Abstract
Streptomyces lavendulae produces complestatin, a cyclic peptide natural product that antagonizes pharmacologically relevant protein–protein interactions including formation of the C4b,2b complex in the complement cascade and gp120-CD4 binding in the HIV life cycle. Complestatin, a member of the vancomycin group of natural products, consists of an α-ketoacyl hexapeptide backbone modified by oxidative phenolic couplings and halogenations. The entire complestatin biosynthetic and regulatory gene cluster spanning ca. 50 kb was cloned and sequenced. It consisted of 16 ORFs, encoding proteins homologous to nonribosomal peptide synthetases, cytochrome P450-related oxidases, ferredoxins, nonheme halogenases, four enzymes involved in 4-hydroxyphenylglycine (Hpg) biosynthesis, transcriptional regulators, and ABC transporters. The nonribosomal peptide synthetase consisted of a priming module, six extending modules, and a terminal thioesterase; their arrangement and domain content was entirely consistent with functions required for the biosynthesis of a heptapeptide or α-ketoacyl hexapeptide backbone. Two oxidase genes were proposed to be responsible for the construction of the unique aryl-ether-aryl-aryl linkage on the linear heptapeptide intermediate. Hpg, 3,5-dichloro-Hpg, and 3,5-dichloro-hydroxybenzoylformate are unusual building blocks that repesent five of the seven requisite monomers in the complestatin peptide. Heterologous expression and biochemical analysis of 4-hydroxyphenylglycine transaminon confirmed its role as an aminotransferase responsible for formation of all three precursors. The close similarity but functional divergence between complestatin and chloroeremomycin biosynthetic genes also presents a unique opportunity for the construction of hybrid vancomycin-type antibiotics.
Keywords: Streptomyces, vancomycin, DNA sequence, antibiotic biosynthesis, peptide synthetase
A large group of the low molecular weight bioactive peptides has been found to be biosynthesized nonribosomally by microorganisms in nature (1, 2). Enzymes responsible for assembly of the peptide backbone of these biologically important secondary metabolites are nonribosomal peptide synthetases (NRPSs). These megasynthetases are organized as giant modular multifunctional enzyme complexes and are responsible for the biosynthesis of antibiotics such as cyclosporin, bleomycin, and vancomycin (3–5). The genes encoding many NRPSs have been cloned over the past decade (1, 2).
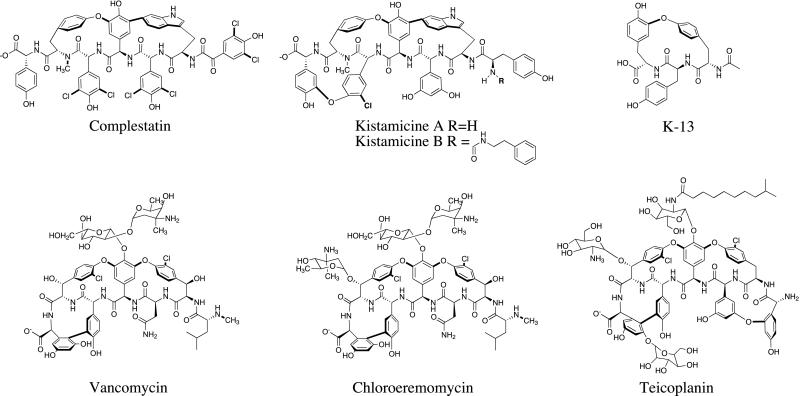
Unlike the cyclic and branched peptides, vancomycin, chloroeremomycin (Cl-E), and complestatin belong to a unique subgroup of peptide natural products biosynthesized by NRPSs (Fig. 1). They are linear nonribosomal peptides further tailored by oxidative phenolic coupling, which yields the rigid crosslinked architecture that allows high-affinity complexation with specific biological targets. Complestatin, which has potent anticomplement activity, was isolated from Streptomyces lavendulae (6). Additionally, it is the first gp120-CD4 binding inhibitor of microbial origin (7), and potentiates fibrinolysis (8). Our choice of the complestatin biosynthetic pathway was dictated by several considerations. First, the lack of a glycosyl moiety in complestatin simplifies its analysis and manipulation, as compared with vancomycin and Cl-E. Second, notwithstanding the obvious relationships between complestatin and vancomycin, there are notable differences that can be exploited via combinatorial biosynthesis. In particular, the choice of residues 1, 2, 3, 5, and 7 in the two heptameric backbones is different (Fig. 1). Moreover, the enzymes catalyzing oxidative coupling and halogenation of the two backbones, which presumably play key roles in stereochemical control, were predicted to be related but distinct. Finally, the established track record of complestatin as a modulator of a variety of pharmacologically interesting protein–protein interactions makes it an attractive target for analoging.
Figure 1.
Vancomycin and complestatin group of natural products.
