Abstract
Background
Blood group genotyping is increasingly utilized for prenatal diagnosis and after recent transfusions, but still lacks the specificity of serology. In whites, the presence of antigen D is predicted, if two or more properly selected RHD-specific polymorphism are detected. This prediction must fail, if an antigen D negative RHD positive allele is encountered. Excluding RHDψ and CdeS frequent only in individuals of African descent, most of these alleles are unknown and the population frequency of any such allele has not been determined.
Methods
We screened 8,442 antigen D negative blood donations by RHD PCR-SSP. RHD PCR positive samples were further characterized by RHD exon specific PCR-SSP or sequencing. The phenotype of the identified alleles was checked and their frequencies in Germans were determined.
Results
We detected 50 RHD positive samples. Fifteen samples harbored one of three new Del alleles. Thirty samples were due to 14 different D negative alleles, only 5 of which were previously known. Nine of the 14 alleles may have been generated by gene conversion in cis, for which we proposed a mechanism triggered by hairpin formation of chromosomal DNA. The cumulative population frequency of the 14 D negative alleles was 1:1,500. Five samples represented a D+/- chimera, a weak D and three partial D, which had been missed by routine serology; two recipients transfused with blood of the D+/- chimera donor became anti-D immunized.
Conclusion
The results of this study allowed to devise an improved RHD genotyping strategy, the false-positive rate of which was lower than 1:10,000. The number of characterized RHD positive antigen D negative and Del alleles was more than doubled and their population frequencies in Europe were defined.
Introduction
The antigen D encoded by the RHD gene is the most important blood group antigen determined by a protein. About 15% of whites are antigen D negative. Antigen D prediction by PCR was initially applied to fetus at risk for hemolytic disease of the newbom [1,2]. If serologic blood group typing cannot be performed with its usual ease, an RHD genotyping with a specificity and sensitivity comparable to serologic methods is of practical importance. For example, the utility of blood group genotyping in patients with recent transfusions was demonstrated by several studies [3,4,5,6].
The two RH genes, RHD and RHCE are about 30,000 bp apart [7], have opposite orientation [7,8] and are homologous retaining more than 90% identity [9]. The most frequent cause for the absence of the antigen D in whites is the lack of the whole RHD gene [10] due to a deletion occurring in the Rhesus box [7]. Therefore, most methods for antigen D prediction in whites probed the presence of RHD specific polymorphism [1,2,11,12,13]. Two RHD positive antigen D negative alleles are frequent in Africans: RHDψ carries a 37 bp insertion at the intron 3/exon 4 boundary and also harbors a stop codon [14]; Cdes is a RHD-CE-D hybrid gene [15,16,17]. In Asians, a major allele may be associated with a G314V missense mutation [18], and several other alleles may represent RHD-CE-D hybrid alleles [18,19,20].
In whites, RHD positive antigen D negative alleles were considered rare. However, the single systematic study [21] indicated a frequency of up to 22% among the rare haplotype Cde, which would render them the major cause of false-positive antigen D prediction by PCR in whites. The majority of RHD positive alleles in D negatives were reported as scattered case reports [22,23,24,25,26,27] with an often incomplete molecular work-up. The relative frequencies of these alleles and their cumulative population frequency remained unknown.
The specificity of RHD genotyping can be improved by a systematic characterization of RHD positive antigen D negative alleles. This rationale prompted us to determine the molecular causes of such alleles and their population frequencies in a random survey among European blood bank donors. We screened more than 8,000 antigen D negative blood donations by RHD PCR, including more than 700 rare Ccddee or ccddEe samples. Nine RHD-CE-D hybrid alleles, 5 other D negative and 3 Del alleles were identified. Five D positive donors missed by routine serology were uncovered. Two anti-D immunizations were traced. We established frequency estimates for RHD positive antigen D negative haplotypes in whites, which allowed us to devise an optimized RHD PCR strategy with an enhanced and defined specificity.
Results
Population surveys
In a first survey, we investigated 1,068 samples of blood donors that were documented as antigen D negative according to routine serologic methods. To cover the whole length of the RHD gene, we tested the RHD promoter, intron 4, exon 7, and the 3' untranslated region of exon 10 by PCR-SSP (Table 1). As antigen D negative RHD gene positive alleles are known to preferentially occur in the Cde and cdE haplotypes [21,22,25], we tested 754 samples with antigen C or antigen E or both along with 314 ccddee samples. We detected 48 donors who carried the RHD gene. All were positive for antigen C or antigen E or both (Table 1).
Table 1.
Population surveys of D negative blood donors documented D negative and screened by RHD PCR-SSP
| Samples (n) | |||
| Documented phenotype | screened | PCR-SSP positive | D positive* |
| Testing as single samples† | |||
| Ccddee | 433 | 34 | 0 |
| ccddEe | 271 | 5 | 2 |
| CCddee | 24 | 4 | 0 |
| CcddEe | 19 | 4 | 2 |
| ccddEE | 6 | 1 | 0 |
| CcddEE | 1 | 0 | 0 |
| ccddee | 314 | 0 | 0 |
| Testing as pools of 20 samples‡ | |||
| ccddee | 7,374 | 2 | 1 |
| Total | 50 | 5 | |
*Samples uncovered on further analysis as weak D, partial D or D+/- chimera. † Positive for at least one of four RHD specific polymorphism tested (promoter, intron 4, exon 7 and 3' UTR). ‡Positive for at least one of three RHD specific polymorphism tested (promoter, intron 4, and 3' UTR).
In a subsequent survey, we checked 7,374 ccddee samples, which were tested in pools of twenty samples for RHD promoter, intron 4 and exon 10. This survey aimed to increase the power of our study for ccddee donors, which represent 92% of all antigen D negative [28]. Two RHD positive donors were detected (Table 1). In summary, 50 RHD positive donors were found in the two population surveys. They were further characterized by a detailed molecular work-up including RHD exon specific PCR-SSP, PCR of intron polymorphism or nucleotide sequencing.
Exclusion of five antigen D positive donors
The molecular and serologic work-up revealed that 5 donors, previously documented as antigen D negative, were weakly antigen D positive (Table 1). Two donors of phenotype CcDEe carried D category VI type I and weak D type 2, respectively. Two donors of phenotype ccDEe carried D category VI type I and the new partial D DIM [29], respectively. One donor of phenotype ccDee was a D+/- chimera.
Molecular analysis of 45 antigen D negative RHD gene positive samples
The remaining 45 samples were investigated by RHD exon specific PCR for exons 3, 4, 5, 6, 7, and 9. Samples with discrepant results for RHD promoter and exon 3 were investigated for intron 1 and intron 2, those with discrepant results for exon 7 and 9 were investigated for intron 7 and intron 8, and those with discrepant results for exon 9 and exon 10 were investigated for intron 9. 24 samples could be assigned to one of nine distinct RHD PCR patterns and 21 samples were positive for all RHD specific polymorphism investigated (Fig. 1). (i) Hybrid alleles. The RHD PCR patterns could be explained by nine RHD-CE-D hybrid alleles (Fig. 1A). Only two of these alleles could be definitively related to prior descriptions of RHD positive antigen D negative alleles: One of the three carriers of the RHD-CE(8-9)-D allele was a donor previously communicated by us as "CCDnex ee", who was negative in an RHD exon 9 PCR [27]. Another pattern was identified as Cdes (Fig. 2 to Fig. 4). We cannot exclude the possibility that some of the seven remaining alleles have been observed previously [18,19,20,21,22,24,25,26]. Because of the limited published data for those observations, we found more than one "compatible" allele in our study for each previous observation. It should be noted that the hybrid structure was predicted from the PCR pattern and alternative explanations like combinations of two hybrid genes or partial deletions of the RHD gene were not formally excluded, (ii) Other alleles. The twenty-one samples positive for the nine RHD specific polymorphism tested were assigned to one of eight different RHD alleles (Fig. 1B). One allele was identical with RHDψ [14], the other seven alleles were novel. Each allele was characterized by nucleotide sequencing of the ten RHD exons in at least one sample. Once a new allele was characterized, the remaining samples were assigned by nucleotide sequencing of the informative exons (Fig. 1B).
