Abstract
In the crowded environment of human cells, folding of nascent polypeptides and refolding of stress-unfolded proteins is error prone. Accumulation of cytotoxic misfolded and aggregated species may cause cell death, tissue loss, degenerative conformational diseases, and aging. Nevertheless, young cells effectively express a network of molecular chaperones and folding enzymes, termed here “the chaperome,” which can prevent formation of potentially harmful misfolded protein conformers and use the energy of adenosine triphosphate (ATP) to rehabilitate already formed toxic aggregates into native functional proteins. In an attempt to extend knowledge of chaperome mechanisms in cellular proteostasis, we performed a meta-analysis of human chaperome using high-throughput proteomic data from 11 immortalized human cell lines. Chaperome polypeptides were about 10 % of total protein mass of human cells, half of which were Hsp90s and Hsp70s. Knowledge of cellular concentrations and ratios among chaperome polypeptides provided a novel basis to understand mechanisms by which the Hsp60, Hsp70, Hsp90, and small heat shock proteins (HSPs), in collaboration with cochaperones and folding enzymes, assist de novo protein folding, import polypeptides into organelles, unfold stress-destabilized toxic conformers, and control the conformal activity of native proteins in the crowded environment of the cell. Proteomic data also provided means to distinguish between stable components of chaperone core machineries and dynamic regulatory cochaperones.
Electronic supplementary material
The online version of this article (doi:10.1007/s12192-013-0413-3) contains supplementary material, which is available to authorized users.
Keywords: SILAC, Mass spectrometry, Proteomics, Molecular chaperones, PPIs, PDIs, Hsp90, Hsp70, Hsp60, CCT, Hsp40, Small HSPs
Introduction
As cellular functions are overwhelmingly carried by proteins, quantitative information from high-throughput proteomics on physiological concentrations and ratios of polypeptides in various compartments of the human cell is of great value to experimental biology and medicine. The relative importance of a given protein for a cellular function may depend on a delicate balance between its activity and its copy number. The rapidly growing proteomic data from various human cells provide a new means to assess degree of generality of particular cellular functions carried by particular proteins. Thus, when a polypeptide is evenly expressed in all cell types and tissues, it is more likely to carry a general housekeeping cellular function than a polypeptide that is only expressed in certain cell types and absent in others. Moreover, knowledge of cellular ratios between various subunits of well-established and less-known functional complexes may refine our understanding of their respective mode of interactions. Whereas equally stoichiometric subunits can serve as stable components of core–protein complexes, substoichiometric subunits cannot and are rather compelled to interact dynamically with core complexes as regulatory subunits. Alternatively, they may differentiate a core–protein complex into several subpopulations with divergent functions.
In the crowded cellular environment, which in eukaryotes reaching protein concentrations of 50–200 mg/ml, folding of nascent polypeptides to their native state and the proper refolding of stress- or mutation-destabilized proteins is a delicate process that is highly error prone (Ellis 2001). Misfolded and aggregated proteins lack their original biological function and, importantly, they also tend to form cytotoxic aggregates that can damage cellular membranes, disturb protein homeostasis, and induce cell death, causing tissue loss in degenerative protein-conformational diseases and aging (Hinault et al. 2006). Detailed knowledge of the true cellular concentrations of potentially toxic polypeptide species and of various cellular machineries that may assist the proper folding of proteins in the different cellular compartments is, therefore, essential to understanding of the causes of proteotoxicity and of the reasons for which the cellular mechanisms that successfully control cellular proteostasis in youth become less effective with age.
Tissieres identified a class of proteins that accumulate in cells during and following heat shock, which were named heat shock proteins (HSPs; Tissieres et al. 1974). Later, the term “molecular chaperone” was used to describe a novel type of cellular function, first demonstrated in the case of bacterial Hsp60, where GroEL from Escherichia coli, assisted by its cochaperone GroES, was shown to promote in the cell proper assembly of phages (Hemmingsen et al. 1988) and of cyanobacterial RubisCO and in vitro to prevent the misfolding and aggregation of artificially unfolded RubisCO and subsequently promote native refolding of active RubisCO oligomers (Goloubinoff et al. 1989a; Goloubinoff et al. 1989b). Several additional molecular chaperones have since been identified as protein-folding aids, in particular, Hsp70, Hsp110, Hsp90, Hsp60, Hsp40, and the small HSPs (Hinault et al. 2006). Noticeably, the attribute “HSP,” which is commonly given to molecular chaperones, is experimentally improper as only a fifth of the human chaperome (33/169 genes) is heat inducible, hence encoding for HSPs in sensu stricto. Moreover, 425 human genes that are not members of the chaperome are significantly induced by heat (Finka et al. 2011).
In eukaryotes, protein synthesis, folding, translocation, and degradation strongly depend on coordinated activity of the chaperone and folding enzyme network (Hinault et al. 2006; Hinault et al. 2011). Hence, in human cells, cotranslational folding of polypeptides is assisted by the ribosome-associated complex (RAC) that includes the J-domain protein zuotin, which is anchored to the large ribosomal subunit RPL31. Zuotin, in turn, recruits cytosolic Hsp70-like molecules, HspA14 and Hsc70, to the nearby extruding nascent chains (Koplin et al. 2010; Albanese et al. 2010). The cytosolic Hsp90 and Hsp70 machineries can also target, import, and mediate native folding of proteins into mitochondria (Neupert and Schatz 1981; Cassina and Casari 2009). In yeast and mammalian cells, post-translational import of cytosolic proteins into endoplasmic reticulum (ER) also involves the J domain-containing Sec63, which is required as a docking site at the translocation pore for luminal Hsp70s to pull and import the cytosolic proteins into the ER lumen by an adenosine triphosphate (ATP)-fueled ratcheting mechanism (Rapoport 2007). Similarly, the post-translational import of many cytosolic proteins into the mitochondria involves TIM14, which J domain serves as a docking site, alongside TIM44, for the mitochondrial Hsp70 (mortalin), enabling the unfolding and post-translational protein import by a mechanism combining entropic pulling and ratcheting (De los Rios et al. 2006; Goloubinoff and De Los Rios 2007). Moreover, the Hsp70/Hsp40/NEF machineries control oligomeric state and thus, the biological activities of many native protein complexes, such as IKapaB (Weiss et al. 2007) or clathrin cages (Schuermann et al. 2008). Cytosolic Hsp70/Hsp40/NEF and Hsp90 machineries are also involved in the activation of native proteins, as in the case of the steroid hormone receptor (Dittmar et al. 1997), or in their inactivation, as in the case of heat-shock transcription factor-1 (Kikis et al. 2010). Noticeably, survival of heat-stressed cells depends on ability of specific heat-induced members of the chaperome to prevent protein misfolding and aggregation during stress and, once the stress is over, to actively unfold stably aggregated proteins that escaped chaperone protection into natively refoldable or protease-degradable polypeptides (Hinault et al. 2006; Hinault et al. 2011). Therefore, Hsp70 with Hsp40 alone (Diamant et al. 2000), but more efficiently in the presence of the Hsp110 cochaperone, can disaggregate and promote the native refolding of stable protein aggregates in vitro and in cells (Shorter 2011; Rampelt et al. 2012). The small HSPs that contain α-crystalline domains can prevent the aggregation of stress-labile proteins and promote their subsequent native refolding by other chaperones, such as Hsp70 and GroEL (Gragerov et al. 1992; Veinger et al. 1998). In animal tissues, onset of proteopathies, such as Alzheimer's disease, Parkinson's disease, amyotrophic lateral sclerosis, and prion and cystic fibrosis, correlate with mutations that destabilize the native conformation of proteins, or with the defective expression, or the stalling of chaperone machineries in stressed cells (Hinault et al. 2006; Coppinger et al. 2012). It is, therefore, of paramount importance to integrate quantitative values to the known qualitative observations and thus, weigh the particular contributions of individual folding, holding, and unfolding chaperone machineries and their regulatory cochaperones to refine the understanding of how the chaperome network can prevent the early onset of protein conformational diseases in youth and to identify reasons for chaperome failure in aging.
Mass spectrometry (MS) analysis coupled with stable isotope labeling with amino acids in cell culture (SILAC) has recently become a method of choice to analyze the human proteome (Schmidt et al. 2011). As polypeptide stoichiometry in enzymes, cofactors, and substrates may unravel molecular mechanisms (Kline and Sussman 2010), we performed a meta-analysis of the human proteome from 11 immortalized cultured cell lines (Geiger et al. 2012), with particular focus on the molecular chaperones and foldases, which together are responsible for proper protein folding, trafficking, translocation, activation, unfolding, disaggregation, and degradation in the different compartments of the cell.
We first developed and validated a method to convert the published intensity-based absolute quantification (iBAQ) numbers for each identified protein into normalized polypeptide copy numbers per cell. In unstressed HeLa cells, the members of the chaperome network contributed 10.4 % to the total protein mass and 8 % to the total polypeptide copy number. Molar ratios between the most prominent members of the Hsp60, Hsp70, small Hsps and Hsp90 chaperone families and their main cochaperones provided new insights on their molecular mechanisms in the cytosol, mitochondria, and ER lumen. Knowledge of physiological amounts and stoichiometry between prominent members of the human chaperome and their clients may contribute to early diagnosis of protein conformational diseases and to design of chaperone-based treatments.
Methods
The raw iBAQ values were obtained for 11 different immortalized human cell lines in triplicates from (http://www.mcponline.org/content/suppl/2012/01/25/M111.014050.DC1/mcp.M111.014050-3.xlsx; Geiger et al. 2012) as a Microsoft Excel spreadsheet. To normalize the iBAQ values, logarithmic values were first converted into linear values. Then, values from each cell (column) were summed up to obtain the total (100 %) iBAQ-derived copy number per cell. In parallel, the iBAQ values were converted into protein mass by dividing the iBAQ numbers by Avogadro's constant and multiplying by the specific molecular weight of each polypeptide, followed by calculation of means, SD, T values, and p values. The values of each column were summed up to obtain the total (100 %) iBAQ-derived protein mass per cell. Then, in each column, the iBAQ-derived protein mass of every individual protein was divided by the total protein mass (Supplementary Figs. 1). The 33 normalized proportional mass and copy number values per cell obtained for each detected protein were than sorted according to their statistical significance. The means, SD, T values, and p values for mass and numeral proportion for each individual polypeptide in the 33 cell samples were calculated using Excel.
We retained for further analysis only 1,497 proteins, which we found to be in statistically significant (p > 0.1) similar polypeptide copy number as well as in similar protein mass per constant cell volume in all the 33 cell samples. To calculate cellular protein crowding and total polypeptide copy number per single HeLa cell, the means and SDs of numeral and mass proportions were separately calculated for each polypeptide based on three samples per cell type and converted to the polypeptide copy number and mass by multiplying proportions with ~2.3 × 109 or 150 pg, respectively, within an average volume of 2,600 μm3 for typical 3-day-old HeLa cells was from Luciani et al. (2001).
