Abstract
Methylation of DNA is an epigenetic mechanism that influences patterns of gene expression. DNA methylation marks contribute to adaptive phenotypic variation but are erased during development. The role of DNA methylation in adaptive evolution is therefore unclear. We propose that environmentally-induced DNA methylation causes phenotypic heterogeneity that provides a substrate for selection via forces that act on the epigenetic machinery. For example, selection can alter environmentally-induced methylation of DNA by acting on the molecular mechanisms used for the genomic targeting of DNA methylation. Another possibility is that specific methylation marks that are environmentally-induced, yet non-heritable, could influence preferential survival and lead to consistent methylation of the same genomic regions over time. As methylation of DNA is known to increase the likelihood of cytosine-to-thymine transitions, non-heritable adaptive methylation marks can drive an increased likelihood of mutations targeted to regions that are consistently marked across several generations. Some of these mutations could capture, genetically, the phenotypic advantage of the epigenetic mark. Thereby, selectively favored transitory alterations in the genome invoked by DNA methylation could ultimately become selectable genetic variation through mutation. We provide evidence for these concepts using examples from different taxa, but focus on experimental data on large-scale DNA sequencing that expose between-group genetic variation after bidirectional selection on honeybees, Apis mellifera.
Introduction
The methylation of DNA in eukaryotes is a chemical modification that involves the addition of a methyl group onto the position 5 of a pyrimidine ring on cytosines (5mC), primarily within cytosine–phosphate–guanine (CpG) dinucleotides. DNA methylation can affect structural changes to chromatin by attracting protein complexes that modify the histone scaffolds holding the DNA coil. DNA methylation in promoter regions can induce a tightly packed form of DNA with attached proteins, called heterochromatin, that restrict the access of the transcriptional machinery. The outcome is the silencing of proximal gene expression (Klose and Bird 2006). In species that contain DNA methylation, the mechanism has been functionally linked to development, behavior, and phenotypic plasticity (Day and Sweatt 2010; Feng et al. 2010; Law and Jacobsen 2010; Boyko and Kovalchuk 2011; Lyko and Maleszka 2011).
DNA methylation is present in genomes across taxa and likely pre-dates the divergence of plants and animals. However, the amount and distribution of DNA methylation in the genome varies widely among species. For instance, >70% of CpGs are methylated in humans, whereas this number is ∼18% in Arabidopsis thaliana and <1% in Apis mellifera (honeybee) (Flores and Amdam 2011). Moreover, some species have lost the enzymes necessary for DNA methylation despite possession of complex development, behavior, and expression of phenotypic plasticity. For example, the fruit fly Drosophila melanogaster has no CpG DNA methylation, but its molecular biology is similar enough to that of mammals to model development, behavior, human disease, and nutrition (Beckingham et al. 2005). In addition, the fly shares complex programs such as metamorphosis with the honeybee, in which methylation of CpGs contributes to developmental outcomes (Kucharski et al. 2008). Thus, it appears that quite dramatic changes in interspecific amounts and distributions of DNA methylation are permitted—and, as we will argue, perhaps accommodated—by evolutionary processes.
Methylation of DNA is unique from other epigenetic mechanisms that affect the structure of chromatin because its usage by genomes carries an evolutionary ramification in the form of increased mutability. For example, the rate of C-to-T mutations is 10-fold to 50-fold higher in humans’ methylated cytosines (Duncan and Miller 1980; Bulmer 1986; Britten et al. 1988; Sved and Bird 1990). Genomes with DNA methylation, overall, show a depletion of CpG dinucleotides that reflects the occurrence of mutations induced by DNA methylation in the germline (Flores and Amdam 2011). It is unclear whether such patterns of depletion include adaptive mutations or reflect neutral and tolerated genomic changes.
Here, we propose a four-stage mechanism that may explain how methylation of DNA can play a role in adaptive evolution: (1) environmental exposures contribute to variability in targeting of DNA methylation, (2) targeting that benefits reproduction and survival are perpetuated over generations when environmental exposures remain unchanged, (3) targeted genomic regions experience increased mutability, and (4) mutations can accommodate the phenotype achieved by methylation targeting and make it available to natural selection.
To arrive at this explanatory framework, we begin by discussing the functional roles of DNA methylation at the cellular and organismic levels. We then discuss studies that exemplify how changes in patterns in genomic DNA methylation can occur in response to environmental variability and the degree to which those changes are transferred to offspring. Thereafter, we build support for the evolutionary role of DNA methylation from genome-wide re-sequencing data of honeybees that were subject to 37 generations of bidirectional selection (Page and Fondrk 1995).
DNA methylation: targeting, gene regulatory functions, and programming
The de novo methylation of DNA in eukaryotic genomes is performed by the DNA methyltransferase DNMT3. The maintenance DNA methyltransferase DNMT1 carries out the methylation of the cytosine on the complementary strand subsequent to de novo methylation and during replication of DNA (Law and Jacobsen 2010). Although DNA methylation is predominantly found on cytosines within CpG dinucleotides, it also occurs to a much lesser extent in the context of CHG and CHH sequences (H = A, C, or T) (Chan et al. 2005; Lister et al. 2008, 2009).
Mechanisms targeting DNA methylation
The targeting of DNA methylation both in plants and mammals is controlled by an RNA-directed mechanism that allows different genomic sites to be independently methylated in response to growth and developmental, and environmental cues (Mette et al. 2000; Aravin and Hannon 2008; Kuramochi-Miyagawa et al. 2008; Morris 2009; Mahfouz 2010). This process involves members of the family of PAZ Piwi domain proteins that are capable of binding to 24- to 26-nt-long RNAs transcribed from non-coding regions, called Piwi-interacting RNAs (piRNAs). The PIWI/piRNA complex is guided to specific sequences in the genome by RNA–DNA or RNA–RNA pairing recognition (Wassenegger et al. 1994; Pélissier and Wassenegger 2000). This PIWI complex then attracts DNMT3 to perform de novo methylation. It is possible that the PIWI/piRNA mechanism of directed DNA methylation may affect the placement of other epigenetic modifications such as H3K4 demethylation, which then attract de novo DNA methyltransferases, but it has been shown that the PIWI/piRNA pathway is at least upstream of de novo DNA methylation (Aravin et al. 2008). Recently, it has also been shown that other small RNAs similarly mediate de novo DNA methylation by associating with PIWI proteins in plants. In these instances, siRNAs or miRNAs that arose from miRNA-coding regions guided DNA methylation at some of their generation sites and in trans at their target sites (Chellappan et al. 2010; Wu et al. 2010).
The genomic functional roles of DNA methylation
DNA methylation is known to affect transcriptional silencing when it occurs in gene-promoter regions, transposons, and repeats. In contrast, intragenic DNA methylation (inside gene bodies) is frequently associated with actively transcribed genes, suggesting that the precise role of DNA methylation in transcriptional regulation may vary between promoter and intragenic regions and between genes (Zhang et al. 2006; Hellman and Chess 2007; Zilberman et al. 2007; Ball et al. 2009; Rauch et al. 2009). Although the conserved regulatory function(s) of intragenic DNA methylation remains elusive, an emerging theory congruent with these findings is that one conserved function of exon methylation is the regulation alternative splicing (Laurent et al. 2010; Lyko et al. 2010; Park et al. 2011). This is especially relevant for the honeybee because over 80% of the DNA methylation in its genome was found to be located within exons (Lyko et al. 2010), and DNA methylation distinctly ends at intron–exon boundaries (Flores and Amdam 2011).
Programming of DNA methylation
The targeting of DNA methylation in the genome is both internally regulated, and as discussed in the next section, sensitive to extrinsic signaling. The internal regulation constitutes a program of DNA methylation which interacts with other dynamic molecular processes, such as transcription. Programmed changes in DNA methylation are thought to help regulate cellular differentiation during development by inducing stable alterations in gene expression (Monk et al. 1987; Kafri et al. 1992; Reik 2007; Sasaki and Matsui 2008; Cedar and Bergman 2009). Recent studies of genome-wide DNA methylation support the tenet that programmed locus-specific changes in DNA methylation correlate with changes in cell phenotype (Lister et al. 2009; Laurent et al. 2010; Li et al. 2010). Experimental perturbation of the intrinsic developmental program of DNA methylation can cause drastic changes in phenotype or be lethal to organisms across eukaryotic taxa, including plants (Lindroth et al. 2001; Cao and Jacobsen 2002a, 2002b; Kankel et al. 2003; Xiao et al. 2006), vertebrates (Li et al. 1992; Okano et al. 1999; Stancheva et al. 2001; Li 2002), and invertebrates (Kucharski et al. 2008; Shi et al. 2011).
