Abstract
Salmonella enterica subsp. houtenae serovar 16:z4, z32:-- str. RKS3027 was isolated from a human in Illinois, USA. S. enterica subsp. houtenae is a facultative aerobic rod-shaped Gram-negative bacterium. Here we describe the features of this organism, together with the draft genome sequence and annotation. The 4,404,136 bp long genome (97 contigs) contains 4,335 protein-coding gene and 28 RNA genes.
Keywords: Salmonella enterica, subspecies, houtenae, genome
Introduction
Salmonella is an important genus of human and animal pathogens [1], and more than 2,600 different serovars have been described. Currently, the genus Salmonella is divided into two species, S. enterica, and S. bongori [2]. S. enterica comprises seven subspecies: I (also called subspecies enterica), II (also called subspecies salamae), IIIa (also called subspecies arizonae), IIIb (also called subspecies diarizonae), IV (also called subspecies houtenae), VI (also called subspecies indica), and VII [3]. Most of Salmonella serovars belong to the S. enterica subspecies I and are responsible for disease in warm-blooded animals and humans [4]. Other serovars were usually isolated from cold-blooded organisms and the environment, but could also cause human disease occasionally. In contrast with S. enterica subspecies I, very limited information is available regarding pathogenicity of the other subspecies. When infecting humans, these serovars usually cause an intestinal infection (e.g., diarrhea), but previous reports in the literature [5] have shown that the serovars of Salmonella subspecies II–IV are capable of causing serious infections, including septicemia and abscesses. There has been an increase in case reports on extraintestinal infections caused by these subspecies [6]. S. enterica subsp. houtenae serovar 16:z4,z32:-- str. RKS3027 is a human isolate. This strain is of interest because of its pathogenicity as well as its divergent phylogenetic position among S. enterica.
Classification and features
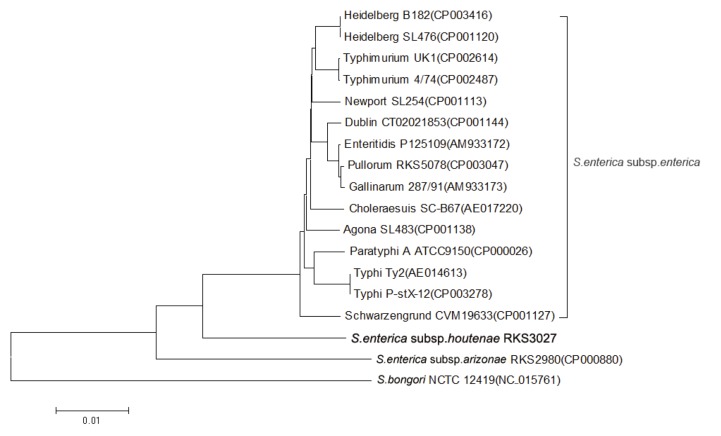
Few 16S rRNA sequences of Salmonella subspecies are available except S. enterica subsp. enterica. Meanwhile, it is increasingly commonplace to construct the phylogenetic tree by using the whole-genome sequence for higher precision and robustness [7,8]. Therefore we used a total of 2,500 orthologs of 18 strains of Salmonella for constructing a genome-scale phylogenetic tree. Genetic relatedness of S. enterica subsp. houtenae strain RKS3027 to other Salmonella subspecies strains was shown in Figure 1. On the tree, all S. enterica subsp. enterica strains were clustered together, and S. enterica subsp. houtenae RKS3027 positioned between S. enterica subsp. enterica and S. bongori.
Figure 1.
Phylogenetic tree highlighting the position of S. enterica subsp. houtenae strain RKS3027 relative to the other types and strains of Salmonella. GenBank accession numbers are indicated in the parentheses. The tree was built based on the comparison of concatenated nucleotide sequences of 2,500 orthologs conserved in all strains. Individual orthologous sequences were aligned by the MAFFT [9] and phylogenetic tree was constructed by using the neighbor-joining method within the MEGA software [10].
