Abstract
Changes in DNA superhelicity during DNA replication are mediated primarily by the activities of DNA helicases and topoisomerases. If these activities are defective, the progression of the replication fork can be hindered or blocked, which can lead to double-strand breaks, elevated recombination in regions of repeated DNA, and genome instability. Hereditary diseases like Werner's and Bloom's Syndromes are caused by defects in DNA helicases, and these diseases are associated with genome instability and carcinogenesis in humans. Here we report a Saccharomyces cerevisiae gene, MGS1 (Maintenance of Genome Stability 1), which encodes a protein belonging to the AAA+ class of ATPases, and whose central region is similar to Escherichia coli RuvB, a Holliday junction branch migration motor protein. The Mgs1 orthologues are highly conserved in prokaryotes and eukaryotes. The Mgs1 protein possesses DNA-dependent ATPase and single-strand DNA annealing activities. An mgs1 deletion mutant has an elevated rate of mitotic recombination, which causes genome instability. The mgs1 mutation is synergistic with a mutation in top3 (encoding topoisomerase III), and the double mutant exhibits severe growth defects and markedly increased genome instability. In contrast to the mgs1 mutation, a mutation in the sgs1 gene encoding a DNA helicase homologous to the Werner and Bloom helicases suppresses both the growth defect and the increased genome instability of the top3 mutant. Therefore, evolutionarily conserved Mgs1 may play a role together with RecQ family helicases and DNA topoisomerases in maintaining proper DNA topology, which is essential for genome stability.
Cells must duplicate their entire genome before dividing to produce daughter cells, but DNA replication does not proceed through all DNA sequences with equal efficiency. Progression of the replication fork is often hindered by exogenous and endogenous DNA damage caused by various agents, including reactive oxygen produced by normal cellular metabolism. In addition, the replication fork is often blocked by proteins bound to DNA and by DNA sequences that form secondary structures such as long inverted repeats. Recent studies suggest that DNA double-strand breaks (DSBs) often occur as a result of replication arrest and/or collapse of the replication fork (1–3).
In E. coli, double-strand breaks are repaired by homologous recombination involving RecBCD and RecA proteins: a DNA 3′ end invades the DNA to form a D-loop as an early intermediate of recombination, and DNA replication is reinitiated from the invading 3′-end (4–7). The D-loop is converted to a Holliday structure, in which two homologous duplex DNA molecules are linked by a single-strand crossover. RuvA, RuvB, and RuvC proteins process Holliday junctions (8–10). The RuvA tetramer binds to Holliday junctions and loads RuvB motor protein onto them. The RuvAB complex drives branch migration in an ATP-dependent manner, and RuvC joins the complex and resolves the junction by its endonucleolytic activity (11–14).
Recent studies have indicated that recombination repairs double-strand breaks and also allows DNA replication to reinitiate at blocked DNA replication forks (2, 3, 7). In the absence of homologous recombination, alternative pathways such as DNA end-joining and error-prone translesion DNA synthesis are used to repair double-strand breaks and to overcome stalled or blocked replication forks, but these pathways can cause gross chromosomal aberration and various types of mutation (15, 16). Therefore, homologous recombination plays a vital role in maintaining genome stability during replication.
Although biochemical studies suggest that Holliday junction-resolving activities are present in eukaryotic cells (17), genes encoding nuclear proteins with such activities have not yet been identified. However, a computational search for proteins homologous to RuvB revealed an ORF in Saccharomyces cerevisiae (YNL218W), whose central region is similar to the corresponding region of RuvB. Subsequent analysis revealed that E. coli possesses an orthologue, ycaJ, with extensive similarity to the S. cerevisiae ORF, which we named MGS1 (Maintenance of Genome Stability 1). A group of highly conserved prokaryotic and eukaryotic genes, the MGS1 orthologues, were identified (39≈45% identity) (Fig. 1). The MGS1 orthologues are likely to be DNA transaction ATPases and may play an important role in DNA metabolism.
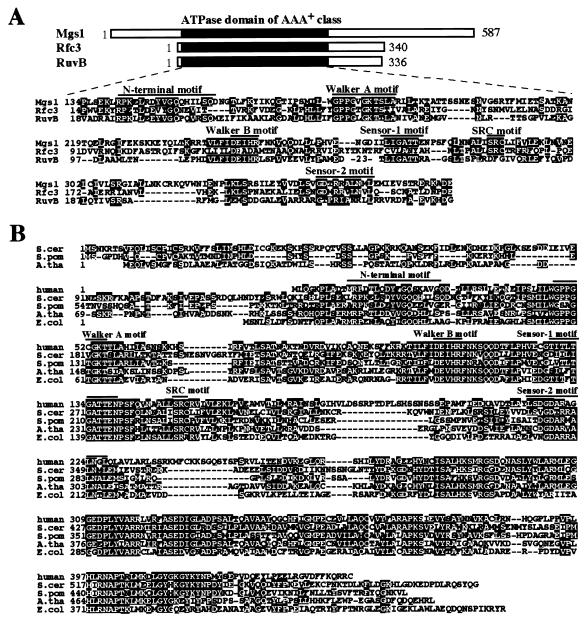
Figure 1.
Mgs1 belongs to the AAA+ class ATPase family, and the Mgs1 orthologues are ubiquitous in prokaryotes and eukaryotes. (A) Alignment of the conserved amino acid sequences of the members of the AAA+ class family (32, 33). Rfc3, S. cerevisiae Rfc3 protein (40). RuvB, E. coli RuvB protein (41). Lines above the sequences indicate the positions of the motifs characteristic of the AAA+ class family. (B) Alignment of amino acid sequences of the Mgs1 orthologues. Human (GenBank accession no. AF218313); S.cer, S. cerevisiae (GenBank accession no. Z71494); S.pom, Schizosaccharomyces pombe (GenBank accession no. T38421); A.tha, Arabidopsis thaliana (GenBank accession no. T00660); E.col, E. coli (GenBank accession no. P45526).
Mutation in E. coli mgs1 reduces the viability of a recA strain by one order of magnitude but does not affect the viability of a wild-type strain (unpublished data). The low viability of strains deficient in recA (≈25% of wild type) (18) may reflect the inability of these cells to bypass a single lethal lesion during DNA replication, which causes the replication fork to collapse. Because the mgs1 mutation reduces the viability of recA mutants, mgs1 may play a role in promoting progression of replication or rescuing an arrested replication fork.