Results
We analyzed the raw quantitative values obtained from SILAC-MS for 11 different immortalized human cell lines in triplicates by Geiger et al. (2012). Whereas 11,731 different polypeptides were identified by SILAC as being present in detectable amounts in at least one of the 33 independent samples, only 8,521 were present in detectable amounts in at least one of the triplicate for each of the 11 different cell lines.
Specific protein quantification using SILAC and other methods have shown that a typical cultured cell may contain up to 130–150 pg of total proteins (Van de Water and Olmsted 1980; Volpe and Eremenko 1970) corresponding to 60–150 mg/ml (Albe et al. 1990) and a total of about 2 × 109 polypeptides (Nagaraj et al. 2011). Although iBAQ are generally considered good proxies to quantify the absolute amount of particular polypeptides in a given cell using SILAC (Schwanhausser et al. 2011), here, the direct iBAQ values could not be directly used because they produced nonphysiologically high and variable values, ranging from 1,000 mg/ml and 3.5 × 1010 polypeptides in K562 cells to 4,800 mg/ml and 15 × 1010 polypeptides in HEK293 cells (Supplementary Fig. 1a and b). The direct non-normalized values predicted that only 123 polypeptides would be present in significantly similar copy numbers per constant cellular volume in all the 11 cell lines, with p values <0.1. As the 11 different human cell types likely share more proteins with a similar copy number per constant volume, we next normalized the raw iBAQ copy number values for each of the 8,521 proteins by the sum of all the 11,731 iBAQ polypeptide copy numbers found in each of the 11 cell types and expressed the copy number for each protein as a fraction of the total polypeptide copies per constant cell volume. Thus normalized, 1,524 different polypeptides were found to be present in significantly similar normalized copy numbers per constant cell volume in all the 11 cell lines with p values <0.1 (Supplementary Table 1).
Confirming the centrality of these 1,524 evenly expressed proteins in all the tested cell lines, the sum of their copy numbers was 63 % of the total polypeptide copy number (100 % = 2.3 × 109 for HeLa), while they were encoded by only 6.5 % of the human genome (1,524/23,438, see the Genome Reference Consortium, version GRCh37; Supplementary Table 1). The validity of this list of proteins expressed in significantly constant copy numbers per constant volume in the 11 different cell lines was confirmed by performing the same normalization process using the calculated mass proportion for each polypeptide (Supplementary Table 2). Once each protein mass was normalized by the sum of all the polypeptides masses in a particular cell and expressed as a fraction of the total protein mass per constant cell volume, 1,564 different polypeptides were found to have similar masses in all the 11 cell lines with p values <0.1 (Supplementary Table 2), which represented 70 % of the total protein mass of cells (Fig. 1a; Supplementary Table 2). Remarkably, both protein lists, by copy number and by mass, overwhelmingly overlapped (Fig. 1a), sharing 1,497 proteins. This confirms that various types of human cells likely express at least 1,500 proteins in constant proportions, most of which likely carrying essential housekeeping functions.
Fig. 1.
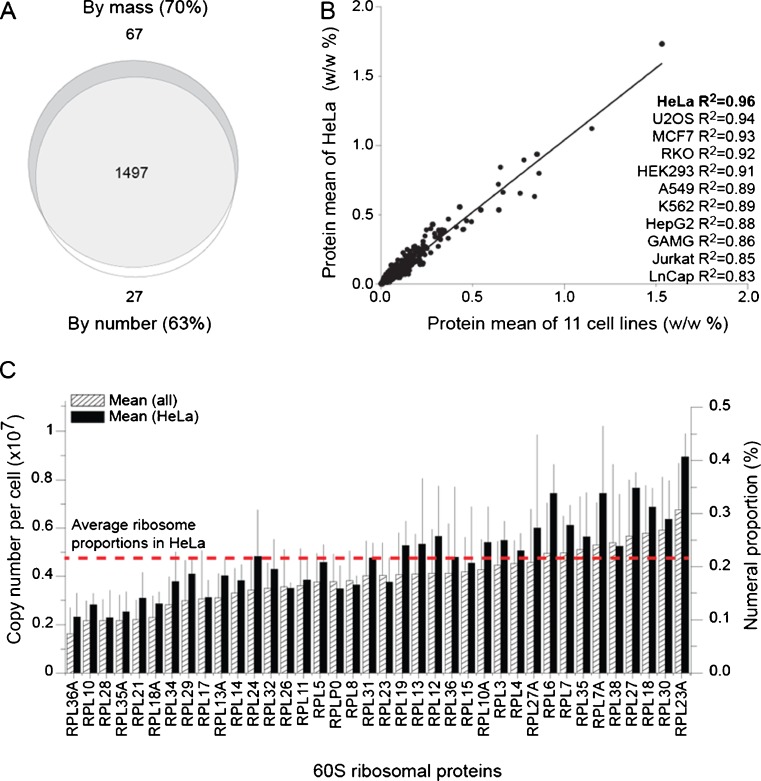
Normalized polypeptide masses and copy numbers in various human cells. a An overlap of 1,497 proteins (light gray), which are significantly similar (with p values < 0.1), both in terms of copy number and proportional mass (deep gray), per constant cell volume, from 33 independent cell measurements in 11 different human cell lines. b The correlation between the mean protein mass proportion (w/w %) of the 1,564 proteins in HeLa cells that are significantly similar in terms of proportional mass per volume against the mean values for the same proteins in the 11 different cell lines. Correlation coefficients for each cell line are listed. c Averaged copy numbers (left vertical axis) of 38 statistically significant 60S ribosomal proteins per HeLa cell from a (plain bars), compared to their numeral proportion (right vertical axis) in the 33 samples from the 11 different cell lines (hatched bars). The red dashed line is the mean value of the 38 ribosomal proteins in HeLa cells. (Geiger et al. 2012)
The relative mass distribution for each polypeptide from the 1,564 protein dataset (by mass) showed best fits to the general means for the HeLa proteins (R2 = 0.96) and the least fits for the LnCap proteins (R2 = 0.86; Fig. 1b). Thus, much of our subsequent analysis focused on the HeLa proteome, as a paradigm for the human proteome.
Classic protein quantification methods have previously shown that an average HeLa cell may contain about 150 pg of total protein (Volpe and Eremenko 1970). Using quantitative precipitation of cytosolic ribosomes and of ribosomal subunits and mass determination by sucrose gradient centrifugation and absorbance measurements of 28S rRNA, Duncan and Hershey independently found that ribosomes contributed 5–10 % to HeLa cells mass and corresponded to 2.6–5.2 × 106 ribosome particles per cell (Duncan and Hershey 1983). These values were recently confirmed by high-throughput SILAC proteomics, with 3.3 x 106 ribosomes, which contributed to about 6 % of the total protein mass and implied the presence of a total of 2 × 109 polypeptides per HeLa cell (Nagaraj et al., 2011). The molecular weight of a ribosome is about 4.3 MDa and the ribosomal protein share is 40 %, corresponding to 3.3 × 106 ribosomes and 9.4 pg of ribosomal protein to the cell. Assuming that 6 % of the total protein mass of the cell corresponded to 9.4 pg of ribosomal protein mass, this implied a total protein mass of 156 pg per HeLa cell, in very good agreement with earlier independent estimates (Volpe and Eremenko 1970).
We next validated our normalization approach for the iBAQ proteomic data from Geiger et al. (2012) by focusing on the proteins from the large ribosomal subunit. Thirty-eight polypeptide subunits of the 60S ribosomal complex belonged to the list of 1,524 significantly evenly expressed proteins per constant cell volume in all the 11 human cell lines (Supplementary Table 1). The normalized mean copy number for these 38 large subunit polypeptides (RPLs), each present in a single copy per ribosome, was 0.175 ± 0.039 % for all the 11 cell types and was slightly higher (0.212 ± 0.036 %) in the case of HeLa cells only (Fig. 1c). As RPL31 was nearest to the mean value, we used it as a proxy to estimate the copy number of ribosome particles per HeLa cell. The dataset showed a mean mass proportion for RPL31 of 0.077 %, relative to the total protein mass in HeLa cells (Supplementary Table 2). Given that the mass of a ribosome is 4,300,000 Da, the net mass of the polypeptides in a ribosome is 1,720,000 Da, and the molecular weight of Rpl31 is 14,436 Da, the normalized data implied that the ribosomes contributed 9.1 % to the total protein mass (0.077 % × 1,720,000/14,630), corresponding to 4.8 × 106 ribosomes. Given that the numeral number proportion of Rpl31 in HeLa cells was slightly higher (0.212 %) than the mean of the 11 cell lines (Fig. 1c), this produced a total of 4.8 × 106 × 100/0.212 = 2.3 × 109 polypeptides per HeLa cell, which was in a good agreement with the previous estimations by Nagaraj et al. (2011). The protein mass and cellular volumes of mammalian cultured cells may decrease with age (Erlinger and Saier 1982; Volpe and Eremenko 1970; Luciani et al. 2001), with cellular volumes of HeLa cells becoming half sized within 1 week of culture (Luciani et al. 2001). Using a mean cellular diameter (Luciani et al. 2001), the volume of 1-week-old HeLa cells should range between 2,600 and 1,100 μm3, which given 150 pg per cell, corresponds to protein crowding of 60–140 mg/ml.
Members of the chaperome are among the most abundant proteins in human cells
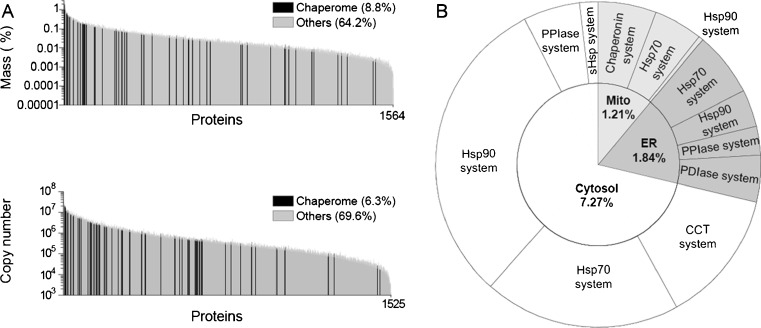
Members of the highly conserved Hsp90 chaperones were found to be the most abundant proteins in the different immortalized human cell lines, averaging alone 2.8 % and, together with the Hsp70s, up to 5.5 % the total protein mass (Supplementary Table 3). Further focusing on the protein homeostasis machineries of the human cells, we established a list of 147 bioinformatically predicted members of the human chaperome according to the criteria previously described (Finka et al. 2011; Kampinga et al. 2009). This list included all the canonical members of the chaperone families Hsp110, Hsp90, Hsp70, Hsp60 and small HSPs, their cochaperones (Hsp10, prefoldins, J-domain proteins, Hsp70-nucleotide exchange factors (NEFs), and Hsp90 cochaperones), and the folding enzymes PPIs and PDIs (Supplementary Table 3). Remarkably, the 147 members of the chaperome contributed up to 7.6 % of total number of polypeptides and up to 10.3 % of the total protein mass in HeLa cells (Fig. 2a, upper panel). As 147 polypeptides were only 1.3 % of 11,731 different SILAC identified human polypeptides, the chaperome polypeptides were thus found to be, on average, eight times more abundant than nonchaperome polypeptides. Moreover, a third of the chaperome (56), principally composed of abundant core members of the ATP-hydrolyzing chaperone machineries Hsp110, Hsp90, Hsp70, and Hsp60, formed two-thirds of the total chaperome mass (8.8 % of the total cellular protein mass) and were consequently 18-fold more abundant than average nonchaperome proteins (Fig. 2a, lower panel). The remaining 91 less abundant chaperome members, which were only 1.5 % of the total cellular mass, were mostly regulatory cochaperones of the Hsp90 and Hsp70 machineries.
