Abstract
Outbreaks of human infection linked to the powdered infant formula (PIF) food chain and associated with the bacterium Cronobacter, are of concern to public health. These bacteria are regarded as opportunistic pathogens linked to life-threatening infections predominantly in neonates, with an under developed immune system. Monitoring the microbiological ecology of PIF production sites is an important step in attempting to limit the risk of contamination in the finished food product. Cronobacter species, like other microorganisms can adapt to the production environment. These organisms are known for their desiccation tolerance, a phenotype that can aid their survival in the production site and PIF itself. In evaluating the genome data currently available for Cronobacter species, no sequence information has been published describing a Cronobacter sakazakii isolate found to persist in a PIF production facility. Here we report on the complete genome sequence of one such isolate, Cronobacter sakazakii SP291 along with its phenotypic characteristics. The genome of C. sakazakii SP291 consists of a 4.3-Mb chromosome (56.9% GC) and three plasmids, denoted as pSP291-1, [118.1-kb (57.2% GC)], pSP291-2, [52.1-kb (49.2% GC)], and pSP291-3, [4.4-kb (54.0% GC)]. When C. sakazakii SP291 was compared to the reference C. sakazakii ATCC BAA-894, which is also of PIF origin, the annotated genome data identified two interesting functional categories, comprising of genes related to the bacterial stress response and resistance to antimicrobial and toxic compounds. Using a phenotypic microarray (PM), we provided a full metabolic profile comparing C. sakazakii SP291 and the previously sequenced C. sakazakii ATCC BAA-894. These data extend our understanding of the genome of this important neonatal pathogen and provides further insights into the genotypes associated with features that can contribute to its persistence in the PIF environment.
Keywords: complete genome, plasmid, Cronobacter sakazakii, stress response, antibiotic resistance, toxic compounds
Introduction
Cronobacter species (formerly Enterobacter sakazakii) is an opportunistic pathogen of the Enterobacteriaceae family. This organism was originally designated as E. sakazakii in 1980 (Farmer et al., 1980). Based on its recently revised taxonomy, the genus was renamed as Cronobacter in 2007 and now consists of seven species, C. sakazakii, C. malonaticus, C. turicensis, C. muytjensii, C. dublinensis (including three subspecies, dublinensis, lausannensis, and lactaridi), C. universalis and C. condimenti (Iversen et al., 2004, 2007, 2008; Joseph et al., 2011). Infections caused by Cronobacter can present as necrotizing enterocolitis, bacteremia and meningitis, with long term complications for those that survive, including delayed neurological development, hydrocephalus and permanent neurological damage. Life-threatening infections have been reported in neonates (of less than 28 days) (Bar-Oz et al., 2001; Gurtler et al., 2005; Mullane et al., 2007), as well as older infants, with lethality rates ranging between 40 and 80% (Bowen and Braden, 2006; Friedemann, 2009), and more recently in immune-compromised adults, mainly the elderly (Gosney et al., 2006; See et al., 2007; Hunter et al., 2008; Tsai et al., 2013).
Cronobacter can be isolated from a wide range of foods and environments (Baumgartner et al., 2009; Chap et al., 2009; El-Sharoud et al., 2009; Jaradat et al., 2009; Schmid et al., 2009). Specifically, contaminated powdered infant formula (PIF) has been epidemiologically linked with many of the neonatal and infant infections (Himelright et al., 2002; Bowen and Braden, 2006; Mange et al., 2006). Previous studies reported the isolation of Cronobacter from PIF, and the PIF production environment (Drudy et al., 2006; Mullane et al., 2008a,b), suggesting that this bacterium has the capacity to adapt to, survive and persist under desiccated environmental conditions. Comparison of environmental and clinical Cronobacter isolates, indicated that the desiccation tolerance exhibited might play a role in the persistence of Cronobacter in PIF and its associated low-moisture ingredients (Walsh et al., 2011; Beuchat et al., 2013). Stress response factors identified previously in Cronobacter, which include heat-shock, cold-stresses, survival in dry conditions, water activity (aw), and pH may contribute to this phenotype (Dancer et al., 2009a,b; Carranza et al., 2010; Chang et al., 2010; Arku et al., 2011). Genome sequencing efforts of Cronobacter species commenced in 2010. To date, 16 genomes are currently available, of which three, C. sakazakii ATCC BAA-894, C. sakazakii ES15 and C. turicensis z3032, are complete (Kucerova et al., 2010; Stephan et al., 2011; Joseph et al., 2012; Shin et al., 2012; Grim et al., 2013).
Following on-going surveillance of a PIF production facility in our laboratory, an interesting isolate C. sakazakii SP291 was identified which exhibited a thermo-adapted phenotype when compared with other Cronobacter and Salmonella species tested under laboratory conditions (Cooney, 2012). In an effort to better understand C. sakazakii SP291, its genome was completely sequenced and compared to that of a PIF isolate C. sakazakii ATCC BAA-894, a whole grain isolate C. sakazakii ES15, a clinical isolate C. turicensis z3032 and other selected draft genomes. Additionally, we interrogated the phenome of C. sakazakii SP291, to determine the functionality of strain-specific genotypic traits that may contribute to its adaption capacity in a PIF production environment.
Materials and methods
Bacterial isolates studied and their culture conditions
Seventeen Cronobacter isolates used in this study are listed in Table 1. Cronobacter sakazakii SP291 was assigned according to the classic rpoB method described previously (Stoop et al., 2009; Lehner et al., 2012). The isolate was cultured routinely in an Isotherm® Forced Convection Laboratory Incubator (Esco GB Ltd., Downton, UK) at 37°C on Trypticase Soy Agar (Oxoid Limited, Hampshire, UK) and stored at −80°C on cryo-beads (Technical Service Consultants Ltd., Lancashire, UK).
Table 1.
Cronobacter species, the strain identifier, source, country of origin, and accession numbers.
| Species | Strain identifiera | Serogroupb | Source | Country of origin | Accession number (GeneBank) |
|---|---|---|---|---|---|
| Cronobacter sakazakii | ATCC BAA-894 | Csak O1 | PIFg | USA | CP000783-CP00785 |
| Cronobacter sakazakii | SP291 | Csak O2 | PIF manufacturing environment | Ireland | CP004091-CP004094 |
| Cronobacter sakazakii | ES15 | NDf | Whole grain | Korea | CP003312 |
| Cronobacter sakazakii | E899 | Csak O2 | Clinical | USA | AFMO01000001-AFMO01000385 |
| Cronobacter sakazakii | 680 | NDf | Clinical | USA | CALG01000001-CALG01000201 |
| Cronobacter sakazakii | 696 | NDf | Clinical | France | CALF01000001-CALF01000569 |
| Cronobacter sakazakii | 701 | NDf | Clinical | France | CALE01000001-CALE01000768 |
| Cronobacter sakazakii | ES35 | Csak O1 | Clinical | Israel | AJLC01000001-AJLC01000183 |
| Cronobacter sakazakii | 2151 | Csak O2 | Clinical, cerebrospinal fluid | USA | AJKT01000001-AJKT01000060 |
| Cronobacter sakazakii | ES713 | Csak O2 | PIF | USA | AJLB01000001-AJLB01000156 |
| Cronobacter sakazakii | E764 | Csak O4 | Clinical | Czech Republic | AJLA01000001-AJLA01000032 |
| Cronobacter malonaticus | LMG 23826 | Cmal O2 | Human, breast abscess | USA | CALC01000001-CALC01000171 |
| Cronobacter turicensis | z3032c | Ctur O1 | Neonate | Switzerland | FN543093-FN543096 |
| Cronobacter dublinensis | CFS 237d | Cdub O1 | PIF | Ireland | CAKZ01000001-CAKZ01000221 |
| Cronobacter mutjensii | ATCC 51329 | CmuyO2 | Unknown | Unknown | AJKU01000001-AJKU01000072 |
| Cronobacter universalis | NCTC 9529 | Cuni O1 | Water | UK | CAKX01000001-CAKX01000231 |
| Cronobacter condimenti | 1330e | NDf | Spiced meat | Slovakia | CAKW01000001-CAKW01000155 |
Strain information was selected from publications (Kucerova et al., 2010; Chen et al., 2011; Stephan et al., 2011; Joseph et al., 2012; Shin et al., 2012; Grim et al., 2013).
Serogroup designations were identified using primers described by Mullane et al. (2008a,b) and Jarvis et al. (2011, 2013).
Cronobacter turicensis species type strain LMG 23827.
Cronobacter dublinensis species type strain LMG 23823.
Cronobacter condimenti species type strain LMG 26250.
Not determined.
Isolate cultured from PIF, of which the PFGE pattern matched the blood sample of an infected neotate in a neonatal intensive care unit (NICU) in Tennessee in 2001. The infection cause the death of the neotate born 20 days previously.
DNA sequencing, annotation, and comparative genomic analysis
Total genomic DNA was purified using a DNeasy Blood and Tissue Kit (QIAGEN, Hilden, Germany) following the manufacturer's instructions. Concentrations were measured using a Nanodrop® (ND 1000) spectrophotometer (Labtech International Ltd., Luton, UK). Purified DNA was maintained at −20°C until required. The whole genome sequencing and assembly methodology is described elsewhere (Power et al., 2013). The complete chromosome and plasmid sequences were uploaded to the RAST (Rapid Annotation using Subsystem Technology) (Aziz et al., 2008) annotation server in a FASTA file format. The RAST server automatically identifies protein-encoding, tRNA and rRNA genes, assigns their functions, predicts the presence of subsystems in the genome, and reconstructs the metabolic network (Aziz et al., 2008). Genome to genome comparative analysis was performed in the SEED viewer as previously reported (Overbeek et al., 2005; Grim et al., 2013). Three complete genomes of C. sakazakii ATCC BAA-894 (Accession number CP000783-CP00785), C. sakazakii ES15 (Accession number CP003312) and C. turicensis z3032 (Accession number FN543093-FN543096) were uploaded and annotated in RAST, and used as reference sequences. Most probable insertion or deletion genome regions of C. sakazakii SP291 were identified as previously reported (Grim et al., 2013). In addition, nitrogen metabolism genes, stress-coding genes, as well as antibiotic and toxic compound resistant genes were determined based on significant identity alignments using BLAST. The genome sequence of C. sakazakii SP291 was deposited in GenBank under the accession numbers CP004091-CP004094. The accession numbers for other genome sequences studied were included in Table 1.
Phenotype microarray analysis
Phenotype microarray (PM) analysis was performed on C. sakazakii ATCC BAA-894 and C. sakazakii SP291 using the OmniLog® automated incubator/reader (Biolog Inc., Hayward, USA) following manufacturer's instruction. All 20 plates (PM-1 through PM-20) inoculated with bacterial cell suspensions, were incubated at 37°C and cell respiration was measured every 15 min for 48 h. The tetrazolium redox dye is reduced when bacteria respire, which provides both amplification and quantitation of the phenotype. Analysis was carried out using OmniLog® phenotype microarray software v1.2 to determine the phenotypic differences. Negative control wells, which contained the inoculated Omnilog™ growth medium, but without any substrate, were measured to normalize differences in inocula and redox dye oxidation between samples. The respiration profiles for both strains were compared using PM's integration function software and a significant divergent phenotype was identified when a difference in Omnilog™ units of 20,000 ± 1800 or greater between the two strains was obtained.
Results and discussion
Cronobacter sakazakii SP291 genome
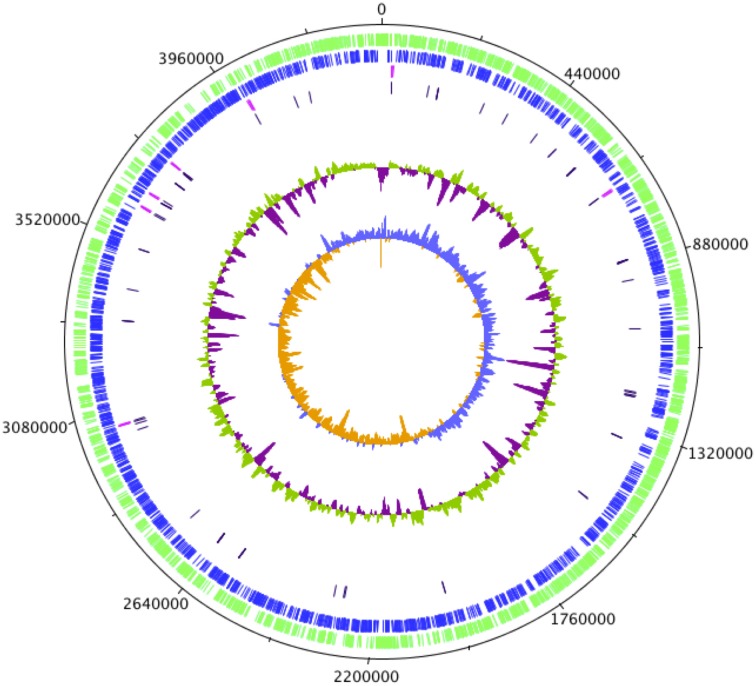
The complete genome sequence of C. sakazakii SP291 is composed of a single, circular chromosome, 4.34 Mb in length with an average GC content of 56.9% along with three plasmids (denoted as pSP291-1, 118.136 kb, 57.2% GC; pSP291-2, 52.134 kb, 49.2% GC and pSP291-3, 4.422 kb, 54.0% GC) (Accession number CP004091-CP004094). The general features of the genome are presented in Table 2. A total of 4129 genes were identified on the chromosome, including 82 tRNA and 22 rRNA genes. The protein coding sequence (CDS) represents 86.3% of the genome and is organized into 4025 CDS, with an average length of 931 nucleic acids (Figure A1). From the annotation of the three plasmids, it was determined that 116 genes cover 87.1% of pSP291-1, 74 genes cover 77.2% of pSP291-2 and 7 genes were located on pSP291-3 and accounts for 48.6% of this structure.
Table 2.
General features of the C. sakazakii SP291 genome.
| Feature | Chromosome | Plasmids | ||
|---|---|---|---|---|
| pSP291-1 | pSP291-2 | pSP291-3 | ||
| Size (bp) | 4,344,092 | 118,136 | 52,134 | 4,422 |
| Predicted CDS | 4025 | 116 | 74 | 7 |
| GC content (%) | 56.9 | 57.2 | 49.2 | 54.0 |
| Coding regions (%) | 86.3 | 87.1 | 77.2 | 48.6 |
| Average CDS length (bp) | 931 | 887 | 544 | 307 |
| tRNA | 82 | nil | nil | nil |
| rRNA | 22 | nil | nil | nil |
Comparative genomic analysis of C. sakazakii SP291 with three other completed Cronobacter genomes
Cronobacter sakazakii SP291 and three other completed genomes: C. sakazakii ATCC BAA-894, C. sakazakii ES15 and C. turicensis z3032 were compared (Figure 1). For the purposes of this comparison, the C. sakazakii ATCC BAA-894 genome was used as the reference. Five genomic regions (denoted as GR-1 through −5, in Figure 1A) were identified and these were present in the other genomes but missing in C. sakazakii SP291 (Table S1). These GRs are discussed in detail below.
