Abstract
The high intracellular oxidative stress in a cancer cell is a biologically relevant stimulus for efficient intracellular delivery of therapeutic genes. In this study, reactive oxygen species (ROS)-responsive poly(amino thioketal) (PATK) was synthesized to achieve efficient and safe intracellular gene delivery in prostate cancer cells. The DNA/PATK polyplexes were efficiently disassembled upon exposure to high levels of ROS in prostate cancer cells, leading to enhanced intracellular release of DNA in the cells. As a result, DNA/PATK polyplexes showed significantly higher gene transfection efficiency than their non-degradable counterparts did. In addition, conjugation of GRP78 protein-targeting peptide to the PATK not only increased its cellular uptake in prostate cancer cells but also enhanced gene transfection efficiency. This study demonstrates that ROS-responsive PATK functionalized with a cancer-targeting peptide is a promising gene carrier for safe, efficient, and cancer-targeted gene delivery.
Keywords: Cancer-targeted delivery, Gene expression, Nanoparticles, Reactive oxygen species
Over the past decade, cancer gene therapy has been one of the most rapidly evolving areas in the clinic due to improved understanding of cancer at the molecular level.[1] The clinical success of cancer gene therapy critically depends on the development of safe, efficient, and targeted gene carriers.[2] In recent years, synthetic nonviral vectors have attracted great attention as a promising alternative to conventional viral vectors due to their lower risk of immunogenicity, larger gene delivery capacity.[3] However, nonviral vectors often suffer from low transfection efficiency due to their poor extra- and intracellular processes as compared to viral vectors.[4] In addition, the lack of target-specificity for nonviral vectors is another critical obstacle that needs to be overcome before their maximized therapeutic efficacy can be achieved at the target site.[4]
Recently, various types of smart polymers capable of responding to the stimuli intrinsic to a tumor environment have emerged as efficient nonviral gene carriers for the treatment of cancer.[5] Most of the stimuli-responsive nonviral vectors rely on the use of extracellular environments of tumors such as low extracellular tumor pH and matrix metalloproteinases (MMPs) for cancer-targeted gene delivery.[6] Few of them has been developed with responses to the unique intracellular stimuli of cancer cells, which could be used to greatly improve their intracellular trafficking and gene transfection.
Recent studies reported that, due to oncogenic transformation, cancer cells constantly generate high levels of intracellular reactive oxygen species (ROS), including H2O2, hydroxyl radical, and superoxide, in comparison with normal cells.[7] Therefore, in this study, the intracellular ROS in cancer cells was utilized as a unique cancer-related stimulus to mediate intracellular gene delivery. We demonstrated efficient and safe gene delivery in cancer cells by developing a novel ROS-responsive, cationic, water-soluble polymer composed of biodegradable thioketal linkages that are readily cleavable in ROS-abundant conditions.[8] It was hypothesized that the ROS-cleavable thioketal-based polymeric carrier could enhance the gene delivery efficiency in cancer cells by facilitating intracellular release of the encapsulated nucleic acids in response to high levels of intracellular ROS (Figure 1a). We further functionalized this polymer with a cancer-targeting peptide to achieve cancer-targeted gene delivery. This modification could enhance the gene delivery efficiency and thus reduce the potential systemic toxicity by avoiding nonspecific accumulation in normal cells.
Figure 1.
(a) Illustration of intracellular delivery of plasmid DNA to the nucleus of a cancer cell using ROS-cleavable poly(amino thioketal) (PATK). After the DNA/PATK polyplexes have been internalized by the cancer cell and escaped from the endosome, ROS-triggered degradation of the PATK facilitates the intracellular release of DNA from the particle, leading to enhanced gene transfection. (b) Synthetic scheme of ROS-responsive PATK.