Fig. 2.
Chaperones are among the most abundant proteins in HeLa cells. a Distributions of mean mass proportions and SD (in percent of total, upper graph), and copy number (lower graph) of each individual protein present in statistically significantly similar amounts per cell. Chaperome proteins are in black and nonchaperome proteins in gray. Means and SD are from three independent measurements from HeLa cells. b Distribution of mass proportions from total HeLa cell proteins of the main chaperome systems as indicated: Hsp90 with cochaperones, Hsp70 with cochaperones, Hsp60 with Hsp10 cochaperone, the small HSPs, and the foldases PPIs and PDIs, in the cytosol (white), the endoplasmic reticulum (ER, deep gray), and the mitochondria (Mito, light gray). (Geiger et al. 2012)
In the cell or in vitro, the formation of stable protein complexes and the cooperative interactions between the various polypeptides of a protein complex strongly depend on the absolute cellular concentrations of its subunits and on their stoichiometry. Thus, we next examined the copy numbers of various components of known molecular chaperone machineries for which there is experimental evidence that they may form stable complexes regulated by various specific cochaperones acting in concert with foldases and other chaperones to control the unfolding and the native refolding and assembly of a large array of client polypeptides in the different compartments of the human cell. We next categorized, on the basis of bioinformatic and experimental evidence, the 147 members of the human chaperome into 109 cytosolic (and nucleus), 13 mitochondrial, and 25 ER polypeptides (Supplementary Table 3), representing 7.42 %, 1.21 %, and 1.84 % respectively, of the total protein mass of HeLa cells (Fig. 2b).
Within each cellular compartment, the chaperome was further subdivided into five functionally distinct chaperone systems: (1) the Hsp60 chaperonins, including the cytosolic CCTs and mitochondrial GroEL and their cochaperones; (2) the Hsp70 chaperones, including the various Hsp70 and Hsp110 isoforms and their J domain and NEFs cochaperones; (3) the Hsp90 chaperones, including the various Hsp90 isoforms and their dozen cochaperones; (4) the α-crystalline-containing small HSP chaperones; and (5) the foldases, including the PPIs and the PDIs. We next analyzed the stoichiometric relationships in each cellular compartment between the various components of these five categories of chaperone systems.
The distribution of the various chaperone systems differed among compartments. Whereas the Hsp90 chaperone system was predominant in the cytoplasm, its relative abundance compared to the Hsp70 chaperone system decreased in the ER to become quantitatively negligible in the mitochondria. The cytosolic Hsp60 chaperonin system (CCTs) was less abundant than both Hsp90 and Hsp70 systems, inexistent in the ER and as abundant as Hsp70 in the mitochondria. Noticeably, whereas the small HSPs are major components of the chaperome in other types of human cells, such as eye lens (Wistow and Piatigorsky 1988), and in plants and yeast (Nakamoto and Vigh 2007), in the HeLa cells they were least abundant chaperones, to be found only in the cytoplasm (Fig. 2b). Expectedly, the PDIs were more present in the oxidative environment of the ER lumen, where misfolded proteins with misaligned disulfide bonds may need to be reshuffled to reach their native state. The PPIs, some of which known to collaborate closely with Hsp90 (Riggs et al. 2003), were present in significant amounts in the cytoplasm and ER, where Hsp90 was also abundant, in contrast to mitochondria.
Quantitative analysis of the Hsp60 chaperonins in the cytosol and mitochondria
In eukaryotes, CCTs are cytosolic chaperonins of class II that are composed of eight different paralogous subunits CCT1–8 (Yebenes et al. 2011) assembled in two base-to-base associated octameric rings (Cong et al. 2010). The CCTs can interact with a large number of different proteins but predominantly assist to the folding of the cytoskeletal actins and tubulins (Melville et al. 2003; Blacque and Leroux 2006; Llorca et al. 1999; Llorca et al. 2001; Dekker et al. 2008; Dekker et al. 2012). The structure of the CCT complexes predicts that each of the eight paralogous CCT subunits is equimolar in the complex.
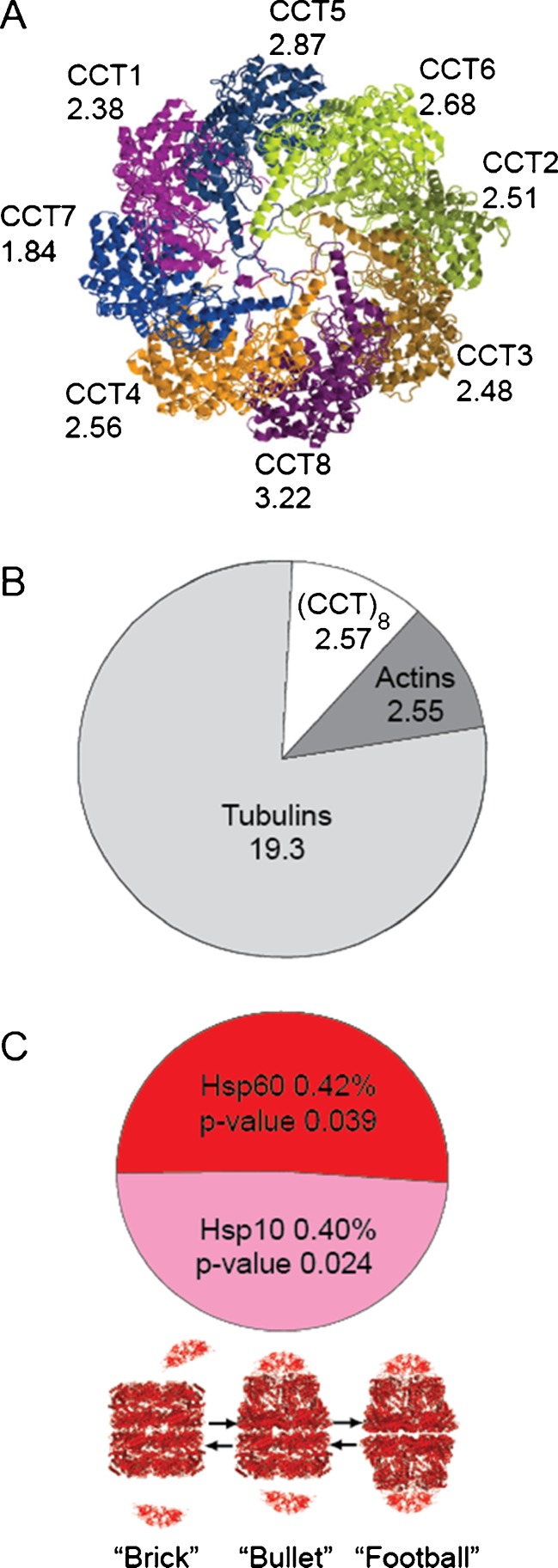
This stoichiometry was remarkably confirmed by the proteomic data in the 11 cell lines. With the exception of CCT7 that was slightly less abundant and CCT8 that was slightly more abundant (with, respectively, 1.84 × 106 and 3.22 × 106 molecules per HeLa cell), the concentration of the remaining subunits CCT1–6 was 2.6 × 106 ± 0.4 (Fig. 3a; Supplementary Table 3). Thus, in addition to the absolute cellular concentrations for the CCT complexes in the human cell (1.3 × 106 CCT16 particles per HeLa cell), high-throughput SILAC proteomics generally confirmed the X-ray evidence about an equimolar stoichiometry between the eight orthologous CCT subunits in the chaperonin complex with an accuracy of ±25 %. Yet, experiments may be needed to exclude the possibility that some chaperonin rings might incorporate a second CCT8 subunit instead of the CCT7. Examination of the cellular stoichiometry between the CCT chaperonins with their two major substrates showed two actin and 16 tubulin molecules per CCT16 complex (Fig. 3b; Supplementary Table 2).
Fig. 3.
Copy numbers in millions and stoichiometry of chaperonins and their substrates. a Average copy number (×106) of each CCT subunit 1–8 per HeLa cell. b Average copy number (×106) of CCT8 rings, actin, and tubulin substrate per HeLa cell. c Average copy number proportions of Hsp60 (red) and Hsp10 (pink) from the 33 independent measures in the 11 different cell types (up). Below Possible mitochondrial chaperonin Hsp60:Hsp10 heterooligomeric complexes allowed by a stoichiometry of 1:1 between Hsp60 and Hsp10 protomers, including free Hsp107, the brick-shaped Hsp6014 core chaperonin, the single capped bullet-shaped Hsp6014Hsp107, and the doubled capped symmetrical football-shaped Hsp107Hsp6014Hsp107 complexes. (Azem et al. 1995; Diamant et al. 1995; Torok et al. 1996)
It has been estimated that the relative mass proportion of cytosolic CCT complex chaperonins range between 0.8 % in HEp-2 cells (Lewis et al. 1992) and 6.4 % in mice testicular cells, supporting our estimation of 1.3 % of CCT complexes in HeLa cell (Supplementary Fig. 1). Initial estimations of purified cytosolic actins and tubulins from different cell types yielded between 0.05 % and 0.2 % of actins (Pollard 1984) and about 2.7 % tubulins (Van de Water and Olmsted 1980). These values are compatible with our estimation of actin and tubulin mass proportion being, respectively, between 0.1 % in HeLa cells and 2.0 % of GaMG (glioma) cells (Supplementary Table 2). Recent protein quantitation of HeLa cells gave an insight of the stoichiometry between the CCT complexes, actins, and tubulins in HeLa cells of 1:2:12 (Nagaraj et al. 2011), demonstrating the robustness of MS. Such stoichiometric relationships suggest that once every chaperonin octameric ring has mediated the folding of one actin and eight tubulins, they may be free to assist the folding of less specific substrates (Dekker et al. 2008). The prefoldin complex, phosducin-like regulatory proteins, and the tubulin folding cofactors have been involved in assembly of tubulins and actin (Lundin et al. 2010). We find here that prefoldin (Siegers et al. 1999) and the tubulin folding auxiliary proteins are between one and two orders of magnitude less abundant than CCT (not shown), suggesting that they may regulate the folding and assembly of tubulin only by way of dynamic interactions with the CCT chaperonins.
