Abstract
Staufen1 (STAU1)-mediated mRNA decay (SMD) is an mRNA degradation process in mammalian cells that is mediated by the binding of STAU1 to a STAU1-binding site (SBS) within the 3'-untranslated region (3'UTR) of target mRNAs. During SMD, STAU1, a double-stranded (ds) RNA-binding protein, recognizes dsRNA structures formed either by intramolecular base-pairing of 3'UTR sequences or by intermolecular base-pairing of 3'UTR sequences with a long noncoding RNA (lncRNA) via partially complementary Alu elements. Recently, STAU2, a paralog of STAU1, has also been reported to mediate SMD. Both STAU1 and STAU2 interact directly with the ATP-dependent RNA helicase UPF1, a key SMD factor, enhancing its helicase activity to promote effective SMD. Moreover, STAU1 and STAU2 form homodimeric and heterodimeric interactions via domain-swapping. Since both SMD and the mechanistically related nonsense-mediated mRNA decay (NMD) employ UPF1, SMD and NMD are competitive pathways. Competition contributes to cellular differentiation processes, such as myogenesis and adipogenesis, placing SMD at the heart of various physiologically important mechanisms.
Introduction
All organisms maintain homeostasis by dynamically regulating gene expression. The regulation of protein-encoding genes occurs during both the synthesis and the decay of their product messenger (m)RNAs. mRNA formation involves nuclear processes that include 5' capping, the removal of introns if the mRNA derives from splicing, 3'-end formation and, possibly, RNA editing and other co- or post-transcriptional nucleotide modifications. Should any one of these processes occur inaccurately or inefficiently, the transcript will be degraded by quality-control mechanisms that function within nuclei1. Cytoplasmic quality-control pathways include nonsense-mediated mRNA decay (NMD)2,3, non-stop mRNA decay (NSD)4 and no-go decay (NGD)5, each of which depends on mRNA translation. Cellular homeostasis is also achieved by cis-acting regulatory elements that reside within mRNAs – often within mRNA 3'UTRs – and govern cytoplasmic mRNA half-life6,7. Depending on the element, its function may or may not depend on mRNA translation, regulators of which would also affect element function. The importance of mRNA decay to human disease was illustrated nearly 40 years ago with the realization that viral FOS and MYC oncogenes evolved from cellular proto-oncogene counterparts that, during the evolutionary process, lost their destabilizing cis-acting regulatory elements, called AU-rich elements (AREs), so as to acquire potent transforming properties8,9. In fact, many human diseases are due to aberrancies in the regulation of mRNA half-life10–12. Thus, the regulation of mRNA decay has become the subject of intensive research7,13.
There are various pathways of cytoplasmic mRNA decay that are conditionally used to regulate gene expression besides ARE-mediated decay14. These include micro (mi)RNA-mediated mRNA decay15, NMD, which not only serves as a quality-control pathway but also regulates normal gene expression3,7,13,16,17, and Staufen1 (STAU1)-mediated mRNA decay (SMD), which is the topic of this review. To understand how mRNA decay occurs, many studies have focused on identifying not only the cis-acting sequences but also the trans-acting factors that play crucial roles in the various pathways. Some of these studies have been initiated serendipitously, as exemplified by the discovery of SMD. In this review, we will describe our current understanding of how SMD is regulated by specific cis-acting sequences and trans-acting factors. Furthermore, we will discuss known physiological functions of SMD.
SMD MECHANISM
Discovery of SMD
In 2005, Kim and colleagues used human UPF1, a central factor in NMD, as bait in a yeast two-hybrid analysis to search for additional NMD factors. He unexpectedly obtained the double-stranded(ds) RNA-binding protein human STAU1 as prey18. After corroborating that human STAU1 and human UPF1 interact directly when synthesized in and purified from E. coli, and after demonstrating that the two proteins co-immunoprecipitate in a largely RNase A-resistant manner from human HeLa-cell lysates, the idea emerged that STAU1 might function analogously to how an exon-junction complex (EJC) functions during NMD: it could recruit UPF1 to mRNA 3'UTRs so as to trigger mRNA decay when translation terminates upstream of where UPF1 was recruited18. This idea was confirmed by experimentally tethering MS2-tagged STAU1 to an MS2 coat protein-binding site that was situated downstream of a termination codon: tethered STAU1 was shown to trigger mRNA decay in a mechanism that was inhibited by downregulating UPF1 but not other NMD and/or EJC factors18. Furthermore, STAU1 was shown to bind the 3'UTRs of a number of mRNAs so as to decrease their half-lives in a translation- and UPF1-dependent mechanism18,19. Microarray analyses illustrated that conservatively ~1% of the 11,569 HeLa-cell mRNAs examined were upregulated when STAU1 was downregulated, suggesting that SMD constitutes a significant post-transcriptional regulatory pathway19. How this pathway relates to the finding that tethering STAU1 to an mRNA 5'UTR promotes the translation of the tethered mRNA20 is unclear.
Staufen1-binding sites (SBSs)
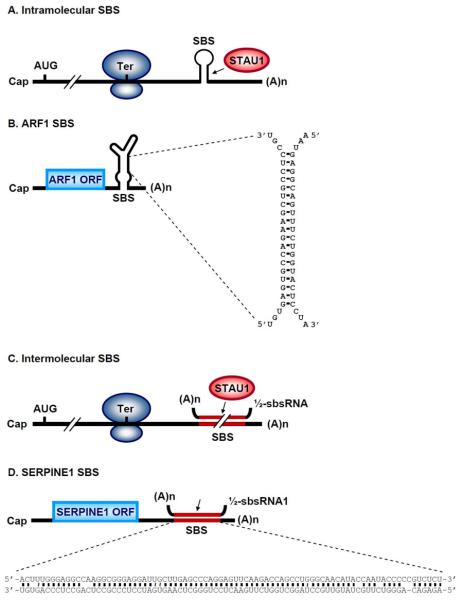
STAU1 contains four dsRNA-binding domains (dsRBDs; Figure 1), of which dsRBD3 and dsRBD4 (but not dsRBD2 or dsRBD5) are responsible for binding dsRNA21. SMD requires the binding of STAU1 to a STAU1-binding site (SBS) that is situated sufficiently downstream of a termination codon in the 3'-untranslated region (3'UTR) of target mRNAs18,19,22, presumably so the footprint of the terminating ribosome does not impinge upon and remove bound STAU1. SBSs may be formed by intramolecular base-pairing within an mRNA 3'UTR (Figure 2A,B) or intermolecular base-pairing between an Alu element within an mRNA 3'UTR and a partially complementary Alu element within one or more long noncoding (lnc)RNAs (Figure 2C,D) that we call ½-sbsRNAs19,22.
Figure 1. Diagrams of human STAU1 and STAU2 isoforms.
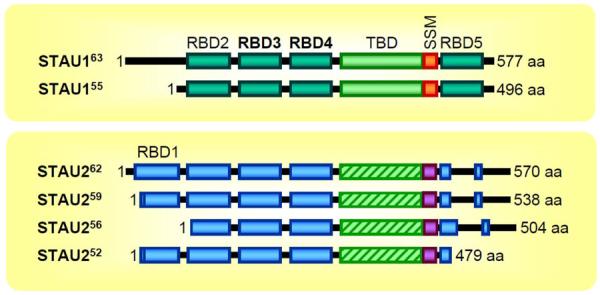
Superscript numbers approximate the mass (kD) of the isoform; boxes represent functional and/or structural domains. 1, N-terminal amino acid; RBD, dsRNA-binding domain (bold RBDs are able to bind dsRNA); TBD, tubulin-binding domain (where hatched STAU2 TBDs are ~18% identical to STAU1 TBDs); SSM, Staufen-swapping motif; aa, amino acids.
Figure 2. Features of STAU1-binding sites.
A. A STAU1-binding site (SBS) that triggers SMD can be a stem-loop structure formed by intramolecular base-paring within an mRNA 3'UTR. (A)n, mRNA poly(A) tail. B. The SBS within ARF1 mRNA provides an example of intramolecular base-paired SBS. Enlarged sequences include the 19-bp stem that largely defines the STAU1-binding sequence. ORF, open reading frame. C. Alternatively, an SBS can be formed by intermolecular base-pairing between a 3'UTR Alu element and a partially complementary Alu element within a long non-coding (lnc) RNA, which we call a ½sbs-RNA. D. Predicted intermolecular base-pairing between the 3'UTR Alu element within SERPINE1 mRNA and the Alu element within ½-sbsRNA1 that forms a functional SBS. Cap, 5' mRNA cap structure, AUG, translation initiation codon; Ter, translation termination codon; (A)n, poly(A) tail.
