Abstract
Background
Cotton (Gossypium spp.) is widely cultivated due to the important economic value of its fiber. However, extreme environmental degradation impedes cotton growth and production. Receptor-like kinase (RLK) proteins play important roles in signal transduction and participate in a diverse range of processes in response to plant hormones and environmental cues. Here, we introduced an RLK gene (GbRLK) from cotton into Arabidopsis and investigated its role in imparting abiotic stress tolerance.
Results
GbRLK transcription was induced by exogenously supplied abscisic acid (ABA), salicylic acid, methyl jasmonate, mock drought conditions and high salinity. We cloned the promoter sequence of this gene via self-formed adaptor PCR. Sequence analysis revealed that the promoter region contains many cis-acting stress-responsive elements such as ABRE, W-Box, MYB-core, W-Box core, TCA-element and others. We constructed a vector containing a 1,890-bp sequence in the 5′ region upstream of the initiation codon of this promoter and transformed it into Arabidopsis thaliana. GUS histochemical staining analysis showed that GbRLK was expressed mainly in leaf veins, petioles and roots of transgenic Arabidopsis, but not in the cotyledons or root hairs. GbRLK promoter activity was induced by ABA, PEG, NaCl and Verticillium dahliae. Transgenic Arabidopsis with constitutive overexpression of GbRLK exhibited a reduced rate of water loss in leaves in vitro, along with improved salinity and drought tolerance and increased sensitivity to ABA compared with non-transgenic Col-0 Arabidopsis. Expression analysis of stress-responsive genes in GbRLK Arabidopsis revealed that there was increased expression of genes involved in the ABA-dependent signaling pathway (AtRD20, AtRD22 and AtRD26) and antioxidant genes (AtCAT1, AtCCS, AtCSD2 and AtCSD1) but not ion transporter genes (AtNHX1, AtSOS1).
Conclusions
GbRLK is involved in the drought and high salinity stresses pathway by activating or participating in the ABA signaling pathway. Overexpression of GbRLK may improve stress tolerance by regulating stress-responsive genes to reduce water loss. GbRLK may be employed in the genetic engineering of novel cotton cultivars in the future. Further studying of GbRLK will help elucidate abiotic stress signaling pathways.
Keywords: Gossypium barbadense, Receptor-like protein kinase, Abscisic acid, Arabidopsis thaliana, Abiotic stress tolerance, Transgene
Background
Cotton is an economically important crop; cotton fiber serves as the primary raw material for the textile industry. However, as environmental degradation continues, cotton is becoming increasingly subjected to extreme drought, high salinity and temperature extremes, each of which can impede plant growth and production [1]. Plant molecular responses to abiotic stresses involve interactions and crosstalk between many molecular pathways. Hormones are important regulators of plant responses to abiotic stress. One of the most important stress-responsive plant hormones is abscisic acid (ABA). ABA is one of the most important phytohormones and is a key regulator of plant responses to various environmental stresses [1]. ABA plays an important role in the tolerance of plants to drought and high salinity, and the mechanisms underlying ABA activity have been studied extensively for several decades [2-4]. The main functions of ABA include regulating plant water balance and increasing osmotic stress tolerance by inducing seed dormancy, delaying seed germination and promoting stomatal closure [1]. To date, many stress-responsive genes have been identified; many of these genes are regulated by ABA. Recently, several ABA-deficient mutants have been reported in Arabidopsis, tobacco, tomato and maize [5]. Without stress treatment, the growth of these mutants is comparable to that of wild-type plants. Under drought stress, ABA-deficient mutants exhibit poor growth or even death [6,7]. Studying these mutants has revealed the presence of two major stress-related pathways, i.e., ABA-dependent and ABA-independent gene pathways [2]. For cotton, high throughput screening techniques such as transcriptome have been used to study the adaptability in drought. Payton et al. [8] found that the majority of stress-responsive transcripts had tissue-specific expression patterns using root and leaf under the water-deficit stress. Genome-wide transcriptomic analysis of cotton under drought stress revealed significant down- regulation of genes and pathways involved in fiber elongation and up-regulation of defense responsive genes under drought stress [9]. Using two accessions G. herbaceum with different tolerant to osmotic stress, Ranjan et al. [10] found that drought tolerance for cotton was not because of a single molecular reason but was rather due to several unique mechanisms.
Plants can receive and recognize signals and stimuli from the environment through various classes of receptors, including receptor-like kinase (RLK) proteins, which play important roles in signal transduction pathways [11]. RLK proteins kinases are encoded by a multigene family, which represents one of the largest gene families in the Arabidopsis genome, comprising at least 610 members [12], with approximately 1,131 members in rice [13]. The size variations of the RLK families have been affected by many factors such as natural and artificial selection, living environments, genome size variation, polyploidization [14]. RLKs are characterized by the presence of a signal sequence, an amino-terminal domain with a transmembrane region and a carboxyl-terminal kinase domain [15,16]. Since the discovery of the first RLK many years ago, substantial efforts have been devoted to examining a few receptor-kinase genes. RLKs participate in a diverse range of processes in response to plant hormones and environmental cues and are involved in the regulation of many physiological changes in plants such as self-incompatibility, endosperm and pollen development, brassinosteroid sensing, anthoptosis and stress and disease resistance [11,12]. Like Arabidopsis brassinosteroid insensitive 1 (BRI1) [17,18], CLAVATA1[19] and HAESA[20] control plant growth and development under normal growth conditions. Tomato Pto[21], rice Xa21[22] and Arabidopsis FLS2[23] take part in the plant defense response and in plant-microbe interactions. Many researches suggested that RLKs can play an important role in optimizing plant responses to drought stress [24-26]. Arabidopsis ARCK1[27], GHR1[28] and RPK1[29] have been implicated in controlling plant abiotic stress tolerance. SnRK2s (SNF1-related protein kinase) is a family of ABA-activated protein kinases [3,6,30]. In Arabidopsis, three members of this family, SRK2D/SnRK2.2, SRK2E/OST1/SnRK2.6 and SRK2I/SnRK2.3, regulate ABA signaling both positively and globally [6]. Rice receptor-like kinase OsSIK1 was also proved to improve drought and salt stress tolerance [31].
Cotton genes encoding protein kinases have been reported, but these genes function mainly in the regulation of cotton fiber development. To our knowledge, RLK genes associated with cotton abiotic stress have not previously been documented. The RLK gene (GbRLK) identified in this study was previously identified among a group of expressed resistance gene analogs that are induced by Verticillium dahliae (VD) in the disease-resistant cotton Gossypium barbasense cv. Hai7124 (L447 published in Gao et al., 2006) [32]. Transgenic cotton and Arabidopsis plants that overexpress GbRLK show increased resistance to VD. In the current study, we cloned the promoter region of GbRLK and analyzed the expression of GbRLK in response to abiotic stress. The results of this study suggest that GbRLK can be induced by ABA, drought stress and salinity. The overexpression of GbRLK in Arabidopsis increased plant tolerance to drought and high salinity and increased sensitivity to ABA. The analysis of this kinase gene will help elucidate the molecular basis of RLK protein function in abiotic stress tolerance in cotton. This study also provides a candidate gene for further research on ABA signaling pathways in cotton and other plants subjected to abiotic stress.
Results
Phylogenetic relationships between GbRLK and other kinase proteins
A functional GbRLK gene [NCBI accession number KC422776], which was induced by Verticillium dahliae, was previously identified among a group of expressed resistance gene analogs from the disease-resistant cotton Gossypium barbasense cv. Hai7124 using RACE [32]. The cDNA was 1536-bp in length and the predicted ORF starts at nucleotide 330 and ends at nucleotide 1409, encoding a predicted protein of 359 amino acids. BLASTx searches revealed that the predicted product shared 68% identity and 75% similarity with the soybean G-type lectin S-receptor-like serine/threonine -protein kinase (E value: 0), 56% identity and 72% similarity with the rice PK3 (E value: 1e-98), 48% and 65% (E value: 4e-73) with the wheat TAK33, and 48% and 64% (E value: 2e-71) with the barley receptor-like protein kinase, respectively. The protein possessed intact signature features of protein tyrosine kinases, and contained conserved motifs characteristic of I-XI of receptor-like protein kinases, which are possessed in the majority of protein kinases. Multiple sequence alignments using BLASTx revealed that the protein shared homology in other regions with protein kinases from various other crop species (Figure 1a). We therefore conclude that this protein may belong to the receptor-like protein kinase family and be a member of the RLK gene family. This gene was thereafter designated as G. barbadense receptor-like protein kinase (GbRLK). GbRLK contains an ATP binding site (residues 32 to 40), a D activation site (at residue 150), four N-glycosylation sites, four N-myristoylation sites, three C terminal phosphorylation sites, one amidation site, four tyrosine kinase II phosphorylation sites, and a membrane-spanning region (residues 28–46).
Figure 1.

