Abstract
Mammalian development is strongly influenced by the epigenetic phenomenon called genomic imprinting, in which either the paternal or the maternal allele of imprinted genes is expressed. Paternally expressed Xist, an imprinted gene, has been considered as a single cis-acting factor to inactivate the paternally inherited X chromosome (Xp) in preimplantation mouse embryos. This means that X-chromosome inactivation also entails gene imprinting at a very early developmental stage. However, the precise mechanism of imprinted X-chromosome inactivation remains unknown and there is little information about imprinted genes on X chromosomes. In this study, we examined whether there are other imprinted genes than Xist expressed from the inactive paternal X chromosome and expressed in female embryos at the preimplantation stage. We focused on small RNAs and compared their expression patterns between sexes by tagging the female X chromosome with green fluorescent protein. As a result, we identified two micro (mi)RNAs–miR-374-5p and miR-421-3p–mapped adjacent to Xist that were predominantly expressed in female blastocysts. Allelic expression analysis revealed that these miRNAs were indeed imprinted and expressed from the Xp. Further analysis of the imprinting status of adjacent locus led to the discovery of a large cluster of imprinted genes expressed from the Xp: Jpx, Ftx and Zcchc13. To our knowledge, this is the first identified cluster of imprinted genes in the cis-acting regulatory region termed the X-inactivation center. This finding may help in understanding the molecular mechanisms regulating imprinted X-chromosome inactivation during early mammalian development.
Introduction
In mammalian development, X-chromosome inactivation (XCI) as well as genomic imprinting is a well-known epigenetic regulatory mechanism [1], [2]. In female mammals, XCI maintains the level of gene expression from both X chromosomes to be the same as in male mammals. Without XCI, female embryos die, indicating the importance of this epigenetic regulation in female mammalian development [3]. During mouse preimplantation development, the paternally inherited X chromosome (Xp) derived from the fertilizing spermatozoon begins to be inactivated selectively by imprinted XCI. This imprinted inactivation is completely established in the extraembryonic tissues after implantation [4]–[7].
To date, Xist, an imprinted gene expressed from the inactive Xp, has been thought to be a single cis-acting regulator of imprinted XCI. However, the mechanism has not been elucidated fully at a molecular level and there is little information about imprinted genes expressed from the inactivated Xp. Therefore, we examined whether there are imprinted genes, other than Xist, expressed from the Xp and only detectable in female preimplantation embryos. If such imprinted genes exist, they are likely candidate genes involved in the process of imprinted XCI.
We have compared the expression patterns of coding genes in wild-type male and female blastocysts using DNA microarrays that cover all known genes. From the comparison, Rhox5 and Fthl17 were identified as imprinted genes predominantly expressed in female embryos and expressed from the inactive Xp [8], [9]. These two genes are the only candidates other than Xist. However, targeted disruption of Rhox5 as well as interfering RNA (RNAi) knockdown of the Fthl17 gene did not affect the birth of female mice, suggesting that these candidate genes are dispensable for the mechanism regulating imprinted XCI [10].
In addition to such coding genes, several studies have indicated the involvement of noncoding RNAs (ncRNA), including small RNAs, in regulating gene expression [11]–[14]. Furthermore, micro (mi)RNAs and small nucleolar (sno)RNAs are imprinted and proposed to be involved in mammalian development [15]. These findings led us to speculate on the possible involvement of imprinted ncRNAs in the regulation of XCI. Using pyrosequencing, we compared expression patterns of small RNAs between male and female preimplantation embryos. Here, we report that two X-linked miRNAs are imprinted and form a large cluster of imprinted genes expressed from the Xp at the preimplantation stage. As far as we know, this is the first identified imprinted gene cluster located within the X-inactivation center that is known to regulate XCI.
Materials and Methods
Animals
The handling and surgical manipulation of all experimental animals were carried out according to the guidelines of the Committee on the Use of Live Animals in Teaching and Research of Tokyo Medical and Dental University, and animal protocols were reviewed and approved by the animal welfare committee of the Tokyo Medical and Dental University (Permit Number: 0130228A). All efforts were made to minimize suffering. The B6C3F1 TgN (act EGFP) Osb CX-38 (G38) transgenic mouse strain expressing the transgene for enhanced green fluorescent protein (EGFP) described elsewhere [8], [9], [16] was used to distinguish between male and female embryos.
Blastocyst Collection for Sequencing Small RNAs
Eight-week-old B6C3F1 female mice were superovulated with intraperitoneal injections of 5 IU pregnant mare serum gonadotropin followed by 5 IU of human chorionic gonadotropin (hCG) 48 h later. These superovulated female mice were mated with XGFPY male mice. Embryos at the 4-cell stage were collected from the oviducts 55 h after the hCG injection, placed in potassium simplex optimization medium and incubated in a humidified atmosphere of 5% CO2 in air at 37°C for an additional 38 h. After collecting early to mid-stage blastocysts, male (EGFP-negative) and female (EGFP-positive) embryos were separated using a fluorescence microscope (M165FC; Leica Microsystems GmbH, Wetzlar, Germany).
Small RNA Library Construction and Sequencing
Small RNA libraries were constructed as described [17]. Briefly, small RNAs (18–26 mer) were isolated from male and female blastocysts: 2000 each. The libraries of male and female blastocysts were constructed using a small RNA cloning kit (TaKaRa Bio Inc., Ohtsu, Japan) and the sequences of small RNAs were determined using a 454 Life Sciences pyrosequencer (Branford, CT). The small RNAs were mapped to the genome using the NCBI BLASTN program (http://blast.ncbi.nlm.nih.gov). Perfect match sequences were selected for further analysis. To annotate small RNAs, we used the following databases: miRNA_mature and miRNA_hairpin from miRBasev16; rRNA, snRNA, snoRNA, lincRNA, miscRNA, Mt_rRNA and Mt_tRNA, from Ensembl (release 6). Of the 397,726 and 473,043 mapped reads from the female and male libraries, only around 10% have been confirmed as noncoding RNAs, the remaining ∼90% are classified “only genomic” (Table 1). Therefore, further annotation was carried out using these “only genomic” sequences against RepeatMasker (mm9 - Jul 2007 - RepeatMasker open-3.2.8 - Repeat Library 20090604) and the mouse tRNA database (tRNAscan-SE Analysis of Mus musculus (mm9 July 2007)). As a result, ∼70% of the sequences were matched to repeat or tRNA sequences, suggesting that some of the small RNAs were degradation products derived from these sequences (Table S1). RNA abundance was measured using reads per kilobase per million reads (RPKM) [18]. The expression data of these annotated small RNAs are supplied as Table S2. The small RNA sequence data derived from male and female blastocysts have been deposited in the DNA Data Bank of Japan (http://www.ddbj.nig.ac.jp) under the accession number DRA001023.