Intramolecular SBSs
During initial studies of SMD18, deletion mapping localized the SBS within mRNA encoding ADP ribosylation factor (ARF)1 to nucleotides 1–300 of the ARF1 3'UTR (where nucleotide 1 is defined as the nucleotide immediately after the normal termination codon). Subsequently, Luc Furic working in the DesGroseillers lab and Yoon Ki Kim working in the Maquat lab, aided by computational analyses performed by Marc Parisien in the Major lab, further delimited the SBS to a 19-base-pair stem formed by nucleotides 75–93 and 212–194 having a 100-nucleotide apex19 (Figure 2B). A series of point-mutations and deletions showed that this stem, the structure of which is conserved among Homo sapiens, Rattus norvegicus and Mus musculus, forms an SBS by virtue of its double-stranded nature and not its explicit primary sequence19. However, a 19-base-pair stem could not be found in the 3'UTRs of other known SMD targets, leaving unanswered the issue of how much of STAU1 binding to dsRNA is sequence-specific vs. recognition of a higher-ordered structure.
Intermolecular SBSs
Subsequently, Gong and Maquat22 noticed that SMD targets encoding either serpin peptidase inhibitor, clade E (nexin, plasminogen activator inhibitor type 1), member 1 (SERPINE1) or sosondowah ankyrin repeat domain family member C (SOWAHC, also known as FLJ21870, which will be used here) harbor a single Alu element in their 3'UTR. Alu elements are short interspersed elements, or SINES, that are particular to primate genomes and present at ~1.4 million copies23. Computational analyses demonstrated that ~4% of HeLa-cell mRNAs contain one or more 3'UTR Alu elements. However, ~13% of HeLa-cell mRNAs that are up-regulated upon STAU1 down-regulation using small interfering (si)RNA, and thus are putative SMD targets, contain a single Alu element. Thus mRNAs containing a 3'UTR Alu element are enriched in SMD targets. Computational analyses also identified ~350 long-noncoding RNAs (lncRNAs) that harbor one or more Alu elements.
The finding that all Alu element-containing lncRNAs that were examined proved to be largely cytoplasmic and polyadenylated raised the intriguing possibility that SBSs could form by intermolecular base-pairing between a 3'UTR Alu element and the partially complementary Alu element of a lncRNA. For example, lncRNA_AF087999 (for which the NCBI accession number is AF087999) was predicted to form a partially complementary duplex with the Alu element in the SERPINE1 or SOWAHC 3'UTR22 (Figure 2D). When the level of this lncRNA, now designated ½-sbsRNA1, was down-regulated using siRNA, the levels of SERPINE1 and SOWAHC mRNAs were up-regulated, phenocopying the effect of STAU1 or UPF1 depletion. Moreover, ½-sbsRNA1 was shown to co-immunoprecipitate with not only SERPINE1 and SOWAHC mRNAs but also STAU1 and UPF1. Depletion of STAU1 reduced the co-immunoprecipitation of SERPINE1 and SOWAHC mRNAs with ½-sbsRNA1, which was shown to depend strictly on their 3'UTR Alu element. This result indicates that the Alu element within ½-sbsRNA1 base-pairs with the Alu element in the 3'UTR of each mRNA in a way that is stabilized by STAU1 binding. Data reveal that individual ½-sbsRNAs regulate a subset of SMD substrates and that individual SMD targets can be regulated by a subset of ½-sbsRNAs, making for a complicated network of post-transcriptional control. Notably, however, not all mRNAs harboring a 3'UTR Alu element are targeted for SMD even in the presence of a complementary ½-sbsRNA, at least in part because messenger ribonucleoprotein (mRNP) structure prevents STAU1 binding24.
STAU1 and STAU2
STAU1, like its paralog STAU2, contains multiple dsRBDs. The STAU1 gene encodes at least two isoforms (63 kDa and 55 kDa), whereas the STAU2 gene encodes at least four isoforms (62 kDa, 59 kDa, 56 kDa and 52 kDa). All isoforms are generated by alternative pre-mRNA splicing and/or polyadenylation (Figure 1). STAU2 and STAU1 are ~50% identical27, and dsRBDs 3 and 4, which are the only dsRBDs that bind dsRNA in STAU1, are 78% and 81% identical, respectively21,28,29 (Figure 1). However, dsRBD2 and the tubulin-binding domain (TBD) of STAU1 and STAU2 are only 48% and 18% identical, respectively. Relative to STAU1, STAU2 has an additional dsRBD, dsRBD1, and only a partial dsRBD530. Even though dsRBDs 3 and 4 are highly similar between STAU1 and STAU2, Furic and colleagues reported that the mRNA content of STAU1 mRNPs and STAU2 mRNPs is largely distinct, manifesting only ~30% overlap27. Moreover, the relative ratio of STAU1 and STAU2 abundance varies among tissues and cells, e.g., STAU2 is enriched in brain and heart29,31. Additionally, there are paralog-specifc RNA granules in the distal dendrites of rat hippocampal neurons29. These differences may explain how the two paralogs can have distinct cellular roles. As examples, STAU1 functions in the late phase of forskolin-induced long-term potentiation32–34 and STAU2 does not, whereas STAU2 functions in metabotropic glutamate receptor-mediated long-term depression34 and STAU1 does not.
However, STAU1 and STAU2 also share a number of biological functions, including the microtubule-dependent transport of RNAs35,36 and the formation and maintenance of the dendritic spines of hippocampal neurons32–34,37,38. Additionally, STAU1 and STAU2 function together in SMD. Both bind to the 3'UTRs of all tested SBSs, including those of mRNAs encoding ARF1, SERPINE1, SOWAHC, growth-associated protein (GAP)43, and v-jun sarcoma virus 17 oncogene homolog (c-JUN)39. Nevertheless, using HeLa-cell lysates, the two paralogs have distinct properties39. Compared to STAU1, STAU2 co-immunoprecipitates ~2-to-5-fold more of each of' the five mRNAs listed above and ~10-fold more UPF139. Furthermore, the co-immunoprecipitation of STAU2 with UPF1 is essentially insensitive to RNase A, whereas the co-immunoprecipitation of STAU1 with UPF1 is only partially insensitive to RNase A39,40. Consistent with these results, in vitro binding assays using purified recombinant proteins indicate that STAU2 interacts with UPF1 ~10-fold more efficiently than does STAU139. Since UPF1 binds to STAU1 amino acids extending from the TBD to just upstream of dsRBD541, variations between STAU1 and STAU2 in this region may explain their different affinities for UPF1. dsRBD2 and/or dsRBD3 of STAU2 have additionally been reported to be responsible for UPF1 binding42, possibly offering another reason for the discrepancy. Despite these differences, using HeLa cells, in which STAU1 and STAU2 are comparably abundant, (i) down-regulating either paralog to the same extent inhibits SMD to the same extent, and (ii) tethering a comparable amount of MS2-tagged STAU1 or MS2-tagged STAU2 downstream of a termination codon triggers SMD to the same extent39 (Figure 3; see below for interpretation). It is unclear39 why Monshausen et al.31 found that tethering rat STAU252 downstream of a termination codon of β-globin 4boxB reporter mRNA reduced reporter mRNA abundance by only ~25% in HeLa cells, and Miki et al. (2011)42 found that Renilla LUC activity from an RLUC-8xMS2bs reporter mRNA was reduced by only ~10% or ~30% when, respectively, MS2-tagged rat STAU259 or MS2-tagged rat STAU155 was tethered reporter mRNA, compared to when MS2 alone was expressed. Since there is no reason to believe that rat STAU isoforms would be functionally distinct from human STAU1 and STAU2 isoforms given their relatively high (~92% and ~93%) identity, respectively39, we conclude that STAU1 and STAU2 indeed have redundant roles in SMD, SMD is better defined as STAU-mediated mRNA decay rather than as STAU1-mediated mRNA decay.
Figure 3. Tethering transiently expressed MS2-tagged STAU1 or MS2-tagged STAU2 to an mRNA 3'UTR triggers SMD in a mechanism that involves interactions with cellular STAU1 and/or cellular STAU2.