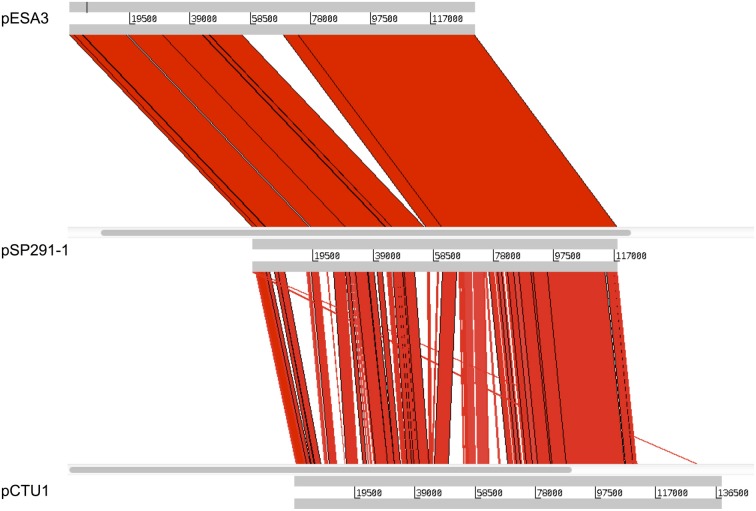
Figure 1.
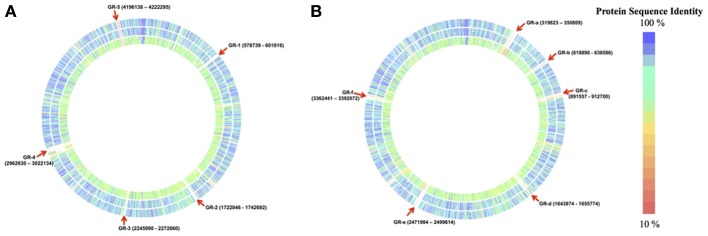
Genomic regions of C. sakazakii SP291 compared to three completed genomes consists of C. sakazakii ATCC BAA 894, C. sakazakii ES15, and C. turicensis z3032. (A) Regions absent in the genome of C. sakazakii SP291 compared to the other three genomes. Reference genome: C. sakazakii ATCC BAA-894; Outer circle: C. sakazakii ES15; Middle circle: C. sakazakii SP291; Inner circle: C. turicensis z3032, see also Table S1. (B) Regions present in the genome of C. sakazakii SP291 compared to the other three genomes. Reference genome: C. sakazakii SP291; Outer circle: C. sakazakii ATCC BAA-894; Middle circle: C. sakazakii ES15; Inner circle: C. turicensis z3032, see also Table S2.
Several unique prophages or phage-like elements of C. sakazakii ATCC BAA-894 were noted in GR-1 (genome positions 578,739…601,816), GR-3 (genome positions 2,245,990…2,272,660) and GR-4 (genome positions 2,962,630…3,022,134). Three specific genes were observed in GR-1, which included the DNA-methyltransferase subunit M and the S subunit of a type I restriction-modification system, along with a zinc binding domain/DNA primase, which is a phage P4-associated/replicative helicase denoted as RepA. A lambda phage portal protein, a large subunit of a terminase enzyme, along with some hypothetical proteins were noted in GR-3. In GR-4, a 1478 bp uncharacterized translocase gene required for O-antigen conversion and two-recombination genes, part of the bacteriophage ninR region, and denoted as ninB and ninG were identified. These annotations confirmed those previous reported (Kucerova et al., 2010). Interstingly, a putative bactoprenol glucosyl transferase was identified in C. sakazakii ATCC BAA-894, and shared with C. sakazakii ES15 and C. turicensis z3032, but not C. sakazakii SP291. Regulatory protein CII along with a phage Kil protein were annotated in C. sakazakii ATCC BAA-894 but not C. sakazakii SP291. Although protein CII was absent in C. turicensis z3032, the Kil protein was present (Stephan et al., 2011). A recently sequenced phage denoted as, phiES15, contained cII and kil (Lee et al., 2012). Unique transposon genes were noted in GR-2 (genome positions 1,722,846…1,742,692) and GR-5 (genome positions 4,196,138…4,222,295). In GR-2, a large part of the region containing tellurium resistance-encoding genes, including terX, terW, terA, terB, terC, and terD, were identified, a feature reported previously (Kucerova et al., 2010; Joseph et al., 2012; Grim et al., 2013). GR-5 contained heavy metal efflux and resistance genes, which consists of cusS, cusR, cusC, cusF, czcB, czcA, cusA, copG, pcoS, pcoB, and pcoA (Kucerova et al., 2010; Joseph et al., 2012). Further detailed information related to the corresponding phenotypes is outlined below (see also Table S1).
Genes unique to C. sakazakii SP291 were also noted and these were used as a reference to interrogate the genomes of the other strains. Six genomic regions (Figure 1B, denoted as GR-a through -f, Table S2) were identified as being unique to C. sakazakii SP291. GR-a (genome regions 319,823…350,809), GR-d (genome regions 1,643,874…165,774) and GR-e (genome regions 2,471,984…2,499,614) contained a set of phage- and phage-related proteins along with some hypothetical proteins. A phage regulatory CII-like protein was identified in C. sakazakii SP291 and mapped within GR-a, which also matched a similar homolog observed in C. turicensis z3032. A holin protein, which controls the length of an infective cycle for bacteriophage (Wang et al., 2000), together with membrane proteins related to metalloendopeptidases were present in C. sakazakii SP291 alone, being located in GR-e. In GR-b (genome regions 618,890…638,586), a YkfI toxin-encoding protein along a YfjZ-antitoxin encoding protein (the corresponding antitoxin to YpjF) were identified and unique to C. sakazakii SP291. This toxin-antitoxin protein pair was also reported in E. coli previously and was shown to regulate cell death through the disruption of essential cellular processes (Brown and Shaw, 2003). It has been proposed by Lewis (2000) that, under some circumstances, it may be evolutionarily advantageous for some cells in a population to undergo programmed cell death in order to provide nutrients for the remainder. Toxin-antitoxin pairs were noted in a previous study as most Cronobacter genomes contain a large number of them, which might be conserved, shared, or unique (Grim et al., 2013). GR-c (genome regions 891,557…912,700) contains seven interesting genes, which includes an uncharacterized protein YeeT, a NgrB protein, an ATP-dependent Clp protease, an ATP-binding subunit ClpA, a small HspC2 heat shock protein, a galactoside O-acetyltransferase-encoding gene and an anti-restriction protein KlcA which have been reported as a component part of a type I DNA restriction system (Serfiotis-Mitsa et al., 2010). A helicase protein, a glycerol dehydrogenase enzyme-encoding gene, and a DNA-cytosine methyltransferase were identified within GR-f (genome regions 3,363,441…3,392,072). Of note, a type I restriction-modification system, specificity the S-subunit-like gene, was identified in C. sakazakii SP291, a feature which was noted earlier in C. sakazakii ATCC BAA-894 (Kucerova et al., 2010; Joseph et al., 2012).
Comparative genomic analysis of C. sakazakii SP291 and selected available genomes within this genus
Two earlier studies described the core genome of Cronobacter (Joseph et al., 2012; Grim et al., 2013). The availability of C. sakazakii SP291 genome has provided an opportunity to re-evaluate the content of the Cronobacter core gemome, comparing it to other currently available genome sequences within the genus. Thus, a comparison between C. sakazakii SP291 and 16 other Cronobacer genomes (Table 1) was performed in SEED viewer server.
Within the 11 Cronobacter sakazakii isolates compared, 57 annotated genes were present in C. sakazakii SP291, but absent in all other genomes, including 41 hypothetical proteins, 12 phage- and prophage-related genes/proteins and four other genes/proteins (Table S3). Among all seven Cronobacter species, there were 154 annotated genes/proteins absent in other species, including 122 hypothetical proteins, 4 phage- and prophage-related genes/proteins and 28 unique genes/proteins (Table S4). Interestingly, a conserved domain protein was identified that was unique to C. sakazakii SP291, which is associated with retron-type reverse transcriptase. Fifteen genes were shared with other species by C. sakazakii SP291, but were absent in all the C. sakazakii genomes compared to date, and these consisted of a retron-type RNA-directed DNA polymerase, a holin protein which controls the timing of bacteriophage infections as mentioned earlier, a topoisomerase IA-encoding protein, and 12 phage- and prophage-related proteins. There were 31 proteins, which are only shared with some of the C. sakazakii genomes by C. sakazakii SP291 and which were absent among the other six species. These included a sodium-dependent phosphate transporter protein, a RelE antibacterial toxin protein, a RelB protein (antitoxin to RelE), a probable poly (beta-D-mannuronate) O-acetylase protein, two putative periplasmic proteins, a possible secretory protein, a GTPase protein, denoted as NgrB, a mobile element protein, a galactoside O-acetyltransferase protein, a mannose-6-phosphate isomerase, class I protein, a different locus type I restriction-modification system, specificity the S subunit-like protein, a predicted transcriptional regulator COGs COG2378, permeases of the major facilitator superfamily, a superfamily II DNA/RNA helicases, SNF2 family, a DNA modification methylase, an IS1 transposase OrfA protein, a probable tonB-dependent receptor yncD precursor, a putative ORF-4 protein, a putative ORF (located using Glimmer/Genemark), seven beta-fimbriae probable major subunits, and four phage related proteins.
Annotated plasmids contained in C. sakazakii SP291
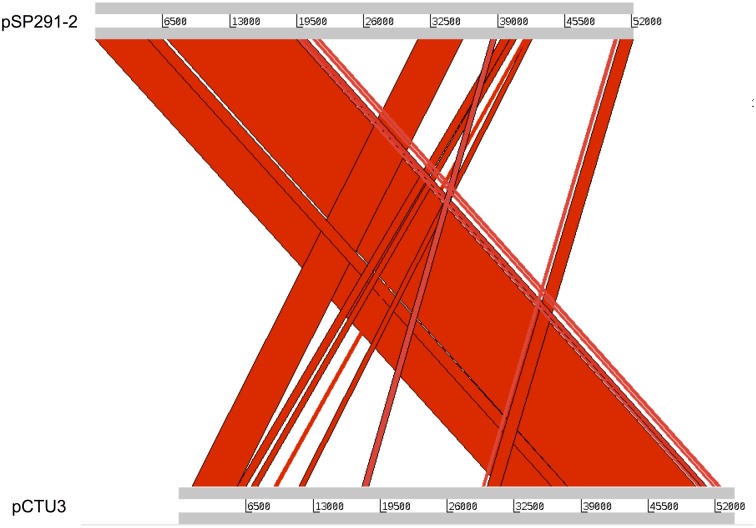
Cronobacter sakazakii SP291 contains three plasmids, including pSP291-1, 118,136 bp (57.2% GC), pSP291-2, 52,134 bp (49.2%) and pSP291-3, 4,422 bp (54.0% GC). The predicted CDS of pSP291-1 was found to be 116, with the average length of 887 bp, while pSP291-2 has 74 CDS with an average length of 544 bp, and pSP291-3 has seven CDS and with the average length of 307 bp (Table 2). Comparision of all three plasmids with five previously published plasmid sequences (including pESA2 and pESA3 of C. sakazakii ATCC BAA-894; along with pCTU1, pCTU2, and pCTU3 of C. turicensis z3032) indicated two closely related plasmid groups. Plasmid group 1, contains pSP291-1, pESA3, and pCTU1, while plasmid group 2, consists of pSP291-2 and pCTU3 (Figures A2, A3, and Table S5).
Several common genes were identified in plasmid group 1, These consisted of a complete ABC transporter (which could function to transport iron; vitamin B12; siderophores and hemin), including the ATP-binding component, the periplasmic substrate-binding module and the permease element. These genes were identified in all three plasmids. An aerobactin siderophore receptor (the IutA/TonB-dependent siderophore receptor) was shared between the three plasmids, while a Cronobacter plasminogen activator (cpa) homolog has only been mapped to pESA3 and pSP291-1, but not pCTU1, which is in agreement with the results reported by Franco et al. (2011) and Grim et al. (2012) (Figure A2). Three arsenical resistance genes were identified on all three plasmids along with pCTU3. Genes corresponding to commonly shared proteins on all three plasmids included a C-terminal helicase protein, a HipA protein previously reported to be required for growth arrest and multi-drug resistance in Escherichia coli (Correia et al., 2006), a hypothetical-encoding gene ycgF reported to be a direct anti-repressor which acts in the blue-light response of E. coli (Tschowri et al., 2009), a starvation sensing protein RspA, a magnesium transporting P-type 1 ATPase protein, transcriptional regulators, including members of ArsR family, GntR family (Kucerova et al., 2010; Joseph et al., 2012), LysR family and TetR-family, a MFS superfamily transporter, a Trk system encoding the potassium uptake protein TrkG, and an uncharacterized protein ImpD. A two-component response regulator protein, a two-component system sensor protein, three uncharacterized proteins ImpB, ImpC, and ImpJ/VasE, a glutathione S-transferase protein, a membrane protein, suppressor for copper-sensitivity ScsB, a hypothetical ABC transport system, periplasmic component, a RND efflux transporter, a suppression of copper sensitivity: putative copper binding protein ScsA were shared between pSP291-1 and pESA3, but not pCTU1, which confirmed the findings from previous studies (Kucerova et al., 2010; Joseph et al., 2012). In plasmid group 2, 15 heavy metal (copper, cobalt, zinc, cadmium, lead, and mercury) resistance genes were shared by both plasmids (Figure A3). An osmosensitive K+ channel histidine kinase protein (KdpD), and a virulence-associated protein vagC were also present in both plasmids. PCR analysis confirmed the presence of a pCTU3 IncH1-like origin of replication gene, repA in C. sakazakii SP291 (data not shown).
Interestingly, pSP291-1 contained two unique proteins, a histone acetyltransferase HPA2 and related acetyltransferases protein, along with an uncharacterized protein ImpH/VasB. Six specific proteins were found in pSP291-2, which included a putative glutathione S-transferase protein, a LysR family transcriptional regulator, a putative phage-associated acyl carrier protein, a S-adenosylmethionine: tRNA ribosyltransferase-isomerase protein, permeases of the major facilitator superfamily and an abortive infection protein. Various pSP291-3 proteins including mobilization proteins MobB, MobC, MobD, and DNA relaxase MbeA, which were not shared with any of the other plasmids, were also identified.
