydrogen bond to Y1786 at the base of the β-hairpin while mutation of the beta-branched side chain of Thr1800 to a glutamate allows sufficient freedom for the backbone peptide to rotate and form a water-mediated hydrogen bond also to Y1786 (Fig. 3a). These extra hydrogen bonds effectively anchor the β-strand in position stabilizing the closed conformation.
We also determined the structures of two mutants I1790Y (yeast V1862Y) and T1789P (yeast T1861P) shown to be second step suppressors based on splicing of the ACT1-CUP1 reporters in yeast. We solved the 1.55 Å structure of the human I1790Y PRP8 RH mutant and observed the formation of a novel hydrogen-bonding interaction between Y1790 and N1797 in the open conformation in which the aromatic side chain of Y1790 spans the displaced loop (Fig. 3b, top). In the closed conformation, the side chains of N1797 and Y1790 project from opposite faces in the β-hairpin region. Therefore, the I-to-Y mutation appears to stabilize the open conformation relative to the closed.
We crystallized and solved the 1.65 Å structure of the human T1789P mutant and the 1.95 Å structure of the T1789P-R1865A double mutant (Supplementary Note 3). The T1789P mutation lies in the β-hairpin and, as expected, this structure is disrupted in the closed but with no discernible effect on the open conformation (Fig. 3b, bottom). We performed a crystallographic mixing experiment where equal concentrations of R1865A and the T1789P-R1865A double mutant were mixed in the crystallization conditions (Supplementary Note 3). The 1.8 Å structure of the resulting crystal shows that only wild-type protein which contains a threonine at position 1789 is visible in the β-hairpin closed conformation. In contrast, within the loop of the open conformation a mixture of threonine and proline is observed at this position (Fig. 3c). This suggests that the T1861P mutation favors the open conformation by destabilizing the closed conformation.
The results of these structural studies complement and rationalize the analysis of mutant prp8 alleles in yeast. They suggest that two first step alleles (V1860D, T1872E) result in the stabilization of the closed conformation of PRP8 RH and imply that this conformation is associated with the first step of splicing. The second step allele V1862Y stabilizes the open conformation while T1861P appears to destabilize the closed and thus favor the open conformation. The structure of the second step allele L1798N (yeast V1870N) can also be modeled to suggest a hydrogen-bonding interaction stabilizing the open conformation (Supplementary Note 4, Supplementary Fig. 3c). These observations argue that the open and metal-binding conformation of PRP8 RH is associated with the second step of splicing.
PRP8 metal binding is coupled to the second step of splicing
In order to examine the role for metal-binding by PRP8 in the context of the spliceosome, we created a panel of five yeast mutants modifying the identity of the inner-sphere ligand at position 1853 (Fig 4a). These are all (with the exception of D1853E) predicted to impair to varying degrees magnesium coordination in the open conformation (Supplementary Fig. 2b, Supplementary Note 5). Moderate effects on growth were observed for a number of these mutants although both D1853S and D1853N exhibited a strong temperature-sensitive phenotype at 37°C. The D1853C mutant was impaired in growth at both 16°C and 30°C and did not grow at 37°C. Mutation of Asp1853 to a hydrophobic residue (leucine or alanine) which would be incapable of either directly or indirectly coordinating Mg2+ or to a positively charged residue (lysine) is lethal (Supplementary Note 6).
Figure 4.
PRP8 Asp1853 mutations selectively impair the second step of splicing. (a) Phenotypic analysis of PRP8 Asp1853 mutations. Serial dilutions of cells from strains harboring the indicated PRP8 alleles were spotted onto YPD medium; plates were incubated for 4 days at 16°C, 2 days at 30°C, or 3 days at 37°C and photographed. (b) Denaturing PAGE analysis of in vitro splicing of 32P-labeled actin pre-mRNA in extracts prepared from strains harboring the indicated Asp1853 PRP8 mutants (top). The positions of the mRNA, pre-mRNA, intron-lariat, and exon-lariat intermediate in the gel are indicated. Quantification of first and second step efficiencies is shown (bottom). (c) D1853C rescue by the second-step allele V1862Y. Growth of a dilution series of the D1853C-V1862Y yeast strain at 16°C, 30°C, and 37°C. Compare to growth of wild-type and D1853C strains in panel a. (d) Denaturing PAGE analysis comparing in vitro splicing of 32P-labeled actin pre-mRNA substrate in extracts prepared from wild-type, D1853C, and D1853C-V1862Y yeast strains.
We prepared splicing extracts derived from all of the D1853 mutant strains and examined their splicing activity in vitro using an actin pre-mRNA substrate (Fig. 4b). Overall, we observe a general trend where residues which should impair Mg2+ ion binding impair splicing substantially with the second step being more strongly affected than the first. We confirmed that mutation of D1853 results in a specific second step defect by preparing extracts from pseudo-diploid strains and specifically immunoprecipitating epitope-tagged mutant PRP8 containing spliceosomes from splicing reactions (Supplementary Note 7, Supplementary Fig. 4a,b,c). In all of these studies, the efficiency of the second step of splicing corresponds with the ability of an amino acid at position 1853 to coordinate or correctly position a Mg2+ ion at the canonical site (Asp and Glu>His and Asn>Ser>>Cys).
Because D1853C was the most impaired viable mutant tested, we further examined the relationship of this mutation to the second step conformation. We created a double mutant based on the V1862Y second step allele (D1853C-V1862Y) and examined growth and splicing activity of extracts prepared from the mutant. Interestingly, V1862Y rescues the severe growth defect at 37°C. As well and predicted for a second step allele, V1862Y partially rescues the second step splicing defect of D1853C in an in vitro assay with actin pre-mRNA (Fig. 4c,d).
