SUMMARY
Mucosal surfaces such as the gut, vagina and oral cavity are colonized by microbiota that are an integral component of the healthy ecosystem. Recent molecular techniques make it feasible to correlate antimicrobial dosing levels with changes in microbiome composition. The objective of this study was to characterize the rat oral plaque microbiome composition at doses of ciprofloxacin that were considerably above and below nominal in vitro minimal inhibitory concentrations (MICs) of a variety of gram positive oral commensal bacteria. We exposed the oral cavities of rats to relatively low (0.1 µg/mL) and high (20 µg/mL) doses of ciprofloxacin in the drinking water over a 3 day period. Plaque microbiota were characterized using 454 pyrosequencing. The rat indigenous community was dominated by Rothia (74.4 %) and Streptococcus genera (4.7%). Dosing at 0.1 µg/mL was associated with changes in Rothia and Streptococcus genera which were not significant, while dosing at 20 µg/mL caused a pronounced (significant) reduction in the relative abundance of the Streptococcus genus. Taxonomic independent analysis indicated that the perturbation in the overall community structure attributed to dosing with ciprofloxacin at either the low or high dose was relatively low. The results suggest that it is feasible to use an antimicrobial dosing regime to selectively target a specific subset of a mucosal microbiome for elimination with minimal perturbation of the entire community.
Keywords: oral, periodontitis, ciprofloxacin, rat, microbiome
INTRODUCTION
Mucosal surfaces such as the gut, vagina and oral cavity are densely colonized by microorganisms that are an integral biological component of healthy humans (Pflughoeft and Versalovic 2012). Besides the well established symbiotic relationships in the gut (Sears 2005), indigenous microbiota that colonize healthy mucosal surfaces compete with putative pathogens (Pultz, et al. 2004, Teughels, et al. 2007, Ubeda, et al. 2010), and contribute to immune homeostasis (Krisanaprakornkit, et al. 2000, Mans, et al. 2009, Neish, et al. 2000, Rakoff-Nahoum, et al. 2004). Concerns about the negative consequences of the disturbance of the commensal mucosal community by conventional antimicrobial treatments have led to characterization of the perturbation of ciprofloxacin dosing on the gut microbiome (Dethlefsen, et al. 2008, Endt, et al. 2012).
Ciprofloxacin is used to treat periodontitis in humans (Otero, et al. 2009). Fluoroquinolones are generally more efficacious against gram-negative bacteria than gram-positive bacteria (Appelbaum and Hunter 2000, Avery 2004) and our previous results suggested that appropriate dosing with this drug might be used to selectively eliminate a periodontal pathogen in an oral biofilm consortium (Suci and Young 2011). Rat models are particularly relevant for characterizing the progression of the disease from health to chronic destructive inflammation (Bezerra, et al., Kawai, et al. 2000, Kawai, et al. 1998, Li, et al. 2010, Schreiner, et al. 2010). Despite evidence that commensal microbes probably play a critical role in determining the outcome of mucosal infections, the rat oral microbiome has not been well characterized using culture independent methods. An unanticipated result of our study was that the high dose of ciprofloxacin selectively affected a subset of the oral microbiome with minimal disturbance of the overall plaque community structure.
MATERIALS AND METHODS
Animals
Fifteen, 6-month-old female rats (Sprague-Dawley, Charles River Laboratories, Hollister, CA) were housed individually at 22 °C under a 12-h light and 12-h dark cycle and fed ad libitum with a soft diet (Purina Laboratory Rodent Chow) following the guidelines and regulations for the use and care of animals at the University of Southern California.
Plaque collection and DNA isolation
Supragingival plaque was acquired from buccal and lingual regions of the upper molars of rats using extrafine paper points (Freire, et al. 2011). Four plaque samples from each of 5 rats were pooled for analysis. Samples were pooled with the intension of reducing the effect of between site and between animal variability in order to detect differences in community structure that could be attributed specifically to dosing. Cells were lysed using a previous protocol (Suci and Young 2011) with an additional bead beating step. DNA was isolated using a GenElute™ Bacterial Genomic DNA kit (Sigma-Aldrich).
16S rRNA Tag Pyrosequencing
Amplicon libraries were generated using HPLC-purified fusion primers that consisted of a 454 adaptor (5'-CCTATCCCCTGTGTGCCTTGGCAGTCTCAG-3') linked to 10 nucleotide identification tag followed by a template specific forward primer (5'-TCACGRCACGAGCTGACGAC-3') or reverse primer (5'-GGATTAGATACCCBRGTAGTC-3') to conserved regions flanking the V5V6 hypervariable region of the 16S rRNA gene (Zaura, et al. 2009). There were 6 identification tags used to discriminate 6 pooled samples. The PCR products were purified using AMPure XP beads (Angercourt). Amplicon quality and concentration were analyzed by BioAnalyzer (Agilent 2100) and Qubit® 2.0 Fluorometer respectively. Purified amplicons were sequenced at the W. M. Keck Center for Comparative and Functional Genomics at the University of Illinois at Urbana-Champaign.
Ciprofloxacin dosing
There were three groups of 5 rats: 1) a "no dose" control group; 2) a "low dose" group (0.1 µg/mL ciprofloxacin); and 3) a "high dose" group (20 µg/mL ciprofloxacin). These concentrations are, respectively, well below and above nominal in vitro minimal inhibitory concentrations (MICs) of a variety of gram positive oral commensal bacteria (Hoogkamp-Korstanje and Roelofs-Willemse 2000). The high dose is approximately 5.8 mg/kg body weight per day, a dose level that we anticipated the rats would tolerate without reduced water consumption (Xue, et al. 2009). Samples were acquired immediately before dosing (day 0) and 72 h later (day 3) yielding 6 pooled samples. We anticipated that dosing over this time period might affect the microbiome taxa in a manner that was comparable to in vitro tests. Rats were dosed with ciprofloxacin (ICN Biomedicals Inc., catalog # 199020) by adding it to the drinking water. In addition, a topical application was made immediately following acquisition of the first plaque sample (day 0) and 24 h after this time point. For this procedure animals were anesthetized with xylazine and ketamine, the mouth was retracted and 500 µL of the ciprofloxacin solution was applied to the buccal and lingual regions of the upper molar over a 5 min period. Water consumption was normal (mean, 64 mL/day water) and not significantly different between the 3 groups of rats.
Data analysis
Sequences were filtered and trimmed using the Ribosomal Data Base (RDP) pyrosequencing pipeline. Taxonomic classifications were made using the RDP Classifier (Wang, et al. 2007) (Release 10, Update 30 which consists of 2,578,902 aligned and annotated 16S rRNA sequences) with a bootstrap cutoff of 80%. Taxonomic independent analysis was performed using MOTHUR (Schloss, et al. 2009). Unique sequences were aligned to the Silva database (Pruesse, et al. 2007), chimeras were removed and operational taxonomic units (OTUs) were determined by clustering at genetic distances ranging from 0.01 to 0.08 using an average neighbor algorithm. OTUs obtained using a similarity (distance) cutoff of 0.03 (3%) were used for further analysis (Dethlefsen, et al. 2008, Zaura, et al. 2009). AMOVA (Excoffier, et al. 1992) as implemented by MOTHUR was used to interpret matrices of Yue & Clayton dissimilarity values (Yue and Clayton 2005). Phylogenic trees were constructed with the program MEGA 4 using the Neighbor-Joining method with Jukes-Cantor distances (Tamura, et al. 2007). Phylogenic trees were analyzed to determine similarity of community structure using UniFrac (Lozupone, et al. 2006).