Phylogenetic analysis of GbRLK. (a) Alignment of deduced amino acid sequences of GbRLK and other kinases. (AAB47421, Pto from tomato; BAB08490, protein kinase from Arabidopsis; AAT70497, protein kinase from peach; BAD37843, protein kinase from rice; AAD44031, PK3 from barley; AAK20741, TAK33 from barley; AAB82755, Xa21 from rice). (b) Phylogenetic relationships between GbRLK and other kinase proteins from Gossypium hirsutum and Gossypium barbadense. (c) Phylogenetic relationships between GbRLK and other reported receptor-like kinase proteins from the eudicot Arabidopsis thaliana (AtANXUR1, AEE74120; AtPBS1, AED91858; AtCERK1, AEE76532; AtPR5K, AED94289; AtACR4, AEE79920; AtWAK1, AEE30079; AtRKF3, AEC10923; AtBRI1, AEE87069; AtERECTA, AEC07825; AtCLV1, AEE35762; AtRLK5, AEE85494; AtPERK1, AEE76920; AtPEK12, AEE30401; AtCRK1, AEE29801; AtSD113, AEE28721; AtRKS1, AEE28720; AtCRK5, EFH43988; AtCRK10, AEE84719; AtRLK3, CAA09731; AtFLS2, AED95370; AtLRK13, AEE78027; AtRLK1, AED97394; AtRLK4, BAC42412; AT4G27300; AT4G11890; AT5G24080; AT4G32300; AT5G01540; AT4GO2410; AT3G59750; AT1G09970; AT1G66880; AT2G33580) and some representative members of Oryza sativa (Xa21, AAB82755), Lycopersicon esculentum (LePTO, BAB08490), and Triticum aestivum (TaLRK10, AAC49629). The phylogenetic tree was constructed using the maximum likelihood method in MEGA 5.0. Numbers on internal nodes are the percentage bootstrap support values (1000 re-sampling); only values exceeding 50% are shown.
Currently the NCBI databases contain 46 cotton sequences predicted to encode protein kinases, and among them, three (AAP47579, AA059488, AAN77146) were known to be involved in fiber development. The functions of the other sequences were unknown. The phylogenetic relationship between the GbRLK protein and the other known cotton protein kinases via the construction of phylogenetic tree revealed that GbRLK protein was most closely related evolutionally to sequences ADJ17363, AEG25668, AEA76434, ADR00582 and AAT64032 that were receptor kinase, somatic embryogenesis receptor-like kinase 1, 2, 3 protein and putative leucine-rich repeat transmembrane protein (Figure 1b).
Phylogenetic analysis was used to investigate the relationship among the GbRLK and other reported receptor-like kinase proteins from eudicot Arabidopsis thaliana and Lycopersicon esculentum monocot Oryza sativa, and Triticum aestivum. GbRLK protein was found to cluster together with other reported receptor-like kinase proteins (Figure 1c).
Temporal and spatial expression analysis of GbRLK under abiotic stress
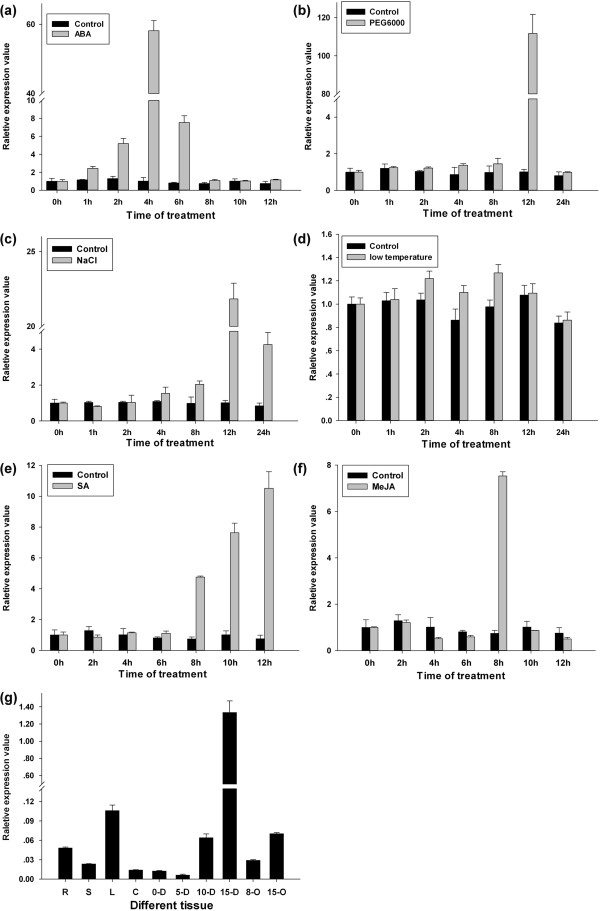
Quantitative real-time (qRT-PCR) was used to analyze the expression patterns of GbRLK in H7124 leaves exposed to ABA, drought, salinity, SA, MeJA and low temperature. As shown in Figure 2, GbRLK was induced in leaves but was differentially expressed in response to SA, MeJA and abiotic stresses. GbRLK expression was gradually up-regulated in leaves exposed to 1 to 4 h of ABA treatment; the transcription level of GbRLK increased and peaked at 4 h, after which the level of mRNA accumulation gradually declined after 6 to 12 h of ABA treatment (Figure 2a). When the seedlings were treated with PEG6000 (Figure 2b) or NaCl (Figure 2c), the transcription level of GbRLK increased rapidly and reached the highest level after 12 h of treatment, followed by a rapid decline, reaching pretreatment levels after 24 h of treatment. The expression of GbRLK was not significantly altered under low temperature treatment, suggesting that GbRLK expression is not affected by cold stress (Figure 2d). For the SA, the expression of GbRLK was induced and up-regulated at 8 h and peaked at 12 h (Figure 2e). When the seedlings were treated with MeJA, the transcription level of GbRLK peaked rapidly at 8 h and declined rapidly and reached pretreatment levels at 10 h (Figure 2f).
Figure 2.
Analysis of GbRLK expression profiles in cotton seedlings in response to different stress treatments. Expression patterns of GbRLK at different time intervals in leaves under ABA (100 μM) treatment (a) for 0, 1, 2, 4, 6, 8, 10 and 12 h; drought (20% w/v PEG6000) (b), salinity stress (200 mM NaCl ) (c) and cold stress (4°C) (d) for 0, 1, 2, 4, 8, 12 and 24 h. The treatment was carried out by spraying the plants with 2 mM SA (e) and 100 μM MeJA (f) solution at 0, 2, 4, 6, 8, 10 and 12 h. Relative gene expression levels were determined using the 2^-ΔΔCT method. The CT (cycle threshold) values for both the target and internal control genes were means of three technical replicates. ΔCt = Ct target gene-Ct EF1α. ΔΔCT = (CTtarget -CTEF1α) xh - (CTtarget - CTEF1α) 0 h. (g) analysis of GbRLK expression profiles in different developmental stages and tissue. R: root, S: stem, L: leaf, C: cotyledons, 5-D, 10-D, 15-D represented the 5, 10, 15 days post anthesis of fiber, respectively; 0-D, 8-O, 15-O represented 0, 10, 15 days ovule, respectively. EF1a (At5g60390; EF-F/R; Additional file 1: Table S1) from cotton was used as an internal control for normalization of different cDNA samples. Error bars represent standard error of means based on three independent reactions.
To further understand the temporal-spatial distribution of GbRLK gene, we analyze the expression patterns of GbRLK in H7124 different tissue. The results indicated that the GbRLK transcripts were difference significant in different tissue. The maximal transcripts of GbRLK were detected at 15 DPA (Figure 2g). This implied that the GbRLK gene may play a multi-functional role in different development stages.
Cloning and analysis of the GbRLK promoter
To further study the characteristics of GbRLK expression, we cloned the promoter sequence of GbRLK. The Sefa-PCR generated a 3,227-bp fragment, including a 347-bp region containing previously identified sequences (Additional file 1: Table S1, Additional file 2: Figure S1a). The transcriptional start site and several regulatory elements were predicted using tools including Neural Network Promoter Prediction [33], PlantCARE [34] and PLACE [35]. The 3,227-bp fragment contained many cis-acting stress-responsive elements, such as ABRE, W-Box, MYB-core, W-Box core, TCA-element and others (Additional file 3: Table S2).
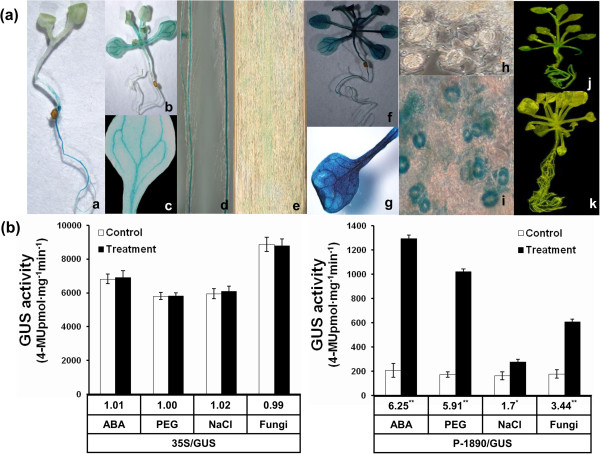
The 1,890-bp fragment (Additional file 2: Figure S1b) on the 5′ region upstream of the initiation codon of the GbRLK promoter transferred into Arabidopsis to address the regulatory mechanisms employed by the GbRLK promoter. The resulting transgenic Arabidopsis lines were designated P-1890/GUS. Histochemical staining revealed that GUS expression occurred mainly in the vascular bundles (Figure 3a-e) and was strictly confined to the leaf veins (Figure 3a-b), petioles (Figure 3a-c) and roots (Figure 3a-d,e), but not or very low in the cotyledons (Figure 3a-a) or root hairs (Figure 3a-d) of transgenic Arabidopsis (Figure 3a). Therefore, the GbRLK promoter displayed a tissue-specific expression pattern. This differed from the expression pattern in plants transformed with the cauliflower mosaic virus 35S (CaMV 35S) promoter controlling GUS, which served as a positive control. These plants exhibited constitutive GUS expression (Figure 3a-j,k).
