Abstract
Bovine papillomavirus (BPV) is an oncogenic virus related to serious livestock diseases. Oncoproteins encoded by BPV are involved in several steps of cellular transformation and have been reported as presenting clastogenic effects in peripheral lymphocytes and primary culture cells. The aim of this study was to evaluate the clastogenic potential of BPV types 1, 2, and 4 by comet assay. Peripheral blood was collected from 37 bovines, 32 infected with different levels of papillomatosis (12 animals have no affection) and five calves, virus free (negative control). The viral identification showed presence of more than one virus type in 59.375% of the infected animals. Comet assay was performed according to alkaline technique. The Kruskal-Wallis test showed statistical difference between the negative control group and infected animals (P = 0.0015). The Dunn post hoc test showed difference comparing the infected animals with calves. Mann-Whitney U test verified no difference between animals infected with only one viral type and animals presenting more than one viral type. The comet assay is considered an efficient tool for assessment of damage in the host chromatin due to viral action, specifically highlighting viral activity in blood cells.
1. Introduction
Bovine papillomavirus (BPV) is a widespread oncogenic virus found worldwide belonging to the Papillomaviridae family, which displays tropism for squamous epithelial and mucosal tissues. These viruses are associated with benign and malignant epithelial lesions. Specifically, BPV presents a double-stranded circular DNA, not coiled, with approximately 8 kb, surrounded by an icosahedral capsid consisting of 360 copies of the L1 protein of 55 kDa, 72 capsomeres arranged in approximately 12 copies of the L2 protein, 39 kDa [1–8]. The papillomavirus genome is divided into three regions: early, late, and noncoding long control region (LCR), separated by two polyadenylation sites [3]. The early control region occupies 50% of the viral genome and encodes E1, E2, E3, E4, E5, E6, and E7 proteins. The late control region occupies 40% of the genome and contains the genes that codify L1 and L2 capsid proteins and LCR, which comprises 10% of the genome, with 850 bp. However, it also contains the origin of replication and the binding sites of multiple transcription factors [3]. Oncoproteins encoded by BPV are involved in several steps of the cell transformation [1, 9].
In cattle, the correlation between papillomavirus and cancer has been investigated in view of the economic costs generated by viral infection [1, 8, 10, 11]. BPV is the etiological agent of bovine papillomatosis, infectious disease, characterized by the presence of hyperproliferative skin lesions (papillomas), causing significant economic loss to livestock ranchers and can progress to cancer with the action of cofactors [8, 10, 12].
Currently, there are 13 BPV virus types described in the literature, although this number may be greater than twenty [7, 9]. The virus types are divided into three genres: Deltapapillomavirus (BPV-1, -2, and -13), Epsilonpapillomavirus (BPV-5 and -8), and Xipapillomavirus (BPV-3, -4, -6, -8, -9, -10, and -12), as well as the BPV-7 that remains not ranked in any genre [7]. Beside these, there are 16 new putative BPVs (BAA-1 to -4, BAPV-2 to -5, BAPV-7 to -10, BAPPV11MY and BPV/BR-UEL-2 to -5) [13]. According to Zhu et al. [13], BAA1 was detected in tongue epithelial papilloma, being designated BPV-12, and BPV/BR-UEL-4 described in ear cutaneous lesions was designated BPV-13 [9].
According to Stocco dos Santos et al. [1], papillomavirus can act on host chromatin causing cytogenetic alterations, such as changes in ploidy, chromatin gaps and breaks, dicentric chromosomes and rings. Significant increase of chromosomal aberrations was detected in animals infected with BPV, affecting genomic stability [4]. However, to date, there are no studies evaluating the BPV clastogenic potential in peripheral blood cells analyzed by comet assay.
Comet assay or single cell gel electrophoresis was introduced by Östling and Johanson [14] and later modified by Singh et al. [15]. The comet assay is a simple and versatile technique that requires few eukaryotic cells, as well as having a vast DNA damage spectrum detection capacity [16–19]. In the test, cells are engulfed by agarose gel and spread over the slide, and then subjected to an electric field that promotes free DNA fragment migration, with the appearance of a comet [15]. The nuclear region causes the head of the comet to fragment, and the length of the tail is directly related to the intensity of the damage.
The objective of this work was to evaluate the clastogenic potential of BPV types 1, 2, and 4 through comet assay in infected animals presenting cutaneous papillomatosis symptoms (hyperproliferative lesions-papillomas), asymptomatic (without papillomas) and calves, not infected, as negative control. The efficacy of the comet assay in clastogenic evaluations justifies this study.
2. Material and Methods
2.1. Ethics Statements
The protocols used in this study was approved by the Ethical Committee in Research of the Universidade Federal de São Paulo (Protocol number 1829/09) assigned by the President of this committee. All efforts were made to minimize any suffering in the animals.