The SGS1 gene is the only member of the RecQ family of DNA helicases in S. cerevisiae (19, 20). The helicase domain of the Sgs1 protein is homologous to the helicase domains of the Werner and Bloom Syndrome proteins (21). A mutation in SGS1 suppresses both the growth defect and the genome instability caused by a mutation in TOP3, which encodes a type-IA topoisomerase (19). Cells from Werner's and Bloom's Syndromes patients and sgs1 and top3 mutants all exhibit genome instability involving increased rates of recombination and chromosomal aberrations (19, 22–24).
In this study, an mgs1 deletion strain of S. cerevisiae was constructed and characterized; the effect of the mutation on genome stability was examined by measuring recombination frequencies during mitosis. mgs1 cells exhibited an increased level of recombination, suggesting that Mgs1 is required for genome stability. The functional relationships among MGS1, SGS1, and TOP3 were also examined. The results provide evidence that these genes are functionally related and probably involved in regulating DNA topology during replication. The role of Mgs1 in maintaining chromosome integrity during replication is discussed.
Materials and Methods
Strains and Plasmids.
A list of S. cerevisiae strains constructed and used in this work is shown in Table 1. Strains derived from W303–1A (25) were used to measure growth rates and recombination in the ribosomal DNA (rDNA) region. For heteroallelic recombination, strains derived from SK1 were used (26). mgs1 deletion strains were constructed as follows. The 3.9-kb HindIII-PstI genomic fragment containing MGS1 was cloned into pUC19 to produce pMG1. The mgs1Δ-generating plasmid was constructed by replacing the AflII-XhoI fragment in the MGS1 coding region with the LEU2 gene. This plasmid was linearized by digestion with FspI and transformed into yeast to generate a strain with deletion of nucleotides +234 to +1,665 of the 1,761-bp MGS1 ORF on chromosome XIV. The plasmids used for generating sgs1Δ (pNS1–27) and top3Δ mutations were kindly provided by T. Enomoto, Tohoku University, Sendai, Japan (27). The 2.4-kb EcoRI fragment containing the 5s rRNA gene, provided by T. Horiuchi, National Institute of Basic Biology, was cloned into pUC19 to produce pRDN1. The URA3 gene was inserted into the unique EcoRV site within DNA of pRDN1 to produce pRDN2. The WD1 strain with the insertion of URA3 in the rDNA cluster was constructed by transformation of the EcoRI-digested pRDN2 into W303–1A. WD1 (MATa) was crossed with W303–1B (MATα) and sporulated, and tetrads were dissected. The Ura+ segregants were designated WD10 (MATa) and WD20 (MATα). A galactose-inducible MGS1 gene was constructed by ligating the NheI-BamHI fragment of pMG1 into p415GAL1 (28), giving pGMG100. The mgs1(K183A) mutant gene was constructed by PCR-mediated site-directed mutagenesis of pMG1, giving pMG2. The NheI-BamHI fragment of pMG2 was cloned into p415GAL1 to produce pGMG183. For overproduction of the Mgs1 protein, the MGS1 gene from pMG1 was cloned into pET-15b (Novagen), a T7 expression vector plasmid, giving pSHR3, which encodes an Mgs1 fusion protein with a 20-aa leader peptide containing six consecutive histidine residues in the N-terminal region. The DNA sequences of recombinant plasmids were confirmed by sequencing the appropriate regions.
Table 1.
Yeast strains
| Strain | Genotype |
|---|---|
| W303-1A | MATa ade2-1 can1-100 his3-11, 15 leu2-3,112 trp1-1 ura3-1 |
| W303-1B | MATα ade2-1 can1-100 his3-11, 15 leu2-3,112 trp1-1 ura3-1 |
| TH10 | mgs1Δ∷Leu2 derivative of W303-1A |
| TH20 | sgs1Δ∷AUR derivative of W303-1A |
| TH30 | top3Δ∷URA3 derivative of W303-1A |
| TH11 | mgs1Δ∷Leu2 sgs1Δ∷AUR derivative of W303-1A |
| TH12 | mgs1Δ∷Leu2 top3Δ∷URA3 derivative of W303-1A |
| TH13 | mgs1Δ∷Leu2 top3Δ∷URA3 sgs1Δ∷AUR derivative of W303-1A |
| WD1 | rDNA∷URA3 derivative of W303-1A |
| WD10 | MATa derivative of WD1 |
| WD20 | MATα derivative of WD1 |
| WD11 | mgs1Δ∷Leu2 sgs1Δ∷AUR derivative of WD10 |
| WD12 | mgs1Δ∷Leu2 top3Δ∷hisG derivative of WD10 |
| WD13 | mgs1Δ∷Leu2 top3Δ∷hisG sgs1Δ∷AUR derivative of WD10 |
| NKY1068 | MATα ho∷LYS2 lys2 ura3 leu2∷hisG his4X-ADE2-his4B ade2∷LK |
| TSK100 | mgs1Δ∷Leu2 derivative of NKY1068 |
| TSK200 | MATa/α ho∷LYS2/ho∷LYS2 lys2/lys2 ura3/ura3 leu2∷hisG/leu2∷hisG his4X/his4B ade2∷LK/ade2∷LK |
| TSK201 | mgs1Δ∷Leu2/mgs1Δ∷Leu2 derivative of TSK200 |
| TSK205 | MGS1/mgs1Δ∷Leu2 derivative of TSK200 |
Determination of Recombination Rates and Frequencies.
Recombination rates were calculated according to the median method described by Lea and Coulson (29). Intrachromosomal recombination was monitored in haploid strains containing his4 heteroallelic sequences, his4B∷ADE2∷his4X (Table 1). Interchromosomal recombination was monitored in diploid strains containing his4B and his4X alleles on different homologous chromosomes (Table 1). For determining recombination rates during mitosis, strains were streaked out on YPAD (yeast extract/peptone/dextrose/adenine sulfate) plates and grown at 30°C. Five colonies for each strain were scraped, suspended in YPAD medium, grown for 12 h, and plated at an appropriate dilution to determine the total cell number on YPAD plates, and the number of His+ or His+ Ade+ recombinants on synthetic complete plates (30) plus 2% glucose minus His [synthetic complete (SC) glucose–His] or SC glucose–His–Ade plates, respectively.