Figure 3.
Histochemical straining and measurement of GUS activity in transgenic Arabidopsis thaliana. (a) a: 3-day-old seedling; b: 7-day-old seedling; c: mature leaf; d: root; e: microscopically observed root; f, g, i: transgenic plants treated with 100 μM ABA for 4 h; h: stomata before treatment with ABA; j: the CaMV 35S (pCAMBIA1301 vector) transformants as the positive control; k: non-transgenic Col-0 as the negative control. (b) GUS activity driven by the GbRLK promoter in above-ground tissues of transgenic Arabidopsis in response to ABA, PEG, NaCl and Verticillium dahliae. GUS activity from the CaMV 35S (pCAMBIA1301 vector) transformants served as the control. Data are mean and standard deviations of three replicates. The numbers below the bars indicate the -fold changes in GUS activity. Standard deviations (bars) are indicated. Significance of the changes produced after each treatment was assessed using Student’s t-tests (*P < 0.05, **P < 0.01).
The expression of GbRLK has been shown to respond to stress stimuli; therefore, our next step was to test whether the application of stress stimuli could trigger the expression of the GbRLK promoter. The induction of GUS activity was observed in ground tissues of P-1890/GUS transgenic plants upon treatment with ABA, PEG, NaCl or V. dahlia. As shown in Figure 2b, the application of ABA, PEG, NaCl or V. dahlia activated GUS expression in the tissues of P-1890/GUS plants, increasing the GUS activity level by 6.25, 5.91, 1.7 and 3.44-fold, respectively, which differed from that of plants containing GUS driven by the 35S (CaMV 35S) promoter (Figure 3b). Interestingly, GUS activity was strongly induced by ABA treatment (Figure 3a-f,g,h,j). Additionally, various stresses rapidly induced increasing transcript level of GUS gene (Additional file 2: Figure S1c).
Overexpression of GbRLK improved salt and drought tolerance in transgenic Arabidopsis
The 35S::GbRLK construct was introduced into Arabidopsis ecotype Col-0 and four independent fertile primary Arabidopsis transformants were regenerated, and positive transgenic plants were confirmed by PCR detection of NPTII and 35S-GbRLK (Additional file 4: Figure S2a). These T0 transgenic lines were self-pollinated to produce four homozygous lines, which were designated K-2, K-3, K-5 and K-6. Southern blotting analysis demonstrated that these lines arose from independent transformants, and each line carried between one and three copies of the GbRLK gene (Additional file 4: Figure S4b). The results of qRT-PCR revealed that GbRLK was expressed in all four transgenic Arabidopsis lines, with the highest level of expression in the K-2 line (Additional file 4: Figure S2c). The four transgenic lines showed no morphological aberrations through the T5 generation. No consistent difference in fresh weight was observed between the control and transgenic Arabidopsis lines (Additional file 4: Figure S2d).
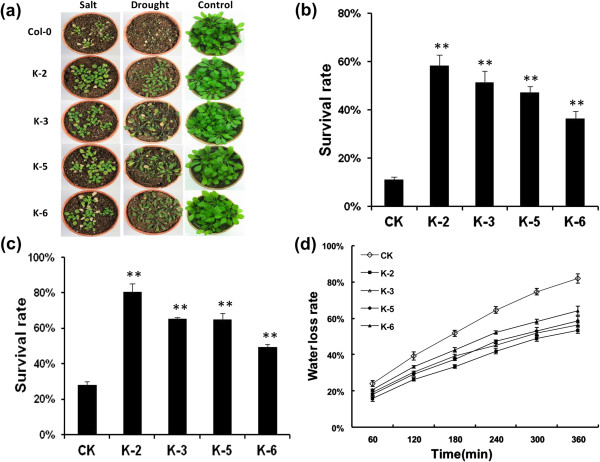
The analysis of the transgenic plants tolerance to drought revealed that the wild-type plants showed more severe symptoms than the transgenic plants (Figure 4a). Only 10% of the WT plants survived to maturity, whereas 58%, 51%, 47% and 36% of the transgenic plants survived in lines K-2, K-3, K-5 and K-6, respectively (Figure 4b). For analysis of high-salt tolerance, 3-week-old seedlings of transgenic and wild-type Arabidopsis grown in pots (with 4–5 leaves per plant ) were irrigated with a solution containing 200 mM NaCl once a week for 5 weeks. As shown in Figure 3a, both the transgenic and wild-type plants showed etiolation and yellowing, but the leaves of the WT plants wilted, and some of the WT plants died. The survival rate was approximately 27% in the WT compared to 80%, 65%, 64% and 49%, respectively in the four transgenic lines (Figure 4c). Additionally, there were also significant differences in fresh weight (FW), rosette diameter, anthocyanin accumulation, chlorophyll content and level of leaf chlorosis between the transgenic and the WT plants suffered drought and salt stresses (Tables 1 and 2). It is concluded that the transgenic Arabidopsis lines are more tolerant to drought and salt stresses compared to the WT, and among these transgenic lines, K-2 showed the highest tolerance, consistent with the expression level of the GbRLK. Overall, the WT plants showed more serious symptoms than the transgenic plants.
Figure 4.
Analysis of wild-type and 35S::GbRLK transgenic Arabidopsis lines subjected to salt and drought treatment. (a) The response of transgenic Arabidopsis to salt and drought stress. Wild-type (Col-0) and transgenic Arabidopsis were grown in pots containing soil mixture (rich soil: vermiculite = 2:1, v/v) and maintained in a growth chamber at 22°C on a 16-h light/8-hr dark regimen. For drought stress treatment, the photograph was taken after withholding water for 42 days. For salt stress treatment, the photograph was taken after treatment with a 150 mM NaCl solution for 35 days at 22°C. The experiments were repeated three times with similar results. (b) and (c) represent the survival ratios of transgenic Arabidopsis lines and Col-0 (CK) under drought-stress treatment for 42 days and salt-stress treatment for 35 days, respectively. (d) Water loss rate in transgenic and non-transgenic plants; standard deviations (bars) are indicated. Comparisons were made between wild-type and individual transgenic lines under control or drought-stress conditions using the paired t-test. ** indicates significant differences compared with the control at P < 0.01.
Table 1.
The symptoms of the transgenicArabidopsisand WT suffered drought stress
| FW(g) | Rosette diameter(cm) | Content of chlorophyll a (nmol/g) | Content of chlorophyll b (nmol/g) | Content of anthocyanin (nmol/g) | |
|---|---|---|---|---|---|
|
N |
0.237 ± 0.07 |
5.24 ± 0.60 |
0.157 ± 0.05 |
0.135 ± 0.03 |
27.522 ± 3.80 |
|
K- 2 |
1.407 ± 0.08** |
7.72 ± 1.34** |
0.755 ± 0.09** |
0.338 ± 0.05** |
16.006 ± 2.29** |
|
K- 6 |
0.664 ± 0.08** |
6.90 ± 1.10* |
0.492 ± 0.09** |
0.249 ± 0.06* |
18.141 ± 2.89** |
|
K- 3 |
0.735 ± 0.12** |
7.74 ± 0.64** |
0.654 ± 0.09** |
0.296 ± 0.02** |
17.295 ± 1.13** |
|
K- 5 |
0.821 ± 0.06** |
7.85 ± 0.94** |
0.684 ± 0.11** |
0.318 ± 0.07** |
17.326 ± 1.83** |
| P | 1.886 ± 0.10 | 9.73 ± 0.58 | 0.967 ± 0.06 | 0.406 ± 0.06 | 9.650 ± 1.84 |
The data were acquired from the 3 week old plants treated by withholding irrigation for 3 weeks. Data represent the mean ± SE (n ≥ 15); similar results were obtained from three independent experiments. N: wild-type Col-0 suffered drought stress, and P: non-treatment wild-type Col-0. Significance of the phenotypic results was assessed using Student’s t tests (*P < 0.05, **P < 0.01).
Table 2.
The symptoms of the transgenicArabidopsisand WT suffered salt stress
| FW(g) | Rosette diameter(cm) | Content of chlorophyll a (nmol/g) | Content of chlorophyll b (nmol/g) | Content of anthocyanin (nmol/g) | Rate of chlorosis (150 mM) | Rate of chlorosis (100 mM) | Germination rates (300 mM) | |
|---|---|---|---|---|---|---|---|---|
|
N |
0.190 ± 0.02 |
2.45 ± 0.39 |
0.211 ± 0.05 |
0.151 ± 0.02 |
21.220 ± 1.45 |
90.2% ± 2.9% |
59.2% ± 3.2% |
3.3% ± 0.9% |
|
K- 2 |
0.588 ± 0.09** |
5.40 ± 0.65** |
0.852 ± 0.06** |
0.365 ± 0.07** |
15.116 ± 3.14** |
58.1% ± 1.4%** |
20.0% ± 4.7%** |
33.3% ± 1.6%** |
|
K- 6 |
0.387 ± 0.03** |
4.48 ± 0.49** |
0.557 ± 0.72** |
0.269 ± 0.05** |
17.428 ± 2.38* |
76.3% ± 2.2%* |
48.3% ± 1.9%* |
23.3% ± 2.1%** |
|
K- 3 |
0.303 ± 0.05** |
4.29 ± 0.29** |
0.691 ± 0.05** |
0.346 ± 0.03** |
16.356 ± 1.30* |
63.3% ± 2.6%** |
32.5% ± 3.2%** |
30.0% ± 2.5%** |
|
K- 5 |
0.397 ± 0.05** |
4.90 ± 0.48** |
0.604 ± 0.12** |
0.325 ± 0.09** |
16.264 ± 1.02* |
66.7% ± 7.2%** |
47.5% ± 5.7%* |
26.7% ± 1.9%** |
| P | 1.886 ± 0.10 | 9.73 ± 0.58 | 0.967 ± 0.06 | 0.406 ± 0.06 | 9.650 ± 1.84 | -- | -- | 1 |
The data of FW, rosette diameter, content of chlorophyll and anthocyanin were acquired from the 3 week old plants treated by submerging the roots of the plants in 200 mM NaCl solution for 3 weeks. The data of rate of chlorosis and germination rates were acquired from plants grown on MS medium containing different concentration NaCl for 2 weeks. Data represent the mean ± SE (n ≥ 15); similar results were obtained from three independent experiments. N: wild-type Col-0 suffered drought stress, and P: non-treatment wild-type Col-0. Significance of the phenotypic results was assessed using Student’s t tests (*P < 0.05, **P < 0.01).
