Abstract
Klebsiella pneumoniae has been frequently associated with nosocomial infections. Efflux systems are ubiquitous transporters that also function in drug resistance. Genome analysis of K. pneumoniae strain NTUH-K2044 revealed the presence of ∼15 putative drug efflux systems. We discuss here for the first time the characterization of a putative SMR-type efflux pump, an ebrAB homolog (denoted here as kpnEF) with respect to Klebsiella physiology and the multidrug-resistant phenotype. Analysis of hypermucoviscosity revealed direct involvement of kpnEF in capsule synthesis. The ΔkpnEF mutant displayed higher sensitivity to hyperosmotic (∼2.8-fold) and high bile (∼4.0-fold) concentrations. Mutation in kpnEF resulted in increased susceptibility to cefepime, ceftriaxone, colistin, erythromycin, rifampin, tetracycline, and streptomycin; mutated strains changed from being resistant to being susceptible, and the resistance was restored upon complementation. The ΔkpnEF mutant displayed enhanced sensitivity toward structurally related compounds such as sodium dodecyl sulfate, deoxycholate, and dyes, including clinically relevant disinfectants such as benzalkonium chloride, chlorhexidine, and triclosan. The prevalence of kpnEF in clinical strains broadens the diversity of antibiotic resistance in K. pneumoniae. Experimental evidence of CpxR binding to the efflux pump promoter and quantification of its expression in a cpxAR mutant background demonstrated kpnEF to be a member of the Cpx regulon. This study helps to elucidate the unprecedented biological functions of the SMR-type efflux pump in Klebsiella spp.
INTRODUCTION
Bacterial multidrug resistance (MDR) is now a global health care problem, and today's armory of antibiotics is increasingly limited. Prominent mechanisms that render bacteria resistant include enzymatic modification of drugs, target site alteration, altered outer membrane permeability, and upregulated multidrug efflux pump activity (1). Antibiotic efflux pumps fall into the ATP-binding cassette, major facilitator super family (MFS), resistance/nodulation/cell division (RND), multidrug and toxic compound extrusion (MATE), and the small MDR (SMR) groups and utilize the energy of the proton motive force or ATP to export antibiotics from the cell (2–5).
Of all of these systems, the SMR proteins are characterized by their short amino acid length (100 to 150 amino acids) resulting in a four-transmembrane (TM)-stranded α-helical protein (6). The SMR protein family can be subdivided into three subclasses namely, small multidrug proteins (SMP), suppressor of groEL mutations, and paired small MDR (PSMR) subclasses (7). Escherichia coli EmrE, Pseudomonas aeruginosa EmrE, Serratia marcescens SsmE, and Acinetobacter baumannii AbeS are studied members from the SMP subclass (8–11). The members of the PSMR subclass require coexpression of two SMR homologues to confer host resistance to antimicrobial compounds and toxic metabolites (12). The experimentally characterized pairs MdtI/MdtJ, YkkC/YkkD, and EbrA/EbrB, belong to the PSMR subclass (13, 14). A study shows that 52% of currently sequenced genomes of bacteria and archaea contain at least one SMR homologue (15), and the functions of many still remain enigmatic.
Klebsiella pneumoniae is a member of the Enterobacteriaceae family, which accounts for the vast majority of hospital- and community-acquired urinary tract infections. It is also a frequent cause of nosocomial bloodstream infections and community-acquired pneumonia among alcoholics (16, 17). Previous studies suggest that serotype K1 K. pneumoniae strains are common pathogens for various type of infections, including pyogenic liver abscess, sometimes complicated by endophthalmitis or meningitis in Korea, Taiwan, China, Japan, Singapore, Vietnam, India, and other Asian countries (18, 19).
The 5.2-Mb genome of one such K. pneumoniae strain NTUH-K2044 from K1 serotype (encoding 4,992 proteins; GC content, 57.7%) is reported to harbor ∼15 open reading frames encoding putative efflux pumps from different families (accession number NC_012731) (20). To date, the following efflux systems have been described in K. pneumoniae: AcrAB and KexD, belonging to the RND family conferring resistance to a wide spectrum of antimicrobial agents (21, 22); KdeA, from the MATE family involved in chloramphenicol, norfloxacin, acriflavine, and ethidium bromide (EtBr) resistance (23); and KmrA, included within the MFS family, providing resistance to several antimicrobial compounds (24). Analysis of completed Klebsiella genomes reveals the presence of several additional novel putative efflux systems from other families (20); however, experimental proof of their role in antimicrobial resistance has never been documented. Previously, we demonstrated that cpxAR two-component signaling system modulates the expression of efflux pumps to mediate antimicrobial resistance in K. pneumoniae (25, 26).
The aim of the present study was to characterize the functions of SMR-type efflux pump (PSMR class, an ebrAB homolog) in Klebsiella physiology, stress response, antimicrobial resistance, and the unprecedented involvement of CpxR, a cell envelope stress response regulator, in its expression.
MATERIALS AND METHODS
Bacterial strains, plasmids, and media.
K. pneumoniae NTUH-K2044 (a strain that resulted in pyogenic liver abscess in a 66-year-old patient) and MGH78578 were kindly provided by Jin Town Wang of the National Taiwan University Hospital, Taipei, Taiwan (27). K. pneumoniae clinical strains K2, K3, K7148, K513, K405, K823, K828, K812, K814, and K140 were kindly provided by Soorya multi-speciality tertiary hospital in Chennai. Escherichia coli SM10-λpir and pUT-Km were used to create isogenic mutants. E. coli KAM32, a highly susceptible strain that lacks major multidrug efflux pumps (ΔacrAB and ΔydhE mutants) was used as a host for heterologous studies (kindly provided by Tomofusa Tsuchiya). Bacterial cultures were grown in Luria-Bertani (LB) broth or on LB agar (Difco/Becton Dickinson, Sparks, MD) at 37°C with constant shaking (220 rpm) and stored at −80°C before use. Genomic DNA, plasmid, restriction digestion, DNA elution (Qiagen), ligation, transformation, conjugation, Southern hybridization, and DNA sequencing (Applied Biosystems) were performed either using standard procedures or as described previously (25, 26). Primers used in the present study were custom synthesized (Eurofins MWG Operons, Germany).
Cloning of kpnEF in E. coli KAM32 for heterologous studies.
The putative ebrAB homolog, KP1_2583/KP1_2584 (designated kpnEF), is located from nucleotide 2483890 to nucleotide 2484568 (kpnE, 363 bp, 120 amino acids [aa], 12.8 kDa; kpnF, 330 bp, 109 aa, 11.67 kDa) in the genome sequence of K. pneumoniae NTUH-K2044 (20). The putative efflux genes kpnE, kpnF, and the entire SMR operon were amplified by a standard PCR protocol using the primer pairs kpnE-F/kpnE-R, kpnF-F/kpnF-R, and kpnE-F/kpnF-R (Table 1), respectively, and cloned into the EcoRI and PstI (New England BioLabs, MA) sites of pUC18. The resulting recombinant plasmid pkpnE, pkpnF, and pkpnEF were transformed into E. coli KAM32 for functional characterization.
Table 1.
Primers used in this study
| Primer | Primer sequence (5′–3′) | Source or reference |
|---|---|---|
| ΔkpnEF-F | GCCCTGTCATATATTTTTCTTGCCTTTGCG | This study |
| ΔkpnEF-R | GCCGCCAGGGACAGGATGCCGTAGATCTTG | This study |
| kpnE-F | GCATCATATGTTTTATTGGATTTTATTAGCTTTA | This study |
| kpnE-R | TTCAGGATCCTCAAACTGCTGCATGGGCCACCTCC | This study |
| kpnF-F | GATACATATGCAGCAGTTTGAGTGGATCCACGCCG | This study |
| kpnF-R | TTCAGAATTCTCAGGCGAGTTTAATTAACACCATG | This study |
| FL kpnEF-F | TACTGAATTCTCCCGACCTCAAACTTCCTTGCCTA | This study |
| FL kpnEF-R | TTCAGAATTCTCAGGCGAGTTTAATTAACACCATG | This study |
| promkpnEF-F | TTAGCCATAGGATCCCGACCTCAAACTTCC | This study |
| promkpnEF-R | CCATTTCATAGACAAGGTGCCGGTAATTTCAGC | This study |
| RTkpnE-F | ATTGCTGAAATTACCGGCAC | This study |
| RTkpnE-R | AAATACCGATCCCTTCCCAC | This study |
| RTrpoBNT | GCGGTTGGTCGTATGAAGTT | This study |
| RTrpoBCT | TGGCGTTGATCATATCCTGA | This study |
| Kprt3706-F | TCAGTCATTACGGCAGAAGA | 47 |
| Kprt3706-R | AGATATAAGGGCATCAACAGATT | 47 |
| Kprt3709-F | CGATGCGTGAGTTTCTACA | 47 |
| Kprt3709-R | TGATATGGTTTCGGCTAATTCC | 47 |
| Kprt3712-F | AGTCATCTTCAGCATATTTCTCT | 47 |
| Kprt3712-R | CAACAATAGCACCTTTCCCT | 47 |
| MagAF1 | GGTGCTCTTTACATCATTGC | 48 |
| MagAR1 | GCAATGGCCATTTGCGTTAG | 48 |
| K2wzy-F1 | GACCCGATATTCATACTTGACAGAG | 48 |
| K2wzy-R1 | CCTGAAGTAAAATCGTAAATAGATGGC | 48 |
| K5wzxF360 | TGGTAGTGATGCTCGCGA | 48 |
| K5wzxR639 | CCTGAACCCACCCCAATC | 48 |
| wzxK54F | CATTAGCTCAGTGGTTGGCT | 48 |
| wzxK54R | GCTTGACAAACACCATAGCAG | 48 |
| wzyK57F | CTCAGGGCTAGAAGTGTCAT | 48 |
| wzyK57R | CACTAACCCAGAAAGTCGAG | 48 |
| wzyK20F | CGGTGCTACAGTGCATCATT | 48 |
| wzyK20R | GTTATACGATGCTCAGTCGC | 48 |
| rmpAF | ACTGGGCTACCTCTGCTTCA | 48 |
| rmpAR | CTTGCATGAGCCATCTTTCA | 48 |
Construction of kpnEF isogenic and complemented mutant in K. pneumoniae.
A 400-bp internal fragment was amplified by PCR using the ΔkpnEF-F and ΔkpnEF-R primers, cloned in EcoRI-digested pUT-Km, and transformed into E. coli SM10-λpir. The recombinant plasmid pUT-kpnEF was mobilized into recipient K. pneumoniae NTUH-K2044 from the donor E. coli SM10-λpir as described previously to construct the isogenic ΔkpnEF mutant (25, 26). The intact kpnEF operon, along with its promoter, was amplified with the FL kpnEF-F and FL kpnEF-R primers and cloned into pCRIITOPO-CAT. The resulting construct was electroporated into the ΔkpnEF mutant and selected on LB agar plates supplemented with 50 μg of kanamycin/ml and 100 μg of chloramphenicol/ml to get the transcomplemented ΔkpnEFΩkpnEF strain. We used a similar strategy to generate K3ΔkpnEF and K3ΔkpnEFΩkpnEF mutant strains from K. pneumoniae K3.