One difficulty with mechanistic investigation of the second step of splicing is that a rate-determining conformational change may mask effects on the chemical step of splicing. An elegant approach to more directly observe the second step of splicing is the separation of the two steps in a bimolecular exon ligation reaction first developed by Anderson and Moore22,23. A pre-mRNA lacking a 3′ splice site undergoes the first but not the second step of splicing; subsequent addition of a 3′ splice site containing oligonucleotide in trans results in a bimolecular ligation effectively separating the second chemical step from the first (Fig. 5a). This approach has been used successfully to investigate the effect, otherwise masked by a presumptive conformational change in the spliceosome, of substrate modifications on second step splicing chemistry24.
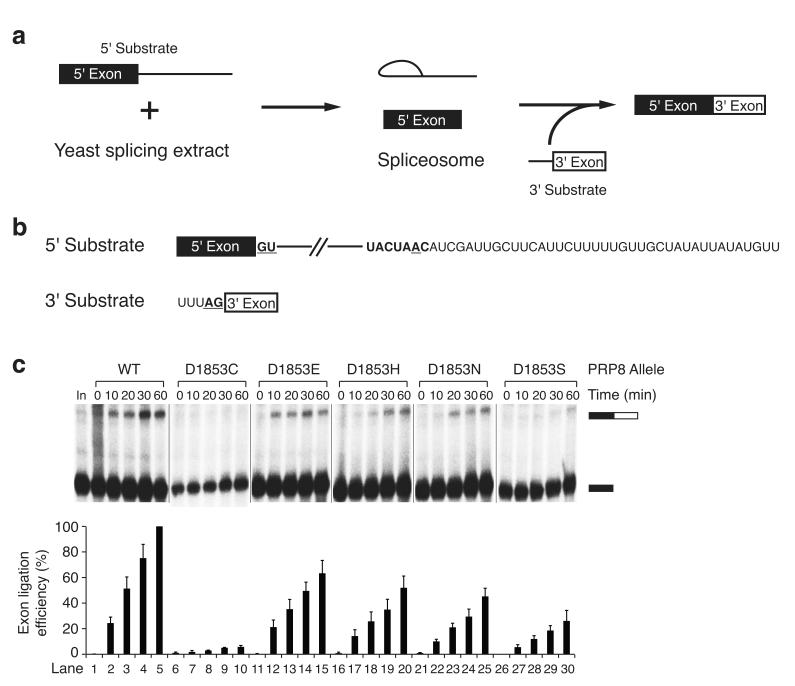
Figure 5.
Bimolecular exon ligation assay implicates PRP8 Asp 1853 in the second transesterification step of pre-mRNA splicing. (a) Schematic representation of bimolecular exon ligation reaction. (b) RNA substrates used in the yeast bimolecular exon ligation. (c) Bimolecular exon ligation assay for analysis of the effect of Asp1853 mutations on the second step of splicing. A 32P-labeled 5′ substrate RNA derived from ACT1 pre-mRNA was incubated under splicing conditions in wild-type and PRP8 Asp1853 mutant yeast extracts, chased with unlabeled RNA containing the 3′ splice site, and the products were analyzed by denaturing PAGE (top; Supplementary Fig. 5c). The positions of the 5′ exon and ligated product are indicated. Quantification of exon ligation efficiency is shown (bottom). Exon ligation efficiency was corrected for the contribution from a degradation product present in the input lane and was normalized to lane 5. Error bars indicate the standard deviation from three independent experiments.
We established a bimolecular exon ligation reaction in yeast extracts in order to directly observe the effects of Asp1853 PRP8 mutants on the second step of splicing (Fig. 5a, Supplementary Fig. 6, Supplementary Note 8). We first synthesized a splicing substrate derived from the ACT1 pre-mRNA containing the 5′ exon, 5′ splice site, branch region and 40 downstream nucleotides but lacking a 3′ splice site. As well, we synthesized a 25 nucleotide 3′ RNA substrate that included the last 5 nucleotides of the intron (UUUAG) representing the 3′ splice site followed by 20 nucleotides of 3′ exon sequence (Fig. 5b). Upon incubation of the 5′ substrate in wild-type extract, we observed an efficient first step of splicing but no second step. Chasing this reaction with the 3′ substrate resulted in bimolecular exon ligation. We then examined the splicing activity of extracts prepared from the panel of Asp1853 mutants using this assay. All of these mutants carried out the first step of splicing with modest impairment corresponding to that observed for the full length ACT1 substrate splicing assay (Supplementary Fig. 5a,b). An impairment of the exon ligation step matching the second step defect observed with the full length splicing substrate was also observed (Fig. 5c, Supplementary Fig. 5c). Strikingly, the efficiency of bimolecular exon ligation in extracts prepared from the D1853C mutant was ~3% of that observed in wild-type extract. This represents an order of magnitude further impairment with respect to the efficiency of exon ligation observed with the intact pre-mRNA. This observation argues that the bimolecular exon ligation has revealed an effect of the D1853C mutation on a rate-limiting step and that this step involves the second transesterification reaction of splicing.
DISCUSSION
In the studies described here, we have observed crystallographically two distinct conformations of the PRP8 RH domain one of which permits coordination of Mg2+ at a canonical and conserved metal-binding site (Fig. 6a). The structures of proteins corresponding to known suppressor alleles argue both that the closed conformation of PRP8 corresponds to a first-step state and that the metal-binding open conformation corresponds to a second-step state. This is consistent with a model whereby PRP8 manifests conformations favoring or facilitating either the first or second transesterification steps of splicing (Fig. 6b).