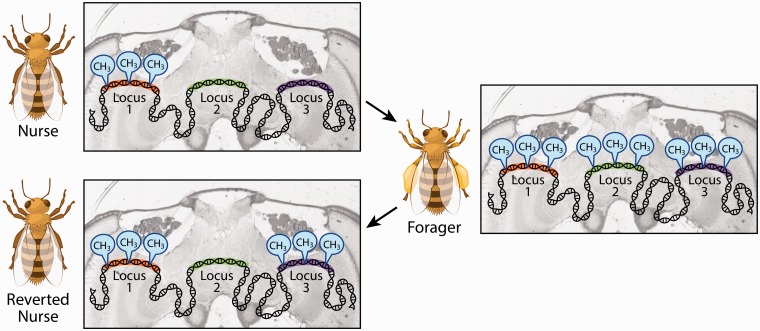
Besides organismal development, programmed changes in DNA methylation are also essential to regulation of synaptic plasticity in memory and of stress-induced behavior (Miller and Sweatt 2007; LaPlant et al. 2010; Miller et al. 2010). For example, various locations in the mouse brain undergo dynamic changes in DNA methylation in connection with neuronal activity (Guo et al. 2011), while inhibition of DNMT enzymes after associative learning in honeybees can interfere with the consolidation of memory (Lockett et al. 2010). Programmed changes in locus-specific DNA methylation also occur in the bee brain during behavioral transitions that are essential for colony fitness (Herb et al. 2012) (Fig. 1). These data suggest that the functional role of the programming of de novo DNA methylation in the brain is conserved between vertebrates and invertebrates.
Fig. 1.
DNA methylation in the honeybee brain is dynamic and associated with an individual’s life history. Honeybee workers usually progress with increasing age from tasks in the nest, such as nursing the brood, to foraging outside the nest. This behavioral switch is essential for colony fitness because it regulates the allocation of workers dedicated to resource-harvesting in response to environmental conditions. When forcing foragers to assume nursing tasks, some, but not all of the patterns of DNA methylation characteristic of foragers will revert to patterns characteristic of nest bees. The patterns of DNA methylation in the brains of nurse bees and foragers differ (Herb et al. 2012).
Changes to the epigenetic code, such as differences in the programming of DNA methylation, can impact fitness by inducing alternative developmental or behavioral phenotypes similar to genetic mutations. Changes in the developmental program of DNA methylation could lead to differences in cellular differentiation and be causal to differences in post-developmental physiology. Changes in the program of neuronal DNA methylation could elicit novel behaviors or behavioral responses to the environment. In the next two sections, we discuss how the environment affects variability in DNA methylation and the epigenetic mechanisms that could transmit such variable DNA methylation to offspring.
The environment as a modifier of DNA methylation
There is increasing evidence that environmental variability can cause variation in the program of DNA methylation in developing offspring. Because DNA methylation also plays a functional role in transcriptional regulation, it is possible that the altered patterns of DNA methylation signaled by the environment may, in turn, signal changes in gene expression. Thereby, variations in DNA-methylation induced by environmental changes may be functional and allow a population to display phenotypic variability despite being genetically homogeneous.
Recent studies in several plant species show that alternative phenotypes can occur in populations with little or no genetic variation, but instead correlate with increased variation in DNA methylation (Lukens and Zhan 2007; Gao et al. 2010; Lira-Medeiros et al. 2010). In the dandelion, Taraxcum officinale, such variability may be induced by environmental stress (Verhoeven et al. 2010). Other data support the view that DNA methylation is required for phenotypic responses to environmental exposures. For example, mutations in the targeting pathway of DNA methylation jin A. thaliana can reduce global genomic DNA methylation along with changes in the plant’s adaptive responses to heat, cold, salt, drought, and flood (Boyko et al. 2010). In animals, moreover, environmental factors such as the maternal diet (Lillycrop et al. 2005, 2007), neonatal diet (Plagemann et al. 2009), rearing behavior (Weaver et al. 2004) and folic acid supplementation (Wolff et al. 1998) can alter de novo programming of DNA methylation during development of the offspring. For example, feeding a protein-restricted diet to pregnant rats results in gene-specific hypo-methylation in the offspring. These differences in DNA methylation in the offspring correlate with changes in their adult phenotype, such as alterations to glucose production in response to stress (Lillycrop et al. 2007) and an increase in systolic blood pressure that may ultimately lead to hypertension (Bogdarina et al. 2007).
Honeybees are an invertebrate species for which the sensitivity of DNA methylation to the environment is conserved. This sensitivity has been socially co-opted to regulate caste fate in female larvae. These larvae can develop into either reproductive queens or sterile workers depending on the diet they receive. The diet is tightly controlled by the larvae’s adult sisters that are nurse bees within the hive. If the rearing of larvae is perturbed, the process of caste differentiation, including the developmental program of DNA methylation is altered, and this process involves changes in the expression of DNMT3 and the locus-specific placement of DNA methylation (Shi et al. 2011). The role of DNA methylation in caste fate was further cemented by results showing that queens can develop from larvae that are artificially reared on a combination of a worker’s diet and silencing DNMT3 with double-stranded RNA (Kucharski et al. 2008).
These studies suggest that DNA methylation is a conserved molecular mechanism in plants, vertebrates, and invertebrates that can be used to convert environmental heterogeneity into phenotypic differences. Thus, we may gain a better understanding of how phenotypic variability arises in populations by studying how the cellular pathways that regulate the genomic targeting of DNA methylation are signaled by environmental change (Fig. 2). Studying these pathways may also allow us to determine how patterns of DNA methylation evolve because genetic changes to these pathways could lead to differences in targeting of DNA methylation and in developmental programming.
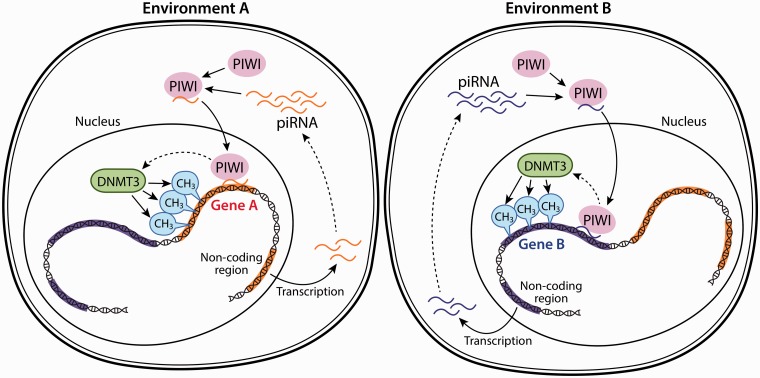
Fig. 2.
Environment and RNA-dependent mechanisms act to influence the methylation of DNA. Patterns of DNA methylation depend on the environment and are directed by piRNAs and the RNA-binding protein PIWI. For simplification, only one gene is shown to be affected under each environment. Under environmental condition A, genomic regions that do not encode a gene (non-coding red part of the DNA) produce piRNAs that bind to PIWI and then attract PIWI to genes that bear near 100% homology on the piRNA sequence. Subsequently, DNMT3 is recruited and the respective gene is methylated. Under environmental condition B, piRNAs derived from another genomic region are produced and a different gene is methylated, leading to an alternate molecular phenotype.
The evolutionary role of environmentally-induced phenotypic heterogeneity mediated by DNA methylation
The capacity for the pathway of de novo DNA methylation to transduce spatial or temporal environmental variation into phenotypic variation implies a potential role for DNA methylation in adaptive evolution. In honeybees, phenotypic variation that is signaled by the environment and mediated by DNA methylation may affect fitness of the colony. Phenotypic heterogeneity among the honeybee workers underlies division of labor within the hive, and an increase in division of labor can increase colony fitness (Waibel et al. 2006; Oldroyd and Fewell 2007). It has been proposed that DNA methylation could be involved in processing the internalization of variations in micro-environments of larvae during development, of workers, thereby helping generate phenotypic heterogeneity in the worker population despite genetic homogeneity (Flores and Amdam 2011). DNA methylation is known to have a functional role in transducing differences in the composition of the diet of larvae between queens and workers (Kucharski et al. 2008). It is possible that DNA methylation may have a similar function among the worker caste, in which altering the larval diet has the capacity to modulate physiological traits that regulate the worker’s heterogeneity, such as the number of ovarioles (Kaftanoglu et al. 2011). Similarly, variability in the environment of queen larvae could signal changes in the queen’s developmental program of DNA methylation. This could lead to an increase in the phenotypic heterogeneity of the worker population that the queen produces if those changes in DNA methylation affect her germline. Alternatively, environmentally signaled DNA methylation could mediate biological effects that are manifest over the lifetime of individual bees, also leading to phenotypic heterogeneity.