2.2. BPV Diagnosis
2.2.1. Animal Selection
37 bovines (Bost aurus) were selected: 32 adults, 12 asymptomatic (without visible cutaneous papillomas) and 20 symptomatic (with visible cutaneous papillomas) and 5 newborn calves that were separated from their mother after birth and did not receive breast milk. The presence of papillomas in others organs was not evaluated due the absence of clinical characteristics that suggest bladder and/or esophageal cancer. The farm does not have the presence of bracken fern Pteridium aquilinum that is involved in oncogenic and mutagenic processes in function of quercetin presence [20]. These calves were isolated in order to obtain virus negative controls. Blood sample. The peripheral blood samples were collected using the EDTA vacutainer. Blood DNA extraction. The extraction of DNA from peripheral blood cells was performed using the GenomicPrep Blood Mini Kit Illustra Spin (GE Healthcare, Buckinghamshire, UK), which uses enzymatic digestion method with protease K, according to the manufacturer. The quality of obtained DNA was verified using PCR by amplifying a bovine β-globin gene fragment [21].
2.2.2. Viral Identification
Viral identification was performed using specific primers for BPV-1 (forward: 5′-GGAGCGCCTGCTAACTATAGGA-3′ and reverse: 5′-ATCTGTTGTTTGGGTGGTGAC-3′), which amplifies the L1 gene, resulting in a 301 bp amplicon, BPV-2 (forward: 5′-GTTATACCACCCAAAGAAGACCCT-3′ and reverse: 5′-CTGGTTGCAAÇAGCTCTCTTTCTC-3′), which amplifies the L2 gene, resulting in a 164 amplicon, and BPV-4 (forward; 5′-GCTGACCTTCCAGTCTTAAT-3′ and reverse; 5′-CAGTTTCAATCTCCTCTTCA-3′), which amplifies the E7 gene, resulting in a 170 bp amplicon. We choose these primers because the virus types are often observed in Brazil and in function of the association with esophageal (BPV-4) and bladder cancer (BPV-1 and -2). In detail, the amplification reactions were performed in a Veriti 96-well thermal cycler (Applied Biosystems, Singapore), with Master Mix (Promega, Madison, USA), under the following conditions: 5 minutes at 95°C, followed by 35 cycles of 1 minute and 30 seconds at 98°C, 2 minutes at 52°C, and 1 minute and 30 seconds at 72°C and a final extension step of 5 minutes at 72°C, for β-globin and specific primers. The PCR products were analyzed in 2% agarose gel electrophoresis stained with GelRed in TAE buffer, visualized under UV light. The images were captured through the software Kodak 1D 3.6.5. Cloned BPV-1, -2 and -4 genomes in Escherichia coli D5Hα were used as positive controls. These clones form part of the biological collection of Genetic Laboratory of Butantan Institute. The fragments were purified using Illustra GFX PCR DNA and Gel Band Purification Kit (GE Healthcare, Buckinghamshire, UK). DNA concentration and purity were determined in a spectrophotometer BioPhotometer plus (Eppendorf, Hamburg, Germany) and submitted to sequencing reactions. Sequencing. The purified amplified products were sequenced in an ABI377 PRISM Genetic Analyzer. The quality of DNA sequences was checked, the overlapping fragments were assembled using the BioEdit package software BioEdit package 7.0.9.0 [22], and the nucleotide sequences were compared through BLAST (http://blast.ncbi.nlm.nih.gov/Blast.cgi).
2.3. Comet Assay
Comet assay was performed according to alkaline technique [15]. An aliquot of 10 μL (0.1 × 105 cells) was transferred to 0.2 mL polypropylene tubes and mixed with 75 μL of low-melting-point agarose (0.7% in PBS) at 37°C, LMA optimum concentration (≤0.8%), without affecting the DNA migration [17], which was spread onto 76 × 26 mm microscope slides precoated with normal-melting-point agarose (1.5% in PBS) at 60°C and dried at 22°C overnight. After the agarose had solidified (4°C for 10 min), the coverslips were carefully removed, and the slides were immersed in lysis solution (2.5 M NaCl, 100 mM Na2EDTA, 10 mM Tris-HCl; pH 10; 1% Triton X-100, and 10% DMSO) for 1 hour at 4°C and then placed into a horizontal electrophoresis apparatus filled with buffer (1 mM Na2EDTA, 300 mM NaOH) at 4°C. The slides were incubated for 40 min in this buffer to unwind the DNA. The electrophoresis was run for 20 min at fixed voltage of 25 V (0.83 V/cm) and 300 mA. At the end of electrophoresis, the slides were washed three times with neutralization buffer (0.4 M Tris-HCl, pH 7.5) and once in 100% ethanol. All steps described above were carried out with the lights out to avoid DNA damage. The slides were stained with 20 μL of propidium iodide (4 μg/mL) and visualized in Carl Zeiss Axio Scope A1 fluorescent microscopy. Nucleoids were classified from 0 (without lesion) to 2 (major damage), and the number of nucleoids observed per class was multiplied to the class value, resulting in a comet score. Statistical analysis. The data were analyzed using the Kruskal-Wallis test, followed by post hoc Dunn test and Mann-Whitney U test, both with significance level of 5%, through BioEstat 5.3. software.
