Abstract
Inflammatory cross talk between perivascular adipose tissue and the blood vessel wall has been proposed to contribute to the pathogenesis of atherosclerosis. We previously reported that human perivascular (PV) adipocytes exhibit a proinflammatory phenotype and less adipogenic differentiation than do subcutaneous (SQ) adipocytes. To gain a global view of the genomic basis of biologic differences between PV and SQ adipocytes, we performed genome-wide expression analyses to identify differentially expressed genes between adipocytes derived from human SQ vs. PV adipose tissues. Although >90% of well-expressed genes were similarly regulated, we identified a signature of 307 differentially expressed genes that were highly enriched for functions associated with the regulation of angiogenesis, vascular morphology, inflammation, and blood clotting. Of the 156 PV upregulated genes, 59 associate with angiogenesis, vascular biology, or inflammation, noteworthy of which include TNFRSF11B (osteoprotegerin), PLAT, TGFB1, THBS2, HIF1A, GATA6, and SERPINE1. Of 166 PV downregulated genes, 21 associated with vascular biology and inflammation, including ANGPT1, ANGPTL1, and VEGFC. Consistent with the emergent hypothesis that PV adipocytes differentially regulate angiogenesis and inflammation, cell culture-derived adipocyte-conditioned media from PV adipocytes strongly enhanced endothelial cell tubulogenesis and monocyte migration compared with media from SQ adipocytes. These findings demonstrate that PV adipocytes have the potential to significantly modulate vascular inflammatory crosstalk in the setting of atherosclerosis by their ability to signal to both endothelial and inflammatory cells.
Keywords: human perivascular fat, global gene expression pattern, inflammation, angiogenesis, atherosclerosis
atherosclerosis-prone blood vessels are surrounded by perivascular adipose tissue (PVAT), which abuts the vascular adventitia and has traditionally been thought to play a role in supporting vascular structure and/or metabolism. Recently, PVAT has been proposed to influence vascular function and the pathogenesis of vascular disease (1, 5, 30, 37). PVAT releases paracrine factors that regulate inflammation, vasoreactivity, and vascular smooth muscle cell growth and migration (11, 13, 24, 27, 38, 44). PVAT, like other adipose depots, contains a mixture of cell types, including mature adipocytes and stromovascular (SV) cells (which include preadipocytes, vascular cells, and inflammatory cells). Very little is known regarding the specific characteristics of perivascular adipocytes and how they might contribute to PVAT's effects on vascular disease.
Prevailing evidence indicates that anatomically separated regional adipose depots are distinct with respect to gene expression patterns and functional characteristics (4, 7–9, 15). These depot-specific features result from inherent differences in the adipocytes, imparted by the distinctive precursor cells (preadipocytes) from which they are derived (12, 19, 40). Thus, in vitro differentiation of preadipocytes isolated from regional adipose depots produces adipocytes that phenotypically resemble their in vivo counterparts (7, 8, 32, 33). In this regard, we demonstrated that in vitro differentiated human coronary artery perivascular (PV) adipocytes exhibit a reduced state of adipogenic differentiation compared with adipocytes from subcutaneous (SQ) or visceral (perirenal) adipose depots derived from the same subjects (8). Secretion of anti-inflammatory adiponectin was markedly reduced, whereas that of proinflammatory cytokines IL-6, IL-8, and monocyte chemoattractant protein-1 (MCP-1), was markedly increased, in PV adipocytes. This unique phenotype of in vitro differentiated human PV adipocytes closely mirrored the phenotype of those residing in the PVAT depot, suggesting that PV adipocytes exhibit a unique gene expression profile that underlies their role in vascular homeostasis.
Here, we performed global gene expression analyses with in vitro differentiated adipocytes from human coronary artery PVAT and subcutaneous adipose tissues derived from unrelated donors. Compared with SQ adipocytes, PV adipocytes exhibit elevated expression of a broad array of genes whose aggregated functions are associated with the regulation of angiogenesis and vascular morphology, inflammation, and blood clotting. Our global gene expression data also indicate that human coronary artery PV adipocytes are more similar to white than classical brown adipocytes, a finding consistent with our previous report (8). Complimentary functional analyses confirm that PV adipocytes are more potent at inducing angiogenesis and monocyte migration compared with SQ adipocytes. These findings provide important insight into the unique molecular characteristics of PV adipocytes and the role of PVAT in regulating vascular function.
MATERIALS AND METHODS
Adipose tissue collection.
SQ and left coronary artery PV adipose tissue samples were collected from human organ donors (Life Center Network, OH) who suffered terminal illnesses or injuries and whose organs were being procured for transplantation. None of the hearts employed in this study were used for transplantation or perfused with preservative solutions. These donors were nondiabetic and had no known metabolic diseases or atherosclerosis. Available donor-specific information of the six subjects in the primary cohort is provided in Table 1 (adapted from Ref. 8). This same cohort was used in our previously published report documenting a proinflammatory phenotype of human PV adipocytes (8).
Table 1.
Donor demography
| Donor | Age | Sex | Race | BMI | Diabetes | Hypertension | Hyperlipidemia |
|---|---|---|---|---|---|---|---|
| 1* | 15 | F | Afa | 22.1 | no | no | no |
| 2* | 26 | F | Afa | 28.4 | no | no | no |
| 3* | 54 | M | Cauc | 25.1 | no | >10 yr | no |
| 4 | 64 | M | Cauc | 26.0 | no | >6 yr | no |
| 5 | 63 | F | Afa | 32.4 | no | no | no |
| 6 | 61 | M | Cauc | 35.8 | no | 5 yr | no |
BMI, body mass index; Afa, African American; Cauc, Caucasian.
Donor samples used in the microarray study.
Subcutaneous adipose tissue (SQAT, collected from beneath the skin of upper abdomen) and PVAT overlying the left coronary artery was carefully dissected out. Adipose tissues were collected in cold DMEM/F12 medium, transported to the laboratory and processed within 30–60 min. Some samples were prepared for histology and examined after hematoxylin and eosin staining as described previously (8). Microarray study was performed with RNA samples from in vitro differentiated preadipocytes (see below) isolated from the first three paired samples (SQAT and PVAT) of the cohort (donors 1–3) shown in Table 1. To validate selected gene expression data of the microarray study, we utilized RNA samples from in vitro differentiated preadipocytes of the entire cohort of six paired samples (SQAT and PVAT) shown in Table 1. Furthermore, RNA samples from in vitro differentiated preadipocytes isolated from SQAT of a cohort of 20 patients undergoing elective abdominal surgery at the University of Cincinnati Medical Center were used to provide independent validation of selected gene expression data of our microarray study. Processing of the tissues and isolation/differentiation of the cells was performed in an identical manner in all cases. Our study was approved by the Institutional Review Board of the University of Cincinnati.
Preadipocyte isolation and in vitro adipogenic differentiation.
Adipose tissue was digested with collagenase, and preadipocytes were isolated according to previously described methods (7, 8). We plated SV fractions (∼106 cells) from 5–8 g of adipose tissue in a T25 flask with DMEM/F12 medium supplemented with 10% fetal bovine serum; the following day, floating and dead cells were removed by washing twice with PBS, and fresh DMEM/F12 supplemented with 10% FBS was applied for one more day. Thereafter, cells were collected by brief trypsinization (to avoid lifting firmly attached macrophages) and transferred to a T75 flask for continued culture. After 3–4 days, nearly confluent preadipocytes from the T75 flask were removed and separated into three T75 flasks. These T75 flasks became near-confluent in 3–4 days, after which the cells were removed and plated for in vitro adipogenic differentiation to evaluate gene expression, functional characterization, and cytoplasmic lipid droplet accumulation. Macrophage contamination was <1%, as determined by F4/80 immunostaining, a finding similar to that reported previously (39).
For in vitro differentiation, preadipocytes (80–90% confluence) were plated onto six-well plates or 10 cm dishes. Upon reaching confluence, the culture medium was replaced with human adipocyte differentiation medium (Cell Applications, cat. no. 811D) for 28 days (8), with replenishment of medium in every 4 days. RNA was extracted from these cells with RNeasy lipid mini kits (Qiagen) and quantified, and quality was determined. We selected RNA from 7-day differentiated cells for Affymetrix Human Genes ST 1.0 gene chip assay. Quantitative RT-PCR validation of gene expression was performed as we described previously (7, 8). Gene expression levels were normalized with ribosomal protein large P0 (RPLP0) as an endogenous control. In some experiments, supernatants from differentiating adipocytes were collected for determination of secreted proteins by ELISA, as we described previously (8). The values were normalized to cellular protein levels (Bio-Rad protein assay kit). Initial plating of equivalent numbers of PV and SQ preadipocytes was routinely confirmed by cell counting and, in some experiments, by measuring cellular DNA content with Hoechst 33342 (Sigma) following differentiation for 7 days. Normalization of data with either cellular protein or DNA content produced similar results.