Furthermore, assays of water loss from detached leaves showed that the transgenic plants lost water much more slowly than the WT plants. At 360 min, the water loss rates were 53.5%, 56.4%, 58.6% and 64.2% in the transgenic lines K-2, K-3, K-5 and K-6, respectively, compared to 81.9% in the WT plants (Figure 4d). These results suggest that the overexpression of GbRLK reduces water loss and increases drought tolerance in transgenic plants. Therefore, GbRLK may function as a positive regulator of plant responses to drought stress.
Transgenic Arabidopsis plants were more sensitive to ABA than WT
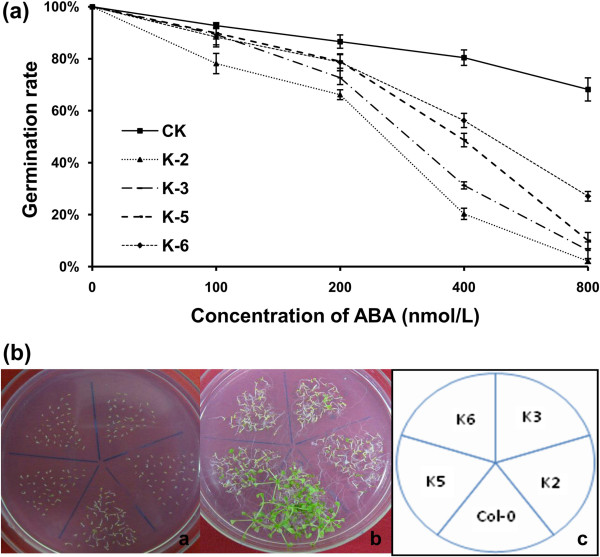
To examine changes in the response of GbRLK transgenic plants to ABA, we investigated the ABA sensitivity of the transgenic plant. Under ABA treatment, the germination rates of the transgenic lines were greatly reduced (Figure 5a). When grown on MS medium containing 100 nmol ABA for 1, 2 and 3 weeks, the transgenic plants exhibited many abnormal phenotypes such as dwarfing, etiolation and malformation (Figure 5b). Furthermore, root elongation in the 35S::GbRLK transgenic lines was hypersensitive to ABA (data not shown). These results demonstrate that ABA suppresses seed germination, root elongation and plant growth rates more strongly in GbRLK transgenic plants than in the WT. This indicates that ABA application activates the GbRLK protein or triggers other responses in transgenic plants. In addition, these results also suggest that the GbRLK protein activates or participates in ABA-dependent signal transduction cascades or plays an important regulatory role in ABA and stress responses.
Figure 5.
Analysis of 35S::GbRLK transgenic lines subjected to ABA treatment. (a) Germination ratios of transgenic lines and Col-0 on MS medium supplemented with 0, 25, 50, 100, 200, 400 and 800 nmol/L ABA. Standard deviations (bars) are indicated. (b) Analysis of the sensitivity of 35S::GbRLK transgenic lines to MS medium supplemented with 100 nmol/L ABA at different time points. The photographs were taken after 7 and 14 days of treatment in a and b , respectively.
GbRLK overexpression altered the expression pattern of stress-responsive genes in transgenic Arabidopsis
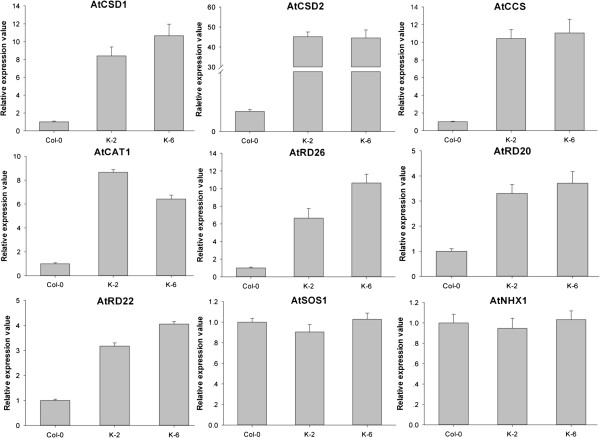
To reveal the possible molecular mechanisms underlying the improvement of stress resistance and tolerance in transgenic Arabidopsis thaliana by GbRLK, we studied the expression patterns of stress-responsive genes (AtRD20, AtRD22 and AtRD26), ion transporter genes (AtNHX1 and AtSOS1) and antioxidant genes (AtCAT1, AtCCS, AtCSD2 and AtCSD1) in transgenic and nontransgenic Arabidopsis thaliana. We examined the expression of these genes in three Arabidopsis lines, including two genetically pure clones, K-6 and K-2, which exhibited differences in stress resistance, and the nontransgenic line Col-0. The expression of all of the genes, except for ion transporter genes (AtNHX1, AtSOS1), was upregulated in transgenic GbRLK plants compared with Col-0 plants, especially antioxidant genes (AtCAT1, AtCCS, AtCSD2 and AtCSD1), which exhibited higher levels of upregulation than other genes (Figure 6).
Figure 6.
The expression of stress-responsive genes in transgenic GbRLK Arabidopsis vs. nontransgenic Col-0. Relative expression levels of stress-responsive genes were determined by qRT-PCR using cDNA synthesized from total RNAs isolated from the leaves of 2-week-old Arabidopsis grown in soil under normal conditions. Relative gene expression levels were determined using the 2^-ΔΔCT method. The CT (cycle threshold) values for both the target and internal control genes were means of three technical replicates. ΔCt = Ct target gene-CtRuBisCo. ΔΔCT = (CTtarget -CTRuBisCo)Transgenic - (CTtarget - CTRuBisCo) Non-transgenic. The large subunit of the RuBisCo gene (AtRuBisCo-F3/R3; Additional file 1: Table S1) from Arabidopsis was used as an internal control for normalization of different cDNA samples. The primers for the amplification of stress-responsive genes (Additional file 1: Table S1) were design based on sequences obtained from NCBI using Primer 5.0 software. Error bars represent standard error of means based on three independent reactions.
Discussion
Since RLKs were first reported, many kinases that play important roles in plant responses to abiotic stress and the ABA signaling network have been identified [27,28]. We have identified a RLK gene, GbRLK from cotton. The deduced protein sequence analysis of this gene revealed the presence of high structural homology with RLK orthologs from other plants. The results of phylogenetic analysis suggested that GbRLK protein was clustered together with other reported receptor-like kinase proteins and displayed the close phylogenetic relationship with Arabidopsis protein AT5G2408 and AT4G32300, which up-regulated with drought treatment [24-26]. This result indicated its potential function on abiotic stresses.
Interestingly, we found the increased expression of GbRLK transcript was observed under ABA, drought and high salt treatment, as shown by qRT-PCR, with highest early induction (4 h) found by ABA, suggesting the stress responsive transcript accumulation of GbRLK in cotton. Moreover GbRLK transcription was first transiently induced by ABA, while transcription induced by NaCl and drought treatment occurred later. In addition, ABA treatment induced higher levels of upregulated GUS expression than NaCl or drought, with NaCl producing the least upregulated GUS expression. We therefore conclude that GbRLK is a member of the ABA signal transduction pathway and activates ABA signal transduction. Drought or high-salt treatment increases ABA levels, which in turn results in the up-regulation of GbRLK. Meanwhile, overexpression of GbRLK can increase drought and high-salt tolerance in transgenic Arabidopsis. These results indicate that this RLK gene from cotton may likely function as a component of a signaling pathway that confers tolerance to abiotic stresses. GbRLK may play an important role in abiotic stress responses and the ABA signaling network.
ABA plays an important role in the tolerance of plants to abiotic stresses. ABA-independent and ABA-dependent signal transduction cascades are involved in the inducible expression of specific genes by abiotic stresses [36]. ABRE, MYB and MYC recognition sites function in ABA-dependent pathways 38]. The analysis of the promoters of such ABA-regulated genes has revealed a conserved cis-element, designated ABA-responsive element (ABRE; PyACGTGGC), which controls ABA-regulated gene expression [37]. In the ABA-independent pathway, gene expression in response to drought is regulated through DREs/CRT cis-acting elements or through NAC recognition sites [38]. Based on the current study, the GbRLK promoter contains two ABREs and some MYB and MYC-core recognition sites and does not contain DREs/CRT cis-acting elements or an NAC recognition site. Moreover, the GbRLK promoter and the GbRLK gene can respond to ABA, drought and high-salt stress. This study revealed a strong correlation between the inducibility of the GbRLK promoter and its internal cis-elements. For example, the W-box [39] and ABRE motifs [37,38], which are defense-responsive elements and cis-acting elements involved in abscisic acid responsiveness, may be the regulatory motifs of the GbRLK promoter that are responsible for the plant responses to the defense-related stimuli of V. dahliae and/or to ABA, PEG and NaCl. This information, combined with the results of the present study, suggests that GbRLK may take part in the ABA-dependent signaling pathway.