The mitochondrial chaperonins, Hsp60 (bacterial GroEL), form two homologous heptameric rings associated back to back in a Hsp6014 complex that become transiently capped by Hsp107 (GroES) homologous heptamers during the folding cycle (Azem et al. 1995; Fig. 3c). Similar to eubacteria (Gross 1996), the copy number values from the 11 different human cell lines showed that the Hsp60 core-chaperonin protomers and the Hsp10 cochaperonin protomers were nearly equimolar (Fig. 3c; Supplementary Table 1; Supplementary Table 3). This stoichiometry strongly favors a chaperonin refolding cycle in which active single-capped bullet-shaped Hsp6014Hsp107 may transiently associate with the free Hsp107 to form transient active doubled-capped symmetrical football-shaped Hsp107Hsp6014Hsp107 complexes (Azem et al. 1995; Diamant et al. 1995; Torok et al. 1996).
The Hsp70 chaperone systems in HeLa cells
Hsp70s belong to a highly conserved family of molecular chaperones present in all the ATP-containing compartments of eukaryotic cells acting as ATP-fueled polypeptide unfolding enzymes and pulling machines. Assisted with J-domain cochaperones and NEFs, Hsp70s can recognize and transiently interact with various protein substrates. Under physiological conditions, Hsp70s may regulate deoligomerization of clathrin cages and triskalion recycling, collaborate with Hsp90 to translocate proteins into the mitochondria and the ER lumen, and mediate targeting of ubiquitinated proteins to the proteasome for degradation (Eisenberg and Greene 2007; Cassina and Casari 2009; Rapoport 2007; Kalia et al. 2011). During and following heat stress, Hsp70, moreover, can use ATP to unfold and solubilize stably misfolded and aggregated proteins (Diamant et al. 2000; Sharma et al. 2010). Recently, Hsp70 was found to act in concert with the Hsp70-like cochaperones, Hsp110s, at mediating the solubilization of stable protein aggregates (Shorter 2011; Rampelt et al. 2012). In the cytosol of HeLa cells, the total copy number of the bioinformatically identified Hsp70s, including Hsp70-like Hsp110s, was 22.2 × 107 (Fig. 4a). Thus, Hsp70s polypeptides were nearly as abundant as Hsp90s. Out of eight different Hsp70s predicted to be expressed in the human cytosol (Hsp12A, HspA14, HspA1A/B, HspA1L, HspA2, HspA6, and HspA8), only two contributed over 99.5 % to the cytosolic Hsp70 pool: about two-thirds being the constitutive HspA8 (Hsc70) and one-third, the heat-inducible HspA1A/B form (Fig. 4a). This confirms that alongside Hsc70, HspA1A/B is involved in housekeeping nearly physiological functions of the different immortalized cells and is not exclusively synthesized to prevent and repair stress-induced conformational damages in proteins.
Fig. 4.
Proteomics of the Hsp70 chaperone machineries. a Average copy number (×107) per HeLa cell of Hsp70-like, Hsp110-like chaperones, J-domain cochaperones, and nucleotide exchange factors (NEFs) in the cytosol (a), the endoplasmic reticulum (b), and the mitochondria (c) as indicated. (Geiger et al. 2012)
J-domain cochaperones specifically target the Hsp70 unfolding machineries to their various native and misfolded protein substrates. Out of the 12 predicted class A and class B soluble J-domain cochaperones in the human cell cytoplasm, DNAJA1, DNAJA2, and DNAJB1 were by far the most abundant Hsp70 cochaperones, suggesting that they are “generalist” cochaperones mainly for HspA8 and HspA1A/B, whereas the other J-domain cochaperones may be more specific, as in the case of auxilin (DNAJC6), which unique known role is to target Hsc70 to clathrin cages for disassembly.
Sequencewise, Hsp110s are Hsp70-like molecules, which are considered to be NEFs, which provide Hsp70 with strong disaggregation activity. Of the three cytosolic Hsp110 proteins, HspH2 (Apg-2) and HspH1 (Hsp105) were the most abundant. Given that Hsc70 and Hsp110 may form stable 1:1 heterodimers (Polier et al. 2008) capable of disaggregating stable protein aggregates (Rampelt et al. 2012; Shorter 2011), the observed stoichiometry in the cytosol suggests that all three types of Hsp110s can form functional disaggregating complexes with 24 % the Hsp70 pool, leaving the remaining 76 % to carry other Hsc70-specific functions, such as deoligomerization of clathrin cages (Sousa and Lafer 2006), inactivating native oligomers (Weiss et al. 2007), and mediating the folding of nascent polypeptides (Koplin et al. 2010). Upon being anchored to the L31 ribosomal subunit near the ribosome exit, the J-domain cochaperone zuotin (DNAJC2) can recruit cytosolic HspA14 (Hsp70L1) and Hsc70 to assist the folding of nascent polypeptides (Fiaux et al. 2010). Examining the cellular amounts showed that whereas zuotin and HspA14 were equimolar, they both were about 45-fold less abundant than the L31 ribosomal anchor (Table 1). Thus, cellular stoichiometry indicates that at any given moment, no more than 2 % of the ribosomes may have a zuotin/HspA14/Hsc70 complex bound at the polypeptide exit site, implying that the RAC complex has highly dynamic interactions with the ribosomes.
Table 1.
Stoichiometry between ribosomes and chaperome complexes. Copy numbers per HeLa cells (×106) of protein RPL31 subunit at the ribosomal exit the ribosomal that is known to anchor the associated complex (RAC) composed of DNAJC2, HspA14, and Hsc70, with corresponding p values from 33 cell measurements in the 11 cell lines according to Geiger et al. (2012). Percentage of ribosomes that may be associated at any given moment with a RAC complex is obtained from the cellular molar ratio between DNAJC2 or HspA14 and RPL31
| Protein | Copy number (×106) | p value | Complex | Association with ribosomes |
|---|---|---|---|---|
| RPL31 | 4.80 | 0.009 | Ribosome | 100 % |
| DNAJC2 (zuotin) | 0.08 | 0.127 | ||
| HspA14 (Hsp70L) | 0.09 | 0.077 | RAC | ~2 % |
The Hsp70 network in the ER was simpler than in the cytosol, with HspA5 (BiP) being the overwhelmingly predominant Hsp70 species, as compared to HspA13. It was nearly ninefold in molar excess over the only lumen-soluble J-domain cochaperone DNAJB11. The ER membrane-anchored J domain-containing Sec63 cochaperone, which is responsible for the post-translational import of a particular subset of cytosolic proteins into the ER lumen (Rapoport 2007), was 30 times less abundant than HspA5, suggesting that at any given moment, less than 4 % of the ER Hsp70s can be recruited to the polypeptide import machinery, allowing the remaining 96 % to be involved in other chaperone duties, such as assisting the proper folding of cotranslationally imported polypeptides or the unfolding of misfolded and aggregated proteins resulting from ER (or UPR) stress (Hetz 2012).
As mortalin (HspA9) is the only mitochondrial Hsp70, the Hsp70 network was even simpler in mitochondria than in the ER. Mortalin was found to be nearly as abundant as HspA5 in the ER lumen. It was 12.3 times more abundant than its only soluble J-domain cochaperone, DNAJA3. Like HspA5, mortalin is an active component of the polypeptide machinery that imports cytosol-synthesized polypeptides with mitochondrial-targeting presequences (Cassina and Casari 2009). Mortalin was 80 times more abundant than the J-domain pore components DNAJC19 (TIM14), 18 times more abundant than the pore-anchoring subunit TIM44, and 25-fold more abundant than the TOM70 dimers. This strongly suggests that, as in the case of the ER lumen, at any given moment, less than 5 % of the mitochondrial Hsp70 can be recruited to the import pore, allowing the remaining 95 % to be involved in other chaperone duties, such as mediating the proper un/folding of imported and of de novo synthesized polypeptides as well as the unfolding of stable protein aggregates (Goloubinoff and De Los Rios 2007).
In the cytosol, the ratio between the sum of Hsp70s and Hsp70-like Hsp110s, relative to their putative J-domain cochaperones was 6.3, as compared to 10:1 in E. coli (Pierpaoli et al. 1997; Gross 1996). In the cytoplasm, Bag2 and Bag3, which were the most abundant NEFs, were 17 times less abundant than their main Hsp70 partners HspA8 and HspA1A/B (Fig. 4). A similar 16-fold ratio was maintained in the mitochondria between mortalin (HspA9) and GRPELE1, in contrast to their respective bacterial homologs, where DnaK is only three times more abundant than GrpE (Pierpaoli et al. 1997; Gross 1996). This is compatible with in vitro observations, indicating that eukaryotic Hsp70 may exchange nucleotides without NEFs more readily than DnaK without GrpE (Rampelt et al. 2011) and implies that eukaryotic Hsp70s are possibly less stringently controlled by NEFs, in general. We found previously using in vitro assays of heat-preaggregated G6PDH that as little as 1:20 GrpE compared to DnaK can mediate half-maximal rates of protein refolding (Ben-Zvi et al. 2004).
Given that the proposed role of J-domain cochaperones is primarily to recognize misfolded or alternatively folded polypeptide substrates and then target the Hsp70-unfolding machineries to these substrates, one can estimate that at any given moment, the cellular pool of J-domain proteins can bind about 0.2 % of the total cellular proteins to be further processed by the Hsp70 chaperone network. In contrast to the J-domain cochaperones recruiting Hsp70s to unfold client proteins, the Hsp70-interacting protein (HIP) recruits Hsp70s to subsequently degrade client proteins by the proteasome (Muller et al. 2012; Kalia et al. 2011). In the cytoplasm, there were about ten times more soluble J-domain cochaperones than CHIP, suggesting that the Hsp70-mediated unfolding for native refolding (rehabilitation) events are more frequent than the Hsp70-mediated unfolding for degradation events, which appears to be a thermodynamically sound equilibrium for cells.
Proteomic analysis of the Hsp90 interactome in HeLa cells
In HeLa cells, cytosolic Hsp90AA1 and Hsp90AB1 were 1.0 % of total polypeptide copy number and 2.3 % of total protein mass (Supplementary Fig. 1). We used the Picard database of cytosolic Hsp90 interactors (http://www.picard.ch/Hsp90Int/index.php, Echeverria et al. 2011) to retrieve a restrictive list of polypeptides for which there are strong direct biochemical (pull down) and genetic evidences that they specifically interact in mammalian cells with Hsp90AA1 and Hsp90AB1 either as cochaperones or as clients (Supplementary Table 3). This list shows that the cytosolic Hsp90s are regulated by over a dozen putative cochaperones, the most studied of which controlling ATP hydrolysis and the chaperone client binding/release activity and modes of collaboration between Hsp90s and Hsp70 chaperones. Much simpler than in the cytosol, the ER Hsp90 (Grp94, Hsp90B) has at most, three putative cochaperones (Marzec et al. 2012) and mitochondrial Hsp90 (TRAP1) has no known cochaperone. The proteomic data showed that all known cytoplasmic Hsp90 cochaperones were by far substoichiometric to the total cytosolic pool of Hsp90AA1 and Hsp90AB1. A substoichiometric molar ratio 1:7 was found for the p23 that inhibits the Hsp90 ATPase activity (Karagoz et al. 2011), 1:9 for valosin-containing protein (VCP), 1:16 for both the Hsp70-interacting protein (HIP, ST13), and the Hsp70/Hsp90-organizing protein (STIP1, HOP), 1:34 for Aha1 that activates the Hsp90 ATPase (AHSA1), 1:41 for the ubiquitin ligase-associated protein CHIP, and 1:46 for the oncogenic cochaperone adaptor CDC37 (Fig. 5a). Such substoichiometric ratios indicate that none of the cochaperones form intrinsic subunits of core Hsp90 dimers and rather associate dynamically to regulate the numerous cellular activities of Hsp90.
