Abstract
Disruption of cell polarity is seen in many cancers; however, it is generally considered a late event in tumor progression. Lethal giant larvae (Lgl) has been implicated in maintenance of cell polarity in Drosophila and cultured mammalian cells. We now show that loss of Lgl1 in mice results in formation of neuroepithelial rosette-like structures, similar to the neuroblastic rosettes in human primitive neuroectodermal tumors. The newborn Lgl1–/– pups develop severe hydrocephalus and die neonatally. A large proportion of Lgl1–/– neural progenitor cells fail to exit the cell cycle and differentiate, and, instead, continue to proliferate and die by apoptosis. Dividing Lgl1–/– cells are unable to asymmetrically localize the Notch inhibitor Numb, and the resulting failure of asymmetric cell divisions may be responsible for the hyperproliferation and the lack of differentiation. These results reveal a critical role for mammalian Lgl1 in regulating of proliferation, differentiation, and tissue organization and demonstrate a potential causative role of disruption of cell polarity in neoplastic transformation of neuroepithelial cells.
Keywords: Cancer, cell polarity, neurogenesis, proliferation, cell–cell adhesion, lethal giant larvae
Cell polarity is severely compromised in a variety of human cancers, and loss of cell polarity remains a critical parameter in the assessment of tumor grade. Although cell polarity is compromised in many solid tumors, it is not known whether loss of polarity is a consequence of uncontrolled proliferation or a causative factor in cancer initiation. Most knowledge about cell polarity mechanisms has been generated in studies using simple, genetically tractable organisms including Drosophila, Caenorhabditis elegans, and yeast. Interestingly, genes responsible for cell polarity also regulate cell proliferation. For example, mutation of the Drosophila protein Lethal (2) giant larvae (D-Lgl) causes loss of apical–basal polarity, loss of epithelial organization, and uncontrolled proliferation (Baek 1999; Bilder 2001; Humbert et al. 2003; Justice and Jan 2003). D-Lgl is in the same genetic pathway as Discs-large (Dlg) and Scribble, which are necessary for establishing and maintaining the basolateral membrane domain and basal protein targeting (Bilder and Perrimon 2000; Ohshiro et al. 2000; Peng et al. 2000; Albertson and Doe 2003; Betschinger et al. 2003; Tanentzapf and Tepass 2003), and functions competitively with Crumbs/stardust and Par-3/Par-6/atypical PKC (aPKC) protein complexes that are necessary for the apical membrane domain (Bilder and Perrimon 2000; Betschinger et al. 2003; Tanentzapf and Tepass 2003). Drosophila Lgl acts together with Numb to regulate the Notch signaling pathway (Justice et al. 2003; Klein 2003), and a genetic screen for genes that cooperate with Ras to promote aggressive cancers in Drosophila identified Scribble, D-Lgl, and Dlg (Pagliarini and Xu 2003).
Mammals have two Lgl homologs, Lgl1 (Mlgl, Hugl) and Lgl2 (NM_008502.1 and NM_145438.1, respectively, for mouse sequences). Lgl1 and Lgl2 directly interact with the Par6/aPKC protein complex and can be phosphorylated by aPKC at a conserved phosphorylation site (Musch et al. 2002; Plant et al. 2003; Yamanaka et al. 2003). Overexpression of a mutant, nonphosphorylatable Lgl in MDCK cells had no detectable phenotype (Musch et al. 2002); however, its expression in mouse embryonic fibroblasts reduced cell polarization in an in vitro wounding assay (Plant et al. 2003), and expression of this mutant in Drosophila neuroblasts inhibited asymmetric localization of Miranda (Betschinger et al. 2003). Reduced aPKC levels in Drosophila suppress most of the Lgl-mediated cell polarity and overproliferation phenotypes (Rolls et al. 2003). The results of these studies suggest that the major function of aPKC is to suppress Lgl activity.
In this study, we decided to determine the functional role of mammalian Lgl homologs in vivo. For this purpose, we have generated and analyzed Lgl1 knockout mice. Lgl1–/– embryos demonstrate a critical role for Lgl1 in the developing brain. Lgl–/– neural progenitor cells display loss of cell polarity, failure of asymmetric localization of Numb, and perturbation of Notch signaling. Loss of polarity and subsequent loss of asymmetric cell divisions of neural progenitor cells cause defects in cell cycle exit and differentiation, resulting in an increased number of proliferating cells that form neuroblastic rosette-like structures resembling primitive neuroectodermal tumors.
Results
Lgl1 and Lgl2 expression pattern in embryos and adult mice
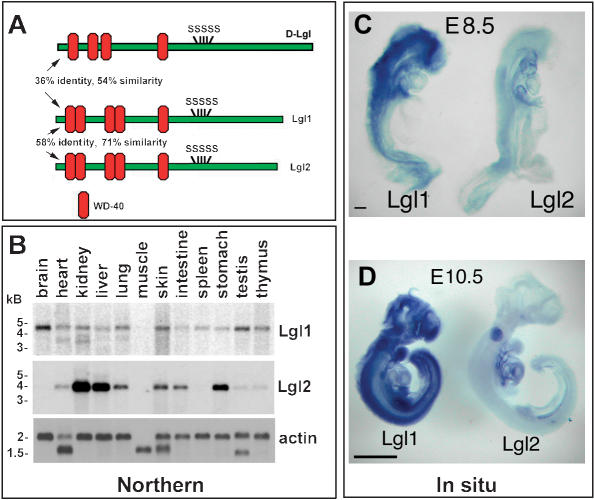
To determine the expression of Lgl transcripts in different organs of adult mice, we performed Northern blot hybridization with Lgl1 and Lgl2 probes (Fig. 1B). The Lgl1 transcripts were broadly expressed, with the highest level of expression in the brain. In contrast, the Lgl2 mRNA was more restricted and was very abundant in kidney, liver, and stomach.
Figure 1.
Mouse Lgl homologs and their expression patterns. (A) Schematic diagram of Drosophila and mouse Lgl proteins. (SSSSS) Conserved phosphorylation sites; (WD-40) conserved WD-40 protein domain used in protein–protein interactions. (B) Northern blot analysis of Lgl1 and Lgl2 expression in adult mouse organs. (C) Whole-mount in situ analysis of Lgl1 and Lgl2 expression in E8.5 and E10.5 embryos. Bars in C represent 1 mm for E10.5 and 0.1 mm for E8.5 embryos.
To determine the expression pattern of Lgl in the developing embryos, we performed whole-mount in situ hybridizations with Lgl1 and Lgl2 probes on embryonic day 8.5 (E8.5) and E10.5 embryos (Fig. 1C,D). At both time points of embryonic development, Lgl1 transcripts were abundantly expressed in most tissues, with the highest levels in the developing brain and spinal cord. The Lgl2 transcripts were detectable at low levels in all organs at E8.5. However, by E10.5 of embryonic development, they were largely restricted to the developing posterior somites, heart, and other internal organs.
We conclude that both Lgl1 and Lgl2 transcripts show a complex pattern of expression in developing embryos and adult animals, with Lgl1 message being almost ubiquitously expressed and Lgl2 expressed in a tissue-specific manner.
Generation of Lgl1–/– mice
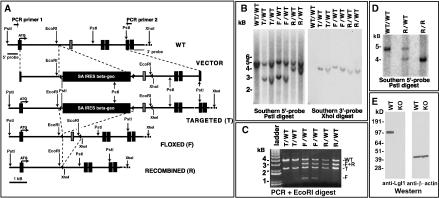
To reveal the in vivo role of Lgl1, we generated Lgl1 knockout mice (Fig. 2). The Lgl1 gene was modified in two independent mouse embryonic stem cell clones using a standard homologs recombination approach. The exon immediately downstream from the exon with the first ATG codon was flanked by LoxP sequences (floxed), and the β-geo selectable marker was deleted by transient transfection of the Cre recombinase gene (Fig. 2A). This modification effectively generated a conditional Lgl1 allele that can be converted to the null allele when exposed to Cre recombinase. Cre-mediated deletion of the floxed exon creates a frame shift in the translation of the protein, and only the first 27 amino acids of Lgl1, lacking known functional domains, will be produced. ES technology was used to generate the Lgl1 conditional mice.