The ROS-cleavable, cationic polymer was synthesized via polymerization of oligoamines with acrylamide thioketal cross-linkers as shown in Figure 1b. Briefly, trifluoroacetate (TFA)- protected N,N'-bis(2-aminoethyl)-1,3-propanediamine was polymerized with the thioketal cross-linkers via Michael addition conjugation, followed by deprotection of the TFA groups, resulting in poly(amino thioketal) (PATK). Figure S1a in the Supporting Information shows the detailed synthesis. Non-degradable poly(amine) without thioketal linkages was also synthesized as an ROS-insensitive counterpart to investigate the effects of ROS-sensitivity on the intracellular release of nucleic acids and consequent gene delivery efficiency (see Figure S1b for detailed synthesis). The formation of PATK was confirmed by the 1H NMR spectroscopy shown in Figure S2a. The molecular weights of PATK were Mn = 5.7 kDa and Mw = 9.4 kDa with a polydispersity index of 1.65, as determined by gel permeation chromatography (GPC).
The degradation profile of PATK under simulated ROS conditions was quantified by 1H NMR spectroscopy. In a typical experiment, PATK was dissolved in D2O containing H2O2 and trace amounts of transition metal ions (e.g., 1.6 μM CuCl2) to trigger its degradation by ROS. 3-(trimethylsilyl)propionic acid sodium salt (TSP) was also added into the D2O solvent as a stable reference. When the PATK was incubated in the solvent at 37 °C for different periods of time, the disappearance of the thioketal linkage peak (δ 1.62 ppm) was monitored and quantified using 1H NMR spectroscopy in comparison with the TSP peak. The 1H NMR spectrum confirmed that the thioketal linkages were efficiently cleaved by ROS, generating acetone (δ 2.16 ppm) as a by-product during the cleavage process (data not shown). This finding was in agreement with a previous study regarding the dethioacetalization using H2O2.[9]. The degradation rate of thioketal linkages was proportional to the concentration of H2O2 as shown in Figure 2a. For example, the half-lives of thioketal linkages in PATK incubated with 100 mM and 200 mM H2O2 solution were approximately 20 h and 11 h, respectively (Figure 2a). The decrease in molecular weight of PATK after incubation with 100 mM H2O2 and 1.6 μM CuCl2 at 37 °C was also confirmed by GPC (Figure S2b).
Figure 2.
(a) Degradation profiles of thioketal linkages in PATK after exposure to H2O2 at various concentrations for different periods of time, as determined by 1H NMR spectroscopy. (b) Transmission electron micrographs of the DNA/PATK polyplexes (N/P = 100) before and after incubation with H2O2 for 12 h at 37 °C. Scale bars indicate 100 nm. (c) Generation of intracellular ROS in PC3 cells confirmed by fluorescence microscopy. Strong green fluorescence throughout the cells demonstrates the high levels of intracellular ROS in PC3 cells. (d) Confocal micrographs illustrating the intracellular localization of DNA/PATK and DNA/non-degradable poly(amine) polyplexes in PC3 cells. Cellular nuclei were counter-stained with DRAQ5 (blue). Yellow dots represent co-localized polymer (green) and DNA (red). Scale bars indicate 10 μm.
Efficient DNA complexation by cationic PATK was confirmed by measuring the particle size and surface charge of DNA/PATK polyplexes. As shown in Figure S3a, PATK efficiently complexed with DNA, forming stable polyplexes with diameters ranging from 130 to 200 nm at amine to phosphate (N/P) ratios of 50 and higher. The zeta potentials of DNA/PATK polyplexes were in the range of +15 to +25 mV at N/P ratios of 50 and higher (Figure S3b). Efficient ROS-triggered dissociation of DNA from DNA/PATK polyplexes was confirmed by an agarose gel electrophoresis assay. Before incubation with a H2O2 solution, DNA was completely retained by PATK at N/P ratios of 20 and higher (Figure S4). However, DNA was efficiently liberated from the PATK polyplexes when the samples were exposed to ROS (Figure S4). TEM images also demonstrated ROS-triggered destabilization of the DNA/PATK polyplexes (Figure 2b). In addition, destabilization of the DNA/PATK polyplexes in response to biologically relevant levels of H2O2 (e.g., 100 μM and 1 mM)[10] was confirmed by ethidium bromide (EtBr) exclusion assay (Figure S5a). The degradation selectivity of the DNA/PATK polyplexes under various ROS, including hydroxyl radical, H2O2, and superoxide, was also examined using the EtBr exclusion assay (Figure S5b). Among the tested ROS, thioketal linkages showed the most efficient degradation in response to the hydroxyl radical.