Comparative phenotypic profiling of C. sakazakii ATCC BAA-894 and SP291
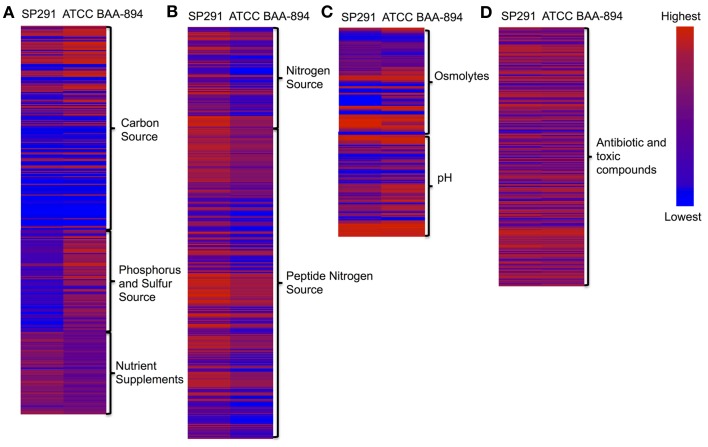
The phenotypic microarray (PM) platform was used previously to support the re-classification of this bacterial genus (Iversen et al., 2008). By comparing the phenotypes of C. sakazakii ATCC BAA-894 and C. sakazakii SP291 expressed across the complete array, interesting differences were observed and these were subsequently assessed in light of comparisons made at the genome level despite of the same PIF orgin. General differences, noted in the phenotypes between the two strains were described in the form of a heat map shown in Figure 2 (the corresponding numerical data is shown in Table S6). Phenotypic differences based on the bacteria's ability to utilize carbon, nitrogen, phosphorous and sulfur sources, as well as other nutrient supplements were noted. Furthermore, growth responses to osmolytes and different pH growth environments, as measured by the array were also observed for both strains. Antibiotic resistance patterns and the ability to respire in the presence of toxic compounds differed.
Figure 2.
(A–D) Heat map of phenotype microarray (PM). Left lane, C. sakazakii SP291; Right lane, C. sakazakii ATCC BAA-894.
Carbon, nitrogen, phosphorous, and sulfur, other nutrient supplement metabolite utilization
Bacteria require a sufficient supply of various biomolecules to support their metabolic activity. In natural environments, where these bacteria are often found, it is to be expected that only limited amounts of these nutrients may be available. To support efficient adaptation and to enable growth in these conditions, bacteria will evolve redundant metabolic systems to support the utilization of a broad range of different substrates, with varying efficiencies. The PM array data gives an insight into how these features differ, between C. sakazakii SP291 and C. sakazakii ATCC BAA-894.
A number of phenotypic differences based on their ability to utilize a range of carbon sources were noted (Figure 2A and Table S6). When compared with C. sakazakii ATCC BAA-894, C. sakazakii SP291 could grow faster in m-inositol and slower in succinic acid, dulcitol, D,L-α-glycerol phosphate, D,L-malic acid, Tween 20, α-ketoglutaric acid, uridine, bromosuccinic acid, glycolic acid, inosine, and dextrin. In contrast there were little or no differences in growth rates when other carbon sources such as methyl pyruvate, mannose, and β-methyl-D-glucuronic acid were compared.
Differences in phenotypes based on the metabolism of carbon sources were compared at the genome level within the carbohydrate subsystem (Table S7). Interestingly, nine inositol catabolism genes were annotated in the C. sakazakii SP291 genome (Table S8), which supported the PM data. Furthermore, a pentose phosphate pathway gene, a lactose utilization gene, and a sucrose utilization gene were also annotated in the C. sakazakii SP291 genome specifically although no evidence to support their activity was found following PM analysis. Similarly, a maltose and maltodextrin utilization gene and a lactate utilization gene were annotated in C. sakazakii ATCC BAA-894 alone, with supporting evidence for the activity lacking from the PM array data. In all, 428 annotated genes related to carbon metabolism were shared between C. sakazakii ATCC BAA-894 and C. sakazakii SP291, which included 10 chitin and N-acetylglucosamine utilization genes, five fructoselysine (amadori product) utilization pathway genes, five dehydrogenase complexes genes, a dihydroxyacetone kinases gene, 14 Entner-Doudoroff pathway genes, and others.
Dancer et al. (2009a,b) reported that for Cronobacter species the availability and utilization of a nitrogen source was an important determinant for biofilm formation when growing in skim milk, and that strong biofilm formers were responsible for coagulation of skim milk (Dancer et al., 2009a). Data from the phenotypic microarray, showed no differences in nitrogen metabolism when C. sakazakii ATCC BAA-894 and C. sakazakii SP291 were compared (Figure 2B and Table S6). Interestingly, when regions of these two genomes known to encode genes associated with nitrogen metabolism were compared, a 16-kb locus, consisting of eight genes was found to be absent in C. sakazakii SP291 compared to C. sakazakii ATCC BAA-894 (Table 3). BLAST analysis of the region facilitated the identification of the corresponding genes located at this position. This locus, contained two nitrate transport proteins NrtB and NrtC, two nitrite reductase proteins NasB and NasA, a respiratory nitrate reductase NarL, a nitrate/nitrite-sensing protein NarX, a nitrite extrusion protein 1 NarK, and a nitrate reductase 1, alpha subunit NarG. This region was also present in C. sakazakii ES15 and C. turicensis z3032. Furthermore, 24 nitrate genes were broadly shared between both of the genomes, which was supported by data from the PM analysis.
Table 3.
Genes associated with nitrogen metabolism, comparing C. sakazakii SP291 and C. sakazakii ATCC BAA 894.
| Function | Gene | Size (bp) | C. sakazakii SP291 | C. sakazakii ATCC BAA-894 |
|---|---|---|---|---|
| Nitrite-sensitive transcriptional repressor NsrR | nsrR | 426 | 166,807…167,232 | 171,773…172,198 |
| Nitrogen regulatory protein P-II | glnB | 339 | 729,723…730,061 | 676,465…676,803 |
| Flavohemoprotein (Hemoglobin-like protein) (Flavohemoglobin) (Nitric oxide dioxygenase) | hmp | 1191 | 731,488…730,298 | 677,084…678,274 |
| Nitrate/nitrite response regulator protein | narP | 645 | 809,507…810,151 | 756,306…756,950 |
| Nitrogen assimilation regulatory protein Nac | nac | 918 | 1,231,697…1,232,614 | 1,195,036…1,195,953 |
| Response regulator NasT | nasT | 1203 | 1,528,174…1,529,376 | 1,450,050…1,451,252 |
| Nitrate ABC transporter, nitrate-binding protein | nrtA | 1251 | 1,530,111…1,531,361 | 1,451,992…1,453,242 |
| Nitrate ABC transporter, permease protein | nrtB | 881 | Absent | 1,453,252…1,454,133 |
| Nitrate ABC transporter, ATP-binding protein | nrtC | 788 | Absent | 1,454,143…1,454,931 |
| Nitrite reductase [NAD(P)H] large subunit | nasB | 4067 | Absent | 1,454,941…1,459,008 |
| Assimilatory nitrate reductase large subunit | nasA | 2702 | Absent | 1,459,005…1,461,707 |
| Nitrate/nitrite response regulator protein | narL | 651 | Absent | 1,463,744…1,464,394 |
| Nitrate/nitrite sensor protein | narX | 1809 | Absent | 1,464,387…1,466,195 |
| Nitrite extrusion protein 1 | narK | 1407 | Absent | 1,466,493…1,467,899 |
| Respiratory nitrate reductase alpha chain | narG | 3747 | Absent | 1,468,317…1,472,063 |
| Respiratory nitrate reductase beta chain | narH | 1311 | 1,531,371…1,532,681 | 1,472,060…1,473,595 |
| Respiratory nitrate reductase delta chain | narJ | 711 | 1,532,678…1,533,388 | 1,473,592…1,474,302 |
| Respiratory nitrate reductase gamma chain | narI | 678 | 1,533,388…1,534,065 | 1,474,302…1,474,979 |
| Oxygen-insensitive NAD(P)H nitroreductase/Dihydropteridine reductase | nfnB | 645 | 1,851,923…1,852,567 | 1,807,325…1,807,969 |
| ABC-type nitrate/sulfonate/bicarbonate transport systems, periplasmic components | 1161 | 1,856,747…1,857,907 | 1,812,312…1,813,310 | |
| Fumarate and nitrate reduction regulatory protein | fnr | 753 | 1,932,047…1,932,799 | 1,887,404… 1,888,156 |
| Hydroxylamine reductase | hcp | 1653 | 2,402,562…2,404,214 | 2,413,065…2,414,717 |
| NADH oxidoreductase hcr | hcr | 969 | 2,404,225…2,405,193 | 2,414,728… 2,415,696 |
| Oxygen-insensitive NADPH nitroreductase | nfsA | 723 | 2,428,664…2,429,386 | 2,434,920…2,434,198 |
| Nitrilotriacetate monooxygenase component B | 618 | 2,510,602…2,511,219 | 2,487,772…2,488,389 | |
| Nitrogen regulatory protein P-II | glnK | 339 | 2,833,540…2,833,878 | 2,792,459…2,792,797 |
| PTS system nitrogen-specific IIA component, PtsN | ptsN | 534 | 3,536,000…3,536,533 | 3,531,047…3,531,580 |
| Phosphocarrier protein, nitrogen regulation associated | ptsO | 273 | 3,537,449…3,537,721 | 3,532,496…3,532,768 |
| Nitrogen regulation protein NR(I) | glnG | 1410 | 4,009,029…4,010,438 | 3,993,113…3,994,522 |
| Nitrogen regulation protein NR(II) | glnL | 1050 | 4,010,447…4,011,496 | 3,994,531…3,995,580 |
| Nitrite reductase [NAD(P)H] small subunit | nirD | 327 | 4,309,983…4,310,309 | 4,334,267…4,334,593 |
| Nitrite reductase [NAD(P)H] large subunit | nirB | 2547 | 4,310,306…4,312,852 | 4,334,590…4,337,136 |
C. sakazakii SP291 was found to grow significantly slower in minimal media supplemented with phosphorous containing compounds (Figure 2A and Table S6), particularly in O-phospho-D-tyrosine, phospho-L-arginine, D,L-α-glycerol phosphate, β-glycerol phosphate, phosphoryl choline, phosphoenol pyruvate, D-glucose-6-phosphate, adenosine 3′-monophosphate, guanosine 2′-monophosphate, guanosine 3′-monophosphate, guanosine 5′-monophosphate, guanosine 2′,3′-cyclic monophosphate, cytidine 2′-monophosphate, cytidine 3′-monophosphate, thymidine 5′-monophosphate, and uridine 5′-monophosphate. Genome annotation provided a conflicting view as determined by the genes identified (Table S9). Twenty-nine phosphorus metabolism genes were broadly shared between C. sakazakii SP291 and C. sakazakii ATCC BAA-894, including eight high affinity phosphate transporters and control of pho-related regulon genes, 18 phosphate metabolism genes, and three polyphosphate genes. Cronobacter species cultured from a PIF production site were compared for their ability to grow in different food matrices (Cooney, 2012). Some demonstrated a slower growth rate compared to others, a feature that might contribute to their enhanced survival in this environment.
No differences in the metabolism of sulfur containing compounds were observed following a comparison of these strains after PM analysis (Figure 2A and Table S6). Forty-nine sulfur metabolism genes were shared by C. sakazakii ATCC BAA-894 and C. sakazakii SP291 (Table S10). These consisted of 17 inorganic sulfur assimilation genes, eight alkanesulfonate assimilation genes, five alkanesulfonates utilization genes, six L-cystine uptake and metabolism genes, four taurine utilization genes, three galactosylceramide and sulfatide metabolism genes, and six thioredoxin-disulfide reductase genes.
Iron is an essential nutrient for bacterial growth and the process of iron acquisition is generally thought to be a prerequisite for a pathogen to establish an infection when entering a host, a feature previously reported in Cronobacter species (Crosa and Walsh, 2002; Franco et al., 2011; Grim et al., 2012). High-affinity iron binding molecules, such as siderophores, and specific iron transport systems function to sequester iron from the environment when bacteria are subjected to iron-limiting growth conditions (Grim et al., 2012). Interestingly, analysis of the PM data showed no major differences between C. sakazakii SP291 and C. sakazakii ATCC BAA-894, in terms of their metabolism of iron or other nutrient supplements. Several transport systems were annotated in C. sakazakii SP291, and which are shared with C. sakazakii ATCC BAA-894 (Kucerova et al., 2010; Joseph et al., 2012), including a ferric hydroxamate ABC transporter denoted as FhuCDBA, 16 ferric enterobactin transporter proteins (including EntA, EntE, EntD, EntB, Fes, EntS, EntF, YbdZ, FepC, FepD, FepG, FepE, FepB, EntC, FepA2, and EntH), a ferrous iron transporter EfeUOB, along with a hemin transporter system, including a ferric reductase protein FhuF and a periplasmic binding protein TonB. A gene summary of iron acquisition and metabolism markers in C. sakazakii SP291 chromosome is shown in Table S11.
Additionally, iron acquisition and metabolism genes were also identified on pSP291-1 (Table S5), which were indistinguishable from that previously reported to be present on pESA3 of C. sakazakii ATCC BAA-894 (Kucerova et al., 2010; Joseph et al., 2012) and pCTU1 of C. turicensis z3032 (Franco et al., 2011; Grim et al., 2012). Target genes from previous reports, such as the RepFIB-like origin of replication gene repA, two plasmid-borne iron acquisition systems (eitCBAD and iucABCD/iutA), as well as the Cronobacter plasminogen activator cpa gene were all present in pSP291-1, with no evidence of the 17-kb type VI secretion system (T6SS) locus identified previously in pESA3 along with a 27-kb region encoding a filamentous hemagglutinin gene (fhaB), its specifc transporter gene (fhaC), and associated putative adhesins (FHA locus) identified in pCTU1 (Kucerova et al., 2010; Franco et al., 2011; Grim et al., 2012, 2013; Joseph et al., 2012). These features support the hypothesis that these plasmids have evolved from a single archetypal backbone that included an iron acquisition system. Our sequence analysis and those of other groups (Joseph et al., 2012; Grim et al., 2013) did not find evidence of plasmid mobilization genes associated with the several plasmid group 1 genomes.
Osmolyte tolerance and survival in different pH environments
When present in different environments, bacteria must develop strategies that promote their survival. Genetic adaptation is derived from modifications of gene expression, via mutations, the acquisition of new and beneficial gene traits, or when these new traits are brought under control of a regulator that was already present in the core genome of the organism's ancestor (Maurelli, 2007). The outcome is that the organism is now better equipped to survive within the new ecological niche. It is generally thought that genes that are no longer compatible with the new lifestyle are selectively inactivated either by point mutation, insertion, or deletion and the contribution of gene loss to an organism's evolution is only now beginning to be appreciated (Maurelli, 2007). Based on our understanding of Cronobacter species epidemiology, these organisms are considered as environmental bacteria. Therefore their ability to survive adverse conditions would be critical. Phenotypes associated with growth in a range of osmolytes and in different pH growth environments were measured by PM analysis (Figure 2C and Table S6) as an indirect reflection of challenging environmental niches. In response to the presence of osmolytes, C. sakazakii SP291 could tolerate 100 mM sodium nitrate compared with C. sakazakii ATCC BAA-894. In contrast, the former grew slower in solutions containing 5% NaCl, 4% potassium chloride, 4% urea, 4–11% sodium lactate, 200 mM sodium phosphate at pH 7 and 20 mM sodium benzoate at pH 5.2. These observations are consistent with what has been suggested previously, in that when a selected adaptation event occurs, and the bacterium enters a new environment such as the human host, phenotypes change (Tall, unpublished observations). Comparing the ability of the environmental isolate C. sakazakii SP291 to survive over a range of different pH growth conditions with that of the PIF isolate C. sakazakii ATCC BAA-894, the former grew faster in a growth condition of pH 9.5 with phenylethylamine, whilst its growth was slower in pH 4.5 with L-proline. This example demonstrates the gain of one phenotype consistent with the inability to survive in the human host (ability to survive in high pH growth conditions) compared to the loss of a sufficient acid resistance response. In this case, the pathoadaptative event that resulted in an increased persistence in the environment comes at the expense of decreased commensal fitness of the microbe (a patent acid response) to survive the acidity of the stomach. However, a greater number of genomes and strains should be evaluated to rule out strain to strain variation.