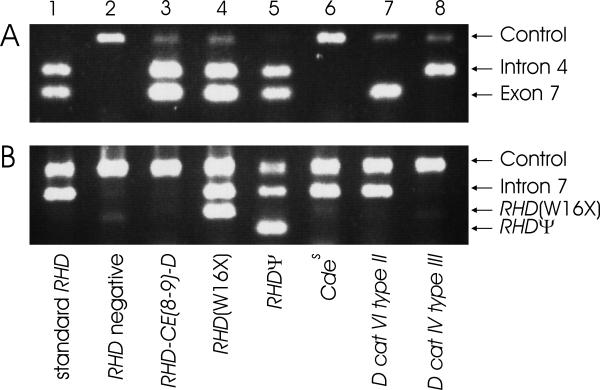
Figure 1.
Predicted molecular structure of the 17 RHD positive, D negative or Del alleles detected. For each haplotype, a schematic representation of the molecular structure is shown along with a designation, haplotype association, phenotype, and numbers of samples observed. Each RHD exon is indicated by a box, intron and promoter polymorphism investigated are shown as circles. White symbols indicate the presence of RHD specific sequences, black symbols their lack as predicted form the RHD exon-specific PCR-SSP results. Exons 1, 2, and 8 are shown in gray, because they are identical in RHD and some RHCE alleles. Panel A: Hybrid alleles. The molecular structures are represented as single hybrid alleles; it should be noted that the PCR patterns could also be caused by combinations of hybrid alleles or by partial RHD deletions. Panel B: Other alleles. The nature of the aberration is indicated, and its position visualized by a vertical bar. The RHD(M295I) allele is similar to weak D type 11 [32] but represents a different haplotype and phenotype.
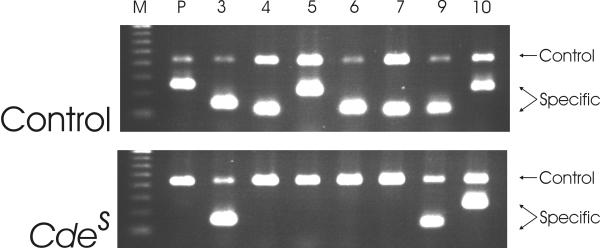
Figure 2.
RHD exon specific PCR-SSP of Cdes. In an RHD positive control, RHD specific PCR products are obtained for the RHD promoter (lane marked P, 255 bp), exon 3 (154 bp), exon 4 (123 bp), exon 5 (228 bp), exon 6 (133 bp), exon 7 (123 bp), exon 9 (119 bp) and exon 10 (232 bp). The 434 bp control product derives from the HGH gene. In the Cdessample, RHD specific amplicons are obtained for exon 3, exon 9, and exon 10, only.
Figure 4.
Molecular cause of the negative RHD promoter PCR in Cdes. The nucleotide sequence of the Cdes, RHD and RHCE promoter reaching from about 1550 to 901 bp 5' of the A of the start codon is shown. The positions of the RHD specific primers re011d and re012 used for the RHD promoter PCR are given. Nucleotides indicating RHD or RHCE origin of the Cdes sequence are highlighted. The Cdes promoter sequence represents RHD. A small DNA stretch of at least 13 bp in the region of re012 is replaced by the corresponding sequence of RHCE. This gene conversion caused the negative result obtained in the RHD promoter PCR-SSP (Fig. 2). RHCE nucleotide sequence is according to GenBank accession number AL031284.
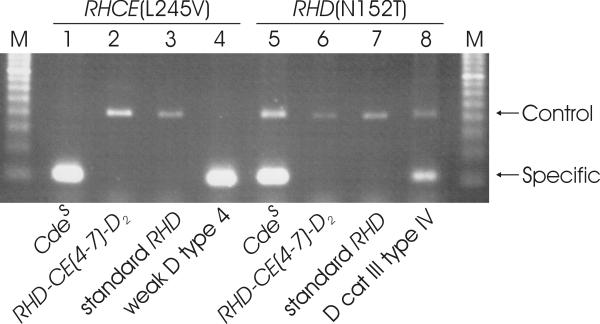
Figure 3.
Demonstration of the RHCE(L245V) and RHD(N152T) substitutions characteristic of Cdes. PCR-SSP were performed to detect single nucleotide polymorphism characteristic for Cdes [16] and indicative of RHCE(L245V) (lanes 1 to 4, 110 bp specific product) and RHD(N152T) (lanes 5 to 8, 120 bp specific product). Both polymorphism were present in the Cdes sample as expected (lanes 1 and 5). The RHD-CE(4-7)-D2 sample was compatible with Cdes according to the RHD exon specific PCR (Fig. 1) but lacked both polymorphism (lanes 2 and 6). Negative controls were standard RHD (lanes 3 and 7), positive controls weak D type 4 (lane 4) and DIII type IV (lane 8), respectively.
Del phenotype
Del is defined by expressing trace amounts of antigen D that can be detected by an adsorption/elution study only [30]. Because current routine serology cannot discriminate D negative from the Del phenotype, at least one RBC sample of each allele (Fig. 1) was tested by adsorption and elution. Three alleles represented the Del phenotype (Fig. 1B) and were characterized by one missense and two splice site mutations, respectively. For each allele, only a single sample was sequenced, and the influence of the mutations on mRNA splicing was not verified by cDNA analysis. Because lack of material, we could not formally exclude the Del phenotype for the two RHD-CE(2-7)-D1 samples (Fig. 1A). However, a Del expressed by this allele was very unlikely, as several other hybrid alleles carrying smaller gene conversions, like RHD-CE(2-7)-D2, were unequivocally D negative.
Population frequencies
The population frequencies of the alleles were calculated (Table 2). The cumulative frequency of all antigen D negative RHD gene positive haplotypes was estimated to be 1:1,537. The most frequent allele was RHD-CE(2-9)-D2 with a frequency of 1:5,682, representing about 27% of antigen D negative RHD gene positive alleles. Hybrid alleles lacking RHD exon 4 to exon 7 accounted for 68% of antigen D negative RHD gene positive alleles. 84% of antigen D negative RHD positive haplotypes carried the antigen C, compared to less than 3% of all D negative haplotypes [28]. The cumulative allele frequency of Del was 1:3,030.
Table 2.