3. Results
3.1. Viral Genotyping
PCR using specific primers to β-globin revealed DNA quality enough for further PCR procedures: all samples resulted in a 450 bp amplicon. We selected the specific primers for BPV-1, -2, and -4 due to their prevalence in the herd, and we could detect the virus sequences in peripheral blood cells collected from the adult animals, with and without skin papillomas (Figure 1(a)). The resulting bands were purified and sequenced to confirm the genotyping of the amplification products. The sequences were aligned through the BioEdit 7.0.9.0, and the nucleotide comparison was done through the BLAST tool, confirming the specificity of the primers employed. Using these primers, coinfection was reported in 59.375% of the infected bovines—19 animals (Table 1(a)). The use of these primers did not detect the presence of BPV in calf peripheral blood samples, with this group being considered a negative control.
Figure 1.
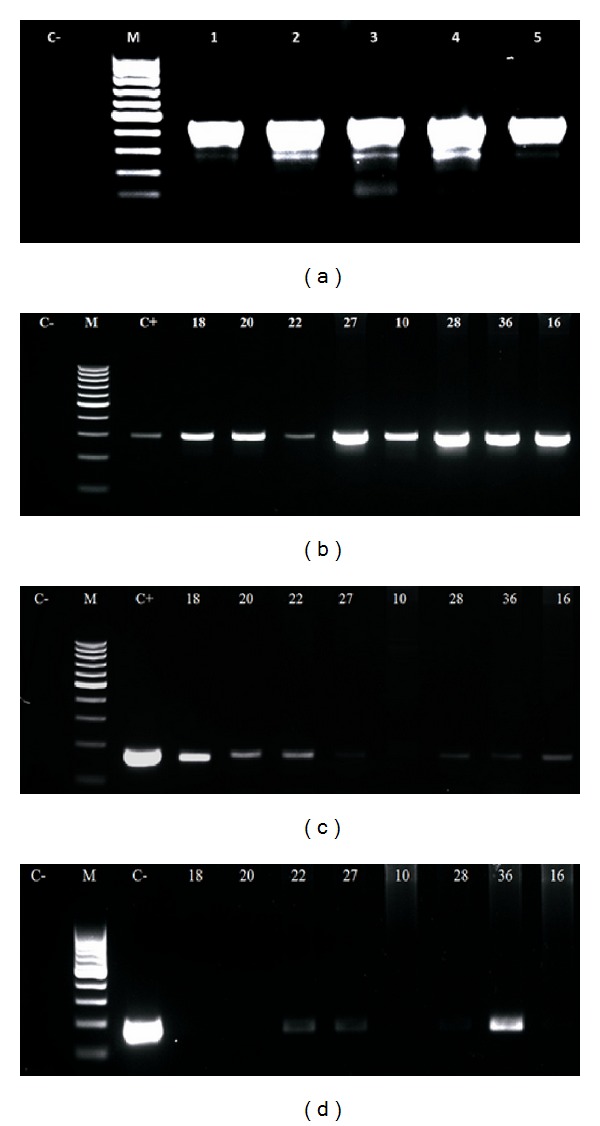
Electrophoresis's images of: (a) (β-globin amplification, resulting in an amplicon of 450 bp), (b) (amplification of L1 gene, using primer to BPV-1, showing an amplicon of 301 bp), (c) (amplification of L2 gene, using primer to BPV-2, showing an amplicon of 164 bp) and (d) (amplification of E7 gene, using primer to BPV-4, showing an amplicon of 170 bp), being C− (negative control) and C+ (positive control).
Table 1.
(a) Results of molecular diagnosis of calves (negative control) and asymptomatic and symptomatic adult cattle, showing the BPV type, the frequency of nucleoids observed per class, and the comet score. (b) Kruskal-Wallis statistic to compare the clastogenicity among calves, asymptomatic and symptomatic cattle, from the comet score. (c) Comparison between comet score observed in monoinfected and coinfected bovines through Mann-Whitney U test.