Water loss in plants is caused by drought and high-salinity conditions. ABA regulates stomatal opening in plants to avoid the affects of water loss due to drought and high salinity [40]. In this study, the water loss rate of the transgenic lines was significantly lower than that of the WT control. Thus, we can speculate that the improvement of resistance to drought and salt stresses may result from the ABA signaling pathway activated by the GRLK gene, which regulated other genes expression to decrease transpiration and water loss.
By contrast, the expression levels of two stress-responsive ion transporter genes, AtNHX1[41,42] and AtSOS1[42,43], were not significantly different in transgenic and nontransgenic Arabidopsis. In addition, GbRLK expression increased in response to Verticillium dahliae infection [32]. This pathogen promotes lignification of the cell wall, the formation of lignitubers and the restriction of vascular spread [44]; these responses not only prevent the pathogen from spreading further, but they also restrict water transport within the vascular system and alter the infected plant’s water balance or flux, causing severe dehydration stress. These results indicate that GbRLK is more highly induced by water deficit than by osmotic stress. Therefore, the main effect of the overexpression of GbRLK is a reduction in the amount of water loss in drought- and high-salt-stressed plants.
The molecular mechanisms underlying the response of higher plants to drought and high-salt stress are complicated. In response to these stress factors, various genes are up-regulated, which can mitigate the effects of stress and lead to an adjustment of plant stress tolerance. RD20 is one of the most highly stress-responsive genes and is often used as a stress marker gene [45-47]. RD26, which encodes an NAC protein, is induced by drought, high salinity and ABA and is probably involved in a novel ABA-dependent stress-signaling pathway [48]. RD22 is another stress-responsive gene [49]. In the current study, the increased expression of RD22, RD20 and RD26 in GbRLK plants compared with WT plants further supports the hypothesis that GbRLK plays a role in dehydration-stress tolerance.
The generation of reactive oxygen species (ROS) is common component of plant responses to abiotic stress and ABA [50]. Plants have evolved a complex antioxidant system to detoxify stress-induced ROS, in which ROS scavenging enzymes such as superoxide dismutase play essential roles [51]. The increased expression of AtCSD1[52], AtCSD2[52], AtCCS and AtCAT1[53] in GbRLK transgenic plants that was observed in the current study may have led to the scavenging of ROS produced under drought stress. These observations have clearly demonstrated the indirect role that GbRLK plays in protecting plants from dehydration stress by scavenging oxygen radicals. This study further clarifies the role played by GbRLK in improving stress tolerance in plants by affecting different stress-related pathways.
Conclusions
This study demonstrates that the manipulation of a GbRLK gene from cotton using a transgenic approach can lead to improved high-salt and drought-stress tolerance in plants. We also obtained similar results by analyzing the promoter of the GbRLK gene. Thus, we hypothesize that GbRLK activates or participate in the ABA signaling pathway, and overexpression of GbRLK may improve stress tolerance by regulating stress-responsive genes to reduce the damage caused by water loss. The results of this study help clarify the role of GbRLK in improving stress tolerance in plants by affecting different stress-related pathways. The GbRLK gene will be a candidate gene for future research on abiotic stress signaling pathways and the genetic engineering of novel cotton cultivars.
Methods
Plant and fungi material, growth conditions
Cotton seeds of G. barbadense L. cv. Hai7124, a Verticillum wilt-resistant cultivar widely used in China for genetic and breeding studies, were delinted with concentrated sulfuric acid. To ensure that the seeds were free of pathogens, the seeds were subjected to 37% formaldehyde fumigation for 24 h. The treated cotton seeds were planted in pots filled with sterile soil using one seed per pot in growth chambers at 27°C 16-h light and 22°C 8-h dark regimen. The relative humidity was approximate 70% (± 5%).
Arabidopsis seeds were surface-sterilized with a solution of NaClO (0.5%, w/v) and Triton X-100 (0.01%, v/v) for 10 min followed by four rinses with sterile water. The seeds were placed into Petri dishes containing MS agar (0.8%, w/v) medium and incubated for 2 days at 4°C, followed by transfer to 22°C for germination. After 7 days of growth on plates, the seedlings were transplanted to pots containing soil mixture (rich soil:vermiculite = 2:1, v/v) and maintained in growth chambers at 22°C under a 16-h light/ 8-h dark regimen. The relative humidity was approximate 70% (± 5%).
V. dahliae cultured on PDA medium (200 g potato, 17 g agar, 20 g sucrose, 1,000 ml H2O) was used to inoculate 100 ml Czapek’s liquid medium (2 g NaNO3, 1 g K2HPO3, 0.5 g MgSO4 · 7H2O, 0.5 g KCl, 0.01 g FeSO4 · 7H2O, 30 g sucrose, 1,000 ml distilled H2O) in a 250-ml flask, which was cultured at 25°C for 8–10 d with shaking. The V. dahliae culture was then passed through four layers of cloth, and the number of spores was counted under a microscope. The final spore concentration was adjusted to 1 × 108 immediately prior to plant inoculation.
Stress treatment
G. barbadense L. cv. Hai7124, seedlings containing two simple leaves and one heart-shaped leaf were subjected to various stress treatments. ABA treatment was carried out by spraying the plants with a 200 μM ABA solution at 0, 1, 2, 4, 6, 8, 10 and 12 h. Dehydration stress and salt-stress treatments were performed by submerging the roots of the plants in 20% PEG6000 or 200 mM NaCl solution under the same temperature and light conditions for 0, 1, 2, 4, 8, 12 and 24 h. Low-temperature stress was applied by transferring the plants to a growth chamber set at 4°C under the same photoperiodic conditions at 0, 1, 2, 4, 8, 12 and 24 h. SA and MeJA treatment were carried out by spraying the plants with 2 mM SA and 100 μM MeJA solution at 0, 2, 4, 6, 8, 10 and 12 h. After exposure to stress, the cotton leaves were immediately frozen in liquid nitrogen prior to expression analysis.
The seedlings were analyzed for ABA sensitivity as described by Fujita [48]. To test the phenotypes of plants grown in soil under salt and drought stress, Arabidopsis seedlings grown on MS medium for seven days were transferred to mixed soil (rich soil:vermiculite = 2:1, v/v) and grown for two week with sufficient watering. Each pot contained five seedlings. To investigate the tolerance to drought and high salt stresses, we supplied enough and equal volume water for every pot plants at the beginning of withholding irrigation and submerging the roots of the plants in 200 mM NaCl solution. The plants were then subjected to drought stress treatment by withholding irrigation for 42 days; the plants were grown at 22°C under a 16-h light/8 h dark regimen. For salt-stress analysis, wild-type and transgenic plants grown in pots at 22°C under a 16-h light/8 h dark regimen were irrigated with 200 mM NaCl solution once a week for 5 weeks. All pots were placed under the same conditions in a growth chamber in random order, and the position of each pot was randomly changed every day to exclude “positional effects”. The experiments were repeated for three times.
To measure GUS activity, Arabidopsis plants grown on MS medium for 14 days were transferred to one of the following solutions: 100 μM ABA, 20% PEG6000, 200 mM NaCl or 1 × 108 spore/ml V. dahliae spore suspension for 4 h, 12 h, 12 h or 96 h. After exposure to stress, the Arabidopsis seedlings were immediately frozen in liquid nitrogen prior to GUS activity analysis.
To measure germination rates and the rate of chlorosis of the transgenic lines, we planted Arabidopsis plants on MS medium containing different concentration NaCl for 14 days, then gathered the data.
Cloning the GbRLK promoter
Self-formed adaptor PCR (SEFA-PCR) was performed to clone the promoter sequence of GbRLK[54]. The detailed procedures were as follows: long and accurate Taq, buffer and deoxynucleoside triphosphates were purchased from TaKaRa Biotechnology Co., Ltd (Shiga, Japan). The PCR mixture included 15 μl of 2 × GC buffer I, 5 μl of 2.5 mM deoxynucleoside triphosphates, 1.5 U of long and accurate Taq enzyme and approximately 1 μg of H7124 template genomic DNA; deionized water was added to a final volume of 30 μl. PCR amplification was performed using three gene-specific internal primers, including SP1, SP2 and SP3 (Additional file 1: Table S1).
PCR amplification
Standard polymerase chain reaction (PCR) was completed using rTaq DNA polymerase (TaKaRa Biotechnology (Dalian) Co., Ltd., China). The reaction system of PCR products contain 2 μl 10 × buffer, 1 μl Mg2+, 0.2 μl rTaq DNA polymerase, 1 μl template and 0.5 μl dNTP in 20 μl volume to PCR thermal cycling conditions were: 4 min at 94°C; 30 cycles of 30s at 94°C, 30 s at 55-62°C, 0.5-1.5 min at 72°C; and finally 7 min at 72°C [55]. The annealing temperature and extending time were changed based on the different primers. The primers used in this paper were listed in Additional file 1: Table S1.