Annotation of the genome suggested that C. sakazakii SP291 contained a repertoire of genes that could function to aid survival under stressed conditions, such as osmolyte tolerance and different pH environments (Table 4). One hundred and fifty-two annotated genes were identified as being involved with various stress responses. Their presence in the genome may provide early insights into how C. sakazakii SP291 adapts to and survives under different stressful growth conditions.
Table 4.
A selection of the stress response-encoding genes, the defined sub-system, together with the gene name, length of the ORF and correspondoing function, identified in C. sakazakii SP291.
| Category | Sub-system | Gene | Size (bp) | Function |
|---|---|---|---|---|
| Osmotic stress | Choline and betaine uptake and betaine biosynthesis | betB | 1472 | Betaine aldehyde dehydrogenase |
| Osmotic stress | Choline and betaine uptake and betaine biosynthesis | betA | 1679 | Choline dehydrogenase |
| Osmotic stress | Choline and betaine uptake and betaine biosynthesis | betI | 608 | HTH-type transcriptional regulator BetI |
| Osmotic stress | Choline and betaine uptake and betaine biosynthesis | betT | 2030 | High-affinity choline uptake protein BetT |
| Osmotic stress | Choline and betaine uptake and betaine biosynthesis | opuCA | 1145 | Glycine betaine/carnitine/choline transport ATP-binding protein OpuCA |
| Osmotic stress | Choline and betaine uptake and betaine biosynthesis | opuCB | 647 | Glycine betaine/carnitine/choline transport ATP-binding protein OpuCB |
| Osmotic stress | Choline and betaine uptake and betaine biosynthesis | opuCC | 905 | Glycine betaine/carnitine/choline transport ATP-binding protein OpuCC |
| Osmotic stress | Choline and betaine uptake and betaine biosynthesis | opuCD | 713 | Glycine betaine/carnitine/choline transport ATP-binding protein OpuCD |
| Osmotic stress | Choline and betaine uptake and betaine biosynthesis | proP | 1506 | L-Proline/Glycine betaine transporter ProP |
| Osmotic stress | Choline and betaine uptake and betaine biosynthesis | proV | 1202 | L-Proline/Glycine betaine ABC transport system permease protein ProV |
| Osmotic stress | Choline and betaine uptake and betaine biosynthesis | proW | 1070 | L-proline glycine betaine ABC transport system permease protein ProW |
| Osmotic stress | Choline and betaine uptake and betaine biosynthesis | proX | 995 | L-proline glycine betaine binding ABC transporter protein ProX |
| Osmotic stress | Osmoprotectant ABC transporter YehZYXW of enterobacteriales | yehX | 941 | Osmoprotectant ABC transporter ATP-binding subunit YehX |
| Osmotic stress | Osmoprotectant ABC transporter YehZYXW of enterobacteriales | yehZ | 908 | Osmoprotectant ABC transporter binding protein YehZ |
| Osmotic stress | Osmoprotectant ABC transporter YehZYXW of enterobacteriales | yehW | 731 | Osmoprotectant ABC transporter inner membrane protein YehW |
| Osmotic stress | Osmoprotectant ABC transporter YehZYXW of enterobacteriales | yehY | 1133 | Osmoprotectant ABC transporter permease protein YehY |
| Osmotic stress | Osmoregulation | aqpZ | 695 | Aquaporin Z |
| Osmotic stress | Osmoregulation | glpF | 848 | Glycerol uptake facilitator protein |
| Osmotic stress | Osmoregulation | osmY | 614 | Osmotically inducible protein OsmY |
| Osmotic stress | Osmoregulation | ompA | 1076 | Outer membrane protein A precursor |
| Osmotic stress | Osmoregulation | yiaD | 662 | Inner membrane lipoprotein yiaD |
| Osmotic stress | Osmotic stress cluster | yciM | 1169 | Heat shock (predicted periplasmic) protein YciM, precursor |
| Osmotic stress | Osmotic stress cluster | osmB | 215 | Osmotically inducible lipoprotein B precursor |
| Osmotic stress | Osmotic stress cluster | pgpB | 764 | Phosphatidylglycerophosphatase B |
| Osmotic stress | Osmotic stress cluster | yciT | 800 | Transcriptional regulatory protein YciT |
| Osmotic stress | Synthesis of osmoregulated periplasmic glucans | mdoH | 2528 | Glucans biosynthesis glucosyltransferase H |
| Osmotic stress | Synthesis of osmoregulated periplasmic glucans | mdoC | 1157 | Glucans biosynthesis protein C |
| Osmotic stress | Synthesis of osmoregulated periplasmic glucans | mdoD | 1715 | Glucans biosynthesis protein D precursor |
| Osmotic stress | Synthesis of osmoregulated periplasmic glucans | mdoG | 1553 | Glucans biosynthesis protein G precursor |
| Osmotic stress | Synthesis of osmoregulated periplasmic glucans | opgC | 1220 | OpgC protein |
| Osmotic stress | Synthesis of osmoregulated periplasmic glucans | mdoB | 2294 | Phosphoglycerol transferase I |
| Cold shock | Cold shock, CspA family of proteins | cspA | 212 | Cold shock protein CspA |
| Cold shock | Cold shock, CspA family of proteins | cspC | 209 | Cold shock protein CspC |
| Cold shock | Cold shock, CspA family of proteins | cspD | 230 | Cold shock protein CspD |
| Cold shock | Cold shock, CspA family of proteins | cspE | 209 | Cold shock protein CspE |
| Cold shock | Cold shock, CspA family of proteins | cspG | 212 | Cold shock protein CspG |
| Heat shock | Heat shock dnaK gene cluster extended | dnaJ | 1139 | Chaperone protein DnaJ |
| Heat shock | Heat shock dnaK gene cluster extended | dnaK | 1700 | Chaperone protein DnaK |
| Heat shock | Heat shock dnaK gene cluster extended | yggX | 275 | FIG001341:,Probable Fe(2+)-trafficking protein YggX |
| Heat shock | Heat shock dnaK gene cluster extended | gshB | 947 | Glutathione synthetase |
| Heat shock | Heat shock dnaK gene cluster extended | grpE | 602 | Heat shock protein GrpE |
| Heat shock | Heat shock dnaK gene cluster extended | rdgB | 593 | Nucleoside 5-triphosphatase RdgB (dHAPTP, dITP, XTP-specific) |
| Heat shock | Heat shock dnaK gene cluster extended | rpoH | 857 | RNA polymerase sigma factor RpoH |
| Heat shock | Heat shock dnaK gene cluster extended | hemN2 | 1136 | Radical SAM family enzyme, similar to coproporphyrinogen III oxidase, oxygen-independent, clustered with nucleoside-triphosphatase RdgB |
| Heat shock | Heat shock dnaK gene cluster extended | rph | 635 | Ribonuclease PH |
| Heat shock | Heat shock dnaK gene cluster extended | rsmE | 731 | 16S rRNA methyltransferase RsmE |
| Heat shock | Heat shock dnaK gene cluster extended | prmA | 881 | Ribosomal protein L11 methyltransferase |
| Heat shock | Heat shock dnaK gene cluster extended | hslR | 401 | Ribosome-associated heat shock protein implicated in the recycling of the 50S subunit (S4 paralog) |
| Heat shock | Heat shock dnaK gene cluster extended | lepA | 1799 | Translation elongation factor LepA |
| Heat shock | Heat shock dnaK gene cluster extended | yraL | 860 | rRNA small subunit methyltransferase I |
| Heat shock | Heat shock dnaK gene cluster extended | smpB | 482 | tmRNA-binding protein SmpB |
| Dessication stress | O-antigen capsule important for environmental persistence | yihT | 875 | Aldolase YihT |
| Dessication stress | O-antigen capsule important for environmental persistence | yihS | 1241 | Aldose-ketose isomerase YihS |
| Dessication stress | O-antigen capsule important for environmental persistence | yihQ | 2030 | Alpha-glucosyltransferase YihQ |
| Dessication stress | O-antigen capsule important for environmental persistence | yihW | 806 | DeoR-type transcriptional regulator YihW |
| Dessication stress | O-antigen capsule important for environmental persistence | yihO | 1430 | Glucuronide transport protein YihO |
| Dessication stress | O-antigen capsule important for environmental persistence | yihP | 1406 | Glucuronide transport protein YihP, homologous to YihO |
| Dessication stress | O-antigen capsule important for environmental persistence | yshA | 686 | Outer membrane sugar transport protein YshA |
| Dessication stress | O-antigen capsule important for environmental persistence | yihU | 899 | Oxidoreductase YihU |
| Dessication stress | O-antigen capsule important for environmental persistence | yihV | 899 | Sugar kinase YihV |
| Dessication stress | O-antigen capsule important for environmental persistence | yihR | 866 | Sugar-1-epimerase YihR |
| Detoxification | D-tyrosyl-tRNA(Tyr) deacylase | dtd | 437 | D-tyrosyl-tRNA(Tyr) deacylase |
| Detoxification | Glutathione-dependent pathway of formaldehyde detoxification | frmA | 1118 | S-(hydroxymethyl)glutathione dehydrogenase |
| Detoxification | Glutathione-dependent pathway of formaldehyde detoxification | yieG | 830 | S-formylglutathione hydrolase YeiG |
| Detoxification | Tellurite resistance: chromosomal determinants | ydsK | 980 | Uncharacterized acetyltransferase ydcK |
| Detoxification | Tellurite resistance: chromosomal determinants | tehB | 593 | Tellurite resistance protein TehB |
| Detoxification | Tellurite resistance: chromosomal determinants | ydcL | 668 | Uncharacterized lipoprotein ydcL |
| Detoxification | Uptake of selenate and selenite | dedA | 659 | DedA protein |
| Detoxification | Uptake of selenate and selenite | cysA | 1094 | Sulfate and thiosulfate import ATP-binding protein CysA |
| Detoxification | Uptake of selenate and selenite | tsgA | 1187 | TsgA protein homolog |
| Oxidative stress | Glutaredoxins | yebA | 1331 | Uncharacterized metalloprotease yebA |
| Oxidative stress | Glutaredoxins | yibP | 1259 | Uncharacterized protein yibP |
| Oxidative stress | Glutaredoxins | hmp | 1190 | Flavohemoprotein (Hemoglobin-like protein) (Flavohemoglobin) (Nitric oxide dioxygenase) |
| Oxidative stress | Glutaredoxins | grxB | 647 | Glutaredoxin 2 |
| Oxidative stress | Glutaredoxins | grxC | 251 | Glutaredoxin 3 (Grx3) |
| Oxidative stress | Glutaredoxins | nrdH | 245 | Glutaredoxin-like protein NrdH, required for reduction of Ribonucleotide reductase class Ib |
| Oxidative stress | Glutaredoxins | grlA | 347 | Probable monothiol glutaredoxin GrlA |
| Oxidative stress | Glutathione: biosynthesis and gamma-glutamyl cycle | ggt | 1766 | Gamma-glutamyltranspeptidase |
| Oxidative stress | Glutathione: biosynthesis and gamma-glutamyl cycle | gshA | 1556 | Glutamate-cysteine ligase |
| Oxidative stress | Glutathione: biosynthesis and gamma-glutamyl cycle | gshB | 947 | Glutathione synthetase |
| Oxidative stress | Glutathione: non-redox reactions | rnhA | 716 | FIG005121: SAM-dependent methyltransferase |
| Oxidative stress | Glutathione: non-redox reactions | gst1 | 668 | Glutathione S-transferase |
| Oxidative stress | Glutathione: non-redox reactions | yghU | 866 | Uncharacterized Glutathione S-transferase like protein yghU |
| Oxidative stress | Glutathione: non-redox reactions | gst | 608 | Glutathione S-transferase |
| Oxidative stress | Glutathione: non-redox reactions | yqjG | 986 | Uncharacterized protein yqjG |
| Oxidative stress | Glutathione: non-redox reactions | gloB | 755 | Hydroxyacylglutathione hydrolase |
| Oxidative stress | Glutathione: non-redox reactions | gloA | 407 | Lactoylglutathione lyase |
| Oxidative stress | Glutathione: non-redox reactions | yfcF | 644 | Probable glutathione S-transferase, YfcF homolog |
| Oxidative stress | Glutathione: non-redox reactions | yfcG | 626 | Probable glutathione S-transferase, YfcG homolog |
| Oxidative stress | Glutathione: non-redox reactions | yibF | 608 | Uncharacterized GST-like protein yibF |
| Oxidative stress | Glutathione: non-redox reactions | yliJ | 626 | Uncharacterized glutathione S-transferase-like protein |
| Oxidative stress | Glutathione: redox cycle | grxB | 635 | Glutaredoxin 2 |
| Oxidative stress | Glutathione: redox cycle | grxC | 251 | Glutaredoxin 3 (Grx3) |
| Oxidative stress | Glutathione: redox cycle | nrdH | 245 | Glutaredoxin-like protein NrdH, required for reduction of Ribonucleotide reductase class Ib |
| Oxidative stress | Glutathione: redox cycle | btuE | 551 | Glutathione peroxidase |
| Oxidative stress | Glutathione: redox cycle | lpd | 1427 | Glutathione reductase |
| Oxidative stress | Glutathione: redox cycle | gor | 1352 | Glutathione reductase |
| Oxidative stress | Glutathionylspermidine and Trypanothione | yjfC | 1187 | Uncharacterized protein yjfC |
| Oxidative stress | Glutathionylspermidine and Trypanothione | ygiC | 1160 | Uncharacterized protein ygiC |
| Oxidative stress | NADPH:quinone oxidoreductase 2 | ytfG | 854 | Uncharacterized oxidoreductase ytfG |
| Oxidative stress | NADPH:quinone oxidoreductase 2 | qorR | 380 | Redox-sensing transcriptional regulator QorR |
| Oxidative stress | Oxidative stress | katG | 2180 | Catalase/peroxidase HPI |
| Oxidative stress | Oxidative stress | katE | 2255 | Hydroperoxidase II |
| Oxidative stress | Oxidative stress | fur | 452 | Ferric uptake regulation protein FUR |
| Oxidative stress | Oxidative stress | dps | 503 | DNA protection during starvation protein |
| Oxidative stress | Oxidative stress | fnr | 752 | Fumarate and nitrate reduction regulatory protein |
| Oxidative stress | Oxidative stress | oxyR | 917 | DNA-binding transcriptional regulator OxyR |
| Oxidative stress | Oxidative stress | dps | 503 | DNA protection during starvation protein |
| Oxidative stress | Oxidative stress | sodA | 626 | Manganese superoxide dismutase |
| Oxidative stress | Oxidative stress | nsrR | 353 | Nitrite-sensitive transcriptional repressor NsrR |
| Oxidative stress | Oxidative stress | dpS | 503 | Non-specific DNA-binding protein Dps |
| Oxidative stress | Oxidative stress | osmC | 428 | Organic hydroperoxide resistance |
| Oxidative stress | Oxidative stress | ohrR | 548 | Organic hydroperoxide resistance transcriptional regulator |
| Oxidative stress | Oxidative stress | yebS | 1283 | Inner membrane protein yebS |
| Oxidative stress | Oxidative stress | pqiA | 1284 | Paraquat-inducible protein A |
| Oxidative stress | Oxidative stress | yebT | 2633 | Uncharacterized protein yebT |
| Oxidative stress | Oxidative stress | ymbA | 563 | Uncharacterized lipoprotein ymbA |
| Oxidative stress | Oxidative stress | pqiB | 1640 | Paraquat-inducible protein B |
| Oxidative stress | Oxidative stress | katG | 2180 | Catalase/peroxidase HPI |
| Oxidative stress | Oxidative stress | soxR | 458 | Redox-sensitive transcriptional activator SoxR |