Estimated population frequencies for antigen D negative RHD positive and Del haplotypes in Europeans
| Frequencies | ||||
| In population | Within haplotype | |||
| Allele | Estimate | 95% confidence interval | Estimate | Haplotype |
| D negative alleles | ||||
| RHD-CE(2-9)-D2 | 1:5,682 | 1:3,046 - 1:13,837 | 1:62 | Cde |
| RHD-CE(2-9)-D1 | 1:15,152 | 1:5,610 - 1:55,568 | 1:167 | Cde |
| RHD-CE(8-9)-D | 1:15,152 | 1:5,610 - 1:55,568 | 1:167 | Cde |
| RHD-CE(4-7)-D1 | 1:18,036 | 1:6,678 - 1:66,145 | 1:101 | cdE |
| RHD-CE(2-7)-D1 | 1:22,727 | 1:6,798 - 1:128,041 | 1:250 | Cde |
| RHD-CE(2-7)-D2 | 1:22,727 | 1:6,798 - 1:128,041 | 1:250 | Cde |
| RHD(W16X) | 1:22,727 | 1:6,798 - 1:128,041 | 1:250 | Cde |
| RHDψ | 1:37,431 | 1:7,032 - 1:733,950 | 1:14,748 | cde |
| RHD-CE(4-7)-D2 | 1:45,455* | 1:8,539 - 1:891,266 | 1:500* | Cde or cdE |
| Cdes | 1:45,455 | 1:8,539 - 1:891,266 | 1:500 | Cedes |
| RHD(G212V) | 1:45,455 | 1:8,539 - 1:891,266 | 1:500 | Cde |
| RHD(Y330X) | 1:45,455 | 1:8,539 - 1:891,266 | 1:500 | Cde |
| RHD(IVS8+1G>A) | 1:45,455 | 1:8,539 - 1:891,266 | 1:500 | Cde |
| RHCE(1-9)-D | 1:54,107 | 1:10,164 - 1:1,060,924 | 1:303 | cdE |
| associated with Cde | 1:1,818 | 1:1,262 -1:2,711 | 1:20 | Cde |
| associated with cdE | 1:13,527 | 1:5,638 - 1:39,610 | 1:75 | cdE |
| associated with cde | 1:37,431 | 1:7,032 - 1:733,950 | 1:14,748 | cde |
| Total | 1:1,537 | not applicable | not applicable | |
| Delalleles | ||||
| RHD(M295I) | 1:6,493 | 1:3,302 - 1:13,837 | 1:71 | Cde |
| RHD(K409K)† | 1:9,091 | 1:4,067 - 1:23,073 | 1:100 | Cde |
| RHD(IVS3+1G>A) | 1:15,152 | 1:5,610 - 1:55,568 | 1:167 | Cde |
| Total | 1:3,030 | 1:1,913 - 1:5,610 | 1:33 | Cde |
* Assuming a Cde haplotype; a cdE haplotype would result in a frequency of 1: 54,107 (95% confidence interval: 1:10,164 - 1:1,060,924; frequency within haplotype 1:303). † Silent mutation adjacent to an intron/exon boundary, probably affecting splicing.
Analysis of SCARF samples
We obtained 2 ccddEE, 1 CcddEe and 1 ccddee (G+) DNA samples through the SCARF Exchange and tested them for promoter, intron 4, exon 7 and the 3' untranslated region by RHD PCR-SSP. Positive reactions were obtained with the CcddEe sample only, which was assigned to the RHD-CE(8-9)-D allele by RHD exon-specific PCR-SSP.
Optimized RHD PCR
Based on the population frequencies, we calculated the expected positive predictive values of a positive result for different RHD PCR strategies (Table 3). RHD PCR based on intron 4 and exon 7 had a considerably higher positive predictive value than testing exon 10 alone. Testing for RHDψ [14] improved specificity. Even greater improvements were effected by testing for other allele like RHD-CE(8-9)-D or RHD(W16X). An optimized PCR strategy would comprise checking for RHD intron 4, exon 7 and intron 7 complemented by the specific detection of RHD(W16X) and RHDψ Antigen prediction in the rare samples positive for either of these alleles necessitates complementary methods, since the allele in trans may be D positive. The five polymorphism tested can be multiplexed in two PCR tubes (Fig. 5). This assay was about twice as reliable as current exon scanning approaches [6,26] requiring, if not multiplexed [26], up to 8 separate PCR tubes [6].
Table 3.
Expected rates of false positive results and expected positive predictive values for differentRHD PCR strategies*
| Rate of | Positive predictive value | Number of | |
| PCR strategy | false positives | of positive result | polymorphism tested |
| Exon 10 only [1,2] | 1:1,276 | 0.999216 | 1 |
| Intron 4/Exon 7 [13] | 1:4,081 | 0.999755 | 2 |
| Intron 4/Exon 7/RHDψ [14] | 1:4,700 | 0.999787 | 3 |
| Intron 4/Exon 7/W16X | 1:5,212 | 0.999808 | 3 |
| Intron 4/Exon 7/Intron 7 | 1:6,051 | 0.999835 | 3 |
| Exons 3, 4, 5, 6, 7, 9 [26] | 1:6,051 | 0.999835 | 6 |
| Exons 2, 3, 4, 5, 6, 7, 9, 10 [27] | 1:6,051 | 0.999835 | 8 |
| Intron 4/Exon 7/W16X/RHDψ | 1:6,267 | 0.999840 | 4 |
| All Exons/RHDψ | 1:7,520 | 0.999867 | 9 |
| Intron 4/Exon 7/Intron 7/W16X | 1:8,921 | 0.999888 | 4 |
| Intron 4/Exon 7/Intron 7/W16X/RHDψ | 1:12,533 | 0.999920 | 5 |
* Rates were calculated based on the population frequencies of different alleles determined in the Table 2. The exact rates are population dependent and may vary according to the prevalence of alleles in the population tested.
Figure 5.
RHD PCR-SSP optimized for specificity. The PCR is performed as a modular system consisting of two multiplex reactions. An RHD intron 4/exon 7 multiplex PCR-SSP (Panel A) is combined with an RHD intron 7 PCR that is multiplexed with reactions for the specific detection of RHD(W16X) and RHDψ (Panel B). Results are shown for a normal D positive sample (lane 1), a normal D negative sample (lane 2), several rare D negative samples (lanes 3 to 6) and major D positive RHD variants (lanes 7 and 8). Standard D positive and D negative samples and D categories VI and IV are recognized in panel A. RHD-CE(8-9)-D is detected in panel B by the absence of the intron 7 band (lane 3). The presence of RHD(W16X) and RHDψ is detected in panel B because of their specific amplicons (lanes 4 and 5). Amplicon size is Panel A, control, 434 bp (HGH gene); intron 4, 226 bp; exon 7, 123 bp; Panel B, control, 659 bp (chromosome 1 genomic sequence about 90,000 bp 5' of Rhesus box); intron 7, 390 bp; RHD(W16X), 248 bp; RHDψ, 154 bp. The internal control amplicons, which were devised to be larger than the specific amplicons, may be suppressed because of competition, if a specific product is amplified.
Gene conversions in cis
The gene conversion of five hybrid alleles observed in this study involved intron 2 (Fig. 1) which was utilized to delineate the allele origin of the RHCE gene segment found in the RHD gene. In all five haplotypes, a gene conversion in cis was likely (Fig. 6A). We proposed that gene conversions in cis occur during hairpin formation, which is favored by the clustered gene arrangement (Fig. 6B).
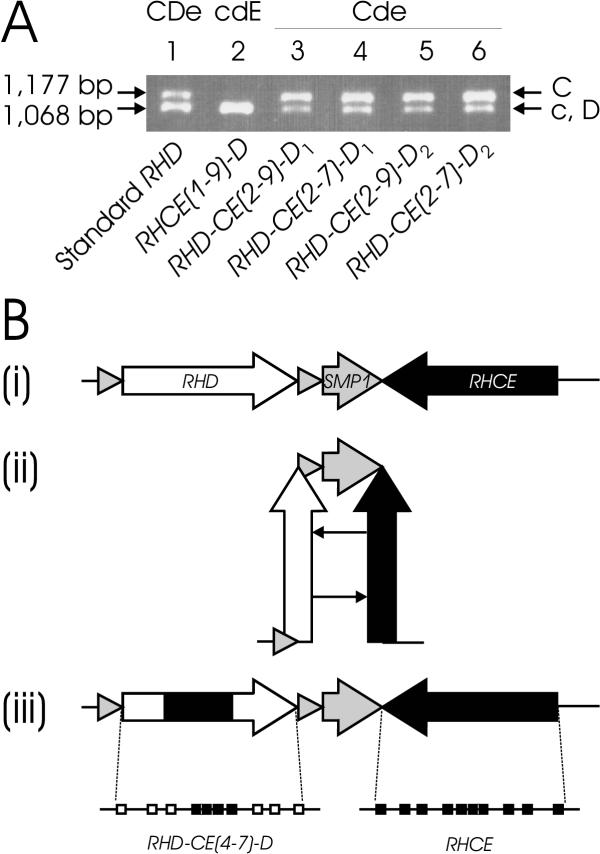
Figure 6.