RNA labeling.
Purified RNAs were analyzed for concentration and integrity using the Agilent Bioanalyzer RNA 6000 Pico Kit (Agilent Technologies, Santa Clara, CA) and then labeled as appropriate for hybridization to Human Genes ST 1.0 GeneChip (Affymetrix, Santa Clara, CA) using the Affymetrix WT labeling kit.
Microarray data analysis.
CEL files were generated from the Affymetrix GeneChip Scanner 3000 7G using Expression Console v1.1.1 (Affymetrix). CEL files were subjected to robust multiarray average (RMA) normalization using Affymetrix CDF probe-set definitions. Relative gene expression levels were calculated based on the ratio of each sample's RMA signal intensity (2**RMA) to that of the median for each probe-set across the six samples. Probe-sets mapping to expressed genes were defined as those for which there were no known cross-hybridizing targets and that had >90% sequence identity to the hg19 genome reference human genome. Differentially expressed genes were identified by selecting probe-sets that demonstrated RMA intensity value >6.0 in more than three samples and that differed between source tissues per a Welch t-test P < 0.05 with (or without) Benjamini-Hochberg false discovery testing correction and fold-change cutoff more than twofold for signature genes or 1.5× for genes used in pathway analysis. The dataset for this manuscript has been uploaded to the National Center for Biotechnology Information (NCBI) Gene Expression Omnibus (GEO) database as record GSE45169 http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE45169.
Transendothelial monocyte migration assay.
Human retinal endothelial cells (HREC) were seeded at 150,000 cells/well in 3 μm Transwell inserts (BD Falcon, cat. #353492) and allowed to form a monolayer. Samples of conditioned medium (250 μl) from in vitro differentiated (7 day) human SQ and PV adipocytes cultured in adipogenic differentiation medium (Cell Applications) were placed in the bottom chambers of the 24 well-plate containing the Transwell inserts, after which THP-1 cells (ATCC) (5 × 104, passages 12–14) suspended in serum-free medium were added to the top chambers and allowed to migrate for 16 h. Cells were removed from the luminal sides of the insert with a cotton swab, and the filter was then fixed in methanol, excised, and stained with 4',6-diamidino-2-phenylindole. Cells on the abluminal side were counted by capturing five low power fields (×10), and the results were normalized to the basal level of THP-1 migration in the absence of a chemokine gradient. Human MCP-1 (R&D) was used as a positive control at the concentration of 20 and 2,000 pg/ml. The choice of HREC is based on our previous study demonstrating the utility of these human endothelial cells to form a stable monolayer for the migration assay (36).
Tubulogenesis assay.
We used enhanced green fluorescent protein (eGFP)-labeled human umbilical vein endothelial cells (HUVEC) for an in vitro tubulogenesis assay to evaluate the angiogenic potential of culture supernatants (conditioned media) from in vitro differentiated human PV and SQ adipocytes. HUVEC, obtained from ATCC, were labeled by infecting with lenti-PGK-eGFP vector; stably expressing eGFP-labeled cells were selected with puromycin and further purified by FACS sorting in our laboratory. These cells (1.5 × 104 cell/well) were subsequently plated onto Matrigel (BD Biosciences, San Jose, CA)-coated μ-slides (Ibidi, Verona, WI) in 10 μl of growth factor-reduced medium (46), to which was added 50 μl of conditioned medium from in vitro differentiated (7 day) human SQ or human PV adipocytes. Medium-incubated without cells served as the control. After 5 or 18 h, eGFP-labeled tube formation was visualized with fluorescent inverted microscope (Olympus), captured with a digital camera. Total tube length was measured as an index of angiogenesis.
Additional data analyses and quality control.
Data are expressed as means ± SE from at least three independent experiments. Statistical significance was evaluated by ANOVA.
RESULTS
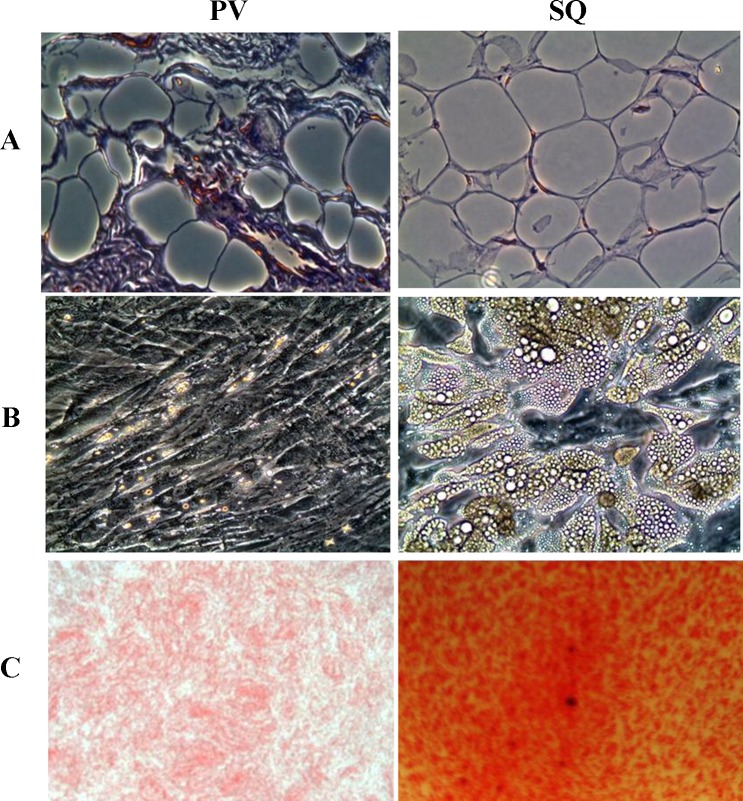
We prepared adipocytes from paired human SQ and PV adipose tissues of unrelated donors using collagenase digestion to release SV cells that then were grown under adipogenic culture conditions (7, 8). Under our defined culture conditions, 80–90% of the adherent SV cells from both sources accumulated cytoplasmic lipid droplets following in vitro adipogenic differentiation. However, adipocytes derived from the two sources were morphologically distinct, with PV adipocytes appearing more irregular in shape and smaller in average size than those from SQ (Fig. 1A). Consistent with their smaller size in situ, and as reported previously, differentiating PV adipocytes accumulated fewer lipid droplets compared with SQ adipocytes (Fig. 1, B and C) (8). Spectrophotometric quantification (not shown) demonstrated nearly 2.5-fold less oil red O-positive lipid accumulation in PV compared with SQ adipocytes (8). RT-qPCR analysis of genes previously identified to be differentially expressed confirmed our earlier findings (8), e.g., reduced expression of classic adipogenic differentiation-associated genes in PV-derived adipocytes compared with those from SQ (not shown).
Fig. 1.
Representative (n = 6) light microscopic (×20) appearance of hematoxylin and eosin-stained sections of left coronary artery perivascular (PV) and subcutaneous (SQ) adipose tissues from human organ donors (A). Preadipocytes (SV cells) were isolated from these tissues and differentiated in vitro as published previously (6, 7). Lipid droplet accumulation by differentiating preadipocytes was examined after 28 days by light microscopy (B) and further verified by imaging after oil red-O staining (C) (×20). Spectrophotometric quantification of the color intensity of oil red-O-positive materials at 510 nm showed that SQ adipocytes accumulate ∼2.6 fold higher levels of cytoplasmic neutral lipids compared with PV adipocytes when normalized to cellular protein levels (7).