Table 1. Classification of annotated small RNAs in male and female blastocysts.
| Female | Male | |
| No. reads | 433,905 | 517,507 |
| Genome mapped* | 397,726 | 473,043 |
| miRNA_mature | 31,488 | 31,608 |
| Mt_tRNA | 3,609 | 4,247 |
| Mt_rRNA | 1,637 | 2,339 |
| snRNA | 1,142 | 1,510 |
| rRNA | 1,026 | 1,180 |
| snoRNA | 1,055 | 1,100 |
| misc_RNA | 381 | 486 |
| lincRNA | 402 | 479 |
| miRNA_hairpin | 130 | 167 |
| (Total ncRNA) | 40,876 | 43,121 |
| Only genomic** | 363,003 | 437,224 |
The number of genes mapped is not always equal to the “Total ncRNA” plus “Only genomic” counts because of multiple annotations for single read sequences.
Abbreviations: miRNA_mature, micro RNA mature form; Mt_tRNA, mitochondrial transfer RNA; Mt_rRNA, mitochondrial ribosomal RNA; snRNA, small nuclear RNA; rRNA, ribosomal RNA; snoRNA, small nucleolar RNA; miscRNA, miscellaneous RNA; lincRNA, large intergenic noncoding RNA; miRNA_hairpin: micro RNA stem-loop sequence obtained from miRBase.
The sequences classified as “Only genomic” were further annotated against RepeatMasker and the tRNA database. The results are shown in Table S1.
Quantitative Reverse Transcription Polymerase Chain Reaction (qRT–PCR) Analysis of Candidate Genes and Small RNAs for Sexually Differential Expression
The total RNA contents of male and female blastocysts were extracted using TRIzol (Invitrogen, Carlsbad, CA) and any contaminating DNA was digested using RNase-free DNase (Nippon Gene, Tokyo, Japan). In the case of qRT–PCR for mRNAs, first-strand cDNA was synthesized using reverse transcriptase (SuperScript III; Invitrogen) and random primers according to the manufacturer’s instructions. Expression levels are reported as the ratios of the number of cDNA copies for each gene to that of cDNA copies for the housekeeping gene β-actin. Primers used for qRT–PCR are summarized in Table S2. For miRNA expression analysis, total RNA was reverse transcribed using the TaqMan MicroRNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA). The TaqMan MicroRNA assays used (with identification numbers in brackets), were as follows: mmu-miR-374-5p (001319); mmu-miR-421-3p (002700); mmu-miR-107-3p (000443); mmu-miR-103-3p (000439); mmu-miR-30c-5p (000419); mmu-miR-7a-5p (000268); mmu-miR-19b-3p (000396); mmu-miR-467c-3p (464331); mmu-miR-295 (000189); snoRNA202 (001232). Mature miRNAs were amplified with TaqMan Universal PCR Master Mix II (Applied Biosystems) using a LightCycler 480 instrument (Roche Diagnostics, Mannheim, Germany). The qRT–PCR reactions were performed using samples collected from three independent RNA preparations.
Verification of Imprinting
Two reciprocal sets of hybrid blastocysts, (JF1× C57BL/6) F1 and (C57BL/6× (JF1× C57BL/6)) F1, were produced by cross-mating appropriately. All samples were recovered from the uteri to avoid the effects of embryo culture. Genomic DNA and RNA were extracted from individual blastocysts. Male and female blastocysts were pooled separately after sex determination by PCR as described [8]. In each experiment, at least 60 blastocysts were pooled by sex. One third of the resulting cDNA samples derived from pooled RNAs were amplified by PCR using the primers listed in Table S2.
In situ Hybridization (ISH)
Whole-mount ISH was performed as described [19]. Briefly, a full-length Ftx variant 2 cDNA clone (RIKEN FANTOM clone ID: 9530061G23) was used for preparation of digoxigenin-labeled sense and antisense RNA probes using the DIG RNA Labeling Mix (Roche, Basel, Switzerland). Specific signals were detected using an alkaline phosphatase-conjugated anti-digoxigenin antibody (Roche).
Statistics
Two-tailed Student’s t-tests were used to analyze the significance of any differences and P<0.01 was considered significant. All quantitative data are presented as the mean ± standard deviation of the mean.