Figure 6.
PRP8 RH undergoes a conformational switch to present a functionally important metal ion to the spliceosome. (a) Structures of closed and open conformations of PRP8 RH. Residues corresponding to first and second step alleles used in this analysis are highlighted in blue, Mg2+ coordinating ligands in red, Mg2+ is purple, Thr1783 in yellow, and the peptide that crosslinks to the 5′ splice site in green5. (b) Distinct conformational states of PRP8 are associated with the first and second transesterification steps of splicing19. (c) Modeling of the PRP8 RH domain closed (yellow) and open (blue) conformations on the overall PRP8 structure (PDB ID 4I43) shown in gray. The Aar2 assembly factor structure has been removed for clarity. (d) The switch from closed (yellow) to open (blue) conformation positions a Mg2+ ion that promotes the second transesterification reaction in the active site of the spliceosome. PRP8 RH domain closed and open structures are modeled on the overall PRP8 structure shown in gray17. The Mg2+ ion is highlighted in purple. Ends of a disordered loop that crosslinks to the branch region of the pre-mRNA before the second step17 are shown in green. (e) Aar2 sequesters the spliceosome active site cavity. The U5 snRNP assembly factor Aar2 (cyan) blocks the surface of the PRP8 RH domain (yellow) partially filling the active site cavity (amino acids 320–344, depicted as spheres) and stabilizes the PRP8 RH β-hairpin by extending the β-sheet17 (amino acids 345–353). Mg2+ coordinating residues in RH are shown in red.
There is a rich and complex provenance to the prp8 suppressor alleles designated here and earlier8,15 as first or second step. For example, the V1870N second-step allele was isolated as a suppressor of the branch site adenosine to guanosine mutation but also acts as a suppressor of 5′ and 3′ splice site mutations.8 This is reminiscent of the observations of Collins and Guthrie with respect to a number of other prp8 suppressor alleles25; these authors interpreted their data to suggest a role for PRP8 at the catalytic core of the spliceosome. A number of prp8 alleles relate to the transition from the U4– U6 snRNA base-pairing that precedes formation of the U2–U6 structure within the mature spliceosome. In a screen for suppressors of the effects of hyperstabilization of the U4–U6 duplex, Kuhn and Brow identified alleles of PRP8 containing V1860D, T1861P, or V1862Y that were able to suppress this cs defect10. It was noted by these authors, as well as by Collins and Guthrie that prp8 alleles tend to suppress mutations either of the pre-mRNA splice sites, or within spliceosomal components such as U4 snRNA10,25,26. Although some prp8 alleles can suppress both types of defects, the general trend of segregation into two groups of prp8 alleles in the context of the results reported here suggest that the PRP8 RH domain plays a role in earlier assembly-pre-catalytic steps within the spliceosome which is distinct from the dynamics reported here. Beyond all of this is the identification of first and second step alleles outside of the PRP8 RH domain8,11,12,19,27 which argues for a larger, concerted conformational change extending beyond that described for the PRP8 RH domain here.
The observation that PRP8 RH metal-binding is only observed in the open conformation combined with our structural and functional analyses of first and second step alleles associates this state with the second step of splicing. Mutagenesis of the inner sphere ligand Asp1853 combined with the effects of mutations at this site on splicing establish the importance of divalent metal binding by PRP8 RH for the second transesterification step of splicing.
The recent structural analysis of a large fragment of yeast PRP8 in complex with the U5 snRNP assembly factor Aar2 suggests the presence of a large cavity in PRP8, which on the basis of both mutational and crosslinking data has been proposed to encompass the active site of the spliceosome5,6,12. This cavity is large enough to contain the Group II intron active site RNA structure which has been proposed as a model for the U2–U6 snRNA components of the spliceosome active site28. The PRP8 RH domain, which also contains the PRP8 sequence that crosslinks to the 5′ splice site faces into this cavity directly across from a disordered loop that crosslinks to the pre-mRNA branch region before the second step of splicing17. Thus, the conformational switch from closed to open conformations directly presents a metal ion, shown here to promote the second catalytic step of splicing, to the active site cavity of the spliceosome (Fig. 6a–d). These observations are consistent with a structural or catalytic role for the PRP8 bound metal ion in the spliceosomal active site for the second step of splicing and suggest that at this step the active site could contain both protein and RNA components. Interestingly, the C-terminal region of Aar2 lies across the PRP8 RH surface and stabilizes the β-hairpin structure by extending the β-sheet. This has the effect of blocking the cavity and locking the PRP8 RH domain in an assembly-pre-catalytic step conformation and is reminiscent of the sequestering of the U6 snRNA in the U4–U6 snRNA duplex that precedes formation of the catalytically active U2–U6 structure and precludes metal binding by U6 snRNA 29,30 (Fig. 6e).
The precise role of the bound metal ion within the open conformation of the PRP8 RH domain remains to be determined and will require further structural and functional analysis. This metal could perform a structural role with respect to stabilizing the second-step conformation of this domain, it could play a role in positioning or stabilizing either substrate or active-site snRNA components, or it could play a direct role in catalysis of the second step.