Here we describe the cloning, sequence analysis, and functional verification of genes involved in complestatin biosynthesis. The potential utility of these genes for manipulating the structures of complestatin and vancomycin is also discussed.
Materials and Methods
Bacterial Strains, Culture Conditions, and Vectors.
Genomic DNA of S. lavendulae was isolated from a culture grown in yeast extract/malt extract medium containing 0.5% glycine (100-ml culture in a 500-ml flask), using the Qiagen genomic DNA purification kit (Qiagen, Chatsworth, CA) with a treatment of lysozyme (Sigma) and proteinase K (GIBCO/BRL) for cell lysis (9). DNA manipulations were performed in Escherichia coli XL1 Blue (Stratagene) by using standard culture conditions. E. coli XL1 Blue MRF′ kan strain was used for the construction of λ phage cosmid library (Stratagene).
Transformation of E. coli and DNA Manipulations.
For the construction of a genomic library, chromosomal DNA of S. lavendulae was partially digested with Sau3AI, and DNA fragments of 30–50 kb were purified by gel electrophoresis. The DNA fragments then were pooled and ligated with SuperCos-1 cosmid vector (Stratagene) previously digested with XbaI, treated with calf intestine alkaline phosphatase, and digested with BamHI. The resulting ligation mixture was packaged into λ phage, followed by phage transfection into E. coli XL1 Blue MRF′ kan strain by using protocols described in the Gigapack III XL Packaging Kit (Stratagene). Individual colonies containing cosmids were separately maintained at 4°C in 50 96-well microtiter plates.
Screening of the S. lavendulae Library.
Marahiel et al. (1) previously reported highly conserved core motifs of the catalytic domains of cyclic and branched peptide synthetases (designated An, Tn, Cn, En, Mn, and TE for adenylation, thiolation, condensation, epimerization, methylation, and thioesterase domains, n = assigned number of the motif). Based on multiple sequence alignments of several reported peptide synthetases and the chloroeremomycin NRPS, the conserved regions C5, A2, A3, A5, A7, A8, M1, M2, M3, T, E2, and E5 were targeted for degenerate primer design. Similar sequence comparisons of the Cl-E P450 oxidase and other homologs gave three conserved regions, P1, P2, and P3. Paired (forward and reverse) combinations of degenerate oligonucleotides derived from these conserved regions were used to amplify probes from S. lavendulae genomic DNA. Of five PCR amplimers, AA, AE, and MT (derived from A3-A7, A8-E2, and M2-T degenerate primers, respectively) showed high homology to NRPS domains, whereas P1P2 and P1P3 (derived P1-P2 and P1-P3 degenerate primers) were highly homologous to P450-related oxidases. The sequences of the positive degenerate oligonucleotides were as follows: A3 forward primers (5′-TAC ACS AGC GGS AGC ACS GG-3′), A7 reverse primers (AVG TCS CCS GTS CKG TAC ATS C-3′), A8 forward primer (5′-CAG GTS AAG RTS MGS GGS TWC MG-3′), E2 reverse primers (5′-GTC SAC SRM SAR GTG GTG-3′), M2 forward primers (5′-AAC GAG YTS AGC RSS TAC MGS TAC-3′), P1 forward primers (5′-GAC CCS CCS GAG CAC ACS MGS YTS MG-3′), P2 reverse primers (5′-GCA STG GTG SAY SCC GTG SCC GAA-3′), P3 reverse primers (5′-ARS CKS ARS SYS GGG AAS CK-3′). The amplimers were labeled with digoxigenin (DIG) and used in Southern blot hybridizations to screen 3,000 cosmids from the library according to the protocols described in the DIG DNA labeling and Detection Kit from Roche Molecular Biochemicals.
Cloning and Sequence Analysis of the Complestatin Gene Cluster.
Three cosmids, designated pHC-E46, pHC-H75, and pHC-D27, hybridized to one or more of the above probes, and mapped onto a single contig. The inserts in each cosmid were digested with various enzymes to deduce a physical map of the contig. Subclones were derived by digestion with restriction enzymes BamHI and SstI, and the resulting double-stranded plasmids were fully sequenced. Each base pair in the complestatin gene cluster was sequenced a minimum of three times on both strands. Compilation and assembly of sequences were done by using the sequencher program (Gene Codes, Ann Arbor, MI). DNA sequences were analyzed by using macvector sequence analysis software (Oxford Molecule Group, Cambridge, U.K.). DNA and protein sequence homology searches were performed by use of the blast server at National Center for Biotechnology Information (Bethesda, MD.).
Cloning, Expression, and Purification of 4-Hydroxyphenylglycine Transaminon (HpgT).