Recombination between the rDNA repeats was determined by the loss of a URA3 gene inserted into the rDNA cluster (Table 1). Strains were streaked out on SC glucose–URA plates and grown at 30°C. Five to nine colonies from each strain were scraped into SC glucose–URA, grown to the stationary phase, and plated at the appropriate dilutions to determine the total cell number (YPAD) and the number of recombinants (SC + 5FOA medium). All strains used to determine the rDNA recombination frequencies are isogenic with WD10 (Table 1).
The Effects of Overproduction of Mgs1 on the Sensitivity to Genotoxoic Agents and Recombination.
Cells harboring p415GAL1 (vector), pGMG100 (MGS1), or pGMG183 (mgs1K183A) were grown overnight in liquid SC glucose–Leu to stationary phase. To assay the effect of Mgs1 overproduction on the sensitivity of the cells to genotoxic agents, 10-fold serial dilutions of cells were spotted onto SC galactose–Leu (induced) or SC glucose–Leu (repressed) plates with 0.005% methyl methanesulfonate (MMS) or 100 mM hydroxyurea (HU). The plates were incubated at 30°C for 3 days. To examine the recombination frequency in an Mgs1-overproducing strain, strain NKY1068 was transformed with p415GAL1, pGMG100, or pGMG183. Five to nine fresh colonies on SC glucose–Leu or SC galactose–Leu plates for each strain were scraped and suspended in liquid SC glucose–Leu medium or SC galactose–Leu medium, respectively. Cells were grown at 30°C for 12 h and plated at an appropriate dilution to determine the total cell number (SC glucose–Leu) and the number of His+ or His+Ade+ recombinants (SC glucose—Leu–His or SC glucose–Leu–His–Ade plates).
Purification of Mgs1 Protein.
E. coli BL21 (DE3) codon+ cells (Stratagene) carrying pSHR3 were grown at 30°C to an OD600 of 0.4–0.6 in 1.5-liter LB broth containing ampicillin (50 μg/ml). Isopropyl-β-d-thiogalactoside was added to a final concentration of 1 mM, and the culture was incubated for 4 h. Cells were collected by centrifugation. All subsequent manipulations were carried out at 4°C. The cells were resuspended in 60 ml of buffer A [20 mM Tris⋅HCl (pH 7.5)/7 mM 2-mercaptoethanol/1 mM EDTA/10% glycerol] containing 50 mM NaCl and disrupted by sonication. The suspension was centrifuged at 39,000 × g for 30 min. Polymin P at pH 7.5 was added to the supernatant to a final concentration of 0.05% (wt/vol). After stirring for 1 h, the suspension was centrifuged at 27,000 × g for 15 min. Proteins were precipitated by ammonium sulfate at 50% saturation, and the precipitate was resuspended in buffer B [50 mM sodium phosphate (pH 7.5)/0.5 M NaCl/7 mM 2-mercaptoethanol] containing 100 mM imidazole. The sample was applied to a nickel–NTA agarose column (Amersham Pharmacia). The column was washed with buffer B, and proteins were eluted with buffer B containing 500 mM imidazole. The eluate containing Mgs1 was dialyzed overnight against buffer A containing 50 mM NaCl. The dialyzed solution was applied to a Hi-trap Heparin column (Amersham Pharmacia) equilibrated with buffer A containing 50 mM NaCl. The bound proteins were eluted with a linear gradient of 50–1,000 mM NaCl in buffer A. The pooled Mgs1 fractions were diluted with buffer C [20 mM Tris⋅HCl (pH 7.0)/7 mM 2-mercaptoethanol/10% glycerol] to 50 mM NaCl. The diluted sample was applied to a cation exchange column, Hi-trap S (Amersham Pharmacia), equilibrated with buffer C containing 50 mM NaCl. Mgs1 protein was eluted with a 20 ml of linear gradient of 50–500 mM NaCl in buffer C. The pooled Mgs1 fractions were dialyzed against storage buffer [30 mM Tris⋅HCl (pH 7.5)/1 mM DTT/100 mM KCl/50% glycerol] and stored at −30°C. Protein concentrations were determined by the Bradford method (Bio-Rad).
ATPase Assay.
ATPase assays were performed at 30°C essentially as described (31). The reaction mixtures (20 μl) containing 20 mM Tris⋅HCl (pH 7.5), 1 mM DTT, 0.01% (wt/vol) BSA, 0.5 mM ATP, the indicated amounts of MgCl2 and NaCl, and 75 μM DNA were prewarmed for 5 min at 30°C and the reactions were started by the addition of Mgs1 (250 nM). Aliquots (5 μl) were taken at the indicated times and added to 5 μl of a stop solution containing 25 mM EDTA and 5 mM ADP.
DNA Annealing Assay.
The indicated amounts of Mgs1 were incubated in a buffer (40 μl) containing 20 mM Tris⋅HCl (pH 7.5), 1 mM DTT, 0.01% (wt/vol) BSA, 5 mM MgCl2, and 25 mM NaCl at 30°C for 5 min. When required, ATP, ADP, or ATPγS was added at a concentration of 1.5 mM. The reactions were started by the addition of boiled 5′-32P-end-labeled pUC19 DNA (5 μM) digested with EcoRI and, 10-μl portions were withdrawn at the indicated times. The reactions were stopped by the addition of 2.5 μl of stop solution containing 2.5% SDS, 4 mg/ml proteinase K, and 100 mM EDTA. Samples were analyzed by electrophoresis on a 0.8% agarose gel, and the reaction products were visualized with a Fuji Bas 1500 image analyzer. As a positive control for the DNA annealing assay, RecA protein (150 nM) was used instead of Mgs1.
Results
Mgs1 Is Highly Conserved in Prokaryotes and Eukaryotes.
S. cerevisiae MGS1 encodes a protein whose central region is similar to the corresponding region of E. coli RuvB (29% identity) (Fig. 1A). Mgs1 is also similar to the clamp loader proteins such as eukaryotic Rfc3 (Fig. 1A). S. cerevisiae Mgs1, E. coli RuvB, and S. cerevisiae Rfc3 share the Walker A and B and sensor I and II motif sequences, which are characteristic of the AAA+ class ATPase family, and they are more similar to each other than to other members of the family (Fig. 1A) (32, 33). Database searching identified Mgs1 orthologues in many eukaryotes and prokaryotes, which are highly conserved in many regions except for a portion of the N-terminal region (Fig. 1B). The S. cerevisiae Mgs1 sequence has 45, 42, 42, and 39% identity in the overlapping region with the human, Arabidopsis thaliana, S. pombe, and E. coli orthologues, respectively (Fig. 1B).