RESULTS
Sequence data
Of the total of 91120 reads, 86513 passed quality control. There were 7234 unique sequences which included 3454 sequences incorporating chimeras which were removed. Chimeric sequences were present at relatively low abundance as shown by the moderate reduction in total number of sequences after chimera removal (80080). The mean length of sequences used for the analysis was 256.52 nucleotides.
Rat plaque community structure
The taxonomic dependent analysis for the four pooled samples from rats not exposed to ciprofloxacin indicated that the population was dominated by Actinobacteria and to a less extent, Firmicutes (Table 1). Of the total of 80080 sequences that were analyzed, 7444 were classified at the genus level and 6303 were unclassified. The genera that contribute by far the most to these phyla are Rothia and Streptococcus, respectively. We report relative abundance data only for taxa represented by more than 5 sequences since the confidence limits for relative abundance determinations of taxa represented by less than 5 sequences are very large (Dethlefsen, et al. 2008).
Table 1.
Taxon relative abundance
| Actinobacteria1 | 82.542 |
| Rothia3 | 74.43 |
| Corynebacterium | 0.18 |
| Microcella | 0.04 |
| Propionibacteria | 0.01 |
| Brachybacterium | 0.01 |
| Janibacter | 0.01 |
| Kocuria | * |
| Micrococcus | * |
| Firmicutes | 7.81 |
| Streptococcus | 4.67 |
| Staphylococcus | 0.36 |
| Exiguobacterium | 0.19 |
| Enterococcus | 0.08 |
| Granulicatella | 0.02 |
| Anaerobacillus | * |
| Gemella | * |
| Proteobacteria | 1.04 |
| Halomonas | 0.55 |
| Shewanella | 0.14 |
| Paramoritella | 0.13 |
| Uruburuella | 0.05 |
| Paracoccus | 0.04 |
| Vibrio | 0.02 |
| Diaphorobacter | 0.02 |
| Neisseria | 0.01 |
| Acinetobacter | 0.01 |
| Lucibacterium | * |
| Sphingomonas | * |
| Cyanobacteria | 0.09 |
| Bacteroidetes | 0.03 |
| Acidobacteria | 0.03 |
| Planctomycetes | 0.02 |
| Alkaliflexus | 0.01 |
| Aquificae | * |
| Unclassified | 8.42 |
Phyla are in bold
Percent relative abundance
Genera are in italics
less than five sequences
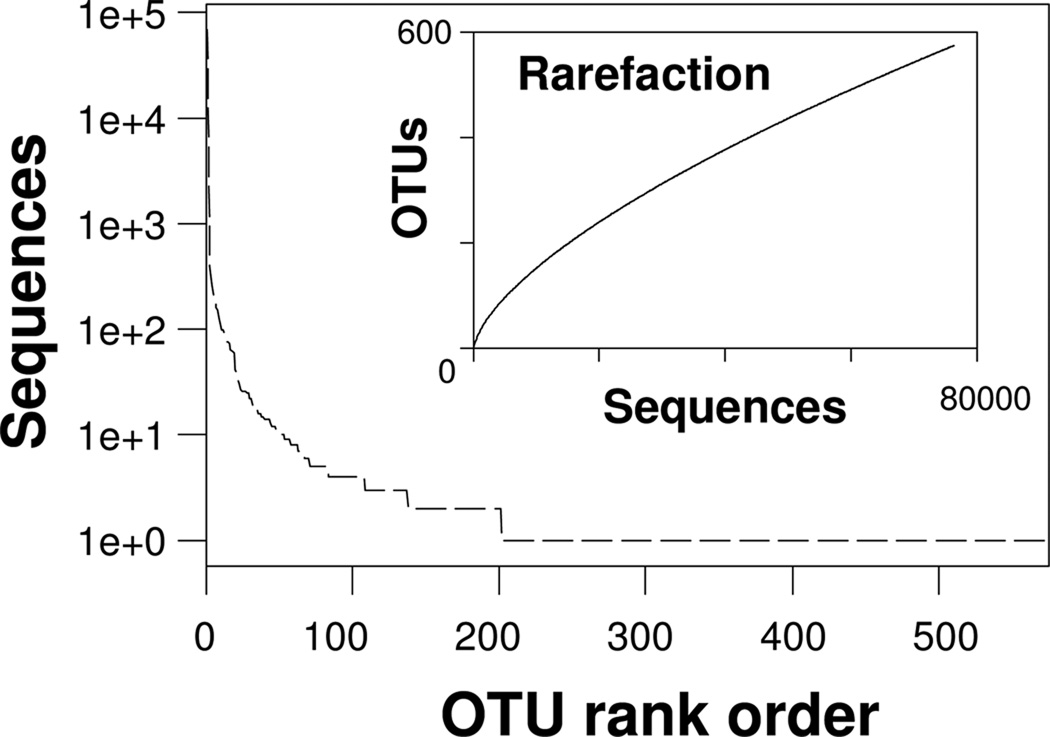
Rank abundance and rarefaction curves obtained from the taxonomic independent analysis exhibit fairly typical shapes (Fig.1). The richness at a genetic distance of 0.03 was 575 OTUs. Of the 575 total OTUs there were 490 OTUs (85.2%) represented by less than 5 sequences. These relatively low abundance OTUs contributed less than 1% to the total relative abundance. According to the Chao I estimator (Chao 1984) the projected richness of the rat plaque community based on the rarefaction analysis was 1648 OTUs at a genetic distance of 0.03 with a 95% confidence range of 1355 to 2050. The estimated community richness of the rat community was considerably less than the oral human supragingival plaque microbiome (reduced by approximately 6.3 %) computed using the same parameters (Chao 1 estimator of richness at a genetic distance of 0.3) (Keijser, et al. 2008).
Figure 1.
Rank abundance curve and rarefaction curve (insert) for sequence data from all 6 conditions combined. OTUs were defined at a distance of 0.03
Effect of ciprofloxacin dosing on the indigenous community
Taxonomic classifications (RDP) were used to assess the effect of ciprofloxacin dosing on individual genera or families. A family was included if: 1) it made up more than 50% of its order and, 2) other genera listed in that family contributed less than 50% abundance to that family. We first looked for evidence for a significant change in relative abundance of a taxonomic group represented by more than 5 sequences for a day 0 condition (~0.04 % relative abundance) (Table 2). A genus or family was excluded from the analysis if it was not present in the no dose day 3 condition or in all day 0 conditions. Sequences associated with these excluded taxa contributed less than 1 % to the total relative abundance. There were 6 taxa for which the relative abundance on day 3 was greater than or equal to one standard deviation unit removed from the mean of the day 0 relative abundance. A single value t test was used to compute p values. For the two dominant genera the most significant effect of ciprofloxacin dosing was the reduction in Streptococcus at the high dose. Rothia, was also perturbed by the high dose more significantly than the low dose. With respect to the low abundance taxa, the pattern of reduction in the relative abundance of the Corynebacterium genus appears to be a consequence of ciprofloxacin dosing, while the changes in the relative abundance of the Microbacteriace family and the Uruburuella genus do not appear to be clearly caused by ciprofloxacin dosing. We also looked for evidence for overgrowth by a genera or family that that was absent or represented by less than 5 sequences for the day 0 low or high dose conditions. The largest increase between day 0 and day 3 in this category was for the Pasteurellaceae family which increased from 0.01 to 0.2 % relative abundance for the low dose condition.
