Abstract
Background
NEUROG3 is a key regulator of pancreatic endocrine cell differentiation in mouse, essential for the generation of all mature hormone producing cells. It is repressed by Notch signaling that prevents pancreatic cell differentiation by maintaining precursors in an undifferentiated state.
Results
We show that, in zebrafish, neurog3 is not expressed in the pancreas and null neurog3 mutant embryos do not display any apparent endocrine defects. The control of endocrine cell fate is instead fulfilled by two basic helix-loop-helix factors, Ascl1b and Neurod1, that are both repressed by Notch signaling. ascl1b is transiently expressed in the mid-trunk endoderm just after gastrulation and is required for the generation of the first pancreatic endocrine precursor cells. Neurod1 is expressed afterwards in the pancreatic anlagen and pursues the endocrine cell differentiation program initiated by Ascl1b. Their complementary role in endocrine differentiation of the dorsal bud is demonstrated by the loss of all hormone-secreting cells following their simultaneous inactivation. This defect is due to a blockage of the initiation of endocrine cell differentiation.
Conclusions
This study demonstrates that NEUROG3 is not the unique pancreatic endocrine cell fate determinant in vertebrates. A general survey of endocrine cell fate determinants in the whole digestive system among vertebrates indicates that they all belong to the ARP/ASCL family but not necessarily to the Neurog3 subfamily. The identity of the ARP/ASCL factor involved depends not only on the organ but also on the species. One could, therefore, consider differentiating stem cells into insulin-producing cells without the involvement of NEUROG3 but via another ARP/ASCL factor.
Keywords: Pancreas, Endocrine, Zebrafish, Ascl1b, Neurod1, Neurog3
Background
The pancreas is a mixed gland of the digestive tract composed of an exocrine compartment (acini and ducts), releasing digestive enzymes into the duodenum, and an endocrine compartment, secreting hormones into the bloodstream in order to control glucose homeostasis. Loss or dysfunction of endocrine insulin-secreting β-cells leads to diabetes, a widespread disease affecting more than 370 million people worldwide.
Outstanding progress has been made to set up new therapies for diabetes through cell therapy (reviewed by [1-5]). Recent efforts have been focused on directing stem cells to differentiate in vitro into pancreatic β cells that could be transplanted to diabetic patients [6]. To achieve that goal, it is essential to understand in detail the molecular mechanisms controlling pancreatic endocrine cell differentiation.
Although much of our knowledge on pancreas organogenesis relies on mouse genetic studies, the use of zebrafish has also significantly contributed to the deciphering of mechanisms involved in the earliest phases of pancreas development [7-12]. In this fish, the endoderm forms two converging sheets of cells by the end of gastrulation (10 hours post fertilization, hpf). Subsequently, these cells condense at the midline to form the endodermal rod which will give rise to the digestive tract and the associated organs [9,13]. Early in development, at the 10 somite stage (10s, 14 hpf), the homeobox Pdx1 factor starts to be expressed in the endodermal region located between the first and the fourth somite [14]. As in mammals, a dorsal and a ventral pancreatic bud will emerge from this pdx1+ region and will later coalesce to form the pancreas [15]. In zebrafish, the first hormone-expressing cells that appear from the dorsal bud are the insulin-producing β-cells, detected from 15 hpf onward. Next appear the somatostatin-secreting δ-cells (17 hpf), the ghrelin ϵ-cells (18 hpf) and finally the glucagon-producing α-cells (21 hpf). This first wave of endocrine cells is followed by a second wave coming from the ventral bud that forms from 32 hpf onwards [15-17]. After that stage, the increase of the endocrine cell mass is believed to result from the differentiation and proliferation of late forming ventral bud-derived endocrine cells [16]. Therefore, while at 2 days post fertilization (dpf), the vast majority of the endocrine cells is generated from the dorsal bud, at 12 days, a majority seems to derive from the ventral bud [16].
Both in zebrafish and mice, the Notch signaling pathway tightly controls pancreatic cell differentiation. Notch prevents commitment to the endocrine cell fate, thereby reserving a population of undifferentiated precursor cells for ongoing proliferation and generation of later-appearing cell lineages [18-21]. Notch pathway is a fundamental and evolutionarily conserved process in metazoan development. There are numerous core players that participate in this process. Briefly, via the Hairy Enhancers-of-split proteins, Notch signaling represses the expression of genes of the Achaete-Scute like (ASCL) family or of the Atonal related protein (ARP) family, this latter being subdivided into Atonal, Neurogenin and Neurod subfamilies. These genes encode basic helix-loop-helix (bHLH) transcription factors and can be classified in two categories, ‘cell fate determinant’ factors and ‘cell differentiation’ factors [22]. The cell fate determinants are transiently expressed at early stages and are both necessary and sufficient to initiate the development of a specific cell lineage. The ‘cell differentiation’ factors are expressed at later stages and implement the differentiation program initiated by the cell fate determinants. For example, in the murine pancreas, NEUROG3 is the cell fate determinant of the pancreatic endocrine lineage [23] as its transient expression initiates the endocrine differentiation program of all endocrine cells [24-26]. Indeed, almost no endocrine pancreatic cells were detected in the Neurog3 knock-out mice [25]. NEUROG3 triggers the sustained expression of the ‘cell differentiation gene’ Neurod1 that maintains the endocrine cell differentiation program [27,28]. Homozygous Neurod1 null mice notably have a striking reduction in the number of insulin-producing β cells and fail to develop mature islets [29].
The neurog3 gene is found in the zebrafish genome but, surprisingly zebrafish neurog3 mRNAs were not detected in the developing pancreas while they were detected in the hypothalamus and intestine [20,30]. In this study, we extensively analyzed neurog3 expression during pancreas development and could not detect any expression at any stages in this tissue. The lack of Neurog3 function in the zebrafish pancreas was further confirmed by analyzing the phenotype of the recently identified sa211 neurog3 null mutant. As neuronal or endocrine cell-fate commitment controlled by Notch is classically carried out via ARP/ASCL factors [31-33], we next searched for other ARP/ASCL factors acting downstream of Notch signaling that would promote the formation of pancreatic endocrine cells. Among the 14 ARP/ASCL factors identified in the zebrafish genome, only ascl1b and neurod1 were found to be strongly expressed at early stages of endocrine cell differentiation. Knock-down analysis reveals that these factors have complementary roles in endocrine cell differentiation and that their simultaneous inactivation leads to a loss of all hormone-secreting cells. These two bHLH factors are, therefore, playing together a role analogous to that described for murine NEUROG3.
Results
neurog3 is not expressed in the pancreatic dorsal bud and null neurog3 mutant embryos do not display any apparent endocrine defects
To determine whether the fundamental role of Neurog3 in endocrine cell differentiation in mice is conserved in zebrafish, we first analyzed neurog3 expression in the pancreas by whole-mount in situ hybridization (WISH). neurog3 expression was never detected in the pancreas at any developmental stages tested (14s, 18s, 24 hpf, 30 hpf, 52 hpf, 72 hpf, 4 dpf and 5 dpf) whereas its expression was found in the hypothalamus and in scattered cells of the intestine (Figure 1A-B), as reported previously [20,30]. To ensure that Neurog3 is actually not involved in pancreatic endocrine cell differentiation as neurog3 expression level could be below the WISH detection limit, we analyzed the pancreas of a novel zebrafish neurog3 mutant recently identified at the Zebrafish Mutant Resource [34]. This neurog3 mutant allele (sa211) harbors an A to T substitution changing codon 74 (Arg) to a premature stop codon and is predicted to give rise to a truncated Neurog3 protein lacking the whole bHLH domain. The sa211 should be, therefore, a null allele. neurog3 homozygous mutant embryos do not display any modifications in the number of pancreatic endocrine α-, β-, δ- and ϵ-cells compared to wild type embryos (Figure 1C). The same result was obtained when we knocked-down neurog3 expression with a morpholino targeting its translation start site (Mo1) or its 5′UTR (Mo2) [see Additional file 1: Figure S1]. As extensive searches in the zebrafish genome did not identify any other neurog3 paralog (see below), these data indicate that the zebrafish Neurog3 factor does not control pancreatic endocrine cell fate, in contrast to the murine NEUROG3, and suggest that another bHLH factor is playing its role in this fish model.
Figure 1.
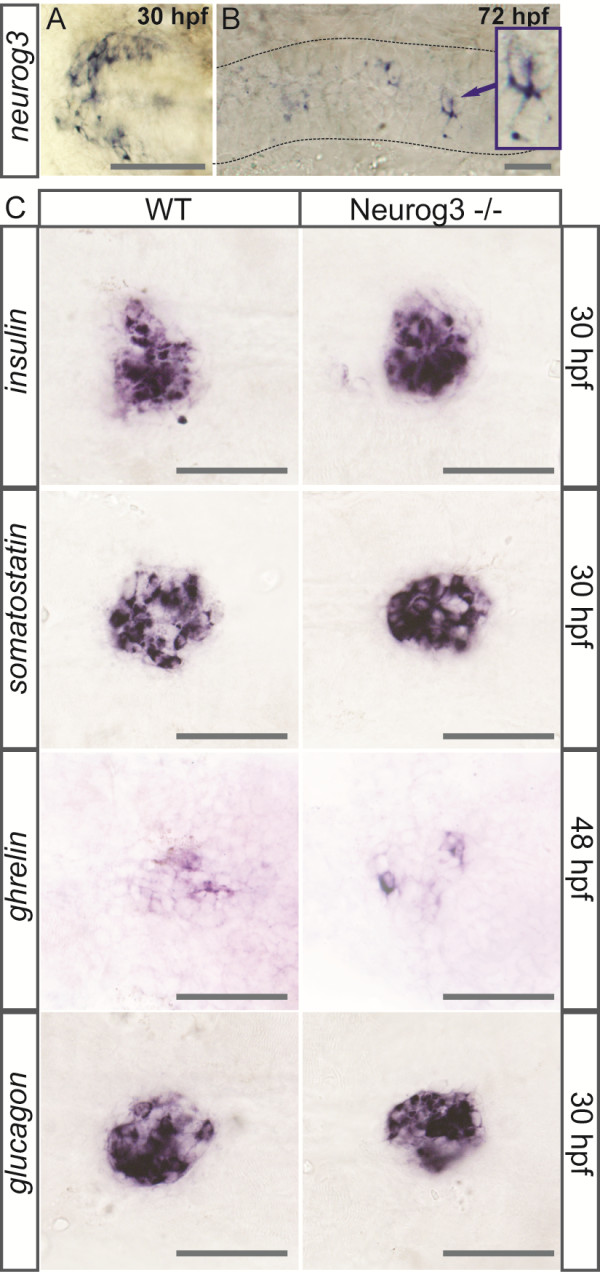
neurog3 is not expressed in the pancreas and null neurog3 mutant embryos do not display any apparent endocrine defects. (A-B) Whole-mount in situ hybridization (WISH) showing expression of neurog3 in the hypothalamus at 30 hpf (A) and in scattered cells of the gut (dotted lines) at 72 hpf (B). (C) WISH showing that the number of cells expressing insulin, somatostatin, ghrelin and glucagon is not changed in the neurog3 sa211 mutants compared to the wt siblings. All views are ventral with the anterior part to the left. Scale bars : 50 µm hpf, hours post fertilization; wt, wild type.
Expression of ARP/ASCL genes in the developing zebrafish pancreas
In order to identify the functional equivalent of murine NEUROG3 in zebrafish, we searched for another ARP/ASCL gene that could assume its role. For that purpose, we performed a phylogenetic analysis of the various ARP/ASCL found in the zebrafish genome based on the bHLH directory described by Wang et al. [35] (Figure 2A). This allowed us to identify 14 ARP/ASCL genes in the zebrafish genome, including neurog3 and another neurogenin gene, neurog1. Extensive in silico searches did not identify any other neurogenin genes in the zebrafish genome (see Material and Methods).
Figure 2.