It has long been believed that the spliceosome is in essence a ribozyme based, in part, on the mechanistic similarity between self-splicing group II introns and the processing of nuclear pre-mRNAs31. There is also considerable evidence for the role of the spliceosomal U2–U6 snRNA structure and particularly U6 snRNA in catalysis of both steps of splicing29,30,32. High resolution structural analyses of the group I and group II introns have supported a general two-metal ion mechanism that was proposed for these catalytic RNAs, as well as the spliceosome, based on analysis of phosphoryl transfer by protein enzymes28,33,34. There is nevertheless scope for the involvement of more or fewer metal ions in similar enzymatic transformations. Berger and coworkers have recently proposed a mechanism for type II and IA topoisomerases where the transition state is stabilized by a single bound metal and conserved arginine35. A detailed characterization of the mechanism of DNA polymerase η argues persuasively for the participation of a third metal ion in the stabilization of an intermediate state36 and three active-site metals have been observed in the structures of other enzymes involved in catalysis of phosphoryl transfer37–40. As well, there is compelling functional evidence for the participation of three metal ions at the active site of the Group I intron41. Thus there is the potential for PRP8 to participate as part of a three metal spliceosomal active site that would include two metals bound by an snRNA catalytic structure, evolutionarily derived from a Group II–like intron active site, and a third contributed by the protein17,33,34.
The conformational change described here represents a specific switch mechanism between the two steps of splicing. Further, the fact that this switch is coupled to binding of a functionally important metal ion implicates it as a key regulatory mechanism in promotion of the second chemical step of splicing.
ONLINE METHODS
Protein expression, purification, and crystallization
hPRP8 aa1769–1990 protein was grown and purified as previously described14; crystals were grown at 23°C using the hanging drop vapor diffusion technique. Crystals of native protein were grown by mixing one μL of 10 mg ml–1 protein solution (10 mM Tris, pH 8.0, 0.1 mM EDTA, 5 mM DTT, 0.02% NaN3) with one μL of precipitant (2.5 M NaCl, 100 mM Tris, pH 7.0, 200 mM MgCl2, or 10–14% (w/v) PEG 4000, 100 mM Tris pH 7.5, 300 mM MgCl2). Crystals were transferred into cryoprotectant (25% (w/v) PEG 4000, 16% (v/v) glycerol, and 100 mM Tris pH 7.5) with the indicated concentration of divalent metal as a chloride salt and frozen in liquid nitrogen for data collection14.
Wild-type and mutant PRP16, PRP22 and PRP43 proteins were grown and purified as previously described42,43.
Data collection and processing
Data were collected at beam line 8.3.1 of the Advanced Light Source, Lawrence Berkeley National Laboratory, beamline CMCF-1 at the Canadian Light Source, University of Saskatchewan, Saskatoon, and on an R-Axis rotating Cu anode X-ray source. For the wild-type and R1865A PRP8 RH crystals grown in 300 mM MgCl2, a single wavelength experiment was performed at 0.97949Å at a temperature of 105K. Data were processed and scaled using the HKL package44; Friedel pairs were not merged when anomalous scattering maps were required. Anomalous scattering maps were calculated using the CCP4 program suite45.
Model building and refinement
The structures were solved by molecular replacement using the program REFMAC46 to refine using PDB entry 3ENB as a starting model. Water molecules were built using ARPWARP47. Iterative cycles of refinement in REFMAC against the merged dataset and manual model building using COOT was used to complete and refine the models48. Ramachandran statistics for the final refined models of wild type and R1865A are: wild-type grown in 300 mM MgCl2, 98.4% favored, 0.0% outliers; R1865A grown in 300 mM MgCl2, 98.2% favored, 0.0% outliers. All additional Ramachandran statistics are given in Supplementary Table 1.
Creation of mutant S. cerevisae strains and splicing extracts
Mutant prp8 genes were created by gap repair of plasmid pJU18649 containing a HIS selectable marker and PRP8 gene. Mutant plasmids were transformed into strain JDY8.06 (ura3-52, leu2-3,-112, ade2, his3-A1, trpl-289, prp8::LEU2, pJU169 (PRP8, URA3, CEN, ARS) gift from J. Beggs, University of Edinburgh, UK), containing wild-type PRP8 on a counter-selectable URA3-marked plasmid50. After transformation with pJU186, cells were selected by growth at 30°C on SDC plates lacking histidine and leucine. Transformants were streaked on media lacking histidine and leucine containing 5-fluoroorotic acid (5-FOA) to select for cells lacking the URA3 plasmid51. Cells from a single colony that survived on 5-FOA plates were grown in media lacking histidine and total DNA was extracted using a DNeasy kit (Qiagen). All mutant PRP8 strains were verified by sequencing. Splicing extracts were prepared as described elsewhere49.
For creation of copper-resistant strains, the ACT1-CUP1 plasmid containing either the wild-type sequence, BSC or BSG mutations was co-transformed along with pJU186 containing wild-type or mutant PRP8 into yJU7511 (MATa, ade2 cup1D::ura3 his3 leu2 lys2 prp8D::LYS2 trp1, pJU169 (PRP8 URA3 CEN ARS)). Transformants were streaked on media lacking histidine and leucine and containing 5-FOA to select for cells lacking the URA3 plasmid. Cells surviving on 5-FOA plates were grown in media lacking histidine and total DNA was extracted using a DNeasy kit (Qiagen). All mutant PRP8 strains were verified by sequencing.
For the creation of pseudo-diploid strains, PRP8 was knocked out with a LEU2-marked fragment in yJPS662, knockout confirmed by PCR and viability rescued with PRP8 on URA3-marked plasmid, generating yJPS1481. To obtain pseudo-diploid strains, yJPS1481 was transformed with wild-type or mutant PRP8 on a HIS3 plasmid and transformants that retained both the wild-type URA3 and the HIS3 PRP8 plasmids were selected for on URA-HIS drop-out media. Mutant PRP8 plasmids were generated from the wild-type plasmid pJU204 (PRP8-HA, HIS3, gift from C. Guthrie) by Quick Change PCR and verified by sequencing.