| Oxidative stress | Oxidative stress | soxS | 323 | Regulatory protein SoxS |
| Oxidative stress | Oxidative stress | sodC | 518 | Superoxide dismutase [Cu-Zn] precursor |
| Oxidative stress | Oxidative stress | zur | 515 | Zinc uptake regulation protein Zur |
| Oxidative stress | Protection from reactive oxygen species | katG | 2180 | Catalase/peroxidase HPI |
| Oxidative stress | Protection from reactive oxygen species | katE | 2255 | Hydroperoxidase II |
| Oxidative stress | Protection from reactive oxygen species | sodA | 626 | Manganese superoxide dismutase |
| Oxidative stress | Protection from reactive oxygen species | sodC | 518 | Superoxide dismutase [Cu-Zn] precursor |
| Oxidative stress | Redox-dependent regulation of nucleus processes | gapA1 | 995 | NAD-dependent glyceraldehyde-3-phosphate dehydrogenase |
| Oxidative stress | Redox-dependent regulation of nucleus processes | gapA2 | 996 | NAD-dependent glyceraldehyde-3-phosphate dehydrogenase |
| Oxidative stress | Redox-dependent regulation of nucleus processes | npdA | 824 | NAD-dependent protein deacetylase of SIR2 family |
| Oxidative stress | Redox-dependent regulation of nucleus processes | pncA | 641 | Nicotinamidase |
| Oxidative stress | Redox-dependent regulation of nucleus processes | pncB | 1202 | Nicotinate phosphoribosyltransferase |
| Periplasmic stress | Periplasmic stress response | htrA | 1427 | HtrA protease/chaperone protein |
| Periplasmic Stress | Periplasmic stress response | skp | 494 | Outer membrane protein H precursor |
| Periplasmic Stress | Periplasmic stress response | degQ | 1367 | Outer membrane stress sensor protease DegQ, serine protease |
| Periplasmic Stress | Periplasmic stress response | degS | 1067 | Outer membrane stress sensor protease DegS |
| Periplasmic Stress | Periplasmic stress response | rseA | 650 | Sigma factor RpoE negative regulatory protein RseA |
| Periplasmic Stress | Periplasmic stress response | rseB | 854 | Sigma factor RpoE negative regulatory protein RseB precursor |
| Periplasmic Stress | Periplasmic Stress response | surA | 1286 | Survival protein SurA precursor (Peptidyl-prolyl cis-trans isomerase SurA) |
| No subcategory | Bacterial hemoglobins | hmp | 1190 | Flavohemoprotein (Hemoglobin-like protein) (Flavohemoglobin) (Nitric oxide dioxygenase) |
| No subcategory | Carbon starvation | 2105 | Carbon starvation protein A | |
| No subcategory | Carbon starvation | cstA | 2153 | Carbon starvation protein A paralog |
| No subcategory | Carbon starvation | csrA | 185 | Carbon storage regulator |
| No subcategory | Carbon starvation | 584 | Starvation lipoprotein Slp paralog | |
| No subcategory | Carbon starvation | rspA | 1316 | Starvation sensing protein RspA |
| No subcategory | Carbon starvation | sspA | 641 | Stringent starvation protein A |
| No subcategory | Carbon starvation | sspB | 491 | Stringent starvation protein B |
| No subcategory | Commensurate regulon activation | marA | 374 | Multiple antibiotic resistance protein MarA |
| No subcategory | Commensurate regulon activation | gpmB | 869 | Probable phosphoglycerate mutase gpmB |
| No subcategory | Commensurate regulon activation | soxS | 324 | Regulatory protein SoxS |
| No subcategory | Commensurate regulon activation | ramA | 344 | Transcriptional activator RamA |
| No subcategory | Flavohaemoglobin | hmp | 1190 | Flavohemoprotein (Hemoglobin-like protein) (Flavohemoglobin) (Nitric oxide dioxygenase) |
| No subcategory | Hfl operon | hflX | 1280 | GTP-binding protein HflX |
| No subcategory | Hfl operon | hflC | 1004 | HflC protein |
| No subcategory | Hfl operon | hflK | 1244 | HflK protein |
| No subcategory | Hfl operon | yjeT | 197 | Putative inner membrane protein YjeT (clustered with HflC) |
| No subcategory | Hfl operon | hfq | 308 | RNA-binding protein Hfq |
| No subcategory | Phage shock protein (psp) operon | pspA | 671 | Phage shock protein A |
| No subcategory | Phage shock protein (psp) operon | pspB | 224 | Phage shock protein B |
| No subcategory | Phage shock protein (psp) operon | pspC | 356 | Phage shock protein C |
| No subcategory | Phage shock protein (psp) operon | pspD | 242 | Phage shock protein D |
| No subcategory | Phage shock protein (psp) operon | pspF | 1001 | Psp operon transcriptional activator |
| No subcategory | Sugar-phosphate stress regulation | sgrR | 1664 | SgrR, sugar-phosphate stress, transcriptional activator of SgrS small RNA |
| No subcategory | Universal stress protein family | uspA | 437 | Universal stress protein A |
| No subcategory | Universal stress protein family | uspB | 335 | Universal stress protein B |
| No subcategory | Universal stress protein family | uspC | 422 | Universal stress protein C |
| No subcategory | Universal stress protein family | uspE | 956 | Universal stress protein E |
| No subcategory | Universal stress protein family | uspG | 428 | Universal stress protein G |
In recent studies involving Salmonella species, a picture of the transcriptome in low-moisture growth conditions has begun to emerge (Frossard et al., 2012; Finn et al., 2013). Allied to this, 25 genes involved in osmotic stress, and covering 16.4% of the stress response genes were identified in C. sakazakii SP291. Interestingly, the osmoprotectant ABC transporter denoted as YehZYXW in the Cronobacter genome, together with the L-proline glycine betaine MFS transporter ProP, and the ABC transporter ProU systems (composed of ProV, ProW, ProX) designed in Escherichia coli (Checroun and Gutierrez, 2004) and Salmonella Typhimurium (Cairney et al., 1985) were identified in the C. sakazakii SP291 genome. Moreover, an osmoregulator transporter including genes opuCA, opuCB, opuCC, and a fourth gene, which was also an ABC transporter denoted as opuCD here, was 77% similar to that of the osmU operon (osmVWXY) in Salmonella (Frossard et al., 2012) at the gene level. Other osmotically functioning genes identified included the betaine/carnitine/choline transporter (BCCT) family, which acts to transport betaine and choline. This operon consists of a high-affinity choline uptake gene betT, a helix-turn-helix (HTH)-type transcriptional regulator betI, which was previously identified in E. coli (Lamark et al., 1991), a betaine aldehyde dehydrogenase betB gene and a choline dehydrogenase gene betA. An in silico assessment of those loci involved in osomotolerance comparing Cronobacter sakazakii ATCC BAA-894 and E. coli K12 MG1655 identified these latter features also (Feeney and Sleator, 2011). Interestingly, several other genes linked to osmotic stress conditions were identified in the genome of C. sakazakii SP291, which included five osmoregulation genes (aqpZ, glpF, osmY, ompA, and yiaD), four osmotic stress cluster genes (yciM, osmB, pgpB, and yciT) and six osmoregulated periplasmic glucan genes (mdoH, mdoC, mdoD, mdoG, opgC and mdoB). None of these genes were identified previously by Feeney and Sleator (2011). Finally, ompA which encodes an outer membrane porin, was identified in C. sakazakii SP291 and is a recognized virulence marker (Kim et al., 2010).
Experiments to investigate the nature of the C. sakazakii responses to cold- and heat-shock conditions have been reported (Shaker et al., 2008; Carranza et al., 2010; Chang et al., 2010; Al-Nabulsia et al., 2011; Gajdosova et al., 2011). Following exposure to extreme temperatures of cold-shock at −20°C, or heat-shock at 47°C, survival of Cronobacter sakazakii was significantly enhanced (Chang et al., 2010). Carranza et al. (2010) reported that when exposed to higher temperatures, several potential virulence factors were up-regulated. The fact that the pathogenic potential of Cronobacter species may be related to its ability to survive at higher temperatures, warrents further investigation. From the genome sequence of C. sakazakii SP291, the cspA family of cold-shock genes (including cspA, cspC, cspD, cspE and cspG) along with 11 other genes annotated as heat-shock genes, part of the dnaK gene cluster, including dnaJ, dnaK, yggX, gshB, grpE, rdgB, rpoH, hemN2, rph, rsmE, prmA, hslR, yraL, and smpB genes were conserved.
Using a top-down proteomics approach, Williams et al. (2005) identified a candidate protein in C. sakazakii, known to be associated with thermotolerance in Methylobacillus flagelatum, and which was denoted as KT. In a recent study, the genomic region containing this presumptive marker of thermotolerance was compared to similar regions in other bacteria (Gajdosova et al., 2011). An in silico analysis showed that this thermotolerace KT-region was present in 4 of 14 isolates consisting of seven Cronobacter species studied by Joseph et al. (2012). Cronobacter sakazakii SP291 can survive desiccation for long periods of time at an average temperature of 56.7°C, similar to that recorded when spray drying is in operation during PIF production (Cooney, 2012). Interestingly, C. sakazakii SP291 was negative for the KT marker, as determined by PCR (data not shown). Apart from the locus between orfA-orfE, when the corresponding region of the SP291 genome was compared to that of C. sakazakii ATCC 29,544, this region was devoid of the KT-encoding homolog (Figure 3). In light of the thermo-adapted phenotype possessed by C. sakazakii SP291, this finding suggests that there may be other thermotolerance survival mechanisms expressed by C. sakazakii SP291.
Figure 3.
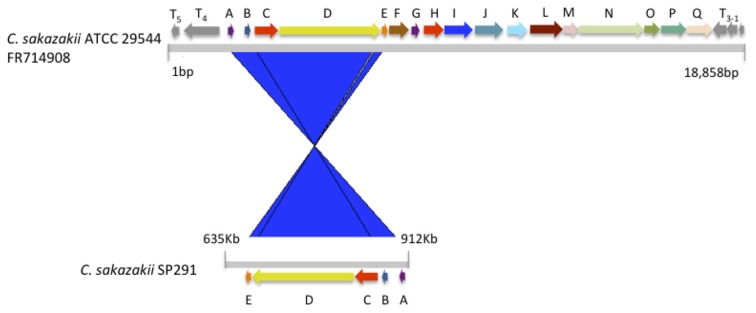
The putative thermotolerance-containing DNA region of C. sakazakii ATCC 29544 compared with C. sakazakii SP291.
As mentioned above, Cronobacter species have the capacity to survive in desiccated environments for long periods, a phenotype that is linked to their epidemiology and routes of infection. As an example of genes linked to this phenotype, the yih-encoding operons, consisted of 10 annotated genes present in the genome of C. sakazakii SP291. Desiccation-related proteins YihU, YihT, YihR, YihS, YihQ and YihV have conserved domains which function in carbohydrate transport and metabolism. YihO is a glucuronide transport protein whilst YihP is homologous to it. YshA is an outer membrane sugar transport protein and YihW is a deoR-type transcriptional regulator, reported to negatively regulate the expression of yihU-oyshA in Salmonella (Gibson et al., 2006). This operon was reported to be up-regulated following desscication stress in Salmonella. Interstingly, the yih operon are conserved in 17 annotated Cronobacter genomes (strain information of the genomes are listed in Table 1) and noted previously (Grim et al., 2013).
The ability of a bacterium to eliminate toxic compounds from the cell is an important survival mechanism. Nine genes involved in detoxification were identified in the C. sakazakii SP291 genome. These included a tellurite resistance-encoding gene tehB, which matches a 593 bp hypothetical protein in C. sakazakii ATCC BAA-894. However, a tellurite resistance region (terACDYZ) was reported only in C. sakazakii ATCC BAA-894 (Table 4), and with the exception of the terC-encoding marker in C. turicensis z3032, was not identified in any of the other Cronobacter species genomes sequenced (Kucerova et al., 2010; Joseph et al., 2012; Grim et al., 2013). Genes involved in the detoxification of organic pollutants, including a D-tyrosyl-tRNA (Tyr) deacylase-encoding dtd, two glutathione-dependent pathway formaldehyde detoxification genes (frmA and yieG), three genes involved in the uptake of selenate and selenite (dedA, cysA, and its homolog tsgA), and two uncharacterized genes (ydsK and ydcL), were also identified in C. sakazakii SP291. This feature supports an earlier report describing the ability of Cronobacter species to detoxify and survive in tannery wastewater effluents (Chandra et al., 2011).
Oxidative stress is an example of an important bacterial stress response, with 62 annotated genes covering 40.8% genome linked to this sub-system. These genes included the zinc uptake regulation zur, which was reported as involved in the oxidative stress response of Streptomyces coelicolor (Shin et al., 2007). Other stress-related genes included seven periplasmic stress related genes, a bacterial hemoglobin gene, seven genes involved in carbon starvation, four commensurate regulon activation genes, a flavohaemoglobin gene, five hfl operon genes, five phage shock protein (psp) operon genes, a sugar-phosphate stress regulation gene, and five universal stress protein family genes.
Resistance to antibiotics and toxic compounds
C. sakazakii was originally reported to be susceptible to a panel of 69 antimicrobial agents (Stock and Wiedemann, 2002). Subsequently, a tetracycline-resistant C. sakazakii cultured from a Chilean freshwater salmon farm (Miranda et al., 2003) was isolated, followed by a report of a trimethoprim and neomycin resistance isolate cultured from fresh domiati cheese (El-Sharoud et al., 2009). More recently, isolates resistant to cephalothin were recovered from dried food (Chon et al., 2012). The emergence of strains that have become resistant to antimicrobial compounds is of great concern to public health (Dumen, 2010; Yan et al., 2012).
Figure 2D shows a heat map comparing C. sakazakii SP291 and C. sakazakii ATCC BAA-894 and Table 5 provides a summary of the significant changes in PM redox measurements, after bacterial growth in microtitre wells containing a number of antimicrobial compounds as part of the phenotypic microarray. Compared to C. sakazakii ATCC BAA-894, C. sakazakii SP291 exhibited more activity in the presence of 5,7-dichloro-8-hydroxyquinoline, 5-nitro-2-furaldehyde semicarbazone, hexamminecobalt (III) chloride, poly-L-lysine, protamine sulfate, ornidazole, tobramycin, streptomycin, apramycin, iodonitro tetrazolium violet, amoxicilin, neomycin, and sisomicin; while exhibiting a reduced activity in the presence of phleomycin, ciprofloxacin, cinoxacin, dichlofluanid, tolylfluanid, guanidine hydrochloride, colistin, methyl viologen, sodium azide, guanazole, rifamycin SV, glycine hydroxamate, D,L-methionine hydroxamate, cefmetazole, and cloxacillin.