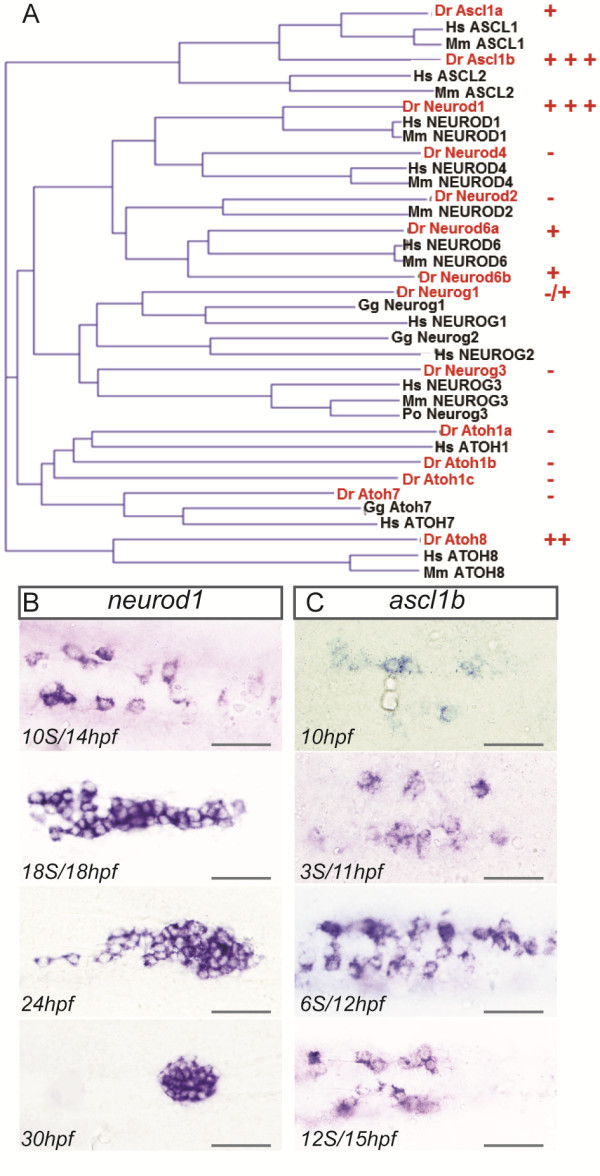
ascl1b and neurod1 are expressed during early pancreas development. (A) Phylogenetic tree calculated with full-length amino acid sequences from vertebrate members of the ARP/ASCL family. The pancreatic expression of zebrafish ARP/ASCL was tested by WISH and the expression levels are depicted by (+++) highly, (++) moderately, (+) weakly, (-/+) very weakly and (-) not expressed in the pancreas. (B-C) WISH showing expression of neurod1(B) and ascl1b(C) in the pancreatic region at the indicated stages. All views are ventral with anterior part to the left. Scale bars : 50 µm WISH, whole-mount in situ hybridization.
The expression of the 14 ARP/ASCL genes was analyzed by WISH at different time points during pancreas development (6 to 8s, 12 to 14s, 18 to 20s, 24 hpf, 30 hpf, 48 hpf, 72 hpf). We observed high expression in the pancreatic region at early stages for only two factors, namely ascl1b and neurod1 (Figure 2B-C). In contrast, only weak expression was detected at early stages for ascl1a, neurog1 and neurod6b within the pancreatic area [see Additional file 2: Figure S2, A-C], while atoh8 and neurod6a genes were detected only at late stages, that is, around two dpf [see Additional file 2: Figure S2, D-E].
neurod1 expression has already been reported in the dorsal pancreatic bud in zebrafish [20,36-38] but its precise expression profile was not determined. In this study, we show that pancreatic neurod1 expression starts around 6s (12 hpf) in a few cells. At 10s (14 hpf), neurod1 is expressed in two rows of cells located on both sites of the midline (Figure 2B). Over the next four hours, the number of neurod1+ cells progressively increases and they start to cluster to form the islet which is completely formed at 30 hpf. neurod1 remains expressed in the pancreas of four dpf larvae as well as in adults (data not shown).
ascl1b is detected before neurod1 as its expression starts as early as 10 hpf (bud stage) in two rows of cells in the prospective pancreatic region (Figure 2C). Over the following two hours, the number of ascl1b-expressing cells increases to reach its maximal level around 12 hpf. Then, ascl1b expression progressively decreases and is turned off at 17 hpf.
ascl1b and neurod1 are both expressed in the pancreatic endocrine precursors and their expression is controlled by Notch signaling
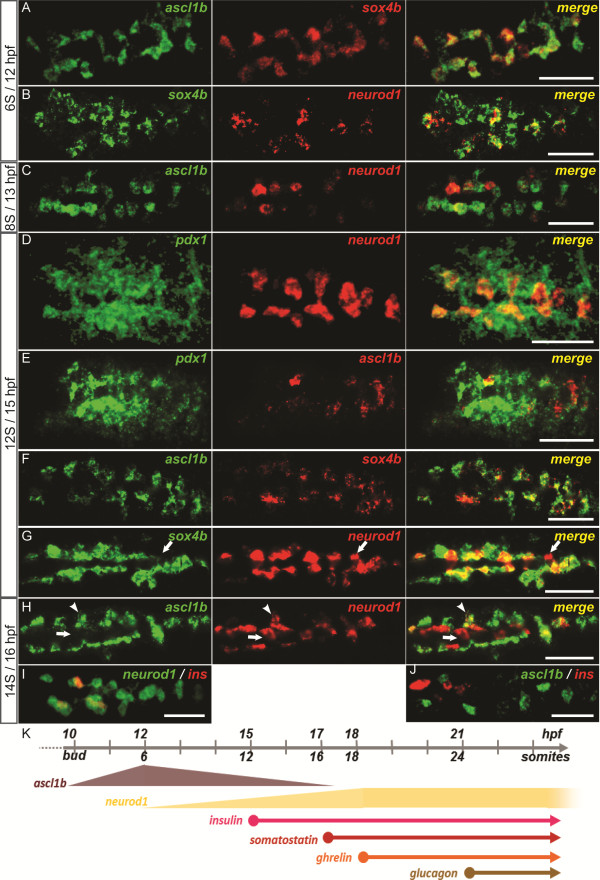
In order to determine in which pancreatic cell types these two bHLH factors are expressed, double fluorescent WISH was performed using probes for other pancreatic factors. As mentioned above, ascl1b expression starts at the end of gastrulation (10 hpf, bud stage) in the prospective pancreatic region where sox4b, the first known marker of endocrine precursors, will appear about two hours later [37,38]. As soon as sox4b is expressed in the pancreatic anlagen, its expression domain overlaps almost perfectly with ascl1b (Figure 3A) and these two factors remain co-expressed over the next five hours (Figure 3F and data not shown) until the expression of ascl1b switches off at 17 hpf. At 12 hpf, neurod1 starts to be expressed within the sox4b/ascl1b expression domain (Figure 3B and C). At 14 hpf, pdx1 expression is initiated in two bilateral rows of endodermic cells adjacent to the midline [14]. ascl1b and neurod1 are expressed within the medial part of the pdx1 expression domain, which includes the pancreatic endocrine precursors (Figure 3D and E) [8,9]. From 15 hpf, the two stripes of pancreatic precursors have begun to coalesce toward the midline starting from the posterior part (Figure 3F and G) [14,37]. At this stage, neurod1+ cells are still largely included in the ascl1b/sox4b domain with the exception of a few cells which are only labeled by neurod1 (arrow in Figure 3G). One hour later, the segregation of neurod1+ cells from the ascl1b/sox4b endocrine precursor cells is more striking (Figure 3H). This separation coincides with the appearance of the first hormone-expressing cells (that is, insulin-producing cells) which co-express neurod1 but not ascl1b (Figure 3I, J). At 17 hpf, the expression of ascl1b turns off. In contrast, neurod1 expression persists and, as shown previously [38], at 24 hpf neurod1 labels the whole endocrine cell lineage including the sox4b+ endocrine precursor cells and the hormone-expressing cells. At 30 hpf, when the majority of endocrine cells are differentiated and clustered into the single islet, neurod1 remains expressed in all endocrine cell types (α, β, δ and ϵ) (data not shown). All these data show that ascl1b is transiently expressed in the pancreatic endocrine precursors and is the earliest pancreatic marker identified so far (Figure 3K). neurod1 expression is initiated two hours later in the endocrine precursors and remains expressed in the mature endocrine cells, in contrast to ascl1b.
Figure 3.
ascl1b and neurod1 are both expressed in pancreatic endocrine precursors. (A-J) Double fluorescent WISH comparing the expression of ascl1b and neurod1 with the expression of the pancreatic factor sox4b(A, B, F, G), pdx1(D, E) or with the insulin hormone (I, J) at early stages (12 to 16 hpf). The arrows point to a cell neurod1+/sox4b- (G) or neurod1+/ascl1b- (H). The arrowheads point to a cell neurod1+/ascl1b+(H). Z-plane confocal images. All views are ventral with anterior part to the left. (K) Diagram illustrating the time windows for the expression of ascl1b, neurod1 and the pancreatic hormones. Scale bars : 50 µm hpf, hours post fertilization; WISH, whole-mount in situ hybridization.
We next asked whether ascl1b and neurod1 could be mediators of Notch signaling by analyzing their expression in mind bomb (mib) mutants. mib mutants lack a Delta-ubiquitin ligase, resulting in the failure to trigger Delta-mediated Notch signaling [39] and leading to an increase of endocrine differentiation at early stages [20,37]. We found that neurod1 and ascl1b expression is strongly increased in mib pancreas (Figure 4), indicating that ascl1b and neurod1 expression is repressed by Notch signaling, further suggesting a role of these two factors in endocrine differentiation.
Figure 4.
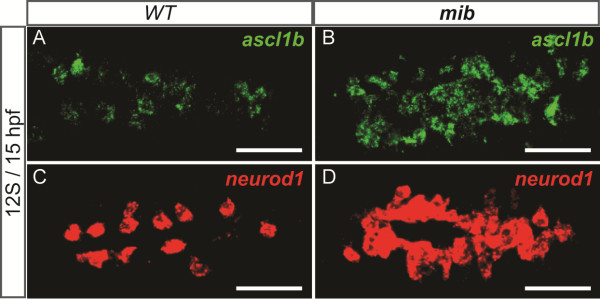
Pancreatic expression of ascl1b and neurod1 is repressed by Notch signaling. Fluorescent WISH showing pancreatic expression of ascl1b(A-B) and neurod1(C-D) in wild-type (wt) embryos (A, C) and mind bomb (mib) mutants (B, D). Confocal projection images. All views are ventral views of 15 hpf embryos with anterior part to the left. Scale bars : 50 µm hpf, hours post fertilization; WISH, whole-mount in situ hybridization.
Impaired endocrine cell differentiation in neurod1 morphants
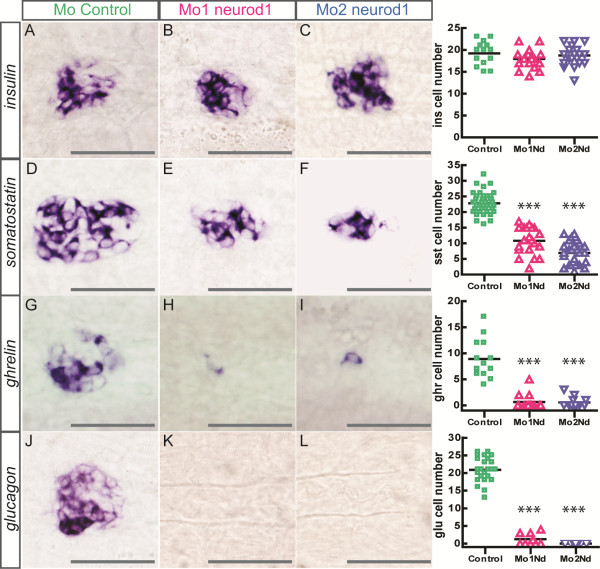
To assess the role of Neurod1 in pancreatic endocrine development, we abrogated Neurod1 protein expression in zebrafish embryos by injecting two distinct antisense morpholinos. The first (Mo1-neurod1) targets the translational start site while the second (Mo2-neurod1) targets the 5′UTR region. Both morpholinos efficiently block the translation of the neurod1 transcript as they completely prevent the expression of GFP from the Tg(neurod1:egfp) transgenic line [40], while they do not disturb the overall morphology of the embryos [see Additional file 3: Figure S3]. In contrast, Neurod1 knockdown leads to pancreatic defects as the injection of each morpholino led to the same phenotype: an almost complete depletion of glucagon (α) and ghrelin (ϵ) cells (Figure 5G-L) together with a severe reduction in the number of somatostatin cells (δ) (Figure 5D-F). In contrast, the number of insulin cells (β) was not significantly decreased (Figure 5A-C). As the Mo2 morpholino was slightly more efficient than Mo1, the following experiments were mainly performed with Mo2.