The proposed coding region for HpgT of the complestatin cluster was amplified by PCR from pHC-E46 with primers HpgTNtermNdeI (5′-GGGAATTCCATATGATACTGGCCCCCATGCAGATC-3′) and HpgTCtermXhoI (5′-CGCCGCTCGAGTCAGTCCGGTGCCGACGACGGCGA-3′). The introduced NdeI and XhoI restriction sites of each primer are underlined. The amplified products were cloned by using the NdeI and XhoI sites (underlined) into pET16b, which encodes an N-terminal deca-histidyl tag. Cultures of BL21(DE3)/pHpgT were grown at 15°C for 72 h. The cells were harvested by centrifugation and resuspended in 100 ml of binding buffer (20 mM Tris⋅HCl, pH 8.0/500 mM NaCl/10 mM imidazole). After sonication, the cell lysates were centrifuged (45 min at 10,000 × g) and loaded on a nickel-chelating Sepharose fast flow (Amersham Pharmacia) column preequilibrated with charge buffer (20 mM Tris⋅HCl, pH 8.0/100 mM NaCl/50 mM nickel sulfate) (150 ml) and binding buffer (150 ml). Cell lysates were loaded onto the column at 5 ml/min. The column was then washed at 5 ml/min with 200 ml of binding buffer, 200 ml of wash buffer (20 mM Tris⋅HCl, pH 8.0/500 mM NaCl/80 mM Imidazole), and 150 ml of binding buffer. The protein was eluted at 5 ml/min with 300 ml of strip buffer (40 mM Tris⋅HCl, pH 8.0/500 mM NaCl/100 mM EDTA). The protein was dialyzed into 10 ml of Tris⋅HCl (pH 7.5), 10 mM NaCl and assayed for activity. Purified ComAT (>90%) could be prepared in 20-mg quantities from 1.5 liters of culture.
HpgT Assays.
All HPLC reactions were analyzed on a Beckman System Gold HPLC using a Vydac C18 small pore column. The solvent system for the analysis of the HpgT reaction used buffer A (water with 0.1% trifluoroacetic acid) and buffer B (acetonitrile with 0.1% trifluoroacetic acid). The profile for separation was a 35-min linear gradient from 0% to 30% buffer B in buffer A. The column was then washed with 100% buffer B and reequilibrated with 100% buffer A. Flow rate for the entire profile was 1 ml/min.
Reactions of HpgT were performed as follows. HpgT (100 μl at 1 mg/ml) was added to a reaction consisting of 1 mM of the amino donor and 1 mM of the amino acceptor and 10 μM of pyridoxal-5-phosphate in 20 mM Tris, pH 7.5. The reactions were allowed to incubate at 25°C for 24 h. The samples were acidified with trifluoroacetic acid to precipitate the protein and protonate the carboxylate-containing compounds. The reactions then were loaded onto an analytical HPLC column with an autosampler using the above conditions.
l-p-Hydroxyphenylglycine and d-p-hydroxyphenylglcyine were purchased from Sigma/Aldrich. 3,5-Dichloro-p-hydroxyphenylglycine was the kind gift of Dewey McCafferty, University of Pennsylvania. The products of the aminotransferase reaction (p-hydroxybenzoylformate and 3,5-dichloro-p-hydroxybenzoylformate) were identified by matrix-assisted laser desorption ionization/time of flight. The presence of l-tyrosine was identified by coinjection with authentic material (Sigma/Aldrich).
Results and Discussions
Cloning of the Complestatin Biosynthetic Gene Cluster.
To clone the complestatin biosynthetic genes, we included characteristics of some biosynthetic genes of the vancomycin and complestatin group in our probe design, so as to differentiate our clones from other NRPS genes that might exist in S. lavendulae. Specifically, we took advantage of the available genetic information regarding the biosynthesis of Cl-E (5). Sequence comparisons between the Cl-E NRPS genes and those involved in other cyclic and branched peptide antibiotic pathways resulted in a set of more “specific” degenerate primers (see Materials and Methods). In addition, degenerate primers were designed based on the conserved sequences of the Cl-E cytochrome P450 oxidases and other related cytochrome P450 oxidases (5). Of the PCR amplimers cloned and sequenced from S. lavendulae, three amplimers (designated AE, AA, and MT) were found to be highly similar (50–65% aa identity) to the Cl-E NRPS genes, and two amplimers (designated P1P2 and P1P3) were homologous (ca. 55% identity at the amino acid level) to the Cl-E cytochrome P450 oxidases.
Two cosmids, pHC-E46 (3′ end, ca. 35-kb insert) and pHC-H75 (5′ end, ca. 40-kb insert, were found to hybridize to all these probes and overlapped to yield a contig of ca. 54 kb (Fig. 2A). The 3′-end sequence of pHC-H75 was homologous to an NRPS adenylation domain. We therefore rescreened the cosmid library of S. lavendulae by using a probe from the 3′ end of pHC-H75. An additional cosmid, pHC-D27 (ca. 34-kb insert), was selected from a group of positive clones, resulting in a three-cosmid contig of ca. 70 kb. As summarized below, this contig covers the entire biosynthetic gene cluster of complestatin.