Bacterial growth curves.
The growth kinetics of the wild-type (WT; control strain NTUH-K2044) and ΔkpnEF and ΔkpnEFΩkpnEF mutant strains were monitored in 96-well clear flat-bottom polystyrene microplates (Costar; Corning) with LB media at different pH levels (3.0, 5.0, 6.0, 7.0, 7.5, 8.0, and 12.0). The optical densities at 600 nm (OD600) were measured for 10 h at 37°C with shaking using a Synergy H1 hybrid microplate reader (BioTek Instruments, Inc., Winooski, VT) and automatically recorded for each well after every 15 min. The experiment was performed with freshly autoclaved medium in triplicates at least three independent times. A growth inactivation assay was performed to assess the impact on drug efflux capacity, as described previously with slight modifications (25, 26). WT and ΔkpnEF cultures at mid-log phase (OD600 = 0.2) were inoculated into LB broth containing antibiotics such as chloramphenicol, erythromycin, or tetracycline (0.05 μg/ml) in independent experiments either alone or with efflux pump inhibitor carbonyl cyanide 3-chlorophenylhydrazone (CCCP; Sigma, St. Louis, MO) at 10 μg/ml. The growth profiles for WT and ΔkpnEF strains at 37°C were then analyzed by measuring the absorbance at OD600 periodically in a Synergy H1 hybrid microplate reader. Efflux pump inhibitors had no intrinsic antibacterial activity against the WT strain at the concentrations used in these experiments. Each of these experiments was performed more than three times.
Hypermucoviscosity, motility, and crystal violet binding assay.
The motility assays were performed as reported previously (28); LB medium-grown K. pneumoniae cultures (OD600 = 1.0) were inoculated with a toothpick onto LB plates with 0.25, 0.45, and 0.7% agar, followed by incubation for 14 h at 37°C. In this growth medium, bacteria can swim through the soft agar and produce a halo. The diameter of the halo is a measure of the ability to swim. The string and precipitation test for hypermucoviscosity was performed as described earlier (25, 26).
A crystal violet binding assay was performed as described previously (28). Bacterial cultures were grown in LB medium and diluted to an OD600 of 0.2; 150 μl of each strain was then placed into glass tubes. The tubes were incubated at 37°C with shaking for 24 h. After incubation, the samples were washed three times with phosphate-buffered saline to remove planktonic growth. The remaining biofilm was fixed with methanol for 15 min. Once the methanol was removed and the tubes were dried, the biofilms were stained with 1% crystal violet for 5 min. The stain was removed by washing with water, and the plates were dried. Biofilm thickness was measured by adding 33% glacial acetic acid and taking a reading at OD580 using an automated plate reader.
Stress challenge assays.
The WT, ΔkpnEF, and ΔkpnEFΩkpnEF strains were exposed to different stress challenges such as bile, sodium chloride (osmotic, NaCl), heat, sodium dodecyl sulfate (SDS), deoxycholate, EtBr, acriflavine, benzalkonium chloride, chlorhexidine, and triclosan, and survival was quantified as described previously (25). The impact of an oxidative stress-inducing agent (H2O2) on the survival of WT, ΔkpnEF, and ΔkpnEFΩkpnEF strains and the growth kinetics in the presence of different nitrosative stress-inducing agents (sodium nitroprusside [SNP] and acidified nitrite) were assessed as described previously (25, 26), with a modification where absorbance was measured a in a Synergy H1 hybrid microplate reader.
Antibiotic susceptibility testing and fluorimetric efflux studies.
Antibiotic susceptibility and MIC levels were examined using commercial discs and E-strips (Hi Media, Mumbai, India) as described previously, and the data were analyzed according to the interpretation criteria recommended by the Clinical and Laboratory Standards Institute (31). Where indicated, the MICs were determined by the agar double-dilution method (31).
The accumulations of ciprofloxacin and EtBr were evaluated as described previously (32). Overnight cultures of WT and mutant cells were diluted to an OD600 of 0.03 in 100 ml of LB broth and grown at 37°C to an OD600 of 1.0. Then, 10 ml of cells was pelleted and resuspended in 5 ml of LB medium. The cells were loaded with EtBr and incubated at 37°C for 15 min. After 5 min in a fluorimeter, cells loaded with EtBr were energized by the addition of glucose, and the efflux of EtBr in mutant and WT cells was monitored continuously by measuring the fluorescence emission at 600 nm upon excitation at 530 nm. The fluorescence was measured by using spectrofluorimeter (Hitachi). After 10 min, 100 μM CCCP was added as indicated to abolish active transport, and fluorescence emission was monitored further. For ciprofloxacin, excitation was set at 275 nm and emission was set at 440 nm. Each data point represents the mean plus the standard deviation for three independent experiments.
RNA isolation and real-time RT-PCR.
Total RNA was extracted from the log-phase cultures using an RNeasy minikit (Qiagen) according to the manufacturer's instructions. Total RNA was digested with DNase I to ensure the removal of contaminating genomic DNA prior to cDNA synthesis. Aliquots of 500 ng of DNase I-treated total RNA served as a template for cDNA synthesis using Superscript III reverse transcriptase (Invitrogen). The quantitative PCRs for different genes were performed using gene specific primers (Table 1). Gene expression levels were monitored by using real-time reverse transcription-PCR (RT-PCR) with a Maxima SYBR green qPCR master mix (Fermentas) in an iCycler thermal cycler (Bio-Rad), and a melting-curve analysis was carried out to confirm amplification of a single product. Total RNA was isolated from at least two separately grown replicate cultures. All real-time RT-PCR experiments were performed more than three times, with rpoB as an internal control.
EMSAs.
In our previous study, the response regulator cpxR was cloned and expressed in pET28C (25). The ability of CpxR to bind the kpnEF promoter was examined by an electrophoretic mobility shift assay (EMSA). Using promkpnEF-F and promkpnEF-R primers, the promoter was amplified (Table 1). Briefly, end-labeled (using [γ-32P]ATP) promoter was incubated with increasing concentrations (in a range of 40 to 500 nM) of protein in binding buffer [10 mM Tris-HCl (pH 8.0), 2 mM EDTA, 0.5 mM dithiothreitol [DTT], 50 mM NaCl, 10% glycerol, 1 μg of poly(dI-dC)]. The complexes were run on 5% polyacrylamide gel (PAGE) for 2 h. The gel was then dried and exposed to a phosphor screen for image analysis. To confirm that the interaction was specific, competition experiments with bovine serum albumin (BSA) as a negative control (noncompetitive) and with 10-fold excess of cold promoter (competitive) were also performed.
Bioinformatic and statistical analysis.
Homology searches, similarity and identity analyses, and conserved domain architecture analyses were performed using the National Center for Biotechnology Information (NCBI) Internet server, Simple Modular Architecture Research Tool (SMART), and NCBI conserved domain search. All data are presented as means ± the standard errors of the mean. Plotting and calculation of the standard deviation was performed in Microsoft Excel. Statistical analysis was performed on crude data by using a paired Student t test. P values of <0.05 were considered significant.
RESULTS
Sequence analysis of the KpnEF SMR-type efflux pump.
The protein sequence deduced from the 678-bp DNA fragment obtained from K. pneumoniae NTUH-K2044 shares 83% identity with the EbrAB efflux system in E. coli, 81% identity with the EbrAB efflux system in Shigella dysenteriae, and 77% identity with the EbrAB efflux system in Salmonella enterica serovar Typhimurium. A homology search in the NCBI database revealed that KpnE and KpnF demonstrated 35 and 30% and 45 and 43% sequence identities with E. coli EmrE and Staphylococcus aureus Smr, respectively. In addition, significant sequence similarities were detected between E. coli, Proteus vulgaris, and Citrobacter freundii KpnEF and SugE. Strikingly, the deduced amino acid sequence of KpnE was 39% identical to that of KpnF. The KpnE has four hydrophobic, presumably transmembrane regions spanning from aa 1 to 22 (22 aa in length), aa 32 to 51 (20 aa in length), aa 57 to 79 (23 aa in length), and aa 83 to 105 (23 aa in length). The KpnF also has four transmembrane regions spanning from aa 7 to 29 (23 aa in length), aa 36 to 58 (23 aa in length), aa 63 to 85 (23 aa in length), and aa 88 to 109 (22 aa in length). According to the SOSUI software, the average hydrophobicities were 1.240834 and 1.055963 for kpnE and kpnF, respectively. Overall, KpnE and KpnF are members of the PSMR family of drug transporters.
Characterization of membrane transporter KpnEF in E. coli KAM32.
The MICs for kpnEF-harboring cells (E. coli KAM32/pkpnEF) were higher—neomycin (1.5-fold), kanamycin (3-fold), erythromycin (1.5-fold), trimethoprim (1.5-fold), amikacin (1.5-fold), nalidixic acid (4-fold), SDS (2-fold), bile (2-fold), chlorhexidine (2-fold), benzalkonium chloride (2-fold), and methyl viologen (2-fold)—compared to those for E. coli KAM32/pUC18 (data not shown). It is significant that pkpnE (E. coli KAM32/pkpnE) or pkpnF (E. coli KAM32/pkpnF) alone showed no change in their susceptibility profiles compared to E. coli KAM32/pUC18.
Contributions of the KpnEF SMR efflux pump to K. pneumoniae cellular physiology. (i) Impact on morphology and hypermucoviscosity.
Interestingly, the WT strain produced bigger (3.0 ± 0.5 mm) and heavily mucoid colonies, whereas the ΔkpnEF strain colonies were smaller (1.15 ± 0.26 mm) and nonmucoid, indicating a direct loss in capsular polysaccharide (CPS) production. Further, the average string lengths for WT and ΔkpnEF mutant strains were 5.0 ± 0.5 cm and 1.2 ± 0.02 cm, respectively. Moreover, upon precipitating the cultures at 4,000 rpm for 10 min, the ΔkpnEF strain formed a compact pellet, in contrast to the WT strain (see Fig. S1 in the supplemental material). The overall results presented here indicate the possible involvement of a kpnEF-like family protein in capsule synthesis, which may contribute to virulence in K. pneumoniae.
(ii) Effect on general growth characteristics.
To evaluate the involvement of kpnEF in cellular growth, the growth profiles of WT, ΔkpnEF, and ΔkpnEFΩkpnEF strains were continuously monitored in LB broth at pH 3.0, 5.0, 6.0, 7.0, 7.5, 8.0, and 12.0. It is interesting that the kpnEF mutant exhibited 1.5-fold (±0.025; P = 0.00618), 1.29-fold (±0.063; P = 0.000987), 1.19-fold (±0.054; P = 0.00660), 1.032-fold (±0.041; P = 0.0912), and 1.5-fold (±0.0021; P = 0.00131) reduced growth compared to the WT strain after 4 h in LB medium at pH 5.0, 6.0, 7.0, 7.5, and 8.0, respectively (Fig. 1). The other tested pH levels—pH 3.0 and 12.0—were toxic to both cultures. These results demonstrate that kpnEF influences the growth of K. pneumoniae.