Growth assays
For spot test analysis, yeast strains were inoculated in YPD media (1% yeast extract, 2% peptone, 2% dextrose) and grown over-night at 30°C with shaking. The next day, the OD at 600 nm was determined, and 10 μL serial dilutions from 1 × 106 to 1.6× 103 cells ml–1 were spotted onto YPD plates. Plates were photographed after 2 days at 30°C, 3 days at 37°C and 4 days at 16°C.
For the copper growth assay, cultures containing the BSG ACT1-CUP1 and PRP8 mutant plasmids were grown overnight in SDC–Leu–His medium, diluted to A600=0.2, and spotted onto SDC–Leu–His agar plates containing the indicated concentrations of CuSO452. Plates were photographed after 3 days at 30°C.
Primer extension
RNA was extracted using hot phenol53. Cultures containing the ACT1-CUP1 and PRP8 mutant plasmids were grown overnight in SDC–Leu–His medium, diluted to 5 ml (A600=0.2), and then grown in SDC–Leu–His medium for an additional 6 hours to A600=1.0. The cells were spun down and resuspended in 400 μL of AE buffer with 10% SDS (50 mM NaOAc, 10 mM EDTA, pH 5.0). Hot phenol/AE solution (400 μL) was immediately added and incubated at 65°C for 30 min. After centrifugation at 21,000g, the upper phase was transferred into clean tubes and the hot phenol/AE treatment was repeated. The upper phase was extracted three-four times with phenol/chloroform/isoamyl alcohol and chloroform followed by ethanol precipitation. The pellet was resuspended in 10 μL ddH2O.
Primer extensions were carried out using the YAC6 primer complementary to exon 2 of ACT154. The primer was end-labeled with [γ-32P]-ATP (3000 Ci/mmol, PerkinElmer). Primer extensions were performed using the RevertAidTM H minus first strand cDNA synthesis kit (Fermentas). First, 12 μL of the mixture (1μg total RNA, 2 pmol labeled primer) was heated to 70°C for 10 min and slowly cooled down to 40°C. The reaction was chilled on ice, supplemented to 20 μL (4 μL reaction buffer, 1 U RNase inhibitor, 1 mM dNTP mix, 10 U reverse transcriptase), then incubated for 5 min at 37°C and 55 min at 42°C. The reaction was terminated and the RNA was degraded with 0.5 M NaOH at 70°C. Extension products were extracted with phenol/chloroform/isoamyl alcohol and chloroform, followed by ethanol precipitation. The pellet was resuspended, separated in 7% polyacrylamide 8 M urea gels and visualized by autoradiography.
Construction of RNA substrates
The ACT1 and UBC4 pre-mRNAs were made by in vitro transcription using T7 RNA polymerase and [α-32P]-ATP using a PCR template amplified from the appropriate plasmid. The 25 μL reaction (40 mM Tris pH 8.0, 6 mM MgCl2, 10 mM NaCl, 2 mM spermidine, 0.5 mM CTP, 0.5 mM GTP, 0.5 mM UTP, 60 μM ATP, 1.32 μM [α-32P]-ATP (PerkinElmer), 5 ng DNA template, 10 mM DTT, and 1 μL T7 RNA polymerase) was incubated at 37°C for 4 h. The reaction was then separated on a 7% 19:1 acrylamide: bis-acrylamide 8 M Urea PAGE gel.
The 5′ ACT1 substrate used in the exon ligation assay was synthesized by in vitro transcription to yield the following sequence: 5′–CUUUUAGAUUUUUCACGCUUACUGCUUUUUUCUUCCCAAGAUCGAAAAU UUACUGAAUUAACAAUGGAUUCUGGUAUGUUCUAGCGCUUGCACCAUCCC AUUUAACUGUAAGAAGAAUUGCACGGUCCCAAUUGCUCGAGAGAUUUCUC UUUUACCUUUUUUUACUAUUUUUCACUCUCCCAUAACCUCCUAUAUUGAC UGAUCUGUAAUAACCACGAUAUUAUUGGAAUAAAUAGGGGCUUGAAAUUU GGAAAAAAAAAAAAAACUGAAAUAUUUUCGUGAUAAGUGAUAGUGAUAUU CUUCUUUUAUUUGCUACUGUGUCUCAUGUACUAACAUCGAUUGCUUCAUU CUUUUUGUUGCUAUAUUAUAUGUU–3′
The 3′ ACT1 substrate used in the exon ligation assay was synthesized (IDT) with the following sequence: 5′–UUUAGAGGUUGCUGCUUUGGUUAUU–3′
Splicing assays
Splicing reactions (10 μL) were perfomed as described54 in reactions containing 2 mM ATP, 2.5 mM MgCl2, 3% (w/v) PEG 8000, 60 mM potassium phosphate pH 7.0, 20 mM KC1, 8 mM HEPES, 8% (v/v) glycerol, 80 μM EDTA, 0.2 mM DTT, and 1 nM pre-mRNA. Reactions were quenched with 10 μL of stop solution (1 mg ml–1 proteinase K, 50 mM EDTA, 1% SDS) and then incubated at 65°C for 15 min. Each sample was extracted with phenol/chloroform/isoamyl alcohol, then chloroform, and was then ethanol precipitated. Pellets were resuspended in loading dye, separated on 7% 19:1 acrylamide:bis-acrylamide 8 M Urea PAGE gels, exposed to a phosphor storage screen, and scanned using a phosphorimager (Molecular Dynamics).