Effects of mgs1 Mutation on Recombination.
To investigate the function of Mgs1 in vivo, an mgs1Δ mutant was constructed by deleting most of the MGS1 coding region. The mgs1Δ strain showed similar sensitivity to MMS, UV light, and HU as the parental wild-type strain (data not shown). To examine the effect of the mgs1Δ mutation on genome stability, the intrachromosomal mitotic recombination rate was measured between heteroallelic DNA sequences by using his4B∷ADE2∷his4X carried on chromosome III (26). In the mgs1Δ mutant, the rate of His+Ade+ recombinants was 3-fold higher than in the wild type, and the rate of His+Ade- recombinants was 5.6-fold higher than in the wild-type strain (Table 2 Upper). In diploids, the interchromosomal recombination rate between chromosomal his4 heteroalleles was 3.7-fold higher in the homozygous mutant, whereas the MGS1/mgs1Δ heterozygote had the same recombination rate as the wild-type strain (Table 2 Lower). The effect of the mgs1 mutation on the stability of the rDNA region was determined by measuring the loss of a URA3 gene inserted into the rDNA cluster. The mgs1 mutation increased the frequency of loss of URA3 by 4-fold (Table 3). However, the drp1 mutation did not affect meiotic recombination (data not shown). These results suggest that the mgs1 mutant has a hyperrecombination phenotype and that Mgs1 is required for genome stability during mitotic cell growth.
Table 2.
Effect of mgs1 mutation on mitotic recombination
| Intrachromosomal recombination rates*
| ||
|---|---|---|
| Genotype | His+ Ade+ (×10−5) | His+ Ade− (×10−5) |
| MGS1 | 4.5 | 1.5 |
| mgs1Δ | 13.3 | 8.3 |
| Interchromosomal recombination rates†
| |
|---|---|
| Genotype | His+ (×10−6) |
| MGS1/MGS1 | 2.5 |
| MGS1/mgs1 | 1.9 |
| mgs1/mgs1 | 9.1 |
Recombination rates were determined between the direct repeat sequences in his4B∷ADE2∷his4X of the haploid strains. His+ rates include both intrachromosomal gene conversion and deletion events. Ade− rates include deletion events between the his4 heteroalleles.
Interchromosomal recombination rates were determined with the heteroalleles his4B and his4X. Recombination rates were calculated according to the median method described by Lea and Coulson (29).
Table 3.
Recombination frequencies in the rDNA region
| Strain | Recombination frequency (×10−4) | Fold increase |
|---|---|---|
| Wild type | 1.1 ± 0.4 | 1.0 |
| mgs1 | 4.4 ± 1.5 | 4.0 |
| sgs1 | 6.2 ± 2.2 | 5.6 |
| top3 | 32 ± 8 | 30 |
| mgs1sgs1 | 9.5 ± 2.9 | 8.6 |
| sgs1top3 | 8.4 ± 2.7 | 7.6 |
| mgs1top3 | ND | ND |
| mgs1top3sgs1 | 42 ± 6.9 | 39 |
The frequency of marker loss was determined as described in Materials and Methods. The data are the averages of five independent measurements. ND, not determined.
Effects of Overexpression of Mgs1 on the Sensitivities of the Cells to Genotoxic Agents and Recombination Frequency.
Overproduction of a wild-type gene product can alter cellular function and provide important information for understanding the biological role of a protein. Mgs1 was overexpressed at a high level from a plasmid carrying a galactose-inducible MGS1 gene. When Mgs1 was induced, the cells became more sensitive to MMS and HU (Fig. 2A) but not UV (data not shown). Overexpression of Mgs1 also increased the spontaneous His+ recombination frequency 5-fold (Fig. 2B). The majority of the increased His+ recombinants were His+Ade− and arose from intrachromatid popout or unequal sister chromatid exchange (Fig. 2B). Treatment of the cells with HU further enhanced the recombination frequency in the Mgs1-overproducing strain (data not shown). However, overexpression of mutant Mgs1 (K183A) protein, which lacks ATPase activity (see below), had no effect on the sensitivity to MMS or HU or on the recombination frequency (Fig. 2); this result suggests that the ATPase activity of Mgs1 is required for these effects. These results indicate that genome instability is caused by either deficiency or overproduction of Mgs1 in yeast and that Mgs1, when overproduced, interferes with DNA repair and replication.
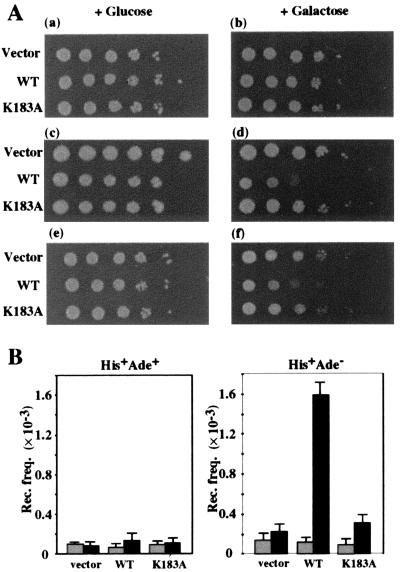
Figure 2.
Overproduction of Mgs1 affects cellular sensitivity to genotoxic agents and recombination frequency. (A) Sensitivity to MMS and HU. Cells harboring empty vector (vector) or a plasmid with the galactose-inducible wild-type MGS1 gene (WT) or mutant mgs1 (K183A) gene (K183A) were grown overnight in liquid SC glucose–Leu medium to stationary phase. Cells were diluted and spotted onto SC galactose–Leu or SC glucose–Leu plates with no drug (a and b), 0.005% MMS (c and d), or 100 mM HU (e and f). The plates were incubated at 30°C for 3 days. (B) Overproduction of Mgs1 increases the intrachromosomal recombination frequency. The recombination frequencies were determined between the intrachromosomal heteroalleles in his4B-ADE2-his4X as described in Materials and Methods. The gray and black columns show the recombination frequencies of the Mgs1-repressed and the Mgs1-induced cells, respectively.