Gene conversion in cis. Panel A: Origin of the RHCE gene segments. The allele origin of the RHCE segments in the RHD gene was analyzed by a PCR length polymorphism in intron 2 [31,43]. The 1,177 bp product is specific for the C allele of RHCE, the 1,068 bp product for the c allele of RHCE and for RHD. The CcDee control shows a strong band at the c/D position and a weaker band at the C position (lane 1). The cE associated RHCE(1-9)-D hybrid allele lacks the C band (lane 2), indicating that the intron 2 of the hybrid allele derives from c. In contrast, all Ce associated hybrid alleles involving intron 2 show a strong C band and a weaker c/D band (lanes 3 to 6), indicating that the introns 2 of those hybrids derive from C. Panel B: Proposed mechanism of gene conversion in cis. (i) The RHD and RHCE genes are inversely orientated [7] as typical for clustered genes. (ii) A putative hairpin formation of the chromosome allows the close proximity of homologous segments in identical orientation. This structural feature is instrumental for gene conversion events in cis. (iii) Resolving the hairpin yields an RHD-CE-D hybrid gene structure, many of which have been observed to date at the RH gene locus. As an example, the RHD-CE(4-7)-D hybrid exon structure is shown. Symbols are according to Fig. 1.
Anti-D immunizations
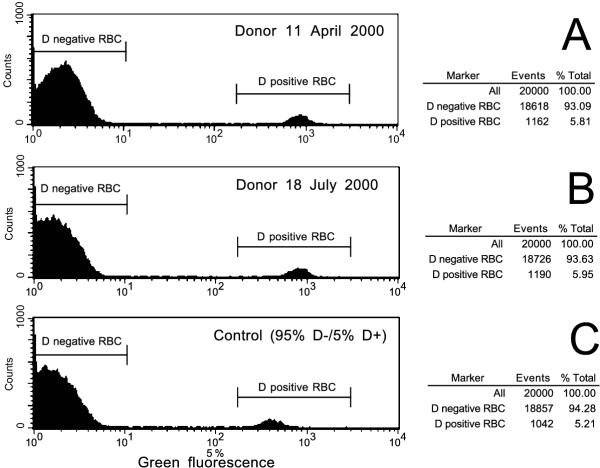
A flow cytometry study of the RBC from the D+/- chimera revealed 94% D negative RBC and an admixture of 6 % D positive RBC (Fig. 7). This chimera was confirmed in a 3 month follow-up. The 24 year old donor was healthy and had no twin. A look-back of this donor revealed 13 units that had been issued as D negative. Two D negative recipients were traced and available for an antibody screen, both of whom were anti-D immunized.
Figure 7.
Flow cytometric analysis of a D+/- chimera. The fluorescence histograms obtained by indirect immunofluorescence with a polyclonal anti-D are shown for the index donation of the chimerical donor (Panel A), a second donation three month later (Panel B) and a control mixture containing 5% D positive RBC (Panel C). There are two peaks separated by a large gap indicating that two different RBC populations are present. The left peak represents D negative RBC, the right peak D positive RBC carrying a normal strength antigen D. The positive RBC population of the donor was about 6%.
Discussion
In a systematic population survey including more than 8,000 antigen D negative blood donations, we identified 14 different RHD positive antigen D negative and 3 different Del haplotypes, the majority of which were novel. The molecular bases were alleles comprising RHD/RHCE hybrids, stop codons, missense mutations and splice site mutations. The cumulative frequency of RHD gene positive antigen D negative haplotypes was about 1:1,500; that of the Del alleles was about 1:3,000. We determined the specificity of antigen D prediction by PCR and devised an optimized RHD PCR strategy with a calculated positive predictive value greater than 0.9999. Five antigen D positive samples missed by routine D typing were uncovered and two anti-D immunizations traced.
For practical purposes, two groups of RHD alleles that do not express antigen D can be distinguished. RHD alleles of the first group lack some or many RHD specific polymorphism and usually represent RHD/CE hybrids. For alleles of this group, a correct antigen D prediction may be accomplished by a prudent selection of the RHD specific polymorphism utilized for RHD genotyping. RHD alleles of the second group carry all RHD specific polymorphism and most often harbor point mutations. For alleles of this group, a correct antigen D prediction necessitates the specific detection of an aberration that is usually unique to the allele. The identification of four new alleles in this group increased the number of known alleles from 3 to 7 and was critical for improving RHD genotyping.
The data of this study allowed for the first time to calculate population frequencies of RHD positive antigen D negative and Del alleles. This information was indispensable to derive rational RHD typing strategies and will be essential for establishing cost-efficient approaches. The majority of samples belonging to the first group of D negative alleles (probable RHD/CE hybrids) was compatible with RHD-CE-D hybrid alleles, in which the DNA segment derived from the RHCE gene encompassed at least exon 4 to exon 7. These samples would be correctly typed, if exon 4/intron 4 and exon 7 were used for RHD genotyping, as proposed previously [13]. With the exception of RHD exon 9, testing additional RHD exons would not have improved the specificity of antigen D prediction. Improving this specificity, however, became possible by the specific detection of frequent alleles of the second group, like RHDψ and RHD(W16X). We demonstrated that testing 5 carefully selected polymorphism would have resulted in an assay yielding false positive results at a rate less than 1:12,000, and hence would have doubled the specificity compared to contemporary approaches testing all informative RHD exons [6,26]. Further improvements may be achieved by the specific detection of additional alleles, that might become practical in massively parallel molecular assays.
The detailed analysis including intron polymorphism revealed that the first group of alleles (probable RHD/CE hybrids) represented at least 9 different molecular events. We proposed that the proximity and inverse orientation of both RH genes favored gene conversions occurring in cis (Fig. 6), which have also been noted in partial D [31]. An exact definition of the molecular bases of the RHD/CE hybrids would allow their specific detection, even if they were positioned in trans to the regular RHD allele. Such a detection would be necessary, if molecular RH zygosity testing is expected to achieve the same specificity as antigen D prediction.
A considerable proportion of seemingly D negative samples carrying the RHD gene presented a Del phenotype. Interestingly, RHD(M295I) coded for weak D, if associated with a ce haplotype [32], but for Del, if associated with a Ce haplotype; this observation may be explained by the suppressive effect of C in cis [33].
The nature and frequency of RHD gene positive antigen D negative alleles differ among populations. Apart from a probably lower absolute frequency, we detected in Europeans many parallels to oriental populations: Both populations shared the diverse nature of RHD haplotypes of the first group (probable RHD/CE hybrids) [18,19,20], the preferential occurrence of RHD positive antigen D negative alleles in Cde haplotypes [18], and the comparatively frequent observation of Del phenotypes [19]. In contrast, RHDψ and Cdes are predominant in African populations [14]. Still another situation may be present in the middle-west USA, where 6 of 26 RHD gene positive antigen D negative samples had aberrations limited to a single exon yet detectable by PCR [34].