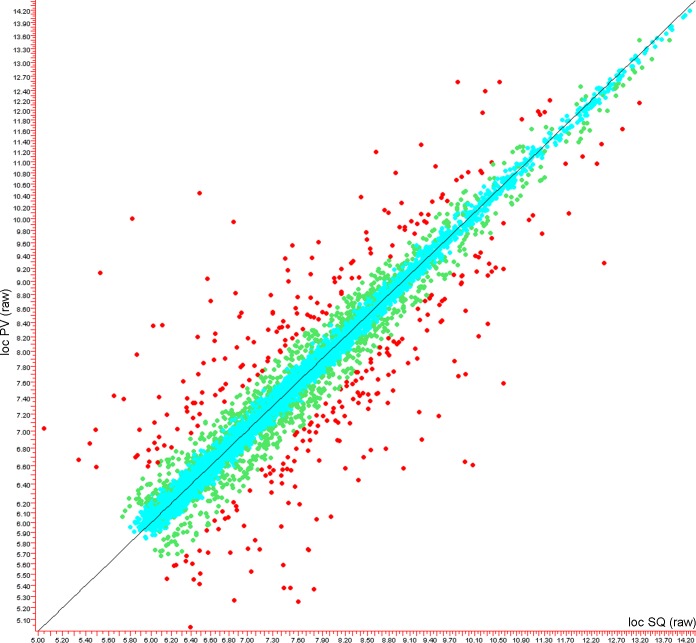
We next performed global gene expression analyses in the PV and SQ adipocyte samples, using the Affymetrix Human Genes ST 1.0 gene chip assay. We identified 15,721 gene-aligned probe-sets whose expression was >6.0 RMA units in at least three samples. As shown in the scatter-gram in Fig. 2, the overall gene expression pattern of these transcripts was highly similar between PV and SQ adipocytes, with >90% of these not differing by more than twofold between any individuals or tissue of origin (blue circles). For example, only 454 probe-sets were identified that differed as a function of the individual using a Welch P value cut-off of 0.05, and fewer than 90 of which exhibited >1.5-fold variation compared with other individuals. In contrast, 1,599 probe-sets differed significantly (P < 0.05) as a function of tissue of origin (green circles), and of these, 405 probe-sets (red circles) corresponding to 307 different genes, were more than 1.5× different between adipocytes from the two different tissues of origin. The relative patterning and distribution of these are shown in the heat map in Fig. 3, and the most differentially regulated of these are listed in Table 2.
Fig. 2.
Scatter plot showing the overall similarity of global gene expression patterns in human PV and SQ adipocytes differentiated in vitro for 7 days. Each dot represents a gene; the y-axis is the average for all PV samples, and the x-axis is that for SQ. Axis units are robust multiarray average (RMA) values (RMA = log2 scale). Genes that are insignificantly different are aqua, P < 0.05 are green, P < 0.05 and >2× are red. Note that most of the genes are quite similarly expressed in the 2 different sample types, but that a select population of genes is substantially different.
Fig. 3.

Heat map showing differential and reproducible gene expression pattern differences between human PV and SQ adipocytes from 3 unrelated donors. The y-axis represents 405 probe-sets that correspond to 307 unique genes.
Table 2.
List of differentially expressed genes in PV and SQ adipocytes
| Common | Description | PV/SQ Ratio | P Value | PV RMA (log2) | SQ RMA (log2) | AFFY ST 1.0 Probe Set ID |
|---|---|---|---|---|---|---|
| TNFRSF11B | tumor necrosis factor receptor superfamily, member 11b | 18.36 | 1.70E-02 | 10.00 | 5.81 | 8152512 |
| KIAA1199 | KIAA1199 | 15.69 | 2.07E-03 | 10.44 | 6.48 | 7985317 |
| ITGA8 | integrin, alpha 8 | 12.33 | 2.07E-03 | 9.13 | 5.52 | 7932254 |
| C7 | complement component 7 | 8.67 | 2.13E-02 | 9.94 | 6.84 | 8105084 |
| SERPINE2 | serpin peptidase inhibitor, clade E, member 2 | 6.86 | 1.42E-02 | 12.58 | 9.82 | 8059376 |
| MEST | mesoderm specific transcript homolog (mouse) | 6.01 | 1.73E-02 | 11.19 | 8.61 | 8136248 |
| TM4SF1 | transmembrane 4 L six family member 1 | 5.62 | 1.89E-02 | 9.04 | 6.57 | 8091411 |
| TCF21 | transcription factor 21 | 5.12 | 4.68E-02 | 8.35 | 6.01 | 8122176 |
| PAMR1 | peptidase domain containing associated with muscle regeneration 1 | 4.83 | 4.18E-02 | 8.36 | 6.10 | 7947512 |
| SERPINE1 | serpin peptidase inhibitor, clade E, member 1 | 4.39 | 1.24E-02 | 12.39 | 10.27 | 8135069 |
| PRUNE2 | prune homolog 2 (Drosophila) | 4.37 | 2.58E-03 | 8.71 | 6.60 | 8161884 |
| PLK2 | polo-like kinase 2 | 4.34 | 4.44E-02 | 7.96 | 5.86 | 8112202 |
| THBS2 | thrombospondin 2 | 4.24 | 9.11E-04 | 12.58 | 10.51 | 8130867 |
| CFH | complement factor H | 4.19 | 1.96E-02 | 11.32 | 9.27 | 7908459 |
| PLAT | plasminogen activator, tissue | 4.15 | 2.63E-02 | 9.56 | 7.53 | 8150509 |
| PDE5A | phosphodiesterase 5A, cGMP-specific | 4.04 | 8.42E-03 | 6.95 | 4.95 | 8102532 |
| CSGALNACT1 | chondroitin sulfate N-acetylgalactosaminyltransferase 1 | 4.00 | 7.53E-04 | 7.03 | 5.04 | 8149574 |
| PTGS1 | prostaglandin-endoperoxide synthase 1 | 3.93 | 2.77E-02 | 10.37 | 8.41 | 8157650 |
| IL6 | interleukin 6 (interferon, beta 2) | 3.92 | 2.83E-02 | 8.82 | 6.87 | 8131803 |
| HEG1 | HEG homolog 1 (zebrafish) | 3.80 | 5.41E-04 | 10.80 | 8.89 | 8090193 |
| DKK1 | dickkopf homolog 1 (Xenopus laevis) | 3.79 | 9.64E-03 | 9.35 | 7.44 | 7927631 |
| LAMA1 | laminin, alpha 1 | 3.48 | 3.15E-04 | 7.42 | 5.64 | 8022176 |
| ALCAM | activated leukocyte cell adhesion molecule | 3.42 | 2.36E-03 | 9.61 | 7.85 | 8081431 |
| F2R | coagulation factor II (thrombin) receptor | 3.38 | 3.48E-02 | 8.20 | 6.46 | 8106393 |
| CYP1B1 | cytochrome P450, family 1, subfamily B, polypeptide 1 | 3.35 | 3.32E-03 | 11.95 | 10.22 | 8051583 |
| AHR | aryl hydrocarbon receptor | 3.28 | 1.00E-02 | 9.17 | 7.47 | 8131614 |
| GCNT4 | glucosaminyl (N-acetyl) transferase 4, core 2 | 3.24 | 3.16E-02 | 6.50 | 4.81 | 8112668 |
| PLA2G4A | phospholipase A2, group IVA (cytosolic, calcium-dependent) | 3.19 | 3.24E-02 | 7.39 | 5.73 | 7908351 |
| SLC7A11 | solute carrier family 7, member 11 | 3.07 | 3.16E-02 | 8.54 | 6.94 | 8102800 |
| MARCH3 | membrane-associated ring finger (C3HC4) 3 | 3.06 | 2.08E-02 | 8.25 | 6.65 | 8113790 |
| MXRA5 | matrix-remodelling associated 5 | 3.05 | 8.30E-03 | 9.37 | 7.77 | 8171172 |
| CD9 | CD9 molecule | 3.01 | 3.30E-02 | 8.48 | 6.91 | 7953291 |
| PTGER2 | prostaglandin E receptor 2 (subtype EP2), 53 kDa | 2.92 | 1.28E-02 | 7.01 | 5.48 | 7974366 |
