Abstract
Membrane fusion between vesicles and target membranes involves the zippering of a four-helix bundle generated by constituent helices derived from t- and v-SNAREs found on the target and vesicular membranes. In neurons the protein complexin clamps otherwise spontaneous fusion by SNARE proteins, allowing neurotransmitters and other mediators to be secreted when and where they are needed as this clamp is released. The membrane-proximal accessory helix of complexin is necessary for clamping, but its mechanism of action is unknown. Here, we present experiments using a reconstituted fusion system that suggest a simple model in which the complexin accessory helix forms an alternative four-helix bundle with the t-SNARE near the membrane, preventing the v-SNARE from completing its zippering.
Intracellular membrane fusion is catalyzed by the assembly of SNARE complexes between membranes, forcing their bilayers together (1, 2). Because SNARE complex assembly is strongly favored energetically (3), fusion will occur spontaneously in the absence of additional proteins that may be provided to prevent this from happening. However, in synaptic transmission and hormone release, fusion does not occur until calcium enters the presynaptic cytoplasm when the action potential terminates at the nerve ending. Though synaptic vesicles are primed and ready, with their SNAREs largely zippered (4), they are unable to complete fusion without calcium (5-7). Complexin (CPX) can function as a clamp (8) by binding the helical bundle of SNAREs mediating neurotransmitter release (and related forms of exocytosis) (9-11) at a late stage of zippering but before fusion is completed (12). This, in turn, enables the primary calcium sensor for synaptic transmission, synaptotagmin (SYT, (13-15)), to release the clamp and activate fusion when calcium appears (12).
How, at a molecular level, does CPX clamp SNARE proteins? CPX binds the cis-SNARE complex (the post-fusion, fully assembled four-helix bundle SNARE complex (16) by a helical region near the middle of the CPX polypeptide chain (9, 10). This helix consists of distinct though contiguous domains (Figure 1 A), termed the “central helix” (residues 48-70 in the human sequence) and the “accessory helix” (residues 26–47). The central helix lies along the interface between the v- and t-SNARE, making numerous contacts with both, positioning CPX about half-way along the bundle. The accessory helix (in the post-fusion state) continues away from the bundle towards the membrane and, interestingly, is well-defined in the crystal structure even though it lacks contacts with the SNARE proteins (9, 10). The location of the accessory helix prior to fusion is not known because of the lack of a structure of CPX with a trans-SNARE complex, however, it has been found to be required for clamping in vitro and in vivo (17-19) even though it is dispensable for CPX binding to the cis-SNARE complex (9, 11).
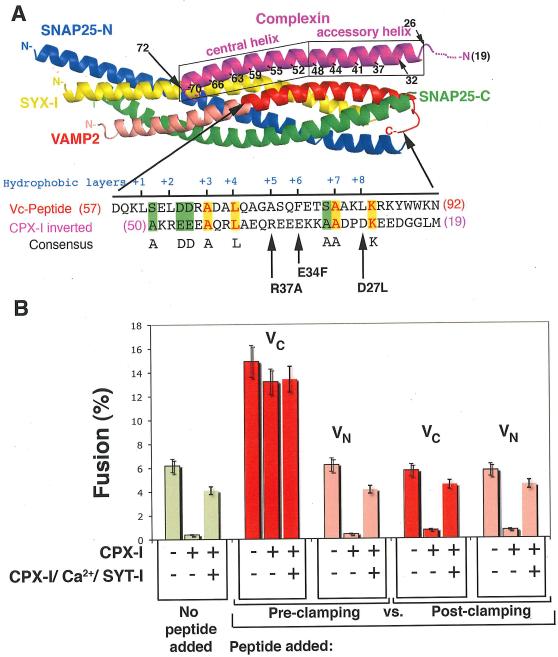
FIGURE 1. CPX-I clamping mechanism.