Functional Relationships Among MGS1, TOP3, and SGS1.
Sgs1 is the only member of the RecQ DNA helicase family in S. cerevisiae (19, 20). Sgs1 has homology to the RecQ helicase domain of Wrn protein (21), and mutations in the SGS1 gene increase mitotic recombination and suppress the slow growth phenotype of a top3 mutant (19). The TOP3 gene encodes a type-IA topoisomerase and is highly conserved among prokaryotes and eukaryotes. The top3 mutant exhibits genomic instability, manifested by hyperrecombination between repeated sequences, which is also suppressed by sgs1 (19, 24). Thus, Mgs1, Sgs1, and Top3 are highly conserved from bacteria to mammals and are all involved in maintaining genome stability.
Experiments were carried out to examine the functional relationships among Mgs1, Sgs1, and Top3. In contrast to the sgs1Δ top3Δ strain, the mgs1Δ top3Δ strain had a severe growth defect (Fig. 3 A and B). In addition, the mgs1Δ mutation was a synthetic lethal mutation when combined with top3Δ in an SK1 strain (data not shown). The mgs1Δ top3Δ sgs1Δ triple mutation had a growth rate similar to the top3Δ single mutant (Fig. 3B), indicating that sgs1Δ partially suppressed the severe growth defect of the double mutant.
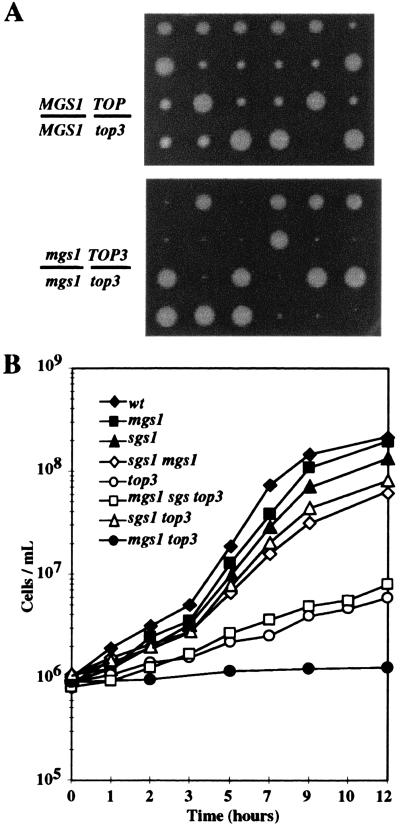
Figure 3.
Effect of mgs1 mutation on growth of sgs1 and top3 strains. (A) Tetrad analysis of a genetic cross between an mgs1 strain and an mgs1 top3 strain (Lower). As a reference, tetrad analysis of a cross between a top3 strain and a TOP3 strain is shown (Upper). The genotypes of the diploids are shown on the Left. All small colonies with either mgs1 (minute) or MGS1 (small) allele were top3, which was confirmed by Ura+ prototropy. The spore clones were grown at 30°C for 4 days. (B) Growth curves of strains with mgs1, sgs1, or top3 mutation, or combined mutations. Yeast cells were grown to the 1 × 107 cells/ml in YPAD, diluted 10-fold, and cultured at 30°C for 12 h. The cell number at each indicated time was counted under a microscope.
The effect of these three mutations on recombination in the rDNA region was examined by measuring the loss of a URA3 marker inserted into the rDNA cluster, as described above. The mgs1, sgs1, or top3 mutations increased the recombination frequency (Table 3). The mgs1Δ top3Δ double mutant lost the URA3 marker at a very high frequency. Haploid cells were formed from diploid cells homozygous for rDNA∷URA3 and heterozygous for mgs1Δ and top3Δ, but the mgs1Δ top3Δ haploid cells failed to form colonies on the SC–URA plates. Because these cells could not be recovered, the exact frequency of URA3 loss could not be measured. These results suggest that the mgs1Δ mutation is synergistic with top3Δ and destabilizes the rDNA locus. The sgs1Δ mgs1Δ top3Δ triple mutation suppressed the increased rDNA recombination frequency of the mgs1Δ top3Δ mutant to a level similar to that of the top3Δ mutant (Table 2); this result agrees with the effect of sgs1Δ on the growth defect of the double mutant described above. Thus, Mgs1 and Sgs1 are functionally related to Top3 and have opposite effects on the Top3 function in growth and genome stability: Mgs1 deficiency exacerbates the phenotype of Top3 deficiency, whereas Sgs1 deficiency ameliorates the phenotype of Top3 deficiency.
Relationships Among MGS1, TOP1, and SGS1.
top1 mutants have a normal growth rate, but sgs1 top1 mutants have a reduced growth rate (34). Deletion or overexpression of MGS1 does not affect the growth rate of top1 mutants (data not shown). Camptothecin (CPT) inhibits DNA replication and targets topoisomerase I during S phase (35, 36). Sensitivity to CPT increased in cells overexpressing Mgs1 and in cells deficient in Sgs1 (Fig. 4), indicating that Mgs1 and Sgs1 have opposite effects on cells exposed to this drug.
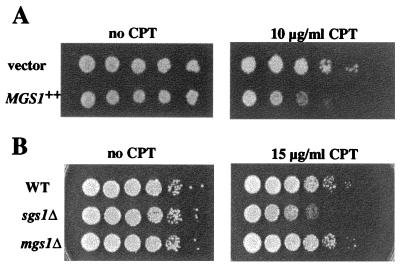
Figure 4.
Effects of sgs1Δ and mgs1Δ mutations on the sensitivity to CPT. (A) Mgs1 overexpression increases the sensitivity of the cells to CPT. Mgs1 was overexpressed as described in the legend of Fig. 2. Cells were 10-fold serially diluted and spotted onto SC galactose–Leu plates buffered with 25 mM Hepes (pH 7.2) and containing 10 μg/ml of CPT (Right) or no CPT as a control (Left). (B) The sgs1Δ mutant is sensitive to CPT. Wild-type, sgs1Δ, and mgs1Δ cultures were 10-fold serially diluted and spotted onto YPAD plates containing 15 μg/ml CPT (Right) or no CPT as a control plates (Left).
Mgs1 Has DNA-Dependent ATPase Activity.