| ADAMTS15 | ADAM metallopeptidase with thrombospondin type 1 motif, 15 | 0.37 | 4.91E-02 | 7.34 | 8.78 | 7945232 |
| CYB5A | cytochrome b5 type A (microsomal) | 0.36 | 3.90E-02 | 9.75 | 11.25 | 8023855 |
| CHN1 | chimerin (chimaerin) 1 | 0.35 | 2.92E-03 | 5.52 | 7.04 | 8056890 |
| AOC2 | amine oxidase, copper containing 2 (retina-specific) | 0.34 | 1.28E-02 | 5.26 | 6.86 | 8007414 |
| HOXC8 | homeobox C8 | 0.33 | 1.93E-03 | 6.57 | 8.19 | 7955869 |
| PLA2G2A | phospholipase A2, group IIA (platelets, synovial fluid) | 0.32 | 1.70E-03 | 10.08 | 11.75 | 7913216 |
| USP18 | ubiquitin specific peptidase 18 | 0.30 | 1.10E-02 | 6.70 | 8.44 | 8074606 |
| MX2 | myxovirus (influenza virus) resistance 2 (mouse) | 0.30 | 2.52E-02 | 6.78 | 8.54 | 8068697 |
| GFRA1 | GDNF family receptor alpha 1 | 0.29 | 3.93E-02 | 6.04 | 7.82 | 7936494 |
| AVPR1A | arginine vasopressin receptor 1A | 0.29 | 3.72E-02 | 5.59 | 7.41 | 7964660 |
| ANGPTL1 | angiopoietin-like 1 | 0.27 | 1.02E-02 | 7.87 | 9.77 | 7922598 |
| CACNB2 | calcium channel, voltage-dependent, beta 2 subunit | 0.27 | 8.03E-03 | 6.44 | 8.37 | 7926506 |
| APCDD1 | adenomatosis polyposis coli down-regulated 1 | 0.26 | 3.51E-02 | 8.38 | 10.32 | 8020141 |
| ANGPT1 | angiopoietin 1 | 0.26 | 7.32E-03 | 6.06 | 8.01 | 8152297 |
| TNC | tenascin C | 0.26 | 1.31E-02 | 8.21 | 10.16 | 8163637 |
| GPD1L | glycerol-3-phosphate dehydrogenase 1-like | 0.26 | 2.68E-02 | 6.80 | 8.75 | 8078386 |
| CILP | cartilage intermediate layer protein, nucleotide pyrophosphohydrolase | 0.26 | 1.70E-02 | 5.74 | 7.72 | 7989750 |
| CALCRL | calcitonin receptor-like | 0.25 | 9.29E-03 | 5.73 | 7.73 | 8057578 |
| PAPPA2 | pappalysin 2 | 0.25 | 6.66E-03 | 7.23 | 9.23 | 7907572 |
| RP1-170O19.21 | homeobox A9 | 0.23 | 6.67E-05 | 5.37 | 7.50 | 8138749 |
| DPT | dermatopontin | 0.23 | 1.09E-02 | 7.67 | 9.84 | 7922130 |
| VIT | vitrin | 0.21 | 1.50E-02 | 7.70 | 9.95 | 8041467 |
| PTPN22 | protein tyrosine phosphatase, nonreceptor type 22 (lymphoid) | 0.21 | 4.75E-04 | 4.14 | 6.42 | 7918657 |
| HOXC10 | homeobox C10 | 0.20 | 1.02E-04 | 5.25 | 7.60 | 7955858 |
| SLC7A6 | solute carrier family 7 (amino acid transporter), member 6 | 0.20 | 2.74E-03 | 7.18 | 9.52 | 7996772 |
| FAM213A | chromosome 10 open reading frame 58 | 0.19 | 3.25E-02 | 6.90 | 9.27 | 7928695 |
| FAM134B | family with sequence similarity 134, member B | 0.19 | 2.64E-03 | 6.58 | 9.01 | 8111136 |
| QPCT | glutaminyl-peptide cyclotransferase | 0.19 | 3.49E-03 | 5.36 | 7.79 | 8041508 |
| TRHDE | thyrotropin-releasing hormone degrading enzyme | 0.18 | 1.67E-02 | 4.77 | 7.23 | 7957221 |
| MAB21L1 | mab-21-like 1 (C. elegans) | 0.15 | 2.30E-03 | 4.80 | 7.56 | 7970949 |
| CIDEC | cell death-inducing DFFA-like effector c | 0.13 | 3.58E-02 | 7.58 | 10.57 | 8085244 |
| TXNIP | thioredoxin interacting protein | 0.11 | 1.78E-03 | 9.28 | 12.44 | 7904726 |
| ABCD2 | ATP-binding cassette, sub-family D (ALD), member 2 | 0.10 | 1.67E-02 | 6.64 | 9.94 | 7962312 |
| NRN1 | neuritin 1 | 0.09 | 1.65E-02 | 6.61 | 10.06 | 8123739 |
PV, perivascular; SV, stromovascular; RMA, robust multiarray average.
Among the most differentially expressed genes that were strikingly higher in PV adipocytes included tumor necrosis factor receptor superfamily, member 11b (osteoprotegerin), integrin alpha 8, complement component 7, as well as many genes associated with vascular, cardiac, and hemostatic biology.
Among the genes expressed at much lower levels in PV adipocytes compared with SQ adipocytes were cell death-inducing DFFA-like effector c (Cidec), neuritin 1, dermatopontin, homeobox A9, C8, C9, C10, neurocalcin delta, zinc finger homeobox 4, and GDNF family receptor alpha 1, arginine vasopressin receptor 1A, and angiopoietin-like 1.
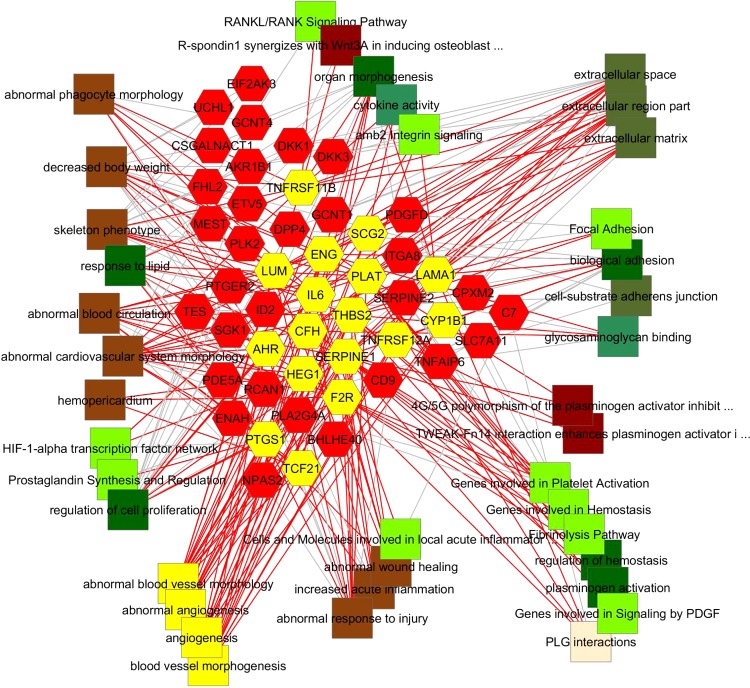
A summary of the features that are highly enriched in the two different lists are shown in Table 3. These results indicate that there are highly significant biological differences between the two distinct adipocyte cell types, which map to early developmental differences, cardiovascular biology-related functions, and features involving the control of inflammation. PV adipocyte overexpressed genes are strikingly enriched for gene functions associated with the regulation of vascular development and the control of vascular morphology. As shown in the network diagram in Fig. 4, the genes that are associated with these specific biological processes, pathways, and mouse gene knockout phenotypes, as well as the functional features themselves, are shown highlighted in yellow and connected by red line edges. What is also illustrated by red lines coming from these genes in to other concepts are the other molecular and functional associations of these genes, and by gray lines coming into them, the other properties, gene ontology pathway, and mutant mouse gene knockout phenotype associations that are also enriched among these PV-overexpressed genes. Thus it is possible to see in this view that the preponderance of these vascular modifying gene-associated features involve in the control of macrophage functions, inflammation, blood coagulation, hemostasis, and fibrinolysis.
Table 3.
