Abstract
Caries is a multifactorial disease and little is still known about the host genetic factors influencing susceptibility. Our previous genome-wide linkage scan has identified the interval 5q12.1–5q13.3 as linked to low caries susceptibility in Filipino families. Here we fine-mapped this region in order to identify genetic contributors to caries susceptibility. Four hundred and seventy-seven subjects from 72 pedigrees with similar cultural and behavioral habits and limited access to dental care living in the Philippines were studied. DMFT scores and genotype data of 75 single-nucleotide polymorphisms were evaluated in the Filipino families with the Family-Based Association Test. For replication purposes, a total 1,467 independent subjects from five different populations were analyzed in a case-control format. In the Filipino cohort, statistically significant and borderline associations were found between low caries experience and four genes spanning 13 million base pairs (PART1, ZSWIM6, CCNB1, and BTF3). We were able to replicate these results in some of the populations studied. We detected PART1 and BTF3 expression in whole saliva, and the expression of BTF3 was associated with caries experience. Our results suggest BTF3 may have a functional role in protecting against caries.
Keywords: Dental caries susceptibility, Epidemiology, Genetics, Immune response genes, Saliva
Currently, identifying children at risk for caries prior to the occurrence of the disease still depends on classic methods, such as questions about diet, inspection of oral hygiene level, and detection of Streptococcus mutans in saliva, but those attempts have had limited success in determining caries risk at the population level, and the most promising ones reach sensitivity and specificity of around 80% [Gao et al., 2010]. Previous caries experience continues to be the best predictor for future disease [Powell, 1998]. A child’s resistance or susceptibility to caries can occur, regardless of exposure to external risk factors. These facts suggest there is a biological influence on the disease susceptibility, likely controlled by genetic factors.
Although caries is a complex disease with little known about the host genetic factors influencing susceptibility, studies on twins and families have provided strong evidence for the role of genetic inheritance for caries and have estimated the genetic contribution to caries development as 40–60% [Conry et al., 1993; Bretz et al., 2005, 2006; Wang et al., 2010]. The importance of genetic factors in caries is also supported by animal model studies that have identified some chromosomal loci for caries susceptibility [Suzuki and Kurihara, 1998; Nariyama et al., 2004]. Regarding the mode of inheritance, both a mouse crossbreeding study [Nariyama et al., 2004] and a family segregation analysis in humans [Werneck et al., 2011] suggest a major gene dominant effect for caries. Our previous genome-wide linkage scan for caries using 46 Filipino families, which included 624 individuals, identified three loci for low caries susceptibility (5q13.3, 14q11.2, and Xq27.1) and two loci for high caries susceptibility (13q31.1 and 14q24.3). The 5q13.3 locus was the only one that had significant results under a dominant model [Vieira et al., 2008].
Here we present the fine-mapping effort of 5q12.1–13.3 and the follow-up experiments interrogating the levels of gene expression in whole saliva of genes identified in the locus as associated with caries experience in humans. Our hypothesis is that genetic factors located in 5q12.1–13.3 underlie biological mechanisms that influence caries experience in humans.
Materials and Methods
The study sites, subject recruitment and DMFT data collection were the same as that outlined in our original genome-wide linkage study [Vieira et al., 2008], but the sample was increased for further fine-mapping. DNA samples were extracted from peripheral blood according to standard protocols. In an attempt to reduce the influence of environmental factors, the study sample consisted of 477 subjects (224 females and 253 males) from 72 pedigrees living in the same area in the Philippines. Families included were part of existing studies of other craniofacial defects. The mean age of the subjects was 22.6 years and ranged from 2 to 72 years and the mean DMFT score was 9.7 and ranged from 0 to 32. This study protocol was approved by the University of Pittsburgh and H.O.P.E. Foundation International Institutional Review Boards, and appropriate written informed consent was obtained from all participants. Age-appropriate assent documents were used for children between 7 and 14 years of age and allowed us to obtain informed, written consent from the child as well as from the parents. Caries experience data (DMFT scores) were collected by a single examiner who was calibrated by one of the authors (A.R.V.). Intraexaminer agreement was assessed by a second clinical exam in 10 families after 2 weeks, with a κ of 1.0. Three criteria of caries experience level based on age and DMFT scores were used for statistical analysis (table 1 ). DMFT cutoffs of criterion 1 were the same used in the original genome-wide linkage study [Vieira et al., 2008]. To create criteria 2 and 3, DMFT cutoffs were moved one DMFT value from criterion 1. Individuals with low caries experience were compared with individuals with high caries experience.
Table 1.
Single marker association results for caries susceptibility in Filipino families and detailed criteria used for analysis of caries phenotypes
| Position (base pair) |
Marker | Gene | Region | Allele | Frequency | Criterion 1 p value |
Criterion 2 p value |
Criterion 3 p value |
Effect |
|---|---|---|---|---|---|---|---|---|---|
| 59837591 | rs27565 | PART1 | intron 3–4 | T | 0.56 | 0.00021 | 0.0022 | 0.00038 | protective |
| 60621839 | rs4700418 | ZSWIM6 | 5′upstream | G | 0.49 | 0.0089 | 0.0041 | 0.0760 | protective |
| 68469923 | rs875459 | CCNB1 | intron 3–4 | G | 0.61 | 0.0046 | 0.0204 | 0.0019 | protective |
| 72798995 | rs6862039 | BTF3 | intron 4–5 | T | 0.51 | 0.0084 | 0.0461 | 0.0026 | risk |
| Caries experience level | n | Criterion 1 |
Criterion 2 |
Criterion 3 |
|||
|---|---|---|---|---|---|---|---|
| DMFT | n | DMFT | n | DMFT | n | ||
| Children, up to 12 years of age | 115 | ||||||
| Low caries experience | 0–2 | 26 | 0–1 | 19 | 0–3 | 34 | |
| High caries experience | 3 or higher | 89 | 2 or higher | 99 | 4 or higher | 81 | |
| Teenagers,13–19 years of age | 104 | ||||||
| Low caries experience | 0–5 | 44 | 0–4 | 40 | 0–6 | 52 | |
| High caries experience | 6 or higher | 60 | 5 or higher | 64 | 7 or higher | 52 | |
| Adults, 20 years of age and older | 258 | ||||||
| Low caries experience | 0–8 | 109 | 0–7 | 87 | 0–9 | 114 | |
| High caries experience | 9 or higher | 149 | 8 or higher | 161 | 10 or higher | 134 | |
Global p < 0.05 is presented in bold. Underlined p value indicates statistical significance after correction for multiple tests (significance set at p < 0.00035). Definition of caries experience level based on age and DMFT scores in Filipino families.