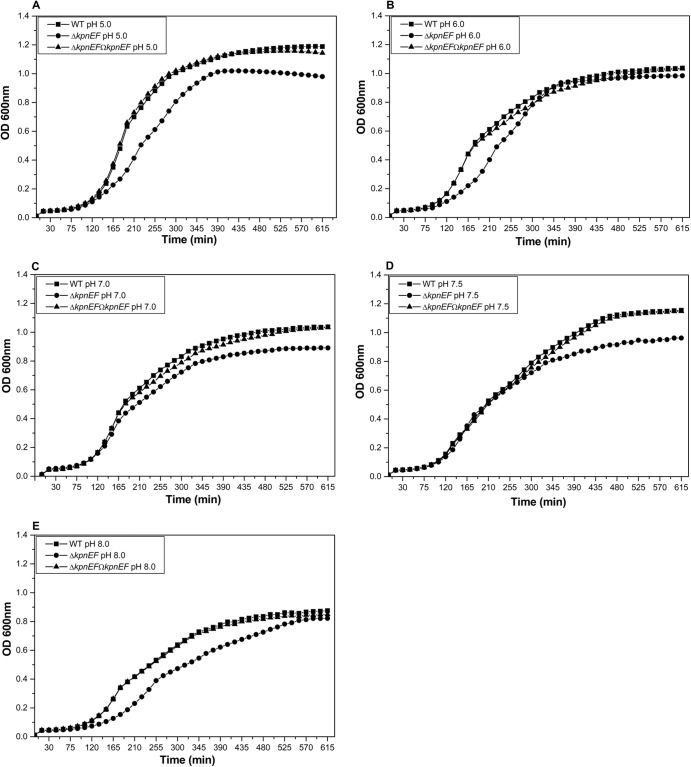
Fig 1.
Effect of kpnEF mutations on general growth characteristics. The effect on bacterial growth was monitored in WT, ΔkpnEF, and ΔkpnEFΩkpnEF strains in LB medium at pH 3.0, 5.0, 6.0, 7.0, 7.5, 8.0, and 12.0. The patterns for the representative pH values 5.0 (A), 6.0 (B), 7.0 (C), 7.5 (D), and 8.0 (E) are shown. After 10 h of growth, the kpnEF mutant exhibited 1.2-fold, 1.06-fold, 1.16-fold, 1.2-fold, and 1.06-fold stunted growth in LB medium at pH 5.0, 6.0, 7.0, 7.5, and 8.0, respectively, which was restored upon complementation. The data presented reflect triplicate determinations.
(iii) Alterations in motility behavior and biofilm formation.
When the cultures were grown in a gradient agar for 14 h, WT cells formed concentric motility rings around the points of inoculation; the migration distances were 37 mm (0.25%), 30 mm (0.45%), and 26 mm (0.7%), indicating robust motility. In stark contrast, the kpnEF mutant cells had impaired motility, and the migration distances were 13 mm (0.25%), 18 mm (0.45%), and 19 mm (0.7%), indicating a loss in motility (Fig. 2A). We found that the in vitro biofilm-forming ability of the ΔkpnEF mutant was ∼1.155-fold less compared to the WT strain (Fig. 2B). The evidence obtained indicated that the kpnEF mutant influences both motility and capsule synthesis, which are known to help K. pneumoniae to establish colonization during the development of a pyogenic liver abscess (17).
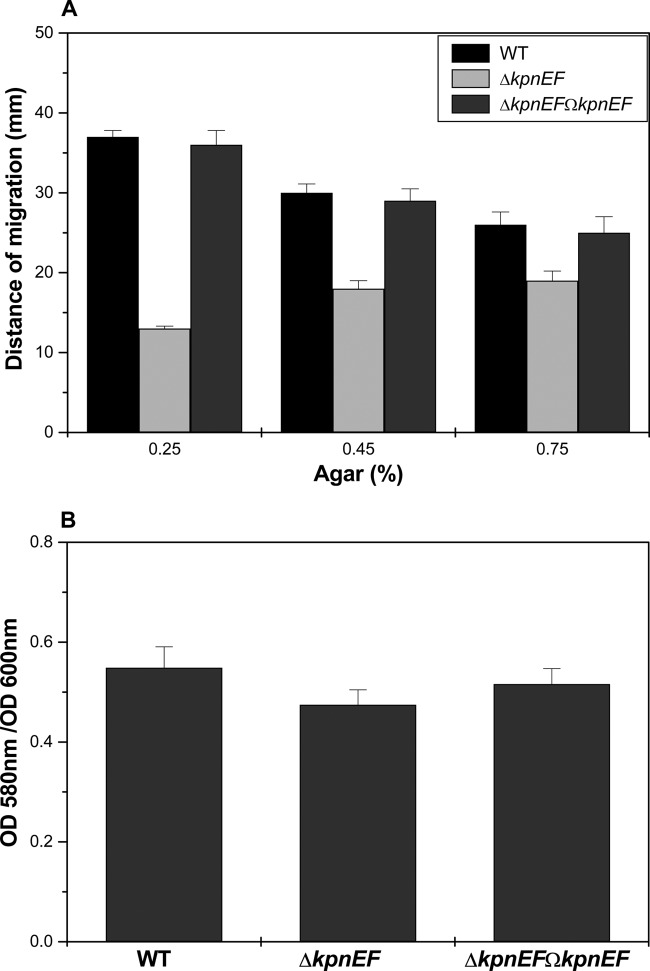
Fig 2.
Alterations in motility behavior and biofilm formation. (A) The average diameters of swimming halos from three different experiments are plotted, along with the standard deviations. The P values for the differences between WT, ΔkpnEF, and ΔkpnEFΩkpnEF strains were all <0.01. The results are given in millimeters, considering K. pneumoniae NTUH-K2044 as the WT. (B) The K. pneumoniae ΔkpnEF mutant is defective in biofilm formation on glass tubes. After 24 h of incubation, biofilm formation was measured by staining with 0.5% crystal violet, and the OD580 was determined for the WT strain and for the ΔkpnEF mutant. The data are means of measurements performed three times. Significant differences with respect to NTUH-K2044 are indicated by an asterisk (*, P < 0.01).
(iv) Effect on gastrointestinal stress response.
The ability of the WT to grow in the presence of different concentrations of bile (the physiological concentration is 0.2 to 2% [33]), e.g., in 0.5% bile it was 1.4-fold (±0.025), in 0.75% bile it was 1.7-fold (±0.035), in 1% bile it was 2.5-fold (±0.024), and in 2% bile it was 4.0-fold (±0.05) greater compared to the ΔkpnEF mutant, whereas the transcomplemented ΔkpnEFΩkpnEF strain restored the ability to tolerate the stress (P = 0.02191) (Fig. 3A). The ability of the WT to grow in the presence of NaCl (physiological concentration, 150 mM [29, 30]) in 0.25 M NaCl was ∼1.629-fold (±0.025) greater, in 0.5 M NaCl it was ∼1.32-fold (±0.012) greater, and in 0.75 M NaCl it was ∼2.75-fold (±0.23) greater compared to the ΔkpnEF mutant, regardless of the inoculum size (P = 0.01944) (Fig. 3B). The temperature-dependent assay showed that the kpnEF mutant displayed 33% reduced survival than the WT at 42°C, clearly demonstrating the response of the K. pneumoniae kpnEF mutant in temperature stress (P = 0.31157) (Fig. 3C). The results presented here clearly indicate the contributory role of kpnEF in varied stress tolerance in K. pneumoniae.
Fig 3.
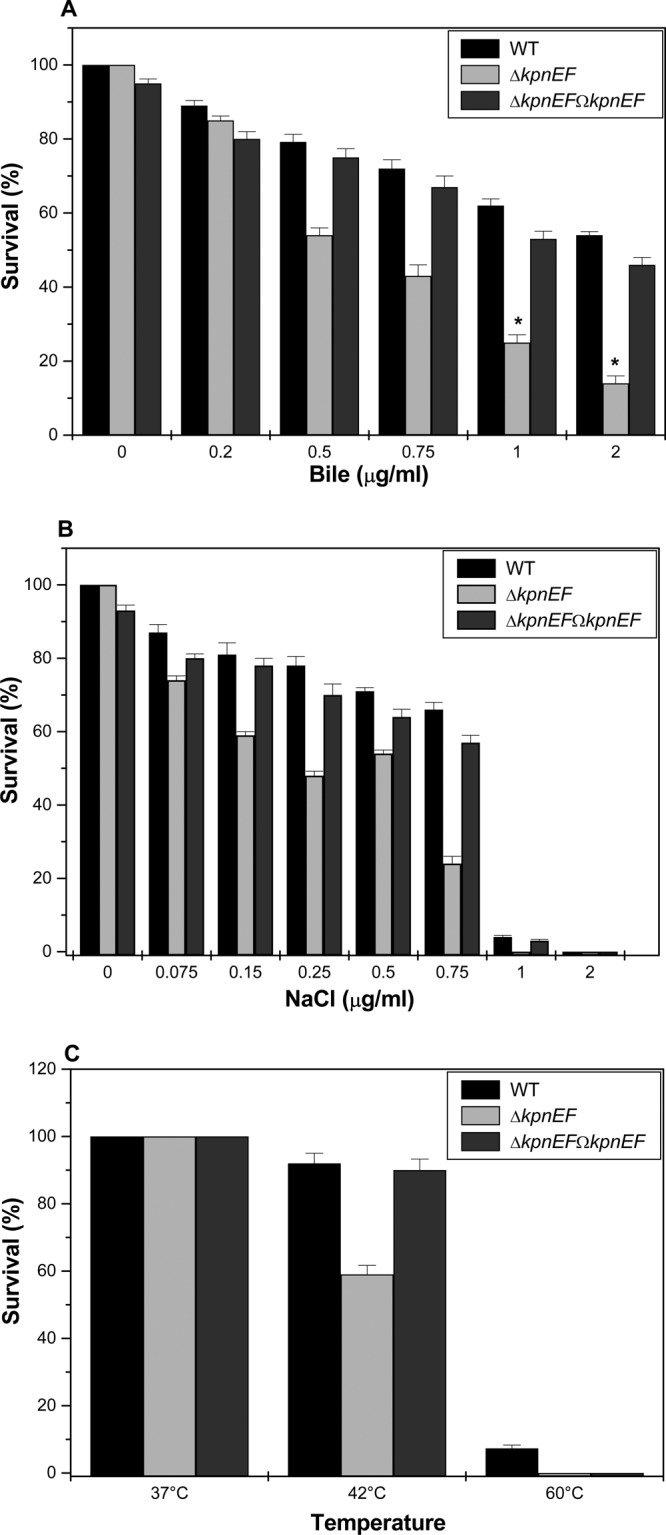
Impact on gastrointestinal stress response. (A) Sensitivities of WT, ΔkpnEF, and ΔkpnEFΩkpnEF strains to different concentrations (0.2, 0.5, 0.75, 1.0, and 2.0%) of bile. The survival ability of the WT strain in 0.5% bile was 1.4-fold (±0.025) greater, in 0.75% bile it was 1.7-fold (±0.035) greater, in 1% bile it was 2.5-fold (±0.024) greater, and in 2% bile it was 4.0-fold (±0.05) greater compared to the ΔkpnEF strain. The complemented ΔkpnEFΩkpnEF strain displayed the same phenotype as the WT. (B) Sensitivities of WT, ΔkpnEF, and ΔkpnEFΩkpnEF strains to different concentrations (0.075, 0.15, 0.25, 0.5, 0.75, 1.0, and 2.0 M) of NaCl. The survival ability of the WT strain in 0.25 M NaCl was ∼1.629-fold (±0.025) greater, in 0.5 M NaCl it was ∼1.32- fold(±0.012) greater, and in 0.75 M NaCl it was ∼2.75-fold (±0.23) greater compared to the ΔkpnEF strain. (C) Survival of the WT, ΔkpnEF, and ΔkpnEFΩkpnEF strains at 37, 42, and 60°C after 1 h of incubation. The percentage of resistance to different stress agents was calculated compared to the numbers of viable cells in LB medium alone. Significant differences with respect to the WT strain are indicated by an asterisk.