Table 5.
Comparison of the differential phenotypes expressed by C. sakazakii SP291 and C. sakazakii ATCC BAA-894 related to antimicrobial and toxic compounds as determined by phenotype microarray.
| Test compound | Differencea | Mode of action |
|---|---|---|
| PHENOTYPES GAINED BY C. sakazakii SP291 RELATIVE TO C. sakazakii ATCC BAA-894- | ||
| Amoxicillin | 20,446 | Wall, lactam |
| Neomycin | 47,681 | Protein synthesis, 30S ribosomal subunit, aminoglycoside |
| Sisomicin | 40,297 | Protein synthesis, 30S ribosomal subunit, aminoglycoside |
| Tobramycin | 33,297 | Protein synthesis, 30S ribosomal subunit, aminoglycoside |
| Sodium arsenate | 22,155 | Toxic anion, PO4 analog |
| Sodium metaborate | 65,231 | Toxic anion |
| EDTA | 89,134 | Chelator, hydrophilic |
| 5,7-Dichloro-8-hydroxyquinoline | 21,638 | Chelator, lipophilic |
| 5-Nitro-2-furaldehyde semicarbazone | 40,348 | DNA damage, multiple sites, nitrofuran analog |
| Protamine sulfate | 27,720 | Membrane, non-specific binding |
| Streptomycin | 40,297 | Protein synthesis, 30S ribosomal subunit, aminoglycoside |
| Potassium tellurite | 27,025 | Toxic anion |
| Sodium tungstate | 50,849 | Toxic anion, molybdate analog |
| Poly-L-lysine | 43,324 | Membrane, detergent, cationic |
| Sodium m-arsenite | 28,927 | Toxic anion |
| Sodium periodate | 47,820 | Toxic anion, oxidizing agent |
| Antimony (III) chloride | 35,457 | Toxic cation |
| Iodonitro tetrazolium violet | 22,427 | Respiration |
| Hexamminecobalt (III) Chloride | 30,202 | DNA synthesis |
| Apramycin | 43,717 | Protein synthesis, 30S ribosomal subunit, aminoglycoside |
| Ornidazole | 22,594 | Protein glycosolation |
| PHENOTYPES LOST BY C. sakazakii SP291 RELATIVE TO C. sakazakii ATCC BAA-894- | ||
| Cloxacillin | −20,939 | Wall, lactam |
| Colistin | −35,638 | Membrane, transport |
| Guanidine hydrochloride | −47,347 | Membrane, chaotropic agent |
| Cefmetazole | −21,713 | Wall, cephalosporin second generation |
| Phleomycin | −45,183 | DNA damage, oxidative, ionizing ratiation |
| Methyl viologen | −81,565 | Oxidizing agent |
| Sodium azide | −22,375 | Respiration, uncoupler |
| Dichlofluanid | −42,205 | Fungicide, phenylsulphamide |
| Cinoxacin | −18,744 | DNA unwinding, gyrase (GN), topoisomerase (GP), quinolone |
| Rifamycin SV | −22,130 | RNA polymerase |
| Glycine hydroxamate | −64,946 | tRNA synthetase |
| D,L−Methionine hydroxamate | −38,317 | tRNA synthetase |
| Sodium bromate | −27,123 | Toxic anion |
| Guanazole | −33,164 | Ribonucleotide DP reductase |
| Ciprofloxacin | −22,172 | DNA unwinding, gyrase (GN), topoisomerase (GP), fluoroquinolone |
| Tolylfluanid | −18,317 | Fungicide, phenylsulphamide |
Denotes the following: a positive number indicates faster growth in C. sakazakii SP291 compared to C. sakazakii ATCC BAA-894; a negative number indicates faster growth in C. sakazakii ATCC BAA-894 compared to C. sakazakii SP291.
Careful analysis of the PM data showed an interesting phenotype, related to bioactive and toxic anions. C. sakazakii SP291 survived significantly better in sodium metaborate, potassium tellurite, sodium m-arsenite, sodium tungstate, sodium periodate, sodium arsenate, and antimony (III) chloride compared with C. sakazakii ATCC BAA-894. In contrast the latter bacterium, exhibited a distinct phenotype in sodium bromate. These observations suggested that C. sakazakii SP291 elaborates a greater ability to counter the effects of a broader range of heavy metals, a characteristic that could be facilitate adaptation in powered infant formula manufacturing environments where metallic compositions such as quaternary containing disinfectants are used for decontamination. This resistance phenotype may be globally regulated as well. Together, this information may explain how this organism adapted to the manufacturing environment.
Based in part on these phenotypes, a total of 44 genes were shared between C. sakazakii ATCC BAA-894 and C. sakazakii SP291. These consisted of adaptation to D-cysteine related genes (include yecC, yecS, and dcyD), a β-lactamase-encoding ampC gene, three cobalt-zinc-cadmium resistance genes (including feiF, zitB and a MerR family transcriptional regulator), 11 copper homeostasis genes (include cueO, yobA, copA, zntA, ccmF, ccmH, cutE, cutF, corC, cutA, and a copper resistance protein D gene), a fosfomycin resistance gene fosA, a lysozyme inhibitor mliC, a tripartite multidrug resistance system found in Gram-negative bacteria, 11 multidrug resistance efflux pumps (including macB, macA, acrA, acrE, norM, acrD, acrB, acrF, acrR, envR, and tolC), four genes encoding resistance to fluoroquinolones (including gyrA, gyrB, parC, and parE), and a multidrug resistance cluster (consisting of mdtB, mdtC, mdtD, mdtA, baeR, and baeS). All of these genes mapped to the bacterial chromosome. In addition three arsenic resistance genes were identified on pSP291-1 (Table S5). This latter feature confirmed the previous report on the copper/silver resistance determinants in Cronobacter species (Kucerova et al., 2010; Sivamaruthi et al., 2011; Joseph and Forsythe, 2012; Joseph et al., 2012; Grim et al., 2013). In contrast, eight cobalt-zinc-cadmium resistance genes (include cusA, cusC, cusF, czcA, czcB, cusS, cusR, and pcoS) and three copper homeostasis genes (including copG, pcoB and pcoA) were unique to C. sakazakii ATCC BAA-894, of which cusRCFBA/silRECBA and pcoABCDR were indicated as two copper and silver resistance regions. The previous region was shared among C. sakazakii, C. malonaticus, and C. turicensis; while the latter region was shared among C. sakazakii, C. malonaticus, C. turicensis and C. universalis (Joseph et al., 2012) (Table 6).
Table 6.
Genes related to resistance to antibiotics and toxic compounds annotated in C. sakazakii SP291 and C. sakazakii ATCC BAA-894.
| Subsystem | Start | Stop | Size (bp) | Gene | Role |
|---|---|---|---|---|---|
| RESISTANCE TO ANTIBIOTIC AND TOXIC COMPUNDS GENES SHARED BY C. sakazakii SP291 AND C. sakazakii ATCC BAA-894- | |||||
| Adaptation to D-cystine | 1,345,012 | 1,345,764 | 752 | yecC | Cystine ABC transporter, ATP-binding protein |
| Adaptation to d-cystine | 1,344,347 | 1,345,015 | 668 | yecS | Cystine ABC transporter, permease protein |
| Adaptation to d-cystine | 1,343,343 | 1,344,323 | 980 | dcyD | D-cystine desulfhydrase |
| Beta-lactamase | 1,853,746 | 1,852,619 | 1127 | ampC | Beta-lactamase |
| Cobalt-zinc-cadmium resistance | 4,087,380 | 4,088,282 | 902 | fieF | Cobalt-zinc-cadmium resistance protein |
| Cobalt-zinc-cadmium resistance | 733,136 | 733,498 | 362 | Transcriptional regulator, MerR family | |
| Cobalt-zinc-cadmium resistance | 2,570,193 | 2,571,155 | 962 | zitB | Zinc transporter ZitB |
| Copper homeostasis | 2,643,555 | 2,645,219 | 1664 | cueO | Blue copper oxidase CueO precursor |
| Copper homeostasis | 1,433,730 | 1,434,104 | 374 | yobA | Copper resistance protein C precursor |
| Copper homeostasis | 1,434,109 | 1,434,978 | 869 | Copper resistance protein D | |
| Copper homeostasis | 2,766,031 | 2,768,538 | 2507 | copA | Copper-translocating P-type ATPase |
| Copper homeostasis | 4,209,899 | 4,207,683 | 2216 | zntA | Zinc/cadmium/mercury/lead-transporting ATPase |
| Copper homeostasis | 924,321 | 926,270 | 1949 | ccmF | Cytochrome c heme lyase subunit CcmF |
| Copper homeostasis | 926,821 | 927,282 | 461 | ccmH | Cytochrome c heme lyase subunit CcmH |
| Copper homeostasis: copper tolerance | 2,651,936 | 2,653,477 | 1541 | cutE | Copper homeostasis protein CutE |
| Copper homeostasis: copper tolerance | 3,053,870 | 3,053,172 | 698 | cutF | Copper homeostasis protein CutF precursor |
| Copper homeostasis: copper tolerance | 2,651,056 | 2,651,931 | 875 | corC | Magnesium and cobalt efflux protein CorC |
| Copper homeostasis: copper tolerance | 130,149 | 129,802 | 347 | cutA | Periplasmic divalent cation tolerance protein CutA |
| Fosfomycin resistance | 1,712,546 | 1,712,959 | 413 | fosA | Fosfomycin resistance protein FosA |
| Lysozyme inhibitors | 1,973,845 | 1,973,522 | 323 | mliC | Membrane-bound lysozyme inhibitor of c-type lysozyme |
| Multidrug resistance, tripartite systems found in gram negative bacteria | 2,468,114 | 2,466,543 | 1517 | emrB2 | Inner membrane component of tripartite multidrug resistance system |
| Multidrug resistance, tripartite systems found in gram negative bacteria | 2,469,268 | 2,468,111 | 1157 | Membrane fusion component of tripartite multidrug resistance system | |
| Multidrug resistance, tripartite systems found in gram negative bacteria | 2,466,541 | 2,465,000 | 1541 | nodT | Outer membrane component of tripartite multidrug resistance system |
| Multidrug resistance efflux pumps | 2,397,113 | 2,395,170 | 1943 | macB | Macrolide export ATP-binding/permease protein MacB |
| Multidrug resistance efflux pumps | 2,398,222 | 2,397,110 | 1112 | macA | Macrolide-specific efflux protein MacA |
| Multidrug resistance efflux pumps | 2,799,127 | 2,800,332 | 1205 | acrA | Membrane fusion protein of RND family multidrug efflux pump |
| Multidrug resistance efflux pumps | 3,599,205 | 3,600,347 | 1142 | acrE | Membrane fusion protein of RND family multidrug efflux pump |
| Multidrug resistance efflux pumps | 1,999,315 | 2,000,688 | 1373 | norM | Multi antimicrobial extrusion protein (Na(+)/drug antiporter), MATE family of MDR efflux pumps |
| Multidrug resistance efflux pumps | 809,372 | 806,256 | 3116 | acrD | Probable aminoglycoside efflux pump |
| Multidrug resistance efflux pumps | 2,800,354 | 2,803,503 | 3149 | acrB | Acriflavine resistance protein B |
| Multidrug resistance efflux pumps | 3,600,357 | 3,603,476 | 3119 | acrF | Acriflavine resistance protein F |
| Multidrug resistance efflux pumps | 2,798,982 | 2,798,317 | 665 | acrR | HTH-type transcriptional regulator acrR |
| Multidrug resistance efflux pumps | 3,598,827 | 3,598,180 | 647 | envR | Transcription repressor of multidrug efflux pump acrAB operon, TetR (AcrR) family |
| Multidrug resistance efflux pumps | 374,805 | 373,318 | 1487 | tolC | Type I secretion outer membrane protein, TolC precursor |
| Resistance to fluoroquinolones | 1,021,753 | 1,024,389 | 2636 | gyrA | DNA gyrase subunit A |
| Resistance to fluoroquinolones | 3,931,538 | 3,929,124 | 2414 | gyrB | DNA gyrase subunit B |
| Resistance to fluoroquinolones | 383,132 | 385,402 | 2270 | parC | Topoisomerase IV subunit A |
| Resistance to fluoroquinolones | 377,546 | 379,438 | 1892 | parE | Topoisomerase IV subunit B |
| The mdtABCD multidrug resistance cluster | 1,153,120 | 1,149,998 | 3122 | mdtB | Multidrug transporter MdtB |
| The mdtABCD multidrug resistance cluster | 1,149,997 | 1,146,908 | 3089 | mdtC | Multidrug transporter MdtC |
| The mdtABCD multidrug resistance cluster | 1,146,904 | 1,145,489 | 1415 | mdtD | Multidrug transporter MdtD |
| The mdtABCD multidrug resistance cluster | 1,154,367 | 1,153,120 | 1247 | mdtA | Probable RND efflux membrane fusion protein |
| The mdtABCD multidrug resistance cluster | 1,144,071 | 1,143,349 | 722 | baeR | Response regulator BaeR |
| The mdtABCD multidrug resistance cluster | 1,145,492 | 1,144,068 | 1424 | baeS | Sensory histidine kinase BaeS |
| RESISTANCE TO ANTIBIOTIC AND TOXIC COMPUNDS GENES SPECIFIC TO C. sakazakii ATCC BAA-894- | |||||
| Cobalt-zinc-cadmium resistance | 4,206,746 | 4,209,892 | 3146 | cusA | Cation efflux system protein CusA |
| Cobalt-zinc-cadmium resistance | 4,203,563 | 4,204,948 | 1385 | cusC | Cation efflux system protein CusC precursor |
| Cobalt-zinc-cadmium resistance | 4,204,976 | 4,205,329 | 353 | cusF | Cation efflux system protein CusF precursor |
| Cobalt-zinc-cadmium resistance | 4,206,746 | 4,209,892 | 3146 | czcA | Cobalt-zinc-cadmium resistance protein CzcA |
| Cobalt-zinc-cadmium resistance | 4,205,443 | 4,206,735 | 1292 | czcB | Cobalt/zinc/cadmium efflux RND transporter, membrane fusion protein, CzcB family |
| Cobalt-zinc-cadmium resistance | 4,202,505 | 4,201,219 | 1286 | cusS | Copper sensory histidine kinase CusS |
| Cobalt-zinc-cadmium resistance | 4,203,373 | 4,202,693 | 680 | cusR | Copper-sensing two-component system response regulator CusR |
| Cobalt-zinc-cadmium resistance | 4,219,774 | 4,221,174 | 1400 | pcoS | Heavy metal sensor histidine kinase |
| Copper homeostasis | 4,209,979 | 4,210,419 | 440 | CopG | CopG protein |
| Copper homeostasis | 4,217,266 | 4,217,688 | 422 | pcoB | Copper resistance protein B |
| Copper homeostasis | 4,214,975 | 4,216,792 | 1817 | pcoA | Multicopper oxidase |
Conclusions
Stress responses and resistance to antibiotic and toxic compounds are interesting phenotypes identified in C. sakazakii SP291, when compared to C. sakazakii ATCC BAA-894, based on the comparative phenotypic microarray analysis in parallel with the annotation of its genome. Given this the fact that the PIF production environment is a stressful ecological niche, the osmoprotectant ABC transporters including YehZYXW, ProP, ProU, and OpuCABCD can be expected to play a role to support bacterial survival, as reported in other microorganisms previously (Cairney et al., 1985; Checroun and Gutierrez, 2004; Frossard et al., 2012; Finn et al., 2013). Notably, C. sakazakii SP291 possesses a greater ability to survive in a broader range of heavy metals, as quaternary containing disinfectants which include metallic compositions are often used for PIF manufacturing environment disinfection. In conclusion, genome analysis of C. sakazakii SP291 along with its metabolome highlighted a number of potential features, which might be considered as candidates for future studies to extend our understanding of the persistence and virulence mechanisms deployed by this bacterium in the production environment. These data provide an early insight into how a factory isolate may survive in a desiccation condition and adapt to the PIF production environment and associated food matrices.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Drs. Marta Martins and Orla Condell for their assistance with the PM data analysis and to Ms. Sarah Finn for critical reading of the manuscript.