(a)
| Controls | Clinical | Virus type | Class of nucleoids | Score | ||
|---|---|---|---|---|---|---|
| 0 | 1 | 2 | ||||
| 1 | Asymptomatic | — | 93 | 4 | 3 | 10 |
| 2 | Asymptomatic | — | 92 | 5 | 3 | 11 |
| 3 | Asymptomatic | — | 96 | 1 | 3 | 7 |
| 4 | Asymptomatic | — | 90 | 8 | 2 | 12 |
| 5 | Asymptomatic | — | 95 | 3 | 2 | 7 |
|
| ||||||
| Infected bovines | ||||||
|
| ||||||
| 6 | Asymptomatic | BPV-1 | 15 | 60 | 25 | 110 |
| 7 | Asymptomatic | BPV-2 | 57 | 36 | 7 | 50 |
| 8 | Asymptomatic | BPV-1 | 75 | 19 | 6 | 31 |
| 9 | Asymptomatic | BPV-1, -2 | 43 | 17 | 40 | 97 |
| 10 | Asymptomatic | BPV-1 | 73 | 19 | 8 | 35 |
| 11 | Asymptomatic | BPV-1, -2 | 47 | 26 | 27 | 80 |
| 12 | Asymptomatic | BPV-1 | 38 | 26 | 36 | 98 |
| 13 | Asymptomatic | BPV-1, -2 | 62 | 13 | 25 | 63 |
| 14 | Asymptomatic | BPV-2 | 72 | 21 | 7 | 35 |
| 15 | Asymptomatic | BPV-1, -2 | 63 | 12 | 25 | 62 |
| 16 | Asymptomatic | BPV-1, -2 | 58 | 8 | 34 | 76 |
| 17 | Asymptomatic | BPV-1, -2 | 40 | 42 | 18 | 78 |
| 18 | Symptomatic | BPV-1, -2 | 61 | 30 | 9 | 48 |
| 19 | Symptomatic | BPV-2 | 22 | 13 | 65 | 143 |
| 20 | Symptomatic | BPV-1, -2 | 65 | 12 | 23 | 58 |
| 21 | Symptomatic | BPV-1, -2 | 57 | 21 | 22 | 65 |
| 22 | Symptomatic | BPV-1, -2, -4 | 31 | 31 | 38 | 107 |
| 23 | Symptomatic | BPV-1 | 89 | 6 | 5 | 16 |
| 24 | Symptomatic | BPV-1 | 92 | 2 | 6 | 14 |
| 25 | Symptomatic | BPV-1 | 69 | 25 | 6 | 37 |
| 26 | Symptomatic | BPV-1 | 41 | 10 | 49 | 108 |
| 27 | Symptomatic | BPV-1, -2, -4 | 70 | 21 | 9 | 39 |
| 28 | Symptomatic | BPV-1, -2, -4 | 65 | 26 | 9 | 44 |
| 29 | Symptomatic | BPV-2 | 87 | 8 | 5 | 18 |
| 30 | Symptomatic | BPV-1, -2 | 0 | 66 | 34 | 134 |
| 31 | Symptomatic | BPV-1, -2 | 60 | 31 | 9 | 49 |
| 32 | Symptomatic | BPV-2 | 52 | 23 | 25 | 73 |
| 33 | Symptomatic | BPV-1, -2 | 72 | 9 | 19 | 47 |
| 34 | Symptomatic | BPV-1, -2 | 91 | 7 | 2 | 11 |
| 35 | Symptomatic | BPV-1, -2 | 74 | 15 | 11 | 37 |
| 36 | Symptomatic | BPV1, -2, -4 | 82 | 13 | 5 | 23 |
| 37 | Symptomatic | BPV-1, -2 | 36 | 22 | 42 | 106 |
(b)
| Groups | Average post | Z calculated | P value |
|---|---|---|---|
| Calves and asymptomatic | 3.300 | 3.5348 | <0.05 |
| Calves and symptomatic | 23.666 | 3.1097 | <0.05 |
| Asymptomatic and symptomatic | 20.125 | 0.8961 | n.s. |
H = 12.9714, P = 0.0015, R of calves = 16.5, R of asymptomatic = 284.0 and R of symptomatic = 402.5.
(c)
| Monoinfected sample | Score | Coinfected sample | Score |
|---|---|---|---|
| 6 | 110 | 9 | 97 |
| 7 | 50 | 11 | 80 |
| 8 | 31 | 13 | 63 |
| 10 | 35 | 15 | 62 |
| 12 | 98 | 16 | 76 |
| 14 | 35 | 17 | 78 |
| 19 | 143 | 18 | 48 |
| 23 | 16 | 20 | 20 |
| 24 | 14 | 21 | 21 |
| 25 | 37 | 22 | 22 |
| 26 | 108 | 27 | 39 |
| 29 | 18 | 28 | 44 |
| 32 | 74 | 30 | 134 |
| — | — | 31 | 49 |
| — | — | 33 | 47 |
| — | — | 34 | 11 |
| — | — | 35 | 37 |
| — | — | 36 | 23 |
| — | — | 37 | 106 |
Mann-Whitney U test results: U = 102.5, Z(U) = 0.80, P value = 0.21.