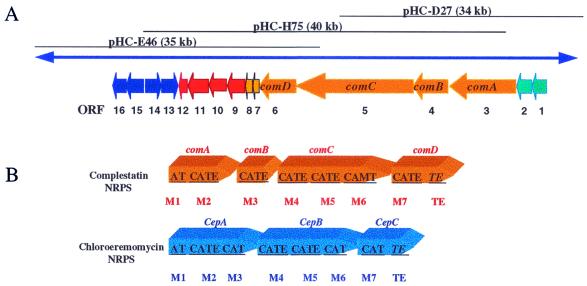
Figure 2.
(A) The three cosmids containing genes involved in the biosynthesis of complestatin. The figure is not drawn to scale. For proposed functions of assigned ORFs, see Table 1. (B) Arrangement of genes encoding NRPSs in the biosynthesis of complestatin and chloroeremomycin. A, Adenylation domain; T, thiolation domain (peptidyl carrier domain); C, condensation domain; E, epimerization domain; M, methylation domain; TE, thioesterase domain; m: NRPS module. The domain assignments and boundaries were determined by sequence comparisons of known NRPS genes. The figure is not drawn to scale.
Organization of the Genes Involved in the Biosynthesis of Complestatin.
The biosynthetic gene cluster of complestatin covers 48.7 kb and includes 16 ORFs. The organization and nature of these ORFs, which are colinear in most part, is shown in Fig. 2 and Table 1. On either side of the complestatin gene cluster, noncoding DNA stretching over at least 2 kb was observed, suggesting that the ends of the gene cluster had been physically mapped. Below we describe the properties of the gene products encoded by this gene cluster.
Table 1.
Summary of genes identified on cosmids pHC-D27, pHC-H75, and pHC-E46 obtained from S. lavandulae
| ORF number | Protein name | Cep homologue | Proposed function | Position start-stop, bp | Cosmid | Number base pairs | Number amino acids |
|---|---|---|---|---|---|---|---|
| 1 | ComG | Transcriptional regulator | 2020-3066 | D27 | 1,047 | 348 | |
| 2 | ComL | Cep M | ABC transporter | 3491-5617 | D27 | 2,127 | 708 |
| 3 | ComA | Cep A | Peptide synthetase | 5664-12012 | D27 | 6,351 | 2,116 |
| 4 | ComB | Cep A | Peptide synthetase | 12011-16606 | D27 & H75 | 4,596 | 1,531 |
| 5 | ComC | Cep B | Peptide synthetase | 16705-314-1 | H75 | 14,697 | 4,898 |
| 6 | ComD | Cep C | Peptide synthetase | 31493-37933 | H75 & E46 | 6,495 | 2,164 |
| 7 | ComE | Cep D | Hypothetical protein in M. tuberculosis | 37938-38159 | H75 & E46 | 222 | 73 |
| 8 | ComF | Integral membrane ion antiporter | 38198-39475 | H75 & E46 | 1,278 | 425 | |
| 9 | ComH | Cep H | Nonheme halogenase | 39504-40997 | H75 & E46 | 1,494 | 497 |
| 10 | ComI | Cep F, G | P450-related oxidase | 41015-42208 | H75 & E46 | 1,194 | 397 |
| 11 | ComJ | Cep E | P450-related oxidase | 42222-43493 | H75 & E46 | 1,272 | 423 |
| 12 | ComK | Ferredoxin | 43538-43756 | H75 & E46 | 219 | 72 | |
| 13 | Hmo | Hmo | p-Hydroxymandelate oxidase | 44952-43834 | H75 & E46 | 1,119 | 372 |
| 14 | HmaS | HmaS | p-Hydroxymandelate synthase | 45955-44939 | H75 & E46 | 1,017 | 338 |
| 15 | HpgT | HpgT | p-Hydroxypenylglycine aminotransferase | 46193-47539 | E46 | 1,347 | 448 |
| 16 | PD | PD | Prephenate dehydrogenase | 47536-48684 | E46 | 1,149 | 382 |
The NRPS region.
The gene cluster includes a region of about 32.3 kb, encoding four proteins with homology to known NRPSs. These colinear genes were designated comA, comB, comC, and comD (Fig. 2B and Table 1). Together, they include a loading module (module 1), followed by six downstream extension modules (modules 2–7), and a C-terminal thioesterase domain. The number of modules corresponds precisely to the number of monomers, one α-ketoacid, and 6 aa in the α-ketoacyl hexapeptide backbone. Moreover, the domain arrangement of each module is also in an exact agreement with the enzymatic functions that would be required to construct the peptide backbone of complestatin. For example, the unique methyltransferase domain in module 6 would be responsible for the N-methylation of the tyrosine6 residue. Likewise, the complestatin backbone is made up of five d-amino acids; the number and location of epimerase domains (E domains) is exactly as predicted for activation of l-monomers and epimerization on the assembly line.