Plasmid construction and Arabidopsis transformation
To construct the 35S::GbRLK vector, the entire coding region of GbRLK was amplified by PCR using Sma I and Xba I linker primers (GbRLK1-F/R) (Additional file 1: Table S1) and ligated into vector pBI121. To construct the pGbRLK::GUS vector, a 1,890-bp fragment of the GbRLK promoter was chosen. This fragment was amplified by PCR using EcoR I and Bgl II linker primers (pGbRLK1-F/R; Additional file 1: Table S1) and ligated into vector pCAMBIA1301. Specifically, the PCR-amplified fragment was digested with EcoR I and Bgl II (Promega, Madison, WI, USA) and purified with a TIAN-quick Midi Purification Kit (TIANGEN, Beijing, People’s Republic of China). The fragment was then fused to the GUS reporter gene of the pCAMBIA1301 vector, which was previously digested with EcoR I and Bgl II to release the 35S promoter. The resulting vector, named P-1890/GUS, was confirmed by sequencing; the pCAMBIA1301 vector served as a control and was named 35S/GUS.
The constructs were introduced into Agrobacterium tumefaciens (strain GV3101) and transferred into Arabidopsis (Col-0) using the floral dip method [56]. Transgenic GbRLK plants were confirmed by examining the segregation ratio of the kanamycin selectable marker and by PCR analysis of NPTII and 35S-GbRLK using the primers NPTII-F/R and 35S-GbRLK (Additional file 1: Table S1). The primers GbRLK2-F/R (Additional file 1: Table S1) were designed to evaluate the expression level of GbRLK in Arabidopsis by qRT-PCR. Southern blotting analysis of transgenic Arabidopsis plants was conducted using the 3′ 407-bp sequence of the full-length cDNA, which was amplified using GbRLK3-F/R primers (Additional file 1: Table S1), as a probe. Hybridization was conducted according to the instructions of the DIG High Prime DNA labeling and Detection Kit (Roche, Switzerland). Transgenic Arabidopsis plants with P-1890/GUS were confirmed using primers designed from the promoter sequence (pGbRLK1-F/R) and GUS (GUS-F/R; Additional file 1: Table S1). The transgenic lines used for analysis in this study were T5 homozygous plants.
Quantitative real time reverse transcription polymerase chain reaction (qRT-PCR)
Total RNA was isolated using methods developed from the cetyltrimethyammonium brodmide (CTAB) method [57]. First-strand cDNA was synthesized from 1 μg total RNA using the M-MLV reverse transcriptase (Promega, USA) according to the manufacturer’s instructions. Quantitative PCR (qRT-PCR) was used in order to evaluate the expression levels of GbRLK gene under different stress treatments using the leaves of G. barbasense L. cv. Hai7124 and the ABI 7500 Real Time System (PE Applied Biosystems, USA). Gene-specific primers were designed according to GbRLK sequences using Primer Premier 5.0 (Additional file 1: Table S1) and synthesized by GenScript Corp (Nanjing, China). The transcript encoding cotton elongation factor (EF1α) was amplified as a standard control for qRT-PCR analysis of cotton, and the large subunit of the RuBisCo gene (AtRuBisCo-F3/R3; Additional file 1: Table S1) [58] from Arabidopsis was used as an internal control for normalization of different Aarabidopsis cDNA samples Melt curve analysis was performed to check the specificity of amplified product and relative gene expression levels were determined using the 2-ΔΔCT method [59]. The CT (cycle threshold) values for both the target and internal control genes were means of three technical replicates. ΔCt = Ct target gene-Ct EF1α. ΔΔCT = (CTtarget - CT control)treatment/transgenic - (CTtarget - CT control)control.
Measurement of transpiration rates
For water loss measurement, rosette leaves were detached from 4-week-old plants and weighted immediately on a piece of weighting paper, which was then placed on a laboratory bench (at a relative humidity of 40% to 50% and a temperature of 22°C to 23°C). The weight loss of each sample was measured at designated time points. The proportion of water loss was calculated on the basis of the initial fresh weight of the sample; water loss was expressed as a percentage of the initial fresh weight. All tests were repeated at least three times.
Histochemical and fluorometric GUS assay
For histochemical staining of GUS, fresh tissue samples were obtained from Arabidopsis plants and immediately exposed to X-Gluc solution. After overnight incubation at 37°C, stained samples were bleached with 70% (v/v) ethanol and observed under OLYMPUS BX51 and SZX12 microscopes. A fluorometric GUS assay was performed as described by Jefferson et al.[60].
The fluorescence of 4-methylumbelliferone (4-MU), the product of GUS-catalyzed hydrolysis, was measured using the TECAN CENios system. The protein concentration in the supernatant was assessed by the Bradford [61] method, using bovine serum albumin (BSA) as a standard. GUS activity was normalized to the protein concentration of each supernatant extract and calculated as pmol of 4-MU per milligram of soluble protein per minute.
Measurement of anthocyanin and chlorophyll content
Anthocyanin was extracted and quantified as described by Gareth et al.[62] from shoot tissues of 10–15 plantlets 3 weeks after treatment with salt and drought. Results are expressed as absorbance at 530 nm with a spectrophotometer (Shimadzu UV-1600, Japan). Chlorophyll was extracted and measured in triplicate as described by Lichtenthaler [63]. About 100 mg of fine powder of leaf tissue was homogenized in 1 ml of 80% acetone and kept for 15 min at room temperature in dark. The crude extract was centrifuged for 20 min at 10000 rpm (rotation per minute) at room temperature, and the resultant supernatant was used for assessing absorbance at 663 and 645 nm with a spectrophotometer (Shimadzu UV-1600, Japan). Chlorophyll and anthocyanin content were computed in terms of fresh weight (FW).
Abbreviations
RLK: Receptor-like kinase; ABA: Abscisic acid; GUS: β-Glucuronidase gene; MS: Murashige and skoog medium; SEFA-PCR: Self-formed adaptor PCR; SA: Salicylic acid; JA: Jasmonic acid.
Competing interests
The authors declare that they have no competing interests.
Authors’ contributions
ZTZ designed the study and wrote the manuscript; ZJ performed the experiments and analyzed the data; GYL cloned the gene; ZZY measured the GUS activity; CTZ transformed the Arabidopsis; GWZ contributed reagents/materials/analysis tools. All authors read and approved the final manuscript.
Supplementary Material
The sequences of primers employed in this study.
Cloning and analysis of GbRLK promoter. (a) Agarose gel electrophoresis of GbRLK promoter region by the Sefa -PCR method. Lane M: DNA Marker; Lane 1: the product of the first round of nested PCR; Lane 2: the product of the second round of nested PCR. (b) The sequence of the GbRLK promoter and cis-acting element analysis. The red background represents the initiation codon of GbRLK. The A nucleotide of the initiation codon ATG is numbered +1. The core promoters (TATA-box and CAAT-box) are indicated with a blue background, the putative transcriptional initiation site (TSS) A is shown in red and larger font and the cis-acting elements involved in light, regulation of plant growth and hormones are indicated by a boxed, yellow background. (c) GUS expression in P-1890/GUS transgenic Arabidopsis plants after treatment with different stressors.
Cis-acting element analysis of 5′ flanking sequences of GbRLK.
Molecular analysis of independent transgenic Arabidopsis lines. (a) PCR tests of transgenic plants. (b) Southern blot analysis of four independent transgenic lines. P: positive control pBI121, N: non-transformed plant; Transgenic lines K-2, K-3, K-5 and K-6. (c) qRT–PCR analysis of GbRLK expression in homozygous transgenic Arabidopsis lines. The large subunit of the RuBisCo gene (AtRuBisCo-F3/R3; Additional file 1: Table S1) from Arabidopsis was used as an internal control for normalization of different cDNA samples. (d) Fresh weight of transgenic and nontransgenic Arabidopsis at different time points.
Contributor Information
Jun Zhao, Email: sxzhaojun88@yahoo.com.cn.
Yulong Gao, Email: gyl3000@163.com.
Zhiyuan Zhang, Email: 2009101154@njau.edu.cn.
Tianzi Chen, Email: actzi0503@gmail.com.
Wangzhen Guo, Email: moelab@njau.edu.cn.
Tianzhen Zhang, Email: cotton@njau.edu.cn.
Acknowledgments
This program was financially supported in part by 973 Program (2011CB109300), NSFC (31171590), MOE (20090097110010), NSF in Jiangsu (BK2010065) and the Priority Academic Program Development of Jiangsu Higher Education Institutions.