Further studies are needed to test whether variability in the environments of larvae or adults (e.g., diet or temperature) could imprint differences in patterns of DNA methylation in workers’ brains, thereby altering behavioral regulation that is critical for division of labor, such as the transition from nurse to forager.
Inheritance of DNA methylation
The capacity to transmit environmentally-induced DNA methylation marks from parent to offspring can be evolutionarily advantageous because it may prepare the offsprings’ phenotype for the environmental stress that the parent(s) may have endured (Mousseau and Fox 1998; Jablonka and Lamb 2005; Youngson and Whitelaw 2008; Jablonka and Raz 2009). A genome-wide erasure of DNA methylation during development would prevent the transfer of DNA methylation marks, and any phenotypic traits caused by them, from parent(s) to offspring. Patterns of DNA methylation are reprogrammed genome-wide during plant and mammalian development, thereby limiting the capacity for transgenerational inheritance of DNA methylation, at least on a genome-wide scale.
DNA methylation reprogramming during plant and mammalian development
The degree to which DNA methylation is erased and then re-established during development differs between plants and mammals; evidence for DNA demethylation is lacking in insects. The erasure of DNA methylation in plants is carried out by a demethylation pathway, which includes the DNA glycosylases and AP lyases ROS1 (repressor of silencing 1), DME (demeter), DML2 (Demeter-like 2), and DML3 (demeter-like 3), to excise cytosine’s that are methylated. The nucleotide gap in DNA is then presumably filled by DNA repair polymerase and ligase enzymes (Zhu 2009). This demethylation pathway affects genome-wide hypo-methylation in Arabidopsis endosperm, especially within transposable elements (Gehring et al. 2009), and mutation of DME partially restores endosperm DNA methylation to the amount found in other tissues (Hsieh et al. 2009). The expression of DME in maternal-specific cells of the endosperm results in the demethylation, and consequently changes in expression, at specific genes, i.e., maternal allele-specific imprinting (Huh et al. 2008).
In mammals, direct transmission of DNA methylation marks from parent(s) to offspring is limited to specific loci because of the waves of genome-wide demethylation, followed by the re-establishment of DNA methylation that occurs during gametogenesis in primordial germ cells and in the embryo immediately following fertilization (Reik 2007; Surani et al. 2007; Sasaki and Matsui 2008; Hemberger et al. 2009). Demethylation in the embryonic stage has the additional complexity that the paternal genome from the sperm is demethylated, but the maternal genome is not and may be protected from demethylation (Mayer et al. 2000; Oswald et al. 2000; Santos et al. 2002; Nakamura et al. 2007). Similarly, differences in the timing of remethylation of DNA occur between the male and female germs cells, in which maternal-specific methylation is established after the male germ cells are initially methylated (Bartolomei and Ferguson-Smith 2011). A combination of active and passive mechanisms of demethylation may contribute to genome-wide demethylation in mammals. Passive demethylation involves the loss of DNA methylation through the lack of maintenance through cell division, resulting in hemi-methylated substrates during the G2 phase of the cell cycle. Several molecular mechanisms have been proposed for active demethylation in mammals, including 5mC modification enzymes, DNA deaminases, and the base excision repair pathway (Hajkova et al. 2010; Popp et al. 2010). It still remains unclear how extensively the active or passive demethylation pathways contribute to the genome-wide erasure of DNA methylation; however, it has been demonstrated that some parental DNA methylation marks can be transmitted to offspring (Richards 2006; Hitchins et al. 2007).
In any case, DNA methylation patterns of the parents could be re-established after demethylation has occurred. This could be facilitated by other epigenetic mechanisms, such as the differential inheritance of DNA-binding proteins, including modified histones, which have the capacity to mediate DNA methylation (Cedar and Bergman 2009). Molecules involved in the targeting of de novo DNA methylation, such as piRNAs, could be passed through the germ line and lead to de novo DNA methylation at specific loci during development in offspring. It has also been demonstrated that DNA methylation imprinted during the development of offspring can be transgenerationally inherited through recapitulation of maternal traits. For example, an increase in the licking and grooming (LG) of pups and arched-back nursing (ABN) by rat mothers alters the pattern of DNA methylation (compared with low-LG–ABN mothers) in the promoter region of the glucocorticoid receptor (GR) in the offspring’s hippocampus. The GR gene regulates the hypothalamic–pituitary–adrenal (HPA) axis and stress response in the hippocampal neurons, and the offspring of high-LG–ABN mothers have a reduced reactivity and anxiety. These more relaxed offspring are then more likely to adopt the same approach to the rearing of young as did their mothers, thereby perpetuating their GR DNA methylation patterns in the next generation (Weaver et al. 2004; Diorio and Meaney 2007).
Evidence for transgenerational inheritance of DNA methylation
Transgenerational inheritance of DNA methylation is more plausible in plants due to the possibility for asexual vegetative reproduction and because in sexual reproduction gametes are derived from almost fully matured vegetative tissue. Transmission of DNA methylation from parent(s) to offspring may also be more adaptive in plants than in animals, since gametes and vegetative offspring are derived from tissue that has been subject to the environmental stress that occurred during almost the entire life history of the parent generation. The exposure to environmental stress has been shown to induce phenotypic changes that can persist to the next generation in plants and animals (Pembrey et al. 2005; Grant-Downton and Dickinson 2006; Koturbash et al. 2006; Molinier et al. 2006) and DNA methylation may play a critical role in the transgenerational perpetuation of such environmentally-induced phenotypes (Mirouze and Paszkowski 2011). For example, in A. thaliana, transgenerational inheritance of stress-induced responses is dependent on de novo DNA methylation (Boyko et al. 2010). A similar study in the dandelion T. officinale found that environmental stress, specifically chemical induction of defensed against herbivores and pathogens, induces differences in DNA methylation and that most of these differences are inherited by the offspring (Verhoeven et al. 2010).
A key feature of the study of dandelions was that it involved a species with asexual reproduction, allowing the variability in DNA methylation to be associated with variability in environments of the parents instead of with genetic variation. DNA methylation may play a role in evolutionary adaptation by providing an epigenetic layer of inheritance on top of genetic inheritance. However, substantiating this concept will require further studies that control for, or accurately measure, genetic variation on a genome-wide scale in order to negate the possibility that observed heritable changes in DNA methylation are caused by genetic mutations.
DNA methylation increases mutability
Genomic regions that are methylated, either due to programmed or environmental signaling, are subject to an increased mutation rate because methylated cytosines spontaneously deaminate to thymine, which is then followed by replacement of guanine by adenine on the opposite DNA strand due to the mismatch repair of DNA (Duncan and Miller 1980). Evidence for these DNA-methylation-induced mutations (DMIMs) in the form of CpG depletion is found at genomic loci that have presumably been methylated over evolutionary time, i.e., across multiple generations, in germ line cells. For example, approximately half of all honeybee genes are methylated, leaving a pattern in which half of all genes have less CpGs than expected (Elango et al. 2009) (Fig. 3 [top]). This pattern of depletion of CpG in honeybees’ genome is also apparent at the exon level, in concordance with the observation that honeybees’ DNA methylation is largely targeted to exons (Figs. 3 [bottom] and 4). Similar patterns of CpG depletion are found in the genomes of Acyrthosiphon pisum, Ciona intestinalis, and humans, but are absent from species that do not show any significant levels of CpG methylation, such as D. melanogaster (Flores and Amdam 2011).
Fig. 3.
Methylation correlates with depletion of CpG in honeybees’ genes and exons. The CpG[O/E] ratio is used as a measure of the depletion of CpG in a genomic region; it is calculated as the number of observed CpGs divided by the number of expected CpGs based on GC content (Elango et al. 2009; Flores and Amdam 2011). (Top) Honeybees’ genes are separated into methylated and unmethylated categories and the distributions of CpG[O/E] ratios is shown for each category. Methylated genes are more depleted of CpGs than are unmethylated genes, likely because of the increased rate of C/T transitions due to deamination of nucleotides. (Bottom) Honeybees’ exons are separated into methylated and unmethylated categories and the distributions of CpG[O/E] ratios is shown for each category. Methylated exons are more depleted of CpGs than are unmethylated exons. Data on DNA methylation were obtained from bisulfite-sequencing of queens and workers [31].