Features of differentially expressed genes in PV vs. SQ adipocytes
| Adipo-UpReg | Category | ID | Name | P Value | Genes Query, n | Genes Genome, n | Representative Genes |
|---|---|---|---|---|---|---|---|
| PV | GO: Biological Process | GO:0001525 | angiogenesis | 2.28E-06 | 14 | 380 | ADAM17, ADORA2B, AHR, CYP1B1, CYR61, ENG, ERRFI1, F2R, GATA6, HEG1, HIF1A, IL6, ITGAV, PLAT, PTGS1, PTPRJ, SCG2, SERPINE1, SRPX2, TGFB1, THBS2, TNFRSF12A |
| PV | GO: Biological Process | GO:0001936 | regulation of endothelial cell | 2.55E-02 | 3 | 79 | |
| PV | GO: Biological Process | GO:0010574 | regulation of vascular | 3.09E-05 | 4 | 23 | |
| PV | Mouse Phenotype | MP:0000260 | abnormal angiogenesis | 5.91E-04 | 13 | 327 | |
| PV | Mouse Phenotype | MP:0003542 | abnormal vascular endothelial cell development | 7.53E-03 | 3 | 29 | |
| PV | GO: Biological Process | GO:0007596 | blood coagulation | 1.94E-07 | 18 | 521 | CD9, F2R, ITGAV, LRP8, PDE5A, PLA2G4A, PLAT, PTPRJ, RAB27A, SERPINE1, SERPINE2, SLC7A11 |
| PV | GO: Biological Process | GO:0042060 | wound healing | 7.96E-11 | 25 | 656 | ADAM17, CD9, ENG, F2R, HIF1A, ITGAV, LRP8, PLAT, PLA2G4A, PTPRJ, SERPINE1, SERPINE2, TGFB1 |
| PV | GO: Biological Process | GO:0006954 | inflammatory response | 2.88E-04 | 13 | 521 | ADAM17, ADORA2B, AHR, CFH, EIF2AK3, ERRFI1, F2R, FKBP11, GBA, GJA1, HIF1A, IL6, ITGAV, LUM, PDE5A, PLA2G4A, PLAT, PTGS1, PTPRJ, RCAN1, SCG2, SERPINE1, TGFB1, TNFAIP6 |
| PV | Mouse Phenotype | MP:0001853 | heart inflammation | 1.09E-03 | 4 | 33 | |
| PV | Mouse Phenotype | MP:0001846 | increased inflammatory response | 1.92E-03 | 20 | 718 | |
| PV | GO: Biological Process | GO:0042127 | regulation of cell proliferation | 4.73E-11 | 35 | 1252 | ADORA2B, AHR, F2R, GRN, HIF1A, IL6, PLA2G4A, PTGER2, PTGS1, SERPINE1, SERPINE2, TGFB1 |
| SQ | GO: Biological Process | GO:0002520 | immune system development | 1.03E-04 | 16 | 628 | ANGPT1, AR, BCL6, CHN1, DUSP10, FAM213A, GPC3, HMGB2, HOXA7, IL11RA, JAK2, KITLG, MAP1B, NEFL, NFKBIA, PLA2G2A, PLXNA4, PTPN22, PTPRD, PTPRG, ROCK2, SNRK, SSH2, STAT5A, TOB2, TWIST1, UNG, VEGFC, ZEB1 |
| SQ | GO: Biological Process | GO:0030099 | myeloid cell differentiation | 2.07E-05 | 11 | 268 | |
| SQ | GO: Biological Process | GO:0045595 | regulation of cell differentiation | 4.48E-05 | 23 | 1072 | |
| SQ | GO: Biological Process | GO:0007169 | transmembrane receptor protein tyrosine kinase signaling pathway | 7.04E-05 | 17 | 674 | ANGPT1, ANGPTL1, CHN1, TXNIP, SOS1, FGF5, JAK2, CILP, AR, ITSN1, NFKBIA, KIAA1161, GFRA1, PTPRG, STAT5A, VEGFC, NET1 |
| SQ | GO: Molecular Function | GO:0008289 | lipid binding | 1.42E-04 | 18 | 791 | ANXA6, APBB1IP, APOL2, AR, FARP1, HMGB2, HSD17B4, ING2, ITSN1, MAP1B, NET1, OPHN1, PLA2G2A, RASA1, RASA3, ROCK2, SNTB2, SOS1 |
| SQ | GO: Biological Process | GO:0010594 | regulation of endothelial cell | 2.78E-03 | 4 | 68 | ANGPT1, CCDC23, PPAP2B, RASA1, SOS1, STAT5A, VEGFC |
| SQ | Pathway | angiopoietin receptor pathway | angiopoietin receptor Tie2-mediated signaling | 1.34E-02 | 3 | 49 | |
| SQ | Pathway | lymphangiogenesis pathway | VEGFR3 signaling in lymphatic endothelium | 2.26E-02 | 2 | 23 |
GO, Gene Ontology.
Fig. 4.
Biological network-based representation of functional associations of PV-overexpressed genes. The gene-associated features (pathways, mouse knockout phenotypes, gene ontology categories) that are directly related to angiogenesis and determine blood vessel morphology and morphogenesis are shown as yellow squares that are connected to their corresponding genes (yellow hexagons), from which red edges representing all of their connections are also displayed in the network diagram. Thus, other categories to which these genes are connected have red edges coming into them. From this representation it is clear that these PV-upregulated genes achieve angiogenic control in part by their various abilities to, in aggregate, regulate wound response, response to lipids, fibrinolysis, and extracellular matrix/adhesion junctions compared with SQ adipocytes.
Genes downregulated in PV compared with SQ adipocytes are also primarily related to the regulation of macrophages and phagocytes, the response to wounding and external stimuli, and the interaction and involvement with extracellular matrix and cell junctions. A large number of transcriptional regulators that are downregulated in PV adipocytes include HOX genes A6, A7, A8 and C8, C9, C10, and D8, and AR, ZFHX4, STAT5A, and PRDM2 as well as EBF1, EBF3, and RUNX1T1. As shown in red edges, these transcriptional regulators are associated with myeloid cell development, macrophage function, locomotion, and EGF signaling. Interestingly, three regulators of angiogenesis are also present in this group, VEGFC, ANGPT1, ANGPTL1. This further suggests that genes involved in the regulation of vascular development and differentiation are differentially expressed in PV adipocytes, which may be critical for the homeostatic maintenance of coronary blood vessels.
It is interesting to note that some anti-inflammatory genes, such as tumor necrosis factor α-induced protein 6 (TAIP6 or TSG6) and suppressor of cytokine signaling 2 (SOCS2), are also expressed at elevated levels in PV adipocytes. Collectively, these gene expression characteristics are consistent with the notion that PV adipocytes can bidirectionally regulate inflammation, thus contributing to maintenance of vascular homeostasis.
The most differentially expressed inflammatory gene identified in our study was osteoprotegerin, a soluble TNF receptor family member that is also a negative regulator of vascular calcification. We selected this gene for further verification by examining its mRNA levels by RT-qPCR in PV and SQ adipocytes and by performing ELISA of culture medium to evaluate protein secretion. Consistent with our microarray data, differentiating PV adipocytes exhibit elevated mRNA expression (Fig. 5A) and protein secretion of osteoprotegerin (Fig. 5B) compared with differentiating SQ adipocytes under identical culture conditions. Note that baseline osteoprotegerin secretion by PV preadipocytes significantly exceeded that by SQ preadipocytes (day 0); interestingly, osteoprotegerin secretion fell dramatically during differentiation of SQ preadipocytes, while rising during differentiation of PV preadipocytes. Osteoprotegerin and various other differentially expressed inflammatory genes identified in our microarray study, such as MCP-1, IL-6, and IL-8, were verified using paired RNA samples from PVAT and SQ adipocytes isolated from all six donors comprising the primary cohort listed in Table 1 (8).
Fig. 5.
Elevated expression of osteoprotegerin (OPG) mRNA (A) and increased secretion of OPG protein (B) by PV adipocytes. A: RT-qPCR determination of OPG mRNA levels in differentiated adipocytes (7 day) showed nearly 15-fold increased expression of OPG mRNA in PV compared with SQ adipocytes. Values are means ± SE of 3 paired samples from unrelated donors used in the microarray assay; *P < 0.05. B: release of OPG protein into the culture medium by preadipocytes (0 day) and differentiating (4–20 days) human PV and SQ adipocytes. OPG release by PV adipocytes was elevated compared with SQ adipocytes and reached a peak around day 8 of differentiation. Values are normalized with respect to total cellular protein and represent means ± SE of 3 paired samples from unrelated donors used in the microarray assay; *P < 0.05.
Next, we examined the functional importance of altered expression of inflammatory genes by PV adipocytes. Conditioned medium (7 day) from PV and SQ adipocytes was used as a source of chemoattractants in a transendothelial monocyte migration assay. Conditioned medium from PV adipocytes, compared with SQ adipocytes, was approximately twice as potent at promoting monocyte migration (Fig. 6, A and B). These results may help to explain observations that in patients with atherosclerosis, PVAT exhibits markedly increased macrophage and T cell infiltration compared with SQ adipose tissue (14).
Fig. 6.
Conditioned medium (CM; 7 day) from PV adipocytes promotes increased monocyte migration. A: transendothelial migration of THP-1 cells was microscopically visualized following DAPI staining of fixed cells. B: cell numbers were subsequently counted by capturing 5 random low power fields (LPF, ×10). Human MCP-1 (20 and 2,000 pg/ml) was used as a positive control. CM were collected from equivalent numbers of PV or SQ adipocytes, as determined by cellular protein levels. Data are expressed as means ± SE from 3 independent experiments; *P < 0.05.
To evaluate the functional significance of altered expression of genes associated with angiogenesis and vascular development in PV adipocytes, we performed an in vitro angiogenesis assay. Conditioned medium from PV and SQ adipocytes was applied to eGFP-expressing HUVEC seeded in matrigel, and the cells were visualized 5 and 18 h later. Exposure to conditioned medium from PV adipocytes for 5 h led to enhanced tube formation compared with control, and the effects were even more dramatic after 18 h of exposure. In contrast, conditioned medium from SQ adipocytes, at the concentration tested in our assay, produced only a modest angiogenic response at the 5 h time point only (Fig. 7, A and B). These findings point to a differential role of PV adipocytes in promoting angiogenesis via paracrine mechanisms.