Figure 5.
Impaired endocrine cell differentiation in neurod1 morphants. Ventral views with anterior to the left of WISH showing expression of insulin(A-C), somatostatin(D-F), ghrelin(G-I) and glucagon(J-L) in control (A, D, G, J), Mo1-neurod1(B, E, H, K) and Mo2-neurod1(C, F, I, L) morphants at 30 hpf. The quantifications on the right side of the figure represent the number of positive cells per embryo for controls and neurod1 morphants. Asterisks (***) indicate that the difference between cell number in controls and neurod1 morphants is statistically significant by Student’s t-test (P <0.001). Scale bars : 50 µm hpf, hours post fertilization; WISH, whole-mount in situ hybridization.
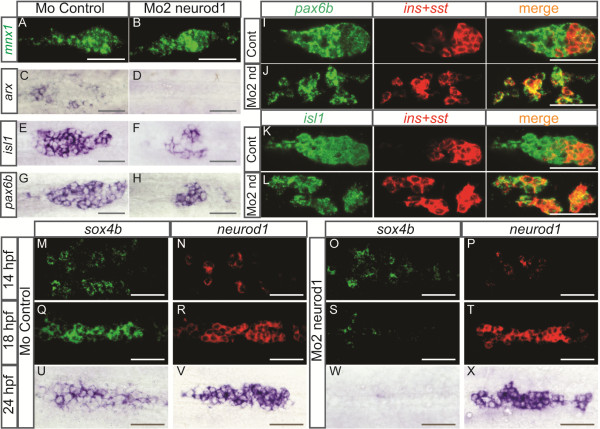
To understand the mechanism by which Neurod1 controls endocrine cell differentiation, we analyzed the expression of various pancreatic transcription factors in the neurod1 morphants at 24 hpf. Consistent with the loss of α-cells, we observed a complete extinction of arx, a factor known to be expressed specifically in all α-cells and essential for their differentiation [41] (Figure 6C and D). In contrast, expression of the mnx1/hb9 gene, specifically involved in β-cell differentiation [36,42] was unchanged, like insulin (Figure 6A and B). The expression domain of pax6b and isl1, which at 24 hpf includes all differentiated hormonal cells [38,43] was clearly reduced (around two-fold, Figure 6E-H), consistent with α-, ϵ-cells loss and a reduced number of δ-cells. Cells still expressing these two factors correspond to β-cells and to the residual δ-cells still present in neurod1 morphants (Figure 6J-L). Since the morpholinos are designed to prevent neurod1 mRNA translation but not the expression of the transcripts, we could highlight neurod1+ cells by WISH in the neurod1 morphants. neurod1+ cells were present in neurod1 morphants at all stages tested (14 hpf, 18 hpf, 24 hpf and 30 hpf) and we did not detect any obvious changes in their number compared to the control morphants (Figure 6P, T, X and data not shown). This suggests that the loss of α cells in the neurod1 morphants is not due to the apoptosis of the neurod1+ cells but due to a blockage in their differentiation process. This was further confirmed by performing terminal deoxynucleotidyl transferase dUTP nick end labeling (TUNEL) assays revealing no apoptotic cells in the pancreatic area of neurod1 and control morphants at different stages (26, 30, 40, 48 and 55 hpf) [see Additional file 4: Figure S4 A-D and data not shown].
Figure 6.
Pancreatic cells are arrested in their differentiation process in neurod1 morphants and the expression of sox4b is not maintained. Ventral views with the anterior to the left of the pancreas of embryos analyzed by visible (C-H, U-X) or fluorescent (A-B, I-T) WISH. (A-H) Pancreatic expression of mnx1, arx, isl1 and pax6b in controls and Mo2-neurod1 morphants at 24 hpf, the number of isl1+ cells in control morphant (50.4 ± 2.1) was reduced about 2.1 fold in the Mo2-neurod1 morphant (24.1 ± 1.2) and the number of pax6b+ cells in control morphant (40.7 ± 1.2) was reduced about 1.8 fold in the Mo2-neurod1 morphant (22.1 ± 1.2). (I-L) Double fluorescent WISH performed with pax6b(I, J) or isl1(K,L) probe together with a mix of insulin (ins) and somatostatin (sst) probes on 24 hpf controls or Mo2-neurod1 morphants. Z-plane confocal images (M-T) Double fluorescent WISH performed with sox4b and neurod1 on 14 hpf and 18 hpf controls or Mo2-neurod1 morphants. Confocal projection images (U-X) WISH performed with sox4b and neurod1 on 24 hpf controls or Mo2-neurod1 morphants. Scale bars : 50 µm hpf, hours post fertilization; WISH, whole-mount in situ hybridization.
To further define neurod1 function in early steps of endocrine cell differentiation, we analyzed the expression of sox4b at early stages of development. While the initiation of sox4b expression is not perturbed in neurod1 morphants (Figure 6M and O) its expression is strongly reduced at 18 hpf (Figure 6Q and S) and is not detectable anymore at 24 hpf (Figure 6U and W). All together, these data show that, in the absence of Neurod1, sox4b expression is correctly initiated. Then, around 17 to 18 hpf, the stage when most β-cells and a minority of δ-cells are already differentiated (see diagram in Figure 3K), sox4b expression is no longer maintained and the differentiation of late-appearing endocrine cells is blocked, leading to a loss of ϵ- and α-cells as well as to a reduction of δ-cells.
Impaired endocrine cell differentiation in ascl1b morphants
Intriguingly, although neurod1 is expressed from 12 hpf (6s) in the pancreatic endocrine precursors, neurod1 morphants do not display any apparent pancreatic defects before 17 hpf. We, therefore, hypothesized that, at early stages, Ascl1b could complement the loss of Neurod1 function. To test this hypothesis, we first defined Ascl1b function in endocrine differentiation by injecting an antisense morpholino targeting the translational start site of ascl1b mRNA as described in [44,45].
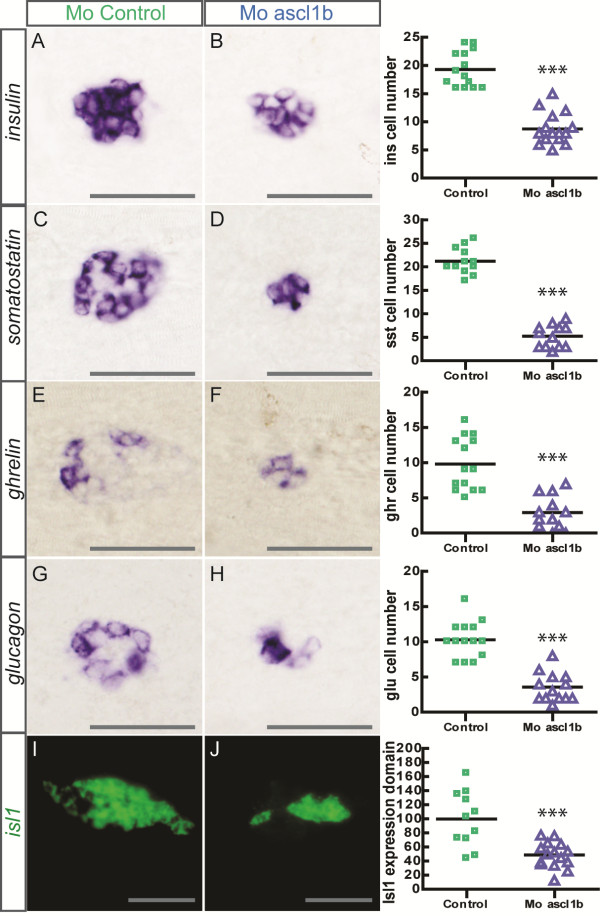
In contrast to neurod1 morphants, in ascl1b morphants we observed a significant reduction in the number of all types of pancreatic hormone-expressing cells (Figure 7). Consistent with this phenotype, we noticed a reduction of the isl1 expression domain, which includes all the mature endocrine cells at 24 hpf (Figure 7I, J). As ascl1a, the paralog of ascl1b, was detected in a few cells of the endocrine pancreas at 18 hpf [see Additional file 2: Figure S2B], we tested if Ascl1a could have redundant functions with Ascl1b. This is not the case as the knockdown of Ascl1b in the ascl1a/pia null mutant [46] did not lead to a further reduction of endocrine cells (data not shown).
Figure 7.
Impaired endocrine cell differentiation in ascl1b morphants. Ventral views with the anterior part to the left of WISH showing expression of insulin(A-B), somatostatin(C-D), ghrelin(E-F) and glucagon(G-H) in control (A, C, E, G) and ascl1b morphants (B, D, F, H) at 30 hpf. The quantifications on the right side of the figure represent the number of positive cells per embryo for controls and ascl1b morphants. (I-J) Confocal projections of ventral views of fluorescent WISH showing expression of isl1 in control (I) and ascl1b morphants (J) at 24 hpf. The quantification of isl1 expression domain is depicted on the right side of the figure. The graph shows the relative volume of isl1+ cells in control and ascl1b morphants, the mean of the volume occupied by the isl1+ cells in control morphants being arbitrarily set to 100%. Asterisks (***) indicate that the difference between controls and ascl1b morphants is statistically significant by Student’s t-test (P <0.001). Scale bars : 50 µm hpf, hours post fertilization; WISH, whole-mount in situ hybridization.
To further define Ascl1b function, we analyzed the initiation of endocrine cell differentiation in ascl1b morphants. At 12 hpf (6s), we could not detect any sox4b+ cells in the ascl1b morphants whereas sox4b was easily detectable in control morphants (Figure 8A and D). At 13 hpf (8s), one or two sox4b+ cells were present in the ascl1b morphants (Figure 8J), and over the next four hours, new sox4b+ cells appeared but their number never reached the control levels (30% compared to the control at 17 hpf, Figure 8P). In contrast, neurod1 expression was correctly initiated in ascl1b morphants (Figure 8E and K). Interestingly, the few sox4b+ cells detected in ascl1b morphants were all seen within the neurod1+ domain (Figure 8L and R) while in control morphants, a majority of sox4b+ cells do not express neurod1 (Figure 8C, I and O). These data suggest that, in ascl1b morphants, sox4b gene expression is switched on by Neurod1. This hypothesis is in accordance with the onset of sox4b that is detected just after neurod1 activation in ascl1b morphants.
Figure 8.
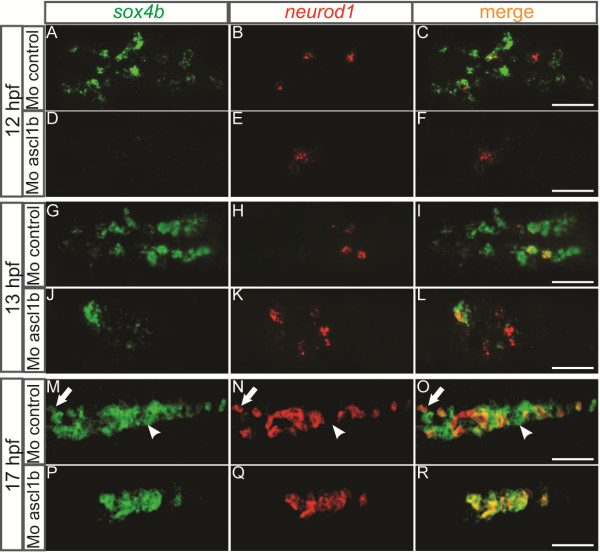
The onset of sox4b expression is delayed in ascl1b morphants and is restricted to neurod1 expressing cells. Ventral views with the anterior part to the left of double fluorescent WISH showing expression of sox4b and neurod1 at 12 hpf (A-F), 13 hpf (G-L) and 17 hpf (M-R) in control and ascl1b morphants. The domain occupied by sox4b+ and neurod1+ cells was reduced respectively 3.3- and 2.6-fold in the Mo-ascl1b at 17 hpf. The arrows point to a cell expressing sox4b and neurod1 while the arrowheads points to a cell expressing only sox4b. Z-plane confocal images. Scale bars : 50 µm hpf, hours post fertilization; WISH, whole-mount in situ hybridization.