To confirm whether cancer cells spontaneously generate ROS, we visualized intracellular ROS generation using 2′,7-dichlorofluoresceindiacetate (DCFDA), a cell permeable fluorescent dye that is rapidly oxidized to a fluorescent molecule by intracellular ROS.[11] The PC3 cell, a human prostate cancer cell line, was used as a model system due to its inherently high levels of ROS.[7a] Fluorescent micrographs of DCFDA-stained PC3 cells clearly demonstrated the generation of intracellular ROS throughout the cell (Figure 2c). To investigate if ROS in PC3 cells trigger efficient intracellular disassembly of DNA/PATK polyplexes, their intracellular disassembly in PC3 cells was examined by confocal laser scanning microscopy (Figure 2d). For comparison, non-degradable poly(amine) and 25 kDa branched PEI (B-PEI) were also complexed with DNA and incubated with PC3 cells. As shown in Figure 2d, a significant amount of free DNA (labeled with Alexa Fluor 568 dye, red color) were disassembled from the Alexa Fluor 488-labeled PATK (green color), demonstrating the efficient intracellular disassembly of DNA/PATK polyplexes in the ROS-generating PC3 cells. In contrast, limited intracellular release of DNA was observed with DNA/non-degradable poly(amine) polyplexes and DNA/B-PEI polyplexes as represented by many yellow dots (Figure 2d and Figure S6).
Enhanced gene transfection by ROS-responsive PATK was investigated by comparing its transfection efficiency with reference to non-degradable poly(amine). As shown in Figure 3a, eGFP expression of the PC3 cells incubated with PATK polyplexes complexing eGFP-encoding plasmid DNA was significantly higher than that of the cells incubated with the DNA/non-degradable poly(amine) polyplexes. The transfection efficiency of PATK was also compared with B-PEI, a commercially available superior transfecting agent. Obviously, DNA/PATK polyplexes transfected PC3 cells more efficiently than DNA/B-PEI polyplexes too (Figure 3a). It is well-known that intracellular release of DNA from the gene carrier is crucial for efficient gene transfection.[12] The confocal micrographs in Figure 2d clearly shows that DNA was efficiently released from PATK polyplexes in PC3 cells. Taken together, enhanced gene transfection by the DNA/PATK polyplexes in comparison with DNA/non-degradable poly(amine) and DNA/B-PEI polyplexes can be attributed to their efficient intracellular disassembly in ROS-generating PC3 cells. In addition to higher gene transfection, ROS-sensitive DNA/PATK polyplexes exhibited lower cytotoxicity than DNA/B-PEI polyplexes (Figure 3b). In general, the cytotoxicity of cationic polymers increases with their charge density and molecular weights.[13] Therefore, the reduced cytotoxicity of PATK can be attributed to its lower molecular weight and lower cationic surface charge (DNA/PATK: ca. 20 mV, DNA/B-PEI: ca. 30 mV) (Figure S3b).
Figure 3.
(a) Transfection efficiency and (b) cell viability for DNA/PATK (N/P = 100), DNA/non-degradable poly(amine) (N/P = 120), and DNA/B-PEI polyplexes (N/P = 9) in PC3 cells. The N/P ratios of the polyplexes were determined by optimized transfection efficiency and cytotoxicity. * indicates p < 0.05. ** indicates p < 0.01.
To investigate if the enhanced gene transfection by PATK is selectively triggered in response to high levels of ROS in cancer cells, its transfection efficiency in PC3 cells was compared with the Chinese hamster ovary (CHO) cells that produce significantly lower levels of ROS (Figure S7a). As shown in Figure 3a and Figure S7b, PATK showed significantly higher transfection efficiency in PC3 cells than in non-cancerous CHO cells, whereas non-degradable B-PEI showed similar transfection efficiencies for both PC3 and CHO cells. Considering that the total cellular uptake of both DNA/PATK and DNA/B-PEI polyplexes was similar for both PC3 and CHO cells (Figure S7c), the noticeably enhanced gene transfection by PATK in PC3 cells can be attributed to more efficient disassembly of the DNA/PATK polyplexes in PC3 cells that are known to have higher levels of ROS. The significantly enhanced gene transfection by PATK in response to high levels of ROS in prostate cancer cells was also consistently observed in LNCaP cells, another prostate cancer cell line showing higher levels of intracellular ROS than those of non-cancerous CHO cells (Figure S7).
