Abstract
Elucidation of tumor-DNA virus associations in many cancer types has enhanced our knowledge of fundamental oncogenesis mechanisms and provided a basis for cancer prevention initiatives. RNA-Seq is a novel tool to comprehensively assess such associations. We interrogated RNA-Seq data from 3,775 malignant neoplasms in The Cancer Genome Atlas database for the presence of viral sequences. Viral integration sites were also detected in expressed transcripts using a novel approach. The detection capacity of RNA-Seq was compared to available clinical laboratory data. Human papillomavirus (HPV) transcripts were detected using RNA-Seq analysis in head-and-neck squamous cell carcinoma, uterine endometrioid carcinoma, and squamous cell carcinoma of the lung. Detection of HPV by RNA-Seq correlated with detection by in situ hybridization and immunohistochemistry in squamous cell carcinoma tumors of the head and neck. Hepatitis B virus and Epstein-Barr virus (EBV) were detected using RNA-Seq in hepatocellular carcinoma and gastric carcinoma tumors, respectively. Integration sites of viral genes and oncogenes were detected in cancers harboring HPV or hepatitis B virus but not in EBV-positive gastric carcinoma. Integration sites of expressed viral transcripts frequently involved known coding areas of the host genome. No DNA virus transcripts were detected in acute myeloid leukemia, cutaneous melanoma, low- and high-grade gliomas of the brain, and adenocarcinomas of the breast, colon and rectum, lung, prostate, ovary, kidney, and thyroid. In conclusion, this study provides a large-scale overview of the landscape of DNA viruses in human malignant cancers. While further validation is necessary for specific cancer types, our findings highlight the utility of RNA-Seq in detecting tumor-associated DNA viruses and identifying viral integration sites that may unravel novel mechanisms of cancer pathogenesis.
INTRODUCTION
The association between infection with DNA viruses and neoplasia is well established in a variety of cancer types (1). The oncogenic potential of DNA viruses is variable, and their role in cancer pathogenesis is mediated through various mechanisms, including, for example, mutagenic integration into the host genome and expression of oncogenic viral proteins (2, 3). The elucidation of such mechanisms has played a key role in enhancing our understanding of cancer pathogenesis even as novel aspects of DNA virus biology continue to be unraveled (4). Furthermore, since eradication of viral infections through prevention and vaccination initiatives could have a drastic epidemiologic impact on the incidence of virus-associated cancers, there has been a vigorous search for tumor-associated viruses in neoplasms where such an association has remained elusive.
One of the best understood causal relationships is between human papillomavirus (HPV) infection and squamous neoplasia of the anogenital and head-and-neck regions. HPV is a small, 50- to 55-nm-diameter, nonenveloped, double-stranded DNA virus that carries out its life cycle in either mucosal or cutaneous stratified squamous epithelia (5). The viral genome (8 kb in size) is amplified initially as extrachromosomal circular elements (episomes) but may eventually integrate into the host genome. Over 120 types of HPV have been identified, of which those capable of infecting humans are designated high risk or low risk on the basis of their association with human neoplasms and oncogenic potential. The oncoproteins E5, E6, and E7 are the primary agents responsible for initiation and progression of HPV-associated cancers, and they operate primarily by abrogating negative growth regulators and inducing genomic instability. The integration of HPV DNA into the host cell genome is considered an important step in malignant progression and is commonly identified in noninvasive and invasive carcinomas associated with high-risk types HPV16 and HPV18 (6–8). HPV integration sites, with a predilection for sites of known genomic fragility, have been found to be distributed randomly over the whole genome in one study (9), and the majority of integrated HPV genomes appear to be actively transcribed (10, 11).
Hepatocellular carcinoma (HCC) is one of the leading causes of cancer-related mortality in the world and is strongly associated with chronic hepatitis B virus (HBV) infection. The HBV virion consists of partially double-stranded DNA packaged with a core protein (HBcAg) and DNA polymerase within envelope proteins (HBsAg) (12). Integration of viral DNA into the genome of HCC cells has been demonstrated in several studies, and insertional mutagenesis has been identified as a critical step in HBV-mediated HCC pathogenesis (13, 14). Integration sites were initially thought to be distributed randomly throughout the host genome, but data supporting a more deliberate process that preferentially involves transcribed regions of critical genes have been reported.(15–19).
The Epstein-Barr virus (EBV) (also known as human herpesvirus 5; HHV5) is a DNA virus that infects over 90% of the world's population before adolescence. It has been associated with a wide variety of human malignancies of epithelial, hematolymphoid, and mesenchymal derivation (2, 20, 21). Gastric carcinoma associated with EBV appears to comprise a distinct entity that is predominant in younger male individuals (22, 23). This subset of gastric carcinoma, 8 to 10% of cases, is more prevalent in Caucasian and Hispanic patients than Asians, and it shows no association with Helicobacter pylori infection (22). In these cases, EBV appears to play a direct oncogenic role through genome-wide alteration of promoter methylation (24), microRNA (miRNA) expression, and expression of genes involved in cell motility and transformation pathways (25). The integration status of EBV in gastric carcinoma remains poorly understood.
The discovery of novel cancer-associated viruses and the genomic effects of known viruses on the cancer cell genome remains incomplete, technically complex (26), and an arena in which significant resources have been and continue to be consumed. To provide an overview of the landscape of DNA virus-tumor associations in malignant cancers, we developed a method to conduct a comprehensive survey of The Cancer Genome Atlas (TCGA) RNA-Seq database for viral transcripts and, where applicable, their integration sites within the host genome.
MATERIALS AND METHODS
The Cancer Genome Atlas data.
Annotated RNA-Seq data were downloaded from the Cancer Genomics Hub (CGHub; https://cghub.ucsc.edu) and the TCGA Data Portal (https://tcga-data.nci.nih.gov/tcga/) at various time points between 18 May and 3 August 2012. Patient enrollment and utilization of data were conducted in accordance with TCGA human subjects protection and data access policies (http://cancergenome.nih.gov/PublishedContent/Files/pdfs/6.3.1_TCGA_Human_Subjects_and_Data_Access_policies_FINAL_011211.pdf).
RNA sequencing.
Total RNA for each sample was converted into a template molecule library for sequencing on an Illumina HiSeq 2000 according to the Illumina TruSeq sample preparation kit protocol. Briefly, poly(A) mRNA was purified from total RNA (1 μg) using poly(T) oligonucleotide-attached magnetic beads. The mRNA then was fragmented, and the first strand of cDNA was synthesized from the cleaved RNA fragments using reverse transcriptase and random primers. Following the synthesis of the second strand of cDNA, end repair was performed on overhangs using T4 DNA polymerase and Klenow DNA polymerase, followed by adenylation of the 3′ ends and ligation of sequencing adapters to the ends of the cDNA fragments. The purified cDNA templates were enriched by 15 cycles of PCR amplification and validated using BioAnalyzer (Agilent, Santa Clara, CA) to assess size, purity, and concentration of the purified cDNA libraries.
The cDNA libraries were placed on an Illumina c-Bot for paired-end (PE) cluster generation according to the protocol outlined in the Illumina HiSeq analysis user guide (part 11251649, RevA). The template cDNA libraries (1.5 μg) were hybridized to a flow cell, amplified, linearized, and denatured to create a flow cell with single-stranded DNA (ssDNA) ready for sequencing. Each flow cell was sequenced on an Illumina HiSeq Genome Analyzer. Each sample underwent one lane of PE sequencing according to the protocol outlined in the Illumina HiSeq user guide (part 11251649, RevA). After completion of the 50-cycle PE sequencing run, bases and quality values (FASTQ files) were generated for each read with the current Illumina pipeline.
Mapping/alignment.
We first performed quality checks on sequencing data using the HTSeq package (http://www-huber.embl.de/users/anders/HTSeq/doc/count.html). The raw paired-end (PE) reads in FASTQ format were then aligned to the human reference genome, GRCh37/hg19 (hg19), using MOSAIK open-source alignment software. MOSAIK works with PE reads and uses both a hashing scheme and the Smith-Waterman algorithm to produce gapped optimal alignments and to map exon junction-spanning reads with a local alignment option for RNA-Seq data.