Endocrine precursor differentiation is abolished in the double ascl1b and neurod1 morphants
To test the hypothesis that sox4b expression can be switched on either through Ascl1b or Neurod1 action, we performed a co-injection of the morpholinos targeting both genes. As shown on Additional file 5: Figure S5, this co-injection does not affect the general morphology of the embryos. In these double ascl1b/neurod1 morphants, sox4b expression was never detected, not at 17 hpf (Figure 9C-D) nor later on (data not shown). Moreover, this double knock-down abolished all endocrine cell differentiation as we did not detect any hormone-producing cells at early or late stages (17 or 38 hpf) (Figure 10A-J). In addition, isl1 expression was not detected in the endocrine islet at 38 hpf while isl1 was correctly expressed in the adjacent mesodermal tissue [10] (Figure 9A-B). Similarly, the expression of pax6b and mnx1 was completely lost in the endocrine pancreas while pax6b remains expressed in the neural tube and mnx1 gene in the motoneurons and the hypochord (Figure 9G-H). In contrast to these transcription factors, the ascl1b transcripts were still detected in the double morphants and the number of ascl1b positive cells was drastically increased at 13 hpf (Figure 9E-F) suggesting that these cells are blocked in their differentiation process. neurod1 transcript was also detected in the double morphants (Figure 9I-J), confirming the above observations that neurod1 initiation can occur independently of Ascl1b. As expected, based on the increase of the number of cells expressing the ascl1b transcripts, we did not observe any apoptosis at 14s (16 hpf) and 19s (18.5 hpf) in these double morphants [see Additional file 4: Figure S4E-H].
Figure 9.
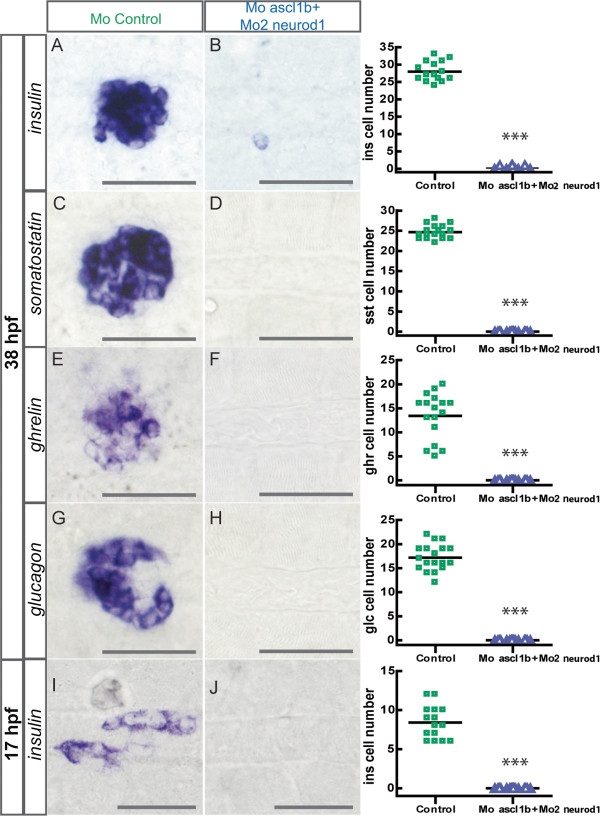
Endocrine cell differentiation is abolished in the double ascl1b and neurod1 morphants. Ventral views of WISH showing expression of insulin(A-B, I-J), somatostatin(C-D), ghrelin(E-F) and glucagon(G-H) in control (A,C,E,G,I) and Mo-ascl1b/Mo2-neurod1(B,D,F,H,J) morphants at 38 hpf (A-H) or 17 hpf (I-J). The quantifications on the right side of the figure represent the number of positive cells per embryo for controls and double ascl1b/neurod1 morphants. Asterisks (***) indicate that the difference between cell number in controls and double ascl1b/neurod1 morphants is statistically significant by Student’s t-test (P <0.001). hpf, hours post fertilization; WISH, whole-mount in situ hybridization.
Figure 10.
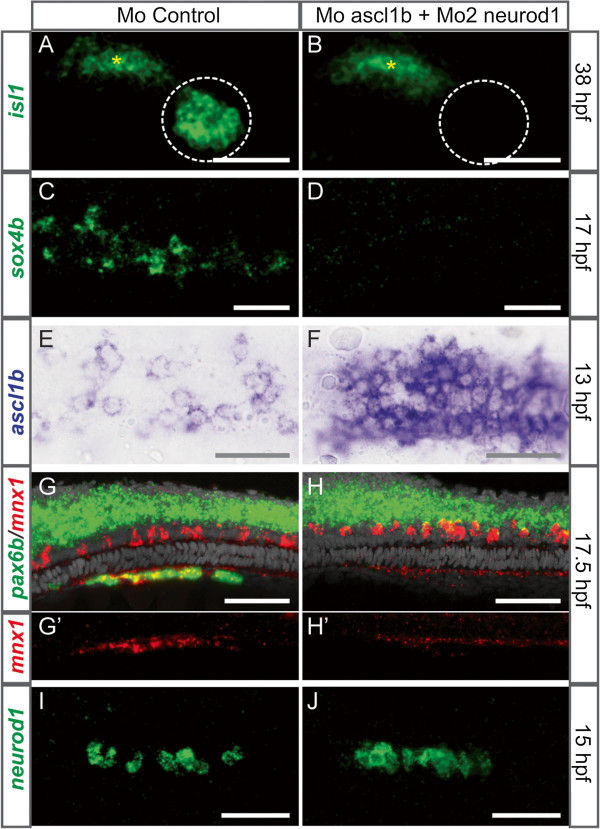
Simultaneous knock-down of Ascl1b and Neurod1 blocks the differentiation process of the precursors. Ventral views with the anterior part to the left of the pancreas of control (A,C,E,G,I) or Mo-ascl1b/Mo2-neurod1 morphants (B,D,F,H,J) analyzed by fluorescent (A-D, G-J) or visible (E-F) WISH. (A-B) Confocal projections of isl1 expression in the pancreatic islet (dotted line) and in the adjacent mesoderm (*) at 38 hpf (C-D) and of sox4b expression at 17 hpf. (E-F)ascl1b transcript expression at 13 hpf (8s). (G-H) Confocal projections of pax6b (green) and mnx1 (red) expression at 17.5 hpf showing that the pancreatic expression of pax6b and mnx1 is completely lost in the double morphants. In contrast, the expression of pax6b in the neural tube and the expression of mnx1 gene in the motoneurons and the hypochord is not perturbed. (I-J)neurod1 transcript expression at 15 hpf (12s). hpf, hours post fertilization; WISH, whole-mount in situ hybridization.
Altogether, our data indicate that the simultaneous loss of ascl1b and neurod1 expression abolishes the formation of all pancreatic hormone-expressing cells by interfering with the initiation of their differentiation process as shown by the loss of the pancreatic expression of sox4b, isl1, pax6b, mnx1 and arx genes.
ascl1b and neurod1 are also expressed in the endocrine cells of the ventral bud but not neurog3
As endocrine cells are also generated at later stages from the intrapancreatic ducts [16,17,47], we investigated whether Neurog3 could be involved in the generation of these late endocrine cells. To detect them more easily, we treated wt and neurog3 null mutant embryos from three to five dpf with the Notch-signaling inhibitor LY411575, treatment known to drastically increase the number of endocrine cells coming from the intrapancreatic ducts (Figure 11A) [17,47,48]. As shown on Figure 11B, the homozygous neurog3 mutants are able to generate endocrine cells efficiently from the intrapancreatic ducts (n = 6), suggesting that Neurog3 is not involved in the formation of the second wave endocrine cells, as it is in the first wave. This is in agreement with the absence of neurog3 expression in the intrapancreatic ducts of LY411575-treated embryos (data not shown). In contrast, we detected a strong expression of ascl1b in the intrapancreatic ducts of the LY411575-treated embryos (Figure 11C) as well as of neurod1 as previously described [48,49], suggesting that the couple Neurod1 and Ascl1b could also play an important role in the formation of second wave endocrine cells. The verification of this hypothesis will require the generation of ascl1b and neurod1 null mutant as the knock-down of Neurod1 is no longer fully efficient after 48 hpf.
Figure 11.
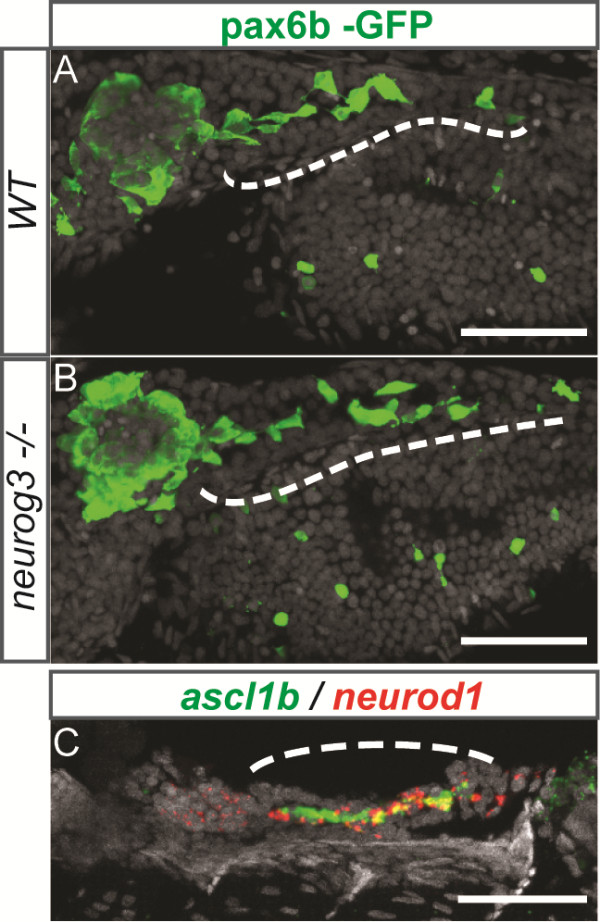
Neurog3 is not involved in the formation of the second wave endocrine cells. (A-B) Ventral views of immunohistochemistry against GFP of five dpf wt(A) or neurog3 null mutant (B) tg(pax6b:GFP) embryos treated from three to five dpf with the Notch signaling inhibitor LY411575. The generation of pax6b endocrine cells from the intrapancreatic duct (indicated by a dash line) is not perturbed in the neurog3 null mutant (n = 6). (C) Confocal projections of ascl1b (green) and neurod1 (red) expression of four dpf embryos treated from three to four dpf with the Notch signaling inhibitor LY411575 showing a strong expression in the intrapancreatic duct. Anterior part to the left. dpf, days post fertilization; wt, wild type.
Discussion
In this study, we show that, in zebrafish, Neurog3 does not control pancreatic endocrine cell fate as it does in the mouse. This function is fulfilled by two ASCL/ARP factors, Ascl1b and Neurod1.
Ascl1b starts to be expressed at bud stage (10 hpf) in the prospective pancreatic region about two hours before sox4b, that was previously the earliest known marker of endocrine precursors [37,38,50]. We show that Ascl1b initiates the pancreatic endocrine cascade and induces the formation of the first sox4b+ endocrine pancreatic precursors (see the model in Figure 12A). Two hours after ascl1b onset, neurod1 expression appears within the ascl1b/sox4b domain. Neurod1 pursues the endocrine differentiation program initiated by Ascl1b notably by maintaining the expression of sox4b in the precursors and allowing their subsequent differentiation. Indeed, in Neurod1 morphants, sox4b expression is properly initiated by Ascl1b but, around 17 to 18 hpf, when ascl1b expression turns off, it is no longer maintained (Figure 12B). The endocrine differentiation does not proceed further, thereby preventing the differentiation of later endocrine cells, that is, the majority of δ (somatostatin), ϵ (ghrelin) and α (glucagon) cells, normally appearing, respectively, at 17 hpf, 18 hpf and 21 hpf (Figure 12B).