Figure 2.
Generation of Lgl1 knockout mice. (A) Schematic representations of the wild-type Lgl1 allele (WT), the targeting vector (VECTOR), the desired targeted allele (T), the desired conditional flanked by LoxP sequences (F), and the null recombined allele (R) after Cre-mediated recombination of the conditional allele. Triangles indicate LoxP sequences; black boxes indicate the exons of the Lgl1 gene and the promoterless selectable marker SA IRES β-geo; the gray box indicates the 98-bp-long exon of the Lgl1 gene selected for conditional removal. The 5′ and 3′ Southern probes corresponding to sequences flanking the targeting vector arms are denoted by gray bars. PCR primers flanking the targeting vector are denoted by small opposing arrows. (B) Southern analysis of DNA extracted from targeted, floxed, and recombined ES clones. WT/WT, T/WT, F/WT, and R/WT denote the recombination status of the Lgl1 locus. Note that the wild-type fragment recognized by the 3′-probe is not detected, because it is too big for efficient transfer to the membrane. (C) PCR and restriction digest analysis of DNA isolated from the ES cell clones containing targeted, floxed, and recombined Lgl1 alleles. T/WT, F/WT, and R/WT denote the recombination status of the Lgl1 locus. Bands resulting from the wild-type (WT), targeted (T), floxed (F), and recombined (R) alleles are indicated.(D) Southern blot analysis of tail DNA from wild-type (WT), heterozygous (R/WT), and Lgl1 knockout (R/R) mice. DNA was digested with PstI and hybridized with a 5′-probe outside the targeting vector. (E) Western blot analysis of total protein extracts from the wild-type (WT) and Lgl1-knockout (KO) mice with anti-Lgl1 and anti-β-actin antibodies. Numbers on the left indicate DNA and protein standards.
To obtain the Lgl1-null allele, the Lgl1-conditional animals were bred with MORE mice expressing Cre in epiblast cells (Tallquist and Soriano 2000). Cre deleted the floxed exon and the resulting Lgl+/–/Cre+ animals were bred to C57BL/6J mice to generate the Lgl+/–/Cre– animals that were crossed to each other to obtain the Lgl–/– mice. As predicted, the wild-type (WT) allele of Lgl1 disappeared in the Lgl–/– animals (Fig. 2D). Western blot analysis with antibodies recognizing the Lgl1 protein demonstrated the absence of the protein in the Lgl–/– mice, confirming the generation of Lgl1 knockout mice (Fig. 2E).
To determine whether some features of the Lgl–/– phenotype (see below) are caused by the loss of Lgl1 in the extraembryonic tissues, rather than in the embryo itself, we have generated and analyzed mice homozygous for the Lgl1-floxed allele, that are also positive for MORE-Cre. The Cre-mediated recombination and deletion of Lgl1 does not occur in the extraembryonic tissues in these animals. Nevertheless, the phenotype of the Lgl–/– embryos obtained using this approach (data not shown) was indistinguishable from the phenotype of Lgl–/– mice obtained by breeding of animals with the heterozygous Lgl1+/– allele (see below). We conclude that the loss of Lgl1 in the extraembryonic tissues does not contribute to the phenotype of Lgl1–/– embryos.
Histologic abnormalities in the brain of Lgl1–/– mice
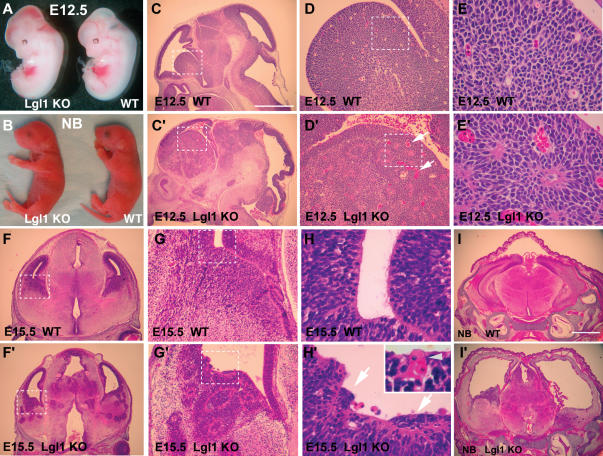
Mice homozygous for the Lgl1-conditional allele or heterozygous for the Lgl1-null allele show no detectable phenotype. In contrast, mice homozygous for the Lgl1-mutant allele are born alive, but die within 24 h after birth. The newborn Lgl1–/– pups appear similar to wild-type and heterozygous animals except for a dome-like shape to their heads (Fig. 3B). All Lgl1–/– embryos show moderate but clearly visible hemorrhage in the ventricles of the brain that can be detected as early as E12.5 (Fig. 3A,B).
Figure 3.
General appearance and histologic analysis of Lgl–/– mice. (A,B) General appearance of Lgl–/– and wild-type E12.5 embryos and newborn pups. Note dome-like appearance of the head of the mutant pup. (C–E′) Histologic appearance of brain from control (C–E) and Lgl–/– (C′–E′) E12.5 embryos. The areas in the dashed squares in C, C′, D, and D′ are shown at higher magnification in D, D′, E and E′, respectively. (F–H′) Histologic appearance of brain from control (F–H) and Lgl–/– (F′–H′) E15.5 embryos. The areas in the dashed squares in F, F′, G, and G′ are shown at higher magnification in G, G′, H, and H′, respectively. The inset in H′ shows a blood vessel that is aberrantly localized to the ventricular wall. The arrows in H′ indicate groups of apolar cells protruding and expanding over the cells that maintain cell polarity. (I,I′) Severe hydrocephalus in the newborn Lgl1–/– mice. Coronal sections through the head region of the wild-type (I) and Lgl1–/– (I′) pups. Note massive dilation of the ventricles and severe damage to the ventricular zone cells in the mutant animal. The bar in C represents 1 mm in C, C′, F, and F′; 0.7 mm in I, I′; 200 μm in D, D′, G, and G′; and 50 μm in E, E′, H, and H′.
Lgl1–/– embryos appear normal until E9.5, but by E12.5 they develop extensive morphologic abnormalities in the brain (Fig. 3C–E′). Overall, there is an expansion of the striatum region (Fig. 3C,C′), where the neuroepithelial cell layer extends into the zones normally occupied by differentiating cells and forms aberrant rosette-like structures. The affected areas are often rich in dilated blood vessels (arrows in Fig. 3D′). The rosettes persist throughout all regions of the brain as development proceeds (Fig. 3F–I′). Rosette-like structures in the brain of Lgl1–/– mice are morphologically similar to the neuroblastic rosettes found in human primitive neuroectodermal tumors (Burger and Scheithauer 1994; Sangueza et al. 1994; Graham and Lantos 2002). High magnification images of the ventricular wall in the wild-type animals reveal polarization of the cells at the ventricle interface. These cells display a prominent eosinophilic apical membrane domain facing the lumen (Fig. 3H). In contrast, the Lgl1–/– brains show groups of cells that have lost polarity and have no visible apical membrane domain (arrows in Fig. 3H′). These groups of cells protrude into the ventricle and extend over the cell layers that maintain an apical membrane domain. Disruption of the normal architecture of the ventricular zone is likely to be responsible for the mislocalization of blood vessels to the ventricle surface causing intraventricular hemorrhage (Fig. 3H′, inset). Although it is possible that the hemorrhage in the Lgl1–/– brain is caused by abnormalities intrinsic to the vasculature, we did not find vascular abnormalities outside of the central nervous system and suggest that the vascular abnormalities might be secondary to the abnormal growth of neuroepithelial cells; however, a conditional disruption of Lgl1 specific to the nervous system will be required to distinguish between these two possibilities.