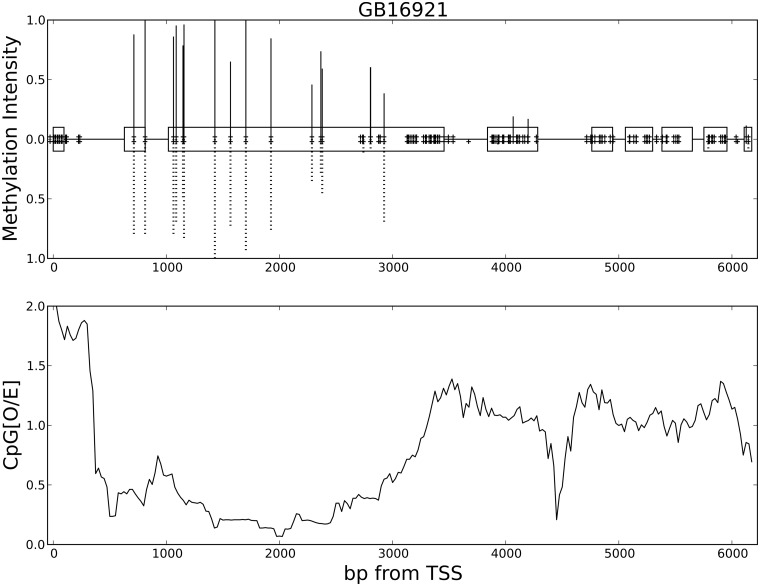
Fig. 4.
Exon-specific depletion of CpG reflects exon-specific DNA methylation in the honeybee gene GB16921 (homolog of Ptip, part of the H3K4 methyltransferase complex). (Top) Base-pair resolution of the intensity of methylation is shown for the length of the gene GB16921. Solid (dotted) lines above (below) the x-axis indicate the intensity of methylation in the queen (worker). The nine exons (1–9 from left to right) for this gene are shown on the x-axis as open boxes. The x-axis is labeled as base-pairs (bp) from translation start site (TSS). Plus signs just above (below) the x-axis indicate CpG coverage from bisulfite-sequencing data in queens (workers). Exons 2 and 3 are heavily methylated in both queens and workers. (Bottom) The CpG[O/E] ratio is calculated with a 200-bp sliding window along the length of the gene (see Fig. 3 for discussion of CpG[O/E]). These data show that depletion of CpG in GB16921 is confined to exons that are heavily methylated (compare with top panel). Data on DNA methylation were obtained at base-pair resolution from genome-wide bisulfite-sequencing of honeybee queens and workers [31].
Because the developmental program of DNA methylation is sensitive to the environment, the level of methylation in some genomic regions in offspring may be a probabilistic function of the environment. In genomic regions at which methylation only occurs during an environmental change, mechanisms of inheritance of transgenerational DNA methylation may help to perpetuate methylation in these genomic regions for several generations after the environmental change has occurred. If this methylation is causal to a phenotype that has a selective advantage during a period of environmental change, then DMIMs could allow that phenotype to become fixed (i.e., genetically programmed). The fixation of phenotypes may be caused by DMIMs that emulate the function of the DNA methylation that was induced by the environmental change. One way this might occur is if DMIMs alter the propensity for a genomic region to be methylated, causing a locus-specific change in DNA methylation (Fig. 5). Alternatively, DMIMs could fix an adaptive phenotype by changing the sequence of a gene-coding region, such as an exon if a DMIM is caused by exon-targeted DNA methylation, or a gene regulatory sequence, such as a transcription factor-binding site if a DMIM is caused by promoter DNA methylation.
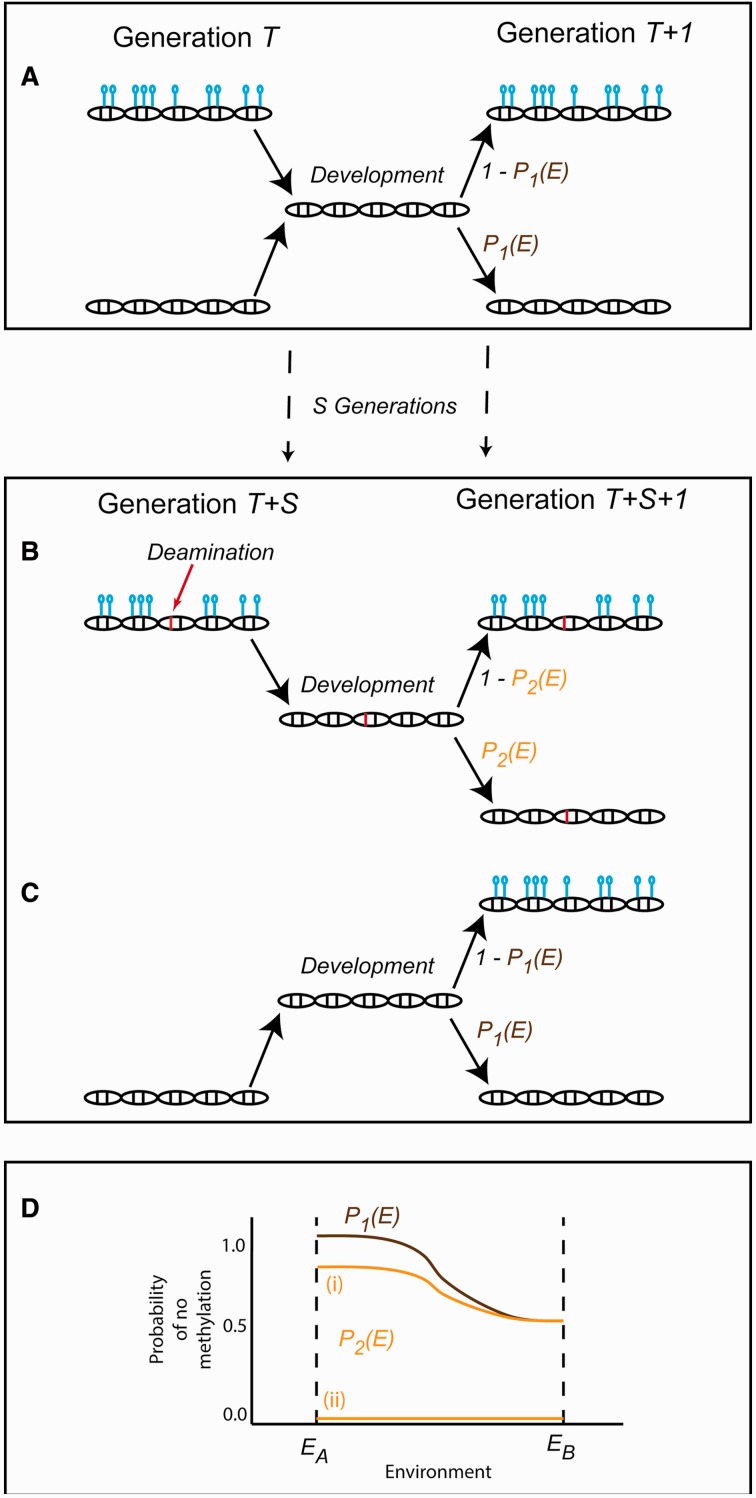
Fig. 5.
The effect of DNA methylation induced mutations on environmentally sensitive methylation patterns. (A) Methylation patterns (open circles attached to lines) between the parent (Generation T) and offspring (Generation T + 1) are erased and then re-established during development whether or not the parent generation is methylated. The mechanism that re-establishes methylation patterns can be sensitive to the environment. Here, EA and EB represent the normal and new environmental conditions, respectively. The probability function P1(E) describes a genotype in which there is no methylation when the environment is EA, and the chance of methylation is random (probability = 0.5) in environment EB. After a number of generations S, deamination in the germline may (B) or may not (C) occur, resulting in a genetic mutation (C → T transition) that changes the probability of methylation during development to a different function P2(E). (D) Scenario (i) represents a case in which P2(E) is more sensitive to a new environment, leading to a higher chance of methylation with less environmental change. Scenario (ii) represents a case in which P2(E) is completely insensitive to the environment and methylation always occurs.
Circumstantial evidence for the evolutionary role of DNA methylation
DMIMs could ultimately circumvent the demethylation waves in the animal germ line and lead to a stable transmission of differences. Experimental evidence supporting the idea of DMIMs comes from studies of the honeybee. Honeybees can be bidirectionally selected for the amount of pollen (source of protein) versus nectar (source of carbohydrate) that is stored by colonies. This colony-level selection for high and low pollen-hoarding was first described by Hellmich et al. (1985) and was subsequently perfected by Page and Fondrk [14]. Page and Amdam (2007) described differences between the resulting genotypes in several traits such as workers’ lifespan, sucrose sensitivity, and ovariole number. Interestingly, in wild-type (unselected) honeybees, similar suites of differences in traits are distributed between sister worker bees that share 0.75 genetic identity (Page and Amdam 2007). This suggests that the bidirectional selection on honeybee food-hoarding captured phenotypic variability that normally is expressed as heterogeneity by genetically similar individuals.