Appendix
Figure A1.
Circular representation of the C. sakazakii SP291 genome. The first circle represents the scale in kilobases starting with the origin of replication at position 0. The second circle shows the distribution of CDS (Goldman and Green, 2009) in forward strand. The third circle indicates the distribution of CDS (blue) in reverse strand. rRNA operons are colored in red on the fourth circle. tRNA operons are colored in purple on the fifth circle. The sixth circle indicates the deviation of the GC content average, with values greater than zero in green and those less than zero in purple. The innermost circle displays the GC skew ([G + C]/[G − C]), with values greater than zero in light blue and those less than zero in orange. The figure was generated using DNAPlotter (Carver et al., 2009).
Figure A2.
Comparision of plasmid group 1 (pESA3, pSP291-1, and pCTU1) using Artemis comparison tool (ACT).
Figure A3.
Comparision of plasmid group 2 (pSP291-2 and pCTU3) using Artemis comparison tool (ACT).
Supplementary material
The Supplementary Material for this article can be found online at: http://www.frontiersin.org/Food_Microbiology/10.3389/fmicb.2013.00256/abstract
Genomic regions missing in C. sakazakii SP291 when compared to the other three completed genomes of C. sakazakii ATCC BAA-894 (as the reference sequence), C. sakazakii E15 and C. turicensis z3032.
Genomic regions present only in C. sakazakii SP291 (as the reference sequence) when compared to the other three completed genomes of C. sakazakii ATCC BAA-894, C. sakazakii ES15 and C. turicensis z3032.
Genome comparison of C. sakazakii SP291 with other available C. sakazakii genomes, which include C. sakazakii SP291, ATCC BAA-894, ES15, E899, 680, 696, 701, ES35, 2151, ES713, and E764.
Genome comparison of C. sakazakii SP291 with Cronobacter isolate of other six species, which consist of C. dublinensis LMG 23823, C. malonaticus LMG 23826, C. muytjensii ATCC 51329, C. turicensis z3032, C. universalis NCTC 9529, and C. condimenti 1330.
Annotated plasmid clusters of three complete genome, which include pSP291-1, pSP291-2, and pSP291-3 of C. sakazakii SP291, pESA2 and pESA3 of C. sakazakii ATCC BAA-894, as well as pCTU1, pCTU2, and pCTU3 of C. turicensis z3032. Plasmid group 1: pSP291-1, pESA3, and pCTU1; plasmid group 2: pSP291-2 and pCTU3.
Numerical data for all 20 PM plates of C. sakazakii SP291 and C. sakazakii ATCC BAA-894, along with the phenotype differences (greater than 20,000±1,800 unit) generated by Omnilog® software.
Genes assigned to defined bacterial sub-systems and their distribution among these functional categories for C. sakazakii SP291 along with other three completed genome, C. sakazakii ATCC BAA-894, C. sakazakii ES15, and C. turicensis z3032.
Genome annotation: Carbohydrate metabolism of C. sakazakii SP291 and C. sakazakii ATCC BAA-894.
Genome annotation: Phosphorus metabolism of C. sakazakii SP291 and C. sakazakii ATCC BAA-894.
Genome annotation: Sulfur metabolism of C. sakazakii SP291 and C. sakazakii ATCC BAA-894.
Iron acquisition systems identified in the chromosome of C. sakazakii SP291.
References
- Al-Nabulsia A. A., Osailia T. M., Zain Elabedeena N. A., Jaradatb Z. W., Shakera R. R., Kheirallahc K. A., et al. (2011). Impact of environmental stress desiccation, acidity, alkalinity, heat or cold on antibiotic susceptibility of Cronobacter sakazakii. Int. J. Food Microbiol. 146, 137–143 10.1016/j.ijfoodmicro.2011.02.013 [DOI] [PubMed] [Google Scholar]
- Arku B., Fanning S., Jordan K. (2011). Heat adaptation and survival of Cronobacter spp. (formerly Enterobacter sakazakii). Foodborne Pathog. Dis. 8, 975–981 10.1089/fpd.2010.0819 [DOI] [PubMed] [Google Scholar]
- Aziz R. K., Bartels D., Best A. A., DeJongh M., Disz T., Edwards R. A., et al. (2008). The RAST Server: rapid annotations using subsystems technology. BMC Genomics 9:75 10.1186/1471-2164-9-75 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bar-Oz B., Preminger A., Peleg O., Block C., Arad I. (2001). Enterobacter sakazakii infection in the newborn. Acta Paediatr. 90, 356–358 10.1080/080352501300067857 [DOI] [PubMed] [Google Scholar]
- Baumgartner A., Grand M., Liniger M., Iversen C. (2009). Detection and frequency of Cronobacter spp. (Enterobacter sakazakii) in different categories of ready-to-eat foods other than infant formula. Int. J. Food Microbiol. 136, 189–192 10.1016/j.ijfoodmicro.2009.04.009 [DOI] [PubMed] [Google Scholar]
- Beuchat L. R., Komitopoulou E., Beckers H., Betts R. P., Bourdichon F., Fanning S., et al. (2013). Low-water activity foods: increased concern as vehicles of foodborne pathogens. J. Food Prot. 76, 150–172 10.4315/0362-028X.JFP-12-211 [DOI] [PubMed] [Google Scholar]
- Bowen A. B., Braden C. R. (2006). Invasive Enterobacter sakazakii disease in infants. Emerg. Infect. Dis. 12, 1185–1189 10.3201/eid1708.051509 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown J. M., Shaw K. J. (2003). A novel family of Escherichia coli toxin-antitoxin gene pairs. J. Bacteriol. 185, 6600–6608 10.1128/JB.185.22.6600-6608.2003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cairney J., Booth I. R., Higgins C. F. (1985). Osmoregulation of gene expression in Salmonella typhimurium: proU encodes an osmotically induced betaine transport system. J. Bacteriol. 164, 1224–1232 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carranza P., Grunau A., Schneider T., Hartmann I., Lehner A., Stephan R., et al. (2010). A gel-free quantitative proteomics approach to investigate temperature adaptation of the food-borne pathogen Cronobacter turicensis 3032. Proteomics 10, 3248–3261 10.1002/pmic.200900460 [DOI] [PubMed] [Google Scholar]
- Carver T., Thomson N., Bleasby A., Berriman M., Parkhill J. (2009). DNAPlotter: circular and linear interactive genome visualization. Bioinformatics 25, 119–120 10.1093/bioinformatics/btn578 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chandra R., Bharagava R. N., Kapley A., Purohit H. J. (2011). Bacterial diversity, organic pollutants and their metabolites in two aeration lagoons of common effluent treatment plant (CETP) during the degradation and detoxification of tannery wastewater. Bioresour. Technol. 102, 2333–2341 10.1016/j.biortech.2010.10.087 [DOI] [PubMed] [Google Scholar]
- Chang C. H., Chiang M. L., Chou C. C. (2010). The effect of heat shock on the response of Cronobacter sakazakii to subsequent lethal stresses. Foodborne Pathog. Dis. 7, 71–76 10.1089/fpd.2009.0345 [DOI] [PubMed] [Google Scholar]
- Chap J., Jackson P., Siqueira R., Gaspar N., Quintas C., Park J., et al. (2009). International survey of Cronobacter sakazakii and other Cronobacter spp. in follow up formulas and infant foods. Int. J. Food Microbiol. 136, 185–188 10.1016/j.ijfoodmicro.2009.08.005 [DOI] [PubMed] [Google Scholar]
- Checroun C., Gutierrez C. (2004). Sigma(s)-dependent regulation of yehZYXW, which encodes a putative osmoprotectant ABC transporter of Escherichia coli. FEMS Microbiol. Lett. 236, 221–226 10.1016/j.femsle.2004.05.046 [DOI] [PubMed] [Google Scholar]
- Chen Y., Strain E. A., Allard M., Brown E. W. (2011). Genome sequence of Cronobacter sakazakii E899, a strain associated with human illness. J. Bacteriol. 193:5861 10.1128/JB.05913-11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chon J. W., Song K. Y., Kim S. Y., Hyeon J. Y., Seo K. H. (2012). Isolation and characterization of Cronobacter from desiccated foods in Korea. J. Food Sci. 77, M354–M358 10.1111/j.1750-3841.2012.02750.x [DOI] [PubMed] [Google Scholar]
- Cooney S. (2012). A Sudy on the Epidemiology and Behavior of Selected Bacteria Colonizing a Podweed Infant Formula (PIF) Low Moisture Production Environment. PhD thesis, University College Dublin, 185.
- Correia F. F., D'Onofrio A., Rejtar T., Li L., Karger B. L., Makarova K., et al. (2006). Kinase activity of overexpressed HipA is required for growth arrest and multidrug tolerance in Escherichia coli. J. Bacteriol. 188, 8360–8367 10.1128/JB.01237-06 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crosa J. H., Walsh C. T. (2002). Genetics and assembly line enzymology of siderophore biosynthesis in bacteria. Microbiol. Mol. Biol. Rev. 66, 223–249 10.1128/MMBR.66.2.223-249.2002 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dancer G. I., Mah J. H., Kang D. H. (2009a). Influences of milk components on biofilm formation of Cronobacter spp. (Enterobacter sakazakii). Lett. Appl. Microbiol. 48, 718–725 [DOI] [PubMed] [Google Scholar]
- Dancer G. I., Mah J. H., Rhee M. S., Hwang I. G., Kang D. H. (2009b). Resistance of Enterobacter sakazakii (Cronobacter spp.) to environmental stresses. J. Appl. Microbiol. 107, 1606–1614 10.1111/j.1365-2672.2009.04347.x [DOI] [PubMed] [Google Scholar]
- Drudy D., O'Rourke M., Murphy M., Mullane N. R., O'Mahony R., Kelly L., et al. (2006). Characterization of a collection of Enterobacter sakazakii isolates from environmental and food sources. Int. J. Food Microbiol. 110, 127–134 10.1016/j.ijfoodmicro.2006.02.008 [DOI] [PubMed] [Google Scholar]
- Dumen E. (2010). Cronobacter sakazakii (Enterobacter sakazakii): only an infant problem. Kafkas Univ. Vet. Fak. Derg. 16, S171–S178 21261874 [Google Scholar]
- El-Sharoud W. M., O'Brien S., Negredo C., Iversen C., Fanning S., Healy B. (2009). Characterization of Cronobacter recovered from dried milk and related products. BMC Microbiol. 9:24 10.1186/1471-2180-9-24 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Farmer J. J., 3rd, Asbury M. A., Hickman F. W., Brenner D. J. (1980). Enterobacter sakazakii A new species of Enterobacteriaceae isolate from clinical specime. Int. J. Syst. Evol. Microbiol. 30, 569–584 [Google Scholar]
- Feeney A., Sleator R. D. (2011). An in silico analysis of osomotolerancec in the emerging gastrointestinal pathogen Cronobacter sakazakii. Bioeng. Bugs 2, 260–270 10.4161/bbug.2.5.17238 [DOI] [PubMed] [Google Scholar]
- Finn S., Handler K., Condell O., Colgan A., Cooney S., Hinton J., et al. (2013). ProP is required for the survival of Salmonella Typhimurium desiccated on stainless steel. Appl. Environ. Microbiol. 79, 4376–4384 10.1128/AEM.00515-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Franco A. A., Hu L., Grim C. J., Gopinath G., Sathyamoorthy V., Jarvis K. G., et al. (2011). Characterization of putative virulence genes on the related RepFIB plasmids harbored by Cronobacter spp. Appl. Environ. Microbiol. 77, 3255–3267 10.1128/AEM.03023-10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Friedemann M. (2009). Epidemiology of invasive neonatal Cronobacter (Enterobacter sakazakii) infections. Eur. J. Clin. Microbiol. 28, 1297–1304 dio: 10.1007/s10096-009-0779-4 10.1007/s10096-009-0779-4 [DOI] [PubMed] [Google Scholar]
- Frossard S. M., Khan A. A., Warrick E. C., Gately J. M., Hanson A. D., Oldham M. L., et al. (2012). Identification of a third osmoprotectant transport system, the osmU system, in Salmonella enterica. J. Bacteriol. 194, 3861–3871 10.1128/JB.00495-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gajdosova J., Benedikovicova K., Kamodyova N., Tothova L., Kaclikova E., Stuchlik S., et al. (2011). Analysis of the DNA region mediating increased thermotolerance at 58 degrees C in Cronobacter spp. and other enterobacterial strains. Antonie Van Leeuwenhoek 100, 279–289 10.1007/s10482-011-9585-y [DOI] [PubMed] [Google Scholar]
- Gibson D. L., White A. P., Snyder S. D., Martin S., Heiss C., Azadi P., et al. (2006). Salmonella produces an O-antigen capsule regulated by AgfD and important for environmental persistence. J. Bacteriol. 188, 7722–7730 10.1128/JB.00809-06 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goldman E., Green L. H. (2009). Practical Handbook of Microbiology. London: CRC Press [Google Scholar]
- Gosney M. A., Martin M. V., Wright A. E., Gallagher M. (2006). Enterobacter sakazakii in the mouths of stroke patients and its association with aspiration pneumonia. Eur. J. Intern. Med. 17, 185–188 10.1016/j.ejim.2005.11.010 [DOI] [PubMed] [Google Scholar]
- Grim C. J., Kothary M. H., Gopinath G., Jarvis K. G., Beaubrun J. J., McClelland M., et al. (2012). Identification and characterization of Cronobacter iron acquisition systems. Appl. Environ. Microbiol. 78, 6035–6050 10.1128/AEM.01457-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grim C. J., Kotewicz M. L., Power K., Pagotto F., Gopinath G., Mammel M. K., et al. (2013). Pan genome analysis of the emerging foodborne pathogen Cronobacter spp. suggests a species-level bidirectional divergence driven by niche adaption. BMC Genomics. 14:366 10.1186/1471-2164-14-366 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gurtler J. B., Kornacki J. L., Beuchat L. R. (2005). Enterobacter sakazakii: a coliform of increased concern to infant health. Int. J. Food Microbiol. 104, 1–34 10.1016/j.ijfoodmicro.2005.02.013 [DOI] [PubMed] [Google Scholar]
- Himelright I., Harris E., Lorch V., Anderson M., Jones T., Craig A., et al. (2002). Enterobacter sakazakii infections associated with the use of powdered infant formula – Tennessee, 2001. JAMA, 287, 2204–2205 [PubMed] [Google Scholar]
