Background: Retrotransposon LINE-1 causes dozens of genetic diseases.
Results: Human MOV10 diminishes the level of LINE-1 RNA by acting at a post-transcriptional stage.
Conclusion: The host protein suppresses LINE-1 transposition.
Significance: MOV10 contributes to the cellular control of LINE-1 replication.
Keywords: Host-Pathogen Interactions, Reverse Transcription, RNA Helicase, Viral Replication, Virology, LINE-1, MOV10, Post-transcriptional, Restriction, Retrotransposon
Abstract
LINE-1 (long interspersed element 1) is an autonomous non-long terminal repeat retrotransposon. Its replication often causes mutation and rearrangement of host genomic DNA. Accordingly, host cells have evolved mechanisms to control LINE-1 mobility. Here, we report that a helicase named MOV10 effectively suppresses LINE-1 transposition. Mutating the helicase motifs impairs this function of MOV10, suggesting that MOV10 requires its helicase activity to suppress LINE-1 replication. Further studies show that MOV10 post-transcriptionally diminishes the level of LINE-1 RNA. The association of MOV10 with both LINE-1 RNA and ORF1 suggests that MOV10 interacts with LINE-1 RNP and consequently causes its RNA degradation. These data demonstrate collectively that MOV10 contributes to the cellular control of LINE-1 replication.
Introduction
LINE-1 (long interspersed element 1) is a non-long terminal repeat (LTR)4 retrotransposon that needs to convert its RNA into DNA to “jump” from one site in the genome to the other. Human genome contains more than 500,000 copies of LINE-1, among which ∼80–100 copies are capable of retrotransposition (1). Of all transposons in human genome including DNA transposons, LTR retrotransposons (also called endogenous retroviruses) and non-LTR retrotransposons (including LINEs and SINEs (short interspersed elements)), LINE-1 is the only active autonomous transposon in human genome. LINE-1 proteins also support the retrotransposition of SINES including Alu and SVA that do not encode their own proteins. The actions of LINE-1, Alu, and SVA cause dozens of genetic diseases (2, 3).
LINE-1 is ∼6,000 nucleotides in length and carries two open reading frames named ORF1 and ORF2, both of which are essential for LINE-1 replication (4). ORF1 is a nucleic acid chaperone protein and binds to LINE-1 RNA preferentially in cis (5). ORF2 has two enzymatic activities: endonuclease and reverse transcriptase (6, 7). The endonuclease domain is responsible for recognizing the target sequence (AATTTT) in host genome and generating a nick in one DNA strand. The reverse transcriptase then uses the exposed 3′ OH to prime the reverse transcription of LINE-1 RNA into DNA. This mechanism is named target site-primed reverse transcription (4). In cultured cells, the LINE-1 RNA-ORF1-ORF2 complex is observed as cytoplasmic foci that are often found adjacent to stress granules (8–10).
LINE-1 transposition poses various deleterious effects on the structure and function of host genome (11). For example, LINE-1 can cause insertion, deletion, and recombination of host DNA (2). LINE-1 can also modulate the expression of host genes (2). Host cells have therefore evolved a number of mechanisms to control LINE-1 replication (12). One strategy involves DNA methylation that silences LINE-1 RNA synthesis. The 5′-UTR of LINE-1 serves as an internal promoter to drive LINE-1 RNA production (13). This region contains multiple CpG islands that are subject to methylation, which suppresses LINE-1 transcription (12, 14). Indeed, mutation or knock-out of the DNA methyltransferases in mice such as Dnmt1, Dnmt3A, Dnmt3B, and Dnmt3L leads to hypomethylation of LINE-1 and other transposons and concomitantly a high level expression of these elements (15–17). LINE-1 is most active in embryonic stem cells, germ line cells, and certain somatic cells (18–21). One cause for this high LINE-1 activity in embryonic stem cells and germ line cells is the temporal erasure of DNA methylation (22–24). To control massively activated LINE-1, germ line cells express a special type of small RNA named piRNA (piwi-interacting RNA) that arms Piwi (P-element induced wimpy testis) proteins to suppress LINE-1 (25, 26). Furthermore, it has been reported that APOBEC3 (apolipoprotein B mRNA-editing, enzyme-catalytic, polypeptide-like 3) proteins, which inhibit the infection of a broad range of viruses particularly retroviruses (27–31), also provide another mechanism to control LINE-1 (32–34). It appears that cells manage to use the same mechanism to protect themselves from either extracellular or intracellular insults that share common biochemical features. One mechanism that was discovered to inhibit one type of virus is often effective against others. This scenario is again manifested on the discovery of the antiviral function of one cellular helicase, Moloney leukemia virus type 10 protein (MOV10).