An analysis of quantitative trait loci that differentiate between high and low pollen-hoarding genotypes showed that epigenetic modulators such as histones, mSin3A (a core component of a large multiprotein complex that displays histone deacetylase activity), and a PIWI protein could be, at least partly, responsible for their physiological and behavioral divide (Hunt et al. 2007). We predicted that the two genotypes could differ in their patterns of DNA methylation. Indeed, an analysis of genome-wide DNA-methylation levels showed differences between the two genotypes with respect to their patterns of DNA methylation in the brain (Herb et al., unpublished data). In order to determine whether the two genotypes also show evidence for DMIMs, which would support a heritable difference in epigenetic effects, we analyzed the genome sequences of four biological replicates per genotype by testing for an enrichment of C/T transitions at cytosines within CpG dinucleotides versus all other cytosines. We found a significant enrichment of C/T transitions within CpG dinucleotides that was approximately equal (=2.59-fold) in both the high and low pollen-hoarding genotypes (Table 1). We also observed that there is a 38.7% GC content in third codon positions (3GC) in the honeybee reference genome, indicating a directional mutational pressure in the GC to AT direction (Sueoka 1988; Khrustalev and Barkovsky 2009). However, 81.8% of cytosines in the third codon position were not contained in CpG dinucleotides. Thus, it is unlikely that factors causal to the AT-pressure in the honeybee genome, besides nucleotide deamination, contributed to the enrichment of C/T transitions within the CpG dinucleotides that we found. These observations suggest a connection between DNA methylation and mutations in the two selected genotypes.
Table 1.
CpG dinucleotides are enriched for C/T transition in bidirectionally selected strains of honeybees
| High pollen-hoarding strain |
Low pollen-hoarding strain |
|||
|---|---|---|---|---|
| C/T | no C/T | C/T | no C/T | |
| C not within CpG | 22,410 | 63,758,382 | 17,381 | 63,763,411 |
| C within CpG | 9148 | 10,020,985 | 7102 | 10,023,031 |
Compared with the number of C/T transitions at cytosine’s not contained in CpG dinucleotides, there are ∼2.59 times more C/T transitions at CpG dinucleotides in honeybee genotypes that were artificially selected for high or low pollen-hoarding behavior [14]. This enrichment of C/T transitions is statistically significant by Fisher’s exact test in each of the two genotypes (P < 2.2e−16). We re-sequenced the whole genomes of four individuals from each of the low and high pollen-hoarding strains of honeybees, using deep sequencing (HiSeq 2000, Illumina, San Diego, CA). Reads generated from deep sequencing were aligned to Amel_2.0 with Bowtie (version 0.12.7) with default options (Langmead et al. 2009). To determine the presence of genetic transitions between the high and low strains, we performed a standard case/control association analysis using PLINK (version 1.07) (Purcell et al. 2007; Purcell 2010); default options and a P-value cutoff of 0.05 were used to infer the presence of a significant genetic difference. This resulted in a total of 401,804 significant genetic differences. For each genotype, we tallied the number of C/T transitions by counting the number of significant genetic differences in which the genotype was called T and the reference genome was C. Similarly, C/T transitions were counted on the opposite strand if a G/A transition was found on the positive strand.
Remarks and future work
Phenotypic plasticity is a ubiquitous property in plants and animals that enables a population to achieve phenotypic variability with respect to environmental change despite genetic uniformity (West-Eberhard 2003). Epigenetic mechanisms used in the developmental program of an organism can be sensitive to the environment; hence epigenetic variation is expected to occur when phenotypic plasticity is manifest. Epigenetic mechanisms such as DNA methylation provide a means of extending the flexibility of the genome by affecting changes to the transcriptome, and to thus increase phenotypic plasticity. Here, we suggest that, by causing increased mutability, DNA methylation links this flexibility with evolutionary processes, culminating in selectable genetic variability. Interestingly, recent work has also shown that DNA hypo-methylation is associated with a higher frequency of homologous recombination and genomic instability (Li et al. 2012). Thus, it is possible that environmentally-induced hyper-methylation or hypo-methylation of DNA could lead to a higher mutation rate.
The evolutionary role of phenotypic plasticity mediated by DNA methylation remains unclear because patterns of DNA methylation mostly are reset in the gametes of plants and mammals, and in the primordial germ cells (PGCs) of mammals (Feng et al. 2010; Law and Jacobsen 2010). However, despite this limitation, we reviewed several mechanisms whereby methylation patterns are transferred transgenerationally. The assessment of these mechanisms will be facilitated by a clearer understanding of how piRNAs are generated, direct the placement of DNA methylation, and whether they are transferred to eggs or embryos. The transgenerational transference of functional DNA methylation has the potential for contributing to short-term adaptation to environmental changes that cause variation in the methylomes of offspring. This effect can vary ever more dramatically in a population that exhibits genetic variance in master regulators of the machinery of DNA methylation and thus shows a broad range in sensitivity toward environmental perturbations. The resulting variable epigenetic response could confer positive fitness effects with respect to environmental change if it increases the rate at which alternative phenotypes that are only manifested during periods of environmental change become genetically fixed (West-Eberhard 2003). For example, in the genotypes of pollen-hoarding honeybees, it is possible that genetic differences in key epigenetic regulators, such as PIWI proteins, could have led to a difference in overall DNA methylation and thereby to variability in phenotypes such as pollen-hoarding behavior. In this scenario, a difference in behavioral phenotype is then repeatedly selected upon over generations and, over time, leads to DMIMs.
It remains to be uncovered precisely how DMIMs aid in the adaptability of an organism to its environment. We argue that DMIMs could decouple the developmental response from the environment by changing the likelihood that the functional effect of DNA methylation will occur without environmental extremes or by fixing genetic changes that replace the effect of DNA methylation. Thereby, DNA methylation may play a role in evolutionary adaptation due to the increased mutability induced in genomic regions where it is used.
Future research targeted at the fixation and reversibility of DNA methylation will be needed in order to shed more light on the question of how changes in the environment relate to epigenetic patterns and adaptability of organisms. Because of their comparably small genomes and their almost exclusive restriction of DNA methylation to exons, insects provide a good model system to study these highly complex relationships. The advent of widely available and increasingly affordable bisulfite-sequencing of entire genomes is bound to help advance our knowledge in these areas. It will be highly instructive to analyze variations of DNA methylation and C/T transitions in wild-type populations and follow their associations throughout controlled breeding programs, such as the bidirectional selection of pollen hoarding in honeybees, in order to better elucidate the connections between DNA methylation and mutation. Furthermore, it will be important to investigate details of the DNA-targeting mechanism in order to understand how the same regions in the genome get consistently methylated, eventually leading to an increased mutation rate for C/T transitions in these areas, but not in others.
Funding
The Research Council of Norway (nos 180504, 191699, and 213976); the PEW Charitable Trust; and National Institute on Aging (NIA P01 AG22500) (to G.V.A.).
Acknowledgments
We thank Sabine Deviche and the ASU School of Life Sciences Vislab for their help with the figures, and David Galbraith and Erik Rasmussen for helpful comments and suggestions.