Fig. 7.
Increased endothelial tube formation in response to CM (7 day) from PV adipocytes. A: representative microscopic images depicting endothelial tube formation in response to CM from SQ and PV adipocytes, differentiated in vitro for 7 days. CM were collected from equivalent numbers of PV or SQ adipocytes, as determined by cellular protein levels. Tube formation was examined at 5 h (top) and 18 h (bottom) following exposure to CM. We used an image analysis program to measured tube length. Note that endothelial tube structures are maintained for at least 18 h after treatment with CM from PV adipocytes. B: tube formation at 5 h time is quantified as % area referenced to control (no CM). Values are mean of 3 independent experiments ± SE, P < 0.05.
Because mouse PVAT from the thoracic aorta was shown to exhibit a gene expression pattern similar to that of interscapular brown adipose tissue (BAT) (10), we also examined expression of brown adipocyte-related genes in human PV adipocytes compared with human SQ adipocytes, which are prototypical white adipocytes. We previously showed by RT-qPCR analysis that expression of UCP-1, a gene highly enriched in BAT, was expressed at similar levels in human PV and SQ adipocytes, whereas expression of PRDM16, a master regulator of brown adipocyte differentiation, and PGC1α, another brown adipocyte marker, was higher in the PV adipocytes (8). In the present study, none of the genes recently reported to be highly enriched in classical BAT of mice (i.e., Elovl3, Slc27a2, Cox7a1, Cpt1b, Kng2, Acot11, Cidea, Dio2, PRDM16) show any statistically significant difference in expression between SQ and PV adipocytes. Interestingly, TCF21, a selective white adipose tissue marker (43), is actually significantly overexpressed in the PV compared with SQ adipocytes, while leptin, another white adipose tissue marker, is expressed at a much lower level in PV compared with SQ adipocytes. Together, these gene expression characteristics are consistent with the conclusion that human coronary artery PV adipocytes are a highly unique subtype of white adipocytes.
DISCUSSION
Our global gene expression analyses indicate that human SQ and PV adipocytes exhibit distinct biological characteristics that map to both early developmental differences and cardiovascular biology-related functions. The early developmental differences predict that PV and SQ adipocytes originate from distinct precursor cells (12, 19). The findings that PV adipocytes constitutively overexpress genes associated with the control of vascular morphology, inflammation, and hemostasis are consistent with the purported role of this adipose depot in regulating vascular pathophysiology. Functionally, PV adipocytes exhibit increased capacity to attract immune cells and to induce angiogenesis, both of which are observed in atherosclerotic arteries. Together, these findings support a role for PV adipocytes in both vascular homeostasis and the pathogenesis of atherosclerosis.
Until recently, PVAT has been virtually ignored in human and animal studies of cardiovascular disease. However, clues have emerged suggesting that PVAT may be an important contributor to disease pathogenesis. First, in humans, PVAT anatomically colocalizes with atherosclerotic plaques and regions of vascular calcification (22, 25). Second, PVAT surrounding atherosclerotic human blood vessels exhibits a heightened state of inflammation, both in terms of cellular infiltration and proinflammatory gene expression (2, 14, 16, 26). In studies in rodent models fed high-fat diets, PVAT accumulates at sites prone to development of atherosclerosis and secretes potent chemokines that attract monocytes and T cells to the adventitial interface (28). Also, mechanical injury rapidly induces inflammation and perturbs adipokine gene expression profiles in PVAT in mice and pigs (14). These observations suggest that PVAT may contribute to a variety of cardiovascular diseases, including atherosclerosis and postangioplasty restenosis. The data provided in this study identify differentially expressed genes in PV adipocytes that are likely to be mechanistically linked to the observational studies and constitute potential therapeutic targets to modulate “outside-in” signaling in vascular disease.
To identify differentially expressed genes, we isolated adipose depot-specific preadipocytes, expanded them in vitro, and induced in vitro differentiation under well-controlled culture conditions. This approach offers several advantages over studying freshly isolated adipocytes removed from their respective depots. First, freshly isolated adipocytes are mechanically fragile, making it difficult to isolate and maintain equivalent numbers of living cells for comparison purposes. Moreover, their viability diminishes rapidly ex vivo, which compromises collection of conditioned media needed for functional correlations. Third, adipose depots are composed of adipocytes that span a broad spectrum of ages; in vitro differentiation synchronizes the adipocytes and eliminates this important variable, which can influence global gene expression patterns (17, 35). Finally, the process of collagenase isolation acutely changes the gene expression profile of isolated adipocytes and can thus itself complicate data analyses (34, 41). Although in vitro differentiation has obvious limitations (e.g., does not perfectly mimic the in vivo differentiation process), our stringent protocol yields adipocytes that bear striking resemblance to their in vivo counterparts in terms of adipogenic gene expression, adipocytokine expression, and lipid accumulation, thus providing a suitable model to compare adipocyte phenotypes between adipose depots within a donor population (8).
Among the various differentially expressed genes, integrin-α8, which serves as a receptor for fibronectin, vitronectin, tenascin-C fragments, osteopontin, and nephronectin, but not for collagens, is expressed at an elevated levels in PV adipocytes. As shown in Fig. 4 ITGA8 is part of a set of genes overexpressed in PV adipocytes that could modify their interactions with extracellular matrix, adhesion, and integrin signaling, thus having the potential to modify tissue morphology. However, the precise functional significance of elevated expression of integrin-α8 and other adhesion molecules in PV adipocytes remains to be established. Nevertheless, this finding suggests that interaction of PV adipocytes with extracellular matrix components is likely to be distinct from that of the SQ adipocytes.
Complement 7 and complement H are also expressed at much higher levels in PV adipocytes. The complement system is a component of the innate immune system and plays an essential role in eliminating microbes, immune complexes, and damaged cells, in addition to modulating adaptive immune responses. Complement genes are regulated at the transcriptional levels by proinflammatory cytokines such as IL-6 and interferon-γ. Indeed, PV adipocytes express elevated levels of proinflammatory factors, including IL-6, SERPINE2, SERPINE1, and cytosolic PLA2. Elevated expression of genes of the complement system is thus consistent with the proinflammatory phenotype of PV adipocytes.
Inflammation is strongly linked to coronary atherosclerosis and myocardial infarction. In particular, one of the most differentially expressed genes identified in this study was osteoprotegerin, whose levels are positively correlated with atherosclerotic disease in humans (42). Our global gene expression analyses demonstrate that PV adipocytes constitutively express elevated levels of numerous proinflammatory and some anti-inflammatory genes, despite the fact that these cells are derived from healthy individuals with no history of coronary disease. These findings suggest that PV adipocytes can potentially both foster and moderate inflammation, consistent with a homeostatic role in health and disease. Disseminated intravascular infections were common in the preantibiotic era and were typically fatal when involving large arteries, suggesting that the proinflammatory properties of PV adipocytes may have been crucial in controlling such infections in the remote past. In mice fed a short-term high-fat diet, PVAT exhibits upregulated proinflammatory gene expression, suggesting that dietary lipids may, analogous to bacterial membrane lipids (endotoxins), activate innate immune signaling pathways that lead to enhanced vascular inflammation (8). This hypothesis, which may help to explain the increased risk of coronary atherosclerosis associated with obesity, needs to be verified in future investigations.
Increased angiogenesis is also a key feature of advanced atherosclerotic lesions, and it may play an important role in modulating the disease process (3, 31). In particular, adventitial microvessels deliver inflammatory cells to the blood vessel wall, while plaque microvessels may also serve as a conduit for entry of erythrocytes, which are enriched in cholesterol and carry chemokines, into atherosclerotic lesions. In human coronary plaques, cholesterol colocalizes with glycophorin-A, a membrane marker of erythrocytes, thus implicating intraplaque hemorrhage in the progression of atheroma (20, 29). However, the mechanisms responsible for increased angiogenesis in advanced atherosclerotic lesions are unknown. Adipocytes secrete a variety of angiogenic factors (6), and our functional studies with conditioned medium raise the intriguing possibility that PV adipocytes may play a previously unrecognized but important role in promoting angiogenesis in the setting of atherosclerosis. To the best of our knowledge, no previous studies have directly compared the angiogenic potential of differentiated adipocytes from human PVAT and SQAT, although a previous study reported comparable angiogenic potential between human SQAT and visceral omental adipose tissues of severely obese subjects (21). The latter study is not strictly comparable with the present study, however, since omental rather than PV tissues were used, and intact adipose tissues contain numerous other cell types (i.e., stromal cells, mesenchymal stem cells, inflammatory cells) that may have contributed to the angiogeneic responses.