By E15.5, the ventricles in the Lgl1–/– embryos become dilated (Fig. 3F′), and newborn animals develop severe hydrocephalus (Fig. 3I′). Because of accumulation of cerebrospinal fluid and increasing pressure, the lateral ventricles in the newborn Lgl1–/– animals appear drastically dilated, the ventricular surface of cerebral cortex is destroyed, and the subventricular structures are severely damaged. Intraventricular hemorrhage is the most likely cause of the hydrocephalus in the Lgl–/– mice (Guzzetta et al. 1995), as we found no evidence of ventricular system blockage in the mutants.
Perturbation of differentiation in the Lgl1–/– brain
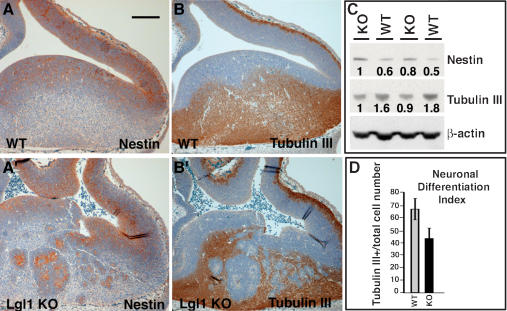
Because Lgl1–/– embryos show expansion of the ventricular zone caused by invagination and rosette formation, it was important to determine the identity of rosette-forming cells using cell-type-specific markers. We concentrated our analysis on the striatum at the ventral side of the lateral ventricle, which is consistently affected in the Lgl1–/– embryos. During neural development, some neural progenitor cells divide asymmetrically, giving rise to one daughter cell that remains a replicating neural progenitor and another daughter cell that terminally differentiates (Lu et al. 2000). Immunostaining with anti-nestin and anti-β-tubulin III antibodies, commonly used neural progenitor and neuronal cell markers, show expansion of the progenitor cell population and reduction of differentiated neuronal cell populations in the knockout brain (Fig. 4A–B′). The neural progenitor cells in the wild-type embryos localize around the ventricles. In contrast, not only the areas around the ventricles, but also the rosette-like structures, are positive for nestin expression in the Lgl1–/– mutants. In addition, rosette structures are negative for β-tubulin III, confirming that cells in the rosettes represent a neural progenitor cell population. Similar staining was obtained with anti-neurofilament antibodies (data not shown). Quantitation of these results shows an overall reduction in the numbers of differentiated neurons in the striatum region of Lgl–/– embryos, suggesting that Lgl1 is necessary for proper regulation of neuronal differentiation (Fig. 4D). Western blot analyses with anti-nestin and anti-tubulin III antibodies demonstrate an overall increase in the levels of nestin and decrease in tubulin III in the Lgl1-null brains (Fig. 4C). We conclude that the rosette-like structures contain neural progenitor cells and that there is an overall increase in the neural progenitor cell domain and decrease in the differentiated neuronal cell domain in the Lgl–/– embryos. It is interesting to note that, although we often observe folding of the Lgl1–/– cortex on tissue sections that occurs during tissue processing and likely reflects overall weakening of the structural integrity of the cortex due to disruption of intercellular adhesion, the cortex does not show the same abnormal nestin or tubulin expression at these stages as is seen in the striatum.
Figure 4.
Changes in neuronal differentiation in Lgl1–/– brains. Expansion of neural progenitor cell domain and reduction of neuronal cell domain in the striatum of Lgl1–/– mice. (A–B′) Immunostaining of sagittal sections from E12.5 wild-type and Lgl1–/– brains with anti-nestin (A,A′) and anti-tubulin III (B,B′) antibodies. The nuclei on the sections were stained by hematoxylin QS. (C) Western blot analysis of total protein extracts from E15.5 wild-type (WT/WT) and Lgl1 knockout (KO/KO) brains with anti-nestin, anti-tubulin III, and anti-β-actin antibodies. (D) Quantitation of experiments shown in C and C′. Neuronal differentiation index determined as a fraction of neurons from the total cell population in the striatum area.
Activation of proliferation, decrease in cell cycle exit, and increase in apoptosis in the Lgl–/– neural progenitor cells
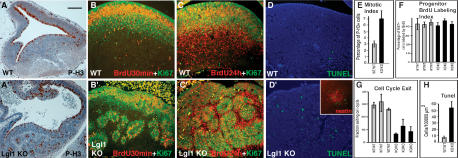
To determine whether loss of Lgl1 leads to changes in cell proliferation, we performed immunostaining with anti-phospho-histone 3 (P-H3) antibodies to identify mitotic cells. As expected, staining of the wild-type embryos with anti-P-H3 antibodies reveals mitotic cells close to the ventricular surface (Fig. 5A). In the Lgl1–/– brains, the P-H3-positive cells are dispersed throughout the brain and localized around the rosette-like structures (Fig. 5A′). Quantitation of the P-H3 staining demonstrates a twofold increase in the number of mitotic cells per total cell number in the striatum of Lgl1–/– mutants (Fig. 5E).
Figure 5.
Activation of proliferation, defects in cell cycle exit, and apoptosis in the Lgl1–/– neural precursors. (A,A′) Immunostaining of sagittal sections from E12.5 wild-type brains with anti-phospho-histone 3 antibodies reveals increase in mitotic cells in the striatum of Lgl1–/– mice. (B,B′) Absence of changes in cell cycle length in the striatum of Lgl–/– animals. The percentage of progenitor cells (Ki67+) labeled with BrdU (BrdU+) after a 30-min pulse is not altered in Lgl–/– animals. (C,C′) Decrease in cell cycle withdrawal in Lgl1–/– brain. Pregnant females were injected with BrdU 24 h before being killed. Sections were stained with anti-BrdU and anti-Ki67 antibodies. Cells that re-enter the cell cycle were labeled with both BrdU and Ki67 antibodies (BrdU+/Ki67+), whereas cells withdrawn from the cell cycle were labeled by BrdU, but not by Ki67 antibodies (BrdU+/Ki67–). (D,D′) Increase in apoptosis in the striatum of Lgl1–/– animals. Mutant brains have more TUNEL-positive (green) cells than the wild-type controls. DNA is counterstained by Hoechst 33342 (blue). The inset in D′ shows prominent localization of TUNEL+ cells (green) to the nestin-positive rosettes (red) in the Lgl1–/– brains. (E) Quantitation of the experiments shown in A and A′. Mitotic index determined as a ratio of mitotic cells (P-H3+) to the total cell number. (F) Quantitation of experiments shown in B and B′. Progenitor BrdU labeling index determined as a percentage of neural progenitors (Ki67+) incorporated BrdU (BrdU+) after 30-min pulse labeling. (G) Quantitation of experiments shown in C and C′. Cell cycle exit is determined as a ratio of the cells that exited the cell cycle (BrdU+/Ki67–) to all cells that incorporated the BrdU (BrdU+) after 24 h labeling. (H) Quantitation of experiments shown in D and D′. The number of TUNEL-positive cells per similar areas of striatum from Lgl1–/– and wild-type mice were counted. The bar in A represents 100 μm for A and A′, and 70 μm for B–D′.
Because we found that E12.5 Lgl1–/– embryos show an expansion of the neural progenitor cell domain and an increase in the number of mitotic cells, it was important to determine whether this phenotype is caused by the shortening of the neural progenitor cell cycle, or is the result of defective cell cycle withdrawal and differentiation. To reveal potential changes in the cell cycle, we counted the proportion of neural progenitor cells labeled by a 30-min pulse of BrdU(Fig. 5B,B′). The neural progenitor cells in this experiment were identified by Ki67 staining (Hartfuss et al. 2001). Because the length of the S phase in mammalian cells remains constant (DiSalvo et al. 1995), the ratio between the BrdU-positive and Ki67-positive cells provides an estimation of the cell cycle length, and a larger proportion of BrdU-positive neural progenitor cells would indicate shortening of the cell cycle. Quantitation of these experiments indicates no change in the length of the cell cycle in the Lgl1–/– neural progenitor cells (n = 3; Fig. 5F).