Results
Identification of Small RNAs Predominantly Expressed in Female Embryos
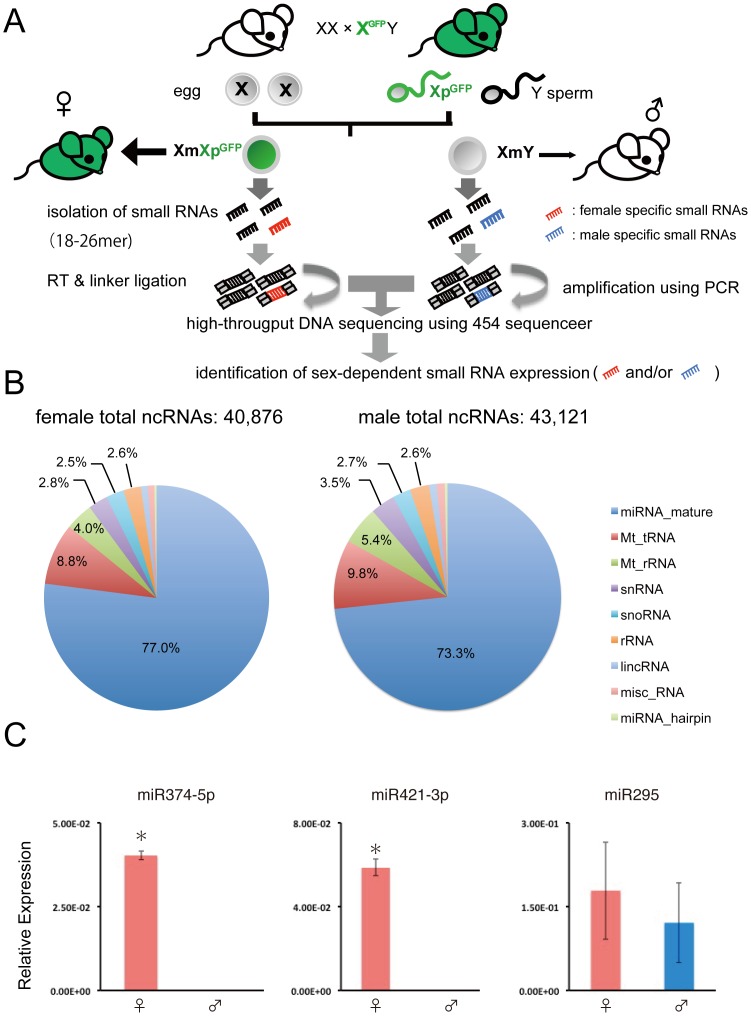
To screen small RNAs predominantly expressed in female embryos, we used a transgenic mouse line in which the X chromosome is tagged with the transgene for EGFP [8], [9], [20]. The sexing methodology is shown in Fig. 1A. The transgenic males (XGFPY) were mated with wild-type females (XX). Because of the technical difficulty of separating early male and female embryos using X-GFP mice, we chose blastocysts in which we could separate the sexes very easily and accurately.
Figure 1. Identification of small RNAs predominantly expressed in female blastocysts.
(A) Scheme of the XGFP sexing system and construction of small RNA libraries. Transgenic male mice (XGFPY) were mated with wild-type female mice (XX). Only the female embryos (XXGFP) fluoresce green because of the paternally inherited XGFP. Small RNAs obtained from male and female blastocysts were sequenced and expression levels were compared between sexes. (B) Annotation of the read sequences from male and female blastocyst libraries. The distribution of the RNA classes is given as a percentage of the total annotated ncRNAs. (C) Expression levels of miR-374-5p, miR-421-3p and miR-295 miRNA in male and female blastocysts. The expression of three miRNAs was measured by qRT–PCR and normalized to that of snoRNA202 examined as a control. The data indicate that the miR-374-5p and miR-421-3p were predominantly expressed in female blastocysts (*P<0.01). Results are expressed as the mean ± SD (n = 3).
First, small RNAs were isolated individually from male and female blastocysts and compared using 454 pyrosequencing followed by sequence annotation. We obtained 397,726 male and 473,043 female small RNA sequences, which were matched with the mouse genome completely. Table 1 shows the annotated small RNA sequences using miRbase (v. 16) and the Ensembl database (release 61). There was no apparent difference in the total distribution of annotated small RNAs between the two sexes (Fig. 1B; Table 1). Therefore, we next compared the expression patterns of individual small RNA sequences and discovered a number of differences between the sexes (Table 2, 3). To confirm the results of pyrosequencing, we carried out qRT–PCR of small RNAs showing more than 10-fold (1000%) changes in sequence reads. We successfully identified two miRNAs, miR-374-5p (Accession number: MI0004125 in miRBase) and miR-421-3p (MI0005496) that were expressed predominantly in female blastocysts(Fig. 1C). There was no significant difference of expressions of other miRNAs between sexes (Fig. S1). We also showed quantification of a microRNA (miR295: MI0000393) not submitted to imprint as control. There was no significant difference of expression between sexes (Fig. 1C). Interestingly, the two mature miRNAs (miR-374-5p and miR-421-3p) we identified were mapped close together on the X chromosome and appeared to be derived from a single primary (pri) miRNA (Fig. 2A). This was predicted to produce two stem–loop structured precursor (pre)-miRNAs. According to miRBase, each of these two pre-miRNAs produces two mature miRNAs, miR-374-5p/miR-374-3p and miR-421-5p/miR-421-3p (Fig. 2B). However, in our study each of the two pre-miRNAs produced a single mature miRNA, miR-374-5p and miR-421-3p, respectively, and neither miR-374-3p nor miR-421-5p was detected at the blastocyst stage.
Table 2. List of differentially expressed small RNAs between male and female blastocysts (1).
| Upregulated small RNAs in female blastocysts. | |||||
| RNA_ID | RNA_TYPE | Female_READS* | Male_READS* | M/F_READS | Map position (chromosome number) |
| mmu-miR-374-5p | miRNA_mature | 46.00 | – | 0.000 | X |
| mmu-miR-421-3p | miRNA_mature | 38.00 | – | 0.000 | X |
| ENSMUST00000143469/Gm13448 | lincRNA | 13.00 | – | 0.000 | 2 |
| RNA_ID | RNA_TYPE | Female_READS ** | Male_READS ** | M/F_READS | Map position (chromosome number) |
| mmu-miR-107 | miRNA_mature | 32.94 | 0.01 | 0.000 | 19 |
| mmu-miR-103 | miRNA_mature | 71.53 | 0.04 | 0.001 | 11 |
| mmu-miR-30c | miRNA_mature | 181.67 | 0.13 | 0.001 | 4 |
| mmu-miR-7a | miRNA_mature | 29.00 | 0.03 | 0.001 | 13 |
| mmu-miR-19b | miRNA_mature | 53.00 | 0.11 | 0.002 | X |
| ENSMUST00000093731 | snoRNA | 95.99 | 0.81 | 0.008 | 11 |
| ENSMUST00000082466 | snoRNA | 48.02 | 0.60 | 0.013 | 19 |
| ENSMUST00000082873 | snoRNA | 25.61 | 0.61 | 0.024 | 11 |
Sequence reads of small RNAs that showed 0 reads in male embryos and more than 10 reads in female embryos.
Small RNAs showing more than a 10-fold (1000%) change in sequence reads are listed. Small RNAs showing fewer than 20 reads have been excluded from this list.