Fig. 5.
Proteomics of the Hsp90 chaperone machineries and clients. a Stoichiometric ratios between various acknowledged and putative Hsp90-cytosolic cochaperones and the cytosolic Hsp90s. The Hsp90-interacting Tomm34 and Tomm70 from the outer membrane of the mitochondria are shown in magenta. b Stoichiometric ratios between various acknowledged and putative cytosolic clients and the cytosolic Hsp90s. The mitochondrial Hsp60 (HspD1, GroEL) is shown in magenta. c Stoichiometric distributions in the cytosol between the Hsp90s (black wedges) and other members of the cytosolic chaperome with which they specifically interact (gray wedges). Empty wedges are members of the cytosolic chaperome for which there is no experimental evidence of interactions with cytosolic Hsp90s
Noticeably, the sum of cytosolic Hsp90s molecules was 2.4 × 107 per HeLa cell, which was slightly less than 3.2 × 107 Hsp70 molecules (Fig. 5b). As both Hsp70-interacting proteins, HIP and HOP, were each 16-fold less abundant, at any given moment, no more that 12 % of the Hsp90 molecules could participate in either, HIP- or HOP-dependent Hsp90/Hsp70 supercomplexes. This further indicates that although Hsp70 has often been found associated to Hsp90 complexes, a majority of the Hsp90s and Hsp70s may act independently of one another.
The total copy number of acknowledged Hsp90 clients summed up to about 10 % of the cellular polypeptides (Fig. 5c). This tenfold molar excess of client polypeptides over the chaperone indicated that in the cytosol, each Hsp90 molecule would need to be able to process during its lifetime at least ten polypeptide substrates. Therefore, Hsp90s would need to be more than mere “holding” chaperones and act as molecular “machines” capable of applying multiple consecutive turnovers onto their client proteins (Fig. 5c). Whereas many Hsp90 clients are involved in cellular signaling, the proteomic analysis showed that the most abundant Hsp90 clients were metabolic enzymes, possibly in correlation with the particular metabolic activity of the immortalized cancer cell lines. Noticeably, the list of Hsp90 interactors showed that cytosolic Hsp90 bound to mitochondrial Hsp60. This is possible given that Hsp90 is involved in the import of mitochondrial proteins, the most abundant of which being Hsp60 and mortalin (mtHsp70). In mammalian cells, the cytosolic Hsp70 and Hsp90 can bind the Tom70 and Tom34 that are outer components of the mitochondrial import pore and facilitate the transfer of a set of preproteins to the receptor for ensuing translocation across the mitochondrial membrane (Young et al. 2003; Cassina and Casari 2009; Salaklang et al. 2008). The proteomic data suggests that at any given moment, at most 3 % of the cytosolic Hsp90 can be recruited to the translocation of polypeptides to the mitochondria.
Discussion
Two polypeptide subunits of an active oligomer with a very high binding constant for each other may yet remain mostly unassociated and inactive when extensively diluted. Addressing the biological impact of such principles of the chemical equilibrium to the experimental cellular biology has, thus far, remained limited for lack of knowledge about the true cellular concentrations of the individual polypeptides that compose the various protein machineries and of their substrates and products. The recent developments of quantitative high-throughput proteomics and metabolomics, combined with existing biochemical knowledge about protein interactions and affinities, permit us to extract new information about various biochemical processes in the crowded environment of the cell. Here, we focused on the chaperome network in the various compartments of the human cells, which apparently needs to recruit at least 10 % of the total protein mass to regulate protein homeostasis. Another reason of interest for the human chaperome is cellular protein crowding, thought to be a primary cause for the formation of toxic misfolded species during folding of de novo synthesized and the refolding of stress-unfolded polypeptides, causal to protein conformational diseases, degenerative tissue loss, and aging (Hinault et al. 2011).
Whereas classical quantitation methods could produce estimates of total protein mass per cellular volume, the precise identity and cellular concentrations of the toxic conformers and of the surrounding native proteins involved in the cellular crowding effects remained elusive. The advent of metabolic labeling through SILAC coupled to MS provides essential new information on the specific cellular concentrations of the potentially toxic, aggregation-prone species, such as α-synuclein and its interactors (Jin et al. 2007). Here, this method permitted us to identify and estimate the cellular concentrations of the various chaperome components that can actively prevent, at least in youth, the formation of proteotoxic species, and which defective expression in aging cells or because of aggregation-promoting mutations leads to cell death and conformational diseases.
The raw non-normalized IBAQ data from the 11 different immortalized human cultured cell lines (Geiger et al. 2012) contained unprecedented indirect information on the copy numbers of almost 12,000 different human proteins, covering, both in terms of protein mass and of polypeptide copy number, the majority of the proteins that are apparently needed for these relatively poorly differentiated cancer cells to proliferate indefinitely in culture. Once normalized, either by the sum of polypeptide masses or by the sum of polypeptide copy numbers per cell, at least 1,500 proteins were found to be significantly evenly expressed in all cell types, which together represented about 70 % of the total cellular proteins. Among them were most of the ribosomal proteins, for which the SILAC-MS-derived values perfectly matched earlier independent estimates using classic 2D gel and spectrophotometric quantitation methods of about 3.5 × 106 ribosomes HeLa cell (Duncan and Hershey 1983; Wolf and Schlessinger 1977). This translated into a predicted total protein crowding of about 150 mg/ml per cell, whereas we obtained a reasonably good match of 60 mg/ml from summing up all the normalized individual mass values from the SILAC-detected proteins in HeLa cells.
This permitted us to focus on the members of the chaperome network in the different crowded cellular compartments of the human cells and to use arguments based on the stoichiometry between main core chaperones, their presumed regulatory cochaperones, and some of their identified protein clients to derive important mechanistic conclusions. It should be noted that following normalization, the cellular concentrations that we obtained for the various members of the chaperome referred to the total cellular volume, whereas it has been estimated that ER and mitochondria each occupy approximately about 10 % of the HeLa cell volume (Cooper 2000; Posakony et al. 1977), implying that the real chaperone concentrations in these compartments might be tenfold higher than their general cellular concentrations.
The data from high-throughput quantitative human proteomics (Geiger et al. 2012) lead to the conspicuous observation that an average unstressed human adult comprising an estimated 6–7 kg of intracellular proteins (Wang et al. 2003) may contain 600–700 g of intracellular chaperones, half of which are Hsp90s and Hsp70s. This demonstrates that the human cells allocate a major fraction of their energy and nutrient resources to maintain complex and abundant protein machineries to control cellular protein homeostasis. Thus, Hsp70s are a major hub of the chaperome network that consume ATP to un/fold de novo synthesized proteins, to import proteins into organelles, to unfold potentially toxic stress-destabilized aggregated polypeptides, to locally unfold or dissociate native protein complexes, and also to control the degradation of short-lived native proteins and of stress-damaged polypeptides (Hinault et al. 2011).
Noticeably, whereas on average, the cytosolic Hsp90s were the most abundant human intracellular proteins in the 11 human cell lines, alongside histone H4 and two cytosolic Hsp70s, this does not imply that this is the case in all human cells, especially in highly specialized tissues. Thus, actins and myosins are, by far, the most abundant proteins in striated muscle cells (Lodish 2000), α-crystalline in eye lens cells (Wistow and Piatigorsky 1988), and hemoglobin in erythrocytes (Lew et al. 1995). Yet, Hsp90s, being the most abundant human intracellular proteins in cell cultures, exacerbates our need to understand what precisely the Hsp90 chaperones are doing to their client proteins. At the present state of knowledge of their structure and chaperone functions, it is commonly accepted that Hsp90s can passively prevent the aggregation of artificially unfolded client proteins upon binding them at as yet undetermined regions of the active Hsp90 dimers. However, preventing client aggregation is, by far, not a unique property of the Hsp90s. The small HSPs, Hsp70s, Hsp60s, and Hsp40s, all can prevent protein aggregation, some more effectively than Hsp90. Likewise, when assisted by its various cochaperones, Hsp90s are known to hydrolyze ATP, while causing “structural changes” of unknown nature to their bound client polypeptides (Csermely et al. 1993; Prodromou 2012). It has been suggested, albeit on the basis of scarce experimental evidence, that ATP-fueled movements within the Hsp90 dimers might cause local unfolding within the bound client proteins (Genest et al. 2011). In contrast, strong experimental evidence exists for Hsp70 and Hsp60 acting as ATP-fueled polypeptide unfolding machineries (Sharma et al. 2010; van der Vaart et al. 2004; Lin et al. 2008). Thus, the very high degree of evolutionary conservation among prokaryotic and eukaryotic Hsp90s, combined with the proteomic data showing very high copy numbers of Hsp90s in the cytoplasm and ER, strongly suggests that Hsp90 carries a unique, yet unknown, chaperone function that cannot be carried by other chaperones.
One possibility is that Hsp90s have evolved a unique set of client polypeptides, the folding of which can be assisted only in collaboration with other chaperones, particularly the Hsp70s, which we find to be nearly as abundant in the cytoplasm and may form transient Hsp90:Hop:Hsp70 complexes. In this line, it has been reported that the pathological form of the cystic fibrosis transmembrane conductance regulator (CFTR) is stalled with a Hsc70/Hsp90 complex during protein maturation showing a ratio of 1:2:2, whereas wild type CFTR have a ratio of 5:1:1, suggesting that the Hsc70/Hsp90 chaperone complexes act transiently more on the early folding steps of the mutant CFTR than in the case of wild type CFTR (Coppinger et al. 2012). Additional examples will likely be found once a larger set of bona fide Hsp70 clients will be identified, as is already the case for Hsp90 (http://www.picard.ch/Hsp90Int/index.php, Echeverria et al. 2011), and efficiently stringent in vitro protein refolding assays for Hsp90 and cochaperone with bona fide clients will be developed, as is the case for Hsp70 and Hsp60 chaperones (Sharma et al. 2010; Goloubinoff et al. 1989a).
It has been traditionally considered that chaperones act principally by way of passively preventing the aggregation of their client polypeptides (Fink 1999). When in excess compared to its clients, Hsp90 could indeed act as a “holding” chaperone, preventing aggregation or inappropriate activity, for example, when it inhibits heat shock factors or steroid hormone receptors (Zuehlke and Johnson 2010). In contrast, when a chaperone is substoichiometric to its clients, as we found in the case of Hsp90, this would imply that it should act as a dynamic machine capable of processing polypeptides consecutively, in multiple turnovers. Although the precise nature of the transformations that are carried by the hsp90s onto their bound client proteins is unclear, nor is it known in what particular way Hsp90s act on its clients, the fact still remains that even under nonstressed conditions, the Hsp90s are systematically found in constant high amounts in the various ATP-containing compartments of the 11 different cell lines. Indeed, Hsp90 inhibitors or decreased cellular concentrations of Hsp90s may cause the expression of cryptic mutant phenotypes (Rutherford and Lindquist 1998; Yeyati et al. 2007). In contrast, Hsp90 overexpression can prevent apoptosis in cancer cells (Lurje and Lenz 2009). This indicates that Hsp90s carry unique essential proteostasis and physiological functions, yet in need to be discovered.