To determine whether loss of Lgl1 causes changes in cell cycle withdrawal, we counted the proportion of cells that exit the cell cycle after 24 h labeling with BrdU(Fig. 5C,C′). The cell cycle exit was estimated as a ratio between the BrdU-positive/Ki67-negative cells and all BrdU-positive cells. Quantitation of these data shows a twofold decrease in the number of cells that withdraw from the cell cycle in the Lgl1–/– animals (n = 3; Fig. 5G). This indicates that Lgl1–/– neural progenitor cells are less likely than wild-type cells to perform an asymmetric cell division and withdraw from the cell cycle. This can explain the expansion of the neural progenitor cell domain in Lgl1–/– animals.
To determine whether loss of Lgl1 leads to changes in apoptosis, we performed TUNEL staining (Fig. 5D,D′). Counting TUNEL-positive cells reveals an eightfold increase in apoptosis in the Lgl1–/– embryos (Fig. 5H). The apoptotic cells in the mutants were localized to the neural progenitor cell domain and were abundantly present in the nestin-positive, rosette-like structures (Fig. 5D′, inset). Therefore, loss of Lgl1 causes not only expansion of proliferating progenitor cells, but also their apoptotic cell death.
Loss of cell polarity in the Lgl1–/– neural progenitor cells
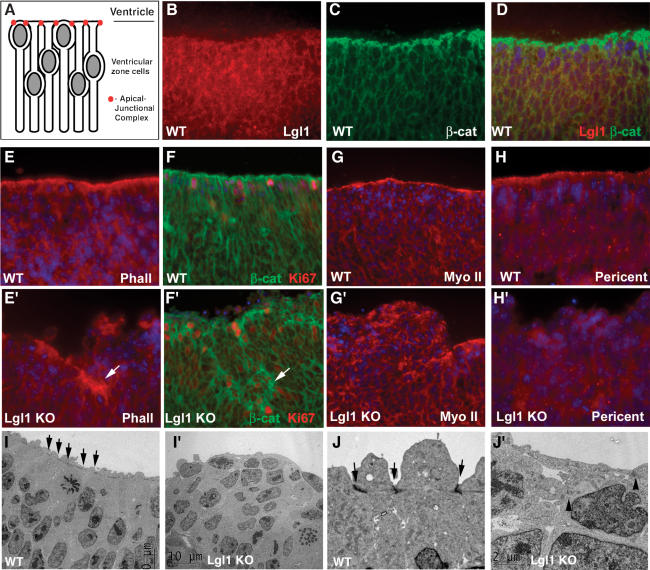
Although regulation of differentiation in neuroepithelia is only beginning to be understood, it is clear that asymmetric cell division and cell polarity play a pivotal role in this process (Shen and Temple 2002; Fishell and Kriegstein 2003). Neural progenitor cells are bipolar, with one cell extension reaching the ventricular surface and another process reaching in the opposite direction (Fig. 6A; Chenn et al. 1998). The apical extensions of neural progenitor cells are connected to each other by adherens junctions, which are especially prominent at the ventricular interface. We decided to determine whether Lgl1 is necessary for maintenance of polarity of the neuroepithelial cells. First we performed staining with anti-Lgl1 antibodies and found that Lgl1 was localized to the cytoplasm of neuroepithelial cells (Fig. 6B–D). As previously described, β-catenin, f-actin, and myosin II-B localize predominantly to the apical–junctional complex in normal neuroepithelium (Fig. 6E–G); whereas the distribution of these proteins shows marked disorganization in the Lgl1–/– neuroepithelium (Fig. 6E′–G′). Similar loss of cell polarity in the Lgl1–/– cells was seen after staining for pericentrin, a marker of centrioles located at the center of the apical extension (Chenn et al. 1998), and shroom, which localizes to the apical cell membrane (Fig. 6H,H′; Hildebrand and Soriano 1999; data not shown). Small fragments of neuroepithelium at the ventricular surface are still capable of maintaining cell polarity, but they are often engulfed by protruding nonpolarized cells that fold back on themselves and become internalized to form rosette-like structures (Fig. 6E′–G′). The loss of cell polarity was especially prominent in the striatum area, whereas the areas showing few rosette-like structures (e.g., cortex) were less affected. We conclude that Lgl1 is necessary for maintaining the apical–junctional complex and overall cell polarity. Engulfment and internalization of small fragments of properly polarized neuroepithelial cells by their apolar neighbors may represent the mechanism of rosette-like structure formation in the Lgl1-null brain.
Figure 6.
Loss of cell polarity and disruption of apical–junctional complex in Lgl1–/– neuroepithelial cells. (A) Model showing intrinsic polarity of neuroepithelial cells maintained by apical–junctional complex of cell–cell adhesion proteins. (B–D) Lgl1 localizes to cytoplasm, but not to apical–junctional complex of neuroepithelial cells. Sections from striatum of E12.5 wild-type mice were stained with anti-Lgl1 (B), anti-β-catenin (C) antibodies. Double staining is shown in D. (E–H′) Lgl1 is required for polarized localization of f-actin (E,E′), β-catenin (green in F,F′; Ki67 staining is in red), myosin IIb (G,G′), and pericentrin (H,H′). Arrows indicate fragments of neuroepithelia in Lgl1–/– brain that maintained cell polarity, which appear to be engulfed by apolar neighbors. (I–J′) Disruption of apical–junctional complex in Lgl1–/– neuroepithelial cells. Striatum areas from wild-type (I,J) and Lgl1–/– (I′,J′) littermates were analyzed by electron microscopy. Arrows in I and J indicate apical–junctional complex. Residual apical–junctional complex in Lgl1–/– cells indicated with arrowheads (J′). Blue staining in D–H′ indicates cell nuclei detected by DAPI. The bar in J′ corresponds to 26 μm in G and G′; 18 μm in B–F′, H, and H′; and 2 μm in J and J′. The bar in I′ corresponds to 10 μm in I and I′.
To understand the nature of the cell–cell adhesion defect in the Lgl1–/– embryos, we used electron microscopy to examine the neuroepithelium of E10.5 mutants, immediately before the formation of rosette-like structures. We found that the apical–junctional complex, prominently visible in the wild-type cells (arrows in Fig. 6I,I′), is disrupted in the mutants (Fig. 6J,J′). Pseudostratified neuroepithelium in the wild-type embryos is replaced by stratified neuroepithelium in the knockout. Therefore, Lgl1 is required for maintenance of the apical–junctional complex in neural progenitor cells.
Loss of polarized localization of Numb and expansion of a cell domain with active Notch signaling in the Lgl1–/– brain
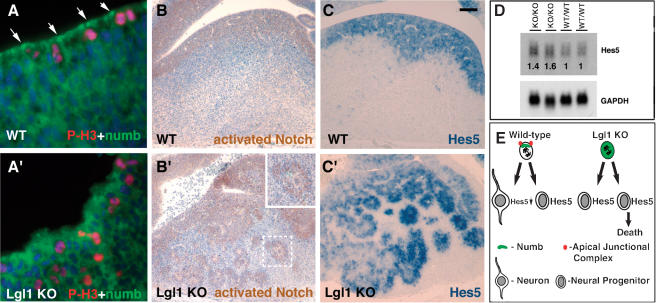
Asymmetric cell division and the asymmetric localization of Numb, an antagonist of Notch signaling, play a pivotal role in the regulation of cell differentiation in neuroepithelia (Lu et al. 2000; Jafar-Nejad et al. 2002; Shen and Temple 2002; Shen et al. 2002; Fishell and Kriegstein 2003). To determine whether Numb is properly localized in Lgl1–/– neuroepithelial cells, we performed staining with anti-Numb antibodies. As expected, in dividing wild-type cells in vivo, Numb is localized to the apical cell membrane (Fig. 7A; Zhong et al. 1996). This asymmetric Numb localization was not observed in the Lgl1–/– cells (Fig. 7A′). As judged by Western blot analysis, the overall levels of Numb protein in the Lgl1–/– mutants were unchanged (data not shown). We conclude that in mouse neuroepithelia, Lgl1 is required to maintain asymmetric localization of Numb.
Figure 7.