For the assay of extracts from pseudo-diploid strains, UBC4 pre-mRNA was pre-incubated for 20 min in 60 uL reactions. Mutant PRP43 protein was added to block release of the excised intron and allow accurate quantification of all splicing species associated with the HA-tagged PRP8. Intermediates associated with PRP8-HA were immunoprecipitated with αHA antibody conjugated to protein-A Sepharose (Sigma) (25 uL slurry in IPP150 (10 mM Tris-HCl pH 8; 150 mM NaCl; 0.01% NP-40 substitute (Fluka)) ). Beads were incubated with the splicing reactions for 90 min at 4°C, washed 3 times with 10 volume equivalents of IPP150, RNA was extracted with phenol/-chloroform-isoamyl alcohol and separated by 15% denaturing PAGE.
For the exon ligation assay, the labeled 5′ portion of the ACT1 pre-mRNA was pre-incubated for 30 min in 10 μL splicing reactions. Exon ligation was initiated by addition of the 3′ portion of the ACT1 RNA substrate to splicing reactions and incubation was then continued for 10, 20, 30 and 60 min. The exon ligation product was amplified by RT-PCR and its identity was confirmed by sequencing. Quantifications were performed using ImageQuant (Molecular Dynamics).
Supplementary Material
ACKNOWLEDGEMENTS
This work was supported by an Operating Grant to AMM from the Canadian Institutes of Health Research (CIHR) and a National Institute of Health (NIH) grant to J.P.S and Joseph Piccirilli (R01GM088656). We would like to thank M. Friis and M. Schultz for helpful advice, J. Beggs (University of Edinburgh, Edinburgh, UK) and C. Guthrie (University of California, San Francisco, San Francisco, California) for providing yeast strains and plasmids, and D. Brow (University of Wisconsin–Madison, Madison, Wisconsin) for providing the PRP8 V1862Y plasmid. Research described in this paper was performed at the Advanced Light Source (Berkeley, CA; supported by the U.S. Department of Energy under Contract No. DE-AC02-05CH11231) and the Canadian Light Source (supported by the Natural Sciences and Engineering Research Council of Canada, the National Research Council Canada, CIHR, the Province of Saskatchewan, Western Economic Diversification Canada, and the University of Saskatchewan).
Footnotes
Accession codes. Accession codes. Coordinates and structure factors for the PRP8 RH domain X-ray structures have been deposited in the Protein Data Bank under accession codes 4JK7, 4JK8, 4JK9, 4JKA, 4JKB, 4JKC, 4JKD, 4JKE, 4JKF, 4JKG, and 4JKH.
Note: Supplementary information is available in the online version of the paper.
AUTHOR CONTRIBUTIONS MJS, TW, DBR, and AMM designed the study; MJS created mutant yeast strains, crystallized protein, collected X-ray diffraction data, solved the structures, and performed in vivo and in vitro assays including development of the bimolecular exon ligation; TW created mutant PRP8 yeast strains, crystallized protein, collected X-ray diffraction data and carried out in vivo and in vitro assays; KAA purified and crystallized proteins; SF and JPS, independently designed and tested mutants and critically analyzed data; DBR designed, crystallized, and analyzed mutants; PL provided technical expertise and helped design experiments in yeast; MJS, TW, and AMM wrote the manuscript.
COMPETING FINANCIAL INTERESTS The authors declare no competing financial interests.
References
- 1.Jurica MS, Moore MJ. Pre-mRNA splicing: awash in a sea of proteins. Mol. Cell. 2003;12:5–14. doi: 10.1016/s1097-2765(03)00270-3. [DOI] [PubMed] [Google Scholar]
- 2.Rappsilber J, Ryder U, Lamond AI, Mann M. Large-scale proteomic analysis of the human spliceosome. Genome Res. 2002;12:1231–1245. doi: 10.1101/gr.473902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Zhou Z, Licklider LJ, Gygi SP, Reed R. Comprehensive proteomic analysis of the human spliceosome. Nature. 2002;419:182–185. doi: 10.1038/nature01031. [DOI] [PubMed] [Google Scholar]
- 4.Madhani HD, Guthrie C. A novel base-pairing interaction between U2 and U6 snRNAs suggests a mechanism for the catalytic activation of the spliceosome. Cell. 1992;71:803–817. doi: 10.1016/0092-8674(92)90556-r. [DOI] [PubMed] [Google Scholar]
- 5.Reyes JL, Gustafson EH, Luo HR, Moore MJ, Konarska MM. The C-terminal region of hPrp8 interacts with the conserved GU dinucleotide at the 5′ splice site. RNA. 1999;5:167–179. doi: 10.1017/s1355838299981785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Turner IA, Norman CM, Churcher MJ, Newman AJ. Dissection of Prp8 protein defines multiple interactions with crucial RNA sequences in the catalytic core of the spliceosome. RNA. 2006;12:375–386. doi: 10.1261/rna.2229706. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Vidal VP, Verdone L, Mayes AE, Beggs JD. Characterization of U6 snRNA-protein interactions. RNA. 1999;5:1470–1481. doi: 10.1017/s1355838299991355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Liu L, Query CC, Konarska MM. Opposing classes of prp8 alleles modulate the transition between the catalytic steps of pre-mRNA splicing. Nat. Struct. Mol. Biol. 2007;14:519–526. doi: 10.1038/nsmb1240. [DOI] [PubMed] [Google Scholar]