High-throughput proteomic data provide new information predicting that substoichiometric subunits likely act as dynamic regulatory cochaperones or differentiate core chaperone machineries into subpopulations with distinct functions. In the case of Hsp70 chaperone network, in vitro chaperone assays show optimal activity for substoichiometric J-domain cochaperones and NEFs (Ben-Zvi et al., 2005) and, therefore, may only interact dynamically and iteratively with the Hsp70 core unfoldases and the substrates during the unfolding/refolding reaction (Sharma et al. 2012).
Nonchaperone human genes are about tenfold less likely to become induced by heat stress than chaperome genes (Finka et al. 2011). Here, we analyzed the chaperome of cultured human cells at physiological growth temperature. Although immortalized cultured cells may be under some mild default stress, possibly expressing higher than usual levels of HSPs, mRNA levels of Hsp genes can still be strongly increased following a mild heat shock (Eymery et al. 2009). Future proteomic analysis should, thus, investigate the effect of heat shock (or fever) on human cells, on the absolute amounts of core chaperones, and on the stoichiometry of their various regulatory cochaperones, ensuing dramatic effects on chaperone activity and specificity. Thus, changes of cellular concentrations and reversible changes in the affinities of J domain and NEFs for Hsp70s could shift substrate binding/release equilibrium: in heat-stressed cells, Hsp70 could transiently become a “holding” chaperone preventing aggregation, and once fever would be over, resume its catalytic polypeptide unfoldase activity, as shown in vitro in the case of DnaK and GrpE (Diamant and Goloubinoff 1998; Sharma et al. 2011; Siegenthaler and Christen 2006).
Moreover, aging mammalian cells or cells expressing aggregation-prone mutant proteins fail to express stress-induced HSPs, in general, and chaperones, in particular (Morimoto 2008). High-throughput proteomic data from young compared to old stressed or unstressed cells could provide important clues about the identity the most limiting components of the chaperome network, which artificial overexpression could restore proteostasis and alleviate symptoms of protein conformational diseases (Hinault et al., 2011).
Electronic supplementary material
Raw polypeptide abundances in 11 imortalized human cell lines. Bar graphs showing high disparity of (A) absolute polypeptide copy numbers per cell and (B) protein concentration, in the 11 different immortalized human cell lines (Geiger et al. 2012). Total polypeptide copy number and SD are from three independent measures. Hatched boxes are 1010 polypeptides and 300 mg/ml per cell as previously reported (Albe et al. 1990). (DOCX 62 kb)
Numeral proportions of 11,731 polypeptides in 11 immortalized human cell lines and individual polypeptide copy number in HeLa cells. The numeral proportions of polypeptides with a p value less than 0.1 are labeled in bold. The polypeptide average copy numbers and SD in HeLa cells are given in a separate sheet (XLSX 7413 kb)
Mass proportions of 11,731 polypeptides in 11 immortalized human cell lines and individual polypeptide mass in HeLa cells. The mass proportions of polypeptides with a p value less than 0.1 are labeled in bold. The average mass and SD (in picograms) of each polypeptide in HeLa cells are given in a separate sheet (XLSX 9413 kb)
List of human 147 chaperome members. Numeral proportions, copy numbers, mass proportions with corresponding p values, and mass of each detected chaperome member in HeLa cells (XLSX 43 kb)
Acknowledgments
This work was supported by the Faculty of Biology and Medicine of Lausanne University of Lausanne and by Grant 31003A-140512/1 from the Swiss National Fund.
Abbreviations
- MDa
Megadaltons
- Pg
Picogram
- ER
Endoplasmic reticulum
- NEF
Nucleotide exchange factor
- Chaperome
Members of the chaperone cochaperone, and the folding enzymes PDIs and PPIs
- PDIs
Peptidyl disulfide isomerases
- PPIs
Peptidylprolyl isomerases
- SILAC
Stable isotope labeling with amino acids in cell culture
- MS
Mass spectrometry
- IBAQ
Intensity-based absolute quantification
- SD
Standard deviation
- Proteostasis
Protein homeostasis
- Proteopathies
Protein conformational diseases
References
- Albanese V, Reissmann S, Frydman J. A ribosome-anchored chaperone network that facilitates eukaryotic ribosome biogenesis. J Cell Biol. 2010;189(1):69–U105. doi: 10.1083/jcb.201001054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Albe KR, Butler MH, Wright BE. Cellular concentrations of enzymes and their substrates. J Theor Biol. 1990;143(2):163–195. doi: 10.1016/S0022-5193(05)80266-8. [DOI] [PubMed] [Google Scholar]
- Azem A, Diamant S, Kessel M, Weiss C, Goloubinoff P. The protein-folding activity of chaperonins correlates with the symmetric GroEL(14)(GroES(7))(2) heterooligomer. Proc Natl Acad Sci USA. 1995;92(26):12021–12025. doi: 10.1073/pnas.92.26.12021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ben-Zvi A, De los Rios P, Dietler G, Goloubinoff P. Active solubilization and refolding of stable protein aggregates by cooperative unfolding action of individual Hsp70 chaperones. J Biol Chem. 2004;279(36):37298–37303. doi: 10.1074/jbc.M405627200. [DOI] [PubMed] [Google Scholar]
- Blacque OE, Leroux MR. Bardet–Biedl syndrome: an emerging pathomechanism of intracellular transport. Cell Mol Life Sci. 2006;63(18):2145–2161. doi: 10.1007/s00018-006-6180-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cassina L, Casari G. The tightly regulated and compartmentalised import, sorting and folding of mitochondrial proteins. Open Biol J. 2009;2:200–221. doi: 10.2174/1874196700902020200. [DOI] [Google Scholar]
- Cong Y, Baker ML, Jakana J, Woolford D, Miller EJ, Reissmann S, Kumar RN, Redding-Johanson AM, Batth TS, Mukhopadhyay A, Ludtke SJ, Frydman J, Chiu W. 4.0-Angstrom resolution cryo-EM structure of the mammalian chaperonin TRiC/CCT reveals its unique subunit arrangement. Proc Natl Acad Sci USA. 2010;107(11):4967–4972. doi: 10.1073/pnas.0913774107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cooper GM. The cell: a molecular approach. 2. Washington: ASM; 2000. [Google Scholar]
- Coppinger JA, Hutt DM, Razvi A, Koulov AV, Pankow S, Yates JR, III, Balch WE. A chaperone trap contributes to the onset of cystic fibrosis. PLoS One. 2012;7(5):e37682. doi: 10.1371/journal.pone.0037682. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Csermely P, Kajtar J, Hollosi M, Jalsovszky G, Holly S, Kahn CR, Gergely P, Soti C, Mihaly K, Somogyi J. ATP induces a conformational change of the 90-kDa heat shock protein (Hsp90) J Biol Chem. 1993;268(3):1901–1907. [PubMed] [Google Scholar]
- De los Rios P, Ben-Zvi A, Slutsky O, Azem A, Goloubinoff P. Hsp70 chaperones accelerate protein translocation and the unfolding of stable protein aggregates by entropic pulling. Proc Natl Acad Sci U S A. 2006;103(16):6166–6171. doi: 10.1073/pnas.0510496103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dekker C, Stirling PC, McCormack EA, Filmore H, Paul A, Brost RL, Costanzo M, Boone C, Leroux MR, Willison KR. The interaction network of the chaperonin CCT. EMBO J. 2008;27(13):1827–1839. doi: 10.1038/emboj.2008.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dekker C, Willison KR, Taylor WR. On the evolutionary origin of the chaperonins. Proteins. 2012;79:1172–1192. doi: 10.1002/prot.22952. [DOI] [PubMed] [Google Scholar]
- Diamant S, Azem A, Weiss C, Goloubinoff P. Effect of free and ATP-bound magnesium and manganese ions on the atpase activity of chaperonin GroEL(14) Biochem-Us. 1995;34(1):273–277. doi: 10.1021/bi00001a033. [DOI] [PubMed] [Google Scholar]
- Diamant S, Ben-Zvi AP, Bukau B, Goloubinoff P. Size-dependent disaggregation of stable protein aggregates by the DnaK chaperone machinery. J Biol Chem. 2000;275(28):21107–21113. doi: 10.1074/jbc.M001293200. [DOI] [PubMed] [Google Scholar]
- Diamant S, Goloubinoff P. Temperature-controlled activity of DnaK-DnaJ-GrpE chaperones: protein-folding arrest and recovery during and after heat shock depends on the substrate protein and the GrpE concentration. Biochem-US. 1998;37(27):9688–9694. doi: 10.1021/bi980338u. [DOI] [PubMed] [Google Scholar]
- Dittmar KD, Demady DR, Stancato LF, Krishna P, Pratt WB. Folding of the glucocorticoid receptor by the heat shock protein (hsp) 90-based chaperone machinery—the role of p23 is to stabilize receptor-hsp90 heterocomplexes formed by hsp90-p60-hsp70. J Biol Chem. 1997;272(34):21213–21220. doi: 10.1074/jbc.272.34.21213. [DOI] [PubMed] [Google Scholar]
- Duncan R, Hershey JWB. Identification and quantitation of levels of protein synthesis initiation factors in crude HeLa cell lysates by two-dimensional polyacrylamide gel electrophoresis. J Biol Chem. 1983;258(11):7228–7235. [PubMed] [Google Scholar]
- Echeverria PC, Bernthaler A, Dupuis P, Mayer B, Picard D. An interaction network predicted from public data as a discovery tool: application to the Hsp90 molecular chaperone machine. PLoS One. 2011;6(10):e26044. doi: 10.1371/journal.pone.0026044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eisenberg E, Greene LE. Multiple roles of auxilin and Hsc70 in clathrin-mediated endocytosis. Traffic. 2007;8(6):640–646. doi: 10.1111/j.1600-0854.2007.00568.x. [DOI] [PubMed] [Google Scholar]
- Ellis RJ. Macromolecular crowding: obvious but underappreciated. Trends Biochem Sci. 2001;26(10):597–604. doi: 10.1016/S0968-0004(01)01938-7. [DOI] [PubMed] [Google Scholar]