(v) Role in oxidative stress tolerance.
A hydrogen peroxide (H2O2) disc diffusion assay showed that the kpnEF mutant exhibited 1.4-fold greater sensitivity to 0.78947 mM H2O2 (inhibition zone = 55 ± 2.0 mm) than the WT (inhibition zone = 39 ± 1.2 mm) (P = 0.512) (Fig. 4A). Upon performing an oxidative survival assay, only 36.9 and 5.39% of the ΔkpnEF cells survived upon treatment with 0.07894 and 0.7894 mM H2O2, respectively, compared to the 89 and 69.76% survival observed in WT cells (P = 0.14665). The survival of WT and ΔkpnEF strains was negligible in other tested concentrations (1.5788, 2.3682, and 3.1576 mM) of H2O2 (Fig. 4B). The results imply that kpnEF had a role in oxidative stress tolerance in K. pneumoniae.
Fig 4.
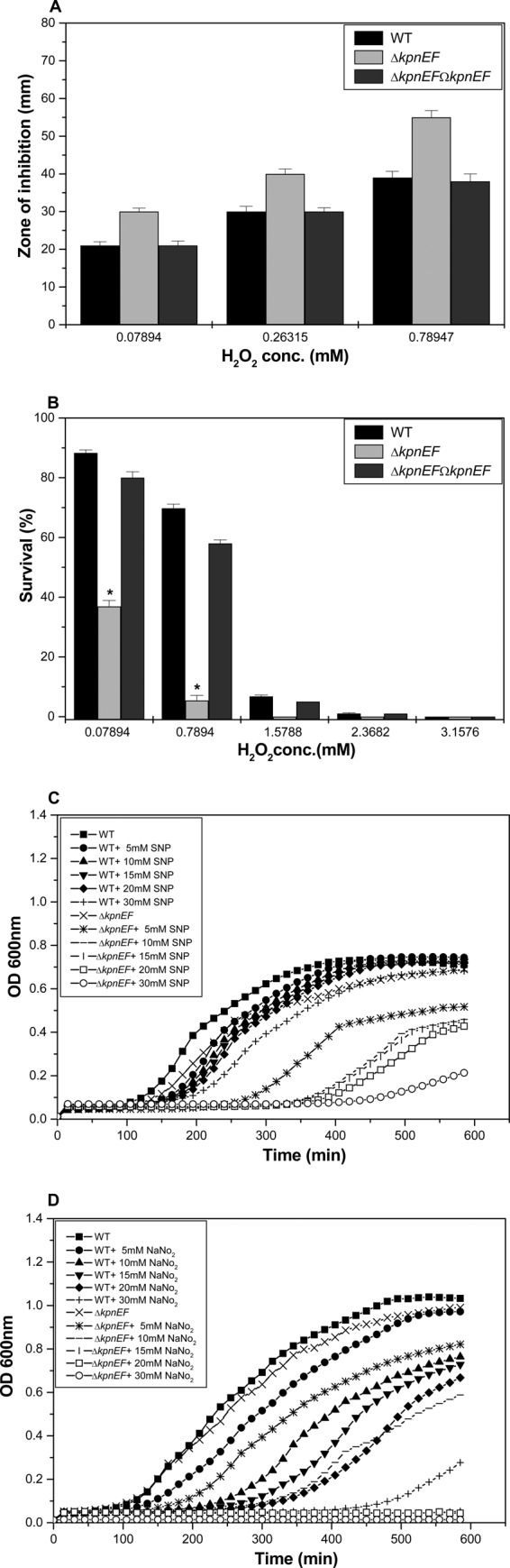
Role in oxidative and nitrosative stress tolerance. (A) Oxidative stress response of WT, ΔkpnEF, and ΔkpnEFΩkpnEF strains. The survival abilities of WT and ΔkpnEF strains to combat different levels of H2O2 stress (0.07894, 0.26315, and 0.78947 mM) were measured by disc diffusion assay. The kpnEF mutant displayed greater sensitivity to 0.78947 mM H2O2 (inhibition zone = 55 ± 2.0 mm) than the WT (39 ± 1.2 mm). The data presented reflect triplicate determinations. (B) Survival of WT, ΔkpnEF, and ΔkpnEFΩkpnEF strains upon exposure to oxidative stress at 0.07894, 0.7894, 1.5788, 2.3682, and 3.1576 mM H2O2. After 1 h of treatment with 0.07894 mM H2O2, only 36.9% of the ΔkpnEF cells survived compared to 89% of the WT cells. The differences between the mutant and its parental WT strain are statistically significant (P < 0.05) for all H2O2 concentrations; the standard errors of the mean from three independent assays are also shown. Significant differences with respect to the WT strain are indicated by an asterisk. (C) Effect of different concentrations of SNP on the growth profiles of the WT and ΔkpnEF strains. The growth kinetics of the ΔkpnEF strain were ∼1.5-fold (P = 0.0039) lower, 1.6-fold (P = 0.0055) lower, 1.62-fold (P = 0.0076) lower, 1.7-fold (P = 0.0095) lower, or 3.2-fold (P = 0.0196) lower than for the WT cells in the presence of 5, 10, 15, 20, or 30 mM concentrations of the NO donor, SNP. (D) Growth patterns of WT and ΔkpnEF strains in the presence of acidified nitrite. The growth kinetics for the ΔkpnEF strain were ∼1.18-fold (P = 0.000424) lower, 1.3-fold (P = 0.00659) lower, 13.75-fold (P = 0.0002) lower, 15.2-fold (P = 0.00013) lower, or 20-fold (P = 0.000202) lower than for WT cells in the presence of 5, 10, 15, 20, or 30 mM concentrations of the acidified nitrite.
(iv) Involvement in nitrosative stress response.
In the presence of an NO donor, SNP at 30 mM, the growth kinetics of the ΔkpnEF mutant were >3.2-fold lower than those of the WT (P = 0.0196) (Fig. 4C). To further evaluate the potential of K. pneumoniae kpnEF against other reactive nitrogen species, we tested the tolerance of ΔkpnEF cells toward acidified nitrite. Protonated nitrite quickly degrades to generate numerous species of nitrogen oxides, e.g., nitric oxide (34). In the presence of 15 mM acidified nitrite at pH 6.0, the growth kinetics of the ΔkpnEF mutant were >13.75-fold lower than for the WT strain (P = 0.00212). In the presence of 30 mM NO donor at pH 6.0, the growth kinetics of the ΔkpnEF mutant were ∼20-fold lower than for the WT (P = 0.000202) (Fig. 4D). Overall, the results imply that kpnEF is involved in nitrosative stress tolerance in K. pneumoniae.
Contributions of the KpnEF efflux pump in K. pneumoniae antimicrobial resistance. (i) Drug susceptibilities and MIC.
Disc diffusion assay results for the ΔkpnEF mutant displayed decreased susceptibility toward cefepime, ceftriaxone, colistin, erythromycin, rifampin, tetracycline, and streptomycin (Fig. 5A). The precise MIC values for WT, ΔkpnEF, and ΔkpnEFΩkpnEF are shown in Table 2.
Fig 5.