A-top): Cartoon of the 3-D structure of the CPX-SNARE complex (9). Actual structure of CPX-I spans from residues 32-72. CPX-I region 26-32 was modeled as α-helix according to secondary structure predictions. A-bottom): Amino acid sequence alignment of the membrane proximal half of the VAMP2 SNARE motif and the inverted sequence of the accessory a-helical region highlighting identical residues in yellow, conserved residues in light blue and similar residues in green. The hydrophobic layers are also indicated in red blue. Arrows point to the missing hydrophobic layer on CPX-I sequence and the corresponding site-directed mutation performed. B)- VC-peptide but not VN-peptide competes with CPX-I-GPI. v-SNARE cells transfected with YFP-nls and CPX-I-GPI, or YFP-nls, CPX-I-GPI and SYT-I or YFP-nls alone (control) were used for fusion experiments. 30 μM of VC-Peptide or VN-Peptide were added to the reaction before the commencement of the reaction (Pre-clamped) or added only during the fusion recovery with SYT-I/Ca (Post-clamped), and their corresponding effect was measured as a percentage of fusion. Results are mean ± SEM of three independent experiments.
The striking arrangement of the accessory helix in the post-fusion state suggests the simple idea that it might function as an on-off “switch” for fusion. In the “on” state (cis-SNARE complex) the accessory helix sticks out and the membrane proximal region of the bundle zippers fully. And (hypothetically) in the “off” state (trans-SNARE complex) the accessory helix could potentially interact with the membrane-proximal region of the SNAREs to prevent them from zippering. To act as a clamp, the “off” state would have to be of lower energy in the context of all reactants. Fusion would be triggered from this state when the accessory helix is switched out by activators like SYT.
A clue came from the similarity of the sequence of the accessory helix of CPX to that of the membrane-proximal region of VAMP2 that is only evident when the sequence of the accessory helix is read backward, reflecting the overall anti-parallel physical orientation of the accessory helix to the SNARE bundle (Fig. 1A). Certain key residues within the SNARE motif of VAMP2 (hydrophobic layer +3, +4, +7, K84, D63 and D64) were also found in the accessory helix of CPX-I, when read backwards (Fig. 1A). These residues are among the most conserved in all complexins and this pattern is not seen in attempted alignments of the inverted accessory helix with either syntaxins or SNAP-25. Although weak, this homology made striking predictions which were confirmed in this work, all of which improved function of the clamp, and which derive their strength from the fact that there are many ways to cripple a protein but few to improve it.
The similarity between the accessory helix and VAMP2 suggested that the accessory helix assembles locally with the three helix t-SNARE as an alternative partner to the v-SNARE VAMP2, forming an alternative non-fusogenic four-helix bundle thereby clamping fusion. The other possible model, that the accessory helix binds up the v-SNARE, can be ruled out because VAMP remains accessible in the clamped state to a proteolytic toxin (BoNTB) which recognizes and cleaves in this same region (4, 12).
Our hypothesis for the mechanism of action of the accessory helix in clamping makes several strong predictions: First, peptides from the aligning region of VAMP2 should functionally compete with CPX and prevent clamping. Second, mutations that “add back” the key hydrophobic residues in the sequence of VAMP2 to the CPX accessory helix should improve clamping, resulting in “super-clamp” CPXs that are more poorly released by Ca ion and SYT. Third, super-clamp complexins should compete better with the v-SNARE peptides than wild-type CPX. Fourth, CPXs harboring mutations in the accessory helix should be found which bind to SNARE complexes but do not clamp, and these should dominantly-interfere with clamping by CPXs.
To test the first prediction, we performed fusion experiments involving cells bearing ‘flipped’ exocytic/synaptic v- or t-SNAREs on their surfaces (20) where peptides coding for the N- or C-terminal half of the SNARE motif (helical region) of VAMP2 (VN-Peptide and VC-Peptide, respectively) were added to the “flipped”- SNARE cell fusion reaction before the commencement of the reaction (Pre-clamping) or only during the fusion recovery by PI-PLC and SYT-I/Ca (Post-clamping), and their corresponding effects were measured as the percentage of CPX-I expressing cells that fused. VC peptide forms a stable complex with t-SNARE by binding to the same region of the t-SNARE as the corresponding region of VAMP2, because VC and VN peptides can simultaneously bind to the t-SNARE (21). For liposome-liposome fusion the VC-Peptide (but not VN-Peptide) binding pre-assembles the otherwise poorly stable membrane-proximal region of the t-SNARE and therefore increases the extent of fusion when added to reactions without CPX-I (2, 21). This same effect was observed with cell fusion mediated by flipped SNAREs (20, 22). When VC-Peptide was added simultaneously with CPX-I at the beginning of the fusion reaction (Pre-clamping, Fig. 1B), fusion is dramatically stimulated and became almost completely resistant to the CPX-I clamping effect. VN-Peptide was without effect. When the VC peptide was added only after cells were allowed to interact (Post-clamping), clamping was not reversed.