5q12.1–5q13.3 covers a 1-LOD interval (upstream and downstream) from the point with the maximum LOD score on chromosome 5 from our previous work in the Philippines [Vieira et al., 2008]. We selected single-nucleotide polymorphisms (SNPs) to study the interval 5q12.1–5q13.3, covering about 18 million base pairs and spanning 50 genes, by using the data from the International HapMap Project on Caucasians and Chinese (www.hapmap.org), viewed through the software Haploview [Barrett et al., 2005]. Based on pairwise linkage disequilibrium, haplotype blocks, and structure of genes, we selected 50 SNPs in the region. Subsequently we increased the density of markers around SNPs with significant results and typed 25 additional SNPs for a total of 75 SNP markers. Genotyping was performed with Taqman chemistry on an ABI 7900HT real-time PCR equipment and the data were read using the ABI SDS software (Applied Biosystems, Calif., USA). Association between caries experience and the SNPs was tested with the transmission disequilibrium test within the statistical package Family-Based Association Test [Horvath et al., 2001]. Correction for multiple comparisons in single marker analyses was performed by the SNP spectral decomposition method [Nyholt, 2004]. We chose this method over the popular Bonferroni correction to avoid overcorrecting our results and reducing power since the genetic markers we studied were purposely chosen to be not in complete linkage disequilibrium. Significance was set at p < 0.00035. Calculations of linkage disequilibrium were computed with the Graphical Overview of Linkage Disequilibrium software [Abecasis and Cookson, 2000].
Replication studies targeted SNPs with significant or suggestive association with caries experience in the Filipino dataset. All DNA samples studied in the replication studies were extracted from whole saliva by the use of Oragene kits (DNA Genotek, Inc., Ottawa, Ont., Canada) according to the manufacturer’s instructions. Subjects were asked to spit 2–4 ml of saliva within 5–10 min. Absorbing sponges provided by the manufacturer were used for children younger than 7 years of age. When the allele frequency was low in a specific replication group, we selected additional markers in the same region. Hardy-Weinberg equilibrium tests were performed to test for deviations in the genotype and allele distributions. We used logistic regression analysis and haplotype analysis (as performed by the PLINK analysis software [Purcell et al., 2007]) to investigate main-effect models to predict caries status. Since the definitions of caries experience are based on age, there was no need to adjust with age in the regression process. For genetic association, a nominal p value of 0.05 was considered statistically significant.
The study sample from Istanbul, Turkey, consisted of 172 unrelated children (93 females and 79 males) from 3 to 6 years of age. All children younger than 6 years of age visiting the Pedodontics Department for treatment during 2007 were invited to participate in the study. Ninety children had a dmft score of 4 or more and 82 children were caries-free [Patir et al., 2008]. Children with dmft scores between 1 and 3 were not included. Both Istanbul University and University of Pittsburgh Institutional Review Boards approved the study of these samples and appropriate written informed consent was obtained from the parents of all participants.
The cohort from Tiquisate, Guatemala consisted of 113 DNA samples from unrelated individuals (70 females and 43 males, mean age of 29 years) who participated in studies conducted by the University of Pittsburgh Center for Craniofacial and Dental Genetics, specifically a 2006 research trip in collaboration with a Children of the Americas medical mission. As part of the research protocol the cohort received a dental examination [Deeley et al., 2008]. No individuals born with clefts were included in the analysis. These samples were collected with the approval of both the University of Pittsburgh Institutional Review Board and the Oversight Ethics Committee of the Hospital Nacional de Tiquisate, and all subjects gave informed, written consent. Age-appropriate assent documents were used for children between 7 and 14 years of age and allowed us to obtain informed, written consent from the child, as well as from the parents.
From Argentina, 143 DNA samples from unrelated individuals living in 12 Patagonia sites were studied (San Carlos de Bariloche, El Bolsón, Esquel, El Maitén, Maquinchao, Ingeniero Jacobacci, Rio Colorado, Choele Choel, Valcheta, Sierra Grande, Santo Antonio Oeste, and General Roca), again as part of the studies at the Center for Craniofacial and Dental Genetics. No individuals born with clefts were included in the analysis. The mean age was 21.7 years (between infants under 1 and 72 years with median of 18 years) and both the Centro de Educación Médica e Investigaciones Clínicas ‘Norberto Quirno’ and University of Pittsburgh Institutional Review Boards approved the study of these samples and appropriate written informed consent was obtained from all participants (parents provided consent for the participation of individuals 17 years of age and under).
From Brazil, two sample data sets were available for this study. The first consisted of 539 unrelated children living in Curitiba (261 females and 278 males) with the mean age of 12 years (between 10 and 14 years). All children registered in the 5th, 6th, or 7th grades of nine public schools during the year 2006 were invited to participate in this study [Brancher et al., 2011]. These samples were used with the approval of both the Pontifical Catholic University of Paraná and the University of Pittsburgh Institutional Review Boards. Age-appropriate assent documents were used for all children and informed, written consent was obtained from the parents. The second group of DNA samples was from 500 unrelated subjects living in Rio de Janeiro participating in studies of other craniofacial anomalies (236 females and 264 males) with a mean age of 9 years (between 2 and 21 years) [Tannure et al., 2012]. These samples were used with the approval of both the Federal University of Rio de Janeiro and the University of Pittsburgh Institutional Review Boards. Age-appropriate assent documents were used for children between 7 and 14 years and informed, written consent was obtained from the child, as well as from the parents.
Caries experience data (DMFT/dmft) was collected according to established protocols. In brief, visible lesions in dentin, as well as visible active lesions in enamel (white spots) were scored as decayed. An explorer was gently used for assessing the smoothness of tooth surfaces. Gauze was used to dry and clean teeth prior to exam. Artificial light and a dental operatory were used for all evaluations, with the exception of the Philippines and Guatemala, where a bed was used to lay down subjects. Exam calibrations were performed according to the following protocol: first, the calibrator presented to the examiner(s) the criteria for caries detection, showing pictures of several situations to be observed in the exam (sound and decayed tooth surfaces, filled teeth with and without secondary lesions, missing teeth due to caries or due to other reasons) and discussing each of these situations in a session that last 1–2 h. Next, the calibrator and examiner(s) examined 10–20 subjects and discussed each case. In the case of data from Turkey, one of the authors (A.P.) carried out the clinical examination after being calibrated by an experienced specialist (F.S.). The intraexaminer agreement was assessed by a second clinical exam in 10% of the sample after 2 weeks, with a κ of 1.0. In the cases of Argentina and Guatemala, data was collected by one single experienced specialist examiner (A.R.V.). Subjects in these projects are seen over a period of no longer than 3–5 days and intraexaminer agreement data could not be generated. Samples from Rio de Janeiro, Brazil, were collected by two examiners (E.C.K. and P.N.T.) and calibrated by an experienced specialist (M.C.C.). The intraexaminer agreement was assessed by a second clinical exam in 20 children after 2 weeks, with a κ of 0.99. Cohen’s kappa values for agreement between examiners were 0.91. Finally, caries experience data from Curitiba, Brazil, were collected by two examiners (J.A.B. and G.D.P.) and calibrated by an experienced specialist (P.C.T.). Inter- and intraexaminer reproducibility was taken on 10% of the sample and κ values were 0.93 for inter- and 0.99 for intraexaminer reliability.