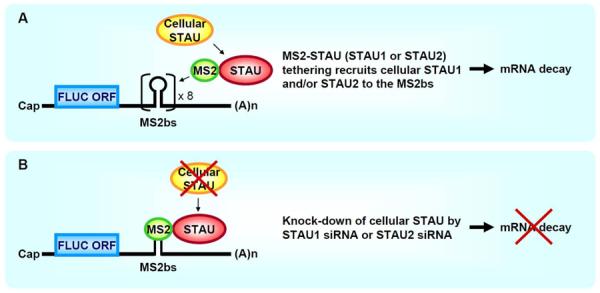
A. Tethering of STAU1 or STAU2, fused to the viral MS2 coat protein (MS2), to eight tandem repeats of the MS2 coat protein-binding site (MS2bs) inserted into the 3'UTR of a firefly luciferase (FLUC) reporter mRNA triggers mRNA decay by recruiting cellular STAU1 and/or STAU2. ORF, open translational reading frame; (A)n, poly(A). B. Knock-down of cellular STAU1 or STAU2 using STAU1 siRNA or STAU2 siRNA, respectively, inhibits mRNA decay brought about by tethering MS2-STAU1 or MS2-STAU2.
STAU1 and STAU2 self-associate and associate with one another
The functional activation of a number of dsRNA-binding proteins requires that they self-associate or associate with other dsRNA-binding proteins. As examples, human immunodeficiency virus trans-activation response RNA-binding protein (TRBP) binds directly to Dicer, which stabilizes Dicer, facilitates Dicer binding to pre-micro(mi)RNA, and thereby leads to miRNA processing, Argonaute 2 binding and miRNA-mediated mRNA decay; protein activator of protein kinase (PK)R (PACT) binds interferon-induced PKR so as to activate the PKR-mediated phosphorylation of eIF2α that inhibits translation; and adenosine deaminase acting on RNA (ADAR)1 or ADAR2 homodimerization is essential for its converting adenosines to inosines within dsRNA43–46.
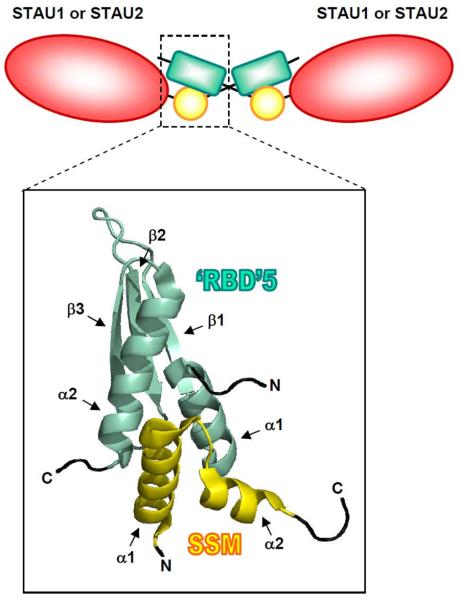
Recently, STAU1 was shown to self-associate39,47,48 (Figure 4). Martel and colleagues47 demonstrated that differentially tagged STAU1 proteins co-immunoprecipitate in HEK293T-cell lysates and are sufficiently close in intact HEK293T cells to transfer bioluminescence resonance energy via dsRBDs 2–dsRBD2 and dsRBD5–dsRBD5 interactions. Subsequent crystallographic studies revealed that the stronger of the two interactions – that involving dsRBD5 – requires a new motif, named the Staufen-swapping motif (SSM), which resides in the region between the TBD and dsRBD548 (Figure 1). The two α-helices of the SSM of one STAU1 molecule interdigitate the two α-helices of the dsRBD5 of another STAU1 molecule to form a STAU1 homodimer, although only the first α-helix of dsRBD5 is required48 (Figure 4). Deleting SSM or generating point-mutations in both the SSM and dsRBD5 that disrupt STAU1 dimerization abolishes not only STAU1 self-association but also the interaction of STAU1 with UPF1 and, thus, SMD48. It follows that STAU1 self-association may serve as a molecular regulator of SMD47. While STAU1 self-association was reduced by point-mutations in dsRBD3 or dsRBD4 that reduced STAU1 binding to dsRNA47, mutations in the SSM or dsRBD5 that disrupt dimerization did not reduce STAU1 binding to dsRNA48. Thus, even though STAU1 self-association is largely resistant to RNase A treatment and occurs in vitro, dsRNA may stabilize STAU1 self-association39,47.
Figure 4. Model for homodimeric and heterodimeric interactions between STAU1 and/or STAU2 via domain swapping.
Simplified diagram of homo- or hetero-dimerization of STAU molecules via domain-swapping between the STAU-swapping motif (SSM) of one molecule and the dsRBD5 of another molecule. Yellow circle, SSM; green rectangle, dsRBD5; red oval; rest of STAU molecule. Enlarged box shows the crystal structure of the interaction, detailing that the two α-helices of the SSM of one STAU molecule interdigitate with the two α-helices of the dsRBD5 of another STAU molecule. Notably, the smallest isoform of STAU2 lacks the C-terminal α-helix of dsRBD5, and only the N-terminal α-helix of dsRBD5 is necessary for the interaction.
Co-immunoprecipitation and in vitro interaction studies revealed that STAU2 likewise self-associates and associates with STAU1 via the same SSM–dsRBD5 α-helical interdigitations that in the dsRBD5 of the smallest isoform of STAU2 are confined to its one and only α-helix39,48. Homo and heterodimers (if not multimers) of STAU1 and STAU2 function in SMD based on the finding that SMD as a result of tethering either MS2-STAU1 or MS2-STAU2 downstream of a termination codon is inhibited by either STAU1 siRNA or STAU2 siRNA39.
STAU1 and STAU2 enhance UPF1 helicase activity without activating UPF1 ATPase activity
The SMD and NMD pathways share the steps of UPF1 phosphorylation49, which has been shown in studies of NMD to trigger translational repression50, mRNA decay51,52 and, most likely, many of the factors that function in these steps. The ATP-dependent RNA helicase activity of UPF1 that functions during NMD53,54 to remodel mRNP prior to its decay55 also appears to function during SMD. In support of this, expressing in HEK293T cells UPF1(G495R,G497E)50, which harbors point-mutations that disrupt ATP binding and therefore helicase activity39,56,57, inhibited the SMD of those reporter and cellular targets that were tested39. Furthermore, in vitro helicase assays of UPF1 demonstrated that both STAU paralogs enhance UPF1 helicase activity without enhancing UPF1 ATPase activity in a mechanism that requires ATP39. While RNA-binding studies of UPF1 in the presence of STAU1 were complicated because STAU1 is not fully soluble under conditions that are conducive to footprinting, STAU2 was found to increase the length of single-stranded RNA that was protected by UPF1 from 11–12 nucleotides to 12–13 nucleotides from cleavage by RNases A + T1, and this length is in-between the length protected by ATP-free UPF1 and UPF1 that is bound by either ATP or an ATP-hydrolytic intermediate39. We conclude from these and related experiments that UPF1 helicase activity is promoted by STAU2 (and based on its conserved function, STAU1), requires ATP binding and/or the energy released from ATP hydrolysis, and changes the conformation of UPF1 to mimic the ATP-bound activated conformation of UPF1 (Figure 5). The ability of STAU paralogs to promote UPF1 helicase activity to the same extent even though STAU2 binds RNA and UPF1 more tenaciously than does STAU1 also explains the comparable abilities of STAU1 and STAU2, when present at the same cellular abundance, to promote SMD to the same extent39. This may be accomplished by STAU1 and STAU2 interacting differently with the relatively large (~129- amino acid) cysteine-and histidine-rich (CH) domain of UPF1, especially since the TBDs that each STAU paralog uses to interact with UPF1 are very different in sequence. Thus, the paralogs may bind to over-lapping or different regions of the CH domain with different affinities, yet still activate UPF1 helicase activity in a binary on-off manner. Provided that STAU1 and STAU2 both bind tightly enough to overcome the activation threshold for UPF1, it is possible for them to manifest different UPF1 binding efficiencies but the same degree of influence on the efficiency of SMD.