Figure 12.
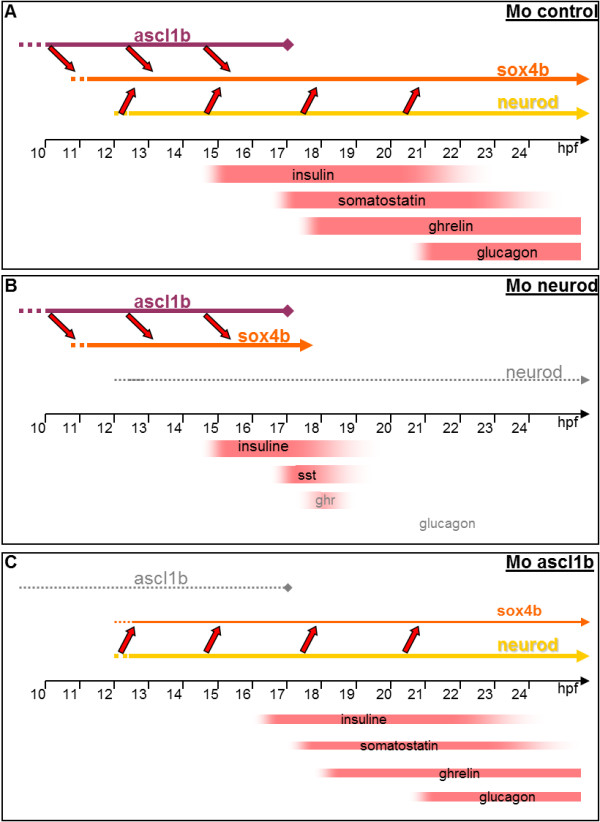
Model describing the function of ascl1b and neurod1 in pancreatic endocrine differentiation. (A) In wt embryos, Ascl1b initiates the expression of sox4b in the endocrine precursors at 11 hpf. Then, around 12 hpf, neurod1 expression starts and maintains sox4b expression in the precursors. The differentiation of the distinct endocrine cell types can take place. (B) In neurod1 morphants, Ascl1b initiates the expression of sox4b in the endocrine precursors at 11 hpf and maintains sox4b expression until 17 hpf. From that stage, ascl1b expression turns off and sox4b expression is no longer maintained. The differentiation of the late-appearing endocrine cells is blocked. (C) In ascl1b morphants, sox4b expression is not initiated at 11 hpf. When Neurod1 starts to be expressed from 12 hpf, it initiates sox4b expression in a reduced number of endocrine precursors. hpf, hours post fertilization; wt, wild type.
The role of Neurod1 as a ‘cell differentiation factor’ required for the maintenance of the endocrine differentiation program in zebrafish is similar to the murine situation where NEUROD1 pursues the endocrine differentiation program initiated by NEUROG3 and participates in the maintenance of the mature islet cells (reviewed by [51,52]). However in zebrafish, if Ascl1b is absent, Neurod1 remains expressed and acts as a cell fate determinant inducing the formation of the sox4b+ precursors. Indeed, in ascl1b morphants, sox4b+ precursor cells are detected only after neurod1 onset and sox4b expression is exclusively induced in neurod1 expressing cells while, in the control morphants, the majority of sox4b+ cells do not express neurod1. Consequently, the pool of endocrine precursors is reduced in Ascl1b morphants, leading to a reduced number of all mature endocrine cells at 30 hpf (Figure 12C). Thus, in zebrafish, neurod1 has the capacity to act as a ‘cell fate determinant’ or a ‘cell differentiation’ factor. In mice, NEUROD1 has been reported to act only as a cell differentiation factor. However, when NEUROD1 is ectopically expressed in the murine pancreatic anlagen under the control of the Pdx1 promoter, it has the same intrinsic capacity as Neurog3 to induce endocrine differentiation [28]. Such functional equivalence highlighted by a gain-of-function approach can probably be explained by the fact that these two factors regulate largely overlapping sets of genes [27].
This dual capacity to promote the selection of precursors and to regulate some differentiation steps is not restricted to Neurod1 but is shared by many ASCL/ARP factors. For example, in the olfactory epithelium, Mash1 and Neurog1 are expressed sequentially in the sensory neuron precursors. MASH1 acts as a cell fate determinant by inducing the olfactory precursors, while Neurog1 acts as a cell differentiation gene by allowing the differentiation of these precursors into sensory neurons. However, NEUROG1 can partly compensate for the loss of the determination function of MASH1 in the olfactory placode, suggesting that NEUROG1 also has the intrinsic capacities to act as a cell determinant or differentiation factor in the same tissue [53]. This suggests that the function of some ARP/ASCL factors is not intrinsically determined by their protein sequence and their structure but could be simply dictated by the context (cell type and timing of expression). This flexibility could be explained by the strong conservation within the bHLH domain. Nine of the ten residues predicted to contact DNA are identical among ARP and ASCL proteins [54] and, based on in vitro studies, there is no indication of DNA binding differences between ARP and ASCl proteins [55,56]. Therefore, these factors can regulate overlapping sets of genes as described for NEUROG3 and NEUROD1 [27] or for the proneural bHLH factors Xath5 and XNeuroD [57].
However, in other cases, the protein sequence can be crucial for the intrinsic properties of ARP/ASCL proteins acting as ‘cell fate determinants’ versus ‘cell differentiation factors.’ For example, in the context of muscle differentiation, the ‘cell differentiation gene’ Myogenin (Myog) is less efficient than the ‘cell fate determinant gene’ Myogenic factor 5 (Myf5) at remodeling chromatin and activating transcription at previously silent loci [58]. Such specificities can result from their interactions with specific cofactors. For example, functional divergence between Ato and Neurog proteins in Drosophila is encoded by three non-conserved residues in the basic domain of these bHLH which are responsible for the differential interaction with Senseless or MyT1 [59]. Such cofactors can affect the interaction of the bHLH with their DNA binding sites and/or modulate their transcriptional activity. Identifying such cofactors will be an important issue to understand better the transcriptional properties of ARP/ASCL factors.
The lack of pancreatic defects in the zebrafish neurog3 mutant contrasts with the crucial function of NEUROG3 in the mouse. This raised the question whether the neurog3 gene studied here is the actual ortholog of the murine Neurog3 gene. This seems to be the case as extensive searches in the zebrafish genome revealed the presence of only two neurogenin genes in zebrafish, neurog3 and neurog1, neurog3 being the most closely related to murine and human NEUROG3 and the locus displaying a conserved synteny with the region of human chromosome 10 containing NEUROG3[30]. As the zebrafish genome is almost completely sequenced [60], it is highly unlikely that another neurogenin gene exists in zebrafish but this possibility cannot be totally excluded. Rather, we show here that the crucial role of murine NEUROG3 in pancreatic endocrine cell differentiation is fulfilled in zebrafish by two bHLH transcription factors, Ascl1b and Neurod1. Indeed, the simultaneous inactivation of these two bHLH factors leads to a complete loss of all hormone-secreting cells, as in Neurog3 null mice. The endocrine cells remain at the precursor stage as they still express the ascl1b transcripts but none of the later pancreatic transcription factors, such as sox4b, pax6b, mnx1, isl1 and arx. During the second endocrine wave coming from the ventral bud, Ascl1b and Neurod1 probably also play an important role as we have observed a strong expression of these two factors in the intrapancreatic ducts after Notch inhibition.
This indicates that NEUROG3 is not the only cell fate determinant that can promote pancreatic endocrine cell differentiation. Even in the mouse, some rare endocrine cells can be produced in a NEUROG3-independent manner as Wang et al. reported a small, yet significant, number of glucagon expressing cells in the Neurog3-/- pancreas before e15.5 [61]. One could, therefore, consider differentiating stem cells into insulin-producing cells without the involvement of NEUROG3 but via another ARP/ASCL factor. Such an assumption is strengthened by the fact that in the murine stomach, it is ASCL1 which is required for the differentiation of all endocrine cell types [62] whereas NEUROG3 is only involved in the differentiation of a subset of these cells [63,64]. In contrast, ASCL1 is only expressed in a few cells of the murine pancreas and its knock-out does not disrupt the pancreatic endocrine differentiation [28]. These results highlight the diversity found in the selection of the ARP/ASCL factors involved in the determination and differentiation of the endocrine cells and show that the choice of these factors depends not only on the organ considered but also on the species.
Methods
Zebrafish maintenance, mutant lines and LY411575 treatment
Zebrafish (Danio rerio) were raised and cared for according to standard protocols [65]. Wild-type embryos from the AB strain were used and staged according to Kimmel (Kimmel et al., 1995). Homozygous mutants were obtained by mating heterozygous fish for the mind bomb (mibta52b) allele [66] and for the neurog3sa211 allele (ZMP, Zebrafish Mutation Project [34]). The genotyping of neurog3sa211 embryos were done on DNA extracted from tails of WISH stained embryos by performing a PCR using primers 0230 (CCAACACATACCCAGTACCTC) and O233 (TGATTTGACCTCTGTCGAAC) followed by a nested PCR, 0231 (GCTTGCAAGAGGTAAGCATC) and O232 (TGTAATTATGCGCGAATCTC) and by subsequent sequencing of the PCR products. The LY411575 treatment was performed by incubating the embryos during the indicated period with a 10 μm LY411575 solution (Medchemexpress) and replacing the media every day.
Search in the zebrafish genome of another neurog3 paralog gene
In order to discover all neurog genes present in the zebrafish genome, we screened the Zv9 Genome assembly using the ensembl genome browser [67] that covers 1.357 Gb in scaffolds placed on chromosomes 1 to 25 and 55 Mb in unassigned scaffolds. This zebrafish sequence is considered to be nearly complete. Indeed, out of a non-redundant set of 21,471 zebrafish cDNAs from ENA/Genbank, only 120 (0.6%) are not currently found in Zv9 (that is, these 120 do not have a match at 90% identity covering at least 10% of their length) [60]). Furthermore, we also screened the nucleotide collection (nt/nr) from the National Center for Biotechnology Information (NCBI) [68] that consists of GenBank + EMBL + DDBJ + PDB + RefSeq sequences. These databanks have been screened by the program ‘tblastn’ with the consensus amino acid sequence of the basic domain of vertebrate Neurog3 (RRXKANDRERNR) or of vertebrate Neurog (RRXKA(D/N)DRERNR), the amino acid sequence of the bHLH domain of murine NEUROG3 and of zebrafish Neurog3 and with the consensus amino acid sequence of the bHLH of all Neurog proteins
(RRXKANXRERXRMHXLNXALDXLRXXLPXFPXDXKLTKIETLRFAXNYIWALXXTXR). All these searches identified only 2 neurog genes in the zebrafish genome, neurog3 (AF181996) and neurog1 (AF017301).
Riboprobes, whole-mount in situ hybridization
Antisense riboprobes were made by transcribing linearized cDNA clones with SP6, T7 or T3 polymerase using digoxigenin or DNP labeling mix (Roche) according to the manufacturer’s instructions. They were subsequently purified on NucAway spin columns (Ambion) and ethanol-precipitated. The zebrafish ascl1a[69], ascl1b[69], sox4b[37], isl1[70], neurod1[71], pax6b[72], neurog3[30], atoh1a[73], atoh1b[73], neurod6a[74], neurod6b[74], atoh1c[75], atoh7[76], atoh8[77], mnx1/hb9[42], arx[78], pdx1[79], insulin[79], somatostatin 2 (PPS2)[80], ghrelin[81] and glucagon[80] probes have been described elsewhere. The neurod2, neurod4 and neurog1 probes were obtained from Imagene clones MAGp998E0411982Q (Pst1, SP6), IRBOp991C31D (Pst1, T7) and IRBOp991BO232D (EcoRI, T7), respectively. Single whole-mount and double fluorescent in situ hybridizations were carried out as described [37].