To test the second prediction - increased affinity of the CPX-I mutants for the t- SNARE complex - we introduced the missing hydrophobic layers on CPX-I by mutating the following residues: R37A (layer +5); E34F (layer +6) and D27L (layer +8), alone or in combination. As predicted, each mutant “super-clamp” (sCPX) clamped better as a soluble protein than wild-type (Fig. 2A) for sCPX E34F/R37A, sCPX D27L/E34F, and wild type. We also tested the clamping ability of these and other designed super-clamps when expressed as GPI-anchored proteins on the surface of v-SNARE expressing cells. Each of the mutants tested clamped the fusion reaction completely (as does wild-type CPX) (Fig. 2B). The super-clamp complexins were also poorly and differentially activated by synaptotagmin and calcium (Fig. 2C). but in every case was activated with the same calcium kinetics as wild-type CPX-I (Fig. S1), indicating that the super-clamping effect is due to a stronger SNARE-“superclamp” CPX-I interaction rather than a failure to interact with its calcium and SYT-I.
FIGURE 2. Mutations on CPX-I designed to mimic hydrophobic layers on VAMP2 stabilize the clamp.
A) Dose dependent inhibition of the cell fusion reaction using different soluble CPX-I mutants (sCPX-I). Increasing concentration of each recombinant sCPX were added at the time the two cell populations were mixed. Cells were allowed to fuse overnight and the fusion efficiency was determined as the percentage of fusion. Results are mean ± SEM of three independent experiments. B) Effect of different cell surface expressed “super-clamp” CPX-I-GPI mutants on cell fusion (blue bars), on the cell fusion recovery after addition of PI-PLC in the absence (green bars) or presence of SYT-I and Calcium (red bars). Experiments are the mean±SEM of three independent experiments. Dashed lines show the maximum cell fusion recovery in the absence (green) or presence of Ca/SYT-I (red) and the total overnight clamping (blue). C) The SYT-I requirement of CPX-I-D27l/E34F-GPI was tested by performing a cell fusion experiment as described in Fig. 2B. In this case the cell fusion recovery was carried out at 200 μM Free Ca2+ and samples were fixed at the indicated time every 5 min. The level of fusion was determined as percentage of transfected v-cells that fused. Results are mean ± SEM of three independent experiments. D) Differential VC-Peptide sensitivity of CPX-I “super-clamp” mutant constructs. Increasing concentrations of VC-peptide were added at the time the two cell populations were mixed. Cells were allowed to fuse overnight and the fusion efficiency was determined as the percentage of fusion. Dashed lines correspond to the basal level of overnight fusion in the absence of CPX-I (green) or in the presence of CPX-I (red). Results are mean ± SEM of three independent experiments
To test the third prediction, we performed titrations of VC-peptide on fusion assays clamped by CPX-I-wt-GPI, CPX-I-D27L/E34F-GPI or not clamped. The VC-peptide competed with the clamp in all cases (Fig. 3D) and clamping by the super-clamp CPX-I-D27L/E34F-GPI was more resistant to VC-peptide than wild-type.
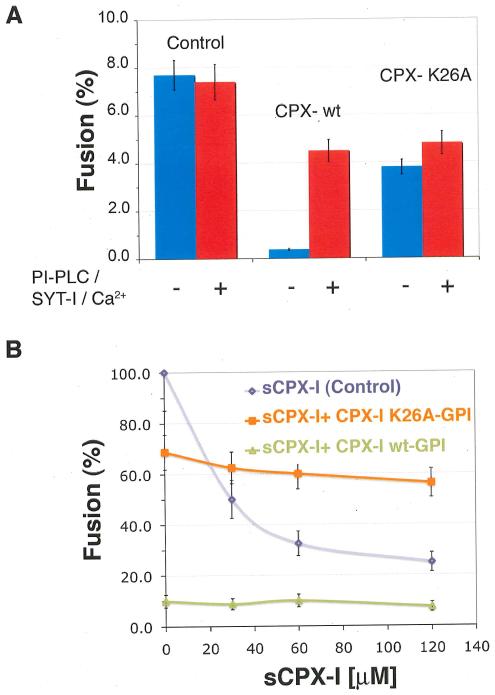
FIGURE 3. Clamping role of the conserved amino acid, K26, in the accessory helix of CPX-I.