3.2. Comet Assay
The samples of peripheral blood cells collected from infected animals and calves (negative control) were evaluated through comet assay, counting 100 nucleoids per sample that were evaluated and classified as 0 (without damage), 1 (medium damage), and 2 (maximum damage) according to Figure 2. The nucleoid value per class was multiplied by the respective class value, resulting in the comet score (Table 1(a)).
Figure 2.
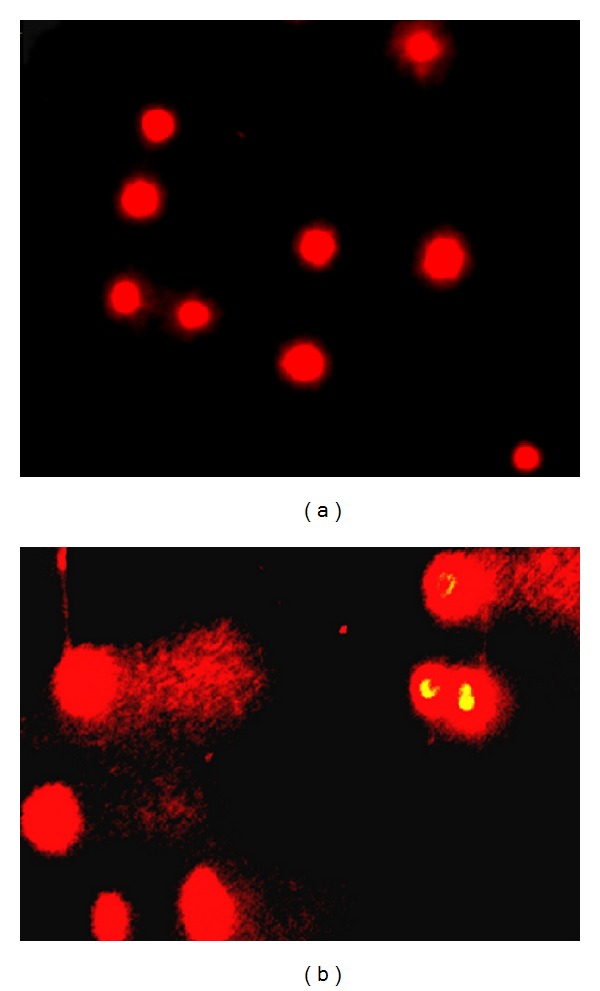

(a) Images of class 0 nucleoids, observed in calves (negative control) and (b) class 2 nucleoids, observed in symptomatic adult bovine, showing DNA fragmentation that is indicative of clastogenicity.
Based on these data, a Kruskal-Wallis test was used, with 5% significance level, through BioEstat 5.3. software to compare the different groups (infected and not infected group). The test reveled statistical differences between the groups (H = 12.9714 and P value = 0.0015). The Dunn post hoc test showed difference in score values among calf and infected animals (asymptomatic and symptomatic) (Table 1(b)). Mann-Whitney U test was done using the same software to compare the level of clastogenicity between animals, which showed just one virus type, and coinfected cattle. The results did not reveal statistical differences between animals infected with only one viral type and animals presenting more than one viral type (Table 1(c)).
4. Discussion
The carcinogenic mechanisms related to BPV are not yet fully elucidated; however, it is known that the malignancy is caused by mutations induced by the virus, associated with the action of viral protein E6, which accelerates the degradation of p53 and E7, which degrades the tumor suppressor protein pRb. These processes change the transcriptional pathway through degradation of transcription factors, activating telomerase, affecting the DNA repair system, and leading to an increase of damage in host genetic material [6, 23–25]. According to You [6], the protein E7 interacts with microtubules in mitosis, causing defects in the alignment of chromosomes during pre-metaphase, resulting in cytogenetic alteration [1, 26]. Wade et al. [25] discussed that the BPV oncoproteins can act on the signal transduction, allowing the return of interphase spinous epithelial cells to synthesis phase, resulting in mutated cell proliferation. Oncoproteins E6 and E7 induce immortalization of transformed cells [3]. According to Primrose and Twyman [27], these oncoproteins are required in the process of viral replication and act in the process of oncogenic transformation of the host cell. Furthermore, the accelerated p53 proteasomal degradation has a prominent role in the carcinogenic action of the virus, since the p53 protein function is to check the integrity of the genome, preventing the proliferation of mutated cells. The p53 accumulates in the cell nucleus, keeping the mitotic cycle in early G1 phase by activating p21 gene, whose gene product inhibits the action of cycling-dependent kinases (CDKs) and activates genes related to repair system [28].