A recent study demonstrated that the gene expression profile of mouse thoracic PVAT is indistinguishable from that of the BAT and is protected from proinflammatory changes during metabolic stress (10). We report that human coronary artery PVAT does not exhibit characteristics of classical BAT; whether PVAT surrounding other vessels in humans, such as the aorta, shares similar characteristics remains to be determined. On the other hand, human coronary artery PV adipocytes could be similar to recently described thermogenic beige adipocytes (43, 45), given their elevated expression of certain beige adipocyte-specific marker genes (8).
Collectively, our findings suggest that human coronary artery PV adipocytes are distinct from SQ adipocytes with respect to gene expression characteristics and functional activities. PV adipocytes exhibit elevated levels of expression of proinflammatory genes and genes associated with angiogenesis, coagulation, and vascular morphology. Under identical adipogenic induction conditions, PV adipocytes achieve a distinct genomic program that is likely reflective of their differential precursor cell programming, their role in regulating cardiac blood vessel morphology and homeostasis, and their potential to mediate outside-in inflammatory and angiogenic signaling. The recent development of catheter-based systems that permit delivery of reagents into the vascular adventitia may soon make it possible to target PV-specific molecular pathways in the treatment of vascular disease in humans (18).
GRANTS
This work is supported in part by the National Institute of Health (NIH) Grants HL-076684, HL-62984 (to N. L. Weintraub), and institutional NIH/NCATS Grant 8UL1TR-000077-04 (B. J. Aronow).
DISCLOSURES
No conflicts of interest, financial or otherwise, are declared by the author(s).
AUTHOR CONTRIBUTIONS
Author contributions: T.K.C. and N.L.W. conception and design of research; T.K.C., W.S.T., Y.T., V.Y.B., D.U., M.G.P., and D.S.W. performed experiments; T.K.C., B.J.A., Y.T., V.Y.B., D.U., D.G.K., and D.Y.H. analyzed data; T.K.C., B.J.A., D.M., A.L.B., S.M.R., D.Y.H., and N.L.W. interpreted results of experiments; T.K.C., B.J.A., and N.L.W. prepared figures; T.K.C., B.J.A., and N.L.W. drafted manuscript; T.K.C., B.J.A., Y.T., V.Y.B., D.U., A.L.B., S.M.R., D.Y.H., and N.L.W. edited and revised manuscript; T.K.C., B.J.A., W.S.T., Y.T., V.Y.B., D.U., A.L.B., M.G.P., D.S.W., S.M.R., D.G.K., D.Y.H., and N.L.W. approved final version of manuscript.
REFERENCES
- 1.Aghamohammadzadeh R, Withers S, Lynch F, Greenstein A, Malik R, Heagerty A. Perivascular adipose tissue from human systemic and coronary vessels: the emergence of a new pharmacotherapeutic target. Br J Pharmacol 165: 670–682, 2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Baker AR, da Silva NF, Quinn DW, Harte AL, Pagano D, Bonser RS, Kumar S, McTernan PG. Human epicardial adipose tissue expresses a pathogenic profile of adipocytokines in patients with cardiovascular disease. Cardiovasc Diabetol 5: 1–7, 2006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Barger AC, Beeuwkes R, 3rd, Lainey LL, Silverman KJ. Hypothesis: vasa vasorum and neovascularization of human coronary arteries. A possible role in the pathophysiology of atherosclerosis. N Engl J Med 310: 175–177, 1984 [DOI] [PubMed] [Google Scholar]
- 4.Boivin A, Brochu G, Marceau S, Marceau P, Hould FS, Tchernof A. Regional differences in adipose tissue metabolism in obese men. Metabolism 56: 533–540, 2007 [DOI] [PubMed] [Google Scholar]
- 5.Briones AM, Nguyen Dinh Cat A, Callera GE, Yogi A, Burger D, He Y, Corrêa JW, Gagnon AM, Gomez-Sanchez CE, Gomez-Sanchez EP, Sorisky A, Ooi TC, Ruzicka M, Burns KD, Touyz RM. Adipocytes produce aldosterone through calcineurin-dependent signaling pathways: implications in diabetes mellitus-associated obesity and vascular dysfunction. Hypertension 59: 1069–1078, 2012 [DOI] [PubMed] [Google Scholar]
- 6.Cao Y. Angiogenesis modulates adipogenesis and obesity. J Clin Invest 117: 2362–2368, 2007 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Chatterjee TK, Idelman G, Blanco V, Blomkalns AL, Piegore MG, Jr, Weintraub DS, Kumar S, Rajsheker S, Manka D, Rudich SM, Tang Y, Hui DY, Bassel-Duby R, Olson EN, Lingrel JB, Ho SM, Weintraub NL. Histone deacetylase 9 is a negative regulator of adipogenic differentiation. J Biol Chem 286: 27836–27847, 2011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Chatterjee TK, Stoll LL, Denning GM, Harrelson A, Blomkalns AL, Idelman G, Rothenberg FG, Neltner B, Romig-Martin SA, Dickson EW, Rudich S, Weintraub NL. Proinflammatory phenotype of perivascular adipocytes: influence of high-fat feeding. Circ Res 104: 541–549, 2009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Drolet R, Bélanger C, Fortier M, Huot C, Mailloux J, Légaré D, Tchernof A. Fat depot-specific impact of visceral obesity on adipocyte adiponectin release in women. Obesity (Silver Spring) 17: 424–30, 2009 [DOI] [PubMed] [Google Scholar]
- 10.Fitzgibbons TP, Kogan S, Aouadi M, Hendricks GM, Straubhaar J, Czech MP. Similarity of mouse perivascular and brown adipose tissues and their resistance to diet-induced inflammation. Am J Physiol Heart Circ Physiol 301: H1425–H1437, 2011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Gao YJ, Takemori K, Su LY, An WS, Lu C, Sharma AM, Lee RM. Perivascular adipose tissue promotes vasoconstriction: the role of superoxide anion. Cardiovasc Res 71: 363–373, 2006 [DOI] [PubMed] [Google Scholar]
- 12.Gesta S, Blüher M, Yamamoto Y, Norris AW, Berndt J, Kralisch S, Boucher J, Lewis C, Kahn CR. Evidence for a role of developmental genes in the origin of obesity and body fat distribution. Proc Natl Acad Sci USA 103: 6676–6681, 2006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Gollasch M. Vasodilator signals from perivascular adipose tissue. Br J Pharmacol 165: 633–642, 2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Henrichot E, Juge-Aubry CE, Pernin A, Pache JC, Velebit V, Dayer JM, Meda P, Chizzolini C, Meier CA. Production of chemokines by perivascular adipose tissue: a role in the pathogenesis of atherosclerosis? Arterioscler Thromb Vasc Biol 25: 2594–2599, 2005 [DOI] [PubMed] [Google Scholar]
- 15.Hou XG, Moser S, Sarr MG, Thompson GB, Que FG, Jensen MD. Visceral and subcutaneous adipose tissue diacylglycerol acyltransferase activity in humans. Obesity (Silver Spring) 17: 1129–1134, 2009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Iacobellis G, Pistilli D, Gucciardo M, Leonetti F, Miraldi F, Brancaccio G, Gallo P, Rosaria C, di Gioia T. Adiponectin expression in human epicardial adipose tissue in vivo is lower in patients with coronary artery disease. Cytokine 29: 251–255, 2005 [DOI] [PubMed] [Google Scholar]
- 17.Jernås M, Palming J, Sjöholm K, Jennische E, Svensson PA, Gabrielsson BG, Levin M, Sjögren A, Rudemo M, Lystig TC, Carlsson B, Carlsson LM, Lönn M. Separation of human adipocytes by size: hypertrophic fat cells display distinct gene expression. FASEB J 20: E832–E839, 2006 [DOI] [PubMed] [Google Scholar]
- 18.Karanian JW, Peregoy JA, Chiesa OA, Murray TL, Ahn C, Pritchard WF. Efficiency of drug delivery to the coronary arteries in swine is dependent on the route of administration: assessment of luminal, intimal, and adventitial coronary artery and venous delivery methods. J Vasc Interv Radiol 21: 1555–1564, 2010 [DOI] [PubMed] [Google Scholar]