Cancer-targeted gene delivery is a key requirement for future cancer gene therapy in the clinic.[14] To demonstrate the feasibility of achieving cancer-targeted gene delivery using the ROS-responsive PATK, we further conjugated it with GRP78-binding peptide (GRP78P, peptide sequence = WIFPWIQL) (Figure S8), which selectively binds to GRP78 proteins over-expressed by many types of tumor cells including the prostate cancer cells.[15] We then complexed fluorescently labeled DNA with the GRP78P-conjugated PATK (GRP78P-PATK) or GRP78P-free PATK to quantify their cellular uptake. Notably, the conjugation of GRP78P significantly increased (ca. 3-fold higher) cellular uptake of the PATK polyplexes in PC3 cells (Figure 4). Consequently, DNA/GRP78P-PATK polyplexes yielded a 2-fold higher gene transfection efficiency in PC3 cells in comparison with DNA/GRP78P-free PATK polyplexes. To confirm that the increased transfection efficiency by DNA/GRP78P-PATK polyplexes was mainly caused by the receptor-mediated cellular uptake of GRP78P, a competitive assay where free GRP78P was pre-incubated with PC3 cells prior to the transfection was conducted. When 200 μM of free GRP78P was pre-incubated with the cells prior to the transfection, the DNA/GRP78P-PATK polyplexes showed drastically reduced DNA transfection (Figure 4). Combined together, these results confirm that DNA/GRP78P-PATK polyplexes are efficiently internalized by the cancer cells via GRP78 receptor-mediated endocytosis, leading to caner-targeted gene delivery.
Figure 4.
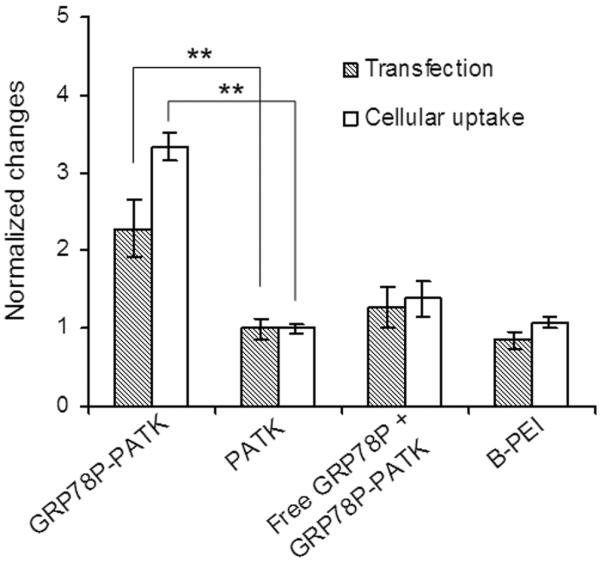
Cancer-targeted gene transfection by DNA/GRP78P-PATK polyplexes in PC3 cells. The results are normalized to the relative eGFP expression and cellular uptake by DNA/PATK polyplexes. Conjugation of cancer cell-targeting GRP78-binding peptide (GRP78P) significantly enhanced the transfection and cellular uptake of DNA/PATK polyplexes. ** indicates p < 0.01.
In conclusion, ROS-responsive thioketal-based PATK was synthesized for safe, efficient, and targeted gene delivery in prostate cancer cells. Degradation of thioketal linkages in PATK under ROS conditions led to efficient intracellular release of the complexed DNA in prostate cancer cells. As a result, the DNA/PATK polyplexes exhibited efficient gene transfection in prostate cancer cells. Incorporation of GRP78-binding peptide to the PATK achieved cancer-targeted gene transfection. Most importantly, this study demonstrates that the high levels of intracellular ROS in cancer cells are unique biological stimuli that can be utilized for efficient and targeted gene delivery in cancer cells.