Melo et al. [26] analyzed peripheral blood lymphocytes obtained from clinically asymptomatic bovines and showed a high level of chromosomal aberrations, suggesting the BPV action in the host chromatin even before the clinical manifestation of papillomatosis. These data also indicate that the skin lesions (papillomas) represent visible clinical manifestations, but the virus or its DNA sequences, detected in peripheral blood, can represent a potential risk to carcinogenesis.
In this study, we verified BPV clastogenic action by comet assay indicating chromatin instability. The comet assay results demonstrated that the BPV is able to induce severe DNA damages, which hinder the repair system, this is because the assay allows to evaluate the DNA double-strand breaks (DSBs) and critical lesions that are involved in genomic instability [29–31].
The DSB is associated with the homologous recombination from the formation of DNA simple strand that invades the template strand, originating a Holliday junction, which migrates to the resolution of heteroduplex [29, 31]. However, unrepaired DSBs leave to apoptosis or cell-cycle arrest, resulting in carcinogenesis [29]. There are lines of evidence that unrepaired DSBs could leave to telomeric breaks and fusion events, also associated with oncogenic process [29, 31].
The viral oncoproteins affect the repair system, allowing an accumulation of stochastic mutations and resulting in increased genomic instability [17]. According to Duensing and Münger [23], breaks in DNA affect the cell-cycle checkpoint that is associated with genomic instability, leading to hyper proliferation, featuring an oncogenic process. According to the European Study Group on Health's Biomarkers, cytogenetic findings, as a high frequency of chromosomal aberrations, including breaks in single or double-stranded, are associated to the carcinogenesis [32]. So, at the same time the presence of BPV is causing DSBs, the virus affects the repair system, favoring oncogenic process associated with unrepaired DSBs.
The virus presence in the blood can suggest one alternative pathway to infection, in which asymptomatic but infected cattle could turn symptomatic from a tissue injury, considering that a lesion causes an inflammatory process with lymphocyte infiltration [33]. The presence of BPV in leukocytes was demonstrated, had been observed the BPV presence in peripheral blood mononuclear cells [33] and the presence of L1 protein in CD4+ and CD8+ leukocytes, representing a potential infection sites to BPV-2 [34]. The possibility of the existence of endogenous pathway of infection has been discussed by Wobeser et al. [33], who suggested that the mononuclear cells act as a source of infection for inflammation sites, as inflamed areas become more susceptible to infection by BPV. Furthermore, it is known that lymphocytes express heparan sulfate, being cells susceptible to infection by papillomavirus [33]. Another observation that supports the possibility of infection was reported by Hartl et al. [35], who found that the spontaneous regression of papillomavirus in transient infections in humans and cattle is accompanied by an accumulation of active lymphocytes CD4+ and CD8+. So, in this pathway, the infiltration of BPV infected cell could develop a tumorigenic process from a clonal evolution started in a histologically normal tissue [35, 36].
5. Conclusion
This study presents direct evidence of DNA damage related to bovine papillomavirus in blood cells, indicating a viral activity in peripheral blood. The levels of damage were analyzed in order to verify if the presence of more than one viral type could increase the clastogenic viral action, but no significant differences could be detected. The results showed the same DNA damage both in presence or absence of cutaneous papillomas, indicating that the presence of bovine papillomatosis just represents clinical symptoms due the BPV presence; however, the BPV presence in peripheral blood leaves to double-stranded breaks, which is associated to carcinogenesis, affecting the healthy animal, as previously reported [1, 26]. Comet assay can be discussed as an interesting technique to evaluate DNA damage which, in this special situation, is related to viral action, demonstrating viral activity in different sites as blood cells.
Conflict of Interests
All the authors declare that there is no conflict of interests regarding the research, authorship, and/or publication of this paper. The authors inform that all materials and trademarks mentioned were used just as a part of experimental protocol and there are no financial gains or favoritism of business.
Acknowledgments
The authors thank the Ministério de Ciência, Tecnologia e Inovação/Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq Proc. 402539/2011-7), Coordenação de Aperfeiçoamento Pessoal de Nível Superior (CAPES), Fundação do Desenvolvimento Administrativo (PAP-FUNDAP) for the financial support, Carolina da Paz Sabino for the editorial support, and Martin W. Breuer for all the efforts and structural support.