Table 2.
Ciprofloxacin dosing effect on biofilm genera
| Day 01 | Day 32 | |||
|---|---|---|---|---|
| no dose | lo dose | hi dose | ||
| Rothia | 70.92 +/− 9.18 | 84.98 (1.5) | 80.39 (1.0) | 93.50 (2.4)* |
| Streptococcus | 4.92 +/− 1.90 | 3.86 (0.6) | 7.49 (1.3) | 0.22 (2.5)* |
| Rhodobacteracea | 0.28 +/− 0.06 | 0.10 (2.9)* | 0.04 (3.9)* | 0.03 (4.0)* |
| Corynebacterium | 0.20 +/− 0.07 | 0.15 (0.7) | 0.02 (2.6)* | 0.02 (2.6)* |
| Microbacteriace | 0.19 +/− 0.04 | 0.05 (3.5)* | 0.05 (3.5)* | 0.02 (4.4)* |
| Uruburuella | 0.07 +/− 0.03 | 0.01 (2.1) | 0.03 (1.4) | 0.00 (2.4)* |
Mean percent relative abundance for Day 0 +/− standard deviation
Mean percent relative abundance for Day 3 (Difference between Day 0 mean in standard deviation units)
p value < 0.05
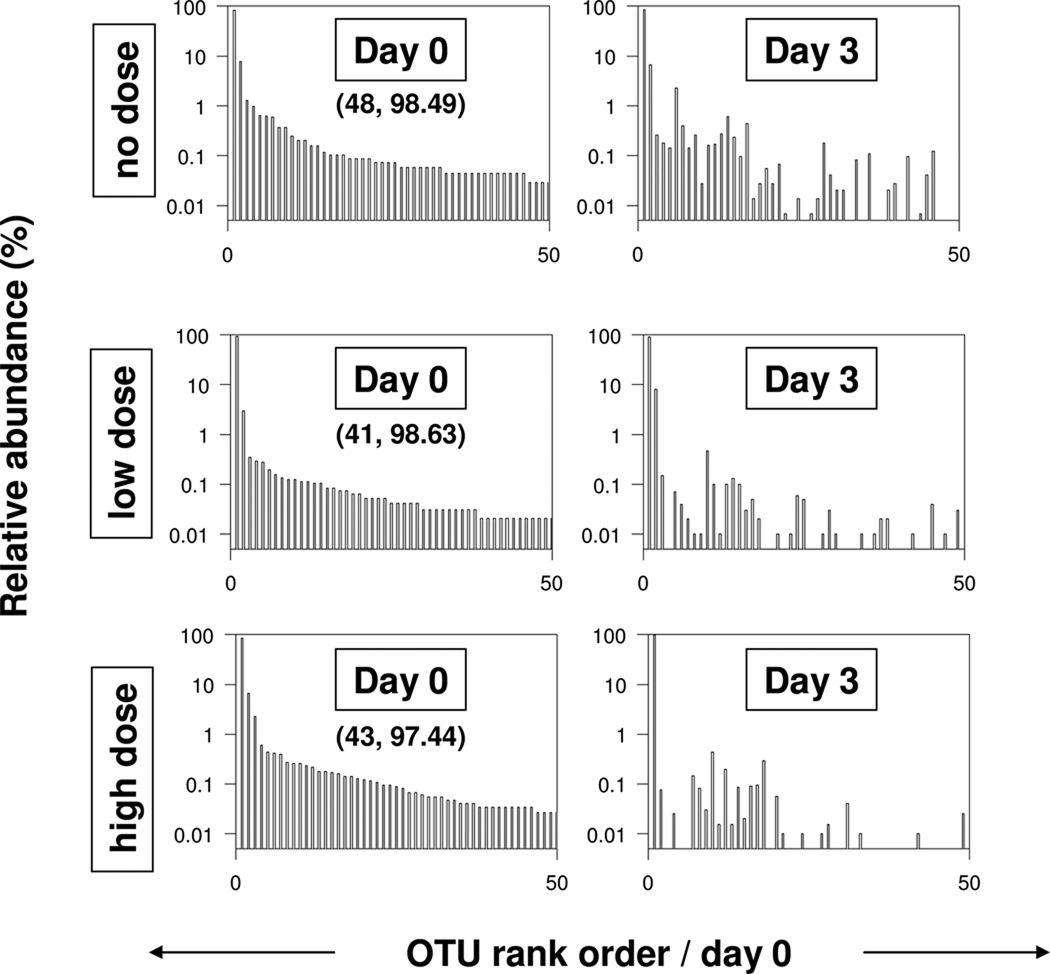
Taxonomy independent analysis indicated that the overall perturbation of the indigenous microbiota caused by ciprofloxacin dosing at either the high or low dose was relatively small. UniFrac analysis based on phylogenic trees indicated that community structure was not significantly different between any of the pairs of conditions (p values > 0.015). Changes in OTU relative abundance over the three day period are shown in Fig.2 for the 50 most abundant OTUs obtained at a distance of 0.03. Most of the sequences were incorporated into shared OTUs. The number of shared OTUs between each pair of day 0/day 3 conditions at the no, low and high dose is shown in parenthesis in the first column of figures (first value in the pair of numbers). Most of the sequences in the community were associated with shared OTUs as indicated by the second value in the pair of numbers, the percentage of sequences represented by these shared OTUs. The beta diversity between pairs of samples acquired on the same day (three pairs each for day 0 or day 3) and between the three pairs of day0/day3 samples at the different dose levels (no, low and high) is relatively low (Fig.3). The values are all less than 0.5 which, by comparison with an extensive study of the human microbiome, suggests modest differences in community structure between each pair of samples (Huttenhower, et al. 2012). The beta diversity between day 0 and day 3 samples for the low dose condition is notably lower than the beta diversity for other pairs of samples, and the beta diversity for the no dose condition pairs is slightly higher than for the high dose pairs. This same trend is evident in the Shannon diversity index, which quantifies the within sample (alpha) diversity (Fig.4). These latter data show that in terms of absolute difference in alpha diversity within each group, the effect of ciprofloxacin was relatively small. Indices of evenness and richness for all 6 conditions are also shown in Fig.4. There is a decrease in the means of each of the indices between day 0 and day 3 which is significant at the 0.05 level only for the richness index. There is no obvious pattern to indicate that any index presented in Fig.4 was perturbed by ciprofloxacin dosing.
Figure 2.
Rank abundance curves of the 50 most abundant OTUs defined at a distance of 0.03 plotted in terms of percent relative abundance for each of the 6 conditions separately. The OTU rank order on the horizontal axes is that obtained from the corresponding day 0 condition for all 3 pairs of plots. Numbers in parentheses below the day 0 label are, respectively, the number of shared OTUs and the percent of sequences in these shared OTUs for each of the day0/day3 pairs of sequences at each of the three dose levels.