- 9.Kuhn AN, Reichl EM, Brow DA. Distinct domains of splicing factor Prp8 mediate different aspects of spliceosome activation. Proc. Natl. Acad. Sci. USA. 2002;99:9145–9149. doi: 10.1073/pnas.102304299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Kuhn AN, Brow DA. Suppressors of a cold-sensitive mutation in yeast U4 RNA define five domains in the splicing factor Prp8 that influence spliceosome activation. Genetics. 2000;155:1667–1682. doi: 10.1093/genetics/155.4.1667. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Umen JG, Guthrie C. Mutagenesis of the yeast gene PRP8 reveals domains governing the specificity and fidelity of 3′ splice site selection. Genetics. 1996;143:723–739. doi: 10.1093/genetics/143.2.723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Grainger RJ, Beggs JD. Prp8 protein: at the heart of the spliceosome. RNA. 2005;11:533–557. doi: 10.1261/rna.2220705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Abelson J. Is the spliceosome a ribonucleoprotein enzyme? Nat. Struct. Mol. Biol. 2008;15:1235–1237. doi: 10.1038/nsmb1208-1235. [DOI] [PubMed] [Google Scholar]
- 14.Ritchie DB, et al. Structural elucidation of a PRP8 core domain from the heart of the spliceosome. Nat. Struct. Mol. Biol. 2008;15:1199–1205. doi: 10.1038/nsmb.1505. [DOI] [PubMed] [Google Scholar]
- 15.Yang K, Zhang L, Xu T, Heroux A, Zhao R. Crystal structure of the beta-finger domain of Prp8 reveals analogy to ribosomal proteins. Proc. Natl. Acad. Sci. USA. 2008;105:13817–13822. doi: 10.1073/pnas.0805960105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Pena V, Rozov A, Fabrizio P, Luhrmann R, Wahl MC. Structure and function of an RNase H domain at the heart of the spliceosome. EMBO J. 2008;27:2929–2940. doi: 10.1038/emboj.2008.209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Galej WP, Oubridge C, Newman AJ, Nagai K. Crystal structure of Prp8 reveals active site cavity of the spliceosome. Nature. 2013;493:638–643. doi: 10.1038/nature11843. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Nowotny M, Gaidamakov SA, Crouch RJ, Yang W. Crystal structures of RNase H bound to an RNA/DNA hybrid: substrate specificity and metal-dependent catalysis. Cell. 2005;121:1005–1016. doi: 10.1016/j.cell.2005.04.024. [DOI] [PubMed] [Google Scholar]
- 19.Query CC, Konarska MM. Suppression of multiple substrate mutations by spliceosomal prp8 alleles suggests functional correlations with ribosomal ambiguity mutants. Mol. Cell. 2004;14:343–354. doi: 10.1016/s1097-2765(04)00217-5. [DOI] [PubMed] [Google Scholar]
- 20.Fouser LA, Friesen JD. Mutations in a yeast intron demonstrate the importance of specific conserved nucleotides for the two stages of nuclear mRNA splicing. Cell. 1986;45:81–93. doi: 10.1016/0092-8674(86)90540-4. [DOI] [PubMed] [Google Scholar]
- 21.Vijayraghavan U, et al. Mutations in conserved intron sequences affect multiple steps in the yeast splicing pathway, particularly assembly of the spliceosome. EMBO J. 1986;5:1683–1695. doi: 10.1002/j.1460-2075.1986.tb04412.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Anderson K, Moore MJ. Bimolecular exon ligation by the human spliceosome. Science. 1997;276:1712–1716. doi: 10.1126/science.276.5319.1712. [DOI] [PubMed] [Google Scholar]
- 23.Anderson K, Moore MJ. Bimolecular exon ligation by the human spliceosome bypasses early 3′ splice site AG recognition and requires NTP hydrolysis. RNA. 2000;6:16–25. doi: 10.1017/s1355838200001862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Gordon PM, Sontheimer EJ, Piccirilli JA. Metal ion catalysis during the exon-ligation step of nuclear pre-mRNA splicing: extending the parallels between the spliceosome and group II introns. RNA. 2000;6:199–205. doi: 10.1017/s1355838200992069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Collins CA, Guthrie C. Allele-specific genetic interactions between Prp8 and RNA active site residues suggest a function for Prp8 at the catalytic core of the spliceosome. Genes Dev. 1999;13:1970–1982. doi: 10.1101/gad.13.15.1970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kuhn AN, Li Z, Brow DA. Splicing factor Prp8 governs U4/U6 RNA unwinding during activation of the spliceosome. Mol. Cell. 1999;3:65–75. doi: 10.1016/s1097-2765(00)80175-6. [DOI] [PubMed] [Google Scholar]
- 27.Siatecka M, Reyes JL, Konarska MM. Functional interactions of Prp8 with both splice sites at the spliceosomal catalytic center. Genes Dev. 1999;13:1983–1993. doi: 10.1101/gad.13.15.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Adams PL, Stahley MR, Kosek AB, Wang J, Strobel SA. Crystal structure of a self-splicing group I intron with both exons. Nature. 2004;430:45–50. doi: 10.1038/nature02642. [DOI] [PubMed] [Google Scholar]
- 29.Yean SL, Wuenschell G, Termini J, Lin RJ. Metal-ion coordination by U6 small nuclear RNA contributes to catalysis in the spliceosome. Nature. 2000;408:881–884. doi: 10.1038/35048617. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Koodathingal P, Novak T, Piccirilli JA, Staley JP. The DEAH box ATPases Prp16 and Prp43 cooperate to proofread 5′ splice site cleavage during pre-mRNA splicing. Mol. Cell. 2010;39:385–395. doi: 10.1016/j.molcel.2010.07.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sontheimer EJ, Sun S, Piccirilli JA. Metal ion catalysis during splicing of premessenger RNA. Nature. 1997;388:801–805. doi: 10.1038/42068. [DOI] [PubMed] [Google Scholar]