References
- Aravin AA, Hannon GJ. Small RNA silencing pathways in germ and stem cells. Cold Spring Harb Symp Quant Biol. 2008;73:283–90. doi: 10.1101/sqb.2008.73.058. [DOI] [PubMed] [Google Scholar]
- Aravin AA, Sachidanandam R, Bourc’his D, Schaefer C, Pezic D, Toth KF, Bestor T, Hannon GJ. A piRNA pathway primed by individual transposons is linked to de novo DNA methylation in mice. Mol Cell. 2008;31:785–99. doi: 10.1016/j.molcel.2008.09.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ball MP, Li JB, Gao Y, Lee J-H, LeProust EM, Park I-H, Xie B, Daley GQ, Church GM. Targeted and genome-scale strategies reveal gene-body methylation signatures in human cells. Nat Biotechnol. 2009;27:361–8. doi: 10.1038/nbt.1533. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bartolomei MS, Ferguson-Smith AC. Mammalian Genomic Imprinting. Cold Spring Harb Perspect Biol. 2011;3 doi: 10.1101/cshperspect.a002592. ( http://cshperspectives.cshlp.org/content/3/7/a002592) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beckingham KM, Armstrong JD, Texada MJ, Munjaal R, Baker DA. Drosophila melanogaster—the model organism of choice for the complex biology of multi-cellular organisms. Gravit Space Biol Bull. 2005;18:17–29. [PubMed] [Google Scholar]
- Bogdarina I, Welham S, King PJ, Burns SP, Clark AJL. Epigenetic modification of the renin-angiotensin system in the fetal programming of hypertension. Circ Res. 2007;100:520–6. doi: 10.1161/01.RES.0000258855.60637.58. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boyko A, Blevins T, Yao Y, Golubov A, Bilichak A, Ilnytskyy Y, Hollunder J, Hollander J, Meins F, Jr, Kovalchuk I. Transgenerational adaptation of Arabidopsis to stress requires DNA methylation and the function of Dicer-like proteins. PLoS One. 2010;5:e9514. doi: 10.1371/journal.pone.0009514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boyko A, Kovalchuk I. Genome instability and epigenetic modification—heritable responses to environmental stress? Curr Opin Plant Biol. 2011;14:260–6. doi: 10.1016/j.pbi.2011.03.003. [DOI] [PubMed] [Google Scholar]
- Britten RJ, Baron WF, Stout DB, Davidson EH. Sources and evolution of human Alu repeated sequences. Proc Natl Acad Sci U S A. 1988;85:4770–4. doi: 10.1073/pnas.85.13.4770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bulmer M. Neighboring base effects on substitution rates in pseudogenes. Mol Biol Evol. 1986;3:322–9. doi: 10.1093/oxfordjournals.molbev.a040401. [DOI] [PubMed] [Google Scholar]
- Cao X, Jacobsen SE. Locus-specific control of asymmetric and CpNpG methylation by the DRM and CMT3 methyltransferase genes. Proc Natl Acad Sci U S A. 2002a;99(Suppl 4):16491–8. doi: 10.1073/pnas.162371599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cao X, Jacobsen SE. Role of the arabidopsis DRM methyltransferases in de novo DNA methylation and gene silencing. Curr Biol. 2002b;12:1138–44. doi: 10.1016/s0960-9822(02)00925-9. [DOI] [PubMed] [Google Scholar]
- Cedar H, Bergman Y. Linking DNA methylation and histone modification: patterns and paradigms. Nat Rev Genet. 2009;10:295–304. doi: 10.1038/nrg2540. [DOI] [PubMed] [Google Scholar]
- Chan SW-L, Henderson IR, Jacobsen SE. Gardening the genome: DNA methylation in Arabidopsis thaliana. Nat Rev Genet. 2005;6:351–60. doi: 10.1038/nrg1601. [DOI] [PubMed] [Google Scholar]
- Chellappan P, Xia J, Zhou X, Gao S, Zhang X, Coutino G, Vazquez F, Zhang W, Jin H. siRNAs from miRNA sites mediate DNA methylation of target genes. Nucleic Acids Res. 2010;38:6883–94. doi: 10.1093/nar/gkq590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Day JJ, Sweatt JD. DNA methylation and memory formation. Nat Neurosci. 2010;13:1319–23. doi: 10.1038/nn.2666. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diorio J, Meaney MJ. Maternal programming of defensive responses through sustained effects on gene expression. J Psychiatry Neurosci. 2007;32:275–84. [PMC free article] [PubMed] [Google Scholar]
- Duncan BK, Miller JH. Mutagenic deamination of cytosine residues in DNA. Nature. 1980;287:560–1. doi: 10.1038/287560a0. [DOI] [PubMed] [Google Scholar]
- Elango N, Hunt BG, Goodisman MAD, Yi SV. DNA methylation is widespread and associated with differential gene expression in castes of the honeybee, Apis mellifera. Proc Natl Acad Sci U S A. 2009;106:11206–11. doi: 10.1073/pnas.0900301106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feng S, Jacobsen SE, Reik W. Epigenetic reprogramming in plant and animal development. Science. 2010;330:622–7. doi: 10.1126/science.1190614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Flores KB, Amdam GV. Deciphering a methylome: what can we read into patterns of DNA methylation? J Exp Biol. 2011;214:3155–63. doi: 10.1242/jeb.059741. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao L, Geng Y, Li B, Chen J, Yang J. Genome-wide DNA methylation alterations of Alternanthera philoxeroides in natural and manipulated habitats: implications for epigenetic regulation of rapid responses to environmental fluctuation and phenotypic variation. Plant Cell Environ. 2010;33:1820–7. doi: 10.1111/j.1365-3040.2010.02186.x. [DOI] [PubMed] [Google Scholar]
- Gehring M, Bubb KL, Henikoff S. Extensive demethylation of repetitive elements during seed development underlies gene imprinting. Science. 2009;324:1447–51. doi: 10.1126/science.1171609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grant-Downton RT, Dickinson HG. Epigenetics and its implications for plant biology 2. The “epigenetic epiphany”: epigenetics, evolution and beyond. Ann Bot. 2006;97:11–27. doi: 10.1093/aob/mcj001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guo JU, Ma DK, Mo H, Ball MP, Jang M-H, Bonaguidi MA, Balazer JA, Eaves HL, Xie B, Ford E, et al. Neuronal activity modifies the DNA methylation landscape in the adult brain. Nat Neurosci. 2011;14:1345–51. doi: 10.1038/nn.2900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hajkova P, Jeffries SJ, Lee C, Miller N, Jackson SP, Surani MA. Genome-wide reprogramming in the mouse germ line entails the base excision repair pathway. Science. 2010;329:78–82. doi: 10.1126/science.1187945. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hellman A, Chess A. Gene body-specific methylation on the active X chromosome. Science. 2007;315:1141–3. doi: 10.1126/science.1136352. [DOI] [PubMed] [Google Scholar]
- Hellmich RL, Kulincevic JM, Rothenbuhler WC. Selection for high and low pollen-hoarding honey bees. J Hered. 1985;76:155–158. [Google Scholar]
- Hemberger M, Dean W, Reik W. Epigenetic dynamics of stem cells and cell lineage commitment: digging Waddington’s canal. Nat Rev Mol Cell Biol. 2009;10:526–37. doi: 10.1038/nrm2727. [DOI] [PubMed] [Google Scholar]
- Herb BR, Wolschin F, Hansen KD, Aryee MJ, Langmead B, Irizarry R, Amdam GV, Feinberg AP. Reversible switching between epigenetic states in honeybee behavioral subcastes. Nat Neurosci. 2012;15:1371–3. doi: 10.1038/nn.3218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hitchins MP, Wong JJL, Suthers G, Suter CM, Martin DIK, Hawkins NJ, Ward RL. Inheritance of a cancer-associated MLH1 germ-line epimutation. N Engl J Med. 2007;356:697–705. doi: 10.1056/NEJMoa064522. [DOI] [PubMed] [Google Scholar]
- Hsieh T-F, Ibarra CA, Silva P, Zemach A, Eshed-Williams L, Fischer RL, Zilberman D. Genome-wide demethylation of Arabidopsis endosperm. Science. 2009;324:1451–4. doi: 10.1126/science.1172417. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huh JH, Bauer MJ, Hsieh T-F, Fischer RL. Cellular programming of plant gene imprinting. Cell. 2008;132:735–44. doi: 10.1016/j.cell.2008.02.018. [DOI] [PubMed] [Google Scholar]
- Hunt GJ, Amdam GV, Schlipalius D, Emore C, Sardesai N, Williams CE, Rueppell O, Guzmán-Novoa E, Arechavaleta-Velasco M, Chandra S, et al. Behavioral genomics of honeybee foraging and nest defense. Naturwissenschaften. 2007;94:247–67. doi: 10.1007/s00114-006-0183-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jablonka E, Lamb MJ. Evolution in four dimensions: genetic, epigenetic, behavioral, and symbolic variation in the history of life. Cambridge (MA): MIT Press; 2005. [Google Scholar]
- Jablonka E, Raz G. Transgenerational epigenetic inheritance: prevalence, mechanisms, and implications for the study of heredity and evolution. Q Rev Biol. 2009;84:131–76. doi: 10.1086/598822. [DOI] [PubMed] [Google Scholar]