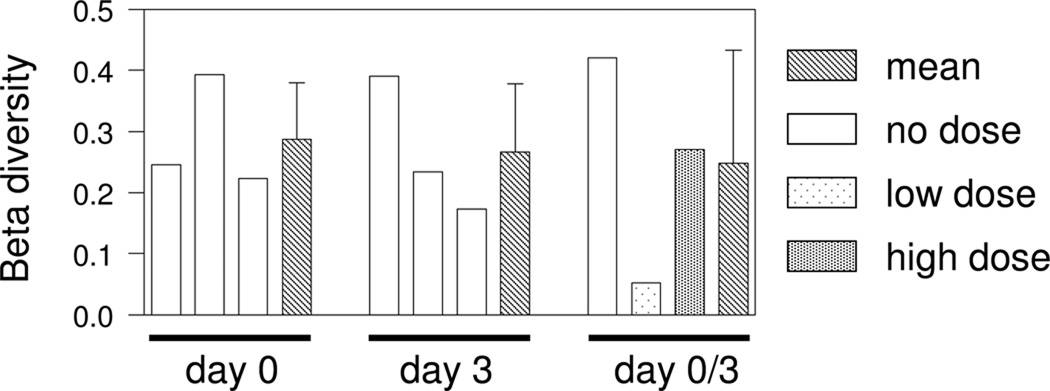
Figure 3.
The Bray-Curtis (beta) diversity between each of the 3 pairs of day 0 and day 3 samples and between each pair of day 0/day 3 samples for each dosing condition. Means are presented for each set of three diversity values. Error bars are standard deviations of the mean.
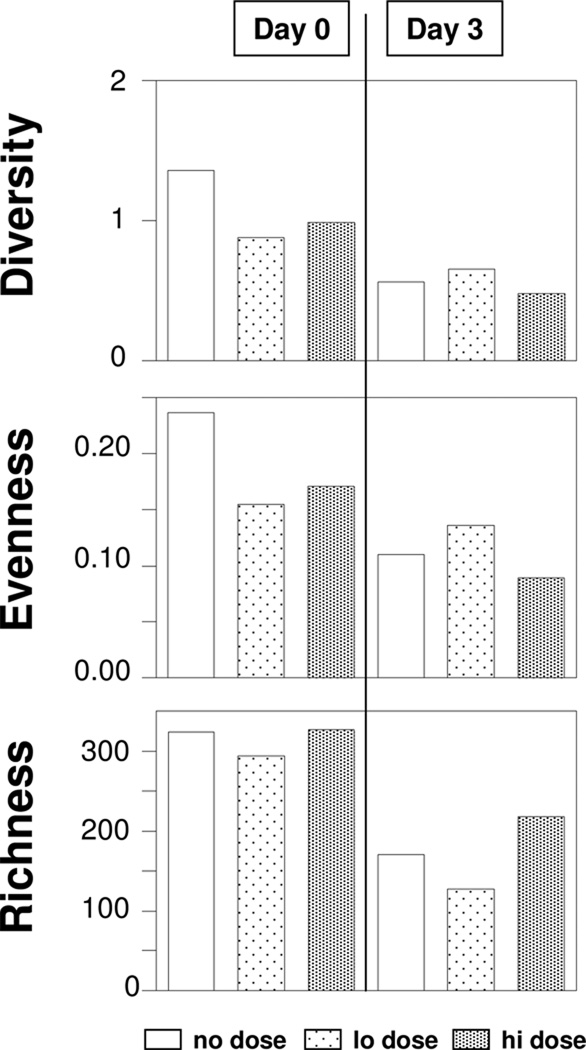
Figure 4.
Shannon diversity, Shannon evenness and richness indices for the 6 conditions. The legend at the bottom applies to all the plots.
To determine whether differences in community structure were associated more strongly with temporal changes over the three day period, or with the effects of ciprofloxacin dosing, a matrix of Yue & Clayton dissimilarity values was computed. There was a relatively good correlation between the Yu & Clayton values and the UniFrac distances (R=0.8790). AMOVA was used to assess the significance of between group differences. When the matrix of Yu & Clayton values was arranged so that group categories were day 0 and day 3 the difference between groups was more significant (p value = 0.10) than when the matrix was arranged so that the dose levels were the group categories (p value=0.76). This indicates that the perturbation in overall community structure caused by ciprofloxacin dosing was less than the temporal changes over the 3 day period.
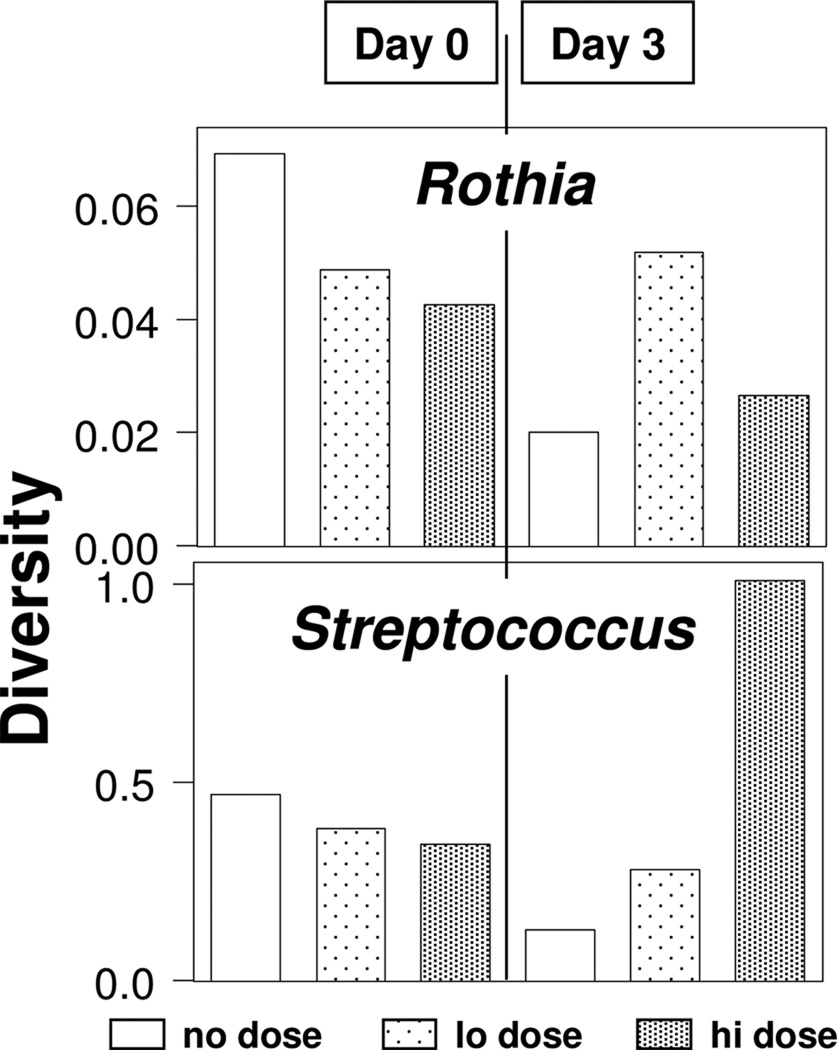
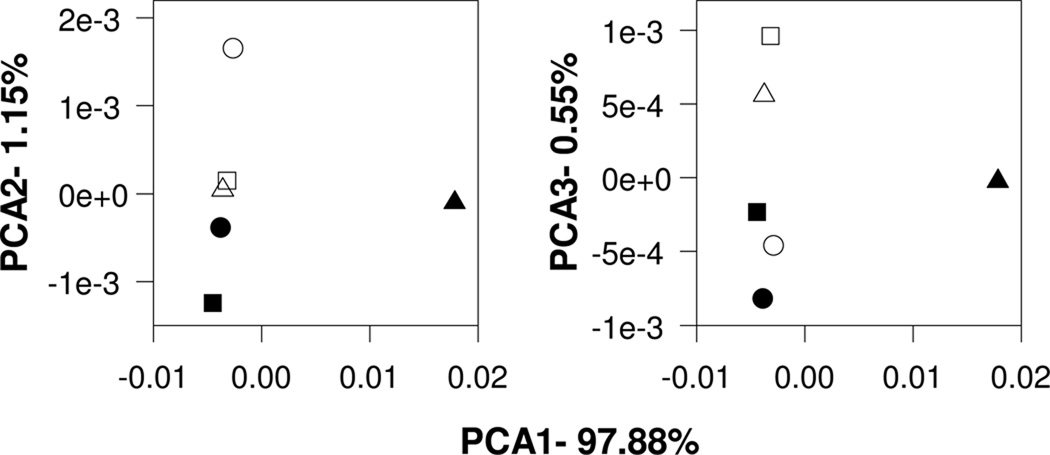
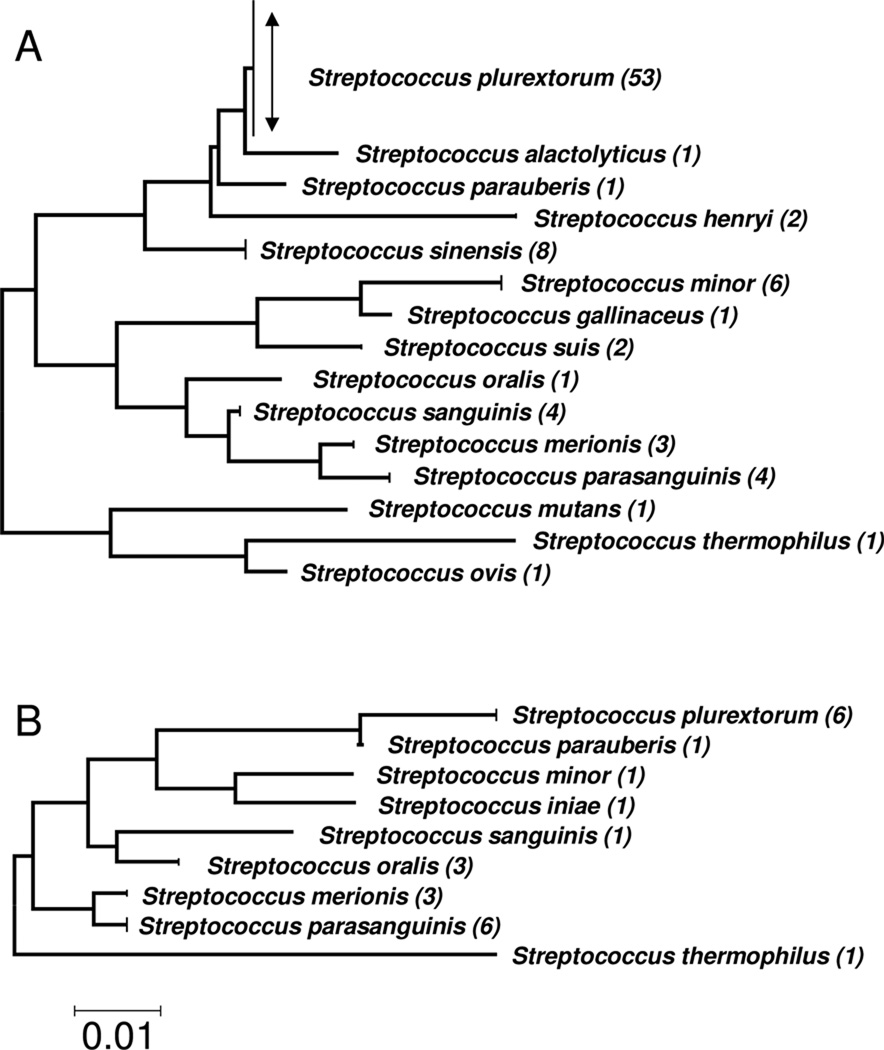
In order to determine whether ciprofloxacin dosing affected the community structure within the two dominant genera, sequences classified by RDP as either Rothia or Streptococcus were analyzed separately. Although the Rothia genus was more abundant than the Streptococcus genus its diversity according to the Shannon diversity index was significantly less (p value < 0.001) (Fig.5). There is no clear pattern in the Shannon diversity index indicating that ciprofloxacin dosing perturbed the community structure of the Rothia genus, while there is a clear trend toward increasing diversity from the no dose to the high dose condition for the Streptococcus population. As above, the community structure was compared between all pairs of the 6 conditions by computing Yue & Clayton dissimilarity values and using AMOVA to assess the significance of between group differences. Similar to the results