- 19.Karastergiou K, Fried SK, Xie H, Lee MJ, Divoux A, Rosencrantz MA, Chang RJ, Smith SR. Distinct developmental signatures of human abdominal and gluteal subcutaneous adipose tissue depots. J Clin Endocrinol Metab 98: 362–371, 2013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kolodgie FD, Gold HK, Burke AP, Fowler DR, Kruth HS, Weber DK, Farb A, Guerrero LJ, Hayase M, Kutys R, Narula J, Finn AV, Virmani R. Intraplaque hemorrhage and progression of coronary atheroma. N Engl J Med 349: 2316–2325, 2003 [DOI] [PubMed] [Google Scholar]
- 21.Ledoux S, Queguiner I, Msika S, Calderari S, Rufat P, Gasc JM, Corvol P, Larger E. Angiogenesis associated with visceral and subcutaneous adipose tissue in severe human obesity. Diabetes 57: 3247–3257, 2008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Lehman SJ, Massaro JM, Schlett CL, O'Donnell CJ, Hoffmann U, Fox CS. Peri-aortic fat, cardiovascular disease risk factors, and aortic calcification: the Framingham Heart Study. Atherosclerosis 210: 656–661, 2010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Lu C, Su LY, Lee RM, Gao YJ. Alterations in perivascular adipose tissue structure and function in hypertension. Eur J Pharmacol 656: 68–73, 2011 [DOI] [PubMed] [Google Scholar]
- 24.Lu C, Su LY, Lee RM, Gao YJ. Mechanisms for perivascular adipose tissue-mediated potentiation of vascular contraction to perivascular neuronal stimulation: the role of adipocyte-derived angiotensin II. Eur J Pharmacol 634: 107–112, 2010 [DOI] [PubMed] [Google Scholar]
- 25.Mahabadi AA, Reinsch N, Lehmann N, Altenbernd J, Kälsch H, Seibel RM, Erbel R, Möhlenkamp S. Association of pericoronary fat volume with atherosclerotic plaque burden in the underlying coronary artery: a segment analysis. Atherosclerosis 211: 195–199, 2010 [DOI] [PubMed] [Google Scholar]
- 26.Mazurek T, Zhang L, Zalewski A, Mannion JD, Diehl JT, Arafat H, Sarov-Blat L, O'Brien S, Keiper EA, Johnson AG, Martin J, Goldstein BJ, Shi Y. Human epicardial adipose tissue is a source of inflammatory mediators. Circulation 108: 2460–2466, 2003 [DOI] [PubMed] [Google Scholar]
- 27.Miao CY, Li ZY. The role of perivascular adipose tissue in vascular smooth muscle cell growth. Br J Pharmacol 165: 643–658, 2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Moos MP, John N, Gräbner R, Nossmann S, Günther B, Vollandt R, Funk CD, Kaiser B, Habenicht AJ. The lamina adventitia is the major site of immune cell accumulation in standard chow-fed apolipoprotein E-deficient mice. Arterioscler Thromb Vasc Biol 25: 2386–2391, 2005 [DOI] [PubMed] [Google Scholar]
- 29.Moreno PR, Purushothaman KR, Sirol M, Levy AP, Fuster V. Neovascularization in human atherosclerosis. Circulation 113: 2245–2252, 2006 [DOI] [PubMed] [Google Scholar]
- 30.Nguyen Dinh Cat A, Briones AM, Callera GE, Yogi A, He Y, Montezano AC, Touyz RM. Adipocyte-derived factors regulate vascular smooth muscle cells through mineralocorticoid and glucocorticoid receptors. Hypertension 58: 479–488, 2011 [DOI] [PubMed] [Google Scholar]
- 31.Okamoto E, Couse T, De Leon H, Vinten-Johansen J, Goodman RB, Scott NA, Wilcox JN. Perivascular inflammation after balloon angioplasty of porcine coronary arteries. Circulation 104: 2228–2235, 2001 [DOI] [PubMed] [Google Scholar]
- 32.Perrini S, Laviola L, Cignarelli A, Melchiorre M, De Stefano F, Caccioppoli C, Natalicchio A, Orlando MR, Garruti G, De Fazio M, Catalano G, Memeo V, Giorgino R, Giorgino F. Fat depot-related differences in gene expression, adiponectin secretion, and insulin action and signalling in human adipocytes differentiated in vitro from precursor stromal cells. Diabetologia 51: 155–164, 2008 [DOI] [PubMed] [Google Scholar]
- 33.Rittig K, Dolderer JH, Balletshofer B, Machann J, Schick F, Meile T, Küper M, Stock UA, Staiger H, Machicao F, Schaller HE, Königsrainer A, Häring HU, Siegel-Axel DI. The secretion pattern of perivascular fat cells is different from that of subcutaneous and visceral fat cells. Diabetologia 55: 1514–1525, 2012 [DOI] [PubMed] [Google Scholar]
- 34.Ruan H, Zarnowski MJ, Cushman SW, Lodish HF. Standard isolation of primary adipose cells from mouse epididymal fat pads induces inflammatory mediators and down-regulates adipocyte genes. J Biol Chem 278: 47585–47593, 2003 [DOI] [PubMed] [Google Scholar]
- 35.Skurk T, Alberti-Huber C, Herder C, Hauner H. Relationship between adipocyte size and adipokine expression and secretion. J Clin Endocrinol Metab 92: 1023–33, 2007 [DOI] [PubMed] [Google Scholar]
- 36.Srinivasan R, Ozhegov E, van den Berg YW, Aronow BJ, Franco RS, Palascak MB, Fallon JT, Ruf W, Versteeg HH, Bogdanov VY. Splice variants of tissue factor promote monocyte-endothelial interactions by triggering the expression of cell adhesion molecules via integrin-mediated signaling. J Thromb Haemost 9: 2087–2096, 2011 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Szasz T, Webb RC. Perivascular adipose tissue: more than just structural support. Clin Sci (Lond) 122: 1–12, 2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Takaoka M, Suzuki H, Shioda S, Sekikawa K, Saito Y, Nagai R, Sata M. Endovascular injury induces rapid phenotypic changes in perivascular adipose tissue. Arterioscler Thromb Vasc Biol 30: 1576–1582, 2010 [DOI] [PubMed] [Google Scholar]
- 39.Tchkonia T, Giorgadze N, Pirtskhalava T, Thomou T, DePonte M, Koo A, Forse RA, Chinnappan D, Martin-Ruiz C, von Zglinicki T, Kirkland JL. Fat depot-specific characteristics are retained in strains derived from single human preadipocytes. Diabetes 55: 2571–25788, 2006 [DOI] [PubMed] [Google Scholar]
- 40.Tchkonia T, Lenburg M, Thomou T, Giorgadze N, Frampton G, Pirtskhalava T, Cartwright A, Cartwright M, Flanagan J, Karagiannides I, Gerry N, Forse RA, Tchoukalova Y, Jensen MD, Pothoulakis C, Kirkland JL. Identification of depot-specific human fat cell progenitors through distinct expression profiles and developmental gene patterns. Am J Physiol Endocrinol Metab 292: E298–E307, 2007 [DOI] [PubMed] [Google Scholar]
- 41.Thompson AC, Nuñez M, Davidson R, Horm T, Schnittker K, Hart MV, Suarez AM, Tsao TS. Mitigation of isolation-associated adipocyte interleukin-6 secretion following rapid dissociation of adipose tissue. J Lipid Res 53: 2797–805, 2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Van Campenhout A, Golledge J. Osteoprotegerin, vascular calcification and atherosclerosis. Atherosclerosis 204: 321–329, 2009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Waldén TB, Hansen IR, Timmons JA, Cannon B, Nedergaard J. Recruited vs. nonrecruited molecular signatures of brown, “brite,” and white adipose tissues. Am J Physiol Endocrinol Metab 302: E19–E31, 2012 [DOI] [PubMed] [Google Scholar]
- 44.Wang P, Xu TY, Guan YF, Su DF, Fan GR, Miao CY. Perivascular adipose tissue-derived visfatin is a vascular smooth muscle cell growth factor: role of nicotinamide mononucleotide. Cardiovasc Res 81: 370–380, 2009 [DOI] [PubMed] [Google Scholar]
- 45.Wu J, Bostrom P, Sparks LM, Ye L, Choi JH, Giang AH, Khandekar M, Virtanen KA, Nuutila P, Schaart G, Huang K, Tu H, van Marken Lichtenbelt WD, Hoeks J, Enerback S, Schrauwen P, Spiegelman B. Beige adipocytes are a distinct type of thermogenic fat cell in mouse and human. Cell 150: 366–376, 2012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Yates-Binder CC, Rodgers M, Jaynes J, Wells A, Bodner RJ, Turner T. An IP-10 (CXCL10)-derived peptide inhibits angiogenesis. PLoS One 7: e40812, 2012 [DOI] [PMC free article] [PubMed] [Google Scholar]