For visible WISH, quantifications of the number of cells were performed by counting the cells under the microscope by focusing successively on each layer of stained cells. For that purpose, the NBT/BCIP staining was carefully monitored in order to avoid an overstaining which would have prevented us from visualizing the individual cell boundaries. This method has been validated by counting the number of isl1-expressing cells at 24 hpf after a visible WISH or after a fluorescent WISH analyzed by confocal microscopy; very similar results were found (a mean of 49 cells counted by the first method and of 47 by the second) (see Figure 7 in [50]). For fluorescent WISH, quantifications were performed by determining the volume occupied by specific cells in control and morphants, 100% being arbitrarily fixed as the mean of the volume occupied by these cells in control morphants. Cell volume has been calculated by the program Imaris (Bitplane) on the confocal images of each embryo.
Whole-mount immunohistochemistry and TUNEL assay
WISH was performed using the following antibodies: rabbit polyclonal antibody against GFP (1:500; Chemicon), guinea pig polyclonal antibody against Insulin (1:500; Biomeda), rabbit polyclonal antibody against Somatostatin (1:1000; MP Biomedicals). The embryos were fixed with 2% formaldehyde in 0.1 M PIPES, 1.0 mM MgSO4 and 2 mM EGTA overnight at 4°C and washed three times with PBS/0.3% Triton X-100. The yolk was manually removed, the embryos permeabilized for one hour with PBS/1% Triton X-100 and then blocked for two hours in PBS/4% BSA/0.3% Triton X-100 at room temperature. Both primary and secondary antibodies were incubated overnight in the same blocking buffer. Washes were done with PBS/0.3% Triton X-100 for three hours to remove excess antibodies.
The fragmented DNA of apoptotic cells was identified by the TUNEL method, using the Apoptag Apoptosis in situ detection kit (Chemicon) in which the fragmented DNA fragments are labeled with digoxigenin-nucleotide, subsequently recognized by peroxidase coupled anti-digoxigenin antibody and revealed by incubating with tyramide-fluorescein isothiocyanate (FITC) substrate. Briefly, after the WISH, the endogenous peroxidases were inactivated by incubation for one hour in 3% H2O2 at room temperature, washed three times in PBS/1% Triton X-100 and incubated in Equilibration Buffer for one hour. The terminal deoxynucleotidyl transferase (TdT) reaction was performed by incubating the embryos in a mixture of 18 μl reaction buffer and 6 μl TdT enzyme overnight at 37°C. The reaction was stopped with 500 μl of stop/wash buffer for 10 minutes. Then, the embryos were rinsed 3 times for 10 minutes with PBS/0.3% Triton X-100. The detection was performed by incubating the embryos with 50 μl anti-digoxigenin conjugate (peroxidase) overnight at 4°C, rinsing three times for 10 minutes with PBS/0.3% Triton X-100 and incubating in 50 μl amplification reagent (Perkin Elmer). + tyramide-FITC (1/5000) for one hour at room temperature. After three 15-minute washes in PBS/0.3% Triton X-100, samples were mounted in Prolong (Invitrogen) and imaged.
Imaging
Microscope pictures were obtained with an Olympus DP70 photocamera fixed on a BX60 Olympus microscope. Confocal imaging was performed using a Leica TCS SP2 inverted confocal laser microscope (Leica Microsystems, Germany) Digitized images were acquired using a 63X (NA 1.2) Plan-Apo water-immersion objective at 1024 X 1024 pixel resolution. For multicolor imaging, FITC was visualized by using an excitation wavelength of 488 nm and the emission light was dispersed and recorded at 500 to 535 nm. Cy3 was detected by using an excitation wavelength of 543 nm and the fluorescence emission was dispersed and recorded at 555 to 620 nm. The acquisition was set up to avoid any cross-talk of the two fluorescence emissions. Series of optical sections were carried out to analyze the spatial distribution of fluorescence, and for each embryo, they were recorded with a Z-step ranging between 1 and 2 μm. Image processing, including background subtraction, was performed with Leica software (version 2.5). Captured images were exported as TIFF and further processed using Adobe Photoshop and Illustrator CS2 for figure mounting.
Morpholino design and injection
All the morpholinos (Mo) were designed by Gene Tools. The neurog3 morpholinos are complementary to either the ATG (Mo1: 5′-GGATCTTGGAGTCATTCTCTTGCAA-3′) or to the 5′UTR (Mo2: 5′-GCTCGCTCAGTAAAACCGAGGTACT-3′). The neurod1 morpholinos are complementary to either the ATG (Mo1: 5′-TTTCCTCGCTGTATGACTTCGTCAT-3′) or the 5′UTR (Mo2: 5′-CCTCTTACCTCAGTTACAATTTATA-3′). The ascl1b morpholino, as described by [44,45], is complementary to the ATG (5′-TCGTAGCGACGACAGTTGCCTCCAT-3′). A standard control Mo, having the sequence 5′-CCTCTTACCTCAGTTACAATTTATA 3′ has also been designed by Gene Tools in a way that it should have no target and no significant biological activity. A morpholino directed against p53 mRNA was used to prevent nonspecific apoptosis [82], observed with the ascl1b morpholino. They were dissolved at a concentration of 3 μg/μl in 1× Danieau buffer containing 0.5% of rhodamine dextran (to follow the microinjection process) and microinjected at the 1 to 2 cells stage at a dose of 2.5 ng for Mo1-neurog3, 3 ng for Mo2-neurog3, 3 ng for Mo1 and Mo2 neurod1 and 6 ng for Mo-ascl1b together with 3 ng Mo-p53. For the double knock-down of Ascl1b and Neurod1, a mix with 6 ng of Mo-ascl1b, 3 ng of Mo2-neurod1 and 3 ng of Mo-p53 was injected. Injected embryos were then grown in the presence of 0.003% 1-phenyl-2-thiourea until the desired stage, fixed overnight in 4% paraformaldehyde and stored in 100% methanol before analysis.
Abbreviations
ARP: Atonal related protein; ASCL: Achaete-Scute like; bHLH: basic helix-loop-helix; BSA: Bovine serum albumin; dpf: days post fertilization; FITC: Fluorescein isothiocyanate; GFP: Green fluorescent protein; hpf: hours post fertilization; Mo: Morpholino; PBS: Phosphate-buffered saline; PCR: Polymerase chain reaction; s: somite; TUNEL: Terminal deoxynucleotidyl transferase dUTP nick end labeling; UTR: Untranslated region; WISH: Whole-mount in situ hybridization; wt: wild type.
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
LCF, BP and MLV designed the research; LCF, JLP, DGS, VVB and IM performed research and analyzed data; LCF, BP and MLV wrote the paper. All authors read and approved the final manuscript.
Supplementary Material
neurog3 morphants do not display any apparent endocrine defects. WISH showing that the number of cells expressing insulin (A-C), somatostatin (D-F) and glucagon (G-I) is not changed in the Mo1-neurog3 (B,E,H) or Mo2-neurog3 (C,F,I) morphants compared to the Mo-control (A,D,G) morphants at 30 hpf. All views are ventral with the anterior part to the left.
ascl1a, neurod6b, neurog1, atoh8 and neurod6a are expressed in the pancreatic area. WISH showing expression of neurog1(A) ascl1a (B), neurod6b (C) neurod6a (D) and atoh8 (E) in the pancreatic region at the indicated stages. All views are ventral with anterior part to the left.
Mo1-neurod1 and Mo2-neurod1 morpholinos efficiently block the expression of GFP from the neurod1: egfp transgenic line. Lateral views of 26 to 30 hpf tg(neurod1:egfp) embryos injected with Mo1-neurod1 (A), Mo2-neurod1 (B) or control morpholinos analyzed by bright field illumination (A,B) or by epifluorescence illumination (A’,B’) showing that both morpholinos efficiently block the expression of GFP from the neurod1:egfp transgenic line [40] without disturbing the general morphology of the embryos.
TUNEL assays on control and morphant embryos. (A–D) Confocal image projections of 30 hpf and 40 hpf control and neurod1 morphants after TUNEL labeling for apoptotic cells (in green) and immunodetection of insulin cells. The arrows highlight individual TUNEL+ cells in the neural tube. No TUNEL+ cells were found in the pancreatic region of control or neurod1 morphants at analyzed stages. (E–H) Confocal image projections of 14S and 19S control and ascl1b/neurod1 double morphants after TUNEL labeling for apoptotic cells (in red) and immunodetection of GFP cells. No TUNEL+ cells were found in the pancreatic region of control or morphants at analyzed stages. Somites 1 (S1) to somites 4 (S4) are shown on the panels G and H.
The double ascl1b/neurod1 morphants do not display general developmental defects. Bright field views of wt embryos injected with Mo-ascl1b and Mo2-neurod1 morpholinos (A) or control morpholinos (B) showing no general developmental defects in the double morphants at 22 hpf.
Contributor Information
Lydie C Flasse, Email: lydie.flasse@sund.ku.dk.
Justine L Pirson, Email: Justine.Pirson@student.ulg.ac.be.
David G Stern, Email: d.stern@student.ulg.ac.be.
Virginie Von Berg, Email: vonbergvirg@hotmail.com.
Isabelle Manfroid, Email: isabelle.manfroid@ulg.ac.be.
Bernard Peers, Email: bpeers@ulg.ac.be.
Marianne L Voz, Email: mvoz@ulg.ac.be.
Acknowledgements
We thank M. Hammerschidt (ascl1b), G. Bellipanni (ascl1a), B.B. Riley (atoh1a and atoh1b), S. Wilson (atoh5), Gong Zhiyuang (neurod6a and neurod6b), M. Hibi (atoh1c) and W. Jia (atoh8) for sending probes. We thank the Sanger Institute Zebrafish Mutation Resource for providing the zebrafish neurog3 knock-out allele sa211. The zebrafish lines were raised with the help of M. Winandy (Zebrafish Facility and Transgenics Platform, GIGA, University of Liège). The results of confocal imaging were obtained thanks to S. Ormenese and G. Moraes (Imaging Platforms, GIGA, University of Liege). The results of sequencing were obtained thanks to Genomic-Sequencing Platform, GIGA, University of Liège. L.C.F was supported by the FNRS-FRS and the Leon Fredericq fund and J.L.P and D.G.S by the FRIA. I.M. was supported by the FNRS-FRS and by the Action de Recherches Concertées (University of Liège). B.P. and M.L.V. are Chercheur qualifié FNRS. This work was funded by the Belgian State’s ‘Interuniversity Attraction Poles’ Program (SSTC, PAI).