Virus detection from RNA-Seq.
Our approach (39) started by computationally subtracting human sequences, followed by generating a set of nonhuman sequences (e.g., viruses) on RNA-Seq. Once raw PE reads from RNA-Seq were aligned to the human genome reference, any read with more than half a read length mapped to the human reference genome is removed along with its paired mate in this subtraction step. Thus, a set of nonhuman sequences was generated after human sequence subtraction. In the second step, our algorithm determined whether the nonhuman sequences match any known viral sequences by searching a comprehensive database that includes all known viral sequences (Genome Information Broker for Viruses [GIB-V]; http://gib-v.genes.nig.ac.jp/) and quantified virus representation by a measure of the virus genome coverage (or overall count of mapped reads) to determine the existence of viruses in human samples. The algorithm subsequently excluded nontranscribed viral genome elements to eliminate/reduce the potential of nonsense reads or inclusion of nontranscribed viral genomic elements. The expression level of each viral transcript was measured by the normalized depth of coverage within each viral transcript. The cutoff of viral gene expression detection was empirically determined by profiling the distribution of viral gene expression levels across multiple cancer-associated viruses (e.g., HPV16, HPV33, and EBV) and multiple patient samples. Any viral expression level below the cutoff was treated as no expression.
Identification of virus integration sites.
The genomes of viruses with expression levels detected in the previous steps were concatenated into a single genome, named chrVirus, with related annotation of each virus in refFlat format. A new hybrid reference genome, named hg19Virus, was built by combining hg19 and chrVirus. All PE reads without computational subtraction were again mapped to this reference (hg19Virus). If the PE reads were uniquely mapped with one end to hg19 and the other to chrVirus, the read pair was reported as a discordant read pair. All discordant reads were annotated by using the genes and viruses defined in the curated refFlat file. Our algorithm subsequently employed BLAT (27) to align the discordant reads against the hybrid reference genome (hg19Virus) and discards any discordant read pair if both discordant reads are aligned to the same gene or virus. It then clustered the remaining discordant read pairs that support the same integration (fusion) event (e.g., HBV-MLL4) and selected them as fusion candidates. A dynamic clustering procedure was implemented to accurately determine the exact fusion junction between a human gene and a virus. Specifically, the boundary for each discordant read cluster of a candidate fusion was estimated on the basis of discordant read mapping locations and orientations with fragment length distribution as a constraint of cluster size, which was measured by using the reads' genomic locations, excluding intronic sizes if mapped reads were located across adjacent exons in a candidate fusion. For the forward-aligned discordant reads in a fusion candidate, the clustering process started with the right-most read, and the genomic coordinate for the right-most read was used to define the in silico fusion junction, excluding outliers within the discordant read cluster. In order to remove outliers within a cluster, our algorithm implemented the robust “extreme studentized deviate” multiple-outlier procedure. If the outliers came from the right end of the cluster, the outliers were removed and the clustering process restarts with a new in silico fusion junction. If the outliers come from the left-most end of the cluster, the cluster size was reset with the in silico fusion junction intact by excluding the outlier reads. For the reverse-aligned discordant reads, the clustering process started with the left-most read, and the genomic coordinate for the left-most read was used to define the in silico fusion junction with the same outlier detection/removal processing step. For either side of the candidate fusion partner (gene or virus), this clustering process was performed independently. This dynamic clustering procedure accurately determined the exact fusion junction between a human gene and a virus. Meanwhile, an in silico sequence was generated using the consensus of reads within discordant read clusters for each fusion candidate to help the PCR primer design, which facilitates quick PCR validation.
RESULTS
RNA sequencing data from 3,775 malignant neoplasms in the TCGA database were interrogated for the presence and, as applicable, integration of site of viral transcripts. The overall median read number per sample was 132,183,938 (Table 1). None of the virus-positive cancers in this analysis harbored more than one type/strain of virus.
Table 1.
Neoplasms in The Cancer Genome Atlas Database surveyed for tumor-associated DNA viruses
| Tumor type | No. of samples analyzed | No. of reads per sample |
Freeze date (mo/day/yr) | ||
|---|---|---|---|---|---|
| Min | Median | Max | |||
| Breast carcinoma | 750 | 61000,388 | 109861,408 | 217446,562 | 8/2/2012 |
| Clear cell renal cell carcinoma | 460 | 30478,126 | 110937,134 | 242361,532 | 8/3/2012 |
| Ovarian serous cystadenocarcinoma | 419 | 102128,078 | 197051,552 | 452149,776 | 8/3/2012 |
| Uterine corpus endometrioid carcinomaa | 254 | 9988,571 | 31894,092 | 38324,550 | 8/3/2012 |
| Head-and-neck squamous cell carcinoma | 239 | 78123,066 | 152296,210 | 243733,050 | 8/2/2012 |
| Lung adenocarcinoma | 225 | 52466,630 | 103258,332 | 270583,684 | 8/3/2012 |
| Lung squamous cell carcinoma | 219 | 54253,346 | 127784,296 | 191438,728 | 8/2/2012 |
| Cutaneous melanoma | 214 | 91516,350 | 158684,346 | 269997,828 | 8/3/2012 |
| Acute myeloid leukemia | 179 | 105463,130 | 147722,794 | 190844,514 | 5/18/2012 |
| Glioblastoma | 168 | 78576,316 | 128354,626 | 240986,684 | 8/3/2012 |
| Thyroid carcinoma | 157 | 96982,636 | 159965,288 | 310309,740 | 8/2/2012 |
| Colon adenocarcinomaa | 138 | 9923,980 | 26751,110 | 37682,149 | 8/2/2012 |
| Gastric adenocarcinoma | 71 | 114694,894 | 184402,162 | 237944,836 | 8/2/2012 |
| Rectal adenocarcinomaa | 66 | 19527,419 | 27027,473 | 33162,004 | 8/2/2012 |
| Prostate adenocarcinoma | 53 | 65688,652 | 132183,938 | 214386,560 | 7/31/2012 |
| Papillary renal cell carcinoma | 47 | 91281,498 | 168971,506 | 243228,666 | 7/24/2012 |
| Lower-grade glioma | 47 | 120564,192 | 160524,446 | 205076,246 | 8/2/2012 |
| Hepatocellular carcinoma | 69 | 68607,776 | 140488,184 | 188335,198 | 8/3/2012 |
Non-paired-end data in TCGA database.
Human papillomavirus detection in malignant cancers.
We analyzed 239 squamous cell carcinomas of the head-and-neck region (HNSCC) available in the TCGA database. We detected HPV transcripts in 36 tumors as the following: 30 tumors with HPV16, 5 tumors with HPV33, and 1 tumor with HPV35. We also detected EBV in 1 tumor (discussed below). Among all cases with HPV transcripts, E7 was expressed in 22, E6 in 20, E1 in 17, and E4 in 8 tumors. In 24 tumors, HPV transcripts encoding key viral proteins/oncoproteins were integrated in the tumor genome, with the majority in association with known genes (Fig. 1 and 2). Tumors with HPV integration harbored the following types: 19 HPV16+, 4 HPV33+, and 1 HPV35+ (Table 2). Of the tumors with HPV integration, 18 have both E6 and E7 integration sites, 4 have only E7 integration sites, and 2 have only E6 integration sites.
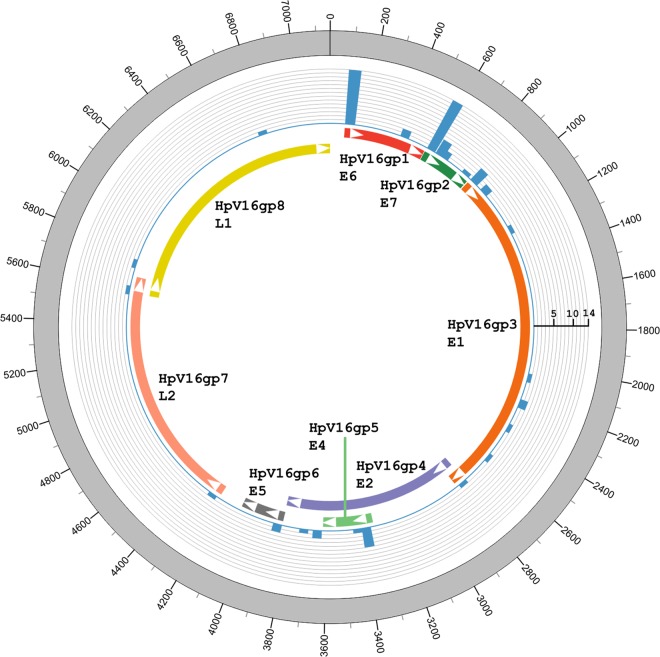
Fig 1.