Lgl1 is required for polarized localization of Numb and regulation of Notch signaling. (A,A′) Loss of asymmetric localization of Numb in dividing Lgl–/– progenitor cells. Double immunostaining of striatum regions of brain from the wild-type (WT) and Lgl–/– (Lgl1 KO) E10.5 mice with anti-Numb (green) and anti-phospho-histone 3 (red) antibodies. Blue indicates cell nuclei detected by DAPI. (B,B′) Expansion of the cell domain with active Notch signaling in the Lgl1-null brain. Immunohistochemical staining of striatum regions of the E12.5 brains with antibodies specifically recognizing cleaved (activated) Notch 1. The area containing the rosette-like structure shown in the dashed square in B′ is magnified in the inset. (C,C′) Expansion of Hes5-positive cell domain in the Lgl–/– brain. Sections from E12.5 wild-type and Lgl1–/– embryos were analyzed by in situ hybridization with the Hes5 probe. (D) Increase in levels of Hes5 transcripts in the brains of Lgl–/– animals. Total RNA was extracted from two E15.5 wild-type (WT/WT) and Lgl–/– (KO/KO) animals and analyzed by Northern blotting with Hes5 and GAPDH probes. The data were quantitated using PhosphorImager. The numbers indicate the relative amounts of Hes5 adjusted by the levels of GAPDH. (E) Schematic representation of brain phenotype in Lgl1 mutants. In Lgl1–/– neural progenitor cells where cell polarity is not maintained, Numb protein is not asymmetrically localized. Loss of asymmetric cell divisions causes expansion of identical neural progenitor cells, some of which undergo apoptosis. The bar in C represents 100 μm in B–C′, and 12 μm in A and A′.
Asymmetric Numb localization during cell division is responsible for differences in Notch activity between daughter cells that result in different cell fate decisions (Lu et al. 2000; Jafar-Nejad et al. 2002; Shen and Temple 2002; Shen et al. 2002; Fishell and Kriegstein 2003). To determine whether loss of Lgl1 influences Notch signaling, we performed immunohistochemical staining with antibodies specifically recognizing the cleaved (activated) Notch (Fig. 7B,B′). In addition, we performed Northern blot and in situ hybridizations with probes to Hes5 (Fig. 7C–D), a gene regulated by Notch. We found that levels of Hes5 mRNA are increased in the brain of Lgl1–/– embryos (Fig. 7D). Immunostaining with antiactivated Notch and in situ hybridization with the Hes5 probe revealed an increase in the number of positive cells, indicating that the increase of active Notch and Hes5 mRNA is caused by an expansion of the cell domain with active Notch signaling (Fig. 7B–C′). We conclude that Lgl1 regulates Notch signaling and subsequent cell fate decisions possibly through the asymmetric inheritance of Numb protein in dividing neural progenitor cells.
Discussion
Lgl1 in neurogenesis
During mammalian brain development, seemingly indistinguishable precursor cells undergo multiple division-linked cell fate decisions to produce an extraordinary complexity of interconnected neurons and glia (Zhong 2003). Whereas early in development (up to E10.5) there is an apparently homogeneous expansion of the precursor cell population, later some of the dividing neural progenitor cells undergo an asymmetric cell division and give rise to committed progeny (Fishell and Kriegstein 2003). The Notch signaling pathway is used for cell fate determination throughout the animal kingdom, and differences in Notch activity between two daughters determine their future fates. Notch can be used multiple times in the same cell lineage, and the outcome of Notch signaling depends on the cellular context. In neural development, Notch signaling appears to inhibit neuronal differentiation and promote the radial glia cell fate (Fishell and Kriegstein 2003; Justice and Jan 2003). During neurogenesis, differences in the Notch activities between two daughter cells may be specified by the asymmetric localization and subsequent inheritance of Numb, a negative regulator of the Notch pathway (Guo et al. 1996; Petersen et al. 2002; Shen et al. 2002). Numb may play multiple functions in developing mammalian brain. During early neurogenesis (E8.5), Numb and Numblike are necessary for progenitor cell maintenance (Petersen et al. 2002). In contrast, the cortex-specific inactivation of Numb and Numblike during later phase of neurogenesis (E12.5–E16.5; Li et al. 2003) leads to a phenotype that is very similar to the Lgl1-null brain: neural progenitor hyperproliferation, delayed cell cycle exit, impaired neuronal differentiation, and defects in cortical morphogenesis.
We have found that Lgl1 is critically involved in regulating asymmetric cell divisions in the developing mammalian brain. Many Lgl1–/– neural progenitors fail to differentiate and, instead, continue to proliferate. This results in an expansion of the neural progenitor cell domain and a decreased domain of differentiated neurons. We found that Lgl1 is required for the localization of Numb during asymmetric divisions of neural progenitor cells. Striking similarities between the Lgl1 knock out brains and the Numb/Numblike cortex-specific knockout (Li et al. 2003) confirms our conclusion that the principal function of mammalian Lgl1 is regulation of asymmetric Numb localization during divisions of neural progenitor cells. The loss of the difference in Numb levels between two daughter cells may be responsible for the failure of Lgl1–/– cells to down-regulate the Notch pathway and execute the neuronal differentiation program. This may explain defective neuronal differentiation in the Lgl1–/– brain. Although it is not known how Lgl1 is involved in the regulation of Numb localization, the role of Lgl1 in the maintenance of cell polarity may be important in this process.
Lgl1 in cell polarity
Most mammalian cells are highly polarized, yet very little is known about the mechanisms responsible for cell polarity in higher eukaryotes. It is not known what might be the consequences of disruption of polarity and whether this plays an important role in cancer development or progression. In this study we determined that Lgl1, the mammalian homolog of Drosophila cell polarity gene D-Lgl, is necessary for the maintenance of cell polarity in developing mammalian brain. Mammalian Lgl proteins have been previously implicated in cell polarity using cells in culture. Expression of the mutant unphosphorylatable Lgl1 (Mlgl) in mouse embryonic fibroblasts decreased their ability to polarize in response to scratch wound in culture (Plant et al. 2003). Overexpression of the wild-type Lgl in the MDCK cells decreased their ability to form epithelial junctions (Yamanaka et al. 2003). However, in a different study the overexpression of the wild-type or mutant unphosphorylatable Lgl in MDCK cells did not produce any detectable cell polarity or cell–cell adhesion phenotypes (Musch et al. 2002). Using a gene-targeting approach, we demonstrate that Lgl1 is required for maintenance of cell polarity in vivo. Interestingly, the brain is the only organ that appears to be affected in the Lgl–/– animals. In addition, although all regions of the brain can be affected, the phenotype is usually more severe in the basal ganglia, thalamus, and areas around the fourth ventricle, whereas the cortex is less affected. Multiple scenarios can explain this tissue-specificity, but we believe that the presence of Lgl2 in other organs and its low but detectable by RT–PCR (data not shown) expression in the brain may compensate for the loss of Lgl1.
It remains to be discovered how Lgl proteins function to maintain cell polarity. Early studies in Drosophila suggested that Lgl interacts and regulates the localization and, possibly, the activity of nonmuscle myosin II (Strand et al. 1994). D-Lgl shows strong negative genetic interaction with myosin II (Strand 1998), and the yeast homologs of D-Lgl, Sro7, and Sro 77, were shown to form a complex and genetically interact with Myo1p, a type II myosin (Kagami et al. 1998). In addition, human Lgl1 (Hugl) was shown to bind to nonmuscle myosin II (Strand et al. 1995). Myosin II is necessary for asymmetric cell division in Drosophila neuroblasts. It promotes asymmetric localization of Numb by excluding it from the apical cortex, rather than by promoting its polarized transport, and D-lgl is required for proper myosin II localization and function (Barros et al. 2003). Interaction between Lgl and myosin II indicated that Lgl might regulate cell polarity through the actin/myosin cytoskeleton. Indeed, the actin cytoskeleton is perturbed in Lgl-mutant Drosophila and yeast (Kagami et al. 1998; Bilder et al. 2000). Recently, the alternative view has been developed. Yeast Sro7 and Sro77 protein were shown to interact with the plasma membrane soluble N-ethylmaleimide-sensitive factor attachment receptor (SNARE), Sec9 (Lehman et al. 1999). Defects in the actin cytoskeleton were not observed in the Sro7/77 mutants in this study; instead, yeast exhibited a defect in exocytosis. The Sro7/77 mutant yeast were filled with exocytic vesicles and found to be defective in fusion of post-Golgi vesicles with the plasma membrane. Later, the mammalian Lgl1 (Mlgl) was found to bind to syntaxin 4, the basolateral t-SNARE protein (Musch et al. 2002). These data suggested that Lgl functions in the maintenance of cell polarity by regulating polarized exocytosis.