- Hunter C. J., Petrosyan M., Ford H. R., Prasadarao N. V. (2008). Enterobacter sakazakii: an emerging pathogen in infants and neonates. Surg. Infect. 9, 533–539 10.1089/sur.2008.006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iversen C., Lehner A., Mullane N., Marugg J., Fanning S., Stephan R., et al. (2007). Identification of “Cronobacter” spp. (Enterobacter sakazakii). J. Clin. Microbiol. 45, 3814–3816 10.1128/JCM.01026-07 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iversen C., Mullane N., McCardell B., Tall B. D., Lehner A., Fanning S., et al. (2008). Cronobacter gen. nov., a new genus to accommodate the biogroups of Enterobacter sakazakii, and proposal of Cronobacter sakazakii gen. nov., comb. nov., Cronobacter malonaticus sp. nov., Cronobacter turicensis sp. nov., Cronobacter muytjensii sp. nov., Cronobacter dublinensis sp. nov., Cronobacter genomospecies 1, and of three subspecies, Cronobacter dublinensis subsp. dublinensis subsp. nov., Cronobacter dublinensis subsp. lausannensis subsp. nov. and Cronobacter dublinensis subsp. lactaridi subsp. nov. Int. J. Syst. Evol. Microbiol. 58, 1442–1447 10.1099/ijs.0.65577-0 [DOI] [PubMed] [Google Scholar]
- Iversen C., Waddington M., On S. L., Forsythe S. (2004). Identification and phylogeny of Enterobacter sakazakii relative to Enterobacter and Citrobacter Species. J. Clin. Microbiol. 42, 5368–5370 10.1128/JCM.42.11.5368-5370.2004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jaradat Z. W., Ababneh Q. O., Saadoun I. M., Samara N. A., Rashdan A. M. (2009). Isolation of Cronobacter spp. (formerly Enterobacter sakazakii) from infant food, herbs and environmental samples and the subsequent identification and confirmation of the isolates using biochemical, chromogenic assays, PCR and 16S rRNA sequencing. BMC Microbiol. 9:225 10.1186/1471-2180-9-225 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jarvis K. G., Grim C. J., Franco A. A., Gopinath G., Sathyamoorthy V., Hu L., et al. (2011). Molecular characterization of Cronobacter lipopolysaccharide O-antigen gene clusters and development of serotype-specific PCR assays. Appl. Environ. Microbiol. 77, 4017–4026 10.1128/AEM.00162-11 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jarvis K. G., Yan Q. Q., Grim C. J., Power K. A., Franco A. A., Hu L., et al. (2013). Identification and characterization of five new molecular serogroups of Cronobacter spp. Foodborne Pathog. Dis. 10, 343–352 10.1089/fpd.2012.1344 [DOI] [PubMed] [Google Scholar]
- Joseph S., Cetinkaya E., Drahovska H., Levican A., Figueras M. J., Forsythe S. J. (2011). Cronobacter condimenti sp. nov., isolated from spiced meat, and Cronobacter universalis sp. nov., a species designation for Cronobacter sp. genomospecies 1, recovered from a leg infection, water and food ingredients. Int. J. Syst. Evol. Microbiol. 62, 1277–1283 10.1099/ijs.0.032292-0 [DOI] [PubMed] [Google Scholar]
- Joseph S., Desai P., Ji Y. M., Cummings C. A., Shih R., Degoricija L., et al. (2012). Comparative analysis of genome seq-uences covering the seven Crono-bacter species. PLoS ONE 7:e49455 10.1371/journal.pone.0049455 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Joseph S., Forsythe S. J. (2012). Insight into the emergent bacterial pathogen Cronobacter spp., generated by multilocus sequence typing and analysis. Front. Microbiol. 3:397 10.3389/fmicb.2012.00397 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim K., Kim K. P., Choi J., Lim J. A., Lee J., Hwang S., et al. (2010). Outer membrane proteins A (OmpA) and X (OmpX) are essential for basolateral invasion of Cronobacter sakazakii. Appl. Environ. Microbiol. 76, 5188–5198 10.1128/AEM.02498-09 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kucerova E., Clifton S. W., Xia X. Q., Long F., Porwollik S., Fulton L., et al. (2010). Genome sequence of Cronobacter sakazakii BAA-894 and comparative genomic hybridization analysis with other Cronobacter Species. PLoS ONE 5:e9556 10.1371/journal.pone.0009556 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lamark T., Kaasen I., Eshoo M. W., Falkenberg P., McDougall J., Strom A. R. (1991). DNA sequence and analysis of the bet genes encoding the osmoregulatory choline – glycine betaine pathway of Escherichia coli. Mol. Microbiol. 5, 1049–1064 10.1111/j.1365-2958.1991.tb01877.x [DOI] [PubMed] [Google Scholar]
- Lee J. H., Choi Y., Shin H., Lee J., Ryu S. (2012). Complete genome sequence of Cronobacter sakazakii temperate bacteriophage phiES15. J. Virol. 86, 7713–7714 10.1128/JVI.01042-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lehner A., Fricker-Feer C., Stephan R. (2012). Identification of the recently described Cronobacter condimenti by a rpoB based PCR system. J. Med. Microbiol. 67, 1034–1035 10.1099/jmm.0.042903-0 [DOI] [PubMed] [Google Scholar]
- Lewis K. (2000). Programmed death in bacteria. Microbiol. Mol. Biol. Rev. 64, 503–514 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mange J. P., Stephan R., Borel N., Wild P., Kim K. S., Pospischil A., et al. (2006). Adhesive properties of Enterobacter sakazakii to human epithelial and brain microvascular endothelial cells. BMC Microbiol. 6:58 10.1186/1471-2180-6-58 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maurelli A. T. (2007). Black holes, antivirulence genes, and gene inactivation in the evolution of bacterial pathogens. FEMS Microbiol. Lett. 267, 1–8 10.1111/j.1574-6968.2006.00526.x [DOI] [PubMed] [Google Scholar]
- Miranda C. D., Kehrenberg C., Ulep C., Schwarz S., Roberts M. C. (2003). Diversity of tetracycline resistance genes in bacteria from chilean salmon farms. Antimicrob. Agents Chemother. 47, 883–888 10.1128/AAC.47.3.883-888.2003 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mullane N., Healy B., Meade J., Whyte P., Wall P. G., Fanning S. (2008a). Dissemination of Cronobacter spp. (Enterobacter sakazakii) in a powdered milk protein manufacturing facility. Appl. Environ. Microbiol. 74, 5913–5917 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mullane N., O'Gaora P., Nally J. E., Iversen C., Whyte P., Wall P. G., et al. (2008b). Molecular analysis of the Enterobacter sakazakii O-antigen gene locus. Appl. Environ. Microbiol. 74, 3783–3794 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mullane N. R., Iversen C., Healy B., Walsh C., Whyte P., Wall P. G., et al. (2007). Enterobacter sakazakii an emerging bacterial pathogen with implications for infant health. Minerva Pediatr. 59, 137–148 [PubMed] [Google Scholar]
- Overbeek R., Begley T., Butler R. M., Choudhuri J. V., Chuang H. Y., Cohoon M., et al. (2005). The subsystems approach to genome annotation and its use in the project to annotate 1000 genomes. Nucleic Acids Res. 33, 5691–5702 10.1093/nar/gki866 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Power K. A., Yan Q. Q., Fox E., Cooney S., Fanning S. (2013). Genome sequence of Cronobacter sakazakii SP291 – a persistent thermo-tolerant powdered infant formula factory-derived pathogen. Genome Announc. 1:e0008213 10.1128/genomeA.00082-13 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmid M., Iversen C., Gontia I., Stephan R., Hofmann A., Hartmann A., et al. (2009). Evidence for a plant-associated natural habitat for Cronobacter spp. Res. Microbiol. 160, 608–614 10.1016/j.resmic.2009.08.013 [DOI] [PubMed] [Google Scholar]
- See K. C., Than H. A., Tang T. (2007). Enterobacter sakazakii bacteraemia with multiple splenic abscesses in a 75-year-old woman: a case report. Age Ageing 36, 595–596 10.1093/ageing/afm092 [DOI] [PubMed] [Google Scholar]
- Serfiotis-Mitsa D., Herbert A. P., Roberts G. A., Soares D. C., White J. H., Blakely G. W., et al. (2010). The structure of the KlcA and ArdB proteins reveals a novel fold and antirestriction activity against Type I DNA restriction systems in vivo but not in vitro. Nucleic Acids Res. 38, 1723–1737 10.1093/nar/gkp1144 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shaker R. R., Osaili T. M., Abu Al-Hasan A. S., Ayyash M. M., Forsythe S. J. (2008). Effect of desiccation, starvation, heat, and cold stresses on the thermal resistance of Enterobacter sakazakii in rehydrated infant milk formula. J. Food Sci. 73, M354–M359 10.1111/j.1750-3841.2008.00880.x [DOI] [PubMed] [Google Scholar]
- Shin H., Lee J. H., Choi Y., Ryu S. (2012). Complete genome sequence of the opportunistic food-borne pathogen Cronobacter sakazakii ES15. J. Bacteriol. 194, 4438–4439 10.1128/JB.00841-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shin J. H., Oh S. Y., Kim S. J., Roe J. H. (2007). The zinc-responsive regulator Zur controls a zinc uptake system and some ribosomal proteins in Streptomyces coelicolor A3(2). J. Bacteriol. 189, 4070–4077 10.1128/JB.01851-06 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sivamaruthi B. S., Ganguli A., Kumar M., Bhaviya S., Pandian S. K., Balamurugan K. (2011). Caenorhabditis elegans as a model for studying Cronobacter sakazakii ATCC BAA-894 pathogenesis. J. Basic Microbiol. 51, 540–549 10.1002/jobm.201000377 [DOI] [PubMed] [Google Scholar]
- Stephan R., Lehner A., Tischler P., Rattei T. (2011). Complete genome sequence of Cronobacter turicensis LMG 23827, a food-borne pathogen causing deaths in neonates. J. Bacteriol. 193, 309–310 10.1128/JB.01162-10 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stock I., Wiedemann B. (2002). Natural antibiotic susceptibility of Enterobacter amnigenus, Enterobacter cancerogenus, Enterobacter gergoviae and Enterobacter sakazakii strains. Clin. Microbiol. Infect. 8, 564–578 10.1046/j.1469-0691.2002.00413.x [DOI] [PubMed] [Google Scholar]
- Stoop B., Lehner A., Iversen C., Fanning S., Stephan R. (2009). Development and evaluation of rpoB based PCR systems to differentiate the six proposed species within the genus Cronobacter. Int. J. Food Microbiol. 136, 165–168 10.1016/j.ijfoodmicro.2009.04.023 [DOI] [PubMed] [Google Scholar]
- Tsai H. Y., Liao C. H., Huang Y. T., Lee P. I., Hsueh P. R. (2013). Cronobacter infection not from infant formula, Taiwan. Emerg. Infect. Dis. 19, 1–3 10.3201/eid1901.120774 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tschowri N., Busse S., Hengge R. (2009). The BLUF-EAL protein YcgF acts as a direct anti-repressor in a blue-light response of Escherichia coli. Genes Dev. 23, 522–534 10.1101/gad.499409 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walsh D., Molloy C., Iversen C., Carroll J., Cagney C., Fanning S., et al. (2011). Survival characteristics of environmental and clinically derived strains of Cronobacter sakazakii in infant milk formula (IMF) and ingredients. J. Appl. Microbiol. 110, 697–703 10.1111/j.1365-2672.2010.04921.x [DOI] [PubMed] [Google Scholar]
- Wang I. N., Smith D. L., Young R. (2000). Holins: the protein clocks of bacteriophage infections. Annu. Rev. Microbiol. 54, 799–825 10.1146/annurev.micro.54.1.799 [DOI] [PubMed] [Google Scholar]
- Williams T. L., Monday S. R., Edelson-Mammel S., Buchanan R., Musser S. M. (2005). A top-down proteomics approach for differentiating thermal resistant strains of Enterobacter sakazakii. Proteomics 5, 4161–4169 10.1002/pmic.200401263 [DOI] [PubMed] [Google Scholar]
- Yan Q. Q., Condell O., Power K., Butler F., Tall B. D., Fanning S. (2012). Cronobacter species (formerly known as Enterobacter sakazakii) in powdered infant formula: a review of our current understanding of the biology of this bacterium. J. Appl. Microbiol. 113, 1–15 10.1111/j.1365-2672.2012.05281.x [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Genomic regions missing in C. sakazakii SP291 when compared to the other three completed genomes of C. sakazakii ATCC BAA-894 (as the reference sequence), C. sakazakii E15 and C. turicensis z3032.
Genomic regions present only in C. sakazakii SP291 (as the reference sequence) when compared to the other three completed genomes of C. sakazakii ATCC BAA-894, C. sakazakii ES15 and C. turicensis z3032.
Genome comparison of C. sakazakii SP291 with other available C. sakazakii genomes, which include C. sakazakii SP291, ATCC BAA-894, ES15, E899, 680, 696, 701, ES35, 2151, ES713, and E764.
Genome comparison of C. sakazakii SP291 with Cronobacter isolate of other six species, which consist of C. dublinensis LMG 23823, C. malonaticus LMG 23826, C. muytjensii ATCC 51329, C. turicensis z3032, C. universalis NCTC 9529, and C. condimenti 1330.
Annotated plasmid clusters of three complete genome, which include pSP291-1, pSP291-2, and pSP291-3 of C. sakazakii SP291, pESA2 and pESA3 of C. sakazakii ATCC BAA-894, as well as pCTU1, pCTU2, and pCTU3 of C. turicensis z3032. Plasmid group 1: pSP291-1, pESA3, and pCTU1; plasmid group 2: pSP291-2 and pCTU3.
Numerical data for all 20 PM plates of C. sakazakii SP291 and C. sakazakii ATCC BAA-894, along with the phenotype differences (greater than 20,000±1,800 unit) generated by Omnilog® software.
Genes assigned to defined bacterial sub-systems and their distribution among these functional categories for C. sakazakii SP291 along with other three completed genome, C. sakazakii ATCC BAA-894, C. sakazakii ES15, and C. turicensis z3032.
Genome annotation: Carbohydrate metabolism of C. sakazakii SP291 and C. sakazakii ATCC BAA-894.
Genome annotation: Phosphorus metabolism of C. sakazakii SP291 and C. sakazakii ATCC BAA-894.
Genome annotation: Sulfur metabolism of C. sakazakii SP291 and C. sakazakii ATCC BAA-894.
Iron acquisition systems identified in the chromosome of C. sakazakii SP291.