References
- Mahajan S, Tuteja N. Cold, salinity and drought stresses: an overview. Arch Biochem Biophys. 2005;444:139–158. doi: 10.1016/j.abb.2005.10.018. [DOI] [PubMed] [Google Scholar]
- Zhang JH, Jia WS, Yang JC, Ismail AM. Role of ABA in integrating plant responses to drought and salt stresses. Field Crops Research. 2006;97:111–119. doi: 10.1016/j.fcr.2005.08.018. [DOI] [Google Scholar]
- Cutler SR, Rodriguez PL, Finkelstein RR, Abrams SR. Abscisic acid: emergence of a core signal network. Ann Rev Plant Biol. 2010;61:651–679. doi: 10.1146/annurev-arplant-042809-112122. [DOI] [PubMed] [Google Scholar]
- Cramer GR, Urano K, Delrot S, Pezzotti M, Shinozaki K. Effects of abiotic stress on plants: a systems biology perspective. BMC Plant Bio. 2011;11:163. doi: 10.1186/1471-2229-11-163. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liotenberg S, North H, Marion-Poll A. Molecular biology and regulation of abscisic acid biosynthesis in plants. Plant Physiol Biochem. 1999;37:341–350. doi: 10.1016/S0981-9428(99)80040-0. [DOI] [Google Scholar]
- Xiong L, Ishitani M, Lee H, Zhu JK. The Arabidopsis LOS5/ABA3 locus encodes a molybdenum cofactor sulfurase and modulates cold stress- and osmotic stress-responsive gene expression. Plant Cell. 2001;13:2063–2083. doi: 10.1105/TPC.010101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fujii H, Zhu JK. Arabidopsis mutant deficient in 3 abscisic acid-activated protein kinases reveals critical roles in growth, reproduction, and stress. Proc Natl Acad Sci USA. 2009;106:8380–8385. doi: 10.1073/pnas.0903144106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Payton P, Kottapalli KR, Kebede H, Mahan JR, Wright RJ, Allen RD. Examining the drought stress transcriptome in cotton leaf and root tissue. Biotechnol Lett. 2011;33:821–828. doi: 10.1007/s10529-010-0499-y. [DOI] [PubMed] [Google Scholar]
- Padmalatha KV, Dhandapani G, Kanakachari M, Kumar S, Dass A, Patil DP, Rajamani V, Kumar K, Pathak R, Rawat B, Leelavathi S, Reddy PS, Jain N, Powar KN, Hiremath V, Katageri IS, Reddy MK, Solanke AU, Reddy VS, Kumar PA. Genome-wide transcriptomic analysis of cotton under drought stress reveal significant down-regulation of genes and pathways involved in fibre elongation and up-regulation of defense responsive genes. Plant Mol Biol. 2012;78:223–246. doi: 10.1007/s11103-011-9857-y. [DOI] [PubMed] [Google Scholar]
- Ranjan A, Nigam D, Asif MH, Singh R, Ranjan S, Mantri S, Pandey N, Trivedi I, Rai KM, Jena SN, Koul B, Tuli R, Pathre UV, Sawant SV. Genome wide expression profiling of two accession of G. herbaceum L. in response to drought. BMC Genomics. 2012;13:94. doi: 10.1186/1471-2164-13-94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shiu SH, Bleecker AB. Plant receptor-like kinase gene family: diversity, function, and signal. Sci.STKE. 2001;113:RE22. doi: 10.1126/stke.2001.113.re22. [DOI] [PubMed] [Google Scholar]
- Shiu SH, Bleecker AB. Receptor-like kinases from Arabidopsis form a monophyletic gene family related to animal receptor kinases. Proc Natl Acad Sci USA. 2001;98:10763–10768. doi: 10.1073/pnas.181141598. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shiu SH, Karlowski WM, Pan R, Tzeng YH, Mayer KF, Li WH. Comparative analysis of the receptor-like kinase family in Arabidopsis and rice. Plant Cell. 2004;16:1220–1234. doi: 10.1105/tpc.020834. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang MP, Wu YH, Lee MK, Liu YH, Rong Y, Santos TS, Wu CC, Xie FM, Nelson RL, Zhang HB. Numbers of genes in the NBS and RLK families vary by more than four-fold within a plant species and are regulated by multiple factors. Nucleic Acids Res. 2010;38:6513–6525. doi: 10.1093/nar/gkq524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walker JC. Structure and function of the receptor-like protein kinases of higher plants. Plant Mol Biol. 1994;26:1599–1609. doi: 10.1007/BF00016492. [DOI] [PubMed] [Google Scholar]
- Torii KU. Receptor kinase activation and signal transduction in plants: an emerging picture. Curr Opin Plant Biol. 2000;3:361–367. doi: 10.1016/S1369-5266(00)00097-2. [DOI] [PubMed] [Google Scholar]
- Tang W. BSKs mediate signal transduction from the receptor kinase BRI1 in Arabidopsis. Science. 2008;321:557–560. doi: 10.1126/science.1156973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim TW. Brassinosteroid signal transduction from cell-surface receptor kinases to nuclear transcription factors. Nature Cell Biol. 2009;11:1254–1260. doi: 10.1038/ncb1970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clark SE. Cell signal at the shoot meristem. Nature Rev Mol Cell Biol. 2001;2:276–284. doi: 10.1038/35067079. [DOI] [PubMed] [Google Scholar]
- Jinn TL, Stone JM, Walker JC. HAESA, an Arabidopsis leucine-rich repeat receptor kinase, controls floral organ abscission. Gene Dev. 2000;14:108–117. [PMC free article] [PubMed] [Google Scholar]
- Martin GB, Brommonschenkel S, Chunwongse J, Frary A, Ganal MW, Spivey R, Wu T, Earle ED, Tanksley SD. Map-based cloning of a protein kinase gene conferring disease resistance in tomato. Science. 1993;262:1432–1436. doi: 10.1126/science.7902614. [DOI] [PubMed] [Google Scholar]
- Song WY, Wang GL, Chen LL, Kim HS, Pi LY, Holsten T, Gadner J, Wang B, Zhai WX, Zhu LH, Fauquet C, Ronald P. A receptor kinase-like protein encoded by the rice disease resistance gene, Xa21. Science. 1995;270:1804–1806. doi: 10.1126/science.270.5243.1804. [DOI] [PubMed] [Google Scholar]
- Gomez-Gomez L, Boller T. FLS2: an LRR receptor-like kinase involved in the perception of bacterial elicitor flagellin in Arabidopsis. Mol Cell. 2000;5:1003–1011. doi: 10.1016/S1097-2765(00)80265-8. [DOI] [PubMed] [Google Scholar]
- Kilian J, Whitehead D, Horak J, Wanke D, Weinl S, Batistic O, D’Angelo C, Bornberg-Bauer E, Kudla J, Harter K. The AtGenExpress global stress expression data set: protocols, evaluation and model data analysis of UV-B light, drought and cold stress responses. Plant J. 2007;50:347–363. doi: 10.1111/j.1365-313X.2007.03052.x. [DOI] [PubMed] [Google Scholar]
- Skirycz A, Claeys H, De Bodt S, Oikawa A, Shinoda S, Andriankaja M, Maleux K, Eloy NB, Coppens F, Yoo SD, Saito K, Inzé D. Pause-and-stop: the effects of osmotic stress on cell proliferation during early leaf development in Arabidopsis and a role for ethylene signaling in cell cycle arrest. Plant Cell. 2011;23:1876–1888. doi: 10.1105/tpc.111.084160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marshall A, Aalen RB, Audenaert D, Beeckman T, Broadley MR, Butenko MA, Caño-Delgado A, De Vries S, Dresselhaus T, Felix G, Graham NS, Foulkes J, Granier C, Greb T, Grossniklaus U, Hammond JP, Heidstra R, Hodgman C, Hothorn M, Inzé D, Østergaard L, Russinova E, Simon R, Skirycz A, Stahl Y, Zipfel C, De Smete I. Tackling drought stress: recepror-like kinase present new approaches. Plant Cell. 2012;24:2262–2278. doi: 10.1105/tpc.112.096677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tanaka H, Osakabe Y, Katsura S, Mizuno S, Maruyama K, Kusakabe K, Mizoi J, Shinozaki K, Yamaguchi-Shinozaki K. Abiotic stress-inducible receptor-like kinases negatively control ABA signal in Arabidopsis. Plant J. 2012;70:599–613. doi: 10.1111/j.1365-313X.2012.04901.x. [DOI] [PubMed] [Google Scholar]
- Hua DP, Wang C, He J, Liao H, Duan Y, Zhu ZQ, Guo Y, Chen ZZ, Gong ZZ. A plasma membrane receptor kinase, GHR1, mediates abscisic acid- and hydrogen peroxide-regulated stomatal movement in Arabidopsis. Plant Cell. 2012;24:2546–2561. doi: 10.1105/tpc.112.100107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Osakabe Y, Mizuno S, Tanaka H, Maruyama K, Osakabe K, Todaka D, Fujita Y, Kobayashi M, Shinozaki K, Yamaguchi-Shinozaki K. Overproduction of the membrane-bound receptor-like protein kinase 1, RPK1, enhances abiotic stress tolerance in Arabidopsis. J Biol Chem. 2010;285:9190–9201. doi: 10.1074/jbc.M109.051938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakashima K, Fujita Y, Kanamori N, Katagiri T, Umezawa T. Three Arabidopsis SnRK2 protein kinases, SRK2D/SnRK2.2, SRK2E/SnRK2.6/OST1 and SRK2I/SnRK2.3, involved in ABA signal are essential for the control of seed development and dormancy. Plant Cell Physiol. 2009;50:1345–1363. doi: 10.1093/pcp/pcp083. [DOI] [PubMed] [Google Scholar]
- Ouyang SQ, Liu YF, Liu P, Lei G, He SJ, Ma B, Zhang WK, Zhang JS, Chen SY. Receptor-like kinase OsSIK1 improves drought and salt stress tolerance in rice (Oryza sativa) plants. Plant J. 2010;62:316–329. doi: 10.1111/j.1365-313X.2010.04146.x. [DOI] [PubMed] [Google Scholar]
- Gao YL, Guo WZ, Wang L, Zhang TZ. Isolation and characterization of resistance and defense gene analogs in cotton (Gossypium barbadense L.) Sci China Life Sci. 2006;36:97–108. doi: 10.1007/s11427-006-2017-y. [DOI] [PubMed] [Google Scholar]