- Erlinger S, Saier MH. Decrease in protein-content and cell volume of cultured dog kidney epithelial cells during growth—importance for transport measurements. Vitro Cell Dev B. 1982;18(3):196–202. doi: 10.1007/BF02618571. [DOI] [PubMed] [Google Scholar]
- Eymery A, Horard B, El Atifi-Borel M, Fourel G, Berger F, Vitte AL, Van den Broeck A, Brambilla E, Fournier A, Callanan M, Gazzeri S, Khochbin S, Rousseaux S, Gilson E, Vourc'h C. A transcriptomic analysis of human centromeric and pericentric sequences in normal and tumor cells. Nucleic Acids Res. 2009;37(19):6340–6354. doi: 10.1093/nar/gkp639. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fiaux J, Horst J, Scior A, Preissler S, Koplin A, Bukau B, Deuerling E. Structural analysis of the ribosome-associated complex (RAC) reveals an unusual Hsp70/Hsp40 interaction. J Biol Chem. 2010;285(5):3227–3234. doi: 10.1074/jbc.M109.075804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fink AL. Chaperone-mediated protein folding. Physiol Rev. 1999;79(2):425–449. doi: 10.1152/physrev.1999.79.2.425. [DOI] [PubMed] [Google Scholar]
- Finka A, Mattoo RU, Goloubinoff P. Meta-analysis of heat- and chemically upregulated chaperone genes in plant and human cells. Cell Stress Chaperones. 2011;16(1):15–31. doi: 10.1007/s12192-010-0216-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geiger T, Wehner A, Schaab C, Cox J. Mann M (2012) Comparative proteomic analysis of eleven common cell lines reveals ubiquitous but varying expression of most proteins. Mol Cell Proteomics. 2012;11(3):M111.014050. doi: 10.1074/mcp.M111.014050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Genest O, Hoskins JR, Camberg JL, Doyle SM, Wickner S. Heat shock protein 90 from Escherichia coli collaborates with the DnaK chaperone system in client protein remodeling. Proc Natl Acad Sci U S A. 2011;108(20):8206–8211. doi: 10.1073/pnas.1104703108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goloubinoff P, Christeller JT, Gatenby AA, Lorimer GH. Reconstitution of active dimeric ribulose bisphosphate carboxylase from an unfolded state depends on 2 chaperonin proteins and Mg-Atp. Nature. 1989;342(6252):884–889. doi: 10.1038/342884a0. [DOI] [PubMed] [Google Scholar]
- Goloubinoff P, De Los Rios P. The mechanism of Hsp70 chaperones: (entropic) pulling the models together. Trends Biochem Sci. 2007;32(8):372–380. doi: 10.1016/j.tibs.2007.06.008. [DOI] [PubMed] [Google Scholar]
- Goloubinoff P, Gatenby AA, Lorimer GH. GroE heat-shock proteins promote assembly of foreign prokaryotic ribulose bisphosphate carboxylase oligomers in Escherichia coli. Nature. 1989;337(6202):44–47. doi: 10.1038/337044a0. [DOI] [PubMed] [Google Scholar]
- Gragerov A, Nudler E, Komissarova N, Gaitanaris GA, Gottesman ME, Nikiforov V. Cooperation of GroEL/GroES and DnaK/DnaJ heat shock proteins in preventing protein misfolding in Escherichia coli. Proc Natl Acad Sci U S A. 1992;89(21):10341–10344. doi: 10.1073/pnas.89.21.10341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gross AC. Function and regulation of the heat shock proteins. In: Neidhardt FC, Curtiss R, editors. Escherichia coli and Salmonella: cellular and molecular biology. 2. Washington: ASM; 1996. [Google Scholar]
- Hemmingsen SM, Woolford C, Vandervies SM, Tilly K, Dennis DT, Georgopoulos CP, Hendrix RW, Ellis RJ. Homologous plant and bacterial proteins chaperone oligomeric protein assembly. Nature. 1988;333(6171):330–334. doi: 10.1038/333330a0. [DOI] [PubMed] [Google Scholar]
- Hetz C. The unfolded protein response: controlling cell fate decisions under ER stress and beyond. Nat Rev Mol Cell Bio. 2012;13(2):89–102. doi: 10.1038/nrm3270. [DOI] [PubMed] [Google Scholar]
- Hinault MP, Ben-Zvi A, Goloubinoff P. Chaperones and proteases—cellular fold-controlling factors of proteins in neurodegenerative diseases and aging. J Mol Neurosci. 2006;30(3):249–265. doi: 10.1385/JMN:30:3:249. [DOI] [PubMed] [Google Scholar]
- Hinault MP, Farina-Henriquez-Cuendet A, Goloubinoff P. Molecular chaperones and associated cellular clearance mechanisms against toxic protein conformers in Parkinson's disease. Neurodegener Dis. 2011;8(6):397–412. doi: 10.1159/000324514. [DOI] [PubMed] [Google Scholar]
- Jin JH, Li GJ, Davis J, Zhu D, Wang Y, Pan C, Zhang J. Identification of novel proteins associated with both alpha-synuclein and DJ-1. Mol Cell Proteomics. 2007;6(5):845–859. doi: 10.1074/mcp.M600182-MCP200. [DOI] [PubMed] [Google Scholar]
- Kalia LV, Kalia SK, Chau H, Lozano AM, Hyman BT, McLean PJ (2011) Ubiquitinylation of alpha-synuclein by carboxyl terminus Hsp70-interacting protein (CHIP) is regulated by Bcl-2-associated athanogene 5 (BAG5). PLoS One 6(2). doi:10.1371/journal.pone.0014695 [DOI] [PMC free article] [PubMed]
- Kampinga HH, Hageman J, Vos MJ, Kubota H, Tanguay RM, Bruford EA, Cheetham ME, Chen B, Hightower LE. Guidelines for the nomenclature of the human heat shock proteins. Cell Stress Chaperones. 2009;14(1):105–111. doi: 10.1007/s12192-008-0068-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karagoz GE, Duarte AMS, Ippel H, Uetrecht C, Sinnige T, van Rosmalen M, Hausmann J, Heck AJR, Boelens R, Rudiger SGD. N-terminal domain of human Hsp90 triggers binding to the cochaperone p23. Proc Natl Acad Sci U S A. 2011;108(2):580–585. doi: 10.1073/pnas.1011867108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kikis EA, Gidalevitz T, Morimoto RI. Protein homeostasis in models of aging and age-related conformational disease. Adv Exp Med Biol. 2010;694:138–159. doi: 10.1007/978-1-4419-7002-2_11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kline KG, Sussman MR. Protein quantitation using isotope-assisted mass spectrometry. Annu Rev Biophys. 2010;39:291–308. doi: 10.1146/annurev.biophys.093008.131339. [DOI] [PubMed] [Google Scholar]
- Koplin A, Preissler S, Ilina Y, Koch M, Scior A, Erhardt M, Deuerling E. A dual function for chaperones SSB-RAC and the NAC nascent polypeptide-associated complex on ribosomes. J Cell Biol. 2010;189(1):57–68. doi: 10.1083/jcb.200910074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lew VL, Raftos JE, Sorette M, Bookchin RM, Mohandas N. Generation of normal human red-cell volume, hemoglobin content, and membrane area distributions by birth or regulation. Blood. 1995;86(1):334–341. [PubMed] [Google Scholar]
- Lewis VA, Hynes GM, Dong Z, Saibil H, Willison K. T-Complex polypeptide-1 is a subunit of a heteromeric particle in the eukaryotic cytosol. Nature. 1992;358(6383):249–252. doi: 10.1038/358249a0. [DOI] [PubMed] [Google Scholar]
- Lin Z, Madan D, Rye HS. GroEL stimulates protein folding through forced unfolding. Nat Struct Mol Biol. 2008;15(3):303–311. doi: 10.1038/nsmb.1394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Llorca O, Martin-Benito J, Grantham J, Ritco-Vonsovici M, Willison KR, Carrascosa JL, Valpuesta JM. The ‘sequential allosteric ring’ mechanism in the eukaryotic chaperonin-assisted folding of actin and tubulin. EMBO J. 2001;20(15):4065–4075. doi: 10.1093/emboj/20.15.4065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Llorca O, McCormack EA, Hynes G, Grantham J, Cordell J, Carrascosa JL, Willison KR, Fernandez JJ, Valpuesta JM. Eukaryotic type II chaperonin CCT interacts with actin through specific subunits. Nature. 1999;402(6762):693–696. doi: 10.1038/45294. [DOI] [PubMed] [Google Scholar]
- Lodish HF. Molecular cell biology. 4. New York: W.H. Freeman; 2000. [Google Scholar]
- Luciani AM, Rosi A, Matarrese P, Arancia G, Guidoni L, Viti V. Changes in cell volume and internal sodium concentration in HeLa cells during exponential growth and following ionidamine treatment. Eur J Cell Biol. 2001;80(2):187–195. doi: 10.1078/0171-9335-00102. [DOI] [PubMed] [Google Scholar]
- Lundin VF, Leroux MR, Stirling PC. Quality control of cytoskeletal proteins and human disease. Trends Biochem Sci. 2010;35(5):288–297. doi: 10.1016/j.tibs.2009.12.007. [DOI] [PubMed] [Google Scholar]
- Lurje G, Lenz HJ. EGFR signaling and drug discovery. Oncology-Basel. 2009;77(6):400–410. doi: 10.1159/000279388. [DOI] [PubMed] [Google Scholar]
- Marzec M, Eletto D, Argon Y. GRP94: An Hsp90-like protein specialized for protein folding and quality control in the endoplasmic reticulum. BBA Mol Cell Res. 2012;1823(3):774–787. doi: 10.1016/j.bbamcr.2011.10.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Melville MW, McClellan AJ, Meyer AS, Darveau A, Frydman J. The Hsp70 and TRiC/CCT chaperone systems cooperate in vivo to assemble the von Hippel–Lindau tumor suppressor complex. Mol Cell Biol. 2003;23(9):3141–3151. doi: 10.1128/MCB.23.9.3141-3151.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morimoto RI. Proteotoxic stress and inducible chaperone networks in neurodegenerative disease and aging. Gene Dev. 2008;22(11):1427–1438. doi: 10.1101/gad.1657108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muller P, Ruckova E, Halada P, Coates PJ, Hrstka R, Lane DP, Vojtesek B (2012) C-terminal phosphorylation of Hsp70 and Hsp90 regulates alternate binding to co-chaperones CHIP and HOP to determine cellular protein folding/degradation balances. Oncogene 1-10. doi:http://www.nature.com/onc/journal/vaop/ncurrent/suppinfo/onc2012314s1.html [DOI] [PubMed]
- Nagaraj N, Wisniewski JR, Geiger T, Cox J, Kircher M, Kelso J, Paabo S, Mann M. Deep proteome and transcriptome mapping of a human cancer cell line. Mol Syst Biol. 2011;7:548. doi: 10.1038/msb.2011.81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakamoto H, Vigh L. The small heat shock proteins and their clients. Cell Mol Life Sci. 2007;64(3):294–306. doi: 10.1007/s00018-006-6321-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Neupert W, Schatz G. How proteins are transported into mitochondria. Trends Biochem Sci. 1981;6(1):1–4. doi: 10.1016/0968-0004(81)90002-5. [DOI] [Google Scholar]