Visualization of HPV16 integration breakpoints in the HPV16 genome. The frequency of integration breakpoints at different loci in the HPV16 genome is shown as a blue histogram. The scale bar indicates the number of tumors. The locations of the genes encoding HPV16 E6 (red), E7 (dark green), E1 (orange), E2 (purple), E4 (green), E5 (gray), L2 (light orange), and L1 (yellow) proteins are shown. Genomic positions are numbered.
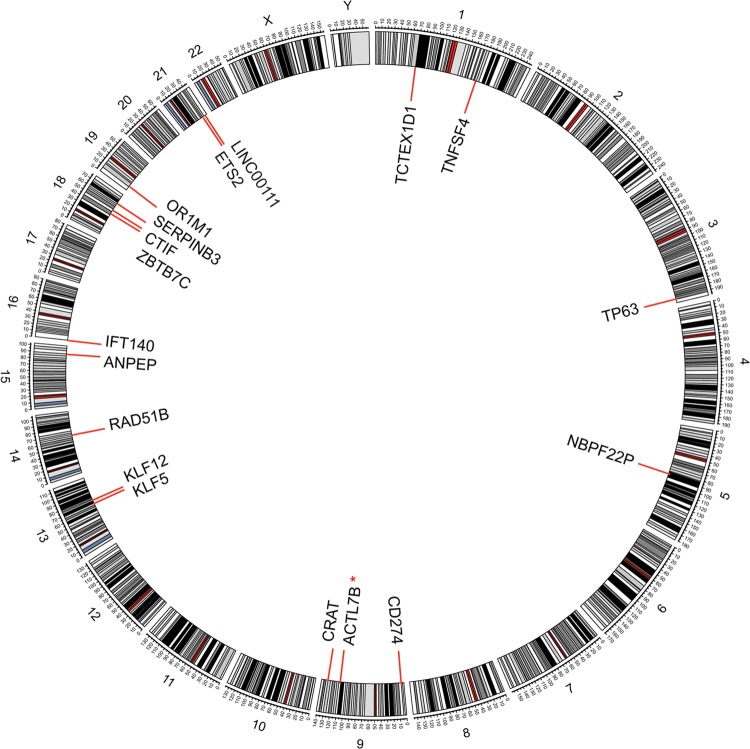
Fig 2.
Integration sites of HPV16 in head-and-neck squamous cell carcinoma tumors in the human genome (hg19). Chromosome numbers are shown (*, detected in two cases).
Table 2.
Head-and-neck squamous cell carcinoma tumors with integrated HPV transcripts
| TCGA ID and virus | Viral transcript | Virus site | Host gene | Integration site | Chromosome no. | Gene position |
|---|---|---|---|---|---|---|
| H736801A | ||||||
| HPV16 | HPV16gp1_E6 | 70 | ACTL7B | 3′ | 9 | 111259747 |
| HPV16 | HPV16gp2_E7 | 559 | ACTL7B | 3′ | 9 | 111241132 |
| HPV16 | HPV16gp3_E1 | 903 | ACTL7B | 3′ | 9 | 111237008 |
| HPV16 | HPV16gp4_E2 | 2769 | ACTL7B | 3′ | 9 | 111541176 |
| H740601A | ||||||
| HPV16 | HPV16gp1_E6 | 106 | ACTL7B | 3′ | 9 | 110495875 |
| HPV16 | HPV16gp2_E7 | 565 | ACTL7B | 3′ | 9 | 110497920 |
| H648101Aa | ||||||
| HPV16 | HPV16gp2_E7 | 812 | ANPEP | 5′ | 15 | 90358960 |
| HPV16 | HPV16gp3_E1 | 978 | ANPEP | 5′ | 15 | 90359954 |
| HPV16 | HPV16gp2_E7 | 649 | ANPEP | Intron 1 | 15 | 90350002 |
| HPV16 | HPV16gp5_E4 | 3355 | ANPEP | Intron 20 | 15 | 90333738 |
| H544301A | ||||||
| HPV16 | HPV16gp2_E7 | 561 | CD274 | Intron 4 | 9 | 5462885 |
| HPV16 | HPV16gp8_L1 | 6758 | CD274 | Intron 4 | 9 | 5464598 |
| H648701Aa | ||||||
| HPV16 | HPV16gp1_E6 | 43 | CRAT | 3′ | 9 | 131856423 |
| HPV16 | HPV16gp3_E1 | 2249 | CRAT | 3′ | 9 | 131856542 |
| HPV16 | HPV16gp2_E7 | 670 | CRAT | Intron 14 | 9 | 131857745 |
| H775401A | ||||||
| HPV16 | HPV16gp2_E7 | 663 | CTIF | 5′ | 18 | 45664322 |
| HPV16 | HPV16gp3_E1 | 1227 | CTIF | 5′ | 18 | 45664232 |
| H696101A | ||||||
| HPV16 | HPV16gp8_L1 | 5573 | ETS2 | Intron 4 | 21 | 40186953 |
| HPV16 | HPV16gp1_E6 | 75 | ETS2 | Intron 7 | 21 | 40191504 |
| HPV16 | HPV16gp2_E7 | 561 | ETS2 | Intron 7 | 21 | 40191445 |
| HPV16 | HPV16gp5_E4 | 3385 | ETS2 | Intron 8 | 21 | 40193588 |
| HPV16 | HPV16gp3_E1 | 2352 | ETS2 | Intron 7 | 21 | 40190767 |
| H647201A | ||||||
| HPV16 | HPV16gp1_E6 | 407 | IFT140 | Intron 18 | 16 | 1612024 |
| HPV16 | HPV16gp2_E7 | 561 | IFT140 | Intron 18 | 16 | 1612023 |
| HPV16 | HPV16gp3_E1 | 2206 | IFT140 | Intron 18 | 16 | 1612004 |
| H736901A | ||||||
| HPV16 | HPV16gp1_E6 | 60 | KLF12 | 3′ | 13 | 73910872 |
| HPV16 | HPV16gp2_E7 | 570 | KLF12 | 3′ | 13 | 73952914 |
| HPV16 | HPV16gp3_E1 | 944 | KLF12 | 3′ | 13 | 73910982 |
| H474101A | ||||||
| HPV16 | HPV16gp1_E6 | 447 | KLF5 | 5′ | 13 | 73620072 |
| H738501A | ||||||
| HPV16 | HPV16gp1_E6 | 119 | LINC00111 | 5′ | 21 | 43089331 |
| HPV16 | HPV16gp2_E7 | 620 | LINC00111 | 5′ | 21 | 43089341 |
| HPV16 | HPV16gp3_E1 | 889 | LINC00111 | 5′ | 21 | 43084796 |
| HPV16 | HPV16gp4_E2 | 3655 | LINC00111 | 5′ | 21 | 43087464 |
| HPV16 | HPV16gp5_E4 | 3355 | LINC00111 | 5′ | 21 | 43090560 |
| HPV16 | HPV16gp7_L2 | 4262 | LINC00111 | 5′ | 21 | 43059265 |
| H597101A | ||||||
| HPV16 | HPV16gp1_E6 | 115 | NBPF22P | 5′ | 5 | 84616300 |
| HPV16 | HPV16gp2_E7 | 567 | NBPF22P | 5′ | 5 | 84624716 |
| HPV16 | HPV16gp3_E1 | 2056 | NBPF22P | 5′ | 5 | 84619865 |
| HPV16 | HPV16gp5_E4 | 3355 | NBPF22P | 5′ | 5 | 84629561 |
| H544201A | ||||||
| HPV16 | HPV16gp1_E6 | 106 | OR1M1 | 5′ | 19 | 9113164 |