The 110-kDa MOV10 protein was first identified by provirus integration in the Mov10 mouse strain as an ATP-binding protein (35) and later classified as a member of the superfamily 1 (SF-1) RNA helicase for consisting of seven highly conserved helicase motifs in its C-terminal region (36). It is well known that helicases are enzymes that have ATP-dependent DNA or RNA “structure remodeling” activity including classical double-stranded nucleic acid unwinding activity. The two largest helicase groups are superfamilies 1 and 2 (SF-1 and SF-2), which carry seven characteristic motifs (I, Ia, II, III, IV, V, and VI) respectively. Motifs I and II (Walker A and B motifs, respectively) are the fingerprints conserved across the SF-1 and SF-2 helicases. Both motifs that present residues interacting with MgATP/MgADP are essential for ATP binding and ATP hydrolysis (37).
MOV10 was reported being associated with argonaute proteins (38). This association endows MOV10 with the role of regulating the activity of microRNA machinery (38). As a result of this function, MOV10 was shown to modulate the synaptic plasticity by suppressing local protein translation at synapses (39). Several lines of evidence implicate that MOV10 also affects a wide range of RNA viruses as a co-factor or restriction factor. For example, MOV10 is required for RNA-directed transcription in hepatitis delta virus (40), whereas it inhibits hepatitis C virus replication (41). Recently, several reports have demonstrated that MOV10 inhibits the replication of lentiviruses including human immunodeficiency virus type 1 (HIV-1). This inhibition depends on the incorporation of MOV10 into HIV-1 particles and the impediment of viral reverse transcription (42–44). In this study, we report that MOV10 also restricts the mobilization of LINE-1, which further demonstrates MOV10 as an important player in host defense against exogenous and endogenous parasitic genetic elements.
EXPREIMENTAL PROCEDURES
Plasmids
CMV-L1-neoRT carries a copy of complete human LINE-1 DNA that bears a neomycin resistance gene as a reporter of LINE-1 retrotransposition (45). The IAP-neoTNF and MusD-neoTNF plasmids express mouse LTR retrotransposons containing a similar neomycin resistance cassette placed in reverse direction (46). The neomycin resistance gene is inactive by the presence of a forward intron that is removed during RNA splicing, resulting in a functional gene after reverse transcription and integration. JM101/L1.3 has a full-length human L1.3 element without any inserted reporter (47). The FLAG/HA-MOV10 DNA clone encodes an N-terminal FLAG/HA-tagged human MOV10 helicase (38). The amino acids Lys-531 and Glu-647 were changed to Arg and Gln, respectively, using site-directed mutagenesis kit (Stratagene) to create two mutants MOV10 (KR) and MOV10 (EQ). The LINE-1 5′-UTR was amplified from CMV-L1-neoRT using primers 5′-AAACCGCTCGAGGAATAGGAACAGCTC-3′ and 5′-TTTCCCAAGCTTCTTTGTGGTTTTATC-3′. The PCR products were digested with restriction enzymes XhoI and HindIII and then inserted to the 5′ end of firefly luciferase gene in the pGL3-Basic vector (Promega) to create L1-Luc reporter DNA.
Cell Colony Assay
HEK293 cells or HeLa cells were seeded in 6-well plates 1 day prior to transfection with 750 ng of the CMV-L1-neoRT, IAP-neoTNF, or MusD-neoTNF reporter DNA. Forty hours after transfection, cells were detached from the plates with trypsin and seeded into 6-well plates at serial dilutions. G418 (0.75 mg/ml) was then added to select for resistant cell colonies. Twelve days into selection when cell colonies were clearly visible, the cells were fixed with methanol at room temperature for 10 min and stained with 0.5% crystal violet (in 25% methanol) for 10 min (48). The p values were calculated with Student's t test.