- Reese MG. Application of a time-delay neural network to promoter annotation in the drosophila melanogaster genome. Comput Chem. 2001;26:51–56. doi: 10.1016/S0097-8485(01)00099-7. [DOI] [PubMed] [Google Scholar]
- Lescot M, De Hais P, Thijs G, Marchal K, Moreau Y, Peer YV, Rouzé P, Rombauts S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002;30:325–327. doi: 10.1093/nar/30.1.325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Higo K, Ugawa Y, Iwamoto M, Korenaga T. Plant cis-acting regulatory DNA elements (PLACE) database. Nucleic Acids Res. 1999;27:297–300. doi: 10.1093/nar/27.1.297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shinozak IK, Yamaguchi-Shinozaki K. Molecular responses to dehydration and low temperature: differences and cross-talk between two stress signal pathways. Curr Opin Plant Biol. 2000;3:217–223. [PubMed] [Google Scholar]
- Busk PK, Pages M. Regulation of abscisic acid-induced transcription. Plant Mol Biol. 1998;37:425–435. doi: 10.1023/A:1006058700720. [DOI] [PubMed] [Google Scholar]
- Yamaguchi-Shinozaki K, Shinozaki K. Organization of cis-acting regulatory elements in osmotic and cold-stress-responsive promoters. Trends Plant Sci. 2005;10:88–94. doi: 10.1016/j.tplants.2004.12.012. [DOI] [PubMed] [Google Scholar]
- Rushton PJ, Reinstädler A, Lipka V, Lippok B, Somssich IE. Synthetic plant promoters containing defined regulatory elements provide novel insights into pathogen- and wound-induced signaling. Plant Cell. 2002;14:749–762. doi: 10.1105/tpc.010412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chinnusamy V, Gong Z, Zhu JK. Abscisic acid-mediated epigenetic processes in plant development and stress responses. J Integr Plant Biol. 2008;50:1187–1195. doi: 10.1111/j.1744-7909.2008.00727.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Apse MP, Sottosanto JB, Blumwald E. Vacuolar cation/H exchange, ion honeostasis, and leaf development are altered in a T-DNA insertional mutant of AtNHX1, the Arabidopsis vacuolar Na+/H + antiporter. Plant J. 2003;36:229–239. doi: 10.1046/j.1365-313X.2003.01871.x. [DOI] [PubMed] [Google Scholar]
- Karan R, Subudhi PK. A stress inducible SUMO conjugating enzyme gene (SaSce9) from a grass halophyte spartina alterniflora enhances salinity and drought stress tolerance in Arabidopsis. BMC Plant Biology. 2012;12:187. doi: 10.1186/1471-2229-12-187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi H, Quintero FJ, Pardo JM, Zhu JK. The putative plasma membrane Na+/H + antiporter SOS1 controls long-distance Na + transport in plants. Plant Cell. 2002;14:465–477. doi: 10.1105/tpc.010371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beckman CH, Talboys PW. In: Fungal wilt diseases in plants. Mace MC, Bell AA, Beckman CH, editor. New York: Academic Press; 1981. Anatomy of resistance; pp. 487–521. [Google Scholar]
- Magnan F, Ranty B, Charpenteau M, Sotta B, Galaud JP, Aldon D. Mutations in AtCML9, a calmodulin-like protein from Arabidopsis thaliana, alter plant responses to abiotic stress and abscisic acid. Plant J. 2008;56:575–589. doi: 10.1111/j.1365-313X.2008.03622.x. [DOI] [PubMed] [Google Scholar]
- Alexandre C, Möller-Steinbach Y, Schönrock N, Gruissem W, Hennig L. Arabidopsis MSI1 is required for negative regulation of the response to drought stress. Mol Plant. 2009;2:675–687. doi: 10.1093/mp/ssp012. [DOI] [PubMed] [Google Scholar]
- Aubert Y, Vile D, Pervent M, Aldon D, Ranty B, Simonneau T, Vavasseur A, Jean-Philippe G. RD20, A stress-inducible caleosin, participates in stomatal control, transpiration and drought tolerance in Arabidopsis thaliana. Plant Cell Physiol. 2010;51(12):1975–1987. doi: 10.1093/pcp/pcq155. [DOI] [PubMed] [Google Scholar]
- Fujita M, Fujita Y, Maruyama K, Seki M, Hiratsu K, Ohme-Takagi M, Tran LP, Yamaguchi-Shinozaki K, Shinozaki K. A dehydration-induced NAC protein, RD26, is involved in a novel ABA-dependent stress-signal pathway. Plant J. 2004;39:863–876. doi: 10.1111/j.1365-313X.2004.02171.x. [DOI] [PubMed] [Google Scholar]
- Yamaguchi-Shinozaki K, Shinozaki K. The plant hormone abscisic acid mediates the drought-induced expression but not the seed-specific expression of rd22, a gene responsive to dehydration stress in Arabidopsis thaliana. Mol Gen Genet. 1993;238:17–25. doi: 10.1007/BF00279525. [DOI] [PubMed] [Google Scholar]
- Huang GT, Ma SL, Bai LP, Zhang L, Ma H, Jia P, Liu J, Zhong M, Guo ZF. Signal transduction during cold, salt, and drought stresses in plants. Mol Biol Rep. 2012;39:969–987. doi: 10.1007/s11033-011-0823-1. [DOI] [PubMed] [Google Scholar]
- Rup Kumar K. Plant responses to water stress role of reactive oxygen species. Plant Signal & Behavior. 2011;6:1741–1745. doi: 10.4161/psb.6.11.17729. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sunkar R, Kapoor A, Zhu JK. Posttranscriptional induction of two Cu/Zn superoxide dismutase genes in Arabidopsis is mediated by down regulation of miR398 and important for oxidative stress tolerance. Plant Cell. 2006;18:2051–2065. doi: 10.1105/tpc.106.041673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Havir EA, McHale NA. Enhanced peroxidatic activity in specific catalase isozymes of tobacco, barley, and maize. Plant Physiol. 1989;91:812–815. doi: 10.1104/pp.91.3.812. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang S, He J, Cui Z, Li S. Self-formed adaptor PCR: a simple and efficient method for chromosome walking. Appl Environ Microbiol. 2007;73:5048–5051. doi: 10.1128/AEM.02973-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Erlich HA, Gelfand DH, Saiki RK. Specific DNA amplification. Nature. 1988;331:461–462. doi: 10.1038/331461a0. [DOI] [Google Scholar]
- Clough SJ, Bent AF. Floral dip: a simplified method for Agrobacterium mediated transformation of Arabidopsis thaliana. Plant J. 1998;16:735–743. doi: 10.1046/j.1365-313x.1998.00343.x. [DOI] [PubMed] [Google Scholar]
- Paterson AH, Lander ES, Hewitt JD. A rapid method for extraction of cotton (Gossypium spp.) genomic DNA suitable for RFLP or PCR analysis. Plant Mol Biol Rep. 1993;11:122–127. doi: 10.1007/BF02670470. [DOI] [Google Scholar]
- Ellendorff U, Fradin EF, Ronnie de J, Thomma J. RNA silencing is required for Arabidopsis defense against Verticillium wilt disease. J Exp Bo. 2009;60:591–602. doi: 10.1093/jxb/ern306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods. 2001;25:402–408. doi: 10.1006/meth.2001.1262. [DOI] [PubMed] [Google Scholar]
- Jefferson RA, Kavanagh TA, Bevan MW. GUS fusions: beta-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J. 1987;6(13):3901–3907. doi: 10.1002/j.1460-2075.1987.tb02730.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bradford MM. A rapid sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- Gareth JD, Sheikh MA, Ratcliffe OJ, Coupland G, Furner LJ. Genetics of homology-dependent gene silencing in Arabidopsis: a role for methylation. Plant J. 1997;12:791–804. doi: 10.1046/j.1365-313X.1997.12040791.x. [DOI] [PubMed] [Google Scholar]
- Lichtenthaler HK. Chlorophylls and carotenoids: pigments of photosynthetic biomembranes. Meth Enzymol. 1987;18:350–382. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
The sequences of primers employed in this study.
Cloning and analysis of GbRLK promoter. (a) Agarose gel electrophoresis of GbRLK promoter region by the Sefa -PCR method. Lane M: DNA Marker; Lane 1: the product of the first round of nested PCR; Lane 2: the product of the second round of nested PCR. (b) The sequence of the GbRLK promoter and cis-acting element analysis. The red background represents the initiation codon of GbRLK. The A nucleotide of the initiation codon ATG is numbered +1. The core promoters (TATA-box and CAAT-box) are indicated with a blue background, the putative transcriptional initiation site (TSS) A is shown in red and larger font and the cis-acting elements involved in light, regulation of plant growth and hormones are indicated by a boxed, yellow background. (c) GUS expression in P-1890/GUS transgenic Arabidopsis plants after treatment with different stressors.
Cis-acting element analysis of 5′ flanking sequences of GbRLK.
Molecular analysis of independent transgenic Arabidopsis lines. (a) PCR tests of transgenic plants. (b) Southern blot analysis of four independent transgenic lines. P: positive control pBI121, N: non-transformed plant; Transgenic lines K-2, K-3, K-5 and K-6. (c) qRT–PCR analysis of GbRLK expression in homozygous transgenic Arabidopsis lines. The large subunit of the RuBisCo gene (AtRuBisCo-F3/R3; Additional file 1: Table S1) from Arabidopsis was used as an internal control for normalization of different cDNA samples. (d) Fresh weight of transgenic and nontransgenic Arabidopsis at different time points.