Supplementary Material
Footnotes
This work was supported by in part by a 2006 NIH Director's Pioneer Award (DP1 OD000798) and start-up funds from Georgia Institute of Technology. The authors thank Dennis Oakley of the Bakewell Neuroimaging Core at Washington University School of Medicine for assistance with some of the confocal microscopy imaging studies.
References
- [1].a) Wu Q, Moyana T, Xiang J. Curr. Gene Ther. 2001;1:101–122. doi: 10.2174/1566523013349002. [DOI] [PubMed] [Google Scholar]; b) Feldman AL, Libutti SK. Cancer. 2000;89:1181–1194. [PubMed] [Google Scholar]; c) Oh Y-K, Park TG. Adv. Drug Deliv. Rev. 2009;61:850–862. doi: 10.1016/j.addr.2009.04.018. [DOI] [PubMed] [Google Scholar]
- [2].a) Mulligan RC. Science. 1993;260:926–932. doi: 10.1126/science.8493530. [DOI] [PubMed] [Google Scholar]; b) Anderson WF. Nature. 1998;392:25–30. doi: 10.1038/32058. [DOI] [PubMed] [Google Scholar]
- [3].a) Somia N, Verma IM. Nat. Rev. Gene. 2000;1:91–99. doi: 10.1038/35038533. [DOI] [PubMed] [Google Scholar]; b) Thomas CE, Ehrhardt A, Kay MA. Nat. Rev. Genet. 2003;4:346–358. doi: 10.1038/nrg1066. [DOI] [PubMed] [Google Scholar]; c) Gabrielson NP, Lu H, Yin L, Li D, Wang F, Cheng J. Angew. Chem. Int. Ed. 2012;51:1143–1147. doi: 10.1002/anie.201104262. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [4].a) Meyer M, Wagner E. Hum. Gene Ther. 2006;17:1062–1076. doi: 10.1089/hum.2006.17.1062. [DOI] [PubMed] [Google Scholar]; b) Merdan T, Kopecek J, Kissel T. Adv. Drug Deliv. Rev. 2002;54:715–758. doi: 10.1016/s0169-409x(02)00046-7. [DOI] [PubMed] [Google Scholar]; c) Park TG, Jeong JH, Kim SW. Adv. Drug Deliv. Rev. 2006;58:467–486. doi: 10.1016/j.addr.2006.03.007. [DOI] [PubMed] [Google Scholar]; d) Pack DW, Hoffman AS, Pun S, Stayton PS. Nat. Rev. Drug Discov. 2005;4:581–593. doi: 10.1038/nrd1775. [DOI] [PubMed] [Google Scholar]; e) Medina-Kauwe LK, Xie J, Hamm-Alvarez S. Gene Ther. 2005;12:1734–1751. doi: 10.1038/sj.gt.3302592. [DOI] [PubMed] [Google Scholar]
- [5].a) Alexander C. Expert Opin. Drug Deliv. 2006;3:573–581. doi: 10.1517/17425247.3.5.573. [DOI] [PubMed] [Google Scholar]; b) Li P, Liu D, Miao L, Liu C, Sun X, Liu Y, Zhang N. Int. J. Nanomedicine. 2012;7:925–939. doi: 10.2147/IJN.S26955. [DOI] [PMC free article] [PubMed] [Google Scholar]; c) Hatakeyama H, Akita H, Kogure K, Oishi M, Nagasaki Y, Kihira Y, Ueno M, Kobayashi H, Kikuchi H, Harashima H. Gene Ther. 2007;14:68–77. doi: 10.1038/sj.gt.3302843. [DOI] [PubMed] [Google Scholar]; d) Burke RS, Pun SH. Bioconjugate Chem. 2010;21:140–150. doi: 10.1021/bc9003662. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [6].a) Lee ES, Gao Z, Bae YH. J. Control. Release. 2008;132:164–170. doi: 10.1016/j.jconrel.2008.05.003. [DOI] [PMC free article] [PubMed] [Google Scholar]; b) Roy R, Yang J, Moses MA. J. Clin. Oncol. 2009;27:5287–5297. doi: 10.1200/JCO.2009.23.5556. [DOI] [PMC free article] [PubMed] [Google Scholar]; c) Hatakeyama H, Akita H, Harashima H. Adv. Drug Deliv. Rev. 2011;63:152–160. doi: 10.1016/j.addr.2010.09.001. [DOI] [PubMed] [Google Scholar]