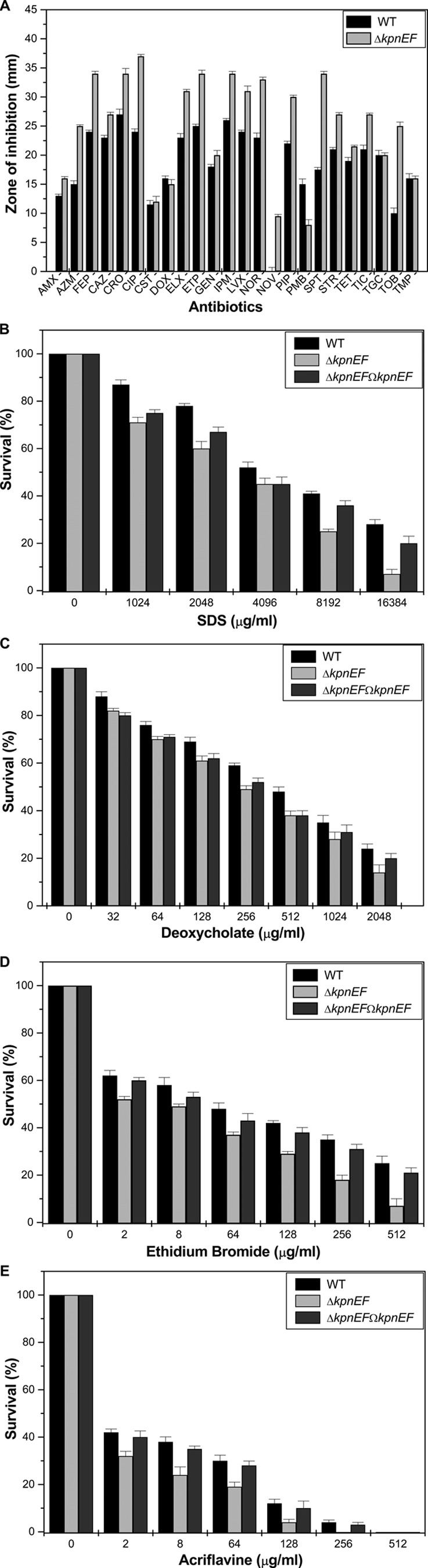
Contributions of the KpnEF efflux pump in K. pneumoniae antimicrobial resistance. (A) A Kirby-Bauer disc diffusion assay was performed with different antibiotics using commercial discs. Antibiotics, abbreviations, and concentrations: amoxicillin, AMX (30 μg/ml); azithromycin, AZM (15 μg/ml); cefepime, FEP (30 μg/ml); ceftazidime, CAZ (30 μg/ml); ceftriaxone, CRO (30 μg/ml); ciprofloxacin, CIP (5 μg/ml); colistin, CST (10 μg/ml); doxycycline, DOX (30 μg/ml); enrofloxacin, ELX (10 μg/ml); ertapenem, ETP (10 μg/ml); gentamicin, GEN (10 μg/ml); imipenem, IPM (10 μg/ml); levofloxacin, LVX (5 μg/ml); norfloxacin, NOR (10 μg/ml); novobiocin, NOV (30 μg/ml); piperacillin, PIP (100 μg/ml); polymyxin B, PMB (300 μg/ml); spectinomycin, SPT (100 μg/ml); streptomycin, STR (10 μg/ml); tetracycline, TET (30 μg/ml); ticarcillin, TIC (75 μg/ml); tigecycline, TGC (15 μg/ml); tobramycin, TOB (10 μg/ml); trimethoprim, TMP (5 μg/ml). The data for representative drugs are shown here. (B) Sensitivities of WT, ΔkpnEF, and ΔkpnEFΩkpnEF strains to different concentrations of SDS. The survival ability of the WT strain in SDS at 1,024 μg/ml was ∼1.22-fold greater, in SDS at 2,048 μg/ml it was ∼1.3-fold greater, in SDS at 4,096 μg/ml it was ∼1.15-fold greater, in SDS at 8,192 μg/ml it was ∼1.64-fold greater, and in SDS at 16,834 μg/ml it was 4-fold greater compared to the ΔkpnEF strain. (C) Susceptibilities of WT, ΔkpnEF, and ΔkpnEFΩkpnEF strains to different concentrations of deoxycholate. The survival ability of the WT strain in deoxycholate at 128 μg/ml was ∼1.13-fold greater, in deoxycholate at 256 μg/ml it was ∼1.2-fold greater, in deoxycholate at 512 μg/ml it was ∼1.26-fold greater, in deoxycholate at 1,024 μg/ml it was ∼1.25-fold greater, and in deoxycholate at 2,048 μg/ml it was 1.71-fold greater compared to the ΔkpnEF strain. (D) Sensitivities of WT, ΔkpnEF, and ΔkpnEFΩkpnEF strains to different concentrations of EtBr. The survival ability of the WT strain in EtBr at 8 μg/ml was ∼1.18-fold greater, in EtBr at 64 μg/ml it was ∼1.29-fold greater, in EtBr at 128 μg/ml it was ∼1.44-fold greater, in EtBr at 256 μg/ml it was ∼1.94-fold greater, and in EtBr at 512 μg/ml it was 3.57-fold greater compared to the ΔkpnEF strain. (E) Susceptibilities of WT, ΔkpnEF, and ΔkpnEFΩkpnEF strains to different concentrations of acriflavine. The survival ability of the WT strain in EtBr at 8 μg/ml was ∼1.58-fold greater, in EtBr at 64 μg/ml it was ∼1.57-fold greater, and in EtBr at 128 μg/ml it was ∼3-fold greater compared to the ΔkpnEF strain. The percent survival was calculated by comparison of the viable cells in the WT. The data are means of measurements made in triplicate and performed three times. *, Significant difference (P < 0.05, Student t test).
Table 2.
Determination of antibiotic MICs for WT, ΔkpnEF, and ΔkpnEFΩkpnEF strainsa
| Antibiotic | MIC (μg/ml) for the WT strain | MIC (μg/ml) for the ΔkpnEF strain | Fold change (WT/ΔkpnEF strain) | MIC (μg/ml) for the ΔkpnEFΩkpnEF strain |
|---|---|---|---|---|
| Cefepime | 2.048 | 1.024 | 2 | 2.048 |
| Ceftazidime | 0.256 | 0.064 | 4 | 0.256 |
| Chloramphenicol | 0.1 | 0.1 | 1 | 0.1 |
| Ciprofloxacin | <0.01 | <0.005 | 2 | <0.01 |
| Colistin | 0.01 | 0.005 | 2 | 0.01 |
| Erythromycin | >4 | 2 | 2 | >4 |
| Kanamycin | 1 | >30 | 30 | 1 |
| Norfloxacin | <0.001 | <0.001 | 1 | <0.001 |
| Polymyxin | 0.1 | 0.05 | 2 | 0.1 |
| Rifampin | 10 | 5 | 2 | 10 |
| Sparfloxacin | <0.01 | <0.01 | 1 | <0.01 |
| Streptomycin | 0.1 | 0.01 | 10 | 0.1 |
| Tetracycline | 5 | 2.5 | 2 | 5 |
| Tobramycin | 0.1 | 0.1 | 1 | 0.1 |
| Vancomycin | >4 | >4 | 1 | >4 |
E-strips were used to determine the precise MICs for different antibiotics, such as amikacin, ampicillin, cefepime, cefotaxime, ceftazidime, ceftriaxone, chloramphenicol, ciprofloxacin, colistin, erythromycin, kanamycin, nalidixic acid, norfloxacin, ofloxacin, polymyxin, rifampin, sparfloxacin, streptomycin, tetracycline, tobramycin, trimethoprim, and vancomycin, according to the CLSI guidelines (31). Representative antibiotics are shown here. The resistance to kanamycin in the ΔkpnEF strain is due to the insertion of a plasmid that carries the Kanr gene. The fold change indicates the ratio of MICs for the WT and ΔkpnEF strains.
(ii) Tolerance to structurally related compounds.
Upon exposing the cells to different concentrations of SDS, it was observed that the total CFU count of the WT at 16,384 μg/ml was 4-fold higher than that of the ΔkpnEF mutant (P = 0.01001) (Fig. 5B). The ability of the ΔkpnEF mutant to withstand deoxycholate at 2,048 μg/ml was reduced by 1.71-fold (P = 0.00054) (Fig. 5C). The ability of the ΔkpnEF mutant to withstand EtBr at 512 μg/ml (P = 0.00263) (Fig. 5D) and acriflavine at 128 μg/ml (P = 0.01789) (Fig. 5E) was reduced by 3.57- and 3.0-fold, respectively. Analysis indicates the wide substrate specificity for kpnEF in K. pneumoniae.
(iii) Susceptibility to hospital-based disinfectants.
In the present study, the susceptibilities of the WT, ΔkpnEF, and ΔkpnEFΩkpnEF strains to different concentrations of disinfectants were tested. Upon exposing the cells to different concentrations of benzalkonium chloride, we observed that the total CFU count for the WT at 25.6 μg/ml was 8.05-fold higher than that for the ΔkpnEF strain (P = 0.05154). When cells were exposed to 12.8 μg of chlorhexidine/ml, it was observed that the total CFU count of WT was 6.39-fold higher than that of the ΔkpnEF strain (P = 0.07508). When cells were exposed to 0.01 μg of triclosan/ml, it was observed that the total CFU count of WT was 3.164-fold higher than that of the ΔkpnEF strain (P = 0.06753) (Fig. 6). Decisively, the results suggest the broad-spectrum antimicrobial resistance property of KpnEF, an SMR-type efflux pump for the first time in K. pneumoniae.
Fig 6.
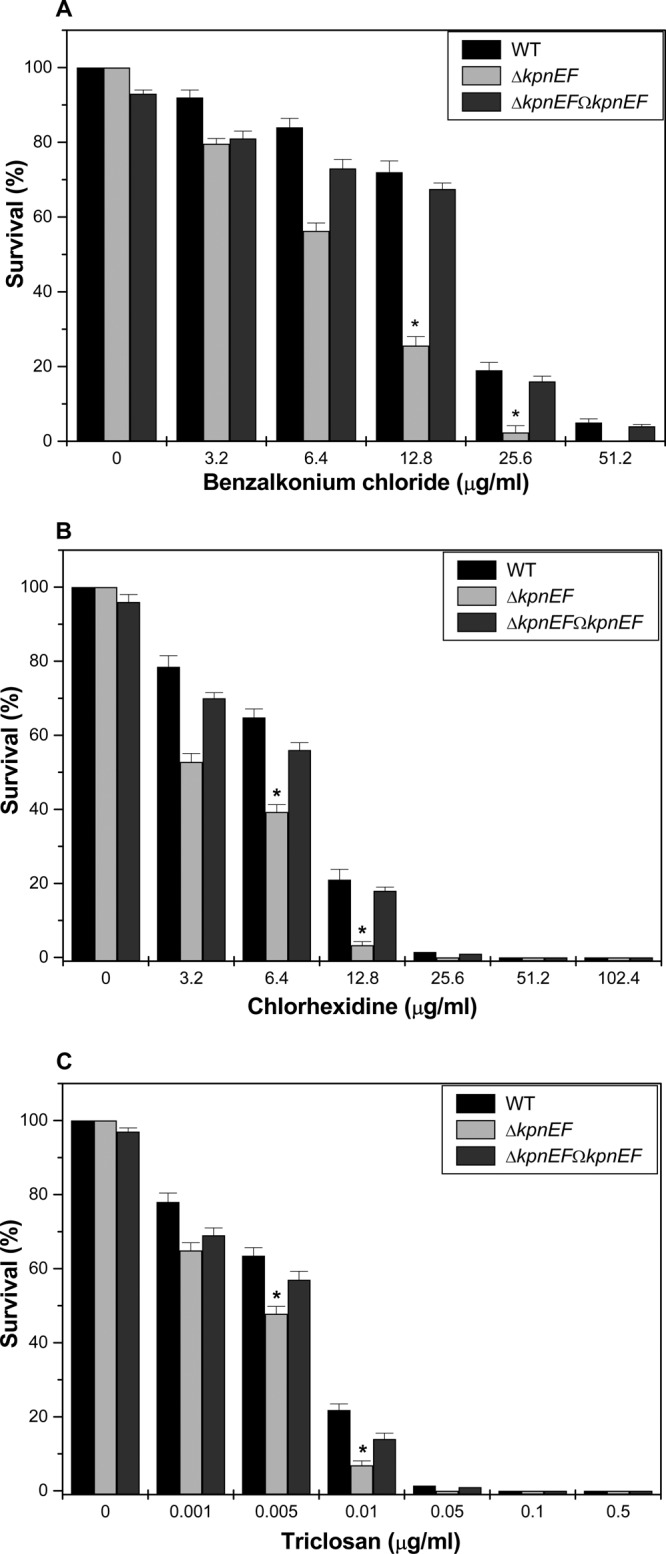
Contributions of kpnEF in disinfectant resistance. (A) Sensitivities of WT, ΔkpnEF, and ΔkpnEFΩkpnEF strains to different concentrations of benzalkonium chloride. The survival ability of the WT strain in benzalkonium chloride at 3.2 μg/ml was ∼1.15-fold greater, in benzalkonium chloride at 6.4 μg/ml it was ∼1.49-fold greater, in benzalkonium chloride at 12.8 μg/ml it was ∼2.81-fold greater, and in benzalkonium chloride at 25.6 μg/ml it was ∼8.05-fold greater compared to the ΔkpnEF mutant. (B) Susceptibilities of WT, ΔkpnEF, and ΔkpnEFΩkpnEF strains to different concentrations of chlorhexidine. The survival ability of the WT strain in chlorhexidine at 3.2 μg/ml was ∼1.48-fold greater, in chlorhexidine at 6.4 μg/ml it was ∼1.64-fold, and in chlorhexidine at 12.8μg/ml it was ∼6.39-fold greater compared to the ΔkpnEF strain. (C) Sensitivities of WT, ΔkpnEF, and ΔkpnEFΩkpnEF strains to different concentrations of triclosan. The survival ability of the WT strain in triclosan at 0.001 μg/ml was ∼1.20-fold greater, in triclosan at 0.005 μg/ml it was ∼1.32-fold greater, and in triclosan at 0.01 μg/ml it was ∼3.16-fold greater compared to the ΔkpnEF strain. The percent survival was calculated by comparison of viable cells in the WT. The data are means of measurements made in triplicate and performed three times. *, Significant difference (P < 0.05, Student t test).