Mutation Analysis for Candidate Genes
We carried out mutation analysis for four genes flanking SNPs that showed significant or suggestive association with caries experience in the Filipinos. These genes included the prostate androgen-regulated transcript 1 (PART1), zinc finger, SWIM-type containing 6 (ZSWIM6), cyclin B1 (CCNB1), and basic transcription factor 3 (BTF3).
Samples carrying two copies of the associated SNP alleles were selected for sequencing and all exons and intron-exon boundaries of these genes were interrogated. Ninety-five DNA samples were sequenced for PART1, 123 for ZSWIM6, 131 for CCNB1, and 130 for BTF3. Reference sequences were obtained from the Ensembl Genome Browser (http://useast.ensembl.org/index.html). The primers used for PCR amplification are shown in table 2. DNA was amplified by 35 cycles of at 95 ° C for 30 s, 55 ° C for 30 s, and 72 ° C for 1 min and PCR products were directly sequenced using an ABI PRISM BigDye™ Terminator Cycle Sequencing Ready Reaction Kit and an ABI 3730xl DNA Analyzers (Applied Biosystems, Calif., USA). Sequences obtained were verified against the sequences in the Ensembl Genome Browser and two unrelated CEPH (Foundation Jean Dausset-Centre d’Etude du Polymorphisme Humain) controls. For sequence variations between high and low caries groups defined by criteria 1 in table 1, χ2 tests were performed to test for deviations in the genotype and allele distributions from Hardy-Weinberg equilibrium.
Table 2.
Primer sets for sequencing of PART1, ZSWIM6, CCNB1, and BTF3 genes
| Gene exon |
Forward primer (5′–3′) | Reverse primer (5′–3′) | Product size (base pair) |
|---|---|---|---|
| PART1 | |||
| 1 | cattaaggcaggaactggca | gcatatctagttgcctcagc | 488 |
| 2 | ccagctcagttacagcttgc | tggtaactagtggagagagg | 368 |
| 3 | ggtgggccctttaagaggtg | gcccagtggagcattcattc | 403 |
| 4–1 | gatgggaaatgagttgtgag | ctggctgtcttaaatactcc | 1023 |
| 4–2 | gataccaactatggcctctc | gtagggtgactctcccatac | 931 |
|
| |||
| ZSWIM6 | |||
| 1 | tagtgccgtttatagggtcc | ttcatttcccactcgcgctg | 945 |
| 2 | cttgctgctgtcaagtccca | gaccaactaagagcaatgctg | 579 |
| 3 | gcatggcctacacagcttta | gtctaagatcacctaaggaa | 666 |
| 4 | gctggtgtgaaggcttgaca | acacgttccatgcatgagca | 448 |
| 5 | ctctctgtgggatttgtggc | gctcctcaagcctcttgaca | 791 |
| 6 | gttcatgaacatctggccac | gccaagcattcatccatctc | 378 |
| 7 | ggagatggatgaatgcttgg | ggcgagtaaagccatcatc | 442 |
| 8 | aaagcctctgttttccccac | cgtgaaagcatgacagtgcc | 384 |
| 9 | catttgcttgtacggggaag | cccatgactatagcaatcagg | 481 |
| 10 | acggatgctataccctgaag | gccaacaagcaaatcacctc | 479 |
| 11 | tgaacttgagtagaggcctg | gacattaacaccactgctgag | 380 |
| 12 | gggagccctagttgtaacac | gttcagctgggaatgaccac | 524 |
| 13 | cttgccaggaggagaatgtg | gctagaaggagccttggaga | 415 |
| 14 | cagcctcctcgtcagtaata | ataccggtgtggctctgttc | 1015 |
|
| |||
| CCNB1 | |||
| 1 | actggcttcactgctctcca | accttcctccccaaatcgctc | 514 |
| 2 | actctgggacccatgttttc | tgcatcaaaggccacagttg | 316 |
| 3 | aggtaactctcttcctgacc | ttgcctcatgctaagcactc | 383 |
| 4 | gccgattcagcagaatactag | cttaagaaatgctgcccacg | 400 |
| 5 | taaatgccccagtgctactg | ggttttactgcgttggccag | 425 |
| 6 | ttcttgggggatatggtgtc | tcccttccccatgtagttag | 475 |
| 7 | ggactttccatgggcatttg | acagagcgagaccttgtctc | 451 |
| 8, 9 | ccatctcttttccactcctg | cgctaccctacagtaggaag | 653 |
|
| |||
| BTF3 | |||
| 1 | ctcgctggaccatcacaaac | cccgcaaagctccaaatttg | 673 |
| 2 | atttggagctttgcggggtg | cgtgaatttccctcgagagg | 338 |
| 3 | ctgcctgctcactgcataag | cagacttaatgagctccctg | 426 |
| 4 | cacctgtgcttcacagtgaa | catatccattctgagccctg | 393 |
| 5 | cccaaatgtcccttatcatgg | cttccctgggtagtttttcc | 278 |
| 6 | cttggggctatgacacagta | cagctcccagtaacttcttc | 223 |
mRNA Expression Analysis for Candidate Genes
Whole saliva RNA samples were obtained from 58 individuals from the Argentinean population described above (19 with high caries experience and 39 with low caries experience) and total RNA from saliva was isolated using RNeasy Micro kit (Qiagen, Calif., USA). mRNA expression analysis of PART1, ZSWIM6, CCNB1, and BTF3 was performed. We also studied the nuclear factor, interleukin-3 regulated (NFIL3), which is predicted to bind to PART1 promoter, and nuclear factor kappaB1 (NFKB1), which is predicted to bind to CCNB1 and BTF3 promoters. These predictions were made by the use of the SABiosciences’ Text Mining Application (Qiagen Inc., Calif., USA). Total RNA from the submaxillary salivary gland epidermoid carcinoma cell line HTB-41™ (American Type Culture Collection, Manassas, Va., USA) was also isolated and studied. One hundred nanograms of total RNA from whole saliva was reverse-transcribed with the High Capacity cDNA Reverse Transcription kit (Applied Biosystems, Calif., USA). The primers used for the RT-PCR and real-time PCR are listed in table 3. β-Actin was used as endogenous control. For real-time PCR analysis, PCR amplification of cDNA was performed by SYBR Green PCR Master Mix (Applied Biosystems, Calif., USA). Products spanned at least one intron, so that cDNA products were distinguishable from potential genomic DNA products. After an initial denaturation at 95 ° C for 5 min, 40 cycles at 95 ° C for 45 s, 55 ° C for 45 s, and 72 ° C for 90 s were performed in a 7900HT real-time PCR system. The quantification of mRNA expression levels relative to β-Actin was performed by 2DDCT method [Livak and Schmittgen, 2001]. Differences between low and high caries experience samples were analyzed by using the Student’s t test; p < 0.05 was considered statistically significant.