| HPV16 | HPV16gp2_E7 | 567 | OR1M1 | 5′ | 19 | 9113071 |
| H407701B | ||||||
| HPV16 | HPV16gp1_E6 | 73 | RAD51B | Intron 7 | 14 | 68758607 |
| HPV16 | HPV16gp3_E1 | 858 | RAD51B | Intron 7 | 14 | 68699906 |
| HPV16 | HPV16gp4_E2 | 3625 | RAD51B | Intron 7 | 14 | 68741641 |
| HPV16 | HPV16gp6_E5 | 3855 | RAD51B | Intron 7 | 14 | 68683627 |
| HPV16 | HPV16gp2_E7 | 565 | RAD51B | Intron 7 | 14 | 68704019 |
| HPV16 | HPV16gp5_E4 | 3434 | RAD51B | Intron 7 | 14 | 68741498 |
| H532301A | ||||||
| HPV16 | HPV16gp3_E1 | 903 | SERPINB3 | 3′ | 18 | 61317170 |
| HPV16 | HPV16gp8_L1 | 5701 | SERPINB3 | 3′ | 18 | 61317182 |
| HPV16 | HPV16gp1_E6 | 130 | SERPINB3 | Exon 8 | 18 | 61322624 |
| HPV16 | HPV16gp2_E7 | 561 | SERPINB3 | Exon 8 | 18 | 61322588 |
| H555901A | ||||||
| HPV16 | HPV16gp5_E4 | 3355 | TCTEX1D1 | Intron 1 | 1 | 67220366 |
| HPV16 | HPV16gp2_E7 | 568 | TCTEX1D1 | Intron 2 | 1 | 67236076 |
| HPV16 | HPV16gp4_E2 | 3717 | TCTEX1D1 | Intron 2 | 1 | 67229043 |
| HPV16 | HPV16gp6_E5 | 3853 | TCTEX1D1 | Intron 2 | 1 | 67229161 |
| HPV16 | HPV16gp3_E1 | 2567 | TCTEX1D1 | Intron 2 | 1 | 67229895 |
| HPV16 | HPV16gp1_E6 | 124 | TCTEX1D1 | Intron 3 | 1 | 67239538 |
| H647301A | ||||||
| HPV16 | HPV16gp1_E6 | 106 | TNFSF4 | 5UTR | 1 | 173176246 |
| HPV16 | HV16gp2_E7 | 577 | TNFSF4 | 5UTR | 1 | 173176043 |
| H474101A | ||||||
| HPV16 | HPV16gp1_E6 | 117 | TP63 | 5′ | 3 | 189226255 |
| HPV16 | HPV16gp2_E7 | 578 | TP63 | 5′ | 3 | 189239853 |
| H775401A | ||||||
| HPV16 | HPV16gp1_E6 | 139 | ZBTB7C | Intron 1 | 18 | 45567460 |
| HPV16 | HPV16gp2_E7 | 567 | ZBTB7C | Intron 1 | 18 | 45567416 |
| H710001A | ||||||
| HPV33 | HPV33gp1_E6 | 509 | EPSTI1 | Intron 5 | 13 | 43537483 |
| HPV33 | HPV33gp5_E4 | 3356 | EPSTI1 | Intron 6 | 13 | 43528079 |
| HPV33 | HPV33gp8_L1 | 5597 | EPSTI1 | Intron 6 | 13 | 43528256 |
| H647101A | ||||||
| HPV33 | HPV33gp1_E6 | 93 | TEAD1 | Intron 5 | 11 | 12897532 |
| HPV33 | HPV33gp2_E7 | 559 | TEAD1 | Intron 5 | 11 | 12897531 |
| HPV33 | HPV33gp3_E1 | 858 | TEAD1 | Intron 5 | 11 | 12897627 |
| H783201A | ||||||
| HPV33 | HPV33gp1_E6 | 116 | MIR802 | 5′ | 21 | 36470118 |
| HPV33 | HPV33gp2_E7 | 560 | MIR802 | 5′ | 21 | 36470162 |
| HPV33 | HPV33gp3_E1 | 858 | MIR802 | 5′ | 21 | 36470118 |
| H710001A | ||||||
| HPV33 | HPV33gp5_E4 | 3347 | PHLDB2 | Intron 1 | 3 | 111451446 |
| HPV33 | HPV33gp3_E1 | 1905 | PHLDB2 | Intron 1 | 3 | 111499049 |
| HPV33 | HPV33gp2_E7 | 671 | PHLDB2 | Intron 1 | 3 | 111496513 |
| HPV33 | HPV33gp3_E1 | 858 | PLCXD2 | Intron 1 | 3 | 111413679 |
| HPV33 | HPV33gp2_E7 | 747 | PLCXD2 | Intron 1 | 3 | 111431547 |
| H759101A | ||||||
| HPV35 | HPV35gp2_E7 | 698 | ACCN1 | Intron 1 | 17 | 32128921 |
| HPV35 | HPV35gp3_E1 | 863 | ACCN1 | Intron 1 | 17 | 32128931 |
| HPV35 | HPV35gp1_E6 | 128 | ACCN1 | Intron 1 | 17 | 32143974 |
Positive for HPV DNA by colorimetric in situ hybridization and/or for p16 overexpression by immunohistochemistry (data unavailable for the other cases).
The clinicopathologic characteristics of HNSCC patients with available data (n = 35) are summarized in Tables 3 and 4. HPV status detected by our method correlated with perfect sensitivity and specificity with known clinicopathologic variables and with established methods for HPV detection (colorimetric in situ hybridization and/or p16ink4a expression) (28). For this HNSCC data set, the sensitivity for HPV16 detection was 100%, with 95% confidence intervals (CI) of 67.6 to 100%, and specificity for HPV16 detection was 100%, with 95% CI of 90.4 to 100%, as reported previously (39).
Table 3.