Detection of LINE-1 DNA by PCR
HeLa cells were first transfected with 750 ng of the CMV-L1-neoRT, IAP-neoTNF, or MusD-neoTNF reporter DNA with or without the same amount of MOV10 DNA. Seventy-two hours later, total cellular DNA was extracted with the QIAamp DNA mini kit (Qiagen). The same amounts of DNA (250 ng) were subject to PCR using a pair of primers 5′-CAGTTCGGCTGGCGCGAGGCC-3′/5′-CAGTTCCGCCCATTCTCCG-3 (49). The primer 5′-CAGTTCGGCTGGCGCGAGGCC-3′ crosses the exon/intron junction within the neomycin resistance gene such that only the spliced and reverse transcribed DNA can be amplified. The PCR was performed with the AccuPrime Taq DNA polymerase (Invitrogen) for 25–38 cycles under the following conditions: 94 °C for 50 s, 60 °C for 50 s, and 68 °C for 1 min. The DNA products were separated by electrophoresis in 1% agarose gels and visualized with ethidium bromide staining.
Detection of LINE-1 RNA by PCR
HEK293 cells were co-transfected with CMV-L1-neoRT and MOV10-expressing plasmid. An equal amount of total cellular RNA was treated with DNase to remove the potential contamination of plasmid DNA. The random primer was used for reverse transcription. The cDNA products were normalized by the GAPDH. The unspliced form of LINE-1 RNA was detected by PCR using the primer Fneo (5′-AATATCACGGGTAGCCAA-3′) and Rintron (5′-CTGGGGTGGCCTAGCTAAC-3′), whereas the spliced form of LINE-1 RNA was detected by PCR using the primer Fneo and Rneo (5′-GAAGCCGGTCTTGTCGATC-3′).
Northern Blotting to Measure LINE-1 RNA
HEK293 cells were transfected with CMV-L1-neoRT, IAP-neoTNF, or MusD-neoTNF reporter DNA. Total cellular RNA was extracted with TRIzol reagent (Invitrogen) 40 h after transfection. The same amounts of RNA samples were subject to electrophoresis in 1% denaturing agarose gels. After being transferred onto nylon membranes, RNA was hybridized to 32P-labeled DNA probes that were amplified using primers 5′-CGGATCAAGCGTATGCAGCC-3′/5′-GCAGGTTCTCCGGCCGCTTG-3′ and recognized the neomycin resistance gene sequence. The same probes bind to the RNA of CMV-L1-neoRT, IAP-neoTNF, and MusD-neoTNF. When JM101/L1.3 was used in transfection, a DNA fragment of L1.3 (nucleotide positions 1199–1533; L19088) was labeled with 32P and used as the probes in Northern blotting. After extensive washing, the radioactive signals were exposed to x-ray films. The GAPDH mRNA was detected as the internal control (50).
Detection of Endogenous LINE-1 RNA by RT-PCR
The endogenous LINE-1 transcript was measured using a RT-PCR protocol described previously (51, 52). In brief, total RNA was extracted from HeLa cells with TRIzol reagent, followed by purifying the poly(A) mRNA using the PolyATract mRNA isolation system III kit (Promega). The first strand cDNA was synthesized with the ORF2(−) primer (5′-CTG TGT CTT TTA ATT GCA GAA TTT AGT CC-3′) using an RT-PCR kit (Promega) in accordance with the manufacturer's protocol. This cDNA was then amplified with the forward primer (5′-AAG CTT GAA TTC GGG GGA GGA GCC AAG ATG GCC G-3′) and the nORF2(−) primer (5′-TTA ATT AAT AGT CCA TTT ATA TTT AAA G-3′). The resulted PCR products were used in the second round of PCR that was performed with the nested forward primer (5′-GGA GCC AAG ATG GCC GAA TAG GAA CAG CT-3′) and the nORF2(−) primer. The final PCR products were analyzed by gel electrophoresis. DNA was excised and purified with the QIAquick gel extraction kit (Qiagen), followed by sequencing to confirm the identity as LINE-1 sequence.
siRNA Knockdown
The endogenous MOV10 protein was knocked down with siRNA oligonucleotides 5′-GAAACCCTGTGGTGACCAA-3′ that target MOV10 mRNA at nucleotide positions 1058–1076 (NM_0011 30079.1) (38). The siRNA oligonucleotides 5′-GCACGGAAGTCCATCTGAA-3′ (siRNA1) and 5′-GCAGGACAAAGATGTATTA-3′ (siRNA2) target Ago2 mRNA at nucleotide positions 1426–1444 and 1083–1101 (NM_001164623.1), respectively (53). The control siRNA was purchased from Qiagen (catalogue number 1027310). HEK293 cells were seeded into 6-well plates 1 day prior to two sequential transfections of siRNA oligonucleotides (25 nm) using Lipofectamine 2000 (Invitrogen). The efficiency of siRNA knockdown was assessed by Western blotting using anti-MOV10 (Abcam; ab80613) or anti-Ago2 (Abnova) antibodies.