Blood group serologists might note the observation of 5 D positive samples in our study with disturbance. In many centers, donors are checked for antigen D by sensitive methods at first and second donations only. On subsequent donations, carriers of partial D, like DvI or DIM, some weak D and D+/- chimerism may pass unnoticed in tests based on direct agglutination, even with the most avid IgM anti-D. Immunizations caused by units of such donors will generally be missed, because the occurrence of an anti-D in a patient is usually not further investigated [35]. For example, the two anti-D immunizations induced by units of the chimerical donor of this study were found only in a look-back triggered by our molecular screen. Chimeras in the Rh system have repeatedly been observed [36,37] and chimeras may be a more widespread phenomenon than anticipated [38]. A lower antigen density threshold for anti-D immunization has not been established yet, and future studies might indicate a need to exclude even Del donors from transfusion to D negative recipients. A routine investigation of all samples by adsorption and elution is not feasible. However, checking D-negative samples, especially those occurring with a C or E or both, for RHD specific sequences by nucleic amplification techniques may become practical in the near future. The knowledge of the detected alleles is also important for fetal genotyping assays using fetal DNA in maternal plasma, because false positive results will be obtained in mothers harboring RHD positive D negative alleles.
Subjects and Methods
Blood samples
EDTA- or citrate-anticoagulated blood samples were collected from blood donors characterized as D negative in routine typing including an antiglobulin test with anti-D. The D antigen determination in antiglobulin technique was performed as part of routine blood donor typing over a period of more than 15 years with varying commercial anti-D in tube or column agglutination. For each donor, this antiglobulin test was done only once, if the donor lacked the antigens C or E, and two times from independent samples, if the donor was either C or E positive. Subsequent donations were checked by direct agglutination using an Olympus PK7200 autoanalyzer only. Samples were collected at random for specific CcEe phenotypes. DNA was isolated by a modified salting-out procedure as described [27,39].
Screening by PCR with sequence specific priming (PCR-SSP)
For the first population survey, 314 ccddee, 433 Ccddee, 271 ccddEe, 19 CcddEe, 24 CCddee, 1 CcddEE and 6 ccddEE samples were tested individually for the presence of RHD specific polymorphism located in the RHD promoter, intron 4, exon 7 and the 3' untranslated region of exon 10 by PCR-SSP. The donor previously reported as "CCDnexee" [27] returned by chance and his allele was further characterized as RHD-CE(8-9)-D.
For the second population survey, 7,374 ccddee samples were analyzed in pools. Equal volumes of 20 samples were mixed. To confirm the sensitivity of the pool testing, 1% of D positive blood was added to an aliquot as positive control. DNA was extracted and checked for RHD promoter, intron 4, and exon 10 using modified PCR-SSP to enhance sensitivity. Repeated testing of donors was minimal, because the mandatory donation interval exceeded the collection period.
The donors were representative for the population currently living in the county of (Land) Baden-Württemberg. The ethnic origin of individual donors, in particular of those carrying Cdes or RHDψ was not identified. Independent of the population surveys, 2 ccddEE, 1 CcddEe and 1 ccddee DNA samples of unknown ethnic backgrounds were obtained from the SCARF Exchange (Hahnemann University, Philadelphia, USA).
Further molecular characterization
All samples positive for any of the above mentioned PCR-SSP assays were further investigated for the presence of RHD specific polymorphism in exon 3, exon 4, exon 5, exon 6, exon 7, and exon 9 by PCR-SSP. Samples positive for all PCR-SSP were sequenced, until they could be assigned to a distinct RHD allele. Samples negative for some PCR-SSP were checked for informative polymorphism in intron 1, intron 2, intron 7, intron 8, and intron 9.
Analysis for Cdes and RHDψ
RHD(N152T) and RHCE(L245V) present in Cdes [16] were checked by PCR-SSP. The 37 bp insertion present in RHDψ was detected by PCR-SSP. The 37 bp insertion, the M218I, F223V and S225F missense mutations and the Y269X nonsense mutation previously described for RHDψ [14] were confirmed by sequencing of all 10 exons; no additional aberrations were detected.
Nucleotide sequencing
The ten RHD exons were sequenced as described [29,32]. The promoter was amplified with primers rend31k (for RHD alleles) or re04 (for RHCE) and rb45 and a DNA stretch encompassing primer re012 was sequenced using primers re08 and re09.
RHD PCR
Most RHD PCR-SSP were similar to the RHD exon specific PCR-SSP previously described [27]. Cycling conditions consisted of an initial denaturation of 2 min at 94°C, followed by ten cycles of 10 s denaturation at 94°C and 1 min annealing/extension at 65°C; and finally 25 cycles of 30 s denaturation at 94°C, 1 min annealing at 61°C and 30 s extension at 72°C. 0.4 U Taq polymerase (Qiagen, Hilden, Germany) were used in a final volume of 10 μl. Primers (Table 4) were re012 and re011d for the promoter; re41 and rb12 for intron 4; ga71 and ga72 for exon 7 in the PCR-SSP screening; rea7 and rr4 for exon 10; ga31 and rb21 for exon 3; ga41 and ga42 for exon 4; rb24 and ga51 for exon 5; ga62 and ga61 for exon 6; rb26 and re71 for exon 7 in the molecular work-up; re83 and re94 for exon 9; rb51 and rb52 for intron 7; RhPsiF and RhPsiB for RHDψ ; Rh152Tb and ga31 for RHD(N152T); and Rh223Vf and Rh245Vb for RHCE(L245V). Primer concentrations were 0.2 μM except for exon 6 (0.1 μM), RHD(N152T)(0.3 μM), and intron 7 and exon 9 (both 0.4 μM). For most samples intron 4/exon 7 was tested as multiplex reaction containing 0.2 μM of exon 7 (ga71/ga72) and 0.1 μM of intron 4 primers. As internal control, two primers amplifying an HGH gene fragment were added in concentrations of 0.05 μM for promoter, intron 4, and exon 7 (ga71/ga72); 0.075 μM for exon 10; 0.1 μM for intron 7, RHDψ, RHD(N152T) and RHCE(L245V); 0.15 μM for exon 3, exon 4, exon 7 (rb26/re71), and exon 9; 0.2 μM for exon 5 and exon 6. Mg2+ concentration was 0.15 μM, except 0.4 μM for intron 7. For exon 6, 20 % solution Q (Qiagen) was added. To enhance sensitivity, the pools were tested with RHD primers in a concentration of 0.3 μM and HGH primers at 0.1 μM.
Intron 1 was tested 1173 and 1174 bp 5' of the intron 1/exon 2 boundary by RHD specific amplification of exon 2 as described [32]. Intron 2 was evaluated by PCR with length polymorphism as described [31]. The BamHI restriction site introduced by the 9 bp deletion in RHD intron 8 position 1114 to 1122 (Genbank accession number AL139426) was checked after amplification with primers re74 and re93 and digestion with BamHI. The 980 bp deletion starting at position 633 in RHD intron 9 (Genbank accession number AL139426) was evaluated using primers re93k and re916.
Optimized RHD PCR-SSP for routine DNA typing
Reaction A contained primers ga71 and ga72 at 0.3 μM, rb12 and re41 at 0.1 μM, and HGH primers at 0.1 μM. Mg2+ was at 0.175 μM. Reaction B contained primers RhPsiF and RhPsiB at 0.5 μM, re11d and RhX1f1 at 0.3 μM, re721 and rb9 at 0.2 μM and as control primers rend9b1 and rend9b2 at 0.2 μM. Mg2+ was at 0.15 μM.