Clinicopathologic characteristics of patients with head-and-neck squamous cell carcinomaa
| Parameter | HPV positive |
HPV negative |
All HNSCC |
P valueb | |||
|---|---|---|---|---|---|---|---|
| n | % | n | % | n | % | ||
| Gender | |||||||
| Female | 4 | 11.4 | 59 | 29.2 | 63 | 26.6 | |
| Male | 31 | 88.6 | 143 | 70.8 | 174 | 73.4 | |
| All | 35 | 100 | 202 | 100 | 237 | 100 | 0.04 |
| Tumor site | |||||||
| Alveolar ridge | 2 | 5.7 | 4 | 2 | 6 | 2.5 | |
| Base of tongue | 5 | 14.3 | 4 | 2 | 9 | 3.8 | |
| Buccal mucosa | 0 | 0 | 7 | 3.5 | 7 | 3 | |
| Floor of mouth | 1 | 2.9 | 21 | 10.4 | 22 | 9.3 | |
| Hard palate | 1 | 2.9 | 2 | 1 | 3 | 1.3 | |
| Hypopharynx | 1 | 2.9 | 0 | 0 | 1 | 0.4 | |
| Larynx | 1 | 2.9 | 70 | 34.8 | 71 | 30.1 | |
| Lip | 0 | 0 | 1 | 0.5 | 1 | 0.4 | |
| Oral cavity | 4 | 11.4 | 31 | 15.4 | 35 | 14.8 | |
| Oral tongue | 3 | 8.6 | 56 | 27.9 | 59 | 25 | |
| Oropharynx | 1 | 2.9 | 2 | 1 | 3 | 1.3 | |
| Tonsil | 16 | 45.7 | 3 | 1.5 | 19 | 8.1 | |
| All | 35 | 100 | 201 | 100 | 236 | 100 | <0.0001 |
| Tumor stagec | |||||||
| I | 1 | 3 | 12 | 6.1 | 13 | 5.7 | |
| II | 5 | 15.2 | 46 | 23.5 | 51 | 22.3 | |
| III | 5 | 15.2 | 40 | 20.4 | 45 | 19.6 | |
| IVA | 22 | 66.7 | 92 | 46.9 | 114 | 49.8 | |
| IVB | 0 | 0 | 4 | 2 | 4 | 1.8 | |
| IVC | 0 | 0 | 2 | 1 | 2 | 0.9 | |
| All | 33 | 100 | 196 | 100 | 229 | 100 | 0.56 |
| Lymph node status | |||||||
| N0 | 12 | 40 | 77 | 38.5 | 89 | 38.7 | |
| N1 | 5 | 16.7 | 32 | 16 | 37 | 16.1 | |
| N2 | 1 | 3.3 | 9 | 4.5 | 10 | 4.3 | |
| N2a | 1 | 3.3 | 4 | 2 | 5 | 2.2 | |
| N2b | 9 | 30 | 38 | 19 | 47 | 20.4 | |
| N2c | 2 | 6.7 | 27 | 13.5 | 29 | 12.6 | |
| N3 | 0 | 0 | 3 | 1.5 | 3 | 1.3 | |
| NX | 0 | 0 | 10 | 5 | 10 | 4.3 | |
| All | 30 | 100 | 200 | 100 | 230 | 100 | 0.73 |
| Metastasis | |||||||
| M0 | 35 | 100 | 199 | 99 | 234 | 99.2 | |
| M1 | 0 | 0 | 2 | 1 | 2 | 0.8 | |
| All | 35 | 100 | 201 | 100 | 236 | 100 | 1 |
| Histologic grade | |||||||
| G1 | 1 | 2.9 | 10 | 5 | 11 | 4.6 | |
| G2 | 14 | 40 | 138 | 68.3 | 152 | 64.1 | |
| G3 | 17 | 48.6 | 49 | 24.3 | 66 | 27.9 | |
| G4 | 1 | 2.9 | 0 | 0 | 1 | 0.4 | |
| GX | 2 | 5.7 | 5 | 2.5 | 7 | 3 | |
| All | 35 | 100 | 202 | 100 | 237 | 100 | 0.0022 |
| vHPV in situ hybridization | |||||||
| Negative | 0 | 0 | 35 | 100 | 35 | 85.4 | |
| Positive | 6 | 100 | 0 | 0 | 6 | 14.6 | |
| All | 6 | 100 | 35 | 100 | 41 | 100 | <0.0001 |
| HPV status by p16 IHC | |||||||
| Negative | 0 | 0 | 36 | 100 | 36 | 83.7 | |
| Positive | 7 | 100 | 0 | 0 | 7 | 16.3 | |
| All | 7 | 100 | 36 | 100 | 43 | 100 | <0.0001 |
Nominal variables grouped by HPV status (integrated and nonintegrated) detected by RNA-Seq data. IHC, immunohistochemistry; IQR, interquartile range.
Determined by Fischer's exact test.
According to AJCC staging criteria.
Table 4.
Continuous variables of clinicopathologic characteristics of patients with head-and-neck squamous cell carcinoma
| Parameter | n | Value for HPV status and smoking historya |
P valuec | ||||
|---|---|---|---|---|---|---|---|
| Min | Mean | Max | IQR | No. NAb | |||
| HPV status | |||||||
| Positive | 35 | 35 | 58 | 82 | 13.5 | 0 | |
| Negative | 202 | 19 | 62 | 87 | 14.5 | 0 | |
| All | 237 | 19 | 62 | 87 | 15 | 0 | 0.05 |
| Smoking history | |||||||
| Positive | 18 | 0 | 30.5 | 102.5 | 28.8 | 17 | |
| Negative | 114 | 0.7 | 48 | 180 | 37.1 | 88 | |
| All | 132 | 0 | 45 | 180 | 32.2 | 105 | 0.0084 |
For HPV status, minimum, mean, and maximum values are given as age in years. For smoking history, minimum, mean, and maximum values are given as pack-years.
Number of cases for which no data were available.
Determined by Mann-Whitney test.
We also detected HPV16 in two tumors of histologic types rarely associated with this virus. From a group of 219 lung squamous cell carcinoma tumors, one case harbored HPV16, where E1, E6, and E7 transcripts were highly expressed and integrated in NROB1 (also known as DAX1), a gene involved in steroidogenesis and cell cycle regulation. In this case, no E2, E4, E5, L1, or L2 transcripts were detected. Additionally, 1 of 253 endometrial carcinoma tumors harbored HPV16, where E1, E2, E4, E5, E6, and E7 transcripts were highly expressed, and no L1 or L2 transcripts were detected. The integration site of the latter could not be determined, because the TCGA endometrial carcinoma RNA-Seq data were not paired end.
Epstein-Barr virus detection in malignant cancers.
We analyzed 71 cases of gastric carcinoma in the TCGA database and detected EBV transcripts in 4 (5.6%) tumors. Of the four tumors with unequivocal EBV association, all harbored transcripts encoding A73, RPMS1, BARF0, BALF3, BALF4, BALF5, LF1, LF2, and BILF1. One tumor additionally had significant levels of transcripts encoding BNLF2a/b, LMP1, and LMP2A (Table 5). Of note, unlike others who demonstrated EBNA1 expression in vitro in gastric carcinoma cell lines (29), we did not identify EBNA1 in any of the gastric carcinoma tumor samples with EBV transcripts. This might be due to biological variation between primary tumor samples and cell lines or low EBNA1 mRNA copy numbers. In the single head-and-neck squamous cell carcinoma tumor associated with EBV, the most abundant transcripts included those that encode BARF1/2, BdRF1, BMRF2, BLF1, BNLF2b, BBLF1, BMRF1, BLRF2, and BNLF2a. None of the tumors analyzed in this study had evidence of EBV integration into the host genome.
Table 5.
EBV transcripts detected in four gastric carcinoma tumors
| Gene symbol | Expression level for TCGA ID no.a: |
|||
|---|---|---|---|---|
| S425301A | S427101A | S429801A | S437601A | |
| EBV_RPMS1 | 66 | 73 | 2 | 65 |
| EBV_BILF1 | 45 | 48 | 1 | 36 |
| EBV_LF1 | 57 | 62 | 2 | 47 |
| EBV_LF2 | 105 | 118 | 4 | 88 |
| EBV_BALF5 | 100 | 112 | 4 | 101 |
| EBV_A73 | 206 | 229 | 7 | 203 |
| EBV_BALF3 | 162 | 174 | 5 | 157 |
| EBV_BALF4 | 160 | 156 | 4 | 138 |
| EBV_BARF0 | 197 | 244 | 8 | 222 |
| EBV_LMP-2A-1 | 10 | 0 | 0 | 0 |
| EBV_LMP-1 | 22 | 0 | 0 | 1 |
| EBV_BNLF2a | 98 | 0 | 0 | 1 |
| EBV_BNLF2b | 106 | 0 | 0 | 1 |
Data refer to the expression level of viral transcripts estimated by the normalized depth of coverage within each viral transcript.
Hepatitis B virus detection in malignant cancers.
We analyzed 69 HCC tumors available in the TCGA database and detected HBV transcripts in 16 (23%) tumors. Eight of these patients had serologic evidence of HBV infection, and one patient was HBV (and hepatitis C virus) seronegative. Serologic data on the remaining patients were either negative or not available. Virus integration was identified in 15 (94%) of these tumors. Our data demonstrate frequent HBV integration within previously identified genes, namely, TERT (5 tumors) and MLL4 (3 tumors), suggesting that these sites are particularly susceptible to HBV insertion. None of the tumors had both MLL4 and TERT involvement. Other insertion sites were restricted to single cases (Table 6). Of the tumors with HBV integration, 11 have X protein integration sites, 8 have S protein integration sites, 6 have core/E antigen integration sites, 4 have pre-S protein integration sites, 4 have polymerase 1 integration sites, and 3 have polymerase 2 integration sites. Integration of two or more HBV genes was detected in 8 tumors, whereas in the remaining 7 tumors integration of only one HBV gene was detected. Interestingly, the latter group included the 3 tumors with MLL4 involvement and 2 with TERT involvement.
Table 6.