RNA Stability Assay
HEK293 cells were co-transfected with CMV-L1-neoRT and the plasmid encoding MOV10. Twenty hours after transfection, the cells were treated with actinomycin D for 0, 0.5, 1, 2, 4, or 8 h, and then the total cellular RNA was extracted and analyzed by Northern blotting using probes specific for LINE-1 and GAPDH, respectively, as described above.
Immunoprecipitation Assay
HEK293 cells were co-transfected with MOV10 DNA and plasmid encoding Myc-tagged ORF1. The cells were collected 48 h post-transfection and then lysed in 350 μl of TNT buffer (20 mm Tris-HCl, pH 7.5, 200 mm NaCl, 1% Triton X-100). Insoluble material was pelleted at 1800 × g for 30 min. Equal amounts of supernatant were incubated with 5 μl of MOV10-specific antibody (Abcam; rabbit) or anti-Myc antibody (Abcam; mouse) for 16 h at 4 °C, followed by the addition of protein A-Sepharose (Amersham Biosciences) for 2 h. The immune-precipitated complex was then washed three times with the TNT buffer and phosphate-buffered saline, followed by Western blot analysis using anti-Myc (mouse) or MOV10-specific antibody, respectively. To detect the possible association of MOV10 with LINE-1 RNA, HEK293 cells were transfected with MOV10 DNA and the CMV-L1-neoRT DNA. The expressed MOV10-FLAG was immune-precipitated with anti-FLAG antibody as described above. The RNA associated with the precipitated complex was extracted with TRIzol agents (Invitrogen) and subjected to RT-PCR using primers that amplify the neo gene as described above.
Immunofluorescence Microscopy
HEK293 cells were seeded in a four-chamber slide 1 day prior to transfection with plasmids expressing FLAG-tagged MOV10, MOV10EQ, or MOV10KR. The cells were fixed with 4% paraformaldehyde (in 1× phosphate-buffered saline) for 10 min at room temperature and permeabilized with 4% paraformaldehyde and 0.1% Triton X-100 for 10 min at room temperature. The cells were then stained for 2 h at room temperature with antibodies against FLAG (1:500 dilution, mouse). After being washed with 1× phosphate-buffered saline, the cells were incubated with either Alexa Fluor 488-conjugated secondary anti-mouse antibody (1:2,000 dilution; Invitrogen). The images were recorded using the Zeiss Pascal laser scanning confocal microscope.
RESULTS
MOV10 Overexpression Diminishes LINE-1 Retrotransposition
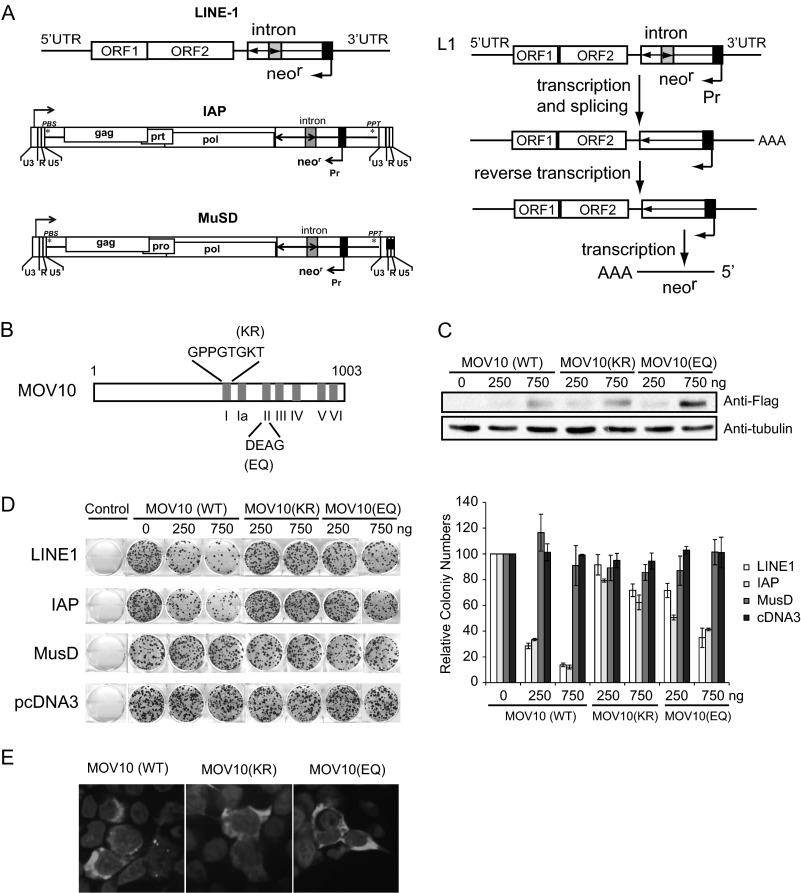
To monitor LINE-1 activity, we used in the following experiments a CMV-L1-neoRT reporter construct that has a neomycin resistance gene inserted in the 3′-UTR of LINE-1 in the opposite direction from the LINE-1 coding sequence (Fig. 1A) (45). Transcription of neomycin resistance gene is driven by the immediate early promoter of CMV. One important feature of this neomycin resistance gene is an intron sequence that has been inserted in the same direction of LINE-1 coding sequence. Such a design ensures that a functional neomycin resistance mRNA can only be produced from the reverse transcribed LINE-1 DNA in which the intron should have been removed during RNA splicing. Therefore, the number of G418-resistant cell colonies reflects the events of LINE-1 retrotransposition (45). We