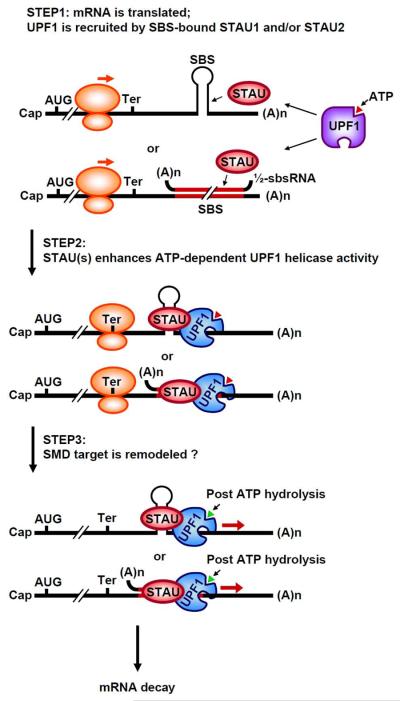
Figure 5. Model for the mechanism of SMD.
STAU1 and/or STAU2, either individually (as shown for simplicity) or more likely as homo- or hetero-dimers if not multimers, bind to a 3'UTR SBS that can be formed by either intramolecular or intermolecular base-pairing19,22. During protein synthesis, when ribosomes reach a termination codon that resides sufficiently upstream of an SBS so that bound STAU1 and/or STAU2 is not removed by the terminating ribosome, UPF1, which is recruited by SBS-bound STAU(s), is activated. Taking cues from the related pathway, NMD, translation termination may facilitate the loading of UPF1 onto the mRNA 3'UTR, as may SBS-bound STAU193. Activation of mRNA decay may involve UPF1 phosphorylation, possibly by SMG1: it has been reported that UPF1 phosphorylation enhances the co-IP of UPF1 with STAU140. STAU promotes a conformational change in UPF1, thereby enhancing UPF1 helicase activity without concomitantly enhancing UPF1 ATPase activity39. That noted, however, ATP hydrolysis is required for helicase activity, which may be required for mRNP remodeling, as is the case for NMD55. Cap, 5' mRNA cap structure; SBS, STAU-binding site; Ter, termination codon; (A)n, poly(A) tail, orange ovals, 80S translationally active ribosome; purple UPF1, inactive form; blue UPF1, conformationally changed and active UPF1.
SMD vs. NMD
Given that UPF1 plays a crucial role in both SMD and NMD, and that STAU1 and the UPF2 NMD factor bind overlapping regions of the UPF1 CH domain, it follows that SMD and NMD are competitive pathways41 (Figure 6). As evidence for this, down-regulating the abundance of cellular STAU1 increases the co-immunoprecipitation of UPF2 and UPF1 and augments the efficiency of NMD, whereas down-regulating the abundance of cellular UPF2 increases the co-immunoprecipitation of STAU1 and UPF1 and augments the efficiency of SMD41. Recently, Cho and colleagues found that the proline-rich nuclear receptor coregulatory protein 2 (PNRC2) functions in both SMD and NMD40,58. PNRC2 interacts with hyper-phosphorylated UPF1 ~5-fold more efficiently than hypo-phosphorylated UPF1, which constitutes the bulk of cellular UPF1. Thus, PNRC2, together with the relative abundance and activities of STAU1, if not also STAU2, and UPF2, may serve as another rheostat for the balance between SMD and NMD (see below).
Figure 6. Model for competition between SMD and NMD.
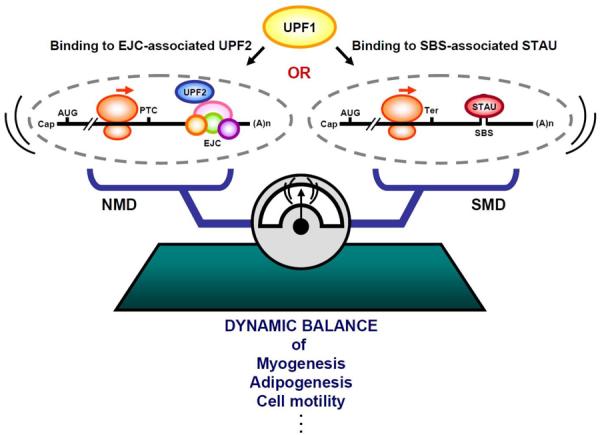
SMD and NMD compete with one another since STAU1 and STAU2, which function in SMD, compete with UPF2, which functions in NMD, for binding to UPF1, which functions in both pathways. Notably, the STAU paralogs function when bound to an intramolecular (shown) or intermolecular (not shown) STAU-binding site (SBS) that is situated sufficiently downstream of a termination codon that is usually the normal termination codon (Ter). In contrast, UPF2 generally functions as a constituent of an exon-junction complex (EJC) that is situated sufficiently downstream of a termination codon that is usually a premature termination codon (PTC). AUG, translation initiation codon; (A)n, poly(A) tail.
Functional Significance of SMD
Myogenesis
Gong and colleagues41 observed that the cellular abundance of STAU1 decreases during the differentiation of mouse skeletal C2C12 myoblasts (MBs) to multinucleated myotubes (MTs). However, the cellular abundance of UPF2 was reduced even more than the reduction of the cellular abundance of STAU1. As a consequence, the co-immunoprecipitation of STAU1 and UPF1 was increased, whereas the co-immunoprecipitation of UPF2 and UPF1 was reduced. Consistent with these findings, the efficiency of SMD was increased, while the efficiency of NMD was decreased. This makes physiological sense considering that paired-box 3 (PAX3) mRNA is an SMD target encoding a transcription factor that inhibits myogenesis59, whereas myogenin mRNA is an NMD target (due to its upstream open reading frame) encoding a basic helix-loop-helix transcription factor that promotes myogenesis60. For these and other reasons49, the changing balance in SMD relative to NMD during myogenesis is critical to this differentiation process. Notably, an increase or decrease in the efficiency of a particular mRNA decay pathway is often coupled to a decrease or an increase, respectively, in the transcriptional efficiency of the gene from which the mRNA derives. For example, downregulating STAU1 in C2C12 MBs using siRNA not only decreased the efficiency of PAX3 SMD and increased the efficiency of myogenin NMD but also decreased the production of PAX3 pre-mRNA and increased the production of myogenin pre-mRNA41. Consistent with this, the transition from MBs to MTs involves PAX3 mRNA decay and a decrease in the production of PAX3 pre-mRNA41.
Adipogenesis
SMD is also involved in adipogenesis, a cellular differentiation process whereby pre-adiopocytes develop into adipocytes. Adipocytes, a component of adipose tissue, play crucial roles in maintaining energy, metabolic homeostasis, the immune response and blood-pressure control61,62. Impairment of the adipose compartment causes metabolic diseases such as obesity, type 2 diabetes, cancer cachexia, and lipodystrophies62. Cho and colleagues40 found that the abundance of STAU1 and PNRC2, which is itself connected to energy homeostasis and obesity63, are increased during the conversion of 3T3-L1 pre-adipocytes to adipocytes40. For example, male PNRC2−/− null mice are lean and resistant to high-fat diet-induced obesity because they consume more oxygen, produce more heat, and have reduced adipose mass and lower levels of leptin than do normal mice63. PNRC2 associates with STAU1 via hyperphosphorylated UPF158,64, and down-regulating PNRC2 up-regulates the abundance of SERPINE1 mRNA, which was used as an assay for the efficiency of SMD40. Thus, as during myogenesis41, the efficiency of SMD increases while the efficiency of NMD decreases40. SMD targets Krüppel-like factor 2 (KLF2) mRNA40. KLF2 encodes an anti-adipogenic factor40,61,65 that induces caveolin-1, the main component of caveolae in the plasma membrane66. KLF2 also inhibits peroxisome proliferator-activated receptor γ (PPARγ), a transcription factor that is essential for lipid uptake and adipogenesis40,67. Thus, down-regulating STAU1, PNRC2 or UPF1 delays the induction of PPARγ, and therefore inhibits adipogenesis. These results, plus the increased interaction of STAU1 with hyper-phosphorylated UPF1 during adipogenesis40, which is essential for efficient SMD39, indicate that SMD facilitates adipogenesis by down-regulating anti-adipogenic factors.
Cutanenous wound-healing
SERPINE1 promotes cutaneous wound-healing68,69 by promoting migration while inhibiting proliferation70,71. Wound-healing is an intricate cellular and molecular process that restores skin integrity after injury and involves inflammation, cell migration, angiogenesis, matrix remodeling and re-epithelialization72,73. The level of SERPINE1 mRNA increases rapidly in response to experimentally “wounding” T2 renal-epithelial cells, which is accomplished by generating a denuded wound-track or “scratch” in a tissue-culture plate of adherent and nearly confluent cells; furthermore, down-regulating the cellular abundance of SERPINE1 impairs wound-healing as monitored by the migration rate of T2 cells into the denuded wound-track69,74.