Hepatocellular carcinoma tumors with integrated HBV transcripts
| TCGA ID | Viral transcript | Virus site | Host gene | Integration site | Chromosome no. | Gene position |
|---|---|---|---|---|---|---|
| L525801A | HBVgp3_X protein | 1742 | MLL4 | Intron 5 | 19 | 36214027 |
| L526201A | HBVgp1_polymerase-1 | 1245 | FOXI2 | 5′ | 10 | 129391432 |
| L526201A | HBVgp3_X protein | 1419 | FOXI2 | 5′ | 10 | 129391427 |
| L526401A | HBVgp2_S protein | 282 | ITGAD | 5′ | 16 | 31402241 |
| L526401A | HBVgp4_core/E antigen | 1991 | ITGAD | Intron 5 | 16 | 31413434 |
| L526401A | HBVgp4_precore/core protein | 1883 | ITGAD | Intron 5 | 16 | 31413431 |
| L526401A | HBVgp3_X protein | 1748 | TEAD1 | Intron 2 | 11 | 12711205 |
| L526401A | HBVgp1_polymerase-1 | 1359 | TECRL | 3′ | 4 | 63651241 |
| L526401A | HBVgp2_S protein | 372 | TECRL | 3′ | 4 | 63647552 |
| L526401A | HBVgp3_X protein | 1373 | TECRL | 3′ | 4 | 63651170 |
| LA10W01A | HBVgp2_S protein | 211 | CLDN10 | Intron 3 | 13 | 96216582 |
| LA10W01A | HBVgp2_pre-S1/pre-S2/S-2 | 2943 | CLDN10 | Intron 3 | 13 | 96229482 |
| LA10W01A | HBVgp2_pre-S2/S-1 | 42 | CLDN10 | Intron 3 | 13 | 96216538 |
| LA10W01A | HBVgp4_core/E antigen | 2290 | CLDN10 | Intron 3 | 13 | 96229477 |
| LA10W01A | HBVgp3_X protein | 1540 | DPY19L2P1 | 3′ | 7 | 35111018 |
| LA10W01A | HBVgp2_S protein | 341 | KCNH1 | Intron 7 | 1 | 210977324 |
| LA10W01A | HBVgp3_X protein | 1800 | KCNH1 | Intron 7 | 1 | 211058920 |
| LA11601A | HBVgp2_S protein | 337 | C19orf55 | Intron 6 | 19 | 36255648 |
| LA11601A | HBVgp2_S protein | 420 | MLL4 | Intron 5 | 19 | 36213891 |
| LA11901A | HBVgp4_core/E antigen | 2097 | MLL4 | Exon 3 | 19 | 36212301 |
| LA1EA01A | HBVgp1_polymerase-2 | 2403 | GLI2 | Intron 1 | 2 | 121585830 |
| LA1EA01A | HBVgp2_S protein | 412 | GLI2 | Intron 1 | 2 | 121585623 |
| LA1EA01A | HBVgp2_pre-S2/S-1 | 23 | GLI2 | Intron 1 | 2 | 121585885 |
| LA1EA01A | HBVgp4_core/E antigen | 1966 | GLI2 | Intron 1 | 2 | 121585602 |
| LA1EA01A | HBVgp2_pre-S2/S-1 | 25 | TERT | Intron 2 | 5 | 1291950 |
| LA1EA01A | HBVgp3_X protein | 1675 | VRK2 | 5′ | 2 | 57459913 |
| LA1EI01A | HBVgp2_S protein | 386 | FN1 | Intron 12 | 2 | 216279398 |
| LA1EI01A | HBVgp4_core/E antigen | 2006 | GALNTL6 | Intron 2 | 4 | 173136585 |
| LA1EL01A | HBVgp3_X protein | 1659 | LOC282980 | 3′ | 10 | 1863160 |
| LA1EL01A | HBVgp3_X protein | 1684 | NCOR2 | Intron 12 | 12 | 124912988 |
| LA1HT01A | HBVgp2_S protein | 406 | TERT | Intron 1 | 5 | 1294653 |
| LA25U01A | HBVgp1_polymerase-1 | 1369 | MIR4457 | 3′ | 5 | 1295462 |
| LA25U01A | HBVgp3_X protein | 1373 | MIR4457 | 3′ | 5 | 1295516 |
| LA25U01A | HBVgp2_S protein | 433 | OBFC2A | 5′ | 2 | 192364684 |
| LA25U01A | HBVgp3_X protein | 1730 | OBFC2A | 5′ | 2 | 192367406 |
| LA25U01A | HBVgp3_X protein | 1569 | TERT | Intron 1 | 5 | 1294676 |
| LA25Z01A | HBVgp3_X protein | 1562 | NDUFA4 | 3′ | 7 | 10697705 |
| LA3CJ01A | HBVgp3_X protein | 1782 | IL-6 | 5′ | 7 | 22735275 |
| LA3CJ01A | HBVgp3_X protein | 1711 | TERT | Intron 1 | 5 | 1295037 |
| LA3CK01A | HBVgp1_polymerase-1 | 1330 | DDX18 | 5′ | 2 | 118543485 |
| LA3CK01A | HBVgp3_X protein | 1584 | DDX18 | 5′ | 2 | 118543232 |
| LA3CK01A | HBVgp1_polymerase-2 | 2735 | RNU6ATAC | 3′ | 9 | 137028935 |
| LA3CK01A | HBVgp3_X protein | 1635 | RNU6ATAC | 3′ | 9 | 137029164 |
| LA3MB01A | HBVgp1_polymerase-2 | 2391 | TERT | Intron 6 | 5 | 1272107 |
| LA3MB01A | HBVgp2_S protein | 406 | TERT | Intron 6 | 5 | 1272167 |
| LA3MB01A | HBVgp3_X protein | 1577 | TERT | Intron 6 | 5 | 1272106 |
| LA3MB01A | HBVgp4_core/E antigen | 2035 | TERT | Intron 6 | 5 | 1272250 |
| LA3MB01A | HBVgp3_X protein | 1503 | CDK15 | Intron 8 | 2 | 202709075 |
| LA3MB01A | HBVgp3_X protein | 1660 | MIR4424 | 5′ | 1 | 178593988 |
We additionally identified HBV transcripts in one case among 460 clear-cell renal cell carcinoma tumors analyzed. In this tumor, as well as in one HCC tumor, we detected HBV S protein transcripts integrated into GLI2, generally considered a marker of activation of the sonic hedgehog signaling pathway, which has been shown to play a role in aggressive types of renal cell carcinoma (30–32) and in HCC (33, 34).
Malignant cancers without detectable DNA viruses.
After confirming the accuracy of our tool (39) in malignancies that are known to be associated with DNA viruses, we turned our attention to screening all remaining TCGA tumors on which RNA-Seq data were available. Our analysis showed no evidence of transcribed viral elements in any of the following neoplasms: acute myeloid leukemia, breast carcinoma (ductal and lobular), colonic and rectal adenocarcinoma, lung adenocarcinoma, prostate adenocarcinoma, cutaneous melanoma, ovarian serous cystadenocarcinoma, papillary renal cell carcinoma, thyroid carcinoma, lower-grade glioma, and glioblastoma multiforme.
DISCUSSION
The ability to identify viral integration points within the host genome using RNA-Seq provides a useful tool to study DNA viruses in cancer. For instance, our findings are consistent with other reports that have demonstrated preferential HBV insertion into MLL4 and TERT in HCC (18, 19, 35). Although such a finding might result from an inherent susceptibility for insertional mutagenesis at these loci, the fact that these two genes are more recurrently involved in HCC might reflect a selection bias, whereby mutation of MLL4 or TERT confers a selective advantage that leads to tumor development. Support for the latter possibility is suggested by our data indicating that among 7 HCC tumors with HBV insertion at a single gene locus, 5 tumors (71%) had involvement of MLL4 or TERT (3 with MLL4 involvement, 2 with TERT involvement). This suggests that the threshold for malignant transformation is lower when these genes, particularly MLL4, are disrupted through HBV insertional mutagenesis.