for the total population, for the Rothia genus the difference between day 0 and day 3 groups was more significant than between dose level groups (p values of 0.53 and 0.992, respectively), In contrast, for the Streptococcus genus the difference was more significant between dose level groups than the day 0 and day 3 groups (p values 0.317 and 0.848, respectively). This statistical result is consistent with the dose-dependent pattern exhibited in Fig.5. This was characterized further using UniFrac distances to perform principle component analysis (PCA) on this subset of sequences (Fig.6). Consistent with the trend in the alpha diversity indices in Fig.5, the subset of Streptococcus sequences belonging to the day 3 high dose condition is an outlier compared to sequences associated with the 5 other conditions with respect to the first principle component that describes 97.88 % of the variation in the data. Inspection of rank abundance curves suggest that the greater diversity exhibited by the subset of Streptococcus sequences belonging to the day 3 high dose condition (Fig.5) originates from a relative reduction in the most abundant OTU. This enhanced the uniformity of the distribution of the sequences among the five different OTU in this relatively small subset of sequences and thus increases the diversity index (data not shown). The interpretation that the high dose of ciprofloxacin produced selective reduction of a subset of phylogenetically similar sequences in the Streptococcus is supported by a comparison of the phylogenetic trees for the day 0 and day 3 high dose conditions constructed from unique sequences (Fig.7). In order to provide some idea of the possible taxonomic identifications within the Streptococcus genus, the trees in Fig.7 are presented in terms of species level identifications obtained by NCBI BLASTN. Sequences identified as S. plurextorum represented, respectively, 88.39 % and 21.74 % of the total sequences in day 0 and day 3 high dose subsets of sequences. With the acknowledgment of the substantial uncertainty in these species level identifications, the data in Fig.7 suggests that ciprofloxacin produced a selective reduction of Streptococcus plurextorum.
Figure 5.
Shannon diversity for sequence data classified as Rothia or Streptococcus for the 6 conditions. The legend at the bottom applies to both plots.