A)- Effect of CPX-I-K26AGPI-mutant construct on cell fusion (blue bars), and on the efficiency of cell fusion recovery after addition of PI-PLC in the presence of SYT-I and Calcium (red bars). B)- Dose dependent inhibition of the cell fusion reaction using soluble CPX-I (sCPX-I) and flipped-SNARE expressing cells co-transfected with the indicated CPX-I-GPI mutant construct or mock-transfected (control). Experiments are the mean ± SEM of three independent experiments.
Finally, we tested the fourth prediction, dominant interference. The residues conserved among different CPX isoforms were identified and systematically and individually replaced by alanine or by serine when alanine was present (Fig. S2). Mutation K26A produced a severe reduction in the clamping efficiency of CPX-I-GPI (Fig. 3A). Mutation A44S, which eliminated one of the conserved hydrophobic layer on CPX-I, partially inhibited the clamping activity (Fig. S2 C). The decrease in clamping activity of the CPX-IK26A-GPI and CPX-IA44S-GPI was not due to a lower binding affinity for cis-SNARE complexes (Fig. S3). As predicted, CPX-IK26A-GPI prevented clamping by soluble wild-type CPX-I (Fig. 3B).
This dominant-interfering mutant confirms the independent roles of the central helix (binding) and accessory helix (clamping). But, these roles are also synergistic, because the binding of CPX by its central helix strategically positions the accessory helix for clamping. The central helix binds to residues in both the v- and t-SNARE motifs present in the central and membrane-distal portions of the SNARE bundle (9, 10), ensuring that CPX can only begin to interfere with SNARE assembly after the SNAREpin has zippered at least half-way. The accessory helix then binds weakly and therefore reversibly to sequences in the membrane-proximal portion of the t-SNARE, ideal for a toggle switch.
Figure 4 presents a highly constrained but still speculative molecular model for the clamped state (details are in SOM) that establishes the structural feasibility of the proposed clamped state. Note that the displaced sequences of VAMP2 include both the cleavage site and the protein recognition sequence for cleavage of VAMP2 by Botulinum-B toxin, which can still act on VAMP2 in the clamped state (4, 12), but the recognition sequence for Tetanus Toxin is assembled into the four-helix bundle in the model, explaining why the complexin-clamped intermediate was found to be resistant to this toxin. We note that, in addition to its role as clamp, complexin is also positively required for fusion in an earlier step that requires the central helix and the N-terminal domain of 26 residues, but not the accessory helix (19). Ultimately, high-resolution structural studies will be needed to confirm the general outline of this model and provide intimate details, though the need for membrane-insertion currently prevents this, necessitating less direct, but we believe still forceful alternative approaches to central mechanistic problems in the control of membrane fusion.
FIGURE 4. Model of the proposed mechanism of clamping of an exocytic SNAREpin by complexin.
showing a hypothetical alternative 3-D structure of the clamped state. The 3-D structure of the CPX-cisSNARE complex (9) was modified in accord with super-clamping mutations analyzed in this study. The color code used to label each protein is as follows: CPX (Magenta), VAMP2 VC (red), VAMP2 VN (pink), Syntaxin1 (yellow), the SNAP25 N-terminal helix (green), and SNAP25 C-terminal helix (blue). Membrane anchors are shown as hypothetical helices (grey). The C-terminal end of VAMP2 was displaced from the CPX-cisSNARE structure to accommodate the CPX accessory helix (residues 26-42), which was docked by superimposing CPX Cα positions 34-42 in inverted direction onto VAMP2 Cα positions 69-77. The clamped CPX linker segment (residues 43-52) was built using the Lego-loop feature and regularization in program O (Alwyn Jones). To allow for this clamped CPX docking, the membrane-proximal segment of the VAMP2 helix (residues 60-85) was kinked away from the t-SNARE three-helix bundle after residue 58 by superimposing both this segment and the remaining N-terminal segment onto the AB helix juncture of lamprey hemoglobin (PDB code 2lhb). Precise positioning of the C-terminal portion of VAMP2 (i.e. VC) is arbitrary. The central and accessory helices of CPX-I and the residues involved in the binding with the SNAREs are labeled. The different recognition/binding regions for Botulinum Neurotoxin-B (BoNT/B, residues 62-71) and Tetanus Toxin (TeNT, 38-47) on VAMP2 are indicated as well as their common cleavage site (residue Q76), showing the accessibility of BoNT/B but not of TeNT (12).