- Kafri T, Ariel M, Brandeis M, Shemer R, Urven L, McCarrey J, Cedar H, Razin A. Developmental pattern of gene-specific DNA methylation in the mouse embryo and germ line. Genes Dev. 1992;6:705–14. doi: 10.1101/gad.6.5.705. [DOI] [PubMed] [Google Scholar]
- Kaftanoglu O, Linksvayer TA, Page RE. Rearing honey bees, Apis mellifera, in vitro 1: effects of sugar concentrations on survival and development. J Insect Sci. 2011;11:1–10. doi: 10.1673/031.011.9601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kankel MW, Ramsey DE, Stokes TL, Flowers SK, Haag JR, Jeddeloh JA, Riddle NC, Verbsky ML, Richards EJ. Arabidopsis MET1 cytosine methyltransferase mutants. Genetics. 2003;163:1109–22. doi: 10.1093/genetics/163.3.1109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khrustalev VV, Barkovsky EV. Mutational pressure is a cause of inter- and intragenomic differences in GC-content of simplex and varicella viruses. Comput Biol Chem. 2009;33:295–302. doi: 10.1016/j.compbiolchem.2009.06.005. [DOI] [PubMed] [Google Scholar]
- Klose RJ, Bird AP. Genomic DNA methylation: the mark and its mediators. Trends Biochem Sci. 2006;31:89–97. doi: 10.1016/j.tibs.2005.12.008. [DOI] [PubMed] [Google Scholar]
- Koturbash I, Baker M, Loree J, Kutanzi K, Hudson D, Pogribny I, Sedelnikova O, Bonner W, Kovalchuk O. Epigenetic dysregulation underlies radiation-induced transgenerational genome instability in vivo. Int J Radiat Oncol Biol Phys. 2006;66:327–30. doi: 10.1016/j.ijrobp.2006.06.012. [DOI] [PubMed] [Google Scholar]
- Kucharski R, Maleszka J, Foret S, Maleszka R. Nutritional control of reproductive status in honeybees via DNA methylation. Science. 2008;319:1827–30. doi: 10.1126/science.1153069. [DOI] [PubMed] [Google Scholar]
- Kuramochi-Miyagawa S, Watanabe T, Gotoh K, Totoki Y, Toyoda A, Ikawa M, Asada N, Kojima K, Yamaguchi Y, Ijiri TW, et al. DNA methylation of retrotransposon genes is regulated by Piwi family members MILI and MIWI2 in murine fetal testes. Genes Dev. 2008;22:908–17. doi: 10.1101/gad.1640708. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Langmead B, Trapnell C, Pop M, Salzberg SL. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 2009;10:R25. doi: 10.1186/gb-2009-10-3-r25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- LaPlant Q, Vialou V, Covington HE, Dumitriu D, Feng J, Warren BL, Maze I, Dietz DM, Watts EL, Iñiguez SD, et al. Dnmt3a regulates emotional behavior and spine plasticity in the nucleus accumbens. Nat Neurosci. 2010;13:1137–43. doi: 10.1038/nn.2619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laurent L, Wong E, Li G, Huynh T, Tsirigos A, Ong CT, Low HM, Kin Sung KW, Rigoutsos I, Loring J, et al. Dynamic changes in the human methylome during differentiation. Genome Res. 2010;20:320–31. doi: 10.1101/gr.101907.109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Law JA, Jacobsen SE. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat Rev Genet. 2010;11:204–20. doi: 10.1038/nrg2719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li E, Bestor TH, Jaenisch R. Targeted mutation of the DNA methyltransferase gene results in embryonic lethality. Cell. 1992;69:915–26. doi: 10.1016/0092-8674(92)90611-f. [DOI] [PubMed] [Google Scholar]
- Li E. Chromatin modification and epigenetic reprogramming in mammalian development. Nat Rev Genet. 2002;3:662–73. doi: 10.1038/nrg887. [DOI] [PubMed] [Google Scholar]
- Li J, Harris RA, Cheung SW, Coarfa C, Jeong M, Goodell MA, White LD, Patel A, Kang S-H, Shaw C, et al. Genomic hypomethylation in the human germline associates with selective structural mutability in the human genome. PLoS Genet. 2012;8:e1002692. doi: 10.1371/journal.pgen.1002692. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Y, Zhu J, Tian G, Li N, Li Q, Ye M, Zheng H, Yu J, Wu H, Jihua Sun, et al. The DNA methylome of human peripheral blood mononuclear cells. PLoS Biol. 2010;8:e1000533. doi: 10.1371/journal.pbio.1000533. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lillycrop KA, Phillips ES, Jackson AA, Hanson MA, Burdge GC. Dietary protein restriction of pregnant rats induces and folic acid supplementation prevents epigenetic modification of hepatic gene expression in the offspring. J Nutr. 2005;135:1382–6. doi: 10.1093/jn/135.6.1382. [DOI] [PubMed] [Google Scholar]
- Lillycrop KA, Slater-Jefferies JL, Hanson MA, Godfrey KM, Jackson AA, Burdge GC. Induction of altered epigenetic regulation of the hepatic glucocorticoid receptor in the offspring of rats fed a protein-restricted diet during pregnancy suggests that reduced DNA methyltransferase-1 expression is involved in impaired DNA methylation and changes in histone modifications. Br J Nutr. 2007;97:1064–73. doi: 10.1017/S000711450769196X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lindroth AM, Cao X, Jackson JP, Zilberman D, McCallum CM, Henikoff S, Jacobsen SE. Requirement of CHROMOMETHYLASE3 for maintenance of CpXpG methylation. Science. 2001;292:2077–80. doi: 10.1126/science.1059745. [DOI] [PubMed] [Google Scholar]
- Lira-Medeiros CF, Parisod C, Fernandes RA, Mata CS, Cardoso MA, Ferreira PCG. Epigenetic variation in mangrove plants occurring in contrasting natural environment. PLoS One. 2010;5:e10326. doi: 10.1371/journal.pone.0010326. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lister R, O’Malley RC, Tonti-Filippini J, Gregory BD, Berry CC, Millar AH, Ecker JR. Highly integrated single-base resolution maps of the epigenome in Arabidopsis. Cell. 2008;133:523–36. doi: 10.1016/j.cell.2008.03.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lister R, Pelizzola M, Dowen RH, Hawkins RD, Hon G, Tonti-Filippini J, Nery JR, Lee L, Ye Z, Ngo Q-M, et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature. 2009;462:315–22. doi: 10.1038/nature08514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lockett GA, Helliwell P, Maleszka R. Involvement of DNA methylation in memory processing in the honey bee. Neuroreport. 2010;21:812–6. doi: 10.1097/WNR.0b013e32833ce5be. [DOI] [PubMed] [Google Scholar]
- Lukens LN, Zhan S. The plant genome’s methylation status and response to stress: implications for plant improvement. Curr Opin Plant Biol. 2007;10:317–22. doi: 10.1016/j.pbi.2007.04.012. [DOI] [PubMed] [Google Scholar]
- Lyko F, Foret S, Kucharski R, Wolf S, Falckenhayn C, Maleszka R. The honey bee epigenomes: differential methylation of brain DNA in queens and workers. PLoS Biol. 2010;8:e1000506. doi: 10.1371/journal.pbio.1000506. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lyko F, Maleszka R. Insects as innovative models for functional studies of DNA methylation. Trends Genet. 2011;27:127–31. doi: 10.1016/j.tig.2011.01.003. [DOI] [PubMed] [Google Scholar]
- Mahfouz MM. RNA-directed DNA methylation: mechanisms and functions. Plant Signal Behav. 2010;5:806–16. doi: 10.4161/psb.5.7.11695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mayer W, Niveleau A, Walter J, Fundele R, Haaf T. Demethylation of the zygotic paternal genome. Nature. 2000;403:501–2. doi: 10.1038/35000656. [DOI] [PubMed] [Google Scholar]
- Mette MF, Aufsatz W, Van der Winden J, Matzke MA, Matzke AJM. Transcriptional silencing and promoter methylation triggered by double-stranded RNA. EMBO J. 2000;19:5194–201. doi: 10.1093/emboj/19.19.5194. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller CA, Gavin CF, White JA, Parrish RR, Honasoge A, Yancey CR, Rivera IM, Rubio MD, Rumbaugh G, Sweatt JD. Cortical DNA methylation maintains remote memory. Nat Neurosci. 2010;13:664–6. doi: 10.1038/nn.2560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller CA, Sweatt JD. Covalent modification of DNA regulates memory formation. Neuron. 2007;53:857–69. doi: 10.1016/j.neuron.2007.02.022. [DOI] [PubMed] [Google Scholar]
- Mirouze M, Paszkowski J. Epigenetic contribution to stress adaptation in plants. Curr Opin Plant Biol. 2011;14:267–74. doi: 10.1016/j.pbi.2011.03.004. [DOI] [PubMed] [Google Scholar]
- Molinier J, Ries G, Zipfel C, Hohn B. Transgeneration memory of stress in plants. Nature. 2006;442:1046–9. doi: 10.1038/nature05022. [DOI] [PubMed] [Google Scholar]