Immunohematology
One sample of each RHD positive allele was evaluated by direct agglutination with two monoclonal anti-D (Seraclone anti-D, clone BS226; Biotest, Dreieich, Germany; and Frekaklon anti-D, clone MS201; Gull, Bad Homburg, Germany). Indirect antiglobulin test was done in a gel matrix test (LISS-Coombs 37°C, DiaMed-ID Micro Typing System, DiaMed, Cressier sur Morat, Switzerland) using an oligoclonal anti-D (Seraclone anti-D blend, clones H41 11B7, BS221 and BS232; Biotest). Samples reactive in gel matrix technique were further investigated using the monoclonal anti-D HM10, HM16, P3x61, P3x35, P3x212 11F1, P3x212 23B10, P3x241, P3x249, P3x290 (Diagast, Loos, France) and H41 11B7 (Biotest). The presence of a Del phenotype was determined by adsorption of 500 μl of a polyclonal anti-D (human incomplete anti-D; Lorne Laboratories, Reading, UK) to 500 μl packed red blood cells (RBC) for 1 h at 37°C and elution using a chloroform technique [35]. A detailed serologic report of the RHD(C285Y) sample, dubbed DIM [29], has been published separately.
Flow cytometry
Flow cytometry was performed as described [40,41] using a polyclonal anti-D (anti-D Molter; Ortho Clinical Diagnostics, Neckarsgmünd, Germany) as primary and goat anti-human IgG, Fab-fragment, FITC-conjugated (Dianova, Hamburg, Germany) as secondary antibody. Markers were set to encompass >99.5% of a D positive control and less than 0.5% of a D negative control. The percentage of cells in the marker area was evaluated.
Haplotype frequencies
For alleles observed more than once, the haplotype association with Cde and cdE was obvious, because of their repeated observations in association with the rare phenotypes Ccddee or ccddEe, respectively. Based on the paucity of RHD positive samples among the ccddee samples, alleles that were observed only once were assumed to be associated with the Cde or cdE haplotype rather than the cde haplotype. An allele occurring in an unique CcddEe sample was counted as Cde. The RHDψ allele was assumed to be associated with the ce(W16C) allele, because RHCE specific sequencing of exon 1 revealed a C/G heterozygosity at position 64 and ce(W16C) is almost absent from the cde haplotypes in our population [27]. The frequency of a given aberrant RHD allele in its haplotype was calculated as the number of observed samples divided by the number of the corresponding haplotypes under observation (500 Cde, 303 cdE). For cde, the haplotype frequency was calculated from the 14,748 haplotypes checked in the second survey. The population frequency of an RHD allele was calculated from the frequency of this allele in its haplotype and the known frequency of the haplotype in the local population [28]. Confidence intervals were calculated according to the Poisson distribution [42]. Donors were not generally checked for kinship. However, the three RHD-CE(8-9)-D donors were siblings; a fourth sample was independently observed in the single RHD positive DNA from the SCARF Exchange.
Table 4.
Primers used
| Genomic | |||||
| Name | Nucleotide sequence | region | Position* | Strandedness | specificity |
| ga31 | ttgtcggtgctgatctcagtgga | exon 3 | 362 to 383 | sense | RHD |
| ga41 | acatgatgcacatctacgtgttcgc | exon 4 | 503 to 527 | sense | RHD/RHCE |
| ga42 | cagacaaactgggtatcgttgctg | exon 4 | 625 to 602 | antisense | RHD/RHCE |
| ga51 | ctgctcaccttgctgatcttccc | intron 5/exon. | 5 8 to 787 | antisense | RHD |
| ga61 | caggtacttggctcccccgac | exon 6 | 936 to 916 | antisense | RHD |
| ga62 | ttatgtgcacagtgcggtgttgg | exon 6 | 804 to 826 | sense | RHD/RHCE |
| ga71 | gttgtaaccgagtgctggggattc | exon 7 | 944 to 967 | sense | RHD/RHCE |
| ga72 | tgccggctccgacggtatc | exon 7 | 1066 to 1048 | antisense | RHD |
| rb12 | tcctgaacctgctctgtgaagtgc | intron 4 | 198 to 175 | antisense | RHD |
| rb21 | aggtccctcctccagcac | intron 3 | 28 to 11 | antisense | RHD/RHCE |
| rb24 | agacctttggagcaggagtg | intron 4 | -53 to -34 | sense | RHD/RHCE |
| rb26 | aggggtgggtagggaatatg | intron 6 | -62 to -43 | sense | RHD/RHCE |
| rb45 | acactgttgrctgaatttcggtgc | intron 1 | 164 to 139 | antisense | RHD/RHCE |
| rb51 | gcatgacgtgttctgcctcttg | intron 7 | -3365 to - 3386 | antisense | RHD |
| rb52 | ccaggttgttaagcattgctgtacc | intron 7 | -3433 to -3409 | sense | RHD |
| re04 | aggtcacatccatttatcccactg | promoter | -2498 to -2474 | sense | RHD/RHCE |
| re08 | gggcttgggacttagttctaac | promoter | -858 to -879 | antisense | RHD/RHCE |
| re09 | cgactgggtgattaaaatctcc | promoter | -1280to-1259 | sense | RHD/RHCE |
| re011d | gcagccaacttcccctgtg | promoter | -883 to -905 | antisense | RHD |
| re012 | tccactttccacctccctgc | promoter | -1148to-1122 | sense | RHD |
| relld | agaagatgggggaatctttttcct | intron 1 | 129 to 106 | antisense | RHD/RHCE |
| re41 | cgatacccagtttgtctgccatgc | exon 4 | 608 to 631 | sense | RHD/RHCE |
| re71 | acccagcaagctgaagttgtagcc | exon 7 | 1,008 to 985 | antisense | RHD |
| re74 | tatccatgaggtgctgggaac | intron 7 | -244 to -224 | sense | RHD/RHCE |
| re721 | ctggaggctctgagaggttgag | intron 7 | -348 to -326 | sense | RHD |
| re83 | gagattaaaaatcctgtgctcca | intron 8 | -54 to - 34 | sense | RHD/RHCE |
| re93 | cacccgcatgtcagactatttggc | intron 9 | 320 to 297 | antisense | RHD/RHCE |
| re93k | gccaaatagtttgacatgcgggtg | intron 9 | 297 to 320 | sense | RHD/RHCE |
| re94 | cttggtcatcaaaatatttagcct | exon 9 | 1216 to 1193 | antisense | RHD |
| re916 | gtttttgaggcaaagtctcgctc | intron 9 | 1689 to 1666 | antisense | RHD/RHCE |
| rea7 | tgttgcctgcatttgtacgtgag | 3' UTR† | 1311 to 1333 | sense | RHD/RHCE |
| rend31k | cctccccaacccagacagaattag | AJ252311‡ | 8506 to 8529 | sense | not applicable |
| rend9b1 | cactgcacttggcaccattgag | AL031432 | 29489 to 29468 | antisense | not applicable |
| rend9b2 | ttccgaaggctgcttttccc | AL031432 | 28840 to 28859 | sense | not applicable |
| Rh152Tb | gatattactgatgaccatcctcatgg | exon3 | 480 to 455 | antisense | RHCE |
| Rh223Vf | ttgtggatgttctggccaagtg | exon 5 | 646 to 667 | sense | RHCE |
| Rh245Vb | gctgtcaccactctgactgctac | exon5 | 755 to 733 | antisense | RHD |
| RhPsiB | tctgatctttatcctccgttccctc | exon 4 | 601 to 577 | antisense | RHD |
| RhPsiF | agacagactaccacatgaacttac | intron 3 | -38 to-15 | sense | RHDψ |
| RhX1f1 | cgctgcctgcccctctga | exon 1 | 31 to 48 | sense | RHD(W16X) |
| rr4 | agcttactggatgaccacca | 3'UTR | 1,541 to 1,522 | antisense | RHD |
*The positions of the synthetic oligonucleotides are indicated relative to their distances from the first nucleotide position of the start codon ATG for all primers in the promoter and in the exons including the 3' untranslated part of exon 10, relative to their adjacent exon/intron boundaries of RHCE for primers in introns; and according to the numbering in the genomic sequences indicated. Primers rh1 [44], ga31 (previously dubbed D-3-383), ga41 (D-4-527), ga42 (D-4-602), ga51 (D-5-787), ga61 (D-6-916), ga62 (D-6-826), ga71 (D-7-967), ga72 (D-7-1048) [27], rb5, rb12, rb24, rh5, rh7 [31], rb21, rb26, re11d, re71, re74, re83, re93, rr4 [32], re012 [29], re011d and rea7 [7] have been published previously. † 5' UTR: 5' untranslated region of exon 1; 3' UTR: 3' untranslated region of exon 10. ‡ Accession number of nucleic acid sequence in EMBL/GenBank/DDBJ; AJ252311 represents upstream Rhesus box; AL031431 Chromosome 1 genomic clone dJ465N24.