Supplementary Material
Acknowledgements
This work is supported by an NIH grant to J.E.R.
REFERENCES
- 1.Sollner T, et al. Nature. 1993;362:318. doi: 10.1038/362318a0. [DOI] [PubMed] [Google Scholar]
- 2.Weber T, et al. Cell. 1998;92:759. doi: 10.1016/s0092-8674(00)81404-x. [DOI] [PubMed] [Google Scholar]
- 3.Li F, et al. Nat Struct Mol Biol. 2007;14:890. doi: 10.1038/nsmb1310. [DOI] [PubMed] [Google Scholar]
- 4.Hua SY, Charlton MP. Nat Neurosci. 1999;2:1078. doi: 10.1038/16005. [DOI] [PubMed] [Google Scholar]
- 5.Wojcik SM, Brose N. Neuron. 2007;55:11. doi: 10.1016/j.neuron.2007.06.013. [DOI] [PubMed] [Google Scholar]
- 6.Koh TW, Bellen HJ. Trends Neurosci. 2003;26:413. doi: 10.1016/S0166-2236(03)00195-4. [DOI] [PubMed] [Google Scholar]
- 7.Sudhof TC. Annu Rev Neurosci. 2004;27:509. doi: 10.1146/annurev.neuro.26.041002.131412. [DOI] [PubMed] [Google Scholar]
- 8.Tang J, et al. Cell. 2006;126:1175. doi: 10.1016/j.cell.2006.08.030. [DOI] [PubMed] [Google Scholar]
- 9.Chen X, et al. Neuron. 2002;33:397. doi: 10.1016/s0896-6273(02)00583-4. [DOI] [PubMed] [Google Scholar]
- 10.Bracher A, Kadlec J, Betz H, Weissenhorn W. J Biol Chem. 2002;277:26517. doi: 10.1074/jbc.M203460200. [DOI] [PubMed] [Google Scholar]
- 11.Pabst S, et al. J Biol Chem. 2000;275:19808. doi: 10.1074/jbc.M002571200. [DOI] [PubMed] [Google Scholar]
- 12.Giraudo CG, Eng WS, Melia TJ, Rothman JE. Science. 2006;313:676. doi: 10.1126/science.1129450. [DOI] [PubMed] [Google Scholar]
- 13.Sudhof TC. Nature. 1995;375:645. doi: 10.1038/375645a0. [DOI] [PubMed] [Google Scholar]
- 14.Geppert M, et al. Cell. 1994;79:717. doi: 10.1016/0092-8674(94)90556-8. [DOI] [PubMed] [Google Scholar]
- 15.Li C, et al. Nature. 1995;375:594. doi: 10.1038/375594a0. [DOI] [PubMed] [Google Scholar]
- 16.Sutton RB, Fasshauer D, Jahn R, Brunger AT. Nature. 1998;395:347. doi: 10.1038/26412. [DOI] [PubMed] [Google Scholar]
- 17.Xue M, et al. Nat Struct Mol Biol. 2007;14:949. doi: 10.1038/nsmb1292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Giraudo CG, et al. J Biol Chem. 2008;283:21211. doi: 10.1074/jbc.M803478200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Maximov A, Tang J, Yang X, Pang ZP, Shudhof TC. Science Submitted. 2008 [Google Scholar]
- 20.Hu C, et al. Science. 2003;300:1745. doi: 10.1126/science.1084909. [DOI] [PubMed] [Google Scholar]
- 21.Melia TJ, et al. J Cell Biol. 2002;158:929. doi: 10.1083/jcb.200112081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Giraudo CG, et al. J Cell Biol. 2005;170:249. doi: 10.1083/jcb.200501093. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.