Table 3. List of differentially expressed small RNAs between male and female blastocysts (2).
| Upregulated small RNAs in male blastocysts. | |||||
| RNA_ID | RNA_TYPE | Female_READS† | Male_READS† | M/F_READS | Map position (chromosome number) |
| mmu-miR-467c* | miRNA_mature | 0.02 | 23.98 | 1372.867 | 2 |
| ENSMUST00000083244 | snRNA | 0.71 | 69.49 | 98.219 | 13 |
| ENSMUST00000083111 | snRNA | 2.72 | 99.33 | 36.580 | 3 |
| ENSMUST00000082989 | snRNA | 7.00 | 137.46 | 19.625 | 4 |
Small RNAs showing more than a 10-fold (1000%) change in sequence reads are listed. Small RNAs showing fewer than 20 reads have been excluded from this list.
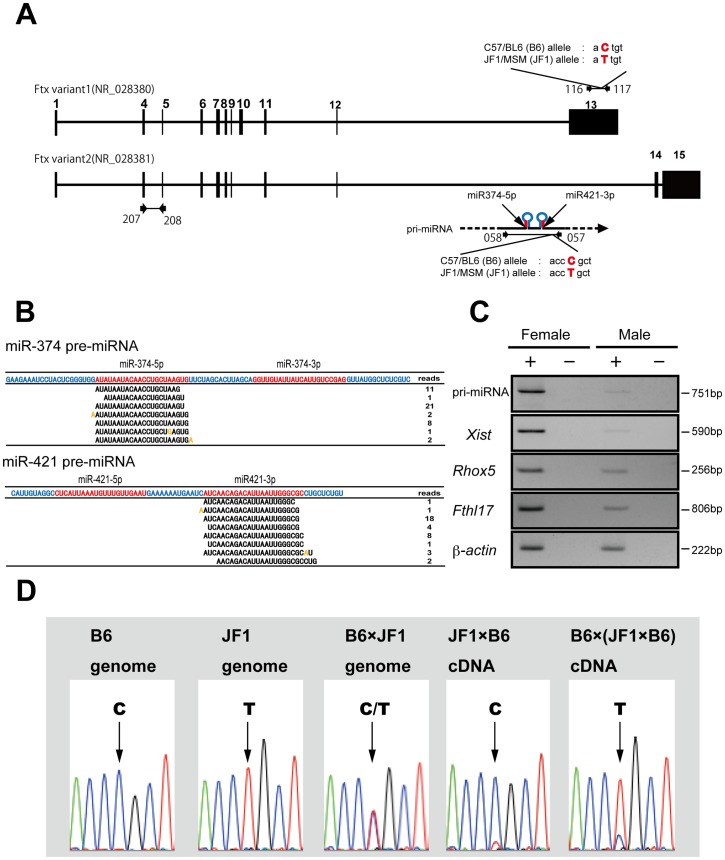
Figure 2. Verification of imprinting of two miRNAs predominantly expressed in female embryos.
(A) Schematic representation of two mature miRNAs, miR-374-5p and miR-421-3p and their preceding transcript pri-miRNA. Both miRNAs were clustered and located in the intron of the Ftx (B230206F22Rik) gene on the X chromosome. Both are thought to be transcribed as pri-miRNAs and then processed to form mature miRNAs. The location of the primers used to assess the pri-miRNA and Ftx expression levels are indicated: qRT–PCR primers 207 and 208 were used for Ftx; primers used for allelic discrimination were 057 and 058 for pri-miRNA, 116 and 117 for Ftx. The exons for Ftx are numbered according to a previous report [22]. DNA polymorphisms used for allelic discrimination are indicated with red characters. PCR primers correspond to the list in Table S3 (B) Production of mature miRNAs from pre-miRNAs in female blastocysts. Although two mature forms of miRNA, miR-374-5p/miR-374-3p and miR-421-5p/miR-421-3p are registered in miRBase, only miR-374-5p and miR-421-3p were detected at the blastocyst stage. Orange coloring shows mismatch sequences compared with the reference sequences, suggesting miRNA editing or misreading by the 454 sequencer. (C) Expression of pri-miRNA in wild-type male and female blastocysts obtained from the uterus. Female-predominant expression was observed as well as previously reported X-linked imprinted genes such as Xist and Rhox5. (D) Verification of imprinting of the pri-miRNA for miR-374-5p and miR-421-3p in preimplantation mouse embryos. Pri-miRNA was expressed from the paternal B6 allele in JF1× C57BL/6 crosses and from the paternal JF1 allele in the reciprocal B6× (JF1× B6) cross. The arrows indicate the polymorphism sites that were used in this assay.
Verification of Imprinting of the Female X Chromosome-expressed miRNA
We next examined whether miR-374-5p and miR-421-3p were imprinted, as is the case for Xist [21]. First, to distinguish paternal expression from maternal expression, we searched for DNA polymorphisms between Mus mus domesticus (C57BL/6: B6) and the Japanese wild mouse M. m. molossinus (JF1/MSM). Mature forms of miRNAs are only 22- and 23-nucleotides (nt) long, making it difficult to find polymorphisms within their sequences. We utilized a polymorphism identified in the pri-miRNA and examined the active alleles in F1 blastocysts. Then, we confirmed that the pri-miRNA in F1 blastocysts was also predominantly expressed in female embryos as well as the mature two miRNAs (Fig. 2C). Furthermore, allelic expression analysis revealed that the pri-miRNA was expressed from the Xp in F1 blastocysts (JF1× B6) derived from mating between JF1/MSM females and C57BL/6 males and F1 blastocysts (B6× (JF1× B6)) derived from the reciprocal cross of C57BL/6 females and F1 males (JF1/MSM × C57/BL6) (Fig. 2D). These data confirmed that miR-374-5p and miR-421-3p are both imprinted and expressed from the Xp chromosome.