The interplay between viral infections and the miRNA machinery is the subject of intensive investigation. In addition to the modulating effect of host miRNAs on viral gene expression, mounting evidence suggests the existence of viral mechanisms leading to reprogramming of host miRNA machinery (36). In a recent study by Wang and colleagues, the HBV X protein was demonstrated to trigger selective miRNA downregulation (37). Among the tumors with HBV insertion in this study, we identified two with involvement of miRNA genes (MIR4457 and MIR4424). The function of these miRNAs in HCC is unknown. Interestingly, both genes have X protein insertion sites, suggesting that insertional mutagenesis comprises another mechanism by HBV which modulates the host miRNA repertoire.
The pathogenetic role of DNA viruses has been well established in some cancers but remains unsettled in others. One of the main findings in our study is the absence of transcribed DNA viral transcripts in some of the most prevalent human cancers, including acute myeloid leukemia, cutaneous melanoma, low- and high-grade gliomas of the brain, and adenocarcinomas of the breast, colon and rectum, lung, prostate, ovary, kidney, and thyroid. In squamous cell carcinoma of the head and neck, our analysis achieved 100% sensitivity and specificity compared to established methods (28) for HPV detection. Furthermore, the vast majority of positive results across all cancers assessed were in line with expected outcomes (e.g., HPV in squamous cell carcinoma of the head and neck, HBV in hepatocellular carcinoma, etc.). On the basis of these results, the probability of false-negative results in this analysis is believed to be low. Indeed, Bexfield and Kellam (38) performed an elegant simulation to estimate the probability of detecting viral sequences based on the viral genome transcript sequence frequency and the number of sequence reads generated. On the basis of their model, 10 million sequence reads gives a >99.99% probability of detecting at least one viral sequence if every cell is infected and the viral genome transcript sequence frequency is 0.0001%. However, 10 million sequence reads gives a >60% probability of detecting at least one viral sequence if only 1 in 10 cells is infected. For our TCGA cohort, the median number of sequence reads is more than 100 million (Table 1). Thus, we have a >99.9% probability of detecting at least one viral sequence in the TCGA cohort even if only 1 in 10 cells is infected with 0.0001% viral genome transcript sequence frequency. However, the probabilities of detecting viral sequence decrease to approximately 84% with 20 million reads. Accordingly, and unless yet-unknown pathogenic DNA viruses are discovered in the future, the yield of future searches for DNA viruses in these types of cancers is likely to be limited.
One of the limitations of this study is the lack of available corresponding tissue samples for direct validation. For instance, further evaluation of the single case of clear cell renal cell carcinoma with HBV transcripts would have been warranted. In addition, it could be argued that RNA-Seq might not detect integration events that are biologically deleterious but do not result in expression of viral transcripts. The latter point, however, does not seem to be a common event (13–15, 18, 19, 26). However, while further validation of specific findings identified in this study is warranted, the overall outcome of the analysis highlights the utility of RNA-Seq in detecting tumor-associated DNA viruses and identifying viral integration sites that may unravel novel mechanisms of cancer pathogenesis.s
ACKNOWLEDGMENTS
This work was supported in part by TCGA grant U24CA143883, National Center for Research Resources grant UL1TR000371 (F.M.-B., J.N.W., X.S.), and The University of Texas MD Anderson Cancer Center support grant P30 CA016672. None of the funding sources had any role in study design, data collection, data analysis, data interpretation, or writing of the manuscript.
J.D.K. and X.S. conceived and designed the study and prepared the manuscript. X.S. designed and developed the bioinformatics algorithm. X.S., Y.C., H.Y., J.Z., and E.J.T. performed sequencing data analysis and statistical data analysis. N.M.T., M.D.W., F.M.-B., and L.J.M. reviewed findings for specific cancer types and also wrote and/or edited the manuscript. J.N.W. reviewed bioinformatics design and results and also wrote and/or edited the manuscript.
None of the authors has conflicts of interest pertaining to this study to declare.
Footnotes
Published ahead of print 5 June 2013
REFERENCES
- 1. Martin D, Gutkind JS. 2008. Human tumor-associated viruses and new insights into the molecular mechanisms of cancer. Oncogene 27(Suppl. 2):S31–S42 [DOI] [PubMed] [Google Scholar]
- 2. Ng SB, Khoury JD. 2009. Epstein-Barr virus in lymphoproliferative processes: an update for the diagnostic pathologist. Adv. Anat. Pathol. 16:40–55 [DOI] [PubMed] [Google Scholar]
- 3. Poreba E, Broniarczyk JK, Gozdzicka-Jozefiak A. 2011. Epigenetic mechanisms in virus-induced tumorigenesis. Clin. Epigenet. 2:233–247 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Boss IW, Plaisance KB, Renne R. 2009. Role of virus-encoded microRNAs in herpesvirus biology. Trends Microbiol. 17:544–553 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Moody CA, Laimins LA. 2010. Human papillomavirus oncoproteins: pathways to transformation. Nat. Rev. Cancer 10:550–560 [DOI] [PubMed] [Google Scholar]
- 6. Alazawi W, Pett M, Arch B, Scott L, Freeman T, Stanley MA, Coleman N. 2002. Changes in cervical keratinocyte gene expression associated with integration of human papillomavirus 16. Cancer Res. 62:6959–6965 [PubMed] [Google Scholar]
- 7. Pett MR, Herdman MT, Palmer RD, Yeo GS, Shivji MK, Stanley MA, Coleman N. 2006. Selection of cervical keratinocytes containing integrated HPV16 associates with episome loss and an endogenous antiviral response. Proc. Natl. Acad. Sci. U. S. A. 103:3822–3827 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Dall KL, Scarpini CG, Roberts I, Winder DM, Stanley MA, Muralidhar B, Herdman MT, Pett MR, Coleman N. 2008. Characterization of naturally occurring HPV16 integration sites isolated from cervical keratinocytes under noncompetitive conditions. Cancer Res. 68:8249–8259 [DOI] [PubMed] [Google Scholar]
- 9. Wentzensen N, Vinokurova S, von Knebel Doeberitz M. 2004. Systematic review of genomic integration sites of human papillomavirus genomes in epithelial dysplasia and invasive cancer of the female lower genital tract. Cancer Res. 64:3878–3884 [DOI] [PubMed] [Google Scholar]
- 10. Ziegert C, Wentzensen N, Vinokurova S, Kisseljov F, Einenkel J, Hoeckel M, von Knebel Doeberitz M. 2003. A comprehensive analysis of HPV integration loci in anogenital lesions combining transcript and genome-based amplification techniques. Oncogene 22:3977–3984 [DOI] [PubMed] [Google Scholar]
- 11. Smith PP, Friedman CL, Bryant EM, McDougall JK. 1992. Viral integration and fragile sites in human papillomavirus-immortalized human keratinocyte cell lines. Genes Chromosomes Cancer 5:150–157 [DOI] [PubMed] [Google Scholar]
- 12. Dandri M, Locarnini S. 2012. New insight in the pathobiology of hepatitis B virus infection. Gut 61(Suppl. 1):i6–i17 [DOI] [PubMed] [Google Scholar]