Figure 6.
Principle components obtained from UniFrac analyses of the phyogenetic tree structure of subsets of sequences classified as Streptococcus by RDP. Symbols are no dose condition (circles), low dose condition (triangles), high dose condition (squares), open symbols (day 0), filled symbols (day 3). Percent variation accounted for by each component is specified.
Figure 7.
Phylogenic trees for unique sequences classified as Streptococcus. Species identifications were obtained using NCBI BLASTN from unique sequences from the high dose groups of rats. Numbers in parentheses are the number of unique sequences classified as the corresponding species. A. Day 0 unique sequences. B. Day 3 unique sequences. The distance scale is the same for each tree.
DISCUSSION
The most outstanding effect of ciprofloxacin dosing was a decrease in the relative abundance of the Streptococcus genus at the high dose as indicated by the taxonomic dependent analysis. The decrease in Streptococci recovered from the plaque after dosing for 3 days at 20 µg/mL is consistent with the susceptibility of a variety of oral Streptococcal species measured in vitro for planktonic cells (MIC range of 0.12 to 16 µg/mL) (Hoogkamp-Korstanje and Roelofs-Willemse 2000), and also with the susceptibility of S. pneumoniae biofilms to levofloxacin (del Prado, et al. 2010). In addition to the decrease in relative abundance of sequences identified by RDP as Streptococcus, ciprofloxacin dosing at the high dose appeared to cause a pronounced difference in the community structure associated with this subset of sequences. Specifically, there was a selective reduction in a particular sequence very tentatively classified as S. plurextorum. S. plurextorum was recognized as a new species recently (Vela, et al. 2009). The absence of S. suis in the day 3 trees presented in Fig.7 B may be related to the reduction in S. plurextorum since these two species are closely related. There is some data indicating that S. suis biofilms are more susceptible to ciprofloxacin than other antimicrobials (Guo, et al. 2012). Pronounced effects on overall community structure that could be readily interpreted as dose dependent were not evident in our data. An interpretation supported by our data analysis is that with respect to common indices of overall community structure, changes in microbiome composition produced by ciprofloxacin dosing were less prominent than relatively modest temporal changes in the microbiome composition that occurred over the three day period.
There was a noticeable increase in the relative abundance of Streptococci at the low dose and a significant decrease in the relative abundance of the Corynebacterium genus. The explanation that Corynebacterium species inhibit Streptococci is not supported by the literature (Hogan, et al. 1987). There is some support for the hypothesis that Streptococcal biofilm was reacting to exposure to sublethal doses of ciprofloxacin by an increase in biomass (Ahmed, et al. 2009).
The dominant Rothia genus was relatively unaffected by ciprofloxacin dosing. There is no published evidence indicating that Rothia species exhibit exceptional resistance to ciprofloxacin (Binder, et al. 1997). It is possible that the biofilm form of Rothia exhibits more phenotypic resistance to ciprofloxacin than the biofilm form of the Streptococcus species. Alternatively, our data are consistent with the interpretation that a portion of the Streptococcal community exhibits uncharacteristically enhanced susceptibility to ciprofloxacin, and that only this subpopulation was selectively reduced.
The oral indigenous plaque community structure of the Sprague-Dawley rats used in this study was relatively simple compared to that of humans (Keijser, et al. 2008, Zaura, et al. 2009) or dogs (Dewhirst, et al. 2012). The predominance of the Rothia genus in the plaque community was unexpected. Rothia species are isolated from the human oral microbiome with moderate frequency (particularly Rothia denticariosa (Pardi, et al. 1996) but also R. mucilaginosa (Zamakhchari, et al. 2011)) but are minor components of the community, while Rothia species were not detected in the (subgingival) plaque of 51 breeds of dogs using clone libraries (Dewhirst, et al. 2012). The oral bacterial composition of Wistar Kyoto rats was previously characterized using culture dependent methods (Isogai, et al. 1985). All locations sampled (saliva, tongue, buccal mucosa and gingival crevice) were dominated by Streptococcus species and Rothia species were not detected. The taxon classification was made using medium selection under anaerobic conditions, which was supplemented with identification using morphological characteristics. Rothia species grow poorly, if at all, under anaerobic conditions (Brown, et al. 1969, Zamakhchari, et al. 2011) and Rothia species have previously been mistaken for Streptococci on the basis of cellular morphology (Brown, et al. 1969). Supragingival plaque samples collected by us from a different group of Sprague-Dawley rats in the same laboratory at a different time and analyzed to obtain preliminary pyrosequencing data were dominated by Rothia and Streptococcus genera in the same rank order abundance as in Table 1, and the classifications at the species level within these groups were almost identical (data not shown). This suggests that at least within this species of rat the oral plaque community structure is fairly stable with respect to the dominant taxa.
There were a number of genera in the plaque community present at low abundance that were previously associated with periodontal disease (Exiguobacterium (Zijnge, et al. 2003), Enterococcus (Sun, et al. 2012), Granulicatella (Colombo, et al. 2009), Neisseria (Uzel, et al. 2012), Acinetobacter (da Silva, et al. 2005) and Micrococcus (Anesti, et al. 2005)). In light of a previous study in Wistar rats, it would be of interest to see if any of these taxa become relatively abundant during ligature induced periodontitis (Duarte, et al. 2010). Some of genera present at low abundance are found most commonly in marine environments (Paramoritella, Alkaliflexus and Janibacter). These may have been introduced by the feed which contained fish meal. The presence of Halomonas was unexpected since this genus survives optimally in high salinity environments (>3 % NaCl) (Stevens, et al. 2009) and has not been previously detected in the oral microbiome (to our knowledge).
In summary, our results indicate that it is possible to reduce a phylogenetically defined portion of the microbiome by an antimicrobial dosing regime without causing a large perturbation in the overall community structure. The idea of selecting antimicrobials on the basis of their ability to target specific periodontal pathogens is not new (Beikler, et al. 2004). Our results suggest that it is worth exploring this treatment option more actively as a possible means to kill periodontal pathogens while preserving potentially beneficial commensals. The concentration of the low dose of ciprofloxacin was 10 times the concentration required to selectively kill the periodontal pathogen Aggregatibacter actinomycetemcomitans (Aa) in biofilms composed of Aa and S. sanguinis (Suci and Young 2011). Therefore, it would be of interest to characterize the ability of ciprofloxacin to selectively kill Aa during the progression of periodontal disease induced by this microbe in rat models (Bezerra, et al., Freire, et al. 2011, Li, et al. 2010, Schreiner, et al. 2010).
ACKNOWLEDGEMENTS
This work was supported by NIH grant 1R21DE019237-01A1 to M.Y.
Footnotes
CONFLICT OF INTEREST
None.