The Salmonella genus belongs to the bacterial family Enterobacteriaceae [11]. The bacteria are rod shaped, Gram-negative, with diameter of 0.7 to 1.5 µm and length of 2 to 5 µm (Table 1). They are facultative anaerobes, non-spore-forming, flagellated, and motile. They grow within the optimal temperature range 35 °C - 37 °C and within an optimal pH range of 7.2-7.6. S. enterica subsp. houtenae is salicin-positive and able to grow in KCB medium, two distinguishing characteristics when compared with S. enterica subsp. enterica. The strain is deposited in the Salmonella Genetic Stock Centre (SGSC), University of Calgary, Canada as S. enterica subsp. houtenae RKS3027 (= SGSC 3086).
Table 1. Classification and general features of S. enterica subsp. houtenae RKS3027 according to the MIGS recommendations [12].
| MIGS ID | Property | Term | Evidence codea |
|---|---|---|---|
| Current classification | Domain Bacteria | TAS [13] | |
| Phylum Proteobacteria | TAS [14] | ||
| Class Gammaproteobacteria | TAS [15,16] | ||
| Order Enterobacteriales | TAS [17] | ||
| Family Enterobacteriaceae | TAS [18-20] | ||
| Genus Salmonella | TAS [18,21-23] | ||
| Species Salmonella enterica | TAS [23,24] | ||
| Subspecies Salmonella enterica subsp. houtenae | TAS [23,24] | ||
| Strain RKS3027 | IDA | ||
| Serovar 16:z3, z32:-- | IDA | ||
| Gram stain | Negative | IDA | |
| Cell shape | Rod-shaped | IDA | |
| Motility | Motile | IDA | |
| Sporulation | Non-sporulating | IDA | |
| Temperature range | Mesophilic | IDA | |
| Optimum temperature | 35 °C - 37 °C | IDA | |
| Carbon source | Glucose | IDA | |
| Energy source | Chemoorganotrophic | IDA | |
| MIGS-6 | Habitat | Reptiles | IDA |
| MIGS-6.3 | Salinity | Medium | IDA |
| MIGS-22 | Oxygen | Facultative anaerobes | IDA |
| MIGS-15 | Biotic relationship | Endophyte | IDA |
| MIGS-14 | Pathogenicity | Pathogenic | IDA |
| MIGS-4 | Geographic location | Illinois, USA | NAS |
| MIGS-5 | Sample collection time | 1986 | NAS |
| MIGS-4.1 | Latitude | Not report | NAS |
| MIGS-4.2 | Longitude | Not report | NAS |
| MIGS-4.3 | Depth | Not report | NAS |
| MIGS-4.4 | Altitude | Not report | NAS |
a) Evidence codes - IDA: Inferred from Direct Assay; TAS: Traceable Author Statement (i.e., a direct report exists in the literature); NAS: Non-traceable Author Statement (i.e., not directly observed for the living, isolated sample, but based on a generally accepted property for the species, or anecdotal evidence). These evidence codes are from the Gene Ontology project [25].
Genome sequencing information
Genome project history
This organism was selected for sequencing on the basis of its phylogenetic position and its serious virulence in humans compared to the reptiles. This Whole Genome Shotgun project has been deposited at DDBJ/EMBL/GenBank under the accession ANHR00000000. The version described in this paper is the first version, ANHR01000000, and the sequence consists of 97 large contigs. Table 2 presents the project information and its association with MIGS version 2.0 compliance [12].
Table 2. Project information.
| MIGS ID | Property | Term |
|---|---|---|
| MIGS-31 | Finishing quality | Draft |
| MIGS-28 | Libraries used | Illumina Paired-End library |
| MIGS-29 | Sequencing platforms | Illumina HiSeq 2000 |
| MIGS-31.2 | Fold coverage | 100 × |
| MIGS-30 | Assemblers | SOAPdenovo v1.05 |
| MIGS-32 | Gene calling method | RAST |
| Genbank ID | ANHR00000000 | |
| GOLD ID | Gi21447 | |
| Project relevance | Evolution in bacteria, human pathogen |
Growth conditions and DNA isolation
S. enterica subsp. houtenae strain RKS3027 was grown Luria Broth (LB) medium at 37°C. The DNA was extracted from the cell, concentrated and purified using the Qiamp kit (Qiagen), as detailed in the manual for the instrument.
Genome sequencing and assembly
The genome of S. enterica subsp. houtenae RKS3027 was sequenced using the Illumina sequencing platform by the paired-end strategy (2×100bp). The details of library construction and sequencing can be found at the Illumina web site [26]. The final coverage reached 100-fold for an estimated genome size of 4.5 Mb. The sequence data from Illumina HiSeq 2000 were assembled with SOAPdenovo v1.05. The final assembly contained 97 large contigs (>3000 bp) in 59 scaffolds generating a genome size of 4.4 Mb.