Growth assays in the presence of efflux pump inhibitors.
The growth rates of the ΔkpnEF strain in the presence of 0.005 μg of chloramphenicol/ml, 0.005 μg of erythromycin/ml, and 0.005 μg of tetracycline/ml were 1.5-fold, 1.4-fold, and 1.3-fold lower, respectively, than that of the WT cells (P > 0.05) (see Fig. S2 in the supplemental material). The stunted growth profile in the mutant is due to the lack of KpnEF efflux pump in its functional form. To distinguish whether this is due to the loss of the efflux pump activity in the kpnEF mutant, the growth profile was monitored with CCCP, an uncoupler known to collapse the membrane energy and block the energy-dependent efflux pump. As expected, a substantial decrease in growth was observed in the ΔkpnEF mutant in the presence of CCCP, clearly illustrating the contribution of KpnEF as an efflux pump in K. pneumoniae. The growth profiles of WT, ΔkpnEF, and ΔkpnEFΩkpnEF strains in the presence of CCCP were equivocal compared to the growth observed in LB medium alone for WT, ΔkpnEF, and ΔkpnEFΩ kpnEF strains (data not shown).
Fluorimetric efflux assay.
To ensure that the antimicrobial resistance was a result of efflux by kpnEF, we tested the activity of the transporter directly by performing whole-cell EtBr transport assays. In the first 5 min, we found a decrease in fluorescence (which originates exclusively from EtBr binding to intracellular nucleic acid) due to active efflux of EtBr from WT cells. However, the addition of glucose energized the cells, which led to a relatively rapid decline in the fluorescence intensity as EtBr was being actively extruded from the WT cells against its concentration gradient. On the contrary, the addition of CCCP increased the fluorescence signal as the inhibitor dissipated the proton electrochemical gradient diminishing active efflux. Since the mutant lacks kpnEF in its functional form, the fluorescence intensity was higher because EtBr was not actively extruded relative to the WT strain. The study with ciprofloxacin yielded a similar conclusion regarding loss of the efflux capability of kpnEF (Fig. 7). Overall, these results suggest that the inactivation of kpnEF distorts active efflux capability in K. pneumoniae.
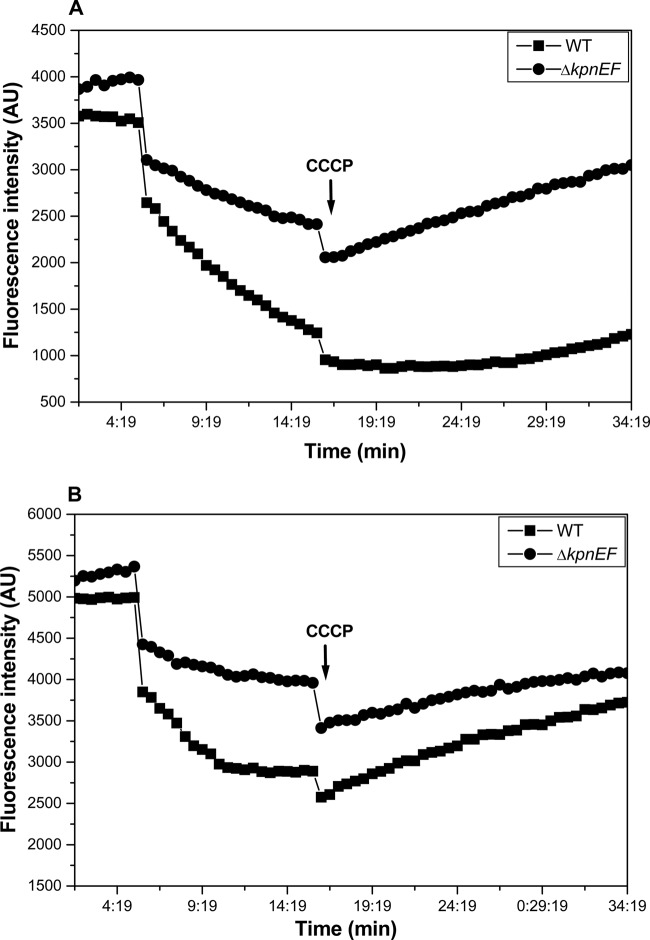
Fig 7.
Fluorimetric efflux assay. Accumulation studies were performed using EtBr (A) and ciprofloxacin (B) with K. pneumoniae and the ΔkpnEF mutant. The efflux of EtBr in mutant and WT cells was monitored continuously by measuring fluorescence emission at 600 nm upon excitation at 530 nm. After 5 min in a fluorimeter, cells loaded with EtBr were energized by the addition of glucose, and the efflux of EtBr was monitored. After 10 min, 100 μM CCCP was added as indicated to abolish active transport, and fluorescence emission was monitored further. The fluorescence was measured using a spectrofluorimeter (Hitachi). For ciprofloxacin, excitation was set at 275 nm and emission was set at 440 nm. Each data point represents the mean plus the standard deviation of three independent experiments.
Prevalence of kpnEF in K. pneumoniae clinical strains.
To check the prevalence of kpnEF in K. pneumoniae clinical isolates of Indian origin, we performed Southern hybridization using genomic DNA of clinical strains, namely, K2 (K1 serotype), K3 (K54 serotype), K7148 (K1 serotype), K823 (K2 serotype), K828 (K5 serotype), K814 (K1 serotype), and K140 (K1 serotype). Analysis revealed the occurrence of kpnEF in >70% of the isolates that were also multidrug resistant (MDR) (see Table S1 and Fig. S3 in the supplemental material).
To decipher the behavior of kpnEF in different strains and serotypes, we constructed the K3ΔkpnEF and K3ΔkpnEFΩkpnEF mutant strains from K. pneumoniae strain K3 (serotype K54) as described in Materials and Methods. The K3ΔkpnEF mutant exhibited decreased susceptibility to antibiotics and antiseptics, such as benzalkonium chloride (4-fold), chlorhexidine (2-fold), and SDS (8-fold) (see Table S2 in the supplemental material). The K3ΔkpnEF mutant exhibited motility, as well as a growth defect, compared to WT strain K3 (see Fig. S4 in the supplemental material). Interestingly, the presence of kpnEF in K. pneumoniae strain MGH78578 (serotype K52) was also determined in southern analysis, and subsequent disruption of the operon made the cells less susceptible to structurally related compounds such as EtBr (3-fold), SDS (6-fold), and benzalkonium chloride (2-fold) (data not shown).
Cell envelope stress response regulator CpxR regulates kpnEF in K. pneumoniae.
We previously demonstrated that the CpxAR two-component system (TCS) is involved in drug resistance in K. pneumoniae NTUH-K2044 (25). The DNA binding feature of CpxR prompted us to analyze the promoter region of kpnEF for the presence of putative CpxR binding sites. Interestingly, our analysis revealed the presence of a conserved putative CpxR binding site spanning the region between bp 328 and 343 from the first methionine of kpnEF (Fig. 8A). To define the possible interaction of CpxR to the promoter of kpnEF, we tested whether CpxR directly interacts with its promoter region. The cpxR gene from K. pneumoniae was cloned into pET-28C in our lab previously (Fig. 8B). Thereafter, gel shift assays were performed with the 32P-labeled kpnEF promoter fragment and CpxR. The protein-DNA complexes after incubation in reaction buffer were resolved on 5% PAGE, and analysis revealed a clear retardation, which was proportional to the protein concentration, as shown in Fig. 8C. The use of various controls, such as competitor inhibitor [specific and nonspecific poly(dI-dC)] and nonspecific protein (BSA), clearly demonstrated the precise DNA-binding ability of CpxR to the promoter of the SMR-type KpnEF efflux pump.
Fig 8.
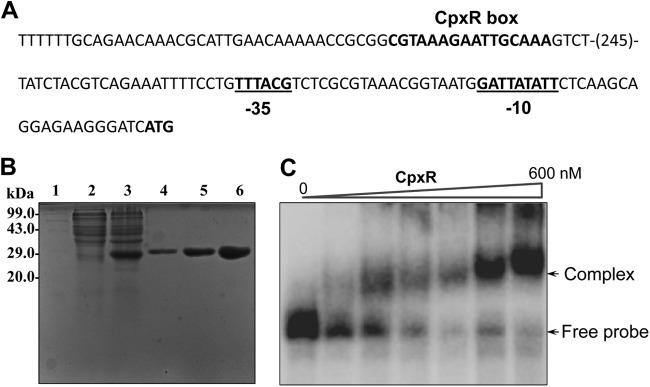
CpxR regulates KpnEF in K. pneumoniae. (A) Promoter region analysis of kpnEF. The numbers in parentheses represent the numbers of nucleotides in the sequence. The −35 and −10 regions in the promoter are underlined. The putative CpxR binding site is indicated in boldface. (B) SDS-PAGE profile of pET-cpxR. Lane 1, medium-size marker; lane 2, pET-cpxR/BL21DE3 uninduced; lane 3, pET-cpxR/BL21DE3, induced; lanes 4 to 6, purified CpxR fractions E1, E2, and E3, respectively. Protein samples after induction were subjected to SDS-PAGE (15% gel), followed by Coomassie brilliant blue staining. (C) Gel shift assays demonstrating the binding of CpxR to the promoter of kpnEF in K. pneumoniae in a concentration-dependent manner. Lane 1 shows the free probe, and lanes 2 to 7 show increasing concentrations of CpxR protein (60 to 600 nM), respectively. Protein-bound DNA complexes and free probe are indicated by arrows. The gels are representative of at least four independent experiments.
Expression analysis of the SMR-type KpnEF efflux pump and capsular genes in K. pneumoniae.