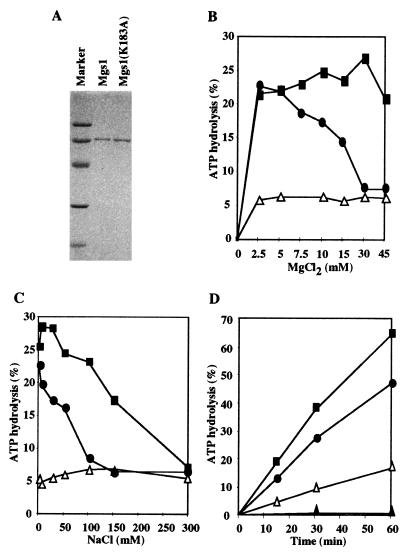
Mgs1 was overproduced as a 67-kDa histidine-tagged protein, purified from E. coli cells and characterized biochemically (Fig. 5A). As predicted for a protein belonging to the AAA+ class, Mgs1 is a DNA-dependent ATPase (Fig. 5 B–D). In the presence of ssDNA, the ATPase was highly active in a broad range of concentrations of MgCl2 and NaCl; however, it was less active in the presence of dsDNA (Fig. 5 B and C). Thus, ssDNA stimulates the ATPase activity more than dsDNA does. A mutant Mgs1(K183A) had no detectable ATPase activity even in the presence of DNA (Fig. 5D). These results suggest that the ATPase activity is an intrinsic property of Mgs1.
Figure 5.
DNA-dependent ATPase activity of Mgs1. (A) Purified Mgs1 proteins were analyzed by 12% SDS/PAGE and stained with Coomassie brilliant blue. (B) MgCl2 titration of ATPase activity at 50 mM NaCl. ATPase activity of Mgs1 was assayed at different MgCl2 concentrations in the absence of DNA (▵), in the presence of ssDNA (■) or dsDNA (●) at 30°C for 30 min. (C) NaCl titration of ATPase activity at 5 mM MgCl2. (D) Time course of ATPase activity in the presence of 7.5 mM MgCl2 and 25 mM NaCl. ATPase activity of wild-type Mgs1 in the absence of DNA (▵), or in the presence of M13mp18 ssDNA (■) or M13mp18 RFI (●). ATPase activity of mutant Mgs1 (K183A) in the presence of ssDNA (▴).
Mgs1 Has DNA Annealing Activity.
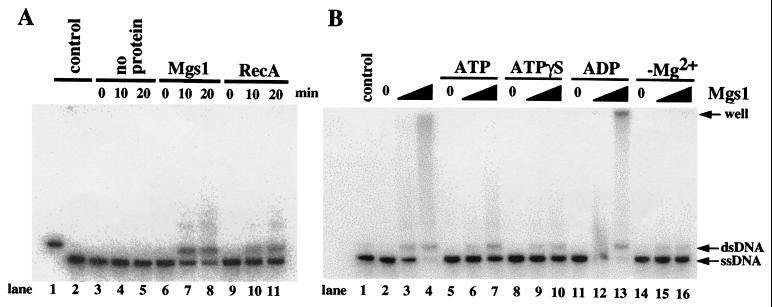
Mgs1 does not have detectable DNA helicase or branch migration activity on various substrates (data not shown). However, Mgs1 promoted annealing of complementary DNA strands in a manner similar to RecA protein (Fig. 6A). This annealing activity required Mg2+ (Fig. 6B, lanes 14–16). ADP slightly enhanced the DNA annealing activity of Mgs1 (Fig. 6B, lanes 11–13), and ATP and ATPγS, a poorly hydrolyzable analogue of ATP, inhibited the activity (Fig. 6B, lanes 5–10). The products of the annealing reaction appeared as a smear on the gel in addition to the unit-length linear duplex and in some cases remained in the well of the gel (Fig. 6B). They are annealing products involving multiple single-strand molecules and are also observed in the reaction with RecA (Fig. 6A) (37). The stimulatory effect of ADP was more pronounced with E. coli Mgs1 than with S. cerevisiae Mgs1 (unpublished data). Therefore, nucleotides appear to function as a molecular switch that regulates the activity of Mgs1 proteins. The observation that Mgs1 and Sgs1 promote reactions that have an opposite effect on DNA winding may be related to the observation that mutants of these two proteins have opposing effects on cellular DNA metabolism in vivo.
Figure 6.
Promotion of annealing of complementary DNA strands by Mgs1. Reactions were performed as described in Material and Methods. (A) Time-course measurements of DNA annealing activity were carried out in the absence of protein (lanes 3–5), or in the presence of 150 μM Mgs1 in the absence of nucleotide cofactor (lanes 6–8) or 150 μM RecA in the presence of 2 mM ATP (lanes 9–11). Lane 1, linear pUC19 dsDNA; lane 2, heat-denatured pUC19 dsDNA. (B) Mgs1 (80 or 200 nM) was incubated with labeled heat-denatured linear pUC19 dsDNA in the absence (lanes 2–4) or presence of ATP (lanes 5–7), ATPγS (lanes 8–10), or ADP (lanes 11–13), or in the absence of Mg2+ (lanes 14–16) at 30°C for 15 min. The resultant samples were analyzed by electrophoresis on a 0.8% agarose gel and fluorography.
Discussion
The Mgs1 orthologues are highly conserved in prokaryotes and eukaryotes and belong to the AAA+ ATPase protein family (32, 33). Indeed, the Mgs1 protein of S. cerevisiae is a DNA-dependent ATPase in vitro (Fig. 5). Mgs1 ATPase activity is strongly enhanced by ssDNA (Fig. 5C) and is likely to be involved in a dynamic aspect of DNA metabolism. RuvB unwinds duplex DNA and promotes branch migration in a complex with RuvA protein, and although Mgs1 has amino acid similarity to RuvB, it has no helicase or branch migration activity with various substrates. Mgs1 does promote Mg2+-dependent annealing of complementary DNA strands (Fig. 6). ADP stimulates this reaction, and ATPγS inhibits it (Fig. 6B). In the presence of ADP, E. coli Mgs1 anneals DNA more efficiently than an identical concentration of E. coli RecA protein in the presence of ATP (unpublished data). These results suggest that Mgs1 efficiently promotes annealing of DNA; in addition, because this activity is regulated by nucleotides, it is likely that it involves dynamic changes in the structure of Mgs1 protein.