Genome annotation
Genes were predicted using RAST (Rapid Annotation using Subsystem Technology) [27] with gene caller GLIMMER3 [28] followed by manual curation. The predicted bacterial protein sequences were compared with the annotated genes from four available Salmonella genomes, i.e., S. enterica subsp. enterica Typhi P-stx-12, S. enterica subsp. enterica Heidelberg B182, S. enterica subsp. enterica Typhimurium UK-1 and S. enterica subsp. enterica Typhimurium 4/74 and searched against the Clusters of Orthologous Groups (COG) databases using BLASTP. The BLAST results were filtered with the following parameters: identities >90% and compared length >70%. CGViewer was used for visualization of genomic features [29].
Genome properties
The genome of S. enterica subsp. houtenae RKS3027 is 4,404,136 bp long (97 contigs) with a 51.68% G + C content (Table 3 and Figure 2). Of the 4,363 predicted genes, 4,335 were protein-coding genes, and 28 were RNAs (1 5S rRNA gene and 27 predicted tRNA genes). A total of 3,378 genes (77.42%) were assigned a putative function. The remaining genes were annotated as hypothetical proteins. The properties and statistics of the genome are summarized in Table 3. The distribution of genes into COGs functional categories is presented in Table 4.
Table 3. Nucleotide content and gene count levels of the genome.
| Attribute | Value | % of totala |
|---|---|---|
| Genome size (bp) | 4,404,136 | |
| DNA coding region (bp) | 3,824,952 | 86.85 |
| DNA G+C content (bp) | 2,276,005 | 51.68 |
| Total genes | 4,363 | 100 |
| RNA genes | 28 | 0.06 |
| Protein-coding genes | 4,335 | 99.36 |
| Genes assigned to COGs | 3,378 | 77.42 |
a) The total is based on either the size of the genome in base pairs or the total number of protein coding genes in the annotated genome.
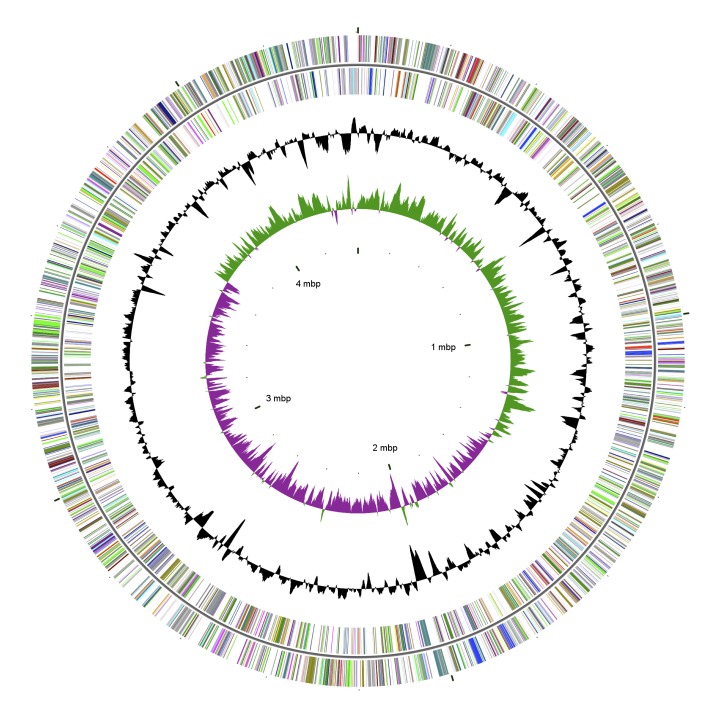
Figure 2.
Graphical circular map of the S. enterica subsp. houtenae strain RKS 3027 genome. From the outside to the center: genes on forward strand (color by COG categories), genes on reverse strand (color by COG categories), GC content, GC skew. The map was generated with the CGviewer software.