- 13. Fujimoto A, Totoki Y, Abe T, Boroevich KA, Hosoda F, Nguyen HH, Aoki M, Hosono N, Kubo M, Miya F, Arai Y, Takahashi H, Shirakihara T, Nagasaki M, Shibuya T, Nakano K, Watanabe-Makino K, Tanaka H, Nakamura H, Kusuda J, Ojima H, Shimada K, Okusaka T, Ueno M, Shigekawa Y, Kawakami Y, Arihiro K, Ohdan H, Gotoh K, Ishikawa O, Ariizumi S, Yamamoto M, Yamada T, Chayama K, Kosuge T, Yamaue H, Kamatani N, Miyano S, Nakagama H, Nakamura Y, Tsunoda T, Shibata T, Nakagawa H. 2012. Whole-genome sequencing of liver cancers identifies etiological influences on mutation patterns and recurrent mutations in chromatin regulators. Nat. Genet. 44:760–764 [DOI] [PubMed] [Google Scholar]
- 14. Sung WK, Zheng H, Li S, Chen R, Liu X, Li Y, Lee NP, Lee WH, Ariyaratne PN, Tennakoon C, Mulawadi FH, Wong KF, Liu AM, Poon RT, Fan ST, Chan KL, Gong Z, Hu Y, Lin Z, Wang G, Zhang Q, Barber TD, Chou WC, Aggarwal A, Hao K, Zhou W, Zhang C, Hardwick J, Buser C, Xu J, Kan Z, Dai H, Mao M, Reinhard C, Wang J, Luk JM. 2012. Genome-wide survey of recurrent HBV integration in hepatocellular carcinoma. Nat. Genet. 44:765–769 [DOI] [PubMed] [Google Scholar]
- 15. Murakami Y, Saigo K, Takashima H, Minami M, Okanoue T, Brechot C, Paterlini-Brechot P. 2005. Large scaled analysis of hepatitis B virus (HBV) DNA integration in HBV related hepatocellular carcinomas. Gut 54:1162–1168 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Dejean A, Bougueleret L, Grzeschik KH, Tiollais P. 1986. Hepatitis B virus DNA integration in a sequence homologous to v-erb-A and steroid receptor genes in a hepatocellular carcinoma. Nature 322:70–72 [DOI] [PubMed] [Google Scholar]
- 17. Wang J, Chenivesse X, Henglein B, Brechot C. 1990. Hepatitis B virus integration in a cyclin A gene in a hepatocellular carcinoma. Nature 343:555–557 [DOI] [PubMed] [Google Scholar]
- 18. Paterlini-Brechot P, Saigo K, Murakami Y, Chami M, Gozuacik D, Mugnier C, Lagorce D, Brechot C. 2003. Hepatitis B virus-related insertional mutagenesis occurs frequently in human liver cancers and recurrently targets human telomerase gene. Oncogene 22:3911–3916 [DOI] [PubMed] [Google Scholar]
- 19. Ferber MJ, Montoya DP, Yu C, Aderca I, McGee A, Thorland EC, Nagorney DM, Gostout BS, Burgart LJ, Boix L, Bruix J, McMahon BJ, Cheung TH, Chung TK, Wong YF, Smith DI, Roberts LR. 2003. Integrations of the hepatitis B virus (HBV) and human papillomavirus (HPV) into the human telomerase reverse transcriptase (hTERT) gene in liver and cervical cancers. Oncogene 22:3813–3820 [DOI] [PubMed] [Google Scholar]
- 20. Deyrup AT. 2008. Epstein-Barr virus-associated epithelial and mesenchymal neoplasms. Hum. Pathol. 39:473–483 [DOI] [PubMed] [Google Scholar]
- 21. Thompson MP, Kurzrock R. 2004. Epstein-Barr virus and cancer. Clin. Cancer Res. 10:803–821 [DOI] [PubMed] [Google Scholar]
- 22. Lee JH, Kim SH, Han SH, An JS, Lee ES, Kim YS. 2009. Clinicopathological and molecular characteristics of Epstein-Barr virus-associated gastric carcinoma: a meta-analysis. J. Gastroenterol. Hepatol. 24:354–365 [DOI] [PubMed] [Google Scholar]
- 23. Takada K. 2000. Epstein-Barr virus and gastric carcinoma. Mol. Pathol. 53:255–261 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Zhao J, Liang Q, Cheung KF, Kang W, Lung RW, Tong JH, To KF, Sung JJ, Yu J. 2012. Genome-wide identification of Epstein-Barr virus-driven promoter methylation profiles of human genes in gastric cancer cells. Cancer 119:304–312 [DOI] [PubMed] [Google Scholar]
- 25. Marquitz AR, Mathur A, Shair KH, Raab-Traub N. 2012. Infection of Epstein-Barr virus in a gastric carcinoma cell line induces anchorage independence and global changes in gene expression. Proc. Natl. Acad. Sci. U. S. A. 109:9593–9598 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Rozenblatt-Rosen O, Deo RC, Padi M, Adelmant G, Calderwood MA, Rolland T, Grace M, Dricot A, Askenazi M, Tavares M, Pevzner SJ, Abderazzaq F, Byrdsong D, Carvunis AR, Chen AA, Cheng J, Correll M, Duarte M, Fan C, Feltkamp MC, Ficarro SB, Franchi R, Garg BK, Gulbahce N, Hao T, Holthaus AM, James R, Korkhin A, Litovchick L, Mar JC, Pak TR, Rabello S, Rubio R, Shen Y, Singh S, Spangle JM, Tasan M, Wanamaker S, Webber JT, Roecklein-Canfield J, Johannsen E, Barabasi AL, Beroukhim R, Kieff E, Cusick ME, Hill DE, Munger K, Marto JA, Quackenbush J, Roth FP, DeCaprio JA, Vidal M. 2012. Interpreting cancer genomes using systematic host network perturbations by tumour virus proteins. Nature 487:491–495 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Kent WJ. 2002. BLAT–the BLAST-like alignment tool. Genome Res. 12:656–664 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Liang C, Marsit CJ, McClean MD, Nelson HH, Christensen BC, Haddad RI, Clark JR, Wein RO, Grillone GA, Houseman EA, Halec G, Waterboer T, Pawlita M, Krane JF, Kelsey KT. 2012. Biomarkers of HPV in head and neck squamous cell carcinoma. Cancer Res. 72:5004–5013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Malik-Soni N, Frappier L. 2012. Proteomic profiling of EBNA1-host protein interactions in latent and lytic Epstein-Barr virus infections. J. Virol. 86:6999–7002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Furler RL, Uittenbogaart CH. 2012. GLI2 regulates TGF-beta1 in human CD4(+) T cells: implications in cancer and HIV pathogenesis. PLoS One 7:e40874. 10.1371/journal.pone.0040874 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Swartling FJ, Savov V, Persson AI, Chen J, Hackett CS, Northcott PA, Grimmer MR, Lau J, Chesler L, Perry A, Phillips JJ, Taylor MD, Weiss WA. 2012. Distinct neural stem cell populations give rise to disparate brain tumors in response to N-MYC. Cancer Cell 21:601–613 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Cao Y, Wang L, Nandy D, Zhang Y, Basu A, Radisky D, Mukhopadhyay D. 2008. Neuropilin-1 upholds dedifferentiation and propagation phenotypes of renal cell carcinoma cells by activating Akt and sonic hedgehog axes. Cancer Res. 68:8667–8672 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Kim Y, Yoon JW, Xiao X, Dean NM, Monia BP, Marcusson EG. 2007. Selective down-regulation of glioma-associated oncogene 2 inhibits the proliferation of hepatocellular carcinoma cells. Cancer Res. 67:3583–3593 [DOI] [PubMed] [Google Scholar]
- 34. Wang Y, Han C, Lu L, Magliato S, Wu T. 2013. Hedgehog signaling pathway regulates autophagy in human hepatocellular carcinoma cells. Hepatology [Epub ahead of print.] 10.1002/hep.26394 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Saigo K, Yoshida K, Ikeda R, Sakamoto Y, Murakami Y, Urashima T, Asano T, Kenmochi T, Inoue I. 2008. Integration of hepatitis B virus DNA into the myeloid/lymphoid or mixed-lineage leukemia (MLL4) gene and rearrangements of MLL4 in human hepatocellular carcinoma. Hum. Mutat. 29:703–708 [DOI] [PubMed] [Google Scholar]
- 36. Skalsky RL, Cullen BR. 2010. Viruses, microRNAs, and host interactions. Annu. Rev. Microbiol. 64:123–141 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Wang Y, Jiang L, Ji X, Yang B, Zhang Y, Fu XD. 2013. Hepatitis B viral RNA directly mediates down-regulation of the tumor suppressor microRNA miR-15a/miR-16-1 in hepatocytes. J. Biol. Chem [Epub ahead of print.] 10.1074/jbc.M113.458158 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Bexfield N, Kellam P. 2011. Metagenomics and the molecular identification of novel viruses. Vet. J. 190:191–198 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Chen Y, Yao H, Thompson EJ, Tannir NM, Weinstein JN, Su X. 2013. VirusSeq: software to identify viruses and their integration sites using next-generation sequencing of human cancer tissue. Bioinformatics 29:226–267 [DOI] [PMC free article] [PubMed] [Google Scholar]