Identification of a Novel Cluster of Imprinted Genes on the X Chromosome
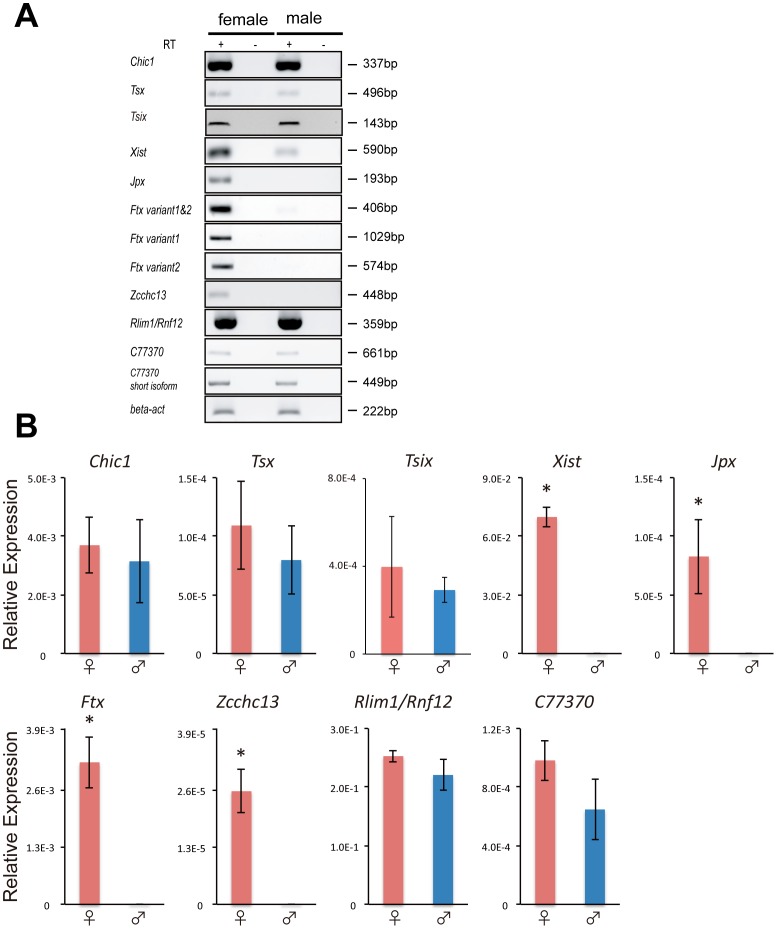
Given that most autosomal imprinted genes exist as clusters, we considered the possibility that these imprinted miRNA might cluster with other imprinted genes on the X chromosome. The two imprinted miRNAs were indeed mapped adjacent to the imprinted Xist gene, strengthening the possibility that they might form a cluster of imprinted genes. Therefore, we analyzed the expression patterns of genes neighboring these miRNAs. From the neighborhood region of approximately 1 Mb of miRNA (UCSC mm9), the expression level of 11 genes–Chic1, Tsx, Xite, Tsix, Xist, Jpx/Enox, Ftx/B230206F22Rik, Zcchc13, Slc16a2, Rlim/Rnf12 and C77370–were compared between male and female blastocysts. In addition to Xist, three genes, Ftx, Jpx and Zcchc13, were expressed predominantly in female blastocysts (Fig. 3A, B). We next examined the allelic expression of these genes to verify whether they were imprinted. Ftx is a polyadenylated ncRNA with two different spliced variants (NR_028380 and NR_028381) [22]. Both variants were predominantly expressed in female embryos (Fig. 3A). Using the DNA polymorphism found in the last exon of Ftx variant 1 (NR_028380), we verified that the Ftx gene was expressed predominantly when it was derived from the father (Fig. 4A). Similarly, the allelic expression of Jpx and Zcchc13 clearly showed that these two genes are also imprinted and expressed from the Xp chromosome (Fig. 4B, C). Taken together, these female-expressed imprinted genes on the X chromosome form a cluster adjacent to the miRNAs, miR-374-5p and miR-412-3p.
Figure 3. Searching for a cluster of imprinted genes on the X chromosome.
(A) qRT–PCR of X-linked genes in male and female blastocysts. It should be noted that very weak expression levels of Xist, Jpx, Ftx, Zcchc13 were detected in male blastocysts when additional PCR cycles were carried out. (B) The expression levels of Chic, Tsx, Xite, Tsix, Xist, Jpx, Ftx, Zcchc13, Rlim1/Rnf12, Slc16a2, C77370 were quantified by qRT–PCR in female and male mouse blastocysts. The ratios of the number of cDNA copies for each gene to that of cDNA copies for a housekeeping gene, β-actin, are indicated as expression levels. Expressions of Xite and Scl16a2 were not detected at the blastocyst stage (data not shown). Results are expressed as the mean ± SD (n = 3). The statistical significance of differences between sample means was determined by Student’s t-test. *P<0.01.
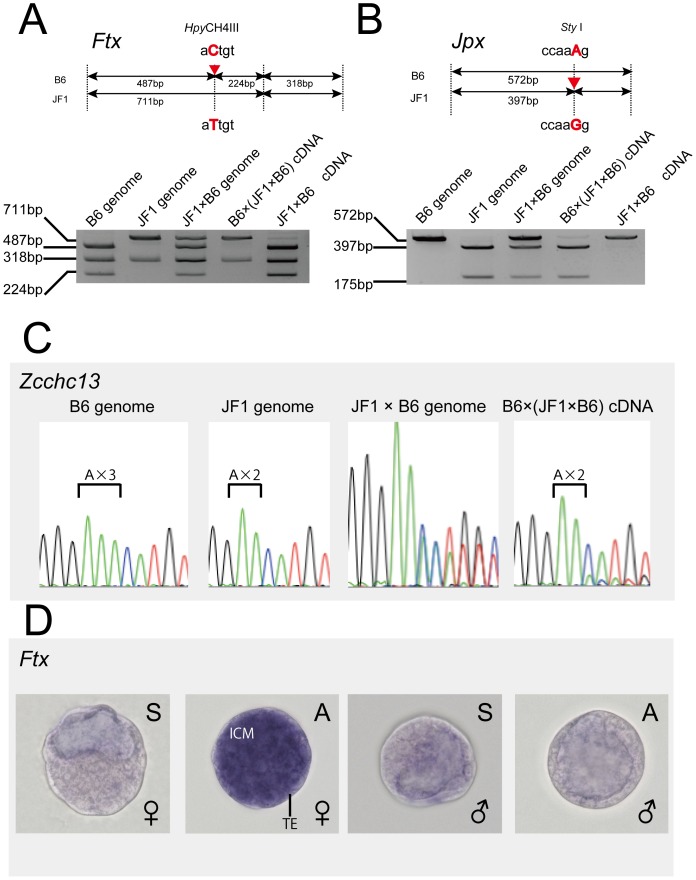
Figure 4. Allelic expression analysis of female-expressed genes at the blastocyst stage.