References
- Desgraz R, Bonal C, Herrera PL. β-cell regeneration: the pancreatic intrinsic faculty. Trends Endocrinol Metab. 2011;22:34–43. doi: 10.1016/j.tem.2010.09.004. [DOI] [PubMed] [Google Scholar]
- Prince VE, Kinkel MD. Recent advances in pancreas development: from embryonic pathways to programming renewable sources of beta cells. F1000 Biol Rep. 2010;2:17. doi: 10.3410/B2-17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tuduri E, Kieffer TJ. Reprogramming gut and pancreas endocrine cells to treat diabetes. Diabetes Obes Metab. 2011;13:53–59. doi: 10.1111/j.1463-1326.2011.01446.x. [DOI] [PubMed] [Google Scholar]
- Zaret KS, Grompe M. Generation and regeneration of cells of the liver and pancreas. Science. 2008;322:1490–1494. doi: 10.1126/science.1161431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stafford D, Hornbruch A, Mueller PR, Prince VE. A conserved role for retinoid signaling in vertebrate pancreas development. Dev Genes Evol. 2004;214:432–441. doi: 10.1007/s00427-004-0420-6. [DOI] [PubMed] [Google Scholar]
- D’Amour KA, Bang AG, Eliazer S, Kelly OG, Agulnick AD, Smart NG, Moorman MA, Kroon E, Carpenter MK, Baetge EE. Production of pancreatic hormone-expressing endocrine cells from human embryonic stem cells. Nat Biotechnol. 2006;24:1392–1401. doi: 10.1038/nbt1259. [DOI] [PubMed] [Google Scholar]
- Chung WS, Andersson O, Row R, Kimelman D, Stainier DY. Suppression of Alk8-mediated Bmp signaling cell-autonomously induces pancreatic beta-cells in zebrafish. Proc Natl Acad Sci U S A. 2010;107:1142–1147. doi: 10.1073/pnas.0910205107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chung WS, Shin CH, Stainier DY. Bmp2 signaling regulates the hepatic versus pancreatic fate decision. Dev Cell. 2008;15:738–748. doi: 10.1016/j.devcel.2008.08.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chung WS, Stainier DY. Intra-endodermal interactions are required for pancreatic beta cell induction. Dev Cell. 2008;14:582–593. doi: 10.1016/j.devcel.2008.02.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manfroid I, Delporte F, Baudhuin A, Motte P, Neumann CJ, Voz ML, Martial JA, Peers B. Reciprocal endoderm-mesoderm interactions mediated by fgf24 and fgf10 govern pancreas development. Development. 2007;134:4011–4021. doi: 10.1242/dev.007823. [DOI] [PubMed] [Google Scholar]
- Naye F, Voz ML, Detry N, Hammerschmidt M, Peers B, Manfroid I. Essential roles of zebrafish bmp2a, fgf10, and fgf24 in the specification of the ventral pancreas. Mol Biol Cell. 2012;23:945–954. doi: 10.1091/mbc.E11-08-0664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stafford D, Prince VE. Retinoic acid signaling is required for a critical early step in zebrafish pancreatic development. Curr Biol. 2002;12:1215–1220. doi: 10.1016/S0960-9822(02)00929-6. [DOI] [PubMed] [Google Scholar]
- Ober EA, Field HA, Stainier DY. From endoderm formation to liver and pancreas development in zebrafish. Mech Dev. 2003;120:5–18. doi: 10.1016/S0925-4773(02)00327-1. [DOI] [PubMed] [Google Scholar]
- Biemar F, Argenton F, Schmidtke R, Epperlein S, Peers B, Driever W. Pancreas development in zebrafish: early dispersed appearance of endocrine hormone expressing cells and their convergence to form the definitive islet. Dev Biol. 2001;230:189–203. doi: 10.1006/dbio.2000.0103. [DOI] [PubMed] [Google Scholar]
- Field HA, Dong PD, Beis D, Stainier DY. Formation of the digestive system in zebrafish. II. Pancreas morphogenesis. Dev Biol. 2003;261:197–208. doi: 10.1016/S0012-1606(03)00308-7. [DOI] [PubMed] [Google Scholar]
- Hesselson D, Anderson RM, Beinat M, Stainier DY. Distinct populations of quiescent and proliferative pancreatic beta-cells identified by HOTcre mediated labeling. Proc Natl Acad Sci U S A. 2009;106:14896–14901. doi: 10.1073/pnas.0906348106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y, Rovira M, Yusuff S, Parsons MJ. Genetic inducible fate mapping in larval zebrafish reveals origins of adult insulin-producing beta-cells. Development. 2011;138:609–617. doi: 10.1242/dev.059097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Esni F, Ghosh B, Biankin AV, Lin JW, Albert MA, Yu X, MacDonald RJ, Civin CI, Real FX, Pack MA, Ball DW, Leach SD. Notch inhibits Ptf1 function and acinar cell differentiation in developing mouse and zebrafish pancreas. Development. 2004;131:4213–4224. doi: 10.1242/dev.01280. [DOI] [PubMed] [Google Scholar]
- Yee NS, Lorent K, Pack M. Exocrine pancreas development in zebrafish. Dev Biol. 2005;284:84–101. doi: 10.1016/j.ydbio.2005.04.035. [DOI] [PubMed] [Google Scholar]
- Zecchin E, Filippi A, Biemar F, Tiso N, Pauls S, Ellertsdottir E, Gnugge L, Bortolussi M, Driever W, Argenton F. Distinct delta and jagged genes control sequential segregation of pancreatic cell types from precursor pools in zebrafish. Dev Biol. 2007;301:192–204. doi: 10.1016/j.ydbio.2006.09.041. [DOI] [PubMed] [Google Scholar]
- Kim W, Shin YK, Kim BJ, Egan JM. Notch signaling in pancreatic endocrine cell and diabetes. Biochem Biophys Res Commun. 2010;392:247–251. doi: 10.1016/j.bbrc.2009.12.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bertrand N, Castro DS, Guillemot F. Proneural genes and the specification of neural cell types. Nat Rev Neurosci. 2002;3:517–530. doi: 10.1038/nrn874. [DOI] [PubMed] [Google Scholar]
- Rukstalis JM, Habener JF. Neurogenin3: a master regulator of pancreatic islet differentiation and regeneration. Islets. 2009;1:177–184. doi: 10.4161/isl.1.3.9877. [DOI] [PubMed] [Google Scholar]
- Apelqvist A, Li H, Sommer L, Beatus P, Anderson DJ, Honjo T, Hrabe de Angelis M, Lendahl U, Edlund H. Notch signalling controls pancreatic cell differentiation. Nature. 1999;400:877–881. doi: 10.1038/23716. [DOI] [PubMed] [Google Scholar]
- Gradwohl G, Dierich A, LeMeur M, Guillemot F. neurogenin3 is required for the development of the four endocrine cell lineages of the pancreas. Proc Natl Acad Sci U S A. 2000;97:1607–1611. doi: 10.1073/pnas.97.4.1607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grapin-Botton A, Majithia AR, Melton DA. Key events of pancreas formation are triggered in gut endoderm by ectopic expression of pancreatic regulatory genes. Genes Dev. 2001;15:444–454. doi: 10.1101/gad.846001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gasa R, Mrejen C, Lynn FC, Skewes-Cox P, Sanchez L, Yang KY, Lin CH, Gomis R, German MS. Induction of pancreatic islet cell differentiation by the neurogenin-neuroD cascade. Differentiation. 2008;76:381–391. doi: 10.1111/j.1432-0436.2007.00228.x. [DOI] [PubMed] [Google Scholar]
- Schwitzgebel VM, Scheel DW, Conners JR, Kalamaras J, Lee JE, Anderson DJ, Sussel L, Johnson JD, German MS. Expression of neurogenin3 reveals an islet cell precursor population in the pancreas. Development. 2000;127:3533–3542. doi: 10.1242/dev.127.16.3533. [DOI] [PubMed] [Google Scholar]
- Naya FJ, Huang HP, Qiu Y, Mutoh H, DeMayo FJ, Leiter AB, Tsai MJ. Diabetes, defective pancreatic morphogenesis, and abnormal enteroendocrine differentiation in BETA2/neuroD-deficient mice. Genes Dev. 1997;11:2323–2334. doi: 10.1101/gad.11.18.2323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang X, Chu LT, He J, Emelyanov A, Korzh V, Gong Z. A novel zebrafish bHLH gene, neurogenin3, is expressed in the hypothalamus. Gene. 2001;275:47–55. doi: 10.1016/S0378-1119(01)00648-5. [DOI] [PubMed] [Google Scholar]
- Fiuza UM, Arias AM. Cell and molecular biology of Notch. J Endocrinol. 2007;194:459–474. doi: 10.1677/JOE-07-0242. [DOI] [PubMed] [Google Scholar]
- Fior R, Henrique D. “Notch-Off”: a perspective on the termination of Notch signalling. Int J Dev Biol. 2009;53:1379–1384. doi: 10.1387/ijdb.072309rf. [DOI] [PubMed] [Google Scholar]
- Ehebauer M, Hayward P, Arias AM. Notch, a universal arbiter of cell fate decisions. Science. 2006;314:1414–1415. doi: 10.1126/science.1134042. [DOI] [PubMed] [Google Scholar]
- Zebrafish Mutant Resource. http://www.sanger.ac.uk/Projects/D_rerio/zmp/
- Wang Y, Chen K, Yao Q, Zheng X, Yang Z. Phylogenetic analysis of zebrafish basic helix-loop-helix transcription factors. J Mol Evol. 2009;68:629–640. doi: 10.1007/s00239-009-9232-7. [DOI] [PubMed] [Google Scholar]
- Dalgin G, Ward AB, Hao le T, Beattie CE, Nechiporuk A, Prince VE. Zebrafish mnx1 controls cell fate choice in the developing endocrine pancreas. Development. 2011;138:4597–4608. doi: 10.1242/dev.067736. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mavropoulos A, Devos N, Biemar F, Zecchin E, Argenton F, Edlund H, Motte P, Martial JA, Peers B. sox4b is a key player of pancreatic alpha cell differentiation in zebrafish. Dev Biol. 2005;285:211–223. doi: 10.1016/j.ydbio.2005.06.024. [DOI] [PubMed] [Google Scholar]
- Soyer J, Flasse L, Raffelsberger W, Beucher A, Orvain C, Peers B, Ravassard P, Vermot J, Voz ML, Mellitzer G, Gradwohl G. Rfx6 is an Ngn3-dependent winged helix transcription factor required for pancreatic islet cell development. Development. 2010;137:203–212. doi: 10.1242/dev.041673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Itoh M, Kim CH, Palardy G, Oda T, Jiang YJ, Maust D, Yeo SY, Lorick K, Wright GJ, Ariza-McNaughton L, Weissman AM, Lewis J, Chandrasekharappa SC, Chitnis AB. Mind bomb is a ubiquitin ligase that is essential for efficient activation of Notch signaling by Delta. Dev Cell. 2003;4:67–82. doi: 10.1016/S1534-5807(02)00409-4. [DOI] [PubMed] [Google Scholar]
- Obholzer N, Wolfson S, Trapani JG, Mo W, Nechiporuk A, Busch-Nentwich E, Seiler C, Sidi S, Sollner C, Duncan RN, Boehland A, Nicolson T. Vesicular glutamate transporter 3 is required for synaptic transmission in zebrafish hair cells. J Neurosci. 2008;28:2110–2118. doi: 10.1523/JNEUROSCI.5230-07.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Collombat P, Mansouri A, Hecksher-Sorensen J, Serup P, Krull J, Gradwohl G, Gruss P. Opposing actions of Arx and Pax4 in endocrine pancreas development. Genes Dev. 2003;17:2591–2603. doi: 10.1101/gad.269003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wendik B, Maier E, Meyer D. Zebrafish mnx genes in endocrine and exocrine pancreas formation. Dev Biol. 2004;268:372–383. doi: 10.1016/j.ydbio.2003.12.026. [DOI] [PubMed] [Google Scholar]
- Delporte FM, Pasque V, Devos N, Manfroid I, Voz ML, Motte P, Biemar F, Martial JA, Peers B. Expression of zebrafish pax6b in pancreas is regulated by two enhancers containing highly conserved cis-elements bound by PDX1, PBX and PREP factors. BMC Dev Biol. 2008;8:53. doi: 10.1186/1471-213X-8-53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nikolaou N, Watanabe-Asaka T, Gerety S, Distel M, Koster RW, Wilkinson DG. Lunatic fringe promotes the lateral inhibition of neurogenesis. Development. 2009;136:2523–2533. doi: 10.1242/dev.034736. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Amoyel M, Cheng YC, Jiang YJ, Wilkinson DG. Wnt1 regulates neurogenesis and mediates lateral inhibition of boundary cell specification in the zebrafish hindbrain. Development. 2005;132:775–785. doi: 10.1242/dev.01616. [DOI] [PubMed] [Google Scholar]
- Pogoda HM, von der Hardt S, Herzog W, Kramer C, Schwarz H, Hammerschmidt M. The proneural gene ascl1a is required for endocrine differentiation and cell survival in the zebrafish adenohypophysis. Development. 2006;133:1079–1089. doi: 10.1242/dev.02296. [DOI] [PubMed] [Google Scholar]