also included in our experiments two mouse endogenous retroviruses IAP-neoTNF and MusD-neoTNF (Fig. 1A) that are LTR retrotransposons and also carry the neomycin resistance gene as the selection marker (46). We noted that the neoRT and neoTNF cassettes differ in the identity of the inserted intron sequence. Because this study does not intend to compare the retrotransposition efficiency between L1, IAP, and MusD but rather to test the effect of MOV10 on each of these three retroelements, interpretation of results in the following studies should not be affected by the difference in neoRT and neoTNF. As a control for the specificity of MOV10 inhibition, a plasmid DNA named pcDNA3.1 (Invitrogen), which carries the neomycin resistance gene, was transfected into cells followed by selection for G418-resistant cell colonies.
FIGURE 1.
Overexpression of MOV10 inhibits LINE-1 retrotransposition in HeLa cells. A, illustration of the CMV-L1-neoRT, IAP-neoTNF, and MusD-neoTNF reporter (left panel) and the generation of a functional neomycin resistance mRNA (right panel). Following transcription from the 5′-UTR promoter of LINE-1, the intron in the neomycin resistance gene is excised. This intronless mRNA is then reverse transcribed into cDNA that is able to produce a functional neomycin resistance mRNA. B, domain structure of the MOV10 helicase. The positions of the mutated amino acids Lys and Glu in the Walk A (GKT) and Walker B (DEAG) motifs are indicated. C, Western blots to show the expression of MOV10 and its mutants in transiently transfected HeLa cells. Tubulin was probed as an internal control. D, G418 resistance colony assays. HeLa cells were transfected with 750 ng of the CMV-L1-neoRT, IAP-neoTNF, MusD-neoTNF, or pcDNA3.1 DNA with or without MOV10 DNA or its mutants. Control represents untransfected HeLa cells that also underwent G418 selection. pcDNA3.1 is a plasmid DNA that carries the neomycin resistance gene and thus confers resistance to G418. Colonies were visualized with crystal violet staining. The results of a representative colony assay are shown. The data from three independent experiments are summarized in the bar graph. The number of cell colonies in the absence of MOV10 is arbitrarily set as 100. E, cellular location of MOV10 and its mutants. FLAG-tagged MOV10 and its mutants (MOV10EQ and MOV10KR) were transiently expressed in HEK293 cells and detected by immunostaining with anti-FLAG antibodies (pseudocolored in green).
When the wild type MOV10 was expressed ectopically from the plasmid DNA, the number of G418-resistant HeLa colonies diminished by 3.5- and 7.3-fold (p < 0.05) with 250 and 750 ng of MOV10 DNA being transfected, respectively (Fig. 1D). A similar degree of inhibition was observed for IAP, whereas no significant change was measured for MusD or pcDNA3.1 (Fig. 1D). These results suggest that MOV10 specifically inhibits LINE-1 and IAP. Because MusD-neoTNF also uses the CMV immediate early promoter to express neomycin phosphotransferase as IAP-neoTNF does, we conclude that the observed inhibition of L1 and IAP by MOV10 does not result from disrupting neomycin phosphotransferase expression.