References
- 1.Stocco dos Santos RC, Lindsey CJ, Ferraz OP, et al. Bovine papillomavirus transmission and chromosomal aberrations: an experimental model. Journal of General Virology. 1998;79(9):2127–2135. doi: 10.1099/0022-1317-79-9-2127. [DOI] [PubMed] [Google Scholar]
- 2.de Freitas AC, de Carvalho C, Brunner O, et al. Viral DNA sequences in peripheral blood and vertical transmission of the virus: a discussion about BPV-1. Brazilian Journal of Microbiology. 2003;34(1):76–78. [Google Scholar]
- 3.Zheng Z-M, Baker CC. Papillomavirus genome structure, expression, and post-transcriptional regulation. Frontiers in Bioscience. 2006;11(1):2286–2302. doi: 10.2741/1971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Freitas AC, Silva MAR, Carvalho CCR, et al. Communicating Current Research and Educational Topics and Trends in Applied Microbiology. 2007. Papillomavirus DNAdetectionin non-epithelial tissue: adiscussion about bovine papillomavirus; pp. 697–704. [Google Scholar]
- 5.Roperto S, Brun R, Paolini F, et al. Detection of bovine papillomavirus type 2 in the peripheral blood of cattle with urinary bladder tumours: possible biological role. Journal of General Virology. 2008;89(12):3027–3033. doi: 10.1099/vir.0.2008/004457-0. [DOI] [PubMed] [Google Scholar]
- 6.You J. Papillomavirus interaction with cellular chromatin. Biochimica et Biophysica Acta. 2010;1799(3-4):192–199. doi: 10.1016/j.bbagrm.2009.09.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Freitas AC, Silva ARM, Jesus ALS, et al. Recent insights into bovine papillomavirus. African Journal of Microbiology Research. 2011;5(33):6004–6012. [Google Scholar]
- 8.de Carvalho C, de Freitas AC, Brunner O, et al. Bovine papillomavirus type 2 in reproductive tract and gametes of slaughtered bovine females. Brazilian Journal of Microbiology. 2003;34(1):82–84. [Google Scholar]
- 9.Lunardi M, Alfieri AA, Otonel RAA, et al. Genetic characterization of a novel bovine papillomavirus member of the Deltapapillomavirus genus. Veterinary Microbiology. 2013;162(1):207–213. doi: 10.1016/j.vetmic.2012.08.030. [DOI] [PubMed] [Google Scholar]
- 10.DeMasi J, Chao MC, Kumar AS, Howley PM. Bovine papillomavirus E7 oncoprotein inhibits anoikis. Journal of Virology. 2007;81(17):9419–9425. doi: 10.1128/JVI.00422-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Lindsey CJ, Almeida ME, Vicari CF, et al. Bovine papillomavirus DNA in milk, blood, urine, semen, and spermatozoa of bovine papillomavirus-infected animals. Genetics and Molecular Research. 2009;8(1):310–318. doi: 10.4238/vol8-1gmr573. [DOI] [PubMed] [Google Scholar]
- 12.Campos SRC, Trindade C, Ferraz OP, et al. Can established cultured papilloma cells harbor bovine papillomavirus? Genetics and Molecular Research. 2008;7(4):1119–1126. doi: 10.4238/vol7-4gmr498. [DOI] [PubMed] [Google Scholar]
- 13.Zhu W, Dong J, Shimizu E, et al. Characterization of novel bovine papillomavirus type 12 (BPV-12) causing epithelial papilloma. Archives of Virology. 2012;157(1):85–91. doi: 10.1007/s00705-011-1140-7. [DOI] [PubMed] [Google Scholar]
- 14.Östling O, Johanson K. Microeletrophoretic study of radiation-induced DNA damages in individual mammalian cells. Biochemical and Biophysical Research Communications. 1984;123:291–298. doi: 10.1016/0006-291x(84)90411-x. [DOI] [PubMed] [Google Scholar]
- 15.Singh NP, McCoy MT, Tice RR, Schneider EL. A simple technique for quantitation of low levels of DNA damage in individual cells. Experimental Cell Research. 1988;175(1):184–191. doi: 10.1016/0014-4827(88)90265-0. [DOI] [PubMed] [Google Scholar]
- 16.Escobar P. New applications of the Comet Assay in genetic toxicology testing. Proceedings of the Genetic Toxicology Association Meeting; September 2008. [Google Scholar]
- 17.Langie SAS, Cameron KM, Waldron KJ, Fletcher KPR, von Zglinicki T, Mathers JC. Measuring DNA repair incision activity of mouse tissue extracts towards singlet oxygen-induced DNA damage: a comet-based in vitro repair assay. Mutagenesis. 2011;26(3):461–471. doi: 10.1093/mutage/ger005. [DOI] [PubMed] [Google Scholar]
- 18.Khoei S, Delfan S, Neshasteh-Riz A, Mahdavi SR. Evaluation of the combined effect of 2ME2 and 60Co on the inducement of DNA damage by IUdR in a spheroid model of the U87MG glioblastoma cancer cell line using alkaline comet assay. Cell Journal. 2011;13(2):83–90. [PMC free article] [PubMed] [Google Scholar]