Our Lgl1–/– animals exhibit a drastic perturbation in the organization of the actin cytoskeleton and localization of myosin II-B (Fig. 6E′,G′). The wild-type neuroepithelial cells display prominent cables of the actin/myosin filaments contacting and reinforcing the cytoplasmic side of the apical–junctional complex. EM and the immunofluorescence show disruption of the apical–junctional complexes in the Lgl1–/– neural progenitor cells (Fig. 6J′). Unlike the studies with Sro7/77 mutants in yeast, we did not observe accumulation of vesicles in the Lgl1–/– neural progenitor cells using EM microscopy (data not shown). The cell polarity defects in the Lgl1–/– brains are similar to the defects seen in the myosin II-B mutant mice (Tullio et al. 2001). These data suggest that, similar to Drosophila neuroblasts, interaction between Lgl1 and myosin II may be an important part of the Lgl function in the maintenance of cell polarity in the mouse brain. Lgl function in the regulation of the actin cytoskeleton and polarized exocytosis are not necessarily exclusive, and may both be used for the maintenance of cell polarity. Interestingly, nonmuscle myosin II-B might be directly involved in exocytosis-dependent membrane trafficking (Togo and Steinhardt 2004). Localization of the apical–junctional complex in neural progenitor cells is highly polarized, and targeted delivery of proteins involved in the formation of this cell–cell adhesion structure should be important for the maintenance of cell polarity. Interestingly, we found that the levels of nonionic detergent-soluble N-cadherin are increased in the Lgl1–/– brains (data not shown). This may indicate a failure to deliver N-cadherin to the apical–junctional complex; however, it could also be a failure to stabilize N-cadherin at the membrane caused by a dysfunctional actin cytoskeleton.
Lgl1 in the regulation of cell proliferation and cancer
Development and maintenance of all tissues in mammals requires a series of cell fate decisions that are accomplished by asymmetric inheritance of molecules involved in cell fate determination. Stem cells, as well as transiently specified progenitor cells, can divide asymmetrically to generate one daughter cell that withdraws from the cell cycle and terminally differentiates. We demonstrate that Lgl1 is necessary for the regulation of asymmetric cell division and cell fate determination in the developing mammalian brain. Lgl1 may exercise this function through control of Numb localization. The loss of Lgl1 causes an expansion of neural progenitor cells and an overall increase in cell proliferation. Interestingly, mutations in Lgl in Drosophila also cause uncontrolled proliferation (Bilder 2001). The imaginal discs in the mutant flies lose polarity, overgrow, and kill the developing larvae. Owing to this phenotype, Lgl, as well as Dlg and Scribble proteins that function in the same pathway, were classified as Drosophila tumor-suppressor proteins. The mechanisms responsible for the function of Lgl in the negative regulation of cell proliferation remain largely unknown. It is believed that mutations in Lgl lead to disruption of overall tissue architecture, which, in turn, cause loss of proliferation control (Bilder et al. 2000). Multiple proteins are involved in the regulation of cell adhesion, and loss of many different proteins can lead to the disruption of tissue architecture. Nevertheless, Lgl, Scribble, and Dlg are unique in their ability to suppress tumor growth and invasion in flies (Pagliarini and Xu 2003). It is remarkable that deletion of Lgl1 in mice causes such a dramatic phenotype that involves not only activation of proliferation and paucity of differentiation, but also massive tissue dysplasia and invasion. The role of Lgl in the regulation of Numb localization and asymmetric cell division demonstrated here may provide a mechanism for the increase in cell proliferation and tissue disorganization in the Lgl mutants. It is also possible that Lgl1 has additional tumor-suppressor functions that remain to be discovered.
Whereas Lgl is a tumor-suppressor protein in flies, it remains to be determined whether it is involved in carcinogenesis in higher eukaryotes. We found that ablation of Lgl1 causes multiple changes in the developing brain that histologically resemble brain cancer. The Lgl1 mutant brain displays neuroblastic rosettes characteristic of a subset of human primitive neuroectodermal tumors (Burger and Scheithauer 1994; Sangueza et al. 1994; Graham and Lantos 2002). The Lgl–/– neural progenitors display hyperproliferation and lack of differentiation. Human Lgl1 localizes to Chromosome 17p11.2. This region is affected in 50% of medulloblastomas; however, the gene involved in medulloblastoma that resides at this locus is not known (Fruhwald et al. 2001). Our data strongly suggest that Lgl1 may be a gene affected in primary medulloblastomas and loss or inactivation of Lgl1 may be involved in medulloblastoma formation.
Whereas similarities between human brain cancers and Lgl1–/– mutant brains are intriguing, it appears to be unlikely that deletion of one gene alone could be sufficient for cancer development. In accordance with this, we found that even though mutation in Lgl1 causes stimulation of cell proliferation, it also results in activation of apoptotic cell death. It is possible that cooperation between Lgl and other oncogenes or tumor-suppressor proteins is necessary for cancer development.
It is possible that Lgl1–/– neural progenitor cells display transformed features only during the active cell proliferation that occurs in neurogenesis, and will exhaust their proliferative potential when neurogenesis is completed. Neonatal lethality of the Lgl1–/– mice precluded us from analyzing this possibility. If this is the case, the Lgl1–/– phenotype may be more similar to severe human brain dysplasia found in periventricular heterotopia, which is characterized by the presence of periventricular lesions consisted of nodules containing differentiated neurons that are oriented in multiple directions (Walsh 1999).
In summary, our studies identify Lgl1 as a critical protein necessary for asymmetric cell divisions and cell cycle withdrawal in the developing mammalian brain. We demonstrate that disruption of cell polarity, which is often seen in human cancer, has a profound affect on cell proliferation and differentiation, and may have a causative role in the initial stages of some central nervous system neoplasias.
Materials and methods
Mice
Mice with a conditional allele of Lgl1 were generated using ES cell technology as described (Hogan et al. 1994; Vasioukhin et al. 2001). The exon downstream from the exon with the first ATG codon was flanked by LoxP sequences, and the β-geo selectable marker was removed by transient expression of Cre recombinase in the ES cells. The Lgl1-null allele was obtained by breeding conditional Lgl1 animals with MORE-Cre mice expressing Cre in the germ line (Tallquist and Soriano 2000). Heterozygous Lgl+/– animals negative for Cre were bred to each other to obtain the Lgl1 knockout mice.
Histology and electron microscopy
Tissues for histology were fixed in 4% paraformaldehyde, processed, and embedded in paraffin. Sections (5 μM) were stained with hematoxylin and eosin, examined, and photographed using the Nikon TE 200 microscope. For transmission electron microscopy (EM), samples were fixed in 2% glutaraldehyde, 4% formaldehyde in 0.05M sodium cacodylate buffer overnight at 4°C and processed for Epon embedding. Samples were visualized with a JEOL 1010 microscope.
Northern, Southern, and Western blot analysis and in situ hybridization
Northern, Southern, and Western blot analyses were carried out according to standard protocols (Sambrook and Russell 2001). PCR-generated fragments of DNA outside the targeting vector and β-geo-containing fragment were used as probes for Southern blot analysis. Whole-mount, in situ, and Northern blot hybridizations were performed using obtained by RT–PCR 400–500-bp probes from the 3′-untranslated regions of mouse Lgl1, Lgl2, and Hes5 transcripts. For in situ hybridization, digoxygenin probes were synthesized using a kit from Roche. Whole-mount in situ hybridizations were carried out as described (Hogan et al. 1994). The in situ hybridizations with paraffin sections were carried out as described (Fijnvandraat et al. 2002).