Table 3.
Primer sets for RT-PCR experiments
| Gene | Forward primer (5′–3′) | Reverse primer (5′–3′) | Product size (base pair) |
|---|---|---|---|
| PART1 | acctggagcaaggtctcttc | ccatctcagcctggataatc | 216 |
| ZSWIM6 | ttggaaagcggctgcgtaga | cggatgcggtataaagacag | 212 |
| CCNB1 | ccaagcccaatggaaacatc | acttcccgacccagtaggta | 212 |
| BTF3 | ggaactgctcgcagaaagaa | ggtttaagatgctgggtagc | 256 |
| ACTB | ggcacccagcacaatgaag | ccgatccacacggagtacttg | 66 |
| NFIL3 | ctccacagcatatgctcaag | ctgcgtgtgttctactgagg | 211 |
| NFKB1 | caagaagtcttaccctcagg | gtatacccaggtttgcgaag | 194 |
PART1 = Prostate androgen-regulated transcript 1; ZSWIM6 = zinc finger, SWIM-type containing 6; CCNB1 = cyclin B1; BTF3 = basic transcription factor 3; ACTB = β-actin; NFIL3 = nuclear factor, interleukin 3 regulated; NFKB1 = nuclear factor kappaB1.
Results
Out of 75 SNPs used for fine-mapping the target chromosomal region, markers in four genes were overtransmitted depending on caries experience (table 1 ). Statistically significant association between the marker rs27565 located in intron 3 of PART1 and caries experience was detected. The T allele of this marker was associated with low caries experience (p = 0.00021 in criterion 1, p = 0.00038 in criterion 3; details on the criteria in table 1), indicating that this allele has a protective effect for caries. Only this marker remained significant after the correction for multiple testing. The G allele of rs4700418 in ZSWIM6 (p = 0.0089 in criterion 1, p = 0.0041 in criterion 2) and the G allele rs875459 in CCNB1 (p = 0.0046 in criterion 1, p = 0.0019 in criterion 3) were suggestively associated with low caries experience and also the T allele of rs6862039 in BTF3 (p = 0.0084 in criterion 1, p = 0.0026 in criterion 3) was suggestively associated with high caries experience. No strong linkage disequilibrium was apparent between these markers based on D’ and r2 values (data not shown).
The study population for replication purposes consisted of 1,467 individuals from five populations including Turkish, Guatemalan, Argentinean, and two Brazilian cohorts. Caries experience was defined as described in table 1. Details regarding the demographics and caries experience of the five study groups are presented in table 4. With the exception of Brazil, drinking water in the regions where samples originated is not artificially fluoridated.
Table 4.
Demographics and caries experience of the replication study populations
| Filipinos | Turkish | Guatemalan | Argentinean | Brazilian Curitiba |
Brazilian Rio de Janeiro |
|
|---|---|---|---|---|---|---|
| Sample size | 477 (9.7±7.3) | 172 (3.8±4.0) | 113 (7.1±6.9) | 143 (7.1±7.8) | 539 (1.4±1.9) | 500 (2.4±3.0) |
| High-caries group | 298 (13.3±6.7) | 92 (7.2±2.3) | 42 (15.0±4.2) | 66 (13.0±7.9) | 117 (4.4±1.5) | 171 (5.8±2.6) |
| Low-caries group | 179 (3.6±2.4) | 80 (0) | 71 (2.4±2.6) | 77 (2.0±2.3) | 410 (0.6±0.9) | 329 (0.6±0.9) |
| Females | 224 | 93 | 70 | 83 | 261 | 236 |
| Males | 253 | 79 | 43 | 60 | 278 | 264 |
| Mean age ± SD, years | 25.8±16.3 | 5.4±0.8 | 29.0±10.5 | 21.7±15.6 | 12.0±0.5 | 9.1±3.1 |
| Pedigrees, n | 72 | unrelated | unrelated | unrelated | unrelated | unrelated |
Figures in parentheses indicate mean DMFT ± SD. High and low caries experience was defined based on criterion 1 in table 1.
In the Turkish case-control cohort, we detected significant association (p = 0.0002) between the G allele of CCNB1 rs875459 and low caries experience. The G allele is the common allele with frequency of 54% in this cohort but was overrepresented in the low caries individuals (56%) as compared to the high caries individuals (36%). The odds ratio for this allele was 2.3 with 95% confidence interval (1.5–3.5). The p value of 0.0002 surpassed the Bonferroni threshold of p = 0.0125 for statistical significance. This supports the same result that was found in the Filipino families.