The boundaries and functions of the 27 domains that comprise the complestatin NRPS were predicted based on sequence alignments with other known NRPSs, especially the Cl-E NRPS (1, 2, 5). Overall, the complestatin NRPS is most similar in sequence and domain organization to the Cl-E NRPS. However, several notable differences can be observed. The complestatin NRPS gene cluster encodes four polypeptides, whereas the Cl-E NRPS gene cluster has only three ORFs. Moreover, there is no internal methylation domain in the Cl-E NRPS. Finally, modules 3 and 7 of the complestatin NRPS contain epimerization domains, whereas their counterparts in the Cl-E NRPS do not. It is not yet experimentally validated whether the A domain of module 1 activates the 3,5-dichloro-4-hydroxybenzoylformate (see below) or 3,5-dichloro-4-hydroxy-l-phenylglycine, which is subsequently oxidatively deaminated.
The tailoring enzyme region: Cytochrome P450-related oxidases.
Downstream of the NRPS genes were two genes, ComI and ComJ, homologous to known cytochrome P450 oxidases (Table 1 and Fig. 2A). In particular, ComI and ComJ show strong sequence similarity to the gene products of two Cl-E ORFs, CepF (58% similarity and 44% identity) and CepE (66% similarity and 53% identity), respectively. The relative positions and orientation of ComI and ComJ are, however, opposite to those of the Cl-E CepF and CepE. The presence of two cytochrome P450-related oxidases agrees with the prediction that there should be two oxidative couplings (C-O-C and C-C) in the complestatin biosynthetic pathway, as judged from the chemical structure of complestatin (Fig. 1). Interestingly, the complestatin cluster lacks a Cl-E CepG homolog, which is more closely related to Cl-E CepF than to Cl-E CepE. Because Cl-E biosynthesis requires two C-O-C couplings and one C-C coupling, we propose that the P450 oxidases encoded by ComJ, Cl-E CepF, and Cl-E CepG catalyze C-O-C couplings, whereas the ComI and Cl-E CepE gene products catalyze C-C couplings.
Heterologous expression of ComI and ComJ in Escherichia coli yielded soluble C-terminally His-tagged proteins. They were purified to homogeneity and judged to contain stoichiometric levels of heme based on their red color and characteristic UV-visible spectrum (data not shown). However, because the natural substrates of these enzymes were unavailable, their precise functions could not be confirmed.
The tailoring enzyme region: Ferredoxin.
The catalytic cycle of cytochrome P450 enzymes requires a ferredoxin and a corresponding NAD(P)H-dependent ferredoxin oxidoreductase. In contrast to the Cl-E gene cluster, which lacks dedicated ferredoxin or oxidoreductase genes (and must therefore “borrow” these activities from other loci in the chromosome of the producing organism), the complestatin gene cluster encodes a ferredoxin (ComK, 216 bp) immediately downstream of the two P450 genes (Fig. 2A and Table 1). The absence of a dedicated oxidoreductase gene, however, suggests that ComK pairs with an endogenous ferredoxin oxidoreductase in S. lavendulae.
The tailoring enzyme region: Halogenase.
Straddled between the NRPS and P450-oxidase genes of the complestatin gene cluster was found comH, a homolog of a family of nonheme halogenase genes. ComH is presumably involved in the dichlorination of hydoxyphenylglycine residues in the complestatin backbone, although the precise timing of this reaction in the biosynthetic cascade remains unknown. (The substrate for the halogenase could be the hydroxyphenylglycine amino acid precursor, an NRPS-bound growing chain, the full-length linear peptide, or even the cyclized peptide.) To date, three types of halogenases have been described—heme-containing, vanadium-containing, and metal-free halogenases (10). Based on sequence analysis the putative halogenase encoded by comH is homologous to the metal-free halogenases. However, it does not contain the catalytic Asp-His-Ser triad, analogous to chloroperoxidases with x-ray structures bearing the α,β-hydrolase fold that are believed to have a serine protease-like mechanism (11). This finding suggests that the complestatin halogenase may use a novel mechanism for its enzymatic activity. blast searching revealed sequence similarity between ComH and known epoxidase genes (e.g., squalene epoxidase AAC32430, 41% similarity and 25% identity in amino acid) in addition to oxygenase and hydroxylase genes. One therefore may propose an enzymatic mechanism for the complestatin halogenase mimicking that of an epoxidase (12). Interestingly, a single halogenase encoded by comH appears to catalyze dichlorination in complestatin biosynthesis, whereas two genes (ORF10 and ORF18) homologous to nonheme halogenases were identified in the Cl-E gene cluster. The Cl-E CepH product is homologous to ComH, whereas the Cl-E ORF18 contains the catalytic Asp-His-Ser triad. Because only one type of (mono-)chlorination occurs in Cl-E biosynthesis, perhaps one of the two Cl-E ORFs is nonfunctional.