- 32.Sashital DG, Cornilescu G, McManus CJ, Brow DA, Butcher SE. U2-U6 RNA folding reveals a group II intron-like domain and a four-helix junction. Nat. Struct. Mol. Biol. 2004;11:1237–1242. doi: 10.1038/nsmb863. [DOI] [PubMed] [Google Scholar]
- 33.Toor N, Keating KS, Taylor SD, Pyle AM. Crystal structure of a self-spliced group II intron. Science. 2008;320:77–82. doi: 10.1126/science.1153803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Steitz TA, Steitz JA. A general two-metal-ion mechanism for catalytic RNA. Proc. Natl. Acad. Sci. USA. 1993;90:6498–6502. doi: 10.1073/pnas.90.14.6498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Schmidt BH, Burgin AB, Deweese JE, Osheroff N, Berger JM. A novel and unified two-metal mechanism for DNA cleavage by type II and IA topoisomerases. Nature. 2010;465:641–644. doi: 10.1038/nature08974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Nakamura T, Zhao Y, Yamagata Y, Hua YJ, Yang W. Watching DNA polymerase eta make a phosphodiester bond. Nature. 2012;487:196–201. doi: 10.1038/nature11181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Le Du MH, et al. Artificial evolution of an enzyme active site: structural studies of three highly active mutants of Escherichia coli alkaline phosphatase. J. Mol. Biol. 2002;316:941–953. doi: 10.1006/jmbi.2001.5384. [DOI] [PubMed] [Google Scholar]
- 38.Kim EE, Wyckoff HW. Reaction mechanism of alkaline phosphatase based on crystal structures. Two-metal ion catalysis. J. Mol. Biol. 1991;218:449–464. doi: 10.1016/0022-2836(91)90724-k. [DOI] [PubMed] [Google Scholar]
- 39.Romier C, Dominguez R, Lahm A, Dahl O, Suck D. Recognition of single-stranded DNA by nuclease P1: high resolution crystal structures of complexes with substrate analogs. Proteins. 1998;32:414–424. [PubMed] [Google Scholar]
- 40.Garcin ED, et al. DNA apurinic-apyrimidinic site binding and excision by endonuclease IV. Nat. Struct. Mol. Biol. 2008;15:515–522. doi: 10.1038/nsmb.1414. [DOI] [PubMed] [Google Scholar]
- 41.Shan S, Yoshida A, Sun S, Piccirilli JA, Herschlag D. Three metal ions at the active site of the Tetrahymena group I ribozyme. Proc. Natl. Acad. Sci. USA. 1999;96:12299–12304. doi: 10.1073/pnas.96.22.12299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Tanaka N, Schwer B. Mutations in PRP43 that uncouple RNA-dependent NTPase activity and pre-mRNA splicing function. Biochemistry. 2006;45:6510–6521. doi: 10.1021/bi052656g. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Mayas RM, Maita H, Semlow DR, Staley JP. Spliceosome discards intermediates via the DEAH box ATPase Prp43p. Proc. Natl. Acad. Sci. USA. 2010;107:10020–10025. doi: 10.1073/pnas.0906022107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Otwinowski ZM, Minor W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 1997;276:307–326. doi: 10.1016/S0076-6879(97)76066-X. [DOI] [PubMed] [Google Scholar]
- 45.Potterton E, Briggs P, Turkenburg M, Dodson E. A graphical user interface to the CCP4 program suite. Acta. Crystallogr. D. Biol. Crystallogr. 2003;59:1131–1137. doi: 10.1107/s0907444903008126. [DOI] [PubMed] [Google Scholar]
- 46.Murshudov GN, Vagin AA, Dodson EJ. Refinement of macromolecular structures by the maximum-likelihood method. Acta. Crystallogr. D. Biol. Crystallogr. 1997;53:240–255. doi: 10.1107/S0907444996012255. [DOI] [PubMed] [Google Scholar]
- 47.Terwilliger TC. Automated main-chain model building by template matching and iterative fragment extension. Acta. Crystallogr. D. Biol. Crystallogr. 2003;59:38–44. doi: 10.1107/S0907444902018036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Emsley P, Cowtan K. Coot: model-building tools for molecular graphics. Acta. Crystallogr. D. Biol. Crystallogr. 2004;60:2126–2132. doi: 10.1107/S0907444904019158. [DOI] [PubMed] [Google Scholar]
- 49.Umen JG, Guthrie C. A novel role for a U5 snRNP protein in 3′ splice site selection. Genes Dev. 1995;9:855–868. doi: 10.1101/gad.9.7.855. [DOI] [PubMed] [Google Scholar]
- 50.Brown JD, Beggs JD. Roles of PRP8 protein in the assembly of splicing complexes. EMBO J. 1992;11:3721–3729. doi: 10.1002/j.1460-2075.1992.tb05457.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Boeke JD, Trueheart J, Natsoulis G, Fink GR. Fluoroorotic acid as a selective agent in yeast molecular genetics. Methods Enzymol. 1987;154:164–175. doi: 10.1016/0076-6879(87)54076-9. [DOI] [PubMed] [Google Scholar]
- 52.Lesser CF, Guthrie C. Mutational analysis of pre-mRNA splicing in Saccharomyces cerevisiae using a sensitive new reporter gene, CUP1. Genetics. 1993;133:851–863. doi: 10.1093/genetics/133.4.851. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Kohrer K, Domdey H. Preparation of high molecular weight RNA. Methods Enzymol. 1991;194:398–415. doi: 10.1016/0076-6879(91)94030-g. [DOI] [PubMed] [Google Scholar]
- 54.Lin RJ, Newman AJ, Cheng SC, Abelson J. Yeast mRNA splicing in vitro. J. Biol. Chem. 1985;260:14780–14792. [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.