(A) Verification of imprinting in the Ftx gene. Upper: schematic representation of the position of the DNA polymorphism in the Ftx gene. The single nucleotide polymorphism detected in C57BL/6: B6 (aCtgt) and JF1/Ms: JF1 (aTtgt) is shown. The Hpy CH4III site in the B6 allele was changed in the JF1 allele. These alleles could be distinguished by digestion with Hpy CH4III. Lower: the imprinted expression of Ftx from the Xp chromosome was determined by qRT–PCR and restriction fragment length polymorphism (RFLP) analysis with inter-subspecific hybrid mouse F1 progenies. (B) Verification of Jpx imprinting. Upper: schematic representation of the position of the DNA polymorphism in the Jpx gene. The Sty I site in the B6 allele (ccaaAg) was changed in the JF1 allele (to ccaaGg). Lower: imprinted expression of Jpx from the Xp chromosome was determined by qRT–PCR and RFLP analysis with inter-subspecific hybrid mouse F1 progenies. (C) Verification of the Zcchc13 imprinting. The single nucleotide insertion polymorphism detected in C57BL/6; B6 (triple A) and JF1/Ms; and JF1 (double A) is shown. The imprinted expression of Zcchc13 from the Xp chromosome was determined by sequencing analysis with inter-subspecific hybrid mice F1 progenies. (D) Whole-mount in situ hybridization of the Ftx gene in female and male blastocysts. ‘S’ indicates the sense strand probe and ‘A’ indicates the antisense strand probe. Abbreviation: ICM, Inner cell mass; TE, trophectoderm.
To investigate the association between the expression patterns of newly identified imprinted genes and X-chromosome inactivation further, whole-mount ISH was carried out on blastocysts. Unfortunately, the expression levels of Jpx and Zcchc13 were too low to be detected by ISH. In contrast, Ftx, the most highly expressed gene among the identified imprinted genes, gave strong signals in female embryos in the trophectoderm where the Xp chromosome is inactivated and the ICM where the Xp chromosome is reactivated (Fig. 4D), whereas a signal was barely detectable in male blastocysts. Notably, Ftx was imprinted and expressed from the Xp chromosome, although this is selectively inactivated in the trophectoderm.
Discussion
A Cluster of Imprinted Genes on the X Chromosome is Expressed in Female Preimplantation Mouse Embryos
Here we found imprinted genes existing as a cluster and expressed from the Xp chromosome in preimplantation embryos. This suggests that these imprinted genes are additional candidates involved in imprinted XCI. Despite the early discovery of various imprinting effects observed including XCI [4], [5], [7], [23]–[26], there have been relatively few reports about imprinted genes on X chromosomes [27]. Xist was the first identified X-linked imprinted gene and is believed to play major roles in imprinted XCI. Adjacent to Xist, here we identified a number of imprinted genes including miR-374-5p and miR-421-3p and the long ncRNAs Jpx and Ftx, together with coding RNA Zcchc13 as a CCHC-type Zinc finger protein. Other than these imprinted genes, Tsix was previously reported to be imprinted and expressed from the maternal X chromosome [28], [29]. Our findings mean that this imprinted gene cluster reaches ∼200 kbp in length and we have named it the Tsix–Zcchc13 imprinted gene cluster (Fig. 5).
Figure 5. Identification of a cluster of imprinted genes in the X-inactivation center (Xic).
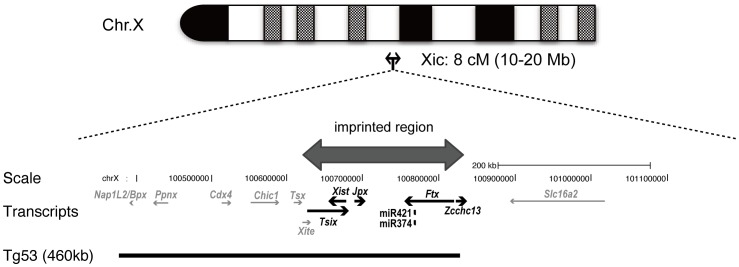
A newly identified cluster of imprinted genes on the X chromosome is shown in the mouse genome (UCSC mm9; http://genome.ucsc.edu/). This cluster is located within the Xic. The arrows show the direction of each transcription unit. Ftx and Zcchc13 are transcribed in opposite directions from the same putative bidirectional promoter. Imprinted genes are shown by the thick black arrows; Xist, Jpx, Ftx, Zcchc13, miR-374-5p and miR-421-3p were paternally expressed, while Tsix was reported previously to be maternally expressed [28], [29]. The thick black bar indicates the position of the Tg53 (460 kb) transgene reported in [35].
Imprinting Mark of the Tsix-Zcchc13 Imprinted Region
Allele-specific expression of imprinted genes requires an epigenetic mark to distinguish paternal from maternal chromosomes. Each cluster of imprinted genes is thought to be controlled by an imprinting control region (ICR) that is marked by DNA methylation during either oogenesis or spermatogenesis. Concerning the Tsix–Zcchc13 imprinted region, it has not been known how the paternal expression of Xist is controlled. However, our results indicate that this might be considered as a problem of how the entire Tsix–Zcchc13 imprinted region is controlled rather than a problem of how the single Xist gene is controlled. Genome-wide DNA methylation profiles from gametes have revealed that six CpG islands located within the Tsix–Zcchc13 region show hypomethylated patterns and have no characteristic of differentially methylated regions that work as ICRs in other autosomal imprinted regions [30], [31]. Given that the paternal expression of Xist does not require DNA methylation [32], these data suggest that the imprinting of this newly identified X-linked region is established by DNA methylation-independent mechanisms. Identification of the imprinted Tsix–Zcchc13 region should help us narrow down the genomic area to search for epigenetic marks distinct from DNA methylation.
Concerning the epigenetic state of this imprinted region, Ftx and Jpx have been reported to partially escape random XCI in differentiated ES cells [22], [33] and also to evade imprinted XCI in trophoblastic stem cells [34]. This suggests that the epigenetic state of this imprinted gene cluster differs from other inactivated X chromosome regions.