- [7].a) Kumar B, Koul S, Khandrika L, Meacham RB, Koul HK. Cancer Res. 2008;68:1777–1785. doi: 10.1158/0008-5472.CAN-07-5259. [DOI] [PubMed] [Google Scholar]; b) Szatrowski TP, Nathan CF. Cancer Res. 1991;51:794–798. [PubMed] [Google Scholar]; c) Senthil K, Aranganathan S, Nalini N. Clin. Chim. Acta. 2004;339:27–32. doi: 10.1016/j.cccn.2003.08.017. [DOI] [PubMed] [Google Scholar]; d) Toyokuni S, Okamoto K, Yodoi J, Hiai H. FEBS Lett. 1995;358:1–3. doi: 10.1016/0014-5793(94)01368-b. [DOI] [PubMed] [Google Scholar]
- [8].a) Wilson DS, Dalmasso G, Wang L, Sitaraman SV, Merlin D, Murthy N. Nat. Mat. 2010;9:923–928. doi: 10.1038/nmat2859. [DOI] [PMC free article] [PubMed] [Google Scholar]; b) Shukla AK, Verma M, Singh KN. Indian J. Chem. B. 2004;43:1748–1752. [Google Scholar]; c) Ganguly NC, Barik SK. Synthesis. 2009;8:1393–1399. [Google Scholar]
- [9].Ganguly NC, Mondal P. Synth. Commun. 2011;41:2374–2384. [Google Scholar]
- [10].Halliwell B, Clement MV, Long LH. FEBS Lett. 2000;486:10–13. doi: 10.1016/s0014-5793(00)02197-9. [DOI] [PubMed] [Google Scholar]
- [11].Genty B, Briantais J, Baker N. Biochem. Biophys. Acta. 1989;990:87–92. [Google Scholar]
- [12].a) Gabrielson NP, Pack DW. Biomacromolecules. 2006;7:2427–2435. doi: 10.1021/bm060300u. [DOI] [PubMed] [Google Scholar]; b) Ou M, Wang X-L, Xu R, Chang C-W, Bull DA, Kim SW. Bioconjugate Chem. 2008;19:626–633. doi: 10.1021/bc700397x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [13].a) Fischer D, Li Y, Ahlemeyer B, Krieglstein J, Kissel T. Biomaterials. 2003;24:1121–1131. doi: 10.1016/s0142-9612(02)00445-3. [DOI] [PubMed] [Google Scholar]; b) Fischer D, Bieber T, Li Y, Elsässer H, Kissel T. Pharm. Res. 1999;16:1273–1279. doi: 10.1023/a:1014861900478. [DOI] [PubMed] [Google Scholar]
- [14].a) Rajendran L, Knölker H-J, Simons K. Nat. Rev. Drug Discov. 2010;9:29–42. doi: 10.1038/nrd2897. [DOI] [PubMed] [Google Scholar]; b) Brannon-Peppas L, Blanchette JO. Adv. Drug Deliv. Rev. 2004;56:1649–1659. doi: 10.1016/j.addr.2004.02.014. [DOI] [PubMed] [Google Scholar]
- [15].a) Arap MA, Lahdenranta J, Mintz PJ, Hajitou A, Sarkis ÁS, Arap W, Pasqualini R. Cancer Cell. 2004;6:275–284. doi: 10.1016/j.ccr.2004.08.018. [DOI] [PubMed] [Google Scholar]; b) Larson N, Ray A, Malugin A, Pike DB, Ghandehari H. Pharm. Res. 2010;27:2683–2693. doi: 10.1007/s11095-010-0267-7. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.