Biosynthesis of the 4-hydroxyphenyl glycine (Hpg) precursor.
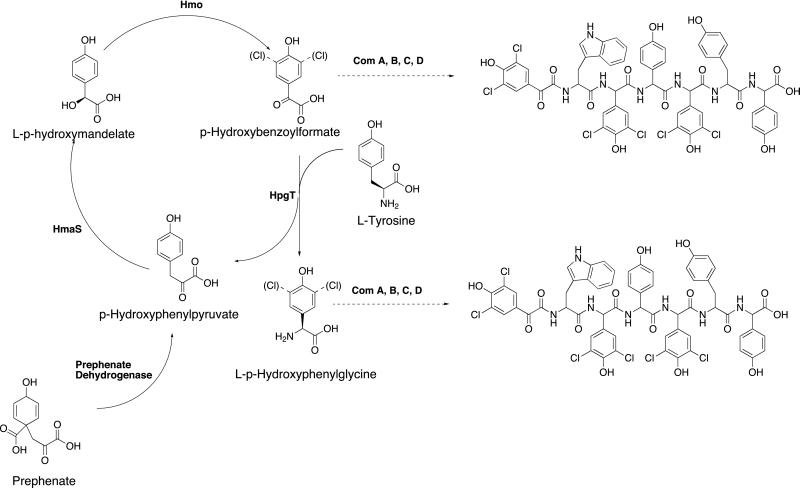
The complestatin gene cluster was also found to contain four ORFs, Hmo, HmaS, HpgT, and PD, homologous to genes encoding p-hydroxymandelate oxidase, p-hydroxymandelic acid synthase, p-hydroxyphenylglycine aminotransferase, and prephenate dehydrogenase, respectively (Fig. 2A and Table 1). Homologs of these genes also are present in the Cl-E gene cluster (5). Hpg is used as a building block for both complestatin and Cl-E biosynthesis. Previous experiments using labeled tyrosine and p-hydroxybenzoylformate showed that tyrosine was incorporated into the Hpg building block of vancomycin and led to the hypothesis that p-hydroxybenzoylformate was an intermediate in this pathway (13). More recently, biochemical analysis of these purified enzymes from the Cl-E gene cluster have led to the definition of the Hpg biosynthetic pathway (Fig. 3) (14). In this unusual pathway l-tyrosine is stoichiometrically converted into Hpg via a catalytic cycle that is primed by prephenate, a commonly available intermediate from the shikimate pathway. Because Hpg is used as a building block in the formation of both the Cl-E and complestatin backbones, we propose that Hmo, HmaS, HpgT, and PD from the complestatin gene cluster are orthologs of the corresponding enzymes from the Cl-E gene cluster.
Figure 3.
Biosynthetic pathway for l-hydroxyphenylglycine. The potential destination of the p-hydroxybenzoylformate (or 3,5-dichloro-p-hydroxybenzoylformate) and Hpg (or 3,5-diClHpg) are shown in the complestatin backbone.
In addition to Hpg, the complestatin peptide contains 3,5-dichlorohydroxyphenylglycine (3,5-diClHpg) and 3,5-dichlorobenzoylformate which are not present in chloroeremomycin. It is possible the catalytic cycle used for the production of Hpg is also responsible for the production of 3,5-diClHpg and 3,5-dichlorbenzoylformate. These three related monomer units constitute five of the seven monomers in the final complestatin backbone (Fig. 1). Because it is unclear at this stage when the halogenation occurs, the specificity of the HpgT could lend insight into the origins of Hpg and the dichlorinated analogues.