REFERENCES
- Ahmed NA, Petersen FC, Scheie AA. AI-2/LuxS Is Involved in Increased Biofilm Formation by Streptococcus intermedius in the Presence of Antibiotics. Antimicrobial Agents and Chemotherapy. 2009;53:4258–4263. doi: 10.1128/AAC.00546-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anesti V, McDonald IR, Ramaswamy M, et al. Isolation and molecular detection of methylotrophic bacteria occurring in the human mouth. Environmental Microbiology. 2005;7:1227–1238. doi: 10.1111/j.1462-2920.2005.00805.x. [DOI] [PubMed] [Google Scholar]
- Appelbaum PC, Hunter PA. The fluoroquinolone antibacterials: past, present and future perspectives. International Journal of Antimicrobial Agents. 2000;16:5–15. doi: 10.1016/s0924-8579(00)00192-8. [DOI] [PubMed] [Google Scholar]
- Avery RK. Adding gram-positive prophylaxis to fluoroquinolone regimens reduces streptococcal bacteraemias, but prophylaxis remains controversial. Cancer Treatment Reviews. 2004;30:385–388. doi: 10.1016/j.ctrv.2004.03.002. [DOI] [PubMed] [Google Scholar]
- Beikler T, Prior K, Ehmke B, et al. Specific antibiotics in the treatment of periodontitis - A proposed strategy. Journal of Periodontology. 2004;75:169–175. doi: 10.1902/jop.2004.75.1.169. [DOI] [PubMed] [Google Scholar]
- Bezerra BD, Andriankaja O, Kang J, et al. A. actinomycetemcomitans-induced periodontal disease promotes systemic and local responses in rat periodontium. Journal of Clinical Periodontology. 2012;39:333–341. doi: 10.1111/j.1600-051X.2011.01847.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Binder D, Zbinden R, Widmer U, et al. Native and prosthetic valve endocarditis caused by Rothia dentocariosa: Diagnostic and therapeutic considerations. Infection. 1997;25:22–26. doi: 10.1007/BF02113502. [DOI] [PubMed] [Google Scholar]
- Brown JM, Georg LK, Waters LC. Laboratory Identification of Rothia Dentocariosa and Its Occurrence in Human Clinical Materials. Applied Microbiology. 1969;17:150-&. doi: 10.1128/am.17.1.150-156.1969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chao A. Nonparametric-Estimation of the Number of Classes in a Population. Scandinavian Journal of Statistics. 1984;11:265–270. [Google Scholar]
- Colombo APV, Boches SK, Cotton SL, et al. Comparisons of Subgingival Microbial Profiles of Refractory Periodontitis, Severe Periodontitis, and Periodontal Health Using the Human Oral Microbe Identification Microarray. Journal of Periodontology. 2009;80:1421–1432. doi: 10.1902/jop.2009.090185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- da Silva RM, Caugant DA, Lingaas PS, et al. Detection of Actinobacillus actinomycetemcomitans but not bacteria of the red complex in aortic aneurysms by multiplex polymerase chain reaction. Journal of Periodontology. 2005;76:590–594. doi: 10.1902/jop.2005.76.4.590. [DOI] [PubMed] [Google Scholar]
- del Prado G, Ruiz V, Naves P, et al. Biofilm formation by Streptococcus pneumoniae strains and effects of human serum albumin, ibuprofen, N-acetyl-L-cysteine, amoxicillin, erythromycin, and levofloxacin. Diagnostic Microbiology and Infectious Disease. 2010;67:311–318. doi: 10.1016/j.diagmicrobio.2010.03.016. [DOI] [PubMed] [Google Scholar]
- Dethlefsen L, Huse S, Sogin ML, et al. The Pervasive Effects of an Antibiotic on the Human Gut Microbiota, as Revealed by Deep 16S rRNA Sequencing. Plos Biology. 2008;6:2383–2400. doi: 10.1371/journal.pbio.0060280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dewhirst FE, Klein EA, Thompson EC, et al. The Canine Oral Microbiome. Plos One. 2012;7 doi: 10.1371/journal.pone.0036067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duarte PM, Tezolin KR, Figueiredo LC, et al. Microbial profile of ligature-induced periodontitis in rats. Archives of Oral Biology. 2010;55:142–147. doi: 10.1016/j.archoralbio.2009.10.006. [DOI] [PubMed] [Google Scholar]
- Endt K, Maier L, Kappeli R, et al. Peroral Ciprofloxacin Therapy Impairs the Generation of a Protective Immune Response in a Mouse Model for Salmonella enterica Serovar Typhimurium Diarrhea, while Parenteral Ceftriaxone Therapy Does Not. Antimicrobial Agents and Chemotherapy. 2012;56:2295–2304. doi: 10.1128/AAC.05819-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Excoffier L, Smouse PE, Quattro JM. Analysis of Molecular Variance Inferred from Metric Distances among DNA Haplotypes - Application to Human Mitochondrial-DNA Restriction Data. Genetics. 1992;131:479–491. doi: 10.1093/genetics/131.2.479. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Freire MO, Sedhgizaden PP, Schaudinn C, et al. Development of an animal model for Aggregatibacter actinomycetemcomitans biofilm-mediated oral osteolytic infection: a preliminary study. Journal of Periodontalogy. 2011;82:778–789. doi: 10.1902/jop.2010.100263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guo DW, Wang LP, Lu CP. In Vitro Biofilm Forming Potential of Streptococcus Suis Isolated from Human and Swine in China. Brazilian Journal of Microbiology. 2012;43:993–1004. doi: 10.1590/S1517-838220120003000021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hogan JS, Pankey JW, Duthie AH. Growth-Responses of Staphylococcus-Aureus and Streptococcus-Agalactiae to Corynebacterium-Bovis Metabolites. Journal of Dairy Science. 1987;70:1294–1301. doi: 10.3168/jds.S0022-0302(87)80144-3. [DOI] [PubMed] [Google Scholar]
- Hoogkamp-Korstanje JAA, Roelofs-Willemse J. Comparative in vitro activity of moxifloxacin against Gram-positive clinical isolates. Journal of Antimicrobial Chemotherapy. 2000;45:31–39. doi: 10.1093/jac/45.1.31. [DOI] [PubMed] [Google Scholar]
- Huttenhower C, Gevers D, Knight R, et al. Structure, function and diversity of the healthy human microbiome. Nature. 2012;486:207–214. doi: 10.1038/nature11234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Isogai E, Isogai H, Sawada H, et al. Microbial Ecology of Plaque in Rats with Naturally-Occurring Gingivitis. Infection and Immunity. 1985;48:520–527. doi: 10.1128/iai.48.2.520-527.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kawai T, Eisen-Lev R, Seki M, et al. Requirement of B7 costimulation for Th1-mediated inflammatory bone resorption in experimental periodontal disease. Journal of Immunology. 2000;164:2102–2109. doi: 10.4049/jimmunol.164.4.2102. [DOI] [PubMed] [Google Scholar]
- Kawai T, Shimauchi H, Eastcott JW, et al. Antigen direction of specific T-cell clones into gingival tissues. Immunology. 1998;93:11–19. doi: 10.1046/j.1365-2567.1998.00408.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keijser BJF, Zaura E, Huse SM, et al. Pyrosequencing analysis of the Oral Microflora of healthy adults. Journal of Dental Research. 2008;87:1016–1020. doi: 10.1177/154405910808701104. [DOI] [PubMed] [Google Scholar]