The results shown here indicate that Mgs1 is required to maintain genome stability. mgs1 mutants have increased rates of spontaneous intrachromosomal and interchromosomal recombination during mitosis (Table 2). The increased recombination frequency in the rDNA cluster region in the mgs1 mutant is additional evidence of genome instability (Table 3). Therefore, a deficiency of Mgs1 causes genome-wide instability. Overexpression of Mgs1 also causes genome instability, increasing spontaneous intrachromosomal recombination and enhancing sensitivity to genotoxic agents such as MMS, HU, and CPT (Figs. 2 and 4). In E. coli, overproduction of Mgs1 protein has an adverse effect on growth, and the inhibitory effect is particularly strong in recA cells. The viability of mgs1 recA cells is one order of magnitude lower than the viability of recA cells (unpublished data). These results suggest that proper levels of Mgs1 expression are required to maintain genome stability in S. cerevisiae and E. coli.
Previous studies demonstrated that Sgs1 interacts with Top3 (19). The sgs1 mutation slightly decreases the growth rate and increases the recombination rate, and the phenotype of the mgs1 sgs1 double mutant is slightly enhanced relative to that of either single mutant (Fig. 3). The sgs1 mutation decreases the severity of the growth defect of the top3 mutant and reduces its high spontaneous recombination rate (Fig. 3), as reported previously (19). However, the mgs1 mutation almost abolishes the ability of the top3 mutant to grow and drastically destabilizes the rDNA locus (Table 2). The opposing effects of Mgs1 and Sgs1 are also observed in relation to topoisomerase I. Cells become more sensitive to CPT when Mgs1 is overproduced and when Sgs1 is deleted (Fig. 4). Thus, Mgs1 and Sgs1 might generate topologically altered DNA structures that topoisomerases function in concert to resolve. It has been shown that the RecQ family of helicases and topoisomerases functionally interact in E. coli, budding yeast, and humans (19, 20, 38, 39). Therefore, orthologues of Mgs1, RecQ helicase, and DNA topoisomerases exist ubiquitously and may act in a coordinated manner to promote and maintain genome stability during replication.
Our findings suggest two possible roles for Mgs1 in maintaining genome stability. First, Mgs1 may play a role in regulating DNA topology during DNA replication, but its function may be redundant with that of other cellular enzymes. Although Mgs1 deficiency is not lethal, it may cause an imbalance in DNA superhelicity and occasional arrest of DNA replication forks, leading to double-strand breaks and a subsequent increase in recombination. Alternatively, Mgs1 may not normally be involved in DNA replication in growing cells but may play a specific role in DNA replication when the replication fork is blocked by DNA damage, DNA secondary structure, or proteins bound to DNA. Mgs1 may be involved in translesion synthesis past DNA damage as a subunit or cofactor of one of the translesion DNA polymerases. If mgs1 mutants are unable to undergo translesion synthesis (or to replicate past a blocked replication fork), double-strand breaks may occur and enhance recombination. Mgs1 may be involved in the formation or resolution of a four-way junction formed by regression of the replication fork followed by annealing of the newly synthesized strands when progression of replication is blocked (3, 7). It is possible that Mgs1 forms a complex with other protein(s) in a manner similar to that in which RuvB forms a complex with RuvA and catalyzes a reaction that was not revealed in the present study. The present findings have implications for the mechanisms by which DNA topology is regulated during replication and recombination and may provide important clues for understanding the molecular pathology of hereditary diseases characterized by genome instability such as Werner's and Bloom's Syndromes.
Acknowledgments
We thank Drs. T. Kobayashi and T. Horiuchi for suggestions for the rDNA recombination experiments and Drs. P. Sung and T. Enomoto for critical reading of the manuscript. T.H. was supported by a Research Fellowship of the Japan Society for the Promotion of Science for Young Scientists. This work was supported by Grants-in-Aid for Scientific Research (nos. 10044206, 08280102, and 08280103) from the Ministry of Education, Science, Sports and Culture of Japan to H.S.
Abbreviations
- SC
synthetic complete
- ssDNA
single-strand DNA
- dsDNA
double-strand DNA
- CPT
camptothecin
- MMS
methyl methanesulfonate
- HU
hydroxyurea
- rDNA
ribosomal DNA
Footnotes
This paper results from the National Academy of Sciences colloquium, “Links Between Recombination and Replication: Vital Roles of Recombination,” held November 10–12, 2000, in Irvine, CA.
References
- 1.Kuzminov A. Mol Microbiol. 1995;16:373–384. doi: 10.1111/j.1365-2958.1995.tb02403.x. [DOI] [PubMed] [Google Scholar]
- 2.Kowalczykowski S C. Trends Biochem Sci. 2000;25:156–165. doi: 10.1016/s0968-0004(00)01569-3. [DOI] [PubMed] [Google Scholar]
- 3.Michel B. Trends Biochem Sci. 2000;25:173–178. doi: 10.1016/s0968-0004(00)01560-7. [DOI] [PubMed] [Google Scholar]