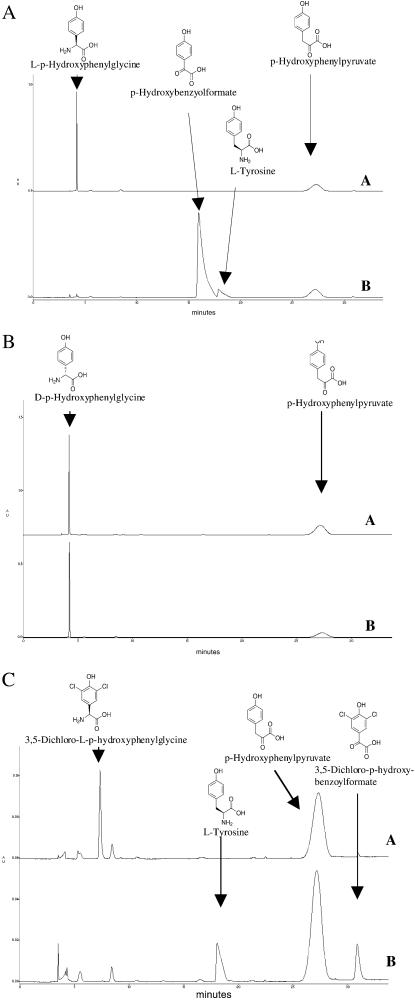
To explore the origins of the Hpg, 3,5-diClHpg, and 3,5-dichlorobenzoylformate, the HpgT gene was overexpressed in E. coli, and the enzyme was purified and reconstituted (see Materials and Methods). The activity of the enzyme was assayed by using l-Hpg, d-Hpg, and and 3,5-dichloro-l-Hpg as substrates. In each case hydroxyphenylpyruvate was used as a cosubstrate, and the reaction was allowed to proceed to equilibrium. Equilibrium concentrations of the two substrates and the two products [l-tyrosine and hydroxybenzoylformate (or its dichloro analog)] were monitored by HPLC. As can be seen in Fig. 4, both l-Hpg and 3,5-dichloro-l-Hpg are substrates for HpgT, whereas d-Hpg is not. The preference for the l-amino acid is consistent with the results from the p-hydroxyphenylglycine aminotransferase in the Cl-E cluster (14).
Figure 4.
Biochemical analysis of HpgT. (A) HPLC trace showing the reactants and products in the l-Hpg reaction. A is the reaction at time 0, and B is the reaction at equlibrium. (B) HPLC trace showing the reactants and products in the d-Hpg reaction. A is the reaction at time 0, and B is the reaction at equilibrium. (C) HPLC trace showing the reactants and products in the 3,5-dichloro-l-Hpg reaction. A is the reaction at time 0, and B is the reaction at equilibrium (see text for details).
These results indicate that l-Hpg, l-3,5-diClHpg, and 3,5-dichlorobenzoylformate can be generated or consumed by HpgT. This leaves open the possibility that all three monomers are generated by this pathway. This result is consistent with the hypothesis that the adenylation domain in module 1 of ComA (Fig. 2A) would activate and load 3,5-dichlorobenzoylformate. The results generated with HpgT are consistent with a single 4-hydroxyphenylglycine biosynthetic pathway (consisting of Hmo, HmaS, HpgT, PD, and ComH) generating five of the seven requisite monomers for the assembly of complestatin.
Proposed Overall Biosynthetic Pathway of Complestatin.
In the metabolic pool of S. lavendulae, the unusual amino acid Hpg is biosynthesized from the shikimate pathway. Derivatives of Hpg, 3,5-dichloro-Hpg, and 3,5-dichloro-hydroxybenzoylformate also are produced concomitantly. These three building blocks, together with tyrosine and tryptophan, are selected and loaded individually on specific NRPS modules. Peptide bond formation occurs sequentially from module 1 to module 7, followed by thioesterase-catalyzed termination of the full-length α-ketoacyl hexapeptide (or heptapeptide) chain. During the course of chain elongation particular aminoacyl-S-thioester domain acyl enzymes are epimerized and methylated as dictated by the presence and microlocation of epimerase and methyltransferase domains. The two cytochrome P450 oxidases encoded by ComI and ComJ then tailor the linear α-ketoacylhexapeptide, carrying out two types of oxidative phenolic couplings. (However, the possibility that the P450 oxidases may cyclize the growing peptide on the NRPS assembly line before chain termination cannot be excluded.) The topology of the crosslinked peptide scaffold from action of these oxidases presumably allows complestatin to adopt a unique β-strand conformation to interact with specific biological targets. A putative transcriptional regulator also is encoded within the complestatin gene cluster and is presumed to regulate its biosynthesis. An ABC transporter homolog, believed to be required for extracellular export of complestatin, also was identified.
Similarities and Differences Between the Complestatin and Chloroeremomycin Gene Clusters: Implications for Combinatorial Biosynthesis.
Among the more than 200 members of the vancomycin group of natural products discovered so far (15), complestatin and Cl-E are the only two whose gene clusters have been cloned and sequenced. With regard to the organization of the NRPS genes, complestatin differs from Cl-E principally in two aspects. First, whereas the first three modules of the complestatin NRPS are distributed over two polypeptides, they reside within a single polypeptide in the Cl-E NRPS. Second, although the heptapeptides of both complestatin and Cl-E are methylated (Fig. 1), this reaction apparently is carried out by an internal methylation domain within an NRPS module of the complestatin NRPS, whereas the Cl-E NRPS lacks a corresponding internal methylation domain. Instead, an independent ORF (ORF16) several genes away from the NRPS genes encodes an external methyltransferase within the Cl-E gene cluster (5). Notwithstanding these functional differences between the complestatin and Cl-E NRPSs, the strong similarity between these two megasynthetases should be invaluable for the construction of novel vancomycin and complestatin analogs by combinatorial biosynthesis. In particular, the linear peptide backbones of the two natural products differ at residues 2, 3, and 7, whereas residue 1 is an N-methyl-d-Leu in vancomycin and a 3,5-dichlor-4-hydroxybenzoylformate residue in complestatin. Thus, combinatorial construction of hybrid NRPS modules at these positions should facilitate the generation of new analogs of either natural product. In parallel, it may be possible to productively exchange the cytochrome P450 enzymes as well, giving rise to alternatively cyclized compounds. Finally, certain tailoring enzymes such as the halogenases and the tyrosine hydroxylase (from Cl-E) may be able to recognize and modify the corresponding substrates in the other pathway.