Human HaCaT keratinocytes also provide an in vitro model for cutaneous wound-healing75. In experiments that examined the possibility that SMD inhibits cell motility and migration by degrading SERPINE1 mRNA, ½-sbsRNA1, which down-regulates SERPINE1 mRNA, was found to inhibit HaCaT-cell wound-healing after scrape-induced injury22. Notably, ½-sbsRNA1 also targets RAB11 family interacting protein (RAB11FIP)1 mRNA for SMD22. Overexpressing RAB11FIP1 enhances the motility of non-tumorigenic epithelial MCF10A cells76. As expected, down-regulating SERPINE1 or RAB11FIP1 delayed HaCaT-cell wound-healing22. It is remarkable that down-regulating the single lncRNA ½-sbsRNA1, which is not the only ½-sbsRNA that is computationally predicted to base-pair with either SERPINE1 mRNA or RAB11FIP1 mRNA, induces keratinocyte motility. This finding likely has relevance to other human-cell motility pathways, should ½-sbsRNA1 be expressed, including axonal regeneration and the epithelial-to mesenchymal transition that accompanies metastatic cancers, and it illustrates the physiological relevance of lncRNAs such as of ½-sbsRNAs.
Conclusion
SMD is a translation-dependent mRNA decay pathway that is mediated by the dsRNA-binding paralogs STAU1 and/or STAU2 (Figure 1). Each binds to an SBS formed by intramolecular or intermolecular base-pairing (Figure 2) either individually or as homo- or hetero-dimers, if not multimers (Figure 4). Should an SBS reside downstream of a translation termination codon, e.g., within an mRNA 3'UTR, then STAU paralog binding to the SBS recruits the ATP-dependent RNA helicase UPF1 to trigger mRNA decay (Figure 5). Since STAU binding to UPF1 competes with binding of the UPF2 NMD factor to UPF1, SMD and NMD are competitive pathways (Figure 6).
Studies of SMD have illustrated its importance to cellular homeostasis, the current data for which are undoubtedly just the “tip of the iceberg”. For example, SMD targets the mRNA encoding c-JUN, which forms the AP-1 early-response transcription factor together with c-FOS77,78. Since AP-1 regulates cell proliferation, differentiation, apoptosis, organogenesis and the response to stress79, it follows that SMD does, too. SMD, which generally reduces mRNA abundance by only an order of magnitude, is viewed to “fine-tune” gene expression, as do, e.g., miRNAs80,81. Despite its modest effect on mRNA abundance, however, SMD contributes in critical ways to the diversity of cell-types and their responses to intra- and extra-cellular changes during development, differentiation and the continually variable environmental milieu. As exemplified by studies of wound-healing, even though an mRNA containing a 3'UTR Alu element is targeted for SMD by multiple ½-sbsRNAs that are expressed simultaneously, down-regulating a single ½-sbsRNA, ½-sbsRNA1, is sufficient to promote keratinocyte motility. This is in part because multiple mRNAs that promote wound-healing are targeted for SMD by ½-sbsRNA182. Additionally, as is generally true for post-transcriptional regulatory pathways, the effect on half-life is “amplified” by concomitant changes in transcription of the gene encoding the targeted transcript6.
Both STAU1 and STAU2 are involved in mRNA transport along microtubules35,36,83,84. mRNA transport leads to the asymmetric localization of STAU-bound mRNAs, thereby regulating their cellular sites of translation85. The localized translation of specific mRNAs plays crucial roles in many cellular processes, including synaptic plasticity, learning and memory, migration during development, cell motility, and asymmetric cell division86–90. Thus, provided an SMD target is translationally repressed until it is transported to a particular cellular locale, SMD would be expected to occur locally. The localization of SMD would provide a burst in protein synthesis immediately followed by mRNA decay akin to the burst in the NMD target that encodes the activity-regulated cytoskeleton-associated protein activity-regulated cytoskeleton-associated protein (ARC)91. Future studies will focus on dissecting the complex network of post-transcriptional events that comprise SMD and how they interface not only with NMD but with the many additional post-transcriptional regulatory pathways. These include post-translational modifications that target STAU1 and STAU2, both of which are known to undergo phosphorylation in ways that could alter STAU multimerization, RNA binding, UPF1 binding and/or the activation of UPF1 helicase activity. Pertinent post-transcriptional regulatory pathways also include those mediating conditional changes in the absolute and relative abundance of STAU1, STAU2 and ½-sbsRNAs, which is known to vary among different cell types22. Given that conservatively ~54% of transcripts are thought to undergo alternative polyadenylation92, consideration should also be given to this process, which could regulate when or not an mRNA isoform maintains an SBS.
Sidebar title: How Staufen protein got its name.
The human STAU paralogs were named after their ortholog in Drosophila melanogaster, which was defined as a maternal-effect loci in a genetic screen undertaken by Trudi Schupbach and Eric Wieschaus25. Although Staufen derives from a strictly maternal gene that is expressed at high levels in the adult female germline, especially in the embryo central nervous system, it is not required for survival of the mother. Homozygotes expressing no Staufen are fertile but give rise to defective embryos that do not properly localize those mRNAs that bind Staufen during oogenesis26. So, where does the name Staufen derive?
There were two German noble families named Staufen. One lived in the area around Freiburg, around which there is still a small and beautiful town by the name of Staufen, but it has nothing to do with the fly mutant after which the Staufen protein was named. The second family was of higher nobility, more often called `Hohenstaufen' (high staufen), after the castle Hohenstaufen that is situated close to the town of Göppingen, east of Stuttgart. There were three German emperors from this family. The first was Friedrich I. von Hohenstaufen, aka `Barbarossa' (red beard). He was elected King of Germany in 1152, crowned emperor of the `Holy Roman Empire of German Nation' in 1155 and died in 1190. The second was Friedrich's son, Heinrich IV, who was made king of Germany in 1169 at the age of four and became emperor after his father's death in 1191. Heinrich died in 1197. Heinrich's son Philipp von Schwaben was King of Germany from 1198 until 1208, but was never crowned emperor. The last was Friedrich II. He was born in 1194, elected king in1211 and crowned emperor in 1220. He died in 1250. All of his male offspring died/were killed, ending the family and the reason for naming the sterile fly mutant Staufen – after what some consider is the most famous family of German emperors.
Acknowledgments
We thank Max Popp and Mike Gleghorn for helpful comments on the manuscript, Mike Gleghorn for the figure showing the SSM–dsRBD5 crystal structure, and Elmar Wahle for sharing his prodigious knowledge of German history. Work on SMD in the Maquat-lab is funded by NIH R01 GM074593 to L.E.M, and E.P. is supported by AHA Founders Affiliate Postdoctoral Fellowship 11POST 7860051.