- 4.Kowalczykowski S C, Dixon D A, Eggleston A K, Lauder S D, Rehrauer W M. Microbiol Rev. 1994;58:401–465. doi: 10.1128/mr.58.3.401-465.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Kogoma T. Microbiol Mol Biol Rev. 1997;61:212–238. doi: 10.1128/mmbr.61.2.212-238.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Kuzminov A. Microbiol Mol Biol Rev. 1999;63:751–813. doi: 10.1128/mmbr.63.4.751-813.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Cox M M, Goodman M F, Kreuzer K N, Sherratt D J, Sandler S J, Marians K J. Nature (London) 2000;404:37–41. doi: 10.1038/35003501. [DOI] [PubMed] [Google Scholar]
- 8.Shinagawa H, Iwasaki H. Trends Biochem Sci. 1996;21:107–111. [PubMed] [Google Scholar]
- 9.West S C. Annu Rev Genet. 1997;31:213–244. doi: 10.1146/annurev.genet.31.1.213. [DOI] [PubMed] [Google Scholar]
- 10.Eggleston A K, Mitchell A H, West S C. Cell. 1997;89:607–617. doi: 10.1016/s0092-8674(00)80242-1. [DOI] [PubMed] [Google Scholar]
- 11.Iwasaki H, Takahagi M, Nakata A, Shinagawa H. Genes Dev. 1992;6:2214–2220. doi: 10.1101/gad.6.11.2214. [DOI] [PubMed] [Google Scholar]
- 12.Iwasaki H, Takahagi M, Shiba T, Nakata A, Shinagawa H. EMBO J. 1991;10:4381–4389. doi: 10.1002/j.1460-2075.1991.tb05016.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Tsaneva I R, Müler B, West S C. Cell. 1992;69:1171–1180. doi: 10.1016/0092-8674(92)90638-s. [DOI] [PubMed] [Google Scholar]
- 14.Dunderdale H J, Benson F E, Parsons C A, Sharples G J, Lloyd R G, West S C. Nature (London) 1991;354:506–510. doi: 10.1038/354506a0. [DOI] [PubMed] [Google Scholar]
- 15.Woodgate R. Genes Dev. 1999;13:2191–2195. doi: 10.1101/gad.13.17.2191. [DOI] [PubMed] [Google Scholar]
- 16.Haber J E. Trends Genet. 2000;16:259–264. doi: 10.1016/s0168-9525(00)02022-9. [DOI] [PubMed] [Google Scholar]
- 17.Constantinou A, Davies A A, West S C. Cell. 2000;104:259–268. doi: 10.1016/s0092-8674(01)00210-0. [DOI] [PubMed] [Google Scholar]
- 18.Cox M M. Genes Cells. 1998;3:65–78. doi: 10.1046/j.1365-2443.1998.00175.x. [DOI] [PubMed] [Google Scholar]
- 19.Gangloff S, McDonald J P, Bendixen C, Arthur L, Rothstein R. Mol Cell Biol. 1994;14:8391–8398. doi: 10.1128/mcb.14.12.8391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Watt P M, Louis E J, Borts R H, Hickson I D. Cell. 1995;81:253–260. doi: 10.1016/0092-8674(95)90335-6. [DOI] [PubMed] [Google Scholar]
- 21.Yu C E, Oshima J, Fu Y H, Wijsman E M, Hisama F, Alisch R, Matthews S, Nakura J, Miki T, Ouais S, et al. Science. 1996;272:258–262. doi: 10.1126/science.272.5259.258. [DOI] [PubMed] [Google Scholar]
- 22.Fukuchi K, Martin G M, Monnat R J J. Proc Natl Acad Sci USA. 1989;86:5893–5897. doi: 10.1073/pnas.86.15.5893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Watt P M, Hickson I D, Borts R H, Louis E J. Genetics. 1996;144:935–945. doi: 10.1093/genetics/144.3.935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Wallis J W, Chrebet G, Brodsky G, Rolfe M, Rothstein R. Cell. 1989;58:409–419. doi: 10.1016/0092-8674(89)90855-6. [DOI] [PubMed] [Google Scholar]
- 25.Thomas B J, Rothstein R. Genetics. 1989;123:725–738. doi: 10.1093/genetics/123.4.725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Bishop D K, Park D, Xu L, Kleckner N. Cell. 1992;69:439–456. doi: 10.1016/0092-8674(92)90446-j. [DOI] [PubMed] [Google Scholar]
- 27.Miyajima A, Seki M, Onoda F, Shiratori M, Odagiri N, Ohta K, Kikuchi Y, Ohno Y, Enomoto T. Mol Cell Biol. 2000;20:6399–6409. doi: 10.1128/mcb.20.17.6399-6409.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Mumberg D, Müler R, Funk M. Nucleic Acids Res. 1994;22:5767–5768. doi: 10.1093/nar/22.25.5767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Lea D E, Coulson C A. J Genet. 1949;49:264–285. doi: 10.1007/BF02986080. [DOI] [PubMed] [Google Scholar]
- 30.Adams A, Gottschling D E, Kaiser C A, Stearns T. Methods in Yeast Genetics: A Cold Spring Harbor Laboratory Course Manual. Plainview, NY: Cold Spring Harbor Lab. Press; 1998. [Google Scholar]
- 31.Hishida T, Iwasaki H, Yagi T, Shinagawa H. J Biol Chem. 1999;274:25335–25342. doi: 10.1074/jbc.274.36.25335. [DOI] [PubMed] [Google Scholar]
- 32.Iwasaki H, Han Y-W, Okamoto T, Ohnishi T, Yoshikawa M, Yamada K, Toh H, Daiyasu H, Ogura T, Shinagawa H. Mol Microbiol. 2000;36:528–538. doi: 10.1046/j.1365-2958.2000.01842.x. [DOI] [PubMed] [Google Scholar]
- 33.Neuwald A F, Aravind L, Spouge J L, Koonin E V. Genome Res. 1999;9:27–43. [PubMed] [Google Scholar]
- 34.Lu J, Mullen J R, Brill S J, Kleff S, Romeo A M, Sternglanz R. Nature (London) 1996;383:678–679. doi: 10.1038/383678a0. [DOI] [PubMed] [Google Scholar]
- 35.Hertzberg R P, Caranfa M J, Hecht S M. Biochemistry. 1989;28:4629–4638. doi: 10.1021/bi00437a018. [DOI] [PubMed] [Google Scholar]
- 36.Hsiang Y H, Hertzberg R, Hecht S, Liu L F. J Biol Chem. 1985;260:14873–14878. [PubMed] [Google Scholar]
- 37.Bryant F R, Menge K L, Nguyen T T. Biochemistry. 1989;28:1062–1069. doi: 10.1021/bi00429a021. [DOI] [PubMed] [Google Scholar]
- 38.Foucault F, Vaury C, Barakat A, Thibout D, Planchon P, Jaulin C, Praz F, Amor-Gueret M. Hum Mol Genet. 1997;6:1427–1434. doi: 10.1093/hmg/6.9.1427. [DOI] [PubMed] [Google Scholar]
- 39.Harmon F G, DiGate R J, Kowalczykowski S C. Mol Cell. 1999;3:611–620. doi: 10.1016/s1097-2765(00)80354-8. [DOI] [PubMed] [Google Scholar]
- 40.Li X, Burgers P M. Proc Natl Acad Sci USA. 1994;91:868–872. doi: 10.1073/pnas.91.3.868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Shinagawa H, Makino K, Amemura M, Kimura S, Iwasaki H, Nakata A. J Bacteriol. 1988;170:4322–4329. doi: 10.1128/jb.170.9.4322-4329.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]