We next determined whether the helicase motifs in MOV10 are required for its inhibition on the replication of LINE-1 and IAP. As a helicase, MOV10 bears the GKT (Walker A) and the DEAG (Walker B) motifs (43). The Walker A motif has the consensus sequence (G/A) GK (T/S), whereas the Walker B motif has the DEXX consensus sequence. Both motifs are essential for ATP binding and ATP hydrolysis (54). Mutating either motif in other helicase has been shown to severely impair its activity. Accordingly, we generated two MOV10 mutants, MOV10 (KR) and MOV10 (EQ), that had the GKT and the DEAG motifs mutated (Fig. 1B), respectively, and measured the effects of these two mutants on LINE-1 retrotransposition. These two mutants were expressed at levels similar to that of the wild type MOV10 (Fig. 1C). The results of colony assays showed that MOV10 (KR) mutant virtually almost lost its ability to inhibit LINE-1 or IAP (Fig. 1D). The results also showed that the MOV10 (EQ) mutant was still able to inhibit LINE-1 or IAP transposition by 2-fold (Fig. 1D), although compared with wild type MOV10, the inhibition activity of MOV10 (EQ) was markedly attenuated. Similar to our findings, Goodier et al. (55) reported that similar mutations in the GKT and the DEAG motifs abolished anti-LINE-1 activity of MOV10. These results collectively indicate that the helicase motifs of MOV10 are required for suppressing the replication of LINE-1 and IAP. Given the important role of the GKT and the DEAG motifs in the helicase function, these data suggest that the helicase activity of MOV10 may be involved in the inhibition.
Recent evidence showed that MOV10 co-localized with LINE-1 RNP in stress granules in the cytoplasmic (55). This raises the possibility that the above mutated GKT and DEAG motifs may alter the cytoplasmic localization of MOV10, consequently affecting its activity against LINE-1. In agreement with previous work (55), we found that wide type and mutant forms of MOV10 localized predominantly in the cytoplasmic (Fig. 1E). Notably, two mutated forms of MOV10 failed to form cytoplasmic foci compared with the wild type form. This suggests that the ability of MOV10 to form the cytoplasmic foci is required for its inhibition of LINE-1 activity, providing further evidence supporting the role of co-localization of MOV10 and LINE-1 RNA in the stress granules in controlling LINE-1 by MOV10.
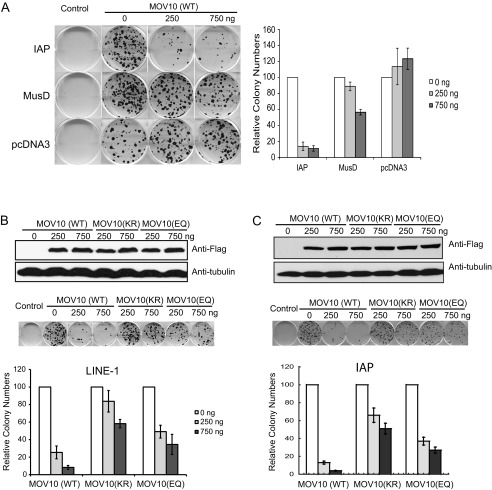
We then asked whether MOV10 was able to suppress LINE-1 activity in cell lines other than HeLa. To this end, we performed similar experiments in human embryonic kidney cell line HEK293. Similarly, wild type MOV10 diminished the activity of LINE-1 or IAP (p < 0.05) by 5–10-fold, yet did not affect the number of G418-resistant colonies that were generated by MusD-neoTNF or pcDNA3.1 (Fig. 2). Again, the MOV10 (KR) mutant lost its ability to inhibit LINE-1 and IAP transposition, whereas the MOV10 (EQ) mutant exerted moderate inhibition (Fig. 2, B and C). Together, these data suggest that MOV10 effectively suppresses the replication of LINE-1 and IAP but not that of MusD and that the GKT and DEAG helicase motifs of MOV10 are important for maximal inhibition.
FIGURE 2.
Overexpression of MOV10 inhibits LINE-1 activity in HEK293 cells. A, effects of MOV10 on the activity of IAP and MusD in transiently transfected HEK293 cells. B and C, effects of MOV10 and its mutants on LINE-1 (B) and IAP (C) retrotransposition. Expression levels of MOV10 and its mutants were assessed by Western blotting (top panels). One representative colony assay is shown (middle panels). The results from three independent experiments are summarized in the bar graph (bottom panels).