References
- 1.Schmid M, Jensen TH. Nuclear quality control of RNA polymerase II transcripts. Wiley Interdiscip Rev RNA. 2010;1:474–485. doi: 10.1002/wrna.24. [DOI] [PubMed] [Google Scholar]
- 2.Maquat LE, Tarn WY, Isken O. The pioneer round of translation: features and functions. Cell. 2010;142:368–374. doi: 10.1016/j.cell.2010.07.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Huang L, Wilkinson MF. Regulation of nonsense-mediated mRNA decay. Wiley Interdiscip Rev RNA. 2012;3:807–828. doi: 10.1002/wrna.1137. [DOI] [PubMed] [Google Scholar]
- 4.Hoshino S. Mechanism of the initiation of mRNA decay: role of eRF3 family G proteins. Wiley Interdiscip Rev RNA. 2012;3:743–757. doi: 10.1002/wrna.1133. [DOI] [PubMed] [Google Scholar]
- 5.Harigaya Y, Parker R. No-go decay: a quality control mechanism for RNA in translation. Wiley Interdiscip Rev RNA. 2010;1:132–141. doi: 10.1002/wrna.17. [DOI] [PubMed] [Google Scholar]
- 6.Ghosh S, Jacobson A. RNA decay modulates gene expression and controls its fidelity. Wiley Interdiscip Rev RNA. 2010;1:351–361. doi: 10.1002/wrna.25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Schoenberg DR, Maquat LE. Regulation of cytoplasmic mRNA decay. Nat Rev Genet. 2012;13:246–259. doi: 10.1038/nrg3160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Meijlink F, Curran T, Miller AD, Verma IM. Removal of a 67-base-pair sequence in the noncoding region of protooncogene fos converts it to a transforming gene. Proc Natl Acad Sci U S A. 1985;82:4987–4991. doi: 10.1073/pnas.82.15.4987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Aghib DF, Bishop JM, Ottolenghi S, Guerrasio A, Serra A, Saglio G. A 3' truncation of MYC caused by chromosomal translocation in a human T-cell leukemia increases mRNA stability. Oncogene. 1990;5:707–711. [PubMed] [Google Scholar]
- 10.Benjamin D, Moroni C. mRNA stability and cancer: an emerging link? Expert Opin Biol Ther. 2007;7:1515–1529. doi: 10.1517/14712598.7.10.1515. [DOI] [PubMed] [Google Scholar]
- 11.Conne B, Stutz A, Vassalli JD. The 3' untranslated region of messenger RNA: A molecular `hotspot' for pathology? Nat Med. 2000;6:637–641. doi: 10.1038/76211. [DOI] [PubMed] [Google Scholar]
- 12.Steinman RA. mRNA stability control: a clandestine force in normal and malignant hematopoiesis. Leukemia. 2007;21:1158–1171. doi: 10.1038/sj.leu.2404656. [DOI] [PubMed] [Google Scholar]
- 13.Reznik B, Lykke-Andersen J. Regulated and quality-control mRNA turnover pathways in eukaryotes. Biochem Soc Trans. 2010;38:1506–1510. doi: 10.1042/BST0381506. [DOI] [PubMed] [Google Scholar]
- 14.von Roretz C, Gallouzi IE. Decoding ARE-mediated decay: is microRNA part of the equation? J Cell Biol. 2008;181:189–194. doi: 10.1083/jcb.200712054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Fabian MR, Sonenberg N. The mechanics of miRNA-mediated gene silencing: a look under the hood of miRISC. Nat Struct Mol Biol. 2012;19:586–593. doi: 10.1038/nsmb.2296. [DOI] [PubMed] [Google Scholar]
- 16.Hwang J, Maquat LE. Nonsense-mediated mRNA decay (NMD) in animal embryogenesis: to die or not to die, that is the question. Curr Opin Genet Dev. 2011;21:422–430. doi: 10.1016/j.gde.2011.03.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Nicholson P, Mühlemann O. Cutting the nonsense: the degradation of PTC-containing mRNAs. Biochem Soc Trans. 2010;38:1615–1620. doi: 10.1042/BST0381615. [DOI] [PubMed] [Google Scholar]
- 18.Kim YK, Furic L, Desgroseillers L, Maquat LE. Mammalian Staufen1 recruits Upf1 to specific mRNA 3'UTRs so as to elicit mRNA decay. Cell. 2005;120:195–208. doi: 10.1016/j.cell.2004.11.050. [DOI] [PubMed] [Google Scholar]
- 19.Kim YK, Furic L, Parisien M, Major F, DesGroseillers L, Maquat LE. Staufen1 regulates diverse classes of mammalian transcripts. EMBO J. 2007;26:2670–2681. doi: 10.1038/sj.emboj.7601712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Dugré-Brisson S, Elvira G, Boulay K, Chatel-Chaix L, Mouland AJ, DesGroseillers L. Interaction of Staufen1 with the 5' end of mRNA facilitates translation of these RNAs. Nucleic Acids Res. 2005;33:4797–4812. doi: 10.1093/nar/gki794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Wickham L, Duchaîne T, Luo M, Nabi IR, DesGroseillers L. Mammalian staufen is a double-stranded-RNA- and tubulin-binding protein which localizes to the rough endoplasmic reticulum. Mol Cell Biol. 1999;19:2220–2230. doi: 10.1128/mcb.19.3.2220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Gong C, Maquat LE. lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3' UTRs via Alu elements. Nature. 2011;470:284–288. doi: 10.1038/nature09701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Batzer MA, Deininger PL. Alu repeats and human genomic diversity. Nat Rev Genet. 2002;3:370–379. doi: 10.1038/nrg798. [DOI] [PubMed] [Google Scholar]
- 24.Gong C, Popp MW, Maquat LE. Biochemical analysis of long non-coding RNA-containing ribonucleoprotein complexes. Methods. 2012;58:88–93. doi: 10.1016/j.ymeth.2012.06.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Schupbach T, Wieschaus E. Germline autonomy of maternal-effect mutations altering the embryonic body pattern of Drosophila. Dev Biol. 1986;113:443–448. doi: 10.1016/0012-1606(86)90179-x. [DOI] [PubMed] [Google Scholar]
- 26.Jansen RP, Niessing D. Assembly of mRNA-protein complexes for directional mRNA transport in eukaryotes--an overview. Curr Protein Pept Sci. 2012;13:284–293. doi: 10.2174/138920312801619493. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Furic L, Maher-Laporte M, DesGroseillers L. A genome-wide approach identifies distinct but overlapping subsets of cellular mRNAs associated with Staufen1- and Staufen2-containing ribonucleoprotein complexes. RNA. 2008;14:324–335. doi: 10.1261/rna.720308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Buchner G, Bassi MT, Andolfi G, Ballabio A, Franco B. Identification of a novel homolog of the Drosophila staufen protein in the chromosome 8q13#x2013;q21.1 region. Genomics. 1999;62:113–118. doi: 10.1006/geno.1999.6015. [DOI] [PubMed] [Google Scholar]
- 29.Duchaîne TF, Hemraj I, Furic L, Deitinghoff A, Kiebler MA, DesGroseillers L. Staufen2 isoforms localize to the somatodendritic domain of neurons and interact with different organelles. J Cell Sci. 2002;115:3285–3295. doi: 10.1242/jcs.115.16.3285. [DOI] [PubMed] [Google Scholar]
- 30.Allison R, Czaplinski K, Git A, Adegbenro E, Stennard F, Houliston E, Standart N. Two distinct Staufen isoforms in Xenopus are vegetally localized during oogenesis. RNA. 2004;10:1751–1763. doi: 10.1261/rna.7450204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Monshausen M, Gehring NH, Kosik KS. The mammalian RNA-binding protein Staufen2 links nuclear and cytoplasmic RNA processing pathways in neurons. Neuromolecular Med. 2004;6:127–144. doi: 10.1385/NMM:6:2-3:127. [DOI] [PubMed] [Google Scholar]
- 32.Lebeau G, Maher-Laporte M, Topolnik L, Laurent CE, Sossin W, Desgroseillers L, Lacaille JC. Staufen1 regulation of protein synthesis-dependent long-term potentiation and synaptic function in hippocampal pyramidal cells. Mol Cell Biol. 2008;28:2896–2907. doi: 10.1128/MCB.01844-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lebeau G, DesGroseillers L, Sossin W, Lacaille JC. mRNA binding protein staufen 1-dependent regulation of pyramidal cell spine morphology via NMDA receptor-mediated synaptic plasticity. Mol Brain. 2011;4:22. doi: 10.1186/1756-6606-4-22. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Lebeau G, Miller LC, Tartas M, McAdam R, Laplante I, Badeaux F, DesGroseillers L, Sossin WS, Lacaille JC. Staufen 2 regulates mGluR long-term depression and Map1b mRNA distribution in hippocampal neurons. Learn Mem. 2011;18:314–326. doi: 10.1101/lm.2100611. [DOI] [PubMed] [Google Scholar]