- Krisanaprakornkit S, Kimball JR, Weinberg A, et al. Inducible expression of human beta-defensin 2 by Fusobacterium nucleatum in oral epithelial cells: Multiple signaling pathways and role of commensal bacteria in innate immunity and the epithelial barrier. Infection and Immunity. 2000;68:2907–2915. doi: 10.1128/iai.68.5.2907-2915.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li Y, Messina C, Bendaoud M, et al. Adaptive immune response in osteoclastic bone resorption induced by orally administered Aggregatibacter actinomycetemcomitans in a rat model of periodontal disease. Molecular Oral Microbiology. 2010;25:275–292. doi: 10.1111/j.2041-1014.2010.00576.x. [DOI] [PubMed] [Google Scholar]
- Lozupone C, Hamady M, Knight R. UniFrac - An online tool for comparing microbial community diversity in a phylogenetic context. Bmc Bioinformatics. 2006;7 doi: 10.1186/1471-2105-7-371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mans JJ, von Lackum K, Dorsey C, et al. The degree of microbiome complexity influences the epithelial response to infection. Bmc Genomics. 2009;10 doi: 10.1186/1471-2164-10-380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Neish AS, Gewirtz AT, Zeng H, et al. Prokaryotic regulation of epithelial responses by inhibition of I kB-alpha ubiquitination. Science. 2000;289:1560–1563. doi: 10.1126/science.289.5484.1560. [DOI] [PubMed] [Google Scholar]
- Otero VN, Posse JL, Carmona IT, et al. Efficacy of Fluoroquinolones Against Pathogenic Oral Bacteria. Mini-Reviews in Medicinal Chemistry. 2009;9:1147–1158. doi: 10.2174/138955709789055243. [DOI] [PubMed] [Google Scholar]
- Pardi G, Acevedo AM, DeIlja RM, et al. Studies over Rothia Dentocariosa present in patients with and without dental caries. Journal of Dental Research. 1996;75:1321–1321. [Google Scholar]
- Pflughoeft KJ, Versalovic J. Human Microbiome in Health and Disease. Annual Review of Pathology: Mechanisms of Disease. 2012;7:99–122. doi: 10.1146/annurev-pathol-011811-132421. [DOI] [PubMed] [Google Scholar]
- Pruesse E, Quast C, Knittel K, et al. SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Research. 2007;35:7188–7196. doi: 10.1093/nar/gkm864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pultz NJ, Hoyen CK, Donskey CJ. Inhibition of methicillin-resistant Staphylococcus aureus by an in vitro continuous-flow culture containing human stool microflora. Fems Microbiology Letters. 2004;241:201–205. doi: 10.1016/j.femsle.2004.10.021. [DOI] [PubMed] [Google Scholar]
- Rakoff-Nahoum S, Paglino J, Eslami-Varzaneh F, et al. Recognition of commensal microflora by toll-like receptors is required for intestinal homeostasis. Cell. 2004;118:229–241. doi: 10.1016/j.cell.2004.07.002. [DOI] [PubMed] [Google Scholar]
- Schloss PD, Westcott SL, Ryabin T, et al. Introducing mothur: Open-Source, Platform-Independent, Community-Supported Software for Describing and Comparing Microbial Communities. Applied and Environmental Microbiology. 2009;75:7537–7541. doi: 10.1128/AEM.01541-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schreiner H, Markowitz K, Miryalkar M, et al. Aggregatibacter actinomycetemcomitans-Induced Bone Loss and Antibody Response in Three Rat Strains. Journal of Periodontology. 2010;82:142–150. doi: 10.1902/jop.2010.100250. [DOI] [PubMed] [Google Scholar]
- Sears CL. A dynamic partnership: Celebrating our gut flora. Anaerobe. 2005;11:247–251. doi: 10.1016/j.anaerobe.2005.05.001. [DOI] [PubMed] [Google Scholar]
- Stevens DA, Hamilton JR, Johnson N, et al. Halomonas, a Newly Recognized Human Pathogen Causing Infections and Contamination in a Dialysis Center Three New Species. Medicine. 2009;88:244–249. doi: 10.1097/MD.0b013e3181aede29. [DOI] [PubMed] [Google Scholar]
- Suci P, Young M. Selective killing of Aggregatibacter actinomycetemcomitans by ciprofloxacin during development of a dual species biofilm with Streptococcus sanguinis. Archives of Oral Biology. 2011;56:1055–1063. doi: 10.1016/j.archoralbio.2011.03.013. [DOI] [PubMed] [Google Scholar]
- Sun J, Sundsfjord A, Song X. Enterococcus faecalis from patients with chronic periodontitis: virulence and antimicrobial resistance traits and determinants. European Journal of Clinical Microbiology & Infectious Diseases. 2012;31:267–272. doi: 10.1007/s10096-011-1305-z. [DOI] [PubMed] [Google Scholar]
- Tamura K, Dudley J, Nei M, et al. MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Molecular Biology and Evolution. 2007;24:1596–1599. doi: 10.1093/molbev/msm092. [DOI] [PubMed] [Google Scholar]
- Teughels W, Haake SK, Sliepen I, et al. Bacteria interfere with A. actinomycetemcomitans colonization. Journal of Dental Research. 2007;86:611–617. doi: 10.1177/154405910708600706. [DOI] [PubMed] [Google Scholar]
- Ubeda C, Taur Y, Jenq RR, et al. Vancomycin-resistant Enterococcus domination of intestinal microbiota is enabled by antibiotic treatment in mice and precedes bloodstream invasion in humans. Journal of Clinical Investigation. 2010;120:4332–4341. doi: 10.1172/JCI43918. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uzel NG, Teles FR, Teles RP, et al. Microbial shifts during dental biofilm re-development in the absence of oral hygiene in periodontal health and disease. Journal of Clinical Periodontology. 2012;38:612–620. doi: 10.1111/j.1600-051X.2011.01730.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vela AI, Casamayor A, del Rey VS, et al. Streptococcus plurextorum sp nov., isolated from pigs. International Journal of Systematic and Evolutionary Microbiology. 2009;59:504–508. doi: 10.1099/ijs.0.65636-0. [DOI] [PubMed] [Google Scholar]
- Wang Q, Garrity GM, Tiedje JM, et al. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Applied and Environmental Microbiology. 2007;73:5261–5267. doi: 10.1128/AEM.00062-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xue H, Field CJ, Sawyer MB, et al. Prophylactic ciprofloxacin treatment prevented high mortality, and modified systemic and intestinal immune function in tumour-bearing rats receiving dose-intensive CPT-11 chemotherapy. British Journal of Cancer. 2009;100:1581–1588. doi: 10.1038/sj.bjc.6605051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yue JC, Clayton MK. A similarity measure based on species proportions. Communications in Statistics-Theory and Methods. 2005;34:2123–2131. [Google Scholar]
- Zamakhchari M, Wei GX, Dewhirst F, et al. Identification of Rothia Bacteria as Gluten-Degrading Natural Colonizers of the Upper Gastro-Intestinal Tract. Plos One. 2011;6 doi: 10.1371/journal.pone.0024455. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zaura E, Keijser BJF, Huse SM, et al. Defining the healthy "core microbiome" of oral microbial communities. Bmc Microbiology. 2009;9 doi: 10.1186/1471-2180-9-259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zijnge V, Harmsen HJM, Kleinfelder JW, et al. Denaturing gradient gel electrophoresis analysis to study bacterial community structure in pockets of periodontitis patients. Oral Microbiology and Immunology. 2003;18:59–65. doi: 10.1034/j.1399-302x.2003.180110.x. [DOI] [PubMed] [Google Scholar]