In the Brazilian cohort from Rio de Janeiro, we detected a suggestive association (p = 0.03) between the C allele of BTF3 rs703882 and low caries experience. The C allele has a frequency of 49.7% in this cohort but is overrepresented in the low-caries individuals (54%) as compared to the high-caries individuals (32%). The odds ratio for the C allele was 2.41 with 95% confidence interval (1.1–5.31). The p value of 0.03 does not meet the Bonferroni threshold of p = 0.0125 for statistical significance. This suggestive result supports the same result in the Filipino families. No other results from the Filipino families were replicated in the other cohorts (table 5 ). No etiologic variants were detected in PART1, ZSWIM6, CCNB1, and BTF3 by sequencing. Two sequence variants were identified in the exonic region of PART1, although PART1 is not predicted to encode a protein product. The variant 1555A>G (rs153152) in exon 4 was associated with caries experience (A:G = 100: 2 in the high-caries experience group, A:G = 78: 10 in the low-caries experience group, p = 0.0079). The variant 354T>C (rs26949) in exon 2 did not show statistically significant difference (T:C = 95: 7 in high caries experience, T:C = 76: 12 in low caries experience, p = 0.12). Strong linkage disequilibrium (D’ = 0.88) was found between the 1555A>G mutation in PART1 exon 4 and rs27565 in PART1. We identified two silent synonymous variants in the coding region of ZSWIM6 (c.3411C>A; rs16892374) and CCNB1 (c.966G>A; rs1128761), however, the allele frequency distribution between high- and low-caries samples was not statistically significantly different (synonymous c.3411C>A in ZSWIM6 exon 14, C:A = 167: 1 in high caries experience, C:A = 77: 1 in low caries experience, p = 0.57; and c.966G>A in CCNB1 exon 7, G:A = 105: 21 in high caries experience, G:A = 115: 21 in low caries experience, p = 0.79). No sequence variations were detected in the coding region of BTF3.
Table 5.
Single marker association results for caries susceptibility in replication study
| Marker | Gene | Region | Turkish p value (genotype/allele) |
Guatemalan p value (genotype/allele) |
Argentinean p value (genotype/allele) |
Brazilian, Curitiba p value (genotype/allele) |
Brazilian, Rio de Janeiro p value (genotype/allele) |
|---|---|---|---|---|---|---|---|
| rs27565* | PART1 | intron3–4 | 0.19/0.20 | ||||
| rs40512 | PART1 | intron1–2 | 0.34/0.36 | 0.22/0.23 | 0.98/0.98 | 0.38/0.40 | |
| rs4700418 | ZSWIM6 | 5′ upstream | 0.61/0.60 | 0.84/0.84 | 0.11/0.11 | 0.31/0.31 | 0.79/0.79 |
| rs875459 | CCNB1 | intron3–4 | 0.0007/0.0002 | 0.62/0.62 | 0.68/0.69 | 0.47/0.46 | 0.16/0.21 |
| rs6862039* | BTF3 | intron4–5 | 0.29/0.31 | ||||
| rs703882 | BTF3 | intron5–6 | 0.12/0.11 | 0.98/0.98 | 0.29/0.27 | 0.03/0.03 |
Global p < 0.05 are presented in bold.
Markers originally associated with low caries experience in the Filipinos were not informative in Turkish, and different markers very close and in linkage disequilibrium to the original markers were chosen.
The initial linkage result was replicated by association approaches that narrowed our search to four genes. However, we could not identify any obvious genetic variants that explain individual susceptibility to caries by direct sequencing, nor do the associated SNP markers appear to have a functional role. Hence, we decided to evaluate if these genes are expressed in saliva and if expression of these genes correlated to caries experience. Total RNA from the submaxillary salivary gland epidermoid carcinoma cell line HTB-41 ™ and whole-saliva RNA samples obtained from 58 individuals from the Argentinean population used in the replication step (19 with high caries experience and 39 with low caries experience) were analyzed.
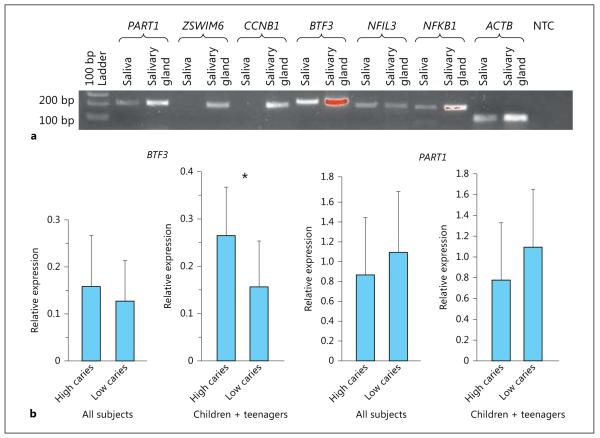
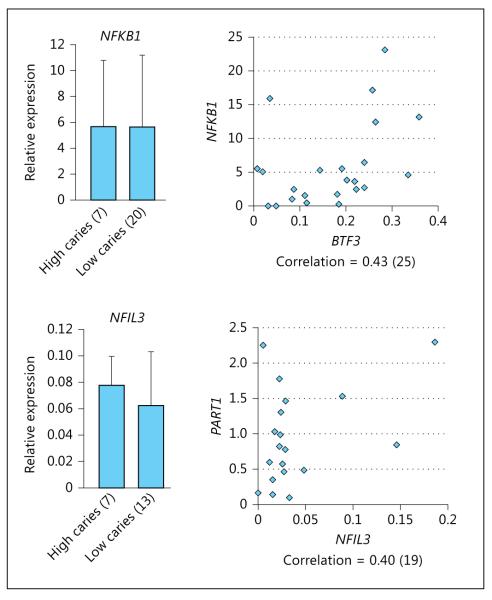
mRNA expression of all genes tested [ PART1, ZSWIM6, CCNB1, BTF3, NFIL3 (which is predicted to bind to PART1 promoter), and NFKB1 (which is predicted to bind to CCNB1 and BTF3 promoters)] was detected in the submaxillary salivary gland cell line. In RNA extracted from whole saliva, only PART1, BTF3, NFIL3, and NFKB1 were detected in a subset of the individuals (fig. 1a). In real-time PCR analysis, mRNA expression of BTF3 in whole saliva was statistically significantly different between children and teenagers with high and low caries experience (fig. 1b). No significant differences were detected in adults. mRNA expression of PART1 did not show significant differences between individuals with high and low caries experience, no matter the age group. PART1 expression showed significant correlation with NFIL3 expression (r = 0.40), and BTF3 expression was significantly correlated with NFKB1 expression (r = 0.43, fig. 2 ).
Fig. 1.
a mRNA expression of PART1, ZSWIM6, CCNB1, BTF3, NFIL3, and NFKB1 in saliva and submaxillary salivary gland by RT-PCR analysis. NTC = No template control for β-Actin (ACTB). b mRNA expression levels of PART1 and BTF3 in saliva from 58 Argentinean individuals (19 with high caries experience and 39 with low caries experience) by real-time PCR analysis. Relative expression values are the means ± SD. * p < 0.05 by t test.