- 35.Kiebler MA, Hemraj I, Verkade P, Kohrmann M, Fortes P, Marion RM, Ortín J, Dotti CG. The mammalian staufen protein localizes to the somatodendritic domain of cultured hippocampal neurons: implications for its involvement in mRNA transport. J Neurosci. 1999;19:288–297. doi: 10.1523/JNEUROSCI.19-01-00288.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Tang SJ, Meulemans D, Vazquez L, Colaco N, Schuman E. A role for a rat homolog of staufen in the transport of RNA to neuronal dendrites. Neuron. 2001;32:463–475. doi: 10.1016/s0896-6273(01)00493-7. [DOI] [PubMed] [Google Scholar]
- 37.Goetze B, Tuebing F, Xie Y, Dorostkar MM, Thomas S, Pehl U, Boehm S, Macchi P, Kiebler MA. The brain-specific double-stranded RNA-binding protein Staufen2 is required for dendritic spine morphogenesis. J Cell Biol. 2006;172:221–231. doi: 10.1083/jcb.200509035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Vessey JP, Macchi P, Stein JM, Mikl M, Hawker KN, Vogelsang P, Wieczorek K, Vendra G, Riefler J, Tubing F, et al. A loss of function allele for murine Staufen1 leads to impairment of dendritic Staufen1-RNP delivery and dendritic spine morphogenesis. Proc Natl Acad Sci U S A. 2008;105:16374–16379. doi: 10.1073/pnas.0804583105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Park E, Gleghorn ML, Maquat LE. Staufen2 functions in Staufen1-mediated mRNA decay by binding to itself and its paralog and promoting UPF1 helicase but not ATPase activity. Proc Natl Acad Sci U S A. 2013;110:405–412. doi: 10.1073/pnas.1213508110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Cho H, Kim KM, Han S, Choe J, Park SG, Choi SS, Kim YK. Staufen1-Mediated mRNA Decay Functions in Adipogenesis. Mol Cell. 2012;46:495–506. doi: 10.1016/j.molcel.2012.03.009. [DOI] [PubMed] [Google Scholar]
- 41.Gong C, Kim YK, Woeller CF, Tang Y, Maquat LE. SMD and NMD are competitive pathways that contribute to myogenesis: effects on PAX3 and myogenin mRNAs. Genes Dev. 2009;23:54–66. doi: 10.1101/gad.1717309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Miki T, Kamikawa Y, Kurono S, Kaneko Y, Katahira J, Yoneda Y. Cell type-dependent gene regulation by Staufen2 in conjunction with Upf1. BMC Mol Biol. 2011;12:48. doi: 10.1186/1471-2199-12-48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Chendrimada TP, Gregory RI, Kumaraswamy E, Norman J, Cooch N, Nishikura K, Shiekhattar R. TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature. 2005;436:740–744. doi: 10.1038/nature03868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Haase AD, Jaskiewicz L, Zhang H, Laine S, Sack R, Gatignol A, Filipowicz W. TRBP, a regulator of cellular PKR and HIV-1 virus expression, interacts with Dicer and functions in RNA silencing. EMBO Rep. 2005;6:961–967. doi: 10.1038/sj.embor.7400509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Peters GA, Hartmann R, Qin J, Sen GC. Modular structure of PACT: distinct domains for binding and activating PKR. Mol Cell Biol. 2001;21:1908–1920. doi: 10.1128/MCB.21.6.1908-1920.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Valente L, Nishikura K. RNA binding-independent dimerization of adenosine deaminases acting on RNA and dominant negative effects of nonfunctional subunits on dimer functions. J Biol Chem. 2007;282:16054–16061. doi: 10.1074/jbc.M611392200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Martel C, Dugré-Brisson S, Boulay K, Breton B, Lapointe G, Armando S, Trepanier V, Duchaîne T, Bouvier M, Desgroseillers L. Multimerization of Staufen1 in live cells. RNA. 2010;16:585–597. doi: 10.1261/rna.1664210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Gleghorn ML, Gong C, Kielkopf CL, Maquat LE. Staufen1 dimerizes via a conserved motif and a degenerate dsRNA-binding domain to promote mRNA decay. Nat Struct Mol Biol. 2013 doi: 10.1038/nsmb.2528. in press. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Maquat LE, Gong C. Gene expression networks: competing mRNA decay pathways in mammalian cells. Biochem Soc Trans. 2009;37:1287–1292. doi: 10.1042/BST0371287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Isken O, Kim YK, Hosoda N, Mayeur GL, Hershey JW, Maquat LE. Upf1 phosphorylation triggers translational repression during nonsense-mediated mRNA decay. Cell. 2008;133:314–327. doi: 10.1016/j.cell.2008.02.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Kashima I, Yamashita A, Izumi N, Kataoka N, Morishita R, Hoshino S, Ohno M, Dreyfuss G, Ohno S. Binding of a novel SMG-1-Upf1-eRF1-eRF3 complex (SURF) to the exon junction complex triggers Upf1 phosphorylation and nonsense-mediated mRNA decay. Genes Dev. 2006;20:355–367. doi: 10.1101/gad.1389006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Yamashita A, Ohnishi T, Kashima I, Taya Y, Ohno S. Human SMG-1, a novel phosphatidylinositol 3-kinase-related protein kinase, associates with components of the mRNA surveillance complex and is involved in the regulation of nonsense-mediated mRNA decay. Genes Dev. 2001;15:2215–2228. doi: 10.1101/gad.913001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Chamieh H, Ballut L, Bonneau F, Le Hir H. NMD factors UPF2 and UPF3 bridge UPF1 to the exon junction complex and stimulate its RNA helicase activity. Nat Struct Mol Biol. 2008;15:85–93. doi: 10.1038/nsmb1330. [DOI] [PubMed] [Google Scholar]
- 54.Chakrabarti S, Jayachandran U, Bonneau F, Fiorini F, Basquin C, Domcke S, Le Hir H, Conti E. Molecular mechanisms for the RNA-dependent ATPase activity of Upf1 and its regulation by Upf2. Mol Cell. 2011;41:693–703. doi: 10.1016/j.molcel.2011.02.010. [DOI] [PubMed] [Google Scholar]
- 55.Franks TM, Singh G, Lykke-Andersen J. Upf1 ATPase-dependent mRNP disassembly is required for completion of nonsense- mediated mRNA decay. Cell. 2010;143:938–950. doi: 10.1016/j.cell.2010.11.043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Bhattacharya A, Czaplinski K, Trifillis P, He F, Jacobson A, Peltz SW. Characterization of the biochemical properties of the human Upf1 gene product that is involved in nonsense-mediated mRNA decay. RNA. 2000;6:1226–1235. doi: 10.1017/s1355838200000546. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Cheng Z, Muhlrad D, Lim MK, Parker R, Song H. Structural and functional insights into the human Upf1 helicase core. EMBO J. 2007;26:253–264. doi: 10.1038/sj.emboj.7601464. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Cho H, Kim KM, Kim YK. Human proline-rich nuclear receptor coregulatory protein 2 mediates an interaction between mRNA surveillance machinery and decapping complex. Mol Cell. 2009;33:75–86. doi: 10.1016/j.molcel.2008.11.022. [DOI] [PubMed] [Google Scholar]
- 59.Epstein JA, Lam P, Jepeal L, Maas RL, Shapiro DN. Pax3 inhibits myogenic differentiation of cultured myoblast cells. J Biol Chem. 1995;270:11719–11722. doi: 10.1074/jbc.270.20.11719. [DOI] [PubMed] [Google Scholar]
- 60.Wright WE, Sassoon DA, Lin VK. Myogenin, a factor regulating myogenesis, has a domain homologous to MyoD. Cell. 1989;56:607–617. doi: 10.1016/0092-8674(89)90583-7. [DOI] [PubMed] [Google Scholar]
- 61.Rosen ED, MacDougald OA. Adipocyte differentiation from the inside out. Nat Rev Mol Cell Biol. 2006;7:885–896. doi: 10.1038/nrm2066. [DOI] [PubMed] [Google Scholar]
- 62.Galic S, Oakhill JS, Steinberg GR. Adipose tissue as an endocrine organ. Mol Cell Endocrinol. 2010;316:129–139. doi: 10.1016/j.mce.2009.08.018. [DOI] [PubMed] [Google Scholar]