Acknowledgments
Acknowledgement
We thank E. Andreas Scharberg, Baden-Baden, Germany for supplying blood samples and Joann M. Moulds, Philadelphia, for rare DNA samples from the SCARF Exchange program, and Beate Wagner, Bremen, Germany, for helpful discussions. We acknowledge the expert technical assistance of Marianne Lotsch, Sabine Zahn, Anita Hacker, Sabine Kaiser and Katharina Schmid. This study was supported by the DRK-Blutspendedienst Baden-Württemberg, Stuttgart, Germany, and the Universitätsklinikum Ulm (Forschungsförderungsprojekt P.531), and the Deutsche Gesellschaft für Transfusionsmedizin und Immunhämatologie (Project DGTI/fle/00-01).
Contributor Information
Franz F Wagner, Email: franz.wagner@medizin.uni-ulm.de.
Alexander Frohmajer, Email: a.frohmajer@12move.de.
Willy A Flegel, Email: willy.flegel@medizin.uni-ulm.de.
References
- Lo Y-MD, Bowell PJ, Selinger M, Mackenzie IZ, Chamberlain P, Gillmer MDG, Littlewood TJ, Fleming KA, Wainscoat JS. Prenatal determination of fetal RhD status by analysis of peripheral blood of rhesus negative mothers. Lancet. 1993;341:1147–1148. doi: 10.1016/0140-6736(93)93161-S. [DOI] [PubMed] [Google Scholar]
- Bennett PR, Le Van Kim C, Colin Y, Warwick RM, Cherif-Zahar B, Fisk NM, Cartron JP. Prenatal determination of fetal RhD type by DNA amplification. N Engl J Med. 1993;329:607–610. doi: 10.1056/NEJM199308263290903. [DOI] [PubMed] [Google Scholar]
- Eshleman JR, Shakin-Eshleman SH, Church A, Kant JA, Spitalnik SL. DNA typing of the human MN and Ss blood group antigens in amniotic fluid and following massive transfusion. Am J Clin Pathol. 1995;103:353–357. doi: 10.1093/ajcp/103.3.353. [DOI] [PubMed] [Google Scholar]
- Legler TJ, Eber SW, Lakomek M, Lynen R, Maas JH, Pekrun A, Repas-Humpe M, Schroter W, Kohler M. Application of RHD and RHCE genotyping for correct blood group determination in chronically transfused patients. Transfusion. 1999;39:852–855. doi: 10.1046/j.1537-2995.1999.39080852.x. [DOI] [PubMed] [Google Scholar]
- Reid ME, Rios M, Powell VI, Charles-Pierre D, Malavade V. DNA from blood samples can be used to genotype patients who have recently received a transfusion. Transfusion. 2000;40:48–53. doi: 10.1046/j.1537-2995.2000.40010048.x. [DOI] [PubMed] [Google Scholar]
- Rozman P, Dove T, Gassner C. Differentiation of autologous ABO, RHD, RHCE, KEL, JK, and FY blood group genotypes by analysis of peripheral blood samples of patients who have recently received multiple transfusions. Transfusion. 2000;40:936–942. doi: 10.1046/j.1537-2995.2000.40080936.x. [DOI] [PubMed] [Google Scholar]
- Wagner FF, Flegel WA. RHD gene deletion occurred in the Rhesus box. Blood. 2000;95:3662–3668. [PubMed] [Google Scholar]
- Suto Y, Ishikawa Y, Hyodo H, Uchikawa M, Juji T. Gene organization and rearrangements at the human Rhesus blood group locus revealed by fiber-FISH analysis. Hum Genet. 2000;106:164–171. doi: 10.1007/s004390051024. [DOI] [PubMed] [Google Scholar]
- Okuda H, Suganuma H, Kamesaki T, Kumada M, Tsudo N, Omi T, Iwamoto S, Kajii E. The analysis of nucleotide substitutions, gaps, and recombination events between RHD and RHCE genes through complete sequencing. Biochem Biophys Res Commun. 2000;274:670–683. doi: 10.1006/bbrc.2000.3206. [DOI] [PubMed] [Google Scholar]
- Colin Y, Cherif-Zahar B, Le Van Kim C, Raynal V, van Huffel V, Cartron JP. Genetic basis of the RhD-positive and RhD-negative blood group polymorphism as determined by southern analysis. Blood. 1991;78:2747–2752. [PubMed] [Google Scholar]
- Aubin JT, Le Van Kim C, Mouro I, Colin Y, Bignozzi C, Brossard Y, Cartron JP. Specificity and sensitivity of RHD genotyping methods by PCR-based DNA amplification. Br J Haematol. 1997;98:356–364. doi: 10.1046/j.1365-2141.1997.2193040.x. [DOI] [PubMed] [Google Scholar]
- Hyland CA, Wolter LC, Saul A. Identification and analysis of RH genes: application of PCR and RFLP typing tests. Transfus Med Rev. 1995;9:289–301. doi: 10.1016/s0887-7963(05)80077-x. [DOI] [PubMed] [Google Scholar]
- Flegel WA, Wagner FF, Muller TH, Gassner C. Rh phenotype prediction by DNA typing and its application to practice. Transfus Med. 1998;8:281–302. doi: 10.1046/j.1365-3148.1998.00173.x. [DOI] [PubMed] [Google Scholar]
- Singleton BK, Green CA, Avent ND, Martin PG, Smart E, Daka A, Narter-Olaga NG, LM Hawthorne, G Daniels. The presence of an RHD pseudogene containing a 37 base pair duplication and a nonsense mutation in africans with the Rh D- negative blood group phenotype. Blood. 2000;95:12–18. [PubMed] [Google Scholar]
- Blunt T, Daniels G, Carritt B. Serotype switching in a partially deleted RHD gene. Vox Sang. 1994;67:397–401. doi: 10.1111/j.1423-0410.1994.tb01280.x. [DOI] [PubMed] [Google Scholar]
- Faas BHW, Becker EAM, Wildoer P, Ligthart PC, Overbeeke MAM, Zondervan HA, von dem Borne AEGK, van der School CE. Molecular background of VS and weak C expression in blacks. Transfusion. 1997;37:38–44. doi: 10.1046/j.1537-2995.1997.37197176949.x. [DOI] [PubMed] [Google Scholar]
- Daniels GL, Faas BH, Green CA, Smart E, Maaskant-van Wijk PA, Avent ND, Zondervan HA, von dem Borne AE, van der School CE. The VS and V blood group polymorphisms in Africans: a serologic and molecular analysis. Transfusion. 1998;38:951–958. doi: 10.1046/j.1537-2995.1998.381098440860.x. [DOI] [PubMed] [Google Scholar]
- Okuda H, Kawano M, Iwamoto S, Tanaka M, Seno T, Okubo Y, Kajii E. The RHD gene is highly detectable in RhD-negative Japanese donors. J Clin Invest. 1997;100:373–379. doi: 10.1172/JCI119543. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun C, Chou C, Lai N, Wang W. RHD gene polymorphisms among RhD-Negative Chinese in Taiwan. Vox Sang. 1998;75:52–57. doi: 10.1159/000030958. [DOI] [PubMed] [Google Scholar]