Depletion of MOV10 Enhances LINE-1 Mobility
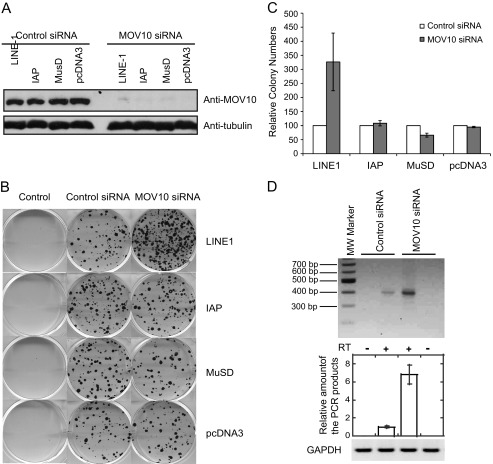
We next depleted the endogenous MOV10 in HEK293 cells using siRNA oligonucleotides and investigated how LINE-1 activity was affected (Fig. 3A). The number of G418-resistant colonies representing LINE-1 transposition increased by 3-fold (p < 0.05) when MOV10 was depleted (Fig. 3, B and C). In contrast, no evident effect of MOV10 knockdown was seen on the activity of IAP, MusD, or pcDNA3.1 (Fig. 3C). These data suggest that LINE-1 retrotransposition is subject to the restriction by the endogenous level of MOV10.
FIGURE 3.
Knockdown of MOV10 increases LINE-1 retrotransposition in HEK293 and HeLa cells. A, Western blots to measure levels of endogenous MOV10 in HEK293 cells that were transfected with control siRNA or siRNA oligonucleotides targeting MOV10 mRNA. B, HEK293 cells were transfected with siRNA oligonucleotides prior to transfection with 750 ng of the CMV-L1-neoRT, IAP-neoTNF, MusD-neoTNF, or pcDNA3.1 DNA. The G418-resistant cell colonies were scored for each transfection. Control represents untransfected HEK293 cells that underwent G418 selection. C, results of three independent transfection experiments shown in B are summarized in the bar graph. D, nest RT-PCR to measure endogenous LINE-1 RNA in HeLa cells. HeLa cells were transfected with siRNA oligonucleotides targeting endogenous MOV10. Equal amounts of total cellular RNA normalized by the GAPDH internal control was used to generate cDNA with (+) or without (−) reverse transcriptase. The cDNA was then amplified using a nest PCR method with primers specific for LINE-1, as described under “Experimental Procedures.” The first lane is molecule weight markers.
To further validate the role of MOV10 in the control of LINE-1 transposition, we investigated the effect of depletion of endogenous MOV10 upon the activity of endogenous LINE-1. We found that MOV10 knockdown resulted in a significant increase (5–7-fold increase) in an approximate 400-bp cDNA product derived from RNA transcript of endogenous LINE-1, which was measured using a nest RT-PCR method (Fig. 3D). Silencing endogenous MOV10 did not affect the level of GAPDH mRNA, suggesting that reduced expression of MOV10 did not result in a general up-regulation of mRNA expression. This provides further evidence on the important role of MOV10 in controlling LINE-1 activity.
IAP is an endogenous retrovirus and differs considerably from non-LTR LINE-1 in terms of genome structure and replication mechanism. Like IAP, HIV-1 was inhibited by the overexpression of MOV10 (43, 44) but insensitive to endogenous MOV10 (56). It is therefore proposed that retroviruses, either of endogenous or exogenous origin, are not inhibited by endogenous MOV10. It is possible that either the level of endogenous MOV10 is not sufficient for inhibiting IAP replication or another host mechanism tightly controls the expression of IAP under normal physiological conditions. Therefore, no significant increases in IAP replication were found even if endogenous MOV10 was absent, whereas overexpression of MOV10 is still able to contribute extra inhibition of IAP replication.