Fig. 2.
mRNA expression levels of NFKB1, which is predicted to bind BTF3 promoter, and NFIL3, which is predicted to bind PART1 promoter, in whole saliva assessed by real-time PCR. Relative expression values are the means ± SD. Correlation between BTF3 and NFKB1, and correlation between PART1 and NFIL3 were also calculated. The number in parentheses indicates the number of individuals. NFKB1 and NFIL3 expression are not different between high- and low- caries experience samples but PART1 expression showed significant correlation with NFIL3 expression, as well as BTF3 expression was significantly correlated with NFKB1 expression.
Discussion
Caries is still a major problem in most industrialized countries, affecting 60–90% of schoolchildren and the vast majority of adults [World Health Organization, 2003]. Our previous genome-wide linkage scan has identified the interval 5q12.1–5q13.3 as linked to low caries susceptibility in Filipino families [Vieira et al., 2008]. We fine-mapped this region with 477 subjects from 72 pedigrees with similar cultural and behavioral habits and limited access to dental care living in the Philippines. For replication purposes a total of 1,467 independent subjects from five different populations were analyzed in a case-control format. Statistically significant associations were found between low caries experience and four genes (PART1, ZSWIM6, CCNB1, and BTF3). We were able to replicate these results in some of the populations studied and we detected PART1 and BTF3 expression in whole saliva. The expression of BTF3 was associated with caries experience. Our results suggest that BTF3 may have a functional role in protecting against caries.
Our study exemplifies the difficulties of interpreting genetic associations. The initial linkage result was replicated by association approaches that narrowed our search to four genes, however, we could not identify any obvious genetic variants that explain individual susceptibility to caries by direct sequencing. Hence, we decided to evaluate if these genes are expressed in saliva and if expression of these genes correlated with caries experience. These studies suggested that BTF3 may be involved in caries susceptibility acting through saliva.
Based on the common disease-common variant theory, we postulated a common variant as the culprit for the observed association. Although we identified two variants in our sequencing effort, it was not clear they are causal. Another possibility, however, is that it is not that a noncausal variant is associated with a causal variant assuming they are in strong linkage disequilibrium, but there is an indirect association between a common variant and at least one and possibly many rarer causal variants with frequencies as low as 0.005 in the population [Dickson et al., 2010]. If that is the case, it is not surprising that multiple common variants are found to be independently associated with the disease phenotype, like it happened in our study. This also could justify the initial linkage results we obtained [Vieira et al., 2008]. This possibility also guided our decision towards directly testing if the expression of the genes flanking the associated markers could be detected in whole saliva.
The purpose of this study was to follow up initial positive results of a genome-wide linkage study and fine-map the 5q12.1–q13.3 chromosomal region for caries susceptibility in 72 multigenerational families (477 total individuals) recruited from the central part of the Philippines, mostly Cebu island. Socioeconomic status is correlated with oral hygiene habits, dietary composition and access to oral health care [Adair et al., 2004]. Socioeconomic status of our subjects is very homogeneous throughout the island. All families are small-scale fishermen or landless rural dwellers, therefore with similar cultural and behavioral habits and limited access to dental care. In addition, the Philippines is one of the countries with the highest caries experience scores at 12 years of age in the world [World Health Organization, 2003] and they are not under the influence of major nongenetic protective factors, such as high quality of oral hygiene, fluoride exposure, and sugar-free diet. Hence, this population is well characterized and suitable for genetic analysis of caries susceptibility.
We tested three variations of caries experience (DMFT scores) cutoff definitions modifying our original study definition [Vieira et al., 2008] and the results suggested that these variations did not importantly affect the findings. Our definition takes into account age, the biggest confounder of DMFT scores, which tend to increase over a lifetime. We found that rs27565 in PART1 was significantly associated with low caries experience, and three markers in ZSWIM6, CCNB1, and BTF3 were suggestively associated with caries experience. These results were replicated in some of the additional study populations (BTF3 in Rio de Janeiro, CCNB1 in Turkey). None of the results were replicated in the Guatemalan or Argentinian samples. There were several differences between these five replication groups and the Filipinos: age for the Turkish and Brazilian (adults vs. children), fluoridated water and lower caries experience in Brazil, socioeconomic status, and cultural norms and behaviors.
In the Brazilian cohort, the Rio de Janeiro cohort showed association with BTF3 and low caries but the Curitiba cohort did not. The Rio de Janeiro sample was younger on average (9 years vs. 12 years) but was more varied (standard deviation was 5 years with a range of 2–21 years vs. Curitiba 6 months standard deviation and a narrower range between 10 and 14). The Rio de Janeiro sample had 59% of their sample under age 10 years (minimum for the Curitiba sample) and 35% of their sample with ages between 10 and 14 years (range of Curitiba sample). These differences may reflect undetected confounders that cannot possibly be easily studied such as temperature variations and their correlation with water consumption, perception of individual oral health, or dental and health literacy. We also cannot exclude the possibility that sample sizes for each individual cohort were not large enough to identify possible genetic contributions. Hence, based on our data, we cannot conclude that a genetic contribution of the 5q locus studied to caries does not exist.
The four genes (PART1, ZSWIM6, CCNB1, and BTF3) span over 13 million base pairs in a region that includes approximately 50 genes, which were covered during finemapping. Direct sequencing of these genes also identified a higher frequency of an exonic sequence variation in PART1 in individuals with lower caries experience. PART1 belongs to the long noncoding RNA (ncRNA) and little is still known about its biological function. PART1 has exhibited increased expression upon exposure to androgens in the LNCaP prostate cancer cell line and has been involved in the androgen receptor-regulated gene network of the human prostate. PART1 expression has been detected predominantly in the prostate with no expression detected in 15 major adult tissues, but with expression detected in salivary gland tissue [Lin et al., 2000]. A number of reports have identified thousands of actively expressed long ncRNA transcripts with distinct properties. The long ncRNAs show differential expression patterns and regulation in a wide variety of cells and tissues, adding significant complexity to the understanding of their biological role. Coexpression of long ncRNA and protein-coding genes suggests regulatory functions [ørom and Shiekhattar, 2011]. Several transcription factors with immune responses are predicted to bind PART1 promoter, including nuclear factor interleukin-3 (NFIL3)-regulated, nuclear factor of activated T cells 1 (NFATC1) and regulatory factor X1 (RFX1) according to information from the National Center for Biotechnology Information (http://www.ncbi.nlm.nih.gov/). Transcription factor NFIL3 plays an important role in immune response, controls IgE production, and generation of the natural killer cell lineage [Rothman, 2010]. Mice homozygous for a Nfil3 knockout allele exhibit decreased immune response [Gascoyne et al., 2009]. We detected mRNA expression of NFIL3 in whole saliva and salivary glands. Although no significant differences were detected in the expression of PART1 mRNA in whole saliva between individuals with low and high caries experience, coexpression of PART1 and NFIL3 or transcription factors with immune responses in saliva and salivary glands may have antimicrobial activity and modulate the levels of oral cariogenic organisms.