- Parsons MJ, Pisharath H, Yusuff S, Moore JC, Siekmann AF, Lawson N, Leach SD. Notch-responsive cells initiate the secondary transition in larval zebrafish pancreas. Mech Dev. 2009;126:898–912. doi: 10.1016/j.mod.2009.07.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ninov N, Borius M, Stainier DY. Different levels of Notch signaling regulate quiescence, renewal and differentiation in pancreatic endocrine progenitors. Development. 2012;139:1557–1567. doi: 10.1242/dev.076000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manfroid I, Ghaye A, Naye F, Detry N, Palm S, Pan L, Ma TP, Huang W, Rovira M, Martial JA, Parsons MJ, Moens CB, Voz ML, Peers B. Zebrafish sox9b is crucial for hepatopancreatic duct development and pancreatic endocrine cell regeneration. Dev Biol. 2012;366:268–278. doi: 10.1016/j.ydbio.2012.04.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Binot AC, Manfroid I, Flasse L, Winandy M, Motte P, Martial JA, Peers B, Voz ML. Nkx6.1 and nkx6.2 regulate alpha- and beta-cell formation in zebrafish by acting on pancreatic endocrine progenitor cells. Dev Biol. 2010;340:397–407. doi: 10.1016/j.ydbio.2010.01.025. [DOI] [PubMed] [Google Scholar]
- Wilson ME, Scheel D, German MS. Gene expression cascades in pancreatic development. Mech Dev. 2003;120:65–80. doi: 10.1016/S0925-4773(02)00333-7. [DOI] [PubMed] [Google Scholar]
- Jensen J. Gene regulatory factors in pancreatic development. Dev Dyn. 2004;229:176–200. doi: 10.1002/dvdy.10460. [DOI] [PubMed] [Google Scholar]
- Cau E, Casarosa S, Guillemot F. Mash1 and Ngn1 control distinct steps of determination and differentiation in the olfactory sensory neuron lineage. Development. 2002;129:1871–1880. doi: 10.1242/dev.129.8.1871. [DOI] [PubMed] [Google Scholar]
- Chien CT, Hsiao CD, Jan LY, Jan YN. Neuronal type information encoded in the basic-helix-loop-helix domain of proneural genes. Proc Natl Acad Sci U S A. 1996;93:13239–13244. doi: 10.1073/pnas.93.23.13239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jarman AP, Grau Y, Jan LY, Jan YN. atonal is a proneural gene that directs chordotonal organ formation in the Drosophila peripheral nervous system. Cell. 1993;73:1307–1321. doi: 10.1016/0092-8674(93)90358-W. [DOI] [PubMed] [Google Scholar]
- Powell LM, Zur Lage PI, Prentice DR, Senthinathan B, Jarman AP. The proneural proteins Atonal and Scute regulate neural target genes through different E-box binding sites. Mol Cell Biol. 2004;24:9517–9526. doi: 10.1128/MCB.24.21.9517-9526.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Logan MA, Steele MR, Van Raay TJ, Vetter ML. Identification of shared transcriptional targets for the proneural bHLH factors Xath5 and XNeuroD. Dev Biol. 2005;285:570–583. doi: 10.1016/j.ydbio.2005.06.033. [DOI] [PubMed] [Google Scholar]
- Gerber AN, Klesert TR, Bergstrom DA, Tapscott SJ. Two domains of MyoD mediate transcriptional activation of genes in repressive chromatin: a mechanism for lineage determination in myogenesis. Genes Dev. 1997;11:436–450. doi: 10.1101/gad.11.4.436. [DOI] [PubMed] [Google Scholar]
- Quan XJ, Denayer T, Yan J, Jafar-Nejad H, Philippi A, Lichtarge O, Vleminckx K, Hassan BA. Evolution of neural precursor selection: functional divergence of proneural proteins. Development. 2004;131:1679–1689. doi: 10.1242/dev.01055. [DOI] [PubMed] [Google Scholar]
- Howe K, Clark MD, Torroja CF, Torrance J, Berthelot C, Muffato M, Collins JE, Humphray S, McLaren K, Matthews L, McLaren S, Sealy I, Caccamo M, Churcher C, Scott C, Barrett JC, Koch R, Rauch GJ, White S, Chow W, Kilian B, Quintais LT, Guerra-Assunção JA, Zhou Y, Gu Y, Yen J, Vogel JH, Eyre T, Redmond S, Banerjee R. et al. The zebrafish reference genome sequence and its relationship to the human genome. Nature. 2013;496:498–503. doi: 10.1038/nature12111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang S, Hecksher-Sorensen J, Xu Y, Zhao A, Dor Y, Rosenberg L, Serup P, Gu G. Myt1 and Ngn3 form a feed-forward expression loop to promote endocrine islet cell differentiation. Dev Biol. 2008;317:531–540. doi: 10.1016/j.ydbio.2008.02.052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kokubu H, Ohtsuka T, Kageyama R. Mash1 is required for neuroendocrine cell development in the glandular stomach. Genes Cells. 2008;13:41–51. doi: 10.1111/j.1365-2443.2007.01146.x. [DOI] [PubMed] [Google Scholar]
- Jenny M, Uhl C, Roche C, Duluc I, Guillermin V, Guillemot F, Jensen J, Kedinger M, Gradwohl G. Neurogenin3 is differentially required for endocrine cell fate specification in the intestinal and gastric epithelium. EMBO J. 2002;21:6338–6347. doi: 10.1093/emboj/cdf649. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee CS, Perreault N, Brestelli JE, Kaestner KH. Neurogenin 3 is essential for the proper specification of gastric enteroendocrine cells and the maintenance of gastric epithelial cell identity. Genes Dev. 2002;16:1488–1497. doi: 10.1101/gad.985002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Westerfield M. The Zebrafish Book: A Guide for the Laboratory Use of Zebrafish (Danio rerio) 3. Eugene, OR: M. Westerfield; 1995. [Google Scholar]
- Haddon C, Jiang YJ, Smithers L, Lewis J. Delta-Notch signalling and the patterning of sensory cell differentiation in the zebrafish ear: evidence from the mind bomb mutant. Development. 1998;125:4637–4644. doi: 10.1242/dev.125.23.4637. [DOI] [PubMed] [Google Scholar]
- Ensembl Genome Browser. http://www.ensembl.org/Danio_rerio/
- BLAST: Basic Local Alignment Search Tool. http://blast.ncbi.nlm.nih.gov/Blast.cgi.
- Allende ML, Weinberg ES. The expression pattern of two zebrafish achaete-scute homolog (ash) genes is altered in the embryonic brain of the cyclops mutant. Dev Biol. 1994;166:509–530. doi: 10.1006/dbio.1994.1334. [DOI] [PubMed] [Google Scholar]
- Korzh V, Edlund T, Thor S. Zebrafish primary neurons initiate expression of the LIM homeodomain protein Isl-1 at the end of gastrulation. Development. 1993;118:417–425. doi: 10.1242/dev.118.2.417. [DOI] [PubMed] [Google Scholar]
- Korzh V, Sleptsova I, Liao J, He J, Gong Z. Expression of zebrafish bHLH genes ngn1 and nrd defines distinct stages of neural differentiation. Dev Dyn. 1998;213:92–104. doi: 10.1002/(SICI)1097-0177(199809)213:1<92::AID-AJA9>3.0.CO;2-T. [DOI] [PubMed] [Google Scholar]
- Krauss S, Johansen T, Korzh V, Moens U, Ericson JU, Fjose A. Zebrafish pax[zf-a]: a paired box-containing gene expressed in the neural tube. EMBO J. 1991;10:3609–3619. doi: 10.1002/j.1460-2075.1991.tb04927.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Millimaki BB, Sweet EM, Dhason MS, Riley BB. Zebrafish atoh1 genes: classic proneural activity in the inner ear and regulation by Fgf and Notch. Development. 2007;134:295–305. doi: 10.1242/dev.02734. [DOI] [PubMed] [Google Scholar]
- Liao J, He J, Yan T, Korzh V, Gong Z. A class of neuroD-related basic helix-loop-helix transcription factors expressed in developing central nervous system in zebrafish. DNA Cell Biol. 1999;18:333–344. doi: 10.1089/104454999315394. [DOI] [PubMed] [Google Scholar]
- Kani S, Bae YK, Shimizu T, Tanabe K, Satou C, Parsons MJ, Scott E, Higashijima S, Hibi M. Proneural gene-linked neurogenesis in zebrafish cerebellum. Dev Biol. 2010;343:1–17. doi: 10.1016/j.ydbio.2010.03.024. [DOI] [PubMed] [Google Scholar]
- Masai I, Stemple DL, Okamoto H, Wilson SW. Midline signals regulate retinal neurogenesis in zebrafish. Neuron. 2000;27:251–263. doi: 10.1016/S0896-6273(00)00034-9. [DOI] [PubMed] [Google Scholar]
- Yao J, Zhou J, Liu Q, Lu D, Wang L, Qiao X, Jia W. Atoh8, a bHLH transcription factor, is required for the development of retina and skeletal muscle in zebrafish. PLoS One. 2010;5:e10945. doi: 10.1371/journal.pone.0010945. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miura H, Yanazawa M, Kato K, Kitamura K. Expression of a novel aristaless related homeobox gene ‘Arx’ in the vertebrate telencephalon, diencephalon and floor plate. Mech Dev. 1997;65:99–109. doi: 10.1016/S0925-4773(97)00062-2. [DOI] [PubMed] [Google Scholar]
- Milewski WM, Duguay SJ, Chan SJ, Steiner DF. Conservation of PDX-1 structure, function, and expression in zebrafish. Endocrinology. 1998;139:1440–1449. doi: 10.1210/en.139.3.1440. [DOI] [PubMed] [Google Scholar]
- Argenton F, Zecchin E, Bortolussi M. Early appearance of pancreatic hormone-expressing cells in the zebrafish embryo. Mech Dev. 1999;87:217–221. doi: 10.1016/S0925-4773(99)00151-3. [DOI] [PubMed] [Google Scholar]
- Pauls S, Zecchin E, Tiso N, Bortolussi M, Argenton F. Function and regulation of zebrafish nkx2.2a during development of pancreatic islet and ducts. Dev Biol. 2007;304:875–890. doi: 10.1016/j.ydbio.2007.01.024. [DOI] [PubMed] [Google Scholar]
- Robu ME, Larson JD, Nasevicius A, Beiraghi S, Brenner C, Farber SA, Ekker SC. p53 activation by knockdown technologies. PLoS Genet. 2007;3:e78. doi: 10.1371/journal.pgen.0030078. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
neurog3 morphants do not display any apparent endocrine defects. WISH showing that the number of cells expressing insulin (A-C), somatostatin (D-F) and glucagon (G-I) is not changed in the Mo1-neurog3 (B,E,H) or Mo2-neurog3 (C,F,I) morphants compared to the Mo-control (A,D,G) morphants at 30 hpf. All views are ventral with the anterior part to the left.
ascl1a, neurod6b, neurog1, atoh8 and neurod6a are expressed in the pancreatic area. WISH showing expression of neurog1(A) ascl1a (B), neurod6b (C) neurod6a (D) and atoh8 (E) in the pancreatic region at the indicated stages. All views are ventral with anterior part to the left.
Mo1-neurod1 and Mo2-neurod1 morpholinos efficiently block the expression of GFP from the neurod1: egfp transgenic line. Lateral views of 26 to 30 hpf tg(neurod1:egfp) embryos injected with Mo1-neurod1 (A), Mo2-neurod1 (B) or control morpholinos analyzed by bright field illumination (A,B) or by epifluorescence illumination (A’,B’) showing that both morpholinos efficiently block the expression of GFP from the neurod1:egfp transgenic line [40] without disturbing the general morphology of the embryos.
TUNEL assays on control and morphant embryos. (A–D) Confocal image projections of 30 hpf and 40 hpf control and neurod1 morphants after TUNEL labeling for apoptotic cells (in green) and immunodetection of insulin cells. The arrows highlight individual TUNEL+ cells in the neural tube. No TUNEL+ cells were found in the pancreatic region of control or neurod1 morphants at analyzed stages. (E–H) Confocal image projections of 14S and 19S control and ascl1b/neurod1 double morphants after TUNEL labeling for apoptotic cells (in red) and immunodetection of GFP cells. No TUNEL+ cells were found in the pancreatic region of control or morphants at analyzed stages. Somites 1 (S1) to somites 4 (S4) are shown on the panels G and H.
The double ascl1b/neurod1 morphants do not display general developmental defects. Bright field views of wt embryos injected with Mo-ascl1b and Mo2-neurod1 morpholinos (A) or control morpholinos (B) showing no general developmental defects in the double morphants at 22 hpf.