A cpxAR-null mutant and the respective complemented strain in K. pneumoniae NTUH-K2044 was constructed in our laboratory (25). Quantitative real-time RT-PCR was used to examine the expression of kpnEF in WT, cpxAR mutant, and complemented strains. Compared to the WT strain, the expression of kpnEF was decreased by 5.03-fold in the cpxAR mutant (P < 0.0003, Student t test) (Fig. 9). Complementation of the cpxAR mutation almost restored the expression of kpnEF (P < 0.0001). Together, these results provide strong evidence for the regulatory role of the CpxAR system on the SMR-type KpnEF efflux pump. Since mutation in kpnEF results in an impairment of capsule synthesis, we monitored the relative expression of capsular synthesis genes (KP1_3706, putative glycosyl transferase wcaI; KP1_3709, GDP-fucose synthetase wcaG; KP1_3712, galactoside O-acetyltransferase atf) in a kpnEF mutant. The ΔkpnEF strain showed a 4-fold (4.66 ± 1.7) increased expression for wcaI, a 2-fold (−2.46 ± 1.2) decreased expression for wcaG, and significant downregulation (−59.71 ± 2.8) for atf relative to the WT.
Fig 9.
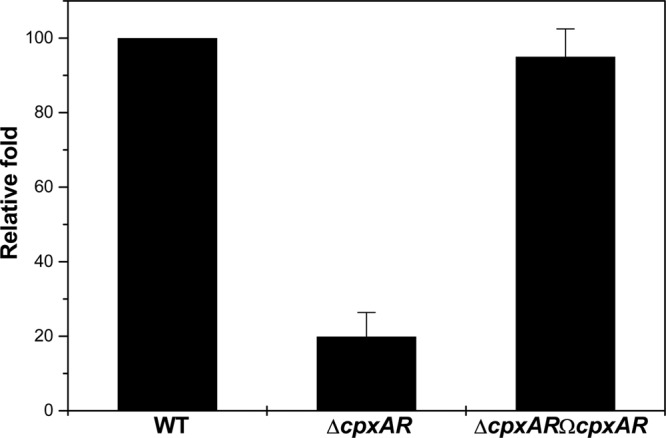
Relative transcriptional level of kpnEF in the ΔcpxAR mutant. The relative transcriptional levels of kpnEF in ΔcpxAR and ΔcpxARΩcpxAR strains determined using real-time RT-PCR are shown compared to the WT strain. The WT expression level is represented as 1-fold. Each bar represents the average value of three independent experiments. Error bars indicate the standard deviations.
DISCUSSION
According to the World Health Organization report 2010, respiratory diseases account for 11% of total deaths from all ages. A recent study documents that 80% of the many reported colorectal cancer-related pyogenic liver abscesses in Eastern Asia—especially in Japan, China, and Korea—are due to the K1 serotype of K. pneumoniae, and multidrug resistance among these isolates is a serious issue of global concern (35). The resistance of K. pneumoniae to β-lactams is mainly associated with acquired carbapenem hydrolyzing β-lactamases. In 2001, the first K. pneumoniae carbapenemase-producing isolate was reported in North Carolina (36). These β-lactamases can be metallo-β-lactamases (IMP and VIM), expanded-spectrum oxacillinases (OXA-48), or Ambler class A enzymes (NMCA, IMI, SME, GES, and K. pneumoniae carbapenemases) (37–39). It is worth noting here that in 2009 Pages et al. detected the masked role of efflux pumps in the β-lactam resistance phenotype in K. pneumoniae clinical isolates (40). Interestingly, analysis of completed K. pneumoniae genome sequences, namely, K. pneumoniae strain 1084 (5.39 Mb, 57.40 %G+C, NC_018522.1), K. pneumoniae strain HS11286 (5.68 Mb, 57.14 %G+C, NC_016845.1), K. pneumoniae strain MGH78578 (5.69 Mb, 57.17 %G+C, NC_009648.1), and K. pneumoniae strain NTUH-K2044 (5.47Mb, 57.39 %G+C, NC_012731.1) pinpoints the presence of chromosomal borne putative efflux pumps (20). Although role of efflux pump genes in multidrug resistance in K. pneumoniae MGH78578 has been demonstrated previously (41), the physiological relevance of efflux pumps in the hypervirulent strain NTUH-K2044 has never been explored. However, it has been reported to contain >15 such open reading frames (20). In addition to this, its genome does not contain the homolog of the well-characterized efflux pump E. coli EmrE; however, it contains a sugE homolog KP1_4280 annotated as a quaternary ammonium compound transport protein, as well as an ebrAB homolog, kpnEF (20). We describe here the functions of kpnEF, a member of Cpx regulon in bacterial physiology, stress response, and antimicrobial resistance.
Bile salts are amphipathic molecules that have a role in host defense due to its antimicrobial properties besides acting as detergents to help lipid digestion (33). Alterations in the outer membrane are considered an added strategy used by many enteric bacteria to combat the deleterious effects of bile; however, efflux still remains a vital mechanism (33). In our study, the kpnEF mutant turned sensitive (4.0-fold) to varied bile challenges, irrespective of the inoculum size, indicating that it facilitates K. pneumoniae colonization in the small intestine. Experimental evidence pinpointing the key role of KpnEF in the survival of the pleomorphic bacillus under conditions mimicking the upper parts of the gastrointestinal tract, where they encounter hyperosmotic conditions in a microaerobic environment, has been provided for the first time in K. pneumoniae.
Efflux pumps are transmembrane transporters, expressed in all types of prokaryotes and eukaryotes with wide substrate specificity that have common amphipathic character and ionizable groups (2, 3). The KpnEF efflux pump mediates resistance to antibiotics and compounds such as cefepime, ceftriaxone, colistin, erythromycin, rifampin, tetracycline, streptomycin, SDS, deoxycholate, EtBr, acriflavine, benzalkonium chloride, chlorhexidine, and triclosan. Moreover, antimicrobial susceptibility testing of the kpnEF mutant constructed in different K. pneumoniae serotypes confirmed its pivotal role in broad-spectrum antimicrobial resistance. Our observations corroborated well with previously defined functions for SMR-type efflux pumps, as in E. coli and P. aeruginosa emrE, Bacillus subtilis ebrAB, S. marcescens ssmE, and A. baumannii abeS (8, 9, 10, and 11). Reports on the role of SMR-type efflux pumps in antimicrobial resistance have been demonstrated in few clinically significant pathogens; however, its regulation has never been looked into in any prokaryotic system thus far.
The Cpx cascade is controlled by the TCS, consisting of the membrane-localized sensor kinase CpxA and the regulator CpxR (42). In a previous study, it was observed that among the nine drug efflux systems in Salmonella, quantitative real-time PCR analysis showed that baeR (a cpxR homolog) induced the expression of RND efflux pumps such as acrD and mdtABC (43). An earlier study in S. Typhimurium has shown that the loss of porin STM1530 and OmpD is regulated by CpxAR/BaeRS, which were important for ceftriaxone resistance (44). Identification of CpxR binding sites in the promoter region of kpnEF and our previous findings together provoked us to decipher the role of CpxR in regulating kpnEF. Strong binding of CpxR to the promoter of kpnEF and quantification of its expression in a cpxAR mutant background demonstrated kpnEF to be a member of the Cpx regulon.
A bacterial biofilm is an intricate, community-like structure that comprises bacterial cells embedded in a self-produced exopolysaccharide matrix, with a slimy mucoid appearance that confer protection against phagocytosis (45). Wu et al. has previously shown that the deletion of SugE, a membrane protein, results in increased biofilm formation associated with higher mucoviscosity and CPS production in this pleomorphic bacillus (46). Ho et al. recently reported that the deletion of capsular genes in K. pneumoniae led to a broad range of transcriptional suppression for 52 upstream genes, which included six multidrug efflux systems (47). However, inactivation of the membrane transporter kpnEF not only decreased biofilm formation but also led to loss in hypermucoviscosity and CPS synthesis, together with the decreased expression of CPS cluster genes. Mutation in the cpxAR TCS had no impact on mucoviscosity (25), biofilm formation, and motility (see Fig. S5 in the supplemental material). In agreement with our previous study, the TCS mutant showed no remarkable change in expression levels for wcaI (1.2 ± 0.82), wcaG (−1.15 ± 0.62), and atf (−2.83 ± 0.76). We propose that the KpnEF efflux pump might help transport polysaccharides to the outer layer of bacterial cell to form the slimy layer and is possibly under additional regulation by other transcriptional factors involved in modulating CPS synthesis and biofilm formation in K. pneumoniae. Given the dearth of knowledge on this topic, studies of this hypothesis are highly warranted. Overall, we report here for the first time the biological characterization of an SMR-type KpnEF efflux system in K. pneumoniae.
Supplementary Material
ACKNOWLEDGMENTS
We are extremely thankful and grateful to our Director Girish Sahni, CSIR-Institute of Microbial Technology (IMTECH), Chandigarh, India, for providing excellent facilities to carry out this work. We are deeply grateful to Jin-Town Wang, National Taiwan University Hospital, for providing K. pneumoniae NTUH-K2044, MGH78578, and other basic vectors. V.B.S. received an Innovative Young Biotechnologist Award from Department of Biotechnology (BT/01/IYBA/2009) for a grant-in-aid/fellowship.
The research was supported by the Department of Biotechnology (BT/01/IYBA/2009) and intramural funds from IMTECH. The funders had no role in the study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Footnotes
Published ahead of print 8 July 2013
Supplemental material for this article may be found at http://dx.doi.org/10.1128/AAC.02284-12.