Table 4. Number of genes associated with the 25 general COG functional categories.
| Code | Value | %agea | Description |
|---|---|---|---|
| J | 163 | 3.76 | Translation |
| A | 1 | 0.02 | RNA processing and modification |
| K | 281 | 6.48 | Transcription |
| L | 176 | 4.06 | Replication, recombination and repair |
| B | 0 | 0.00 | Chromatin structure and dynamics |
| D | 32 | 0.74 | Cell cycle control, mitosis and meiosis |
| Y | 0 | 0.00 | Nuclear structure |
| V | 48 | 1.11 | Defense mechanisms |
| T | 103 | 2.38 | Signal transduction mechanisms |
| M | 235 | 5.42 | Cell wall/membrane biogenesis |
| N | 95 | 2.19 | Cell motility |
| Z | 0 | 0.00 | Cytoskeleton |
| W | 0 | 0.00 | Extracellular structures |
| U | 41 | 0.95 | Intracellular trafficking and secretion |
| O | 138 | 3.18 | Posttranslational modification, protein turnover, chaperones |
| C | 254 | 5.86 | Energy production and conversion |
| G | 343 | 7.91 | Carbohydrate transport and metabolism |
| E | 319 | 7.36 | Amino acid transport and metabolism |
| F | 77 | 1.78 | Nucleotide transport and metabolism |
| H | 131 | 3.02 | Coenzyme transport and metabolism |
| I | 89 | 2.05 | Lipid transport and metabolism |
| P | 175 | 4.04 | Inorganic ion transport and metabolism |
| Q | 47 | 1.08 | Secondary metabolites biosynthesis, transport and catabolism |
| R | 318 | 7.34 | General function prediction only |
| S | 312 | 7.20 | Function unknown |
| - | 957 | 22.08 | Not in COGs |
a) The total is based on the total number of protein coding genes in the annotated genome.
Acknowledgments
This work was supported by grants of the National Natural Science Foundation of China (NSFC30970119, 81030029, 81271786, NSFC-NIH 81161120416) to SLL.
References
- 1.Grassl GA, Finlay BB. Pathogenesis of enteric Salmonella infections. Curr Opin Gastroenterol 2008; 24:22-26 10.1097/MOG.0b013e3282f21388 [DOI] [PubMed] [Google Scholar]
- 2.Reeves MW, Evins GM, Heiba AA, Plikaytis BD, Farmer JJ., III Clonal nature of Salmonella typhi and its genetic relatedness to other salmonellae as shown by multilocus enzyme electrophoresis, and proposal of Salmonella bongori comb. nov. J Clin Microbiol 1989; 27:313-320 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Boyd EF, Wang FS, Whittam TS, Selander RK. Molecular genetic relationships of the salmonellae. Appl Environ Microbiol 1996; 62:804-808 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Li J, Ochman H, Groisman EA, Boyd EF, Solomon F, Nelson K, Selander RK. Relationship between evolutionary rate and cellular location among the Inv/Spa invasion proteins of Salmonella enterica. Proc Natl Acad Sci USA 1995; 92:7252-7256 10.1073/pnas.92.16.7252 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Weiss SH, Blaser MJ, Paleologo FP, Black RE, McWhorter AC, Asbury MA, Carter GP, Feldman RA, Brenner DJ. Occurrence and distribution of serotypes of the Arizona subgroup of Salmonella strains in the United States from 1967 to 1976. J Clin Microbiol 1986; 23:1056-1064 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Abbott SL, Ni FC, Janda JM. Increase in extraintestinal infections caused by Salmonella enterica subspecies II-IV. Emerg Infect Dis 2012; 18:637-639 10.3201/eid1804.111386 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Luo Y, Fu C, Zhang DY, Lin K. BPhyOG: an interactive server for genome-wide inference of bacterial phylogenies based on overlapping genes. BMC Bioinformatics 2007; 8:266 10.1186/1471-2105-8-266 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Jiang LW, Lin KL, Lu CL. OGtree: a tool for creating genome trees of prokaryotes based on overlapping genes. Nucleic Acids Res 2008;36(Web Server issue):W475-80. [DOI] [PMC free article] [PubMed]
- 9.Katoh K, Kuma K, Toh H, Miyata T. MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res 2005; 33:511-518 10.1093/nar/gki198 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 2011; 28:2731-2739 10.1093/molbev/msr121 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Ewing WH. Edwards and Ewing’s identification of Enterobacteriaceae 4th ed. Burgess Publishing Co., New York, NY. 1986. [Google Scholar]