Immunofluorescence and immunohistochemistry
For immunofluorescence staining, tissues were first fixed in 4% paraformaldehyde in PBS for 0.5–3 h at 4°C, washed with PBS, embedded in OCT, and then frozen immediately on dry ice. Sections were subjected to indirect immunostaining with antibodies and analyzed using the Nicon TE 200 microscope equipped with a COOLSNAP HQ digital camera or using the Applied Precision, Inc. Deltavision SA3.1 Deconvolution Microscope. In some cases, tissues were first processed, embedded in paraffin, and sectioned; the resulting sections were deparafinized, dehydrated, and processed as described above. The ABC elite or ABC MOM kits (Vector laboratories) were used for immunohistochemistry. Antibodies were detected with a DAB peroxidase substrate kit, and sections were counterstained with hematoxylin QS (both from Vector laboratories).
Antibodies and apoptosis staining
The antibodies used were anti-β-tubulin III (Sigma), anti-neurofilament (Chemicon), anti-nestin (Developmental Studies Hybridoma Bank), anti-Ki67 (Novocastra), anti-Phospho-H3 (Upstate Biotechnology), anti-β-catenin, phalloidin-TRITC (Sigma), anti-pericentrin, anti-myosin II-B (Covance), anti-Numb (from Yuh Nung Jan, Howard Hughes Medical Institute, and Departments of Physiology and Biochemistry, University of California, San Francisco, San Francisco, CA), anti-active Notch 1 (Cell Signaling), and anti-Notch 1 (Santa Cruz). To generate anti-mLgl1 antibodies, rabbits were immunized with peptide (C)EGSPSSAHSKRADTME-COOH from the C terminus of mLgl1. In some experiments, anti-Lgl1 antibodies from Shigeo Ohno (Department of Molecular Biology, Yokohama City Univerity School of Medicine, Kanazawa-ku, Japan) were used (Yamanaka et al. 2003). Relevant FITC- or TxR-conjugated donkey or goat antibodies (Jackson Laboratories) were used for detection of primary antibodies. Apoptosis was determined using the TUNEL staining kit from Oncogene Research.
Quantitation of cell differentiation, mitosis, cell cycle length, cell cycle exit
For quantitation of data obtained using the immunofluorescent and immunohistochemical stainings, the identical areas in the striatum region of brain of the Lgl–/– and littermate wild-type animals were selected, and the total numbers of cells and cells stained positive with indicated antibodies were counted. Analysis of cell cycle length and cell cycle exit using BrdU labeling was performed as described (Chenn and Walsh 2002).
Acknowledgments
We thank Philippe Soriano for MORE-Cre mice, encouragement, and helpful advice; Shigeo Ohno, Yuh Nung Jan, and the Developmental Studies Hybridoma Bank for generous gifts of antibodies; Alex Spence for comments on Lgl1–/– histology; Nanyan Jiang for injections of ES cells; and Linda Cherepow for help with tissue sectioning. This work was supported by NCI grant 1R01CA098161-01.
The publication costs of this article were defrayed in part by payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 USC section 1734 solely to indicate this fact.
Article and publication are at http://www.genesdev.org/cgi/doi/10.1101/gad.1178004.
References
- Albertson R. and Doe, C.Q. 2003. Dlg, Scrib and Lgl regulate neuroblast cell size and mitotic spindle asymmetry. Nat. Cell Biol. 5: 166–170. [DOI] [PubMed] [Google Scholar]
- Baek K.H. 1999. The first oncogene in Drosophila melanogaster. Mutat. Res. 436: 131–136. [DOI] [PubMed] [Google Scholar]
- Barros C.S., Phelps, C.B., and Brand, A.H. 2003. Drosophila nonmuscle myosin II promotes the asymmetric segregation of cell fate determinants by cortical exclusion rather than active transport. Dev. Cell 5: 829–840. [DOI] [PubMed] [Google Scholar]
- Betschinger J., Mechtler, K., and Knoblich, J.A. 2003. The Par complex directs asymmetric cell division by phosphorylating the cytoskeletal protein Lgl. Nature 422: 326–330. [DOI] [PubMed] [Google Scholar]
- Bilder D. 2001. Cell polarity: Squaring the circle. Curr. Biol. 11: R132–R135. [DOI] [PubMed] [Google Scholar]
- Bilder D. and Perrimon, N. 2000. Localization of apical epithelial determinants by the basolateral PDZ protein Scribble. Nature 403: 676–680. [DOI] [PubMed] [Google Scholar]
- Bilder D., Li, M., and Perrimon, N. 2000. Cooperative regulation of cell polarity and growth by Drosophila tumor suppressors. Science 289: 113–116. [DOI] [PubMed] [Google Scholar]
- Burger P.C. and Scheithauer, B.W. 1994. Atlas of tumor pathology. Tumors of the central nervous system. Armed Forces Institute of Pathology, Washington, DC.
- Chenn A. and Walsh, C.A. 2002. Regulation of cerebral cortical size by control of cell cycle exit in neural precursors. Science 297: 365–369. [DOI] [PubMed] [Google Scholar]
- Chenn A., Zhang, Y.A., Chang, B.T., and McConnell, S.K. 1998. Intrinsic polarity of mammalian neuroepithelial cells. Mol. Cell. Neurosci. 11: 183–193. [DOI] [PubMed] [Google Scholar]
- DiSalvo C.V., Zhang, J.W., and Jacobberger, J.W. 1995. Regulation of NIH-3T3 cell G1 phase transit by serum during exponential growth. Cell Prolif. 28: 511–524. [DOI] [PubMed] [Google Scholar]
- Fijnvandraat A.C., De Boer, P.A., Deprez, R.H., and Moorman, A.F. 2002. Non-radioactive in situ detection of mRNA in ES cell-derived cardiomyocytes and in the developing heart. Microsc. Res. Tech. 58: 387–394. [DOI] [PubMed] [Google Scholar]
- Fishell G. and Kriegstein, A.R. 2003. Neurons from radial glia: The consequences of asymmetric inheritance. Curr. Opin. Neurobiol. 13: 34–41. [DOI] [PubMed] [Google Scholar]
- Fruhwald M.C., O'Dorisio, M.S., Dai, Z., Rush, L.J., Krahe, R., Smiraglia, D.J., Pietsch, T., Elsea, S.H., and Plass, C. 2001. Aberrant hypermethylation of the major breakpoint cluster region in 17p11.2 in medulloblastomas but not supratentorial PNETs. Genes Chromosomes Cancer 30: 38–47. [PubMed] [Google Scholar]
- Graham D.I. and Lantos, P.L. 2002. Greenfield's neuropathology. Arnold, New York.