Roles of the Tsix–Zcchc13 Imprinted Gene Cluster in Imprinted XCI
Regarding the function of the identified imprinted gene cluster, Okamoto et al. indicated a possible involvement of this cluster in the regulation of imprinted XCI. A 460 kb single-copy transgene (Tg53) containing a part of the Tsix–Zcchc13 cluster was reported to induce imprinted cis-inactivation in autosomes at the preimplantation embryo stage when it was inherited paternally [35]. Thus, this 460 kb transgene contains sufficient factors to trigger imprinted inactivation of the paternal chromosome. The 460 kb transgene was reported to encompass the genomic region from Nap1l2/Bpx to Ftx, but not Zcchc13 (Fig. 5). Our research provides evidence that, in addition to the Xist gene, this transgene possesses two imprinted genes and two miRNAs: Jpx, Ftx, miR-374-5p and miR-421-3p that are expressed from the Xp chromosome. Thus, this transgenic analysis supported the idea that these newly identified imprinted genes (Jpx and Ftx and two miRNAs, miR-374-5p and miR-421-3p) function as cis-acting factor(s) and are involved in the mechanisms of imprinted paternal XCI at the preimplantation embryo stage.
Common Machinery for Random and Imprinted XCI
XCI can be separated into two consecutive processes, initial ‘imprinted XCI’ followed by ‘random XCI’. Random XCI is controlled by a cis-acting X-inactivation center termed Xic. [36], [37]. Inside Xic, we found a large cluster of imprinted genes: the Tsix–Zcchc13 cluster (Fig. 5). Among them, Jpx and Ftx were reported to upregulate the expression of Xist in random XCI mechanisms [22], [33]. Given the functions of these ncRNAs as positive regulators, they can also function to activate the expression of Xist in imprinted XCI at the preimplantation stage. Gene targeting and the generation of transgenic mice carrying these imprinted ncRNAs and miRNAs will help answer whether and how they are involved in imprinted XCI. Our finding not only provides much information about X-linked imprinted genes, but also helps in clarifying the molecular mechanisms of imprinted XCI. Functional analysis of these imprinted genes is currently under way using gene targeting technology.
Supporting Information
Validation of the expression of micro RNAs using qRT–PCR. The qRT–PCR analysis of differentially expressed microRNA candidates mmu-miR-7a, mmu-miR-19b, mmu-miR-30c, mmu-miR-103, mmu-miR-107 and mmu-miR-467 that are presented in Tables 2 and 3. The expression of each miRNA was measured and normalized to that of miR295 as a control. Results are expressed as the mean ± SD (n = 3).
(TIF)
Classification of annotated small RNAs in male and female blastocysts against RepeatMasker and tRNA database.
(XLS)
A list of annotated small RNAs expressed in male and female blastocysts.
(XLS)
A list of the PCR primer sets used in this research.
(XLS)
Acknowledgments
We are very grateful to Dr. Takashi Kohda and Dr. Daisuke Endo for their critical reading of the manuscript and useful advice. We also thank Mami Oikawa for technical assistance in collection of embryos.
Funding Statement
This work was supported by the Japan Science and Technology Agency, Precursory Research for Embryonic Science and Technology (PRESTO); a Grant-in-Aid for Scientific Research from The Ministry of Education, Culture, Sports, Science and Technology; and by the Sumitomo Foundation. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1. Fedoriw A, Mugford J, Magnuson T (2012) Genomic imprinting and epigenetic control of development. Cold Spring Harb Perspect Biol 4: 1–15. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Lyon MF (1961) Gene action in the X-chromosome of the mouse (Mus musculus L.). Nature 190: 372–373. [DOI] [PubMed] [Google Scholar]
- 3. Marahrens Y, Panning B, Dausman J, Strauss W, Jaenisch R (1997) Xist-deficient mice are defective in dosage compensation but not spermatogenesis. Genes Dev 11: 156–166. [DOI] [PubMed] [Google Scholar]
- 4. Mak W, Nesterova TB, de Napoles M, Appanah R, Yamanaka S, et al. (2004) Reactivation of the paternal X chromosome in early mouse embryos. Science 303: 666–669. [DOI] [PubMed] [Google Scholar]
- 5. Okamoto I, Otte AP, Allis CD, Reinberg D, Heard E (2004) Epigenetic dynamics of imprinted X inactivation during early mouse development. Science 303: 644–649. [DOI] [PubMed] [Google Scholar]
- 6. Patrat C, Okamoto I, Diabangouaya P, Vialon V, Le Baccon P, et al. (2009) Dynamic changes in paternal X-chromosome activity during imprinted X-chromosome inactivation in mice. Proc Natl Acad Sci U S A 106: 5198–5203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Huynh KD, Lee JT (2003) Inheritance of a pre-inactivated paternal X chromosome in early mouse embryos. Nature 426: 857–862. [DOI] [PubMed] [Google Scholar]
- 8. Kobayashi S, Fujihara Y, Mise N, Kaseda K, Abe K, et al. (2010) The X-linked imprinted gene family Fthl17 shows predominantly female expression following the two-cell stage in mouse embryos. Nucleic Acids Res 38: 3672–3681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Kobayashi S, Isotani A, Mise N, Yamamoto M, Fujihara Y, et al. (2006) Comparison of gene expression in male and female mouse blastocysts revealed imprinting of the X-linked gene, Rhox5/Pem, at preimplantation stages. Curr Biol 16: 166–172. [DOI] [PubMed] [Google Scholar]
- 10. Pitman JL, Lin TP, Kleeman JE, Erickson GF, MacLeod CL (1998) Normal reproductive and macrophage function in Pem homeobox gene-deficient mice. Dev Biol 202: 196–214. [DOI] [PubMed] [Google Scholar]
- 11. Kim VN, Han J, Siomi MC (2009) Biogenesis of small RNAs in animals. Nat Rev Mol Cell Biol 10: 126–139. [DOI] [PubMed] [Google Scholar]
- 12. Krol J, Loedige I, Filipowicz W (2010) The widespread regulation of microRNA biogenesis, function and decay. Nat Rev Genet 11: 597–610. [DOI] [PubMed] [Google Scholar]