The biological function of ZSWIM6 is unknown because the predicted amino acid sequence of the cDNA shows no homology with any other proteins. Hence, we focused our follow-up effort on BTF3, since the relative expression of this gene in whole saliva was significantly associated with caries experience. BTF3 is involved in the initiation of transcription by RNA polymerase and is also involved in cell cycle regulation, apoptosis, and the transcriptional regulation of tumor-associated genes [Kusumawidjaja et al., 2007]. The protein encoded by CCNB1 is a regulatory protein involved in mitosis, and the gene product complexes with p34 (cdc2) protein kinase to form the maturation-promoting factor, which is required for cells to undergo mitosis [Brandeis et al., 1998]. So far, there is no evidence that these two genes are associated with caries susceptibility, however, nuclear factor kappaB1 (NFKB1), which is involved in the regulation of the immune response, is predicted to bind BTF3 and CCNB1 promoters according to National Center for Biotechnology Information. Homozygous Nfkb1 -null mice have abnormal T cell development and decreased number of peripheral T cells, and abnormal humoral responses with decreased immunoglobulin class switching [Sha et al., 1995]. mRNA expression of NFKB1 was detected in whole saliva and salivary gland and correlated with expression of BTF3, therefore coexpression of BTF3 and NFKB1 in whole saliva or salivary gland may have an antimicrobial effect on oral cariogenic organisms.
Caries in general currently lacks a biologically relevant diagnostic scheme. Traditional measurements quantify the sequelae of the disease (the caries lesions) but do not offer much insight into the disease process, particularly if taken in adults. Hence, our work is groundbreaking because it is identifying a biological mechanism that may help elucidate the reasons why people are differently affected by caries. Our study design includes groups with very high caries experience that benefit very little from current knowledge about disease pathogenesis. In these groups (i.e., Filipinos) the disease runs its natural course and most of protective factors may be related to individual biological mechanisms. When compared to groups that may benefit from measures that improve oral health, such as good oral hygiene, fluoride in the drinking water, and better socioeconomic status (i.e., Brazilians), levels of caries experience are not only lower, the identification of biological factors protecting against caries is likely more difficult, since these factors can probably be easily overcome by external factors. Our studies suggest that there are indeed potential functional variants in chromosome 5q12.1–q13.3, possibly influencing BTF3 function, acting as protective factors against caries. Future studies will determine the function of BTF3 and NFKB1 in caries susceptibility and focus on the translation of these findings into a novel tool that can more effectively protect the population against this very prevalent disease.
Acknowledgments
We would like to thank the individuals that participated in this study for their support. Sarah E. Vinski revised the text for grammar and style. This research is supported by NIH grants R01-DE018914 and R01-DE016148. Study subjects in Guatemala were recruited through collaboration with Children of the Americas (http://www.childrenoftheamericas.org/).
Footnotes
The authors have declared that no competing interests exist.
Author Contributions Data analysis: T.S., M.E.C., A.R.V. Study design: T.S., F.A.P., J.C.M., E.C.K., F.S., M.C.C., J.M.G., P.C.T., I.M.O., E.E.C., M.L.M., A.R.V. Manuscript writing: T.S., M.E.C., A.R.V. Data collection: F.A.A., J.A.B., G.D.P., E.C.K., P.N.T., A.L., T.C.S.V., A.P., M.Y., J.C.M., J.M.R., C.A.B., A.R.V. DNA/RNA manipulation/genotyping: T.S., K.D., J.B.-R., I.M.F. Jr., E.C.K. Revising and reviewing the paper: T.S., K.D., J.B.-R., I.M.F. Jr., F.A.P., J.A.B., G.D.P., E.C.K., P.N.T., A.L., T.C.S.V., A.P., M.Y., J.C.M., J.M.R., C.A.B., M.E.C., F.S., M.C.C., J.M.G., P.C.T., I.M.O., E.E.C., M.L.M., A.R.V.
WEB Resources The URLs for data presented herein are as follows: dbSNP, http://www.ncbi.nlm.nih.gov/projects/SNP/
Ensembl genome browser, http://www.ensembl.org/index.html
Gene Cards, http://www.genecards.org/
HapMap Project, http://hapmap.ncbi.nlm.nih.gov/
PLINK, http://pngu.mgh.harvard.edu/~purcell/plink/
UCSC Genome Bioinformatics, http://genome.ucsc.edu/
World Health Organisation, http://www.who.int/oral_health/publications/report03/en/
References
- Abecasis GR, Cookson WO. GOLD – graphical overview of linkage disequilibrium. Bioinformatics. 2000;16:182–183. doi: 10.1093/bioinformatics/16.2.182. [DOI] [PubMed] [Google Scholar]
- Adair PM, Pine CM, Burnside G, Nicoll AD, Gillett A, Anwar S, Broukal Z, Chestnutt IG, Declerck D, Ping FX, Ferro R, Freeman R, Grant-Mills D, Gugushe T, Hunsrisakhun J, Irigoyen-Camacho M, Lo EC, Moola MH, Naidoo S, Nyandindi U, Poulsen VJ, Ramos-Gomez F, Razanamihaja N, Shahid S, Skeie MS, Skur OP, Splieth C, Soo TC, Whelton H, Young DW. Familial and cultural perceptions and beliefs of oral hygiene and dietary practices among ethnically and socio-economically diverse groups. Community Dent Health. 2004;21:102–111. [PubMed] [Google Scholar]
- Barrett JC, Fry B, Maller J, Daly MJ. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics. 2005;21:263–265. doi: 10.1093/bioinformatics/bth457. [DOI] [PubMed] [Google Scholar]