- Lan JC, Chen Q, Wu DL, Ding H, Pong DB, Zhao T. Genetic polymorphism of RhD-negative associated haplotypes in the Chinese. J Hum Genet. 2000;45:224–227. doi: 10.1007/s100380050006. [DOI] [PubMed] [Google Scholar]
- Avent ND, Martin PG, Armstrong-Fisher SS, Liu W, Finning KM, Maddocks D, Urbaniak SJ. Evidence of genetic diversity underlying Rh D negative, weak D (D") and partial D phenotypes as determined by multiplex PCR analysis of the RHD gene. Blood. 1997;89:2568–2577. [PubMed] [Google Scholar]
- Hyland CA, Wolter LC, Saul A. Three unrelated Rh D gene polymorphisms identified among blood donors with Rhesus CCee (r'r') phenotypes. Blood. 1994;84:321–324. [PubMed] [Google Scholar]
- Andrews KT, Wolter LC, Saul A, Hyland CA. The RhD- trait in a white patient with the RhCCee phenotype attributed to a four-nucleotide deletion in the RHD gene. Blood. 1998;92:1839–1840. [PubMed] [Google Scholar]
- Huang CH. Alteration of RH gene structure and expression in human dCCee and DCW- red blood cells: phenotypic homozygosity versus genotypic heterozygosity. Blood. 1996;88:2326–2333. [PubMed] [Google Scholar]
- Faas BHW, Beckers EAM, Simsek S, Overbeeke MAM, Pepper R, van Rhenen DJ, von dem Borne AEGK, van der School CE. Involvement of Ser103 of the Rh polypeptides in G epitope formation. Transfusion. 1996;36:506–511. doi: 10.1046/j.1537-2995.1996.36696269508.x. [DOI] [PubMed] [Google Scholar]
- Maaskant-van Wijk PA, Faas BHW, de Ruijter JAM, Overbeeke MAM, von dem Borne AEGK, van Rhenen DJ, van der Schoot CE. Genotyping of RHD by multiplex polymerase chain reaction analysis of six RHD-specific exons. Transfusion. 1998;38:1015–1021. doi: 10.1046/j.1537-2995.1998.38111299056309.x. [DOI] [PubMed] [Google Scholar]
- Gassner C, Schmarda A, Kilga-Nogler S, Jenny-Feldkircher B, Rainer E, Muller TH, Wagner FF, Flegel WA, Schonitzer D. RhesusD/CE typing by polymerase chain reaction using sequence-specific primers. Transfusion. 1997;37:1020–1026. doi: 10.1046/j.1537-2995.1997.371098016439.x. [DOI] [PubMed] [Google Scholar]
- Wagner FF, Kasuike D, Kerowgan M, Flegel WA. Frequencies of the blood groups ABO, Rhesus, D category VI, Kell, and of clinically relevant high-frequency antigens in South-Western Germany. Infusionsther Transfusionsmed. 1995;22:285–290. doi: 10.1159/000223144. [DOI] [PubMed] [Google Scholar]
- Wagner FF, Frohmajer A, Ladewig B, Eicher NI, Lonicer CB, Müller TH, Siegel MH, Flegel WA. Weak D alleles express distinct phenotypes. Blood. 2000;95:2699–2708. [PubMed] [Google Scholar]
- Okubo Y, Yamaguchi H, Tomita T, Nagao N. A D variant, Del? Transfusion. 1984;24:542. doi: 10.1046/j.1537-2995.1984.24685066827.x. [DOI] [PubMed] [Google Scholar]
- Wagner FF, Gassner C, Müller TH, Schönïtzer D, Schunter F, Flegel WA. Three molecular structures cause Rhesus D category VI phenotypes with distinct immunohematologic features. Blood. 1998;91:2157–2168. [PubMed] [Google Scholar]
- Wagner FF, Gassner C, Müller TH, Schönïtzer D, Schunter F, Flegel WA. Molecular basis of weak D phenotypes. Blood. 1999;93:385–393. [PubMed] [Google Scholar]
- Araszkiewicz P, Szymanski IO. Quantitative studies on the Rh-antigen D. Effect of the C gene. Transfusion. 1987;27:257–261. doi: 10.1046/j.1537-2995.1987.27387235634.x. [DOI] [PubMed] [Google Scholar]
- Allen RW, Ward S, Harris R. Prenatal genotyping for the RhD blood group antigen: considerations in developing an accurate test. Genet Test. 2001;4:377–381. doi: 10.1089/109065700750065126. [DOI] [PubMed] [Google Scholar]
- Flegel WA, Khull S, Wagner FF. Primary anti-D immunization by weak D type 2 RBC. Transfusion. 2000;40:428–434. doi: 10.1046/j.1537-2995.2000.40040428.x. [DOI] [PubMed] [Google Scholar]
- Northoff H, Goldmann SF, Lattke H, Steinbach P. A patient, mosaic for Rh and Fy antigens lacking other signs of chimerism or chromosomal disorder. Vox Sang. 1984;47:164–169. doi: 10.1111/j.1423-0410.1984.tb01578.x. [DOI] [PubMed] [Google Scholar]
- Salaru NN, Lay WH. Rh blood group mosaicism in a healthy elderly woman. Vox Sang. 1985;48:362–365. doi: 10.1111/j.1423-0410.1985.tb00197.x. [DOI] [PubMed] [Google Scholar]
- van Dijk BA, Boomsma DI, de Man AJ. Blood group chimerism in human multiple births is not rare. Am J Med Genet. 1996;61:264–268. doi: 10.1002/(SICI)1096-8628(19960122)61:3<264::AID-AJMG11>3.0.CO;2-R. [DOI] [PubMed] [Google Scholar]
- Miller SA, Dykes DD, Polesky HF. A simple salting-out procedure for extracting DNA from human nucleated cells. Nucl Acids Res. 1988;16:1215–1215. doi: 10.1093/nar/16.3.1215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wagner FF. Influence of Rh phenotype on the antigen density of C, c, and D: flow cytometric study using a frozen standard red cell. Transfusion. 1994;34:671–676. doi: 10.1046/j.1537-2995.1994.34894353461.x. [DOI] [PubMed] [Google Scholar]
- Wagner FF, Flegel WA. Analysis by flow cytometry of chimerism after bone-marrow transplantation and of erythrocyte antigen density. In Aspects of the Flow-Cytometric Analysis of Red Blood Cells (Edited by Gutensohn K, Sonneborn H-H, Kuehnl P) Heidelberg, Clin Lab Publications. 1997. pp. 95–103.
- Sachs L. Angewandte Statistik. 7th edition Berlin, springer. 1992.
- Poulter M, Kemp TJ, Carritt B. DNA-based Rhesus typing: simultaneous determination of RHC and RHD status using the polymerase chain reaction. Vox Sang. 1996;7:164–168. doi: 10.1111/j.1423-0410.1996.tb01316.x. [DOI] [PubMed] [Google Scholar]
- Simsek S, de Jong CAM, Cuijpers HTM, Bleeker PMM, Overbeeke MAM, Goldschmeding R, van der Schoot CE, von dem Borne AEGK. Sequence analysis of cDNA derived from reticulocyte mRNAs coding for Rh polypeptides and demonstration of E/e and C/c polymorphisms. Vox Sang. 1994;67:203–209. doi: 10.1111/j.1423-0410.1994.tb01661.x. [DOI] [PubMed] [Google Scholar]