- 63.Zhou D, Shen R, Ye JJ, Li Y, Tsark W, Isbell D, Tso P, Chen S. Nuclear receptor coactivator PNRC2 regulates energy expenditure and adiposity. J Biol Chem. 2008;283:541–553. doi: 10.1074/jbc.M703234200. [DOI] [PubMed] [Google Scholar]
- 64.Lai T, Cho H, Liu Z, Bowler MW, Piao S, Parker R, Kim YK, Song H. Structural Basis of the PNRC2-Mediated Link between mRNA Surveillance and Decapping. Structure. 2012;20:2025–2037. doi: 10.1016/j.str.2012.09.009. [DOI] [PubMed] [Google Scholar]
- 65.Rosen ED, Spiegelman BM. Molecular regulation of adipogenesis. Annu Rev Cell Dev Biol. 2000;16:145–171. doi: 10.1146/annurev.cellbio.16.1.145. [DOI] [PubMed] [Google Scholar]
- 66.Scherer PE, Lisanti MP, Baldini G, Sargiacomo M, Mastick CC, Lodish HF. Induction of caveolin during adipogenesis and association of GLUT4 with caveolin-rich vesicles. J Cell Biol. 1994;127:1233–1243. doi: 10.1083/jcb.127.5.1233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Jones JR, Barrick C, Kim KA, Lindner J, Blondeau B, Fujimoto Y, Shiota M, Kesterson RA, Kahn BB, Magnuson MA. Deletion of PPARgamma in adipose tissues of mice protects against high fat diet-induced obesity and insulin resistance. Proc Natl Acad Sci U S A. 2005;102:6207–6212. doi: 10.1073/pnas.0306743102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Providence KM, Higgins PJ. PAI-1 expression is required for epithelial cell migration in two distinct phases of in vitro wound repair. J Cell Physiol. 2004;200:297–308. doi: 10.1002/jcp.20016. [DOI] [PubMed] [Google Scholar]
- 69.Providence KM, Higgins SP, Mullen A, Battista A, Samarakoon R, Higgins CE, Wilkins-Port CE, Higgins PJ. SERPINE1 (PAI-1) is deposited into keratinocyte migration “trails” and required for optimal monolayer wound repair. Arch Dermatol Res. 2008;300:303–310. doi: 10.1007/s00403-008-0845-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Qi L, Higgins SP, Lu Q, Samarakoon R, Wilkins-Port CE, Ye Q, Higgins CE, Staiano-Coico L, Higgins PJ. SERPINE1 (PAI-1) is a prominent member of the early G0 --> G1 transition “wound repair” transcriptome in p53 mutant human keratinocytes. J Invest Dermatol. 2008;128:749–753. doi: 10.1038/sj.jid.5701068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Ploplis VA, Balsara R, Sandoval-Cooper MJ, Yin ZJ, Batten J, Modi N, Gadoua D, Donahue D, Martin JA, Castellino FJ. Enhanced in vitro proliferation of aortic endothelial cells from plasminogen activator inhibitor-1-deficient mice. J Biol Chem. 2004;279:6143–6151. doi: 10.1074/jbc.M307297200. [DOI] [PubMed] [Google Scholar]
- 72.Martin P. Wound healing--aiming for perfect skin regeneration. Science. 1997;276:75–81. doi: 10.1126/science.276.5309.75. [DOI] [PubMed] [Google Scholar]
- 73.Santoro MM, Gaudino G. Cellular and molecular facets of keratinocyte reepithelization during wound healing. Exp Cell Res. 2005;304:274–286. doi: 10.1016/j.yexcr.2004.10.033. [DOI] [PubMed] [Google Scholar]
- 74.Providence KM, Kutz SM, Staiano-Coico L, Higgins PJ. PAI-1 gene expression is regionally induced in wounded epithelial cell monolayers and required for injury repair. J Cell Physiol. 2000;182:269–280. doi: 10.1002/(SICI)1097-4652(200002)182:2<269::AID-JCP16>3.0.CO;2-E. [DOI] [PubMed] [Google Scholar]
- 75.Matsuura K, Kuratani T, Gondo T, Kamimura A, Inui M. Promotion of skin epithelial cell migration and wound healing by a 2-benzazepine derivative. Eur J Pharmacol. 2007;563:83–87. doi: 10.1016/j.ejphar.2007.02.014. [DOI] [PubMed] [Google Scholar]
- 76.Zhang J, Liu X, Datta A, Govindarajan K, Tam WL, Han J, George J, Wong C, Ramnarayanan K, Phua TY, et al. RCP is a human breast cancer-promoting gene with Ras-activating function. J Clin Invest. 2009;119:2171–2183. doi: 10.1172/JCI37622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Chinenov Y, Kerppola TK. Close encounters of many kinds: Fos-Jun interactions that mediate transcription regulatory specificity. Oncogene. 2001;20:2438–2452. doi: 10.1038/sj.onc.1204385. [DOI] [PubMed] [Google Scholar]
- 78.Ryseck RP, Bravo R. c-JUN, JUN B, and JUN D differ in their binding affinities to AP-1 and CRE consensus sequences: effect of FOS proteins. Oncogene. 1991;6:533–542. [PubMed] [Google Scholar]
- 79.Shaulian E, Karin M. AP-1 as a regulator of cell life and death. Nat Cell Biol. 2002;4:E131–136. doi: 10.1038/ncb0502-e131. [DOI] [PubMed] [Google Scholar]
- 80.Lai EC. miRNAs: whys and wherefores of miRNA-mediated regulation. Curr Biol. 2005;15:R458–460. doi: 10.1016/j.cub.2005.06.015. [DOI] [PubMed] [Google Scholar]
- 81.Sevignani C, Calin GA, Siracusa LD, Croce CM. Mammalian microRNAs: a small world for fine-tuning gene expression. Mamm Genome. 2006;17:189–202. doi: 10.1007/s00335-005-0066-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Gong C, Maquat LE. “Alu”strious long ncRNAs and their role in shortening mRNA half-lives. Cell Cycle. 2011;10:1882–1883. doi: 10.4161/cc.10.12.15589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Kohrmann M, Luo M, Kaether C, DesGroseillers L, Dotti CG, Kiebler MA. Microtubule-dependent recruitment of Staufen-green fluorescent protein into large RNA-containing granules and subsequent dendritic transport in living hippocampal neurons. Mol Biol Cell. 1999;10:2945–2953. doi: 10.1091/mbc.10.9.2945. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Jeong JH, Nam YJ, Kim SY, Kim EG, Jeong J, Kim HK. The transport of Staufen2-containing ribonucleoprotein complexes involves kinesin motor protein and is modulated by mitogen-activated protein kinase pathway. J Neurochem. 2007;102:2073–2084. doi: 10.1111/j.1471-4159.2007.04697.x. [DOI] [PubMed] [Google Scholar]
- 85.Besse F, Ephrussi A. Translational control of localized mRNAs: restricting protein synthesis in space and time. Nat Rev Mol Cell Biol. 2008;9:971–980. doi: 10.1038/nrm2548. [DOI] [PubMed] [Google Scholar]
- 86.Batish M, van den Bogaard P, Kramer FR, Tyagi S. Neuronal mRNAs travel singly into dendrites. Proc Natl Acad Sci U S A. 2012;109:4645–4650. doi: 10.1073/pnas.1111226109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 87.Doyle M, Kiebler MA. Mechanisms of dendritic mRNA transport and its role in synaptic tagging. EMBO J. 2011;30:3540–3552. doi: 10.1038/emboj.2011.278. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 88.Medioni C, Mowry K, Besse F. Principles and roles of mRNA localization in animal development. Development. 2012;139:3263–3276. doi: 10.1242/dev.078626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Kusek G, Campbell M, Doyle F, Tenenbaum SA, Kiebler M, Temple S. Asymmetric segregation of the double-stranded RNA binding protein Staufen2 during mammalian neural stem cell divisions promotes lineage progression. Cell Stem Cell. 2012;11:505–516. doi: 10.1016/j.stem.2012.06.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Vessey JP, Amadei G, Burns SE, Kiebler MA, Kaplan DR, Miller FD. An asymmetrically localized Staufen2-dependent RNA complex regulates maintenance of mammalian neural stem cells. Cell Stem Cell. 2012;11:517–528. doi: 10.1016/j.stem.2012.06.010. [DOI] [PubMed] [Google Scholar]
- 91.Giorgi C, Yeo GW, Stone ME, Katz DB, Burge C, Turrigiano G, Moore MJ. The EJC factor eIF4AIII modulates synaptic strength and neuronal protein expression. Cell. 2007;130:179–191. doi: 10.1016/j.cell.2007.05.028. [DOI] [PubMed] [Google Scholar]
- 92.Tian B, Hu J, Zhang H, Lutz CS. A large-scale analysis of mRNA polyadenylation of human and mouse genes. Nucleic Acids Res. 2005;33:201–212. doi: 10.1093/nar/gki158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Kurosaki T, Maquat LE. Rules that govern UPF1 binding to mRNA 3' UTRs. Proc Natl Acad Sci U S A. 2013 doi: 10.1073/pnas.1219908110. [DOI] [PMC free article] [PubMed] [Google Scholar]