REFERENCES
- 1.Alekshun MN, Levy SB. 2007. Molecular mechanisms of antibacterial multidrug resistance. Cell 128:1037–1050 [DOI] [PubMed] [Google Scholar]
- 2.Piddock LJ. 2006. Multidrug-resistance efflux pumps: not just for resistance. Nat. Rev. Microbiol. 4:629–636 [DOI] [PubMed] [Google Scholar]
- 3.Piddock LJ. 2006. Clinically relevant chromosomally encoded multidrug resistance efflux pumps in bacteria. Clin. Microbiol. Rev. 19:382–402 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Paulsen IT. 2003. Multidrug efflux pumps and resistance: regulation and evolution. Curr. Opin. Microbiol. 6:446–451 [DOI] [PubMed] [Google Scholar]
- 5.Putman M, Van Veen HW, Konings WN. 2000. Molecular properties of bacterial multidrug transporters. Microbiol. Mol. Biol. Rev. 64:672–693 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bay DC, Rommens KL, Turner RJ. 2008. Small multidrug resistance proteins: a multidrug transporter family that continues to grow. Biochim. Biophys. Acta 1778:1814–1838 [DOI] [PubMed] [Google Scholar]
- 7.Paulsen IT, Skurray RA, Tam R, Saier MH, Jr, Turner RJ, Weiner JH, Goldberg EB, Grinius LL. 1996. The SMR family: a novel family of multidrug efflux proteins involved with the efflux of lipophilic drugs. Mol. Microbiol. 19:1167–1175 [DOI] [PubMed] [Google Scholar]
- 8.Yerushalmi H, Lebendiker M, Schuldiner S. 1995. EmrE, an Escherichia coli 12-kDa multidrug transporter, exchanges toxic cations and H+ and is soluble in organic solvents. J. Biol. Chem. 270:6856–6863 [DOI] [PubMed] [Google Scholar]
- 9.Chung YJ, Saier MH. 2002. Overexpression of the Escherichia coli sugE gene confers resistance to a narrow range of quaternary ammonium compounds. J. Bacteriol. 184:2543–2545 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Minato Y, Shahcheraghi F, Ogawa W, Kuroda T, Tsuchiya T. 2008. Functional gene cloning and characterization of the SsmE multidrug efflux pump from Serratia marcescens. Biol. Pharm. Bull. 31:516–519 [DOI] [PubMed] [Google Scholar]
- 11.Srinivasan VB, Rajamohan G, Gebreyes WA. 2009. Role of AbeS, a novel efflux pump of the SMR family of transporters, in resistance to antimicrobial agents in Acinetobacter baumannii. Antimicrob. Agents Chemother. 53:5312–5316 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Bay DC, Turner RJ. 2009. Diversity and evolution of the small multidrug resistance protein family. BMC Evol. Biol. 9:140. 10.1186/1471-2148-9-140 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Jack DL, Storms ML, Tchieu JH, Paulsen IT, Saier MH., Jr 2000. A broad specificity multidrug efflux pump requiring a pair of homologous SMR-type proteins. J. Bacteriol. 182:2311–2313 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Masaoka Y, Ueno Y, Morita Y, Kuroda T, Mizushima T, Tsuchiya T. 2000. A two-component multidrug efflux pump, EbrAB, in Bacillus subtilis. J. Bacteriol. 182:2307–2310 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Poulsen BE, Rath A, Deber CM. 2009. The assembly motif of a bacterial small multidrug resistance protein. J. Biol. Chem. 284:9870–9875 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Podschun R, Ullmann U. 1998. Klebsiella spp. as nosocomial pathogens: epidemiology, taxonomy, typing methods, and pathogenicity factors. Clin. Microbiol. Rev. 11:589–603 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Highsmith AK, Jarvis WR. 1985. Klebsiella pneumoniae: selected virulence factors that contribute to pathogenicity. Infect. Control 6:75–77 [DOI] [PubMed] [Google Scholar]
- 18.Arnold RS, Thom KA, Sharma S, Phillips M, Kristie Johnson J, Morgan DJ. 2011. Klebsiella pneumoniae carbapenemase-producing bacteria. South Med. J. 104:40–45 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Nordmann P, Cuzon G, Naas T. 2009. The real threat of KPC carbapenemase-producing bacteria. Lancet Infect. Dis. 9:228–2336 [DOI] [PubMed] [Google Scholar]
- 20.Wu KM, Li LH, Yan JJ, Tsao N, Liao TL, Tsai HC, Fung CP, Chen HJ, Liu YM, Wang JT, Fang CT, Chang SC, Shu HY, Liu TT, Chen YT, Shiau YR, Lauderdale TL, Su IJ, Kirby R, Tsai SF. 2009. Genome sequencing and comparative analysis of Klebsiella pneumoniae NTUH-K2044, a strain causing liver abscess and meningitis. J. Bacteriol. 191:4492–4501 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Padilla E, Llobet E, Doménech-Sánchez A, Martínez-Martínez L, Bengoechea JA, Albertí S. 2010. Klebsiella pneumoniae AcrAB efflux pump contributes to antimicrobial resistance and virulence. Antimicrob. Agents Chemother. 54:177–183 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ogawa W, Onishi M, Ni R, Tsuchiya T, Kuroda T. 2012. Functional study of the novel multidrug efflux pump KexD from Klebsiella pneumoniae. Gene 498:177–182 [DOI] [PubMed] [Google Scholar]
- 23.Ping Y, Ogawa W, Kuroda T, Tsuchiya T. 2007. Gene cloning and characterization of KdeA, a multidrug efflux pump from Klebsiella pneumoniae. Biol. Pharm. Bull. 30:1962–1964 [DOI] [PubMed] [Google Scholar]
- 24.Ogawa W, Koterasawa M, Kuroda T, Tsuchiya T. 2006. KmrA multidrug efflux pump from Klebsiella pneumoniae. Biol. Pharm. Bull. 29:550–553 [DOI] [PubMed] [Google Scholar]
- 25.Srinivasan VB, Vaidyanathan V, Mondal A, Rajamohan G. 2012. Role of the two-component signal transduction system CpxAR in conferring cefepime and chloramphenicol resistance in Klebsiella pneumoniae NTUH-K2044. PLoS One 7:e33777. 10.1371/journal.pone.0033777 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Srinivasan VB, Venkataramaiah M, Mondal A, Vaidyanathan V, Govil T, Rajamohan G. 2012. Functional characterization of a novel outer membrane porin KpnO, regulated by PhoBR two-component system in Klebsiella pneumoniae NTUH-K2044. PLoS One 7:e41505. 10.1371/journal.pone.0041505 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Fang CT, Chuang YP, Shun CT, Chang SC, Wang JT. 2004. A novel virulence gene in Klebsiella pneumoniae strains causing primary liver abscess and septic metastatic complications. J. Exp. Med. 199:697–705 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Bazzini S, Udine C, Sass A, Pasca MR, Longo F, Emiliani G, Fondi M, Perrin E, Decorosi F, Viti C, Giovannetti L, Leoni L, Fani R, Riccardi G, Mahenthiralingam E, Buroni S. 2011. Deciphering the role of RND efflux transporters in Burkholderia cenocepacia. PLoS One 6:e18902. 10.1371/journal.pone.0018902 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Coudeyras S, Nakusi L, Charbonnel N, Forestier C. 2008. A tripartite efflux pump involved in gastrointestinal colonization by Klebsiella pneumoniae confers a tolerance response to inorganic acid. Infect. Immun. 76:4633–4641 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Hennequin C, Forestier C. 2009. oxyR, a LysR-type regulator involved in Klebsiella pneumoniae mucosal and abiotic colonization. Infect. Immun. 77:5449–5457 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Clinical and Laboratory Standards Institute 2006. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically; approved standard M7-A7, vol 26-2 Clinical and Laboratory Standards Institute, Wayne, PA [Google Scholar]
- 32.Sigal N, Vardy E, Molshanski-Mor S, Eitan A, Pilpel Y, Schuldiner S, Bibi E. 2005. 3D model of the Escherichia coli multidrug transporter MdfA reveals an essential membrane-embedded positive charge. Biochemistry 44:14870–14880 [DOI] [PubMed] [Google Scholar]
- 33.Gunn JS. 2000. Mechanisms of bacterial resistance and response to bile. Microbes Infect. 2:907–913 [DOI] [PubMed] [Google Scholar]
- 34.Stevanin TM, Ioannidis N, Mills CE, Kim SO, Hughes MN, Poole RK. 2000. Flavohemoglobin Hmp affords inducible protection for Escherichia coli respiration, catalyzed by cytochromes bo′ or bd, from nitric oxide. J. Biol. Chem. 46:35868–35875 [DOI] [PubMed] [Google Scholar]
- 35.Qu K, Liu C, Wang ZX, Tian F, Wei JC, Tai MH, Zhou L, Meng FD, Wang RT, Xu XS. 2012. Pyogenic liver abscesses associated with nonmetastatic colorectal cancers: an increasing problem in Eastern Asia. World J. Gastroenterol. 18:2948–2955 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Yigit H, Queenan AM, Anderson GJ, Domenech-Sanchez A, Biddle JW, Steward CD. 2001. Novel carbapenem-hydrolyzing β-lactamase KPC-1 from a carbapenem-resistant strain of Klebsiella pneumoniae. Antimicrob. Agents Chemother. 45:1151–1161 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Nordmann P, Poirel L. 2002. Emerging carbapenemases in gram-negatives aerobes. Clin. Microbiol. Infect. 8:321–331 [DOI] [PubMed] [Google Scholar]
- 38.Poirel L, Héritier C, Tolun V, Nordmann P. 2004. Emergence of oxacillinase-mediated resistance to imipenem in Klebsiella pneumoniae. Antimicrob. Agents Chemother. 48:15–22 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Queenan AM, Bush K. 2007. Carbapenemases: the versatile beta-lactamases. Clin. Microbiol. Rev. 20:440–458 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Pages JM, Lavigne JP, Leflon-Guibout V, Marcon E, Bert F, Noussair L, Nicolas-Chanoine MH. 2009. Efflux pump, the masked side of beta-lactam resistance in Klebsiella pneumoniae clinical isolates. PLoS One 4:e4817. 10.1371/journal.pone.0004817 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Ogawa W, Li DW, Yu P, Begum A, Mizushima T, Kuroda T, Tsuchiya T. 2005. Multidrug resistance in Klebsiella pneumoniae MGH78578 and cloning of genes responsible for the resistance. Biol. Pharm. Bull. 28:1505–1508 [DOI] [PubMed] [Google Scholar]
- 42.Raivio TL, Silhavy TJ. 1997. Transduction of envelope stress in Escherichia coli by the Cpx two-component system. J. Bacteriol. 179:7724–7733 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Nishino K, Nikaido E, Yamaguchi A. 2007. Regulation of multidrug efflux systems involved in multidrug and metal resistance of Salmonella enterica serovar Typhimurium. J. Bacteriol. 189:9066–9075 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Hu WS, Chen HW, Zhang RY, Huang CY, Shen CF. 2011. The expression levels of outer membrane proteins STM1530 and OmpD, which are influenced by the CpxAR and BaeSR two-component systems, play important roles in the ceftriaxone resistance of Salmonella enterica serovar Typhimurium. Antimicrob. Agents Chemother. 55:3829–3837 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Di Martino P, Cafferini N, Joly B, Darfeuille-Michaud A. 2003. Klebsiella pneumoniae type 3 pili facilitate adherence and biofilm formation on abiotic surfaces. Res. Microbiol. 154:9–16 [DOI] [PubMed] [Google Scholar]
- 46.Wu MC, Lin TL, Hsieh PF, Yang HC, Wang JT. 2011. Isolation of genes involved in biofilm formation of a Klebsiella pneumoniae strain causing pyogenic liver abscess. PLoS One 6:e23500. 10.1371/journal.pone.0023500 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Ho JY, Lin TL, Li CY, Lee A, Cheng AN, Chen MC, Wu SH, Wang JT, Li TL, Tsai MD. 2011. Functions of some capsular polysaccharide biosynthetic genes in Klebsiella pneumoniae NTUH K-2044. PLoS One 6:e21664. 10.1371/journal.pone.0021664 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Turton JF, Perry C, Elgohari S, Hampton CV. 2010. PCR characterization and typing of Klebsiella pneumoniae using capsular type-specific, variable number tandem repeat and virulence gene targets. J. Med. Microbiol. 59:541–547 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.