- Monk M, Boubelik M, Lehnert S. Temporal and regional changes in DNA methylation in the embryonic, extraembryonic and germ cell lineages during mouse embryo development. Development. 1987;99:371–82. doi: 10.1242/dev.99.3.371. [DOI] [PubMed] [Google Scholar]
- Morris KV. RNA-directed transcriptional gene silencing and activation in human cells. Oligonucleotides. 2009;19:299–306. doi: 10.1089/oli.2009.0212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mousseau TA, Fox CW. Maternal effects as adaptations. New York: Oxford University Press; 1998. [Google Scholar]
- Nakamura T, Arai Y, Umehara H, Masuhara M, Kimura T, Taniguchi H, Sekimoto T, Ikawa M, Yoneda Y, Okabe M, et al. PGC7/Stella protects against DNA demethylation in early embryogenesis. Nat Cell Biol. 2007;9:64–71. doi: 10.1038/ncb1519. [DOI] [PubMed] [Google Scholar]
- Okano M, Bell DW, Haber DA, Li E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell. 1999;99:247–57. doi: 10.1016/s0092-8674(00)81656-6. [DOI] [PubMed] [Google Scholar]
- Oldroyd BP, Fewell JH. Genetic diversity promotes homeostasis in insect colonies. Trends Ecol Evol (Amst) 2007;22:408–13. doi: 10.1016/j.tree.2007.06.001. [DOI] [PubMed] [Google Scholar]
- Oswald J, Engemann S, Lane N, Mayer W, Olek A, Fundele R, Dean W, Reik W, Walter J. Active demethylation of the paternal genome in the mouse zygote. Curr Biol. 2000;10:475–8. doi: 10.1016/s0960-9822(00)00448-6. [DOI] [PubMed] [Google Scholar]
- Page RE, Fondrk MK. The effects of colony-level selection on the social organization of honey bee (Apis mellifera L.) colonies: colony-level components of pollen hoarding. Behav Ecol Sociobiol. 1995;36:135–44. [Google Scholar]
- Page RE, Jr, Amdam GV. The making of a social insect: developmental architectures of social design. Bioessays. 2007;29:334–43. doi: 10.1002/bies.20549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park J, Peng Z, Zeng J, Elango N, Park T, Wheeler D, Werren JH, Yi SV. Comparative analyses of DNA methylation and sequence evolution using Nasonia genomes. Mol Biol Evol. 2011;28:3345–54. doi: 10.1093/molbev/msr168. ( http://mbe.oxfordjournals.org/content/early/2011/06/20/molbev.msr168.abstract) [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pélissier T, Wassenegger M. A DNA target of 30 bp is sufficient for RNA-directed DNA methylation. RNA. 2000;6:55–65. doi: 10.1017/s135583820099201x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pembrey ME, Bygren LO, Kaati G, Edvinsson S, Northstone K, Sjostrom M, Golding J. Sex-specific, male-line transgenerational responses in humans. Eur J Hum Genet. 2005;14:159–66. doi: 10.1038/sj.ejhg.5201538. [DOI] [PubMed] [Google Scholar]
- Plagemann A, Harder T, Brunn M, Harder A, Roepke K, Wittrock-Staar M, Ziska T, Schellong K, Rodekamp E, Melchior K, et al. Hypothalamic proopiomelanocortin promoter methylation becomes altered by early overfeeding: an epigenetic model of obesity and the metabolic syndrome. J Physiol (Lond) 2009;587:4963–76. doi: 10.1113/jphysiol.2009.176156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Popp C, Dean W, Feng S, Cokus SJ, Andrews S, Pellegrini M, Jacobsen SE, Reik W. Genome-wide erasure of DNA methylation in mouse primordial germ cells is affected by AID deficiency. Nature. 2010;463:1101–5. doi: 10.1038/nature08829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D, Maller J, Sklar P, De Bakker PIW, Daly MJ, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81:559–75. doi: 10.1086/519795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Purcell S. 2010. PLINK (version 1.07) ( http://pngu.mgh.harvard.edu/purcell/plink/) [Google Scholar]
- Rauch TA, Wu X, Zhong X, Riggs AD, Pfeifer GP. A human B cell methylome at 100-base pair resolution. Proc Natl Acad Sci U S A. 2009;106:671–8. doi: 10.1073/pnas.0812399106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reik W. Stability and flexibility of epigenetic gene regulation in mammalian development. Nature. 2007;447:425–32. doi: 10.1038/nature05918. [DOI] [PubMed] [Google Scholar]
- Richards EJ. Inherited epigenetic variation—revisiting soft inheritance. Nat Rev Genet. 2006;7:395–401. doi: 10.1038/nrg1834. [DOI] [PubMed] [Google Scholar]
- Santos F, Hendrich B, Reik W, Dean W. Dynamic reprogramming of DNA methylation in the early mouse embryo. Dev Biol. 2002;241:172–82. doi: 10.1006/dbio.2001.0501. [DOI] [PubMed] [Google Scholar]
- Sasaki H, Matsui Y. Epigenetic events in mammalian germ-cell development: reprogramming and beyond. Nat Rev Genet. 2008;9:129–40. doi: 10.1038/nrg2295. [DOI] [PubMed] [Google Scholar]
- Shi YY, Huang ZY, Zeng ZJ, Wang ZL, Wu XB, Yan WY. Diet and cell size both affect queen-worker differentiation through DNA methylation in honey bees (Apis mellifera, Apidae) PLoS One. 2011;6:e18808. doi: 10.1371/journal.pone.0018808. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stancheva I, Hensey C, Meehan RR. Loss of the maintenance methyltransferase, xDnmt1, induces apoptosis in Xenopus embryos. EMBO J. 2001;20:1963–73. doi: 10.1093/emboj/20.8.1963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sueoka N. Directional mutation pressure and neutral molecular evolution. Proc Natl Acad Sci U S A. 1988;85:2653–7. doi: 10.1073/pnas.85.8.2653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Surani MA, Hayashi K, Hajkova P. Genetic and epigenetic regulators of pluripotency. Cell. 2007;128:747–62. doi: 10.1016/j.cell.2007.02.010. [DOI] [PubMed] [Google Scholar]
- Sved J, Bird A. The expected equilibrium of the CpG dinucleotide in vertebrate genomes under a mutation model. Proc Natl Acad Sci U S A. 1990;87:4692–6. doi: 10.1073/pnas.87.12.4692. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Verhoeven KJF, Jansen JJ, Van Dijk PJ, Biere A. Stress-induced DNA methylation changes and their heritability in asexual dandelions. New Phytol. 2010;185:1108–18. doi: 10.1111/j.1469-8137.2009.03121.x. [DOI] [PubMed] [Google Scholar]
- Waibel M, Floreano D, Magnenat S, Keller L. Division of labour and colony efficiency in social insects: effects of interactions between genetic architecture, colony kin structure and rate of perturbations. Proc Biol Sci. 2006;273:1815–23. doi: 10.1098/rspb.2006.3513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wassenegger M, Heimes S, Riedel L, Sänger HL. RNA-directed de novo methylation of genomic sequences in plants. Cell. 1994;76:567–76. doi: 10.1016/0092-8674(94)90119-8. [DOI] [PubMed] [Google Scholar]
- Weaver ICG, Cervoni N, Champagne FA, D’Alessio AC, Sharma S, Seckl JR, Dymov S, Szyf M, Meaney MJ. Epigenetic programming by maternal behavior. Nat Neurosci. 2004;7:847–54. doi: 10.1038/nn1276. [DOI] [PubMed] [Google Scholar]
- West-Eberhard MJ. Developmental plasticity and evolution. Oxford: Oxford University Press; 2003. [Google Scholar]
- Wolff GL, Kodell RL, Moore SR, Cooney CA. Maternal epigenetics and methyl supplements affect agouti gene expression in Avy/a mice. FASEB J. 1998;12:949–57. [PubMed] [Google Scholar]
- Wu L, Zhou H, Zhang Q, Zhang J, Ni F, Liu C, Qi Y. DNA methylation mediated by a microRNA pathway. Mol Cell. 2010;38:465–75. doi: 10.1016/j.molcel.2010.03.008. [DOI] [PubMed] [Google Scholar]
- Xiao W, Custard KD, Brown RC, Lemmon BE, Harada JJ, Goldberg RB, Fischer RL. DNA methylation is critical for Arabidopsis embryogenesis and seed viability. Plant Cell. 2006;18:805–14. doi: 10.1105/tpc.105.038836. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Youngson NA, Whitelaw E. Transgenerational epigenetic effects. Annu Rev Genomics Hum Genet. 2008;9:233–57. doi: 10.1146/annurev.genom.9.081307.164445. [DOI] [PubMed] [Google Scholar]
- Zhang X, Yazaki J, Sundaresan A, Cokus S, Chan SW-L, Chen H, Henderson IR, Shinn P, Pellegrini M, Jacobsen SE, et al. Genome-wide high-resolution mapping and functional analysis of DNA methylation in Arabidopsis. Cell. 2006;126:1189–201. doi: 10.1016/j.cell.2006.08.003. [DOI] [PubMed] [Google Scholar]
- Zhu J-K. Active DNA demethylation mediated by DNA glycosylases. Annu Rev Genet. 2009;43:143–66. doi: 10.1146/annurev-genet-102108-134205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zilberman D, Gehring M, Tran RK, Ballinger T, Henikoff S. Genome-wide analysis of Arabidopsis thaliana DNA methylation uncovers an interdependence between methylation and transcription. Nat Genet. 2007;39:61–9. doi: 10.1038/ng1929. [DOI] [PubMed] [Google Scholar]