- 19.Fabiani R, Rosignoli P, de Bartolomeo A, Fuccelli R, Morozzi G. Genotoxicity of alkene epoxides in human peripheral blood mononuclear cells and HL60 leukaemia cells evaluated with the comet assay. Mutation Research. 2012;747:1–6. doi: 10.1016/j.mrgentox.2012.01.004. [DOI] [PubMed] [Google Scholar]
- 20.Dias JDC, Sgnacchiti MDC, Giuriato PGG, Nunes LC, Pereira-Júnior OS. Detecção do papilomavírus bovino tipo 2 em bexigas de bovinos com hematúria enzoótica pela técnica de reação de cadeia de polimerase no Sul do Espírito Santo. Revista Brasileira de Medicina Veterinária. 2012;34:146–151. [Google Scholar]
- 21.Yaguiu A, Dagli MLZ, Birgel EH, Jr., et al. Simultaneous presence of bovine papillomavirus and bovine leukemia virus in different bovine tissues: in situ hybridization and cytogenetic analysis. Genetics and Molecular Research. 2008;7(2):487–497. doi: 10.4238/vol7-2gmr436. [DOI] [PubMed] [Google Scholar]
- 22.Hall TA. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series. 1999;41:95–98. [Google Scholar]
- 23.Duensing S, Münger K. The human papillomavirus type 16 E6 and E7 oncoproteins independently induce numerical and structural chromosome instability. Cancer Research. 2002;62(23):7075–7082. [PubMed] [Google Scholar]
- 24.Liu Y, Baleja JD. Structure and function of the papillomavirus E6 protein and its interacting proteins. Frontiers in Bioscience. 2008;13(1):121–134. doi: 10.2741/2664. [DOI] [PubMed] [Google Scholar]
- 25.Wade R, Brimer N, Vande Pol S. Transformation by bovine papillomavirus type 1 E6 requires paxillin. Journal of Virology. 2008;82(12):5962–5966. doi: 10.1128/JVI.02747-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Melo TC, Diniz N, Campos SRC, et al. Cytogenetic studies in peripheral blood of bovines afflicted by papillomatosis. Veterinary and Comparative Oncology. 2011;9(4):269–274. doi: 10.1111/j.1476-5829.2011.00263.x. [DOI] [PubMed] [Google Scholar]
- 27.Primrose SB, Twyman RM. Principles of Gene Manipulation and Genomics. 7th edition. Oxford, UK: Blackwell Scientific Publications; 2006. [Google Scholar]
- 28.Cavalcanti-Júnior GB, Klumb CE, Maia RC. p53 e as hemopatias malignas. Revista Brasileira de Cancerologia. 2002;48:419–442. [Google Scholar]
- 29.O’Driscoll M, Jeggo PA. The role of double-strand break repair—insights from human genetics. Nature Reviews Genetics. 2006;7(1):45–54. doi: 10.1038/nrg1746. [DOI] [PubMed] [Google Scholar]
- 30.Kanaar R, Hoeijmakers JHJ, van Gent DC. Molecular mechanisms of DNA double-strand break repair. Trends in Cell Biology. 1998;8(12):483–489. doi: 10.1016/s0962-8924(98)01383-x. [DOI] [PubMed] [Google Scholar]
- 31.Shrivastav M, de Haro LP, Nickoloff JA. Regulation of DNA double-strand break repair pathway choice. Cell Research. 2008;18(1):134–147. doi: 10.1038/cr.2007.111. [DOI] [PubMed] [Google Scholar]
- 32.Collins A, Anderson D, Coskun E, et al. Launch of the ComNet (Comet-Network) project on the comet assay in human population studies during the International Comet Assay Workshop Meeting in Kusadasi, Turkey (September 13–16, 2011) Mutagenesis. 2012;27:385–386. doi: 10.1093/mutage/ges014. [DOI] [PubMed] [Google Scholar]
- 33.Wobeser BK, Hill JE, Jackson ML, et al. Localization of Bovine papillomavirus in equine sarcoids and inflammatory skin conditions of horses using laser microdissection and two forms of DNA amplification. Journal of Veterinary Diagnostic Investigation. 2012;24(1):32–41. doi: 10.1177/1040638711425952. [DOI] [PubMed] [Google Scholar]
- 34.Roperto S, Comazzi S, Ciusani E, et al. PBMCS are additional sites of productive infection of bovine papillomavirus type 2. Journal of General Virology. 2011;92(8):1787–1794. doi: 10.1099/vir.0.031740-0. [DOI] [PubMed] [Google Scholar]
- 35.Hartl B, Hainisch EK, Shafti-Keramat S, et al. Inoculation of young horses with bovine papillomavirus type 1 virions leads to early infection of PBMCS prior to pseudo-sarcoid formation. Journal of General Virology. 2011;92(10):2437–2445. doi: 10.1099/vir.0.033670-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Leung CT, Brugge JS. Outgrowth of single oncogene-expressing cells from suppressive epithelial environments. Nature. 2012;482(7385):410–413. doi: 10.1038/nature10826. [DOI] [PMC free article] [PubMed] [Google Scholar]