- 13. Ponting CP, Oliver PL, Reik W (2009) Evolution and functions of long noncoding RNAs. Cell 136: 629–641. [DOI] [PubMed] [Google Scholar]
- 14. Wang KC, Chang HY (2011) Molecular mechanisms of long noncoding RNAs. Mol Cell 43: 904–914. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Girardot M, Cavaille J, Feil R Small regulatory RNAs controlled by genomic imprinting and their contribution to human disease. Epigenetics 7: 1341–1348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Nakanishi T, Kuroiwa A, Yamada S, Isotani A, Yamashita A, et al. (2002) FISH analysis of 142 EGFP transgene integration sites into the mouse genome. Genomics 80: 564–574. [DOI] [PubMed] [Google Scholar]
- 17. Watanabe T, Totoki Y, Sasaki H, Minami N, Imai H (2007) Analysis of small RNA profiles during development. Methods Enzymol 427: 155–169. [DOI] [PubMed] [Google Scholar]
- 18. Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5: 621–628. [DOI] [PubMed] [Google Scholar]
- 19. Yamamoto M, Saijoh Y, Perea-Gomez A, Shawlot W, Behringer RR, et al. (2004) Nodal antagonists regulate formation of the anteroposterior axis of the mouse embryo. Nature 428: 387–392. [DOI] [PubMed] [Google Scholar]
- 20. Isotani A, Nakanishi T, Kobayashi S, Lee J, Chuma S, et al. (2005) Genomic imprinting of XX spermatogonia and XX oocytes recovered from XX<–>XY chimeric testes. Proc Natl Acad Sci U S A 102: 4039–4044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Kay GF, Barton SC, Surani MA, Rastan S (1994) Imprinting and X chromosome counting mechanisms determine Xist expression in early mouse development. Cell 77: 639–650. [DOI] [PubMed] [Google Scholar]
- 22. Chureau C, Chantalat S, Romito A, Galvani A, Duret L, et al. (2011) Ftx is a non-coding RNA which affects Xist expression and chromatin structure within the X-inactivation center region. Hum Mol Genet 20: 705–718. [DOI] [PubMed] [Google Scholar]
- 23. Jamieson RV, Tan SS, Tam PP (1998) Retarded postimplantation development of X0 mouse embryos: impact of the parental origin of the monosomic X chromosome. Dev Biol 201: 13–25. [DOI] [PubMed] [Google Scholar]
- 24. Skuse DH, James RS, Bishop DV, Coppin B, Dalton P, et al. (1997) Evidence from Turner’s syndrome of an imprinted X-linked locus affecting cognitive function. Nature 387: 705–708. [DOI] [PubMed] [Google Scholar]
- 25. Takagi N, Sasaki M (1975) Preferential inactivation of the paternally derived X chromosome in the extraembryonic membranes of the mouse. Nature 256: 640–642. [DOI] [PubMed] [Google Scholar]
- 26. Thornhill AR, Burgoyne PS (1993) A paternally imprinted X chromosome retards the development of the early mouse embryo. Development 118: 171–174. [DOI] [PubMed] [Google Scholar]
- 27.Williamson C, Blake A, Thomas S, Beechey C, Hancock J, et al. (2012) MRC Harwell, Oxfordshire. World Wide Web Site - Mouse Imprinting Data and References - Available: http://www.har.mrc.ac.uk/research/genomic_imprinting/. Accessed 2013 Jul 7.
- 28. Lee JT (2000) Disruption of imprinted X inactivation by parent-of-origin effects at Tsix . Cell 103: 17–27. [DOI] [PubMed] [Google Scholar]
- 29. Sado T, Wang Z, Sasaki H, Li E (2001) Regulation of imprinted X-chromosome inactivation in mice by Tsix. Development 128: 1275–1286. [DOI] [PubMed] [Google Scholar]
- 30. Kobayashi H, Sakurai T, Imai M, Takahashi N, Fukuda A, et al. (2012) Contribution of intragenic DNA methylation in mouse gametic DNA methylomes to establish oocyte-specific heritable marks. PLoS Genet 8: e1002440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Smallwood SA, Tomizawa S, Krueger F, Ruf N, Carli N, et al. (2011 ) Dynamic CpG island methylation landscape in oocytes and preimplantation embryos. Nat Genet 43: 811–814. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Chiba H, Hirasawa R, Kaneda M, Amakawa Y, Li E, et al. (2008) De novo DNA methylation independent establishment of maternal imprint on X chromosome in mouse oocytes. Genesis 46: 768–774. [DOI] [PubMed] [Google Scholar]
- 33. Tian D, Sun S, Lee JT (2010) The long noncoding RNA, Jpx, is a molecular switch for X chromosome inactivation. Cell 143: 390–403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Calabrese JM, Sun W, Song L, Mugford JW, Williams L, et al. (2012) Site-specific silencing of regulatory elements as a mechanism of X inactivation. Cell 151: 951–963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Okamoto I, Arnaud D, Le Baccon P, Otte AP, Disteche CM, et al. (2005) Evidence for de novo imprinted X-chromosome inactivation independent of meiotic inactivation in mice. Nature 438: 369–373. [DOI] [PubMed] [Google Scholar]
- 36. Rastan S (1983) Non-random X-chromosome inactivation in mouse X-autosome translocation embryos–location of the inactivation centre. J Embryol Exp Morphol 78: 1–22. [PubMed] [Google Scholar]
- 37. Rastan S, Robertson EJ (1985) X-chromosome deletions in embryo-derived (EK) cell lines associated with lack of X-chromosome inactivation. J Embryol Exp Morphol 90: 379–388. [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Validation of the expression of micro RNAs using qRT–PCR. The qRT–PCR analysis of differentially expressed microRNA candidates mmu-miR-7a, mmu-miR-19b, mmu-miR-30c, mmu-miR-103, mmu-miR-107 and mmu-miR-467 that are presented in Tables 2 and 3. The expression of each miRNA was measured and normalized to that of miR295 as a control. Results are expressed as the mean ± SD (n = 3).
(TIF)
Classification of annotated small RNAs in male and female blastocysts against RepeatMasker and tRNA database.
(XLS)
A list of annotated small RNAs expressed in male and female blastocysts.
(XLS)
A list of the PCR primer sets used in this research.
(XLS)