- 12.Field D, Garrity G, Gray T, Morrison N, Selengut J, Sterk P, Tatusova T, Thomson N, Allen MJ, Angiuoli SV, et al. The minimum information about a genome sequence (MIGS) specification. Nat Biotechnol 2008; 26:541-547 10.1038/nbt1360 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Woese CR, Kandler O, Wheelis ML. Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya. Proc Natl Acad Sci USA 1990; 87:4576-4579 10.1073/pnas.87.12.4576 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Garrity GM, Bell JA, Lilburn T. Phylum XIV. Proteobacteria phyl. nov. In: Garrity GM, Brenner DJ, Krieg NR, Staley JT (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 2, Part B, Springer, New York, 2005, p. 1. [Google Scholar]
- 15.Validation of publication of new names and new combinations previously effectively published outside the IJSEM. List no. 106. Int J Syst Evol Microbiol 2005; 55:2235-2238 10.1099/ijs.0.64108-0 [DOI] [PubMed] [Google Scholar]
- 16.Garrity GM, Bell JA, Lilburn T. Class III. Gammaproteobacteria class. nov. In: Garrity GM, Brenner DJ, Krieg NR, Staley JT (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 2, Part B, Springer, New York, 2005, p. 1. [Google Scholar]
- 17.Garrity GM, Holt JG. Taxonomic Outline of the Archaea and Bacteria In: Garrity GM, Boone DR, Castenholz RW (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 1, Springer, New York, 2001, p. 155-166. [Google Scholar]
- 18.Skerman VBD, McGowan V, Sneath PHA. Approved Lists of Bacterial Names. Int J Syst Bacteriol 1980; 30:225-420 10.1099/00207713-30-1-225 [DOI] [PubMed] [Google Scholar]
- 19.Rahn O. New principles for the classification of bacteria. Zentralbl Bakteriol Parasitenkd Infektionskr Hyg 1937; 96:273-286 [Google Scholar]
- 20.Judicial Commission Conservation of the family name Enterobacteriaceae, of the name of the type genus, and designation of the type species OPINION NO. 15. Int Bull Bacteriol Nomencl Taxon 1958; 8:73-74 [Google Scholar]
- 21.Lignieres J. Maladies du porc. Bulletin of the Society for Central Medical Veterinarians 1900; 18:389-431 [Google Scholar]
- 22.Le Minor L, Rohde R. Genus IV. Salmonella Lignieres 1900, 389. In: Buchanan RE, Gibbons NE (eds), Bergey's Manual of Determinative Bacteriology, Eighth Edition, The Williams and Wilkins Co., Baltimore, 1974, p. 298-318. [Google Scholar]
- 23.Judicial Commission of the International Committee on Systematics of Prokaryotes The type species of the genus Salmonella Lignieres 1900 is Salmonella enterica (ex Kauffmann and Edwards 1952) Le Minor and Popoff 1987, with the type strain LT2, and conservation of the epithet enterica in Salmonella enterica over all earlier epithets that may be applied to this species. Opinion 80. Int J Syst Evol Microbiol 2005; 55:519-520 10.1099/ijs.0.63579-0 [DOI] [PubMed] [Google Scholar]
- 24.Le Minor L, Popoff MY. Request for an Opinion. Designation of Salmonella enterica sp. nov., nom. rev., as the type and only species of the genus Salmonella. Int J Syst Bacteriol 1987; 37:465-468 10.1099/00207713-37-4-465 [DOI] [Google Scholar]
- 25.Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 2000; 25:25-29 10.1038/75556 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.http://www.illumina.com/technology/sequencing_technology.ilmn
- 27.Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Formsma K, Gerdes S, Glass EM, Kubal M, et al. The RAST Server: rapid annotations using subsystems technology. BMC Genomics 2008; 9:75 10.1186/1471-2164-9-75 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Delcher AL, Bratke KA, Powers EC, Salzberg SL. Identifying bacterial genes and endosymbiont DNA with Glimmer. Bioinformatics 2007; 23:673-679 10.1093/bioinformatics/btm009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Stothard P, Wishart DS. Circular genome visualization and exploration using CGView. Bioinformatics 2005; 21:537-539 10.1093/bioinformatics/bti054 [DOI] [PubMed] [Google Scholar]