- Guo M., Jan, L.Y., and Jan, Y.N. 1996. Control of daughter cell fates during asymmetric divisions: Interaction of Numb and Notch. Neuron 17: 27–41. [DOI] [PubMed] [Google Scholar]
- Guzzetta F., Mercuri, E., and Spano, M. 1995. Mechanisms and evolution of the brain damage in neonatal post-hemorrhagic hydrocephalus. Childs Nerv. Syst. 11: 293–296. [DOI] [PubMed] [Google Scholar]
- Hartfuss E., Galli, R., Heins, N., and Gotz, M. 2001. Characterization of CNS precursor subtypes and radial glia. Dev. Biol. 229: 15–30. [DOI] [PubMed] [Google Scholar]
- Hildebrand J.D. and Soriano, P. 1999. Shroom, a PDZ domain-containing actin-binding protein, is required for neural tube morphogenesis in mice. Cell 99: 485–497. [DOI] [PubMed] [Google Scholar]
- Hogan B., Beddington, R., Constantini, F., and Lacy, E. 1994. Manipulating the mouse embryo. A laboratory manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
- Humbert P., Russell, S., and Richardson, H. 2003. Dlg, Scribble and Lgl in cell polarity, cell proliferation and cancer. Bioessays 25: 542–553. [DOI] [PubMed] [Google Scholar]
- Jafar-Nejad H., Norga, K., and Bellen, H. 2002. Numb: “Adapting” notch for endocytosis. Dev. Cell 3: 155–156. [DOI] [PubMed] [Google Scholar]
- Justice N.J. and Jan, Y.N. 2003. A lethal giant kinase in cell polarity. Nat. Cell Biol. 5: 273–274. [DOI] [PubMed] [Google Scholar]
- Justice N., Roegiers, F., Jan, L.Y., and Jan, Y.N. 2003. Lethal giant larvae acts together with numb in notch inhibition and cell fate specification in the Drosophila adult sensory organ precursor lineage. Curr. Biol. 13: 778–783. [DOI] [PubMed] [Google Scholar]
- Kagami M., Toh-e, A., and Matsui, Y. 1998. Sro7p, a Saccharomyces cerevisiae counterpart of the tumor suppressor l(2)gl protein, is related to myosins in function. Genetics 149: 1717–1727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klein T. 2003. The tumour suppressor gene l(2)giant discs is required to restrict the activity of Notch to the dorsoventral boundary during Drosophila wing development. Dev. Biol. 255: 313–333. [DOI] [PubMed] [Google Scholar]
- Lehman K., Rossi, G., Adamo, J.E., and Brennwald, P. 1999. Yeast homologs of tomosyn and lethal giant larvae function in exocytosis and are associated with the plasma membrane SNARE, Sec9. J. Cell Biol. 146: 125–140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H.S., Wang, D., Shen, Q., Schonemann, M.D., Gorski, J.A., Jones, K.R., Temple, S., Jan, L.Y., and Jan, Y.N. 2003. Inactivation of Numb and Numblike in embryonic dorsal forebrain impairs neurogenesis and disrupts cortical morphogenesis. Neuron 40: 1105–1118. [DOI] [PubMed] [Google Scholar]
- Lu B., Jan, L., and Jan, Y.N. 2000. Control of cell divisions in the nervous system: Symmetry and asymmetry. Annu. Rev. Neurosci. 23: 531–556. [DOI] [PubMed] [Google Scholar]
- Musch A., Cohen, D., Yeaman, C., Nelson, W.J., Rodriguez-Boulan, E., and Brennwald, P.J. 2002. Mammalian homolog of Drosophila tumor suppressor lethal (2) giant larvae interacts with basolateral exocytic machinery in Madin-Darby canine kidney cells. Mol. Biol. Cell 13: 158–168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohshiro T., Yagami, T., Zhang, C., and Matsuzaki, F. 2000. Role of cortical tumour-suppressor proteins in asymmetric division of Drosophila neuroblast. Nature 408: 593–596. [DOI] [PubMed] [Google Scholar]
- Pagliarini R.A. and Xu, T. 2003. A genetic screen in Drosophila for metastatic behavior. Science 302: 1227–1231. [DOI] [PubMed] [Google Scholar]
- Peng C.Y., Manning, L., Albertson, R., and Doe, C.Q. 2000. The tumour-suppressor genes lgl and dlg regulate basal protein targeting in Drosophila neuroblasts. Nature 408: 596–600. [DOI] [PubMed] [Google Scholar]
- Petersen P.H., Zou, K., Hwang, J.K., Jan, Y.N., and Zhong, W. 2002. Progenitor cell maintenance requires numb and numblike during mouse neurogenesis. Nature 419: 929–934. [DOI] [PubMed] [Google Scholar]
- Plant P.J., Fawcett, J.P., Lin, D.C., Holdorf, A.D., Binns, K., Kulkarni, S., and Pawson, T. 2003. A polarity complex of mPar-6 and atypical PKC binds, phosphorylates and regulates mammalian Lgl. Nat. Cell Biol. 5: 301–308. [DOI] [PubMed] [Google Scholar]
- Rolls M.M., Albertson, R., Shih, H.-P., Lee, C.-Y., and Doe, C.Q. 2003. Drosophila aPKC regulates cell polarity and cell proliferation in neuroblasts and epithelia. J. Cell Biol. 163: 1089–1098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sambrook J. and Russell, D.W. 2001. Molecular cloning. A laboratory manual, 3rd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
- Sangueza O.P., Sangueza, P., Valda, L.R., Meshul, C.K., and Requena, L. 1994. Multiple primitive neuroectodermal tumors. J. Am. Acad. Dermatol. 31: 356–361. [DOI] [PubMed] [Google Scholar]
- Shen Q. and Temple, S. 2002. Creating asymmetric cell divisions by skewing endocytosis. Sci. STKE 162: PE52. [DOI] [PubMed] [Google Scholar]
- Shen Q., Zhong, W., Jan, Y.N., and Temple, S. 2002. Asymmetric Numb distribution is critical for asymmetric cell division of mouse cerebral cortical stem cells and neuroblasts. Development 129: 4843–4853. [DOI] [PubMed] [Google Scholar]
- Strand D. 1998. The tumor suppressor l(2)gl; a myosin II-binding protein family. In G proteins, cytoskeleton and cancer (eds. H. Maruta and K. Kohama), pp. 61–78. R.G. Landers, Georgetown, TX.
- Strand D., Jakobs, R., Merdes, G., Neumann, B., Kalmes, A., Heid, H.W., Husmann, I., and Mechler, B.M. 1994. The Drosophila lethal(2) giant larvae tumor suppressor protein forms homo-oligomers and is associated with nonmuscle myosin II heavy chain. J. Cell Biol. 127: 1361–1373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strand D., Unger, S., Corvi, R., Hartenstein, K., Schenkel, H., Kalmes, A., Merdes, G., Neumann, B., Krieg-Schneider, F., Coy, J.F., et al. 1995. A human homolog of the Drosophila tumour suppressor gene l(2)gl maps to 17p11.2–12 and codes for a cytoskeletal protein that associates with nonmuscle myosin II heavy chain. Oncogene 11: 291–301. [PubMed] [Google Scholar]
- Tallquist M.D., and Soriano, P. 2000. Epiblast-restricted Cre expression in MORE mice: A tool to distinguish embryonic vs. extra-embryonic gene function. Genesis 26: 113–115. [DOI] [PubMed] [Google Scholar]
- Tanentzapf G. and Tepass, U. 2003. Interactions between the crumbs, lethal giant larvae and bazooka pathways in epithelial polarization. Nat. Cell Biol. 5: 46–52. [DOI] [PubMed] [Google Scholar]
- Togo T. and Steinhardt, R.A. 2004. Nonmuscle myosin IIA and IIB have distinct functions in the exocytosis-dependent process of cell membrane repair. Mol. Biol. Cell 15: 688–695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tullio A.N., Bridgman, P.C., Tresser, N.J., Chan, C., Conti, M.N., Adelstein, R.S., and Hara, Y. 2001. Structural abnormalities develop in the brain after ablation of the gene encoding nonmuscle myosin II-B heavy chain. J. Comp. Neurol. 433: 62–74. [DOI] [PubMed] [Google Scholar]
- Vasioukhin V., Bauer, C., Degenstein, L., Wise, B., and Fuchs, E. 2001. Hyperproliferation and defects in epithelial polarity upon conditional ablation of alpha-catenin in skin. Cell 104: 605–617. [DOI] [PubMed] [Google Scholar]
- Walsh C.A. 1999. Genetic malformations of the human cerebral cortex. Neuron 23: 19–29. [DOI] [PubMed] [Google Scholar]
- Yamanaka T., Horikoshi, Y., Sugiyama, Y., Ishiyama, C., Suzuki, A., Hirose, T., Iwamatsu, A., Shinohara, A., and Ohno, S. 2003. Mammalian Lgl forms a protein complex with PAR-6 and aPKC independently of PAR-3 to regulate epithelial cell polarity. Curr. Biol. 13: 734–743. [DOI] [PubMed] [Google Scholar]
- Zhong W. 2003, Diversifying neural cells through order of birth and asymmetry of division. Neuron. 37: 11–14. [DOI] [PubMed] [Google Scholar]
- Zhong W., Feder, J.N., Jiang, M.M., Jan, L.Y., and Jan, Y.N. 1996. Asymmetric localization of a mammalian numb homolog during mouse cortical neurogenesis. Neuron 17: 43–53. [DOI] [PubMed] [Google Scholar]