Acknowledgments
This research was supported by a grant from the Schering Plough Research Institute.
Abbreviations
- NRPS
nonribosomal peptide synthetase
- Cl-E
chloroeremomycin
- HpgT
4-hydroxyphenylglycine transaminon
- Hmo
4-hydroxymandelate oxidase
- PD
prephenate dehydrogenase
- HmaS
4-hydroxymadelate synthase
- Hpg
4-hydroxyphenyl glycine
- 3,5-diClHpg
3,5-dichlorohydroxyphenylglycine
Footnotes
Data deposition: The sequence reported in this paper has been deposited in the GenBank database (accession no. AF386507).
References
- 1.Marahiel M A, Stachelhaus T, Mootz H D. Chem Rev. 1997;97:2651–2673. doi: 10.1021/cr960029e. [DOI] [PubMed] [Google Scholar]
- 2.von Döhren H, Keller U, Vater J, Zocher R. Chem Rev. 1997;97:2675–2705. doi: 10.1021/cr9600262. [DOI] [PubMed] [Google Scholar]
- 3.Weber G, Schörgendorfer K, Schneider-Scherzer E, Leitner E. Curr Genet. 1994;26:120–125. doi: 10.1007/BF00313798. [DOI] [PubMed] [Google Scholar]
- 4.Du L, Chen M, Sanchez C, Shen B. FEMS Microbiol Lett. 2000;189:171–175. doi: 10.1111/j.1574-6968.2000.tb09225.x. [DOI] [PubMed] [Google Scholar]
- 5.Wageningen A M A, Kirkpatrick P N, Williams D H, Harris B R, Kershaw J K, Lennard N J, Jones M, Jones S J M, Solenberg P J. Chem Biol. 1998;5:155–162. doi: 10.1016/s1074-5521(98)90060-6. [DOI] [PubMed] [Google Scholar]
- 6.Seto H, Fujioka T, Furihata K, Kaneko I, Takahashi S. Tetrahedron Lett. 1989;30:4987–4990. [Google Scholar]
- 7.Matsuzaki K, Ikeda H, Ogino T, Matsumoto A, Woodruff H B, Tanaka H, Omura S. J Antibiot. 1994;47:1173–1174. doi: 10.7164/antibiotics.47.1173. [DOI] [PubMed] [Google Scholar]
- 8.Tachikawa K, Hasumi K, Endo A. Thromb Haemostasis. 1997;77:137–142. [PubMed] [Google Scholar]
- 9.Hopwood D A, Bibb M J, Chater K F, Kieser T, Bruton C J, Kieser H M, Lydiate D J, Smith C P, Ward J M, Schrempf H. Genetic Manipulation of Streptomyces: A Laboratory Manual. Norwich, U.K.: The John Innes Foundation; 1985. [Google Scholar]
- 10.Littlechild J. Curr Opin Chem Biol. 1999;3:28–34. doi: 10.1016/s1367-5931(99)80006-4. [DOI] [PubMed] [Google Scholar]
- 11.Pelletier I, Altenbuchner J, Mattes R. Biochim Biophys Acta. 1995;1250:149–157. doi: 10.1016/0167-4838(95)00055-y. [DOI] [PubMed] [Google Scholar]
- 12.Tuynman A, Spelberg J L, Kooter I M, Schoemaker H E, Wever R. J Biol Chem. 2000;275:3025–3030. doi: 10.1074/jbc.275.5.3025. [DOI] [PubMed] [Google Scholar]
- 13.Hammond S J, Williamson M P, Williams D H, Boeck L D, Marconi G G. J. Chem. Soc. Chem. Commun. 1982. 344–346. [Google Scholar]
- 14.Hubbard B K, Thomas M G, Walsh C T. Chem Biol. 2000;7:931–942. doi: 10.1016/s1074-5521(00)00043-0. [DOI] [PubMed] [Google Scholar]
- 15.Rao AV R, Gurjar M K, Rao A S. Chem Rev. 1995;95:2135–2138. [Google Scholar]