- Pierpaoli EV, Sandmeier E, Baici A, Schonfeld HJ, Gisler S, Christen P. The power stroke of the DnaK/DnaJ/GrpE molecular chaperone system. J Mol Biol. 1997;269(5):757–768. doi: 10.1006/jmbi.1997.1072. [DOI] [PubMed] [Google Scholar]
- Polier S, Dragovic Z, Hartl FU, Bracher A. Structural basis for the cooperation of Hsp70 and Hsp110 chaperones in protein folding. Cell. 2008;133(6):1068–1079. doi: 10.1016/j.cell.2008.05.022. [DOI] [PubMed] [Google Scholar]
- Pollard TD. Citation classic—actin and myosin and cell-movement. Life Sci. 1984;30:13–13. [Google Scholar]
- Posakony J, England J, Attardi G. Mitochondrial growth and division during the cell cycle in HeLa cells. J Cell Biol. 1977;74(2):468–491. doi: 10.1083/jcb.74.2.468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prodromou C. The ‘active life’ of Hsp90 complexes. BBA Mol Cell Res. 2012;1823(3):614–623. doi: 10.1016/j.bbamcr.2011.07.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rampelt H, Kirstein-Miles J, Nillegoda NB, Chi K, Scholz SR, Morimoto RI, Bukau B (2012) Metazoan Hsp70 machines use Hsp110 to power protein disaggregation. EMBO J 31 (21):4221-4235. doi:http://www.nature.com/emboj/journal/v31/n21/suppinfo/emboj2012264a_S1.html [DOI] [PMC free article] [PubMed]
- Rampelt H, Mayer M, Bukau B. Nucleotide exchange factors for Hsp70 chaperones. In: Calderwood SK, Prince TL, editors. Molecular chaperones. New York: Humana; 2011. pp. 83–91. [DOI] [PubMed] [Google Scholar]
- Rapoport TA. Transport of proteins in and out of the endoplasmic reticulum. FASEB J. 2007;21(5):A207–A207. [Google Scholar]
- Riggs DL, Roberts PJ, Chirillo SC, Cheung-Flynn J, Prapapanich V, Ratajczak T, Gaber R, Picard D, Smith DF. The Hsp90-binding peptidylprolyl isomerase FKBP52 potentiates glucocorticoid signaling in vivo. EMBO J. 2003;22(5):1158–1167. doi: 10.1093/emboj/cdg108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rutherford SL, Lindquist S. Hsp90 as a capacitor for morphological evolution. Nature. 1998;396(6709):336–342. doi: 10.1038/24550. [DOI] [PubMed] [Google Scholar]
- Salaklang J, Steitz B, Finka A, O'Neil CP, Moniatte M, van der Vlies AJ, Giorgio TD, Hofmann H, Hubbell JA, Petri-Fink A. Superparamagnetic nanoparticles as a powerful systems biology characterization tool in the physiological context. Angew Chem Int Edit. 2008;47(41):7857–7860. doi: 10.1002/anie.200800357. [DOI] [PubMed] [Google Scholar]
- Schmidt A, Beck M, Malmstrom J, Lam H, Claassen M, Campbell D, Aebersold R (2011) Absolute quantification of microbial proteomes at different states by directed mass spectrometry. Molecular Systems Biology 7: 510. doi:http://www.nature.com/msb/journal/v7/n1/suppinfo/msb201137_S1.html [DOI] [PMC free article] [PubMed]
- Schuermann JP, Jiang JW, Cuellar J, Llorca O, Wang LP, Gimenez LE, Jin SP, Taylor AB, Demeler B, Morano KA, Hart PJ, Valpuesta JM, Lafer EM, Sousa R. Structure of the Hsp110:Hsc70 nucleotide exchange machine. Mol Cell. 2008;31(2):232–243. doi: 10.1016/j.molcel.2008.05.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwanhausser B, Busse D, Li N, Dittmar G, Schuchhardt J, Wolf J, Chen W, Selbach M. Global quantification of mammalian gene expression control. Nature. 2011;473(7347):337–342. doi: 10.1038/nature10098. [DOI] [PubMed] [Google Scholar]
- Sharma K, Vabulas RM, Macek B, Pinkert S, Cox J, Mann M, Hartl FU (2012) Quantitative proteomics reveals that Hsp90 inhibition preferentially targets kinases and the DNA damage response. Mol Cell Proteomics 11(3). doi:10.1074/mcp.M111.014654 [DOI] [PMC free article] [PubMed]
- Sharma SK, De Los Rios P, Christen P, Lustig A, Goloubinoff P. The kinetic parameters and energy cost of the Hsp70 chaperone as a polypeptide unfoldase. Nat Chem Biol. 2010;6(12):914–920. doi: 10.1038/nchembio.455. [DOI] [PubMed] [Google Scholar]
- Sharma SK, De Los Rios P, Goloubinoff P. Probing the different chaperone activities of the bacterial Hsp70–Hsp40 system using a thermolabile luciferase substrate. Proteins Struct Funct Bioinform. 2011;79(6):1991–1998. doi: 10.1002/prot.23024. [DOI] [PubMed] [Google Scholar]
- Shorter J (2011) The mammalian disaggregase machinery: Hsp110 synergizes with Hsp70 and Hsp40 to catalyze protein disaggregation and reactivation in a cell-free system. PLoS One 6(10). doi:10.1371/journal.pone.0026319 [DOI] [PMC free article] [PubMed]
- Siegenthaler RK, Christen P. Tuning of DnaK chaperone action by nonnative protein sensor DnaJ and thermosensor GrpE. J Biol Chem. 2006;281(45):34448–34456. doi: 10.1074/jbc.M606382200. [DOI] [PubMed] [Google Scholar]
- Siegers K, Waldmann T, Leroux MR, Grein K, Shevchenko A, Schiebel E, Hartl FU. Compartmentation of protein folding in vivo: sequestration of non-native polypeptide by the chaperonin–GimC system. EMBO J. 1999;18(1):75–84. doi: 10.1093/emboj/18.1.75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sousa R, Lafer EM. Keep the traffic moving: mechanism of the Hsp70 motor. Traffic. 2006;7(12):1596–1603. doi: 10.1111/j.1600-0854.2006.00497.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tissieres A, Mitchell HK, Tracy UM. Protein synthesis in salivary glands of Drosophila melanogaster—relation to chromosome puffs. J Mol Biol. 1974;84(3):389–398. doi: 10.1016/0022-2836(74)90447-1. [DOI] [PubMed] [Google Scholar]
- Torok Z, Vigh L, Goloubinoff P. Fluorescence detection of symmetric GroEL(14)(GroES(7))(2) heterooligomers involved in protein release during the chaperonin cycle. J Biol Chem. 1996;271(27):16180–16186. doi: 10.1074/jbc.271.27.16180. [DOI] [PubMed] [Google Scholar]
- Van de Water L, 3rd, Olmsted JB The quantitation of tubulin in neuroblastoma cells by radioimmunoassay. J Biol Chem. 1980;255(22):10744–10751. [PubMed] [Google Scholar]
- van der Vaart A, Ma JP, Karplus M. The unfolding action of GroEL on a protein substrate. Biophys J. 2004;87(1):562–573. doi: 10.1529/biophysj.103.037333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Veinger L, Diamant S, Buchner J, Goloubinoff P. The small heat-shock protein IbpB from Escherichia coli stabilizes stress-denatured proteins for subsequent refolding by a multichaperone network. J Biol Chem. 1998;273(18):11032–11037. doi: 10.1074/jbc.273.18.11032. [DOI] [PubMed] [Google Scholar]
- Volpe P, Eremenko T. Quantitative studies on cell proteins in suspension cultures. Eur J Biochem. 1970;12(1):195–200. doi: 10.1111/j.1432-1033.1970.tb00837.x. [DOI] [PubMed] [Google Scholar]
- Wang ZM, Shen W, Kotler DP, Heshka S, Wielopolski L, Aloia JF, Nelson ME, Pierson RN, Heymsfield SB. Total body protein: a new cellular level mass and distribution prediction model. Am J Clin Nutr. 2003;78(5):979–984. doi: 10.1093/ajcn/78.5.979. [DOI] [PubMed] [Google Scholar]
- Weiss YG, Bromberg Z, Raj N, Raphael J, Goloubinoff P, Ben-Neriah Y, Deutschman CS. Enhanced heat shock protein 70 expression alters proteasomal degradation of I kappa B kinase in experimental acute respiratory distress syndrome. Crit Care Med. 2007;35(9):2128–2138. doi: 10.1097/01.CCM.0000278915.78030.74. [DOI] [PubMed] [Google Scholar]
- Wistow GJ, Piatigorsky J. Lens crystallins—the evolution and expression of proteins for a highly specialized tissue. Annu Rev Biochem. 1988;57:479–504. doi: 10.1146/annurev.bi.57.070188.002403. [DOI] [PubMed] [Google Scholar]
- Wolf SF, Schlessinger D. Nuclear metabolism of ribosomal RNA in growing, methionine-limited, and ethionine-treated HeLa cells. Biochem-Us. 1977;16(12):2783–2791. doi: 10.1021/bi00631a031. [DOI] [PubMed] [Google Scholar]
- Yebenes H, Mesa P, Munoz IG, Montoya G, Valpuesta JM. Chaperonins: two rings for folding. Trends Biochem Sci. 2011;36(8):424–432. doi: 10.1016/j.tibs.2011.05.003. [DOI] [PubMed] [Google Scholar]
- Yeyati PL, Bancewicz RM, Maule J, van Heyningen V. Hsp90 selectively modulates phenotype in vertebrate development. PLoS Genet. 2007;3(3):431–447. doi: 10.1371/journal.pgen.0030043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young JC, Hoogenraad NJ, Hartl FU. Molecular chaperones Hsp90 and Hsp70 deliver preproteins to the mitochondrial import receptor Tom70. Cell. 2003;112(1):41–50. doi: 10.1016/S0092-8674(02)01250-3. [DOI] [PubMed] [Google Scholar]
- Zuehlke A, Johnson JL. Hsp90 and co-chaperones twist the functions of diverse client proteins. Biopolym. 2010;93(3):211–217. doi: 10.1002/bip.21292. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Raw polypeptide abundances in 11 imortalized human cell lines. Bar graphs showing high disparity of (A) absolute polypeptide copy numbers per cell and (B) protein concentration, in the 11 different immortalized human cell lines (Geiger et al. 2012). Total polypeptide copy number and SD are from three independent measures. Hatched boxes are 1010 polypeptides and 300 mg/ml per cell as previously reported (Albe et al. 1990). (DOCX 62 kb)
Numeral proportions of 11,731 polypeptides in 11 immortalized human cell lines and individual polypeptide copy number in HeLa cells. The numeral proportions of polypeptides with a p value less than 0.1 are labeled in bold. The polypeptide average copy numbers and SD in HeLa cells are given in a separate sheet (XLSX 7413 kb)
Mass proportions of 11,731 polypeptides in 11 immortalized human cell lines and individual polypeptide mass in HeLa cells. The mass proportions of polypeptides with a p value less than 0.1 are labeled in bold. The average mass and SD (in picograms) of each polypeptide in HeLa cells are given in a separate sheet (XLSX 9413 kb)
List of human 147 chaperome members. Numeral proportions, copy numbers, mass proportions with corresponding p values, and mass of each detected chaperome member in HeLa cells (XLSX 43 kb)