- Brancher JA, Pecharki GD, Doetzer AD, Medeiros KG, Cordeiro CA, Jr, Sotomaior VS, Bauer P, Trevilatto PC. Analysis of polymorphisms in the lactotransferrin gene promoter and dental caries. Int J Dent. 2011;2011:571726. doi: 10.1155/2011/571726. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brandeis M, Rosewell I, Carrington M, Crompton T, Jacobs MA, Kirk J, Gannon J, Hunt T. Cyclin B2-null mice develop normally and are fertile whereas cyclin B1-null mice die in utero. Proc Natl Acad Sci USA. 1998;95:4344–4349. doi: 10.1073/pnas.95.8.4344. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bretz WA, Corby PM, Melo MR, Coelho MQ, Costa SM, Robinson M, Schork NJ, Drewnowski A, Hart TC. Heritability estimates for dental caries and sucrose sweetness preference. Arch Oral Biol. 2006;51:1156–1160. doi: 10.1016/j.archoralbio.2006.06.003. [DOI] [PubMed] [Google Scholar]
- Bretz WA, Corby PM, Schork NJ, Robinson MT, Coelho M, Costa S, Melo Filho MR, Weyant RJ, Hart TC. Longitudinal analysis of heritability for dental caries traits. J Dent Res. 2005;84:1047–1051. doi: 10.1177/154405910508401115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Conry JP, Messer LB, Boraas JC, Aeppli DP, Bouchard TJ., Jr Dental caries and treatment characteristics in human twins reared apart. Arch Oral Biol. 1993;38:937–943. doi: 10.1016/0003-9969(93)90106-v. [DOI] [PubMed] [Google Scholar]
- Deeley K, Letra A, Rose EK, Brandon CA, Resick JM, Marazita ML, Vieira AR. Possible association of amelogenin to high caries experience in a Guatemalan-Mayan population. Caries Res. 2008;42:8–13. doi: 10.1159/000111744. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dickson SP, Wang K, Krantz I, Hakonarson H, Goldstein DB. Rare variants create synthetic genome-wide associations. PLoS Biol. 2010;8:e1000294. doi: 10.1371/journal.pbio.1000294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao XL, Hsu CY, Xu Y, Hwarng HB, Loh T, Koh D. Building caries risk assessment models for children. J Dent Res. 2010;89:637–643. doi: 10.1177/0022034510364489. [DOI] [PubMed] [Google Scholar]
- Gascoyne DM, Long E, Veiga-Fernandes H, de Boer J, Williams O, Seddon B, Coles M, Kloussis D, Brady HJ. The basic leucine zipper transcription factor E4BP4 is essential for natural killer cell development. Nat Immunol. 2009;10:1118–1124. doi: 10.1038/ni.1787. [DOI] [PubMed] [Google Scholar]
- Horvath S, Xu X, Laird NM. The family based association test method: strategies for studying general genotype-phenotype associations. Eur J Hum Genet. 2001;9:301–306. doi: 10.1038/sj.ejhg.5200625. [DOI] [PubMed] [Google Scholar]
- Kusumawidjaja G, Kayed H, Giese N, Bauer A, Erkan M, Giese T, Hoheise JD, Friess H, Kleeff J. Basic transcription factor 3 (BTF3) regulates transcription of tumor-associated genes in pancreatic cancer cells. Cancer Biol Ther. 2007;6:367–376. doi: 10.4161/cbt.6.3.3704. [DOI] [PubMed] [Google Scholar]
- Lin B, White JT, Ferguson C, Bumgarner R, Friedman C, Trask B, Ellis W, Lange P, Hood L, Nelson PS. PART-1: a novel human prostatespecific, androgen-regulated gene that maps to chromosome 5q12. Cancer Res. 2000;60:858–863. [PubMed] [Google Scholar]
- Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001;25:402–408. doi: 10.1006/meth.2001.1262. [DOI] [PubMed] [Google Scholar]
- Nariyama M, Shimizu K, Uematsu T, Maeda T. Identification of chromosomes associated with dental caries susceptibility using quantitative trait locus analysis in mice. Caries Res. 2004;38:79–84. doi: 10.1159/000075929. [DOI] [PubMed] [Google Scholar]
- Nyholt DR. A simple correction for multiple testing for single-nucleotide polymorphisms in linkage disequilibrium with each other. Am J Hum Genet. 2004;74:765–769. doi: 10.1086/383251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- ørom UA, Shiekhattar R. Long non-coding RNAs and enhancers. Curr Opin Genet Dev. 2011;21:194–198. doi: 10.1016/j.gde.2011.01.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patir A, Seymen F, Yildirim M, Deeley K, Cooper ME, Marazita ML, Vieira AR. Enamel formation genes are associated with high caries experience in Turkish children. Caries Res. 2008;42:394–400. doi: 10.1159/000154785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Powell LV. Caries prediction: a review of the literature. Community Dent Oral Epidemiol. 1998;26:361–371. doi: 10.1111/j.1600-0528.1998.tb01974.x. [DOI] [PubMed] [Google Scholar]
- Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, Maller J, Sklar P, de Bakker PI, Daly MJ, Sham PC. PLINK: a toolset for whole-genome association and population-based linkage analysis. Am J Hum Genet. 2007;81:559–575. doi: 10.1086/519795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothman PB. The transcriptional regulator NFIL3 controls IgE production. Trans Am Clin Climatol Assoc. 2010;121:1156–171. [PMC free article] [PubMed] [Google Scholar]
- Sha WC, Liou HC, Tuomanen EI, Baltimore D. Targeted disruption of the p50 subunit of NF-kappa B leads to multifocal defects in immune responses. Cell. 1995;80:321–330. doi: 10.1016/0092-8674(95)90415-8. [DOI] [PubMed] [Google Scholar]
- Suzuki N, Kurihara Y. Dental caries susceptibility in mice is closely linked to the H-2 region on chromosome 17. Caries Res. 1998;32:262–265. doi: 10.1159/000016462. [DOI] [PubMed] [Google Scholar]
- Tannure PN, Küchler EC, Lips A, Costa MC, Luiz RR, Granjeiro JM, Vieira AR. Genetic variation in MMP20 contributes to higher caries experience. J Dent. 2012;40:381–386. doi: 10.1016/j.jdent.2012.01.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vieira AR, Marazita ML, Goldstein-McHenry T. Genome-wide scan finds suggestive caries loci. J Dent Res. 2008;87:435–439. doi: 10.1177/154405910808700506. [DOI] [PubMed] [Google Scholar]
- Wang X, Shaffer JR, Weyant RJ, Cuenco KT, De-Sensi RS, Crout R, McNeil DW, Marazita ML. Genes and their effects on dental caries may differ between primary and permanent dentitions. Caries Res. 2010;44:277–284. doi: 10.1159/000314676. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Werneck RI, Lázaro FP, Cobat A, Grant AV, Xavier MB, Abel L, Alcaïs A, Trevilatto PC, Mira MT. A major gene effect controls resistance to caries. J Dent Res. 2011;90:735–739. doi: 10.1177/0022034510397614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- World Health Organization The world oral health report 2003. 2003.