MOV10 Diminishes the Level of LINE-1 cDNA
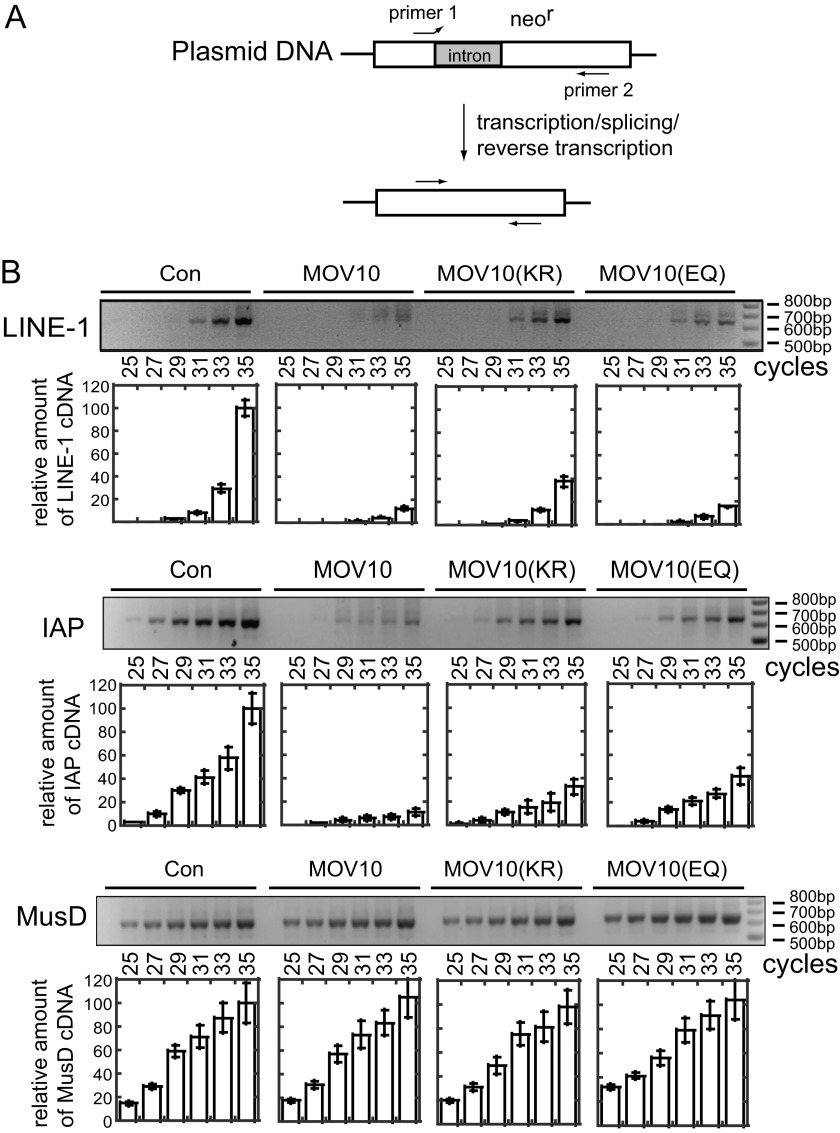
To determine which step of LINE-1 mobilization is affected by MOV10, we first performed PCR to measure the level of LINE-1 cDNA that has been reverse transcribed from LINE-1 RNA, including the newly synthesized and the integrated forms. To eliminate the contamination of PCR by the CMV-L1-neoRT reporter plasmid DNA that was used for transfection in HEK293 cells, we employed a pair of primers that were previously described in Ref. 49. The forward primer was designed to solely bind to the spliced neomycin resistance gene such that only the LINE-1 cDNA that has been reverse transcribed from the spliced RNA, which was produced from either the transfected reporter plasmid or the integrated copy thereof, can be amplified (Fig. 4A). Consistent with the results of the cell colony assays, wild type MOV10 significantly diminished the amounts of cDNA of LINE-1 and IAP by approximately 7–10-fold but did not affect MusD cDNA production (Fig. 4B). The MOV10 (KR) mutant exhibited no discernable effect in this regard, whereas MOV10 (EQ) moderately reduced LINE-1 cDNA level (Fig. 4B). These data further substantiate the deleterious effect of MOV10 on LINE-1 replication.
FIGURE 4.
Overexpression of MOV10 diminishes the production of LINE-1 cDNA. A, location of primers that were used to amplify cDNA of L1-neoRT, IAP-neoTNF, and MusD-neoTNF (49). Primer 1 binds only to cDNA that is generated from spliced neomycin resistance mRNA, not to the plasmid DNA that was used in transfection. B, equal amounts of total cellular DNA normalized by the GAPDH internal control (Con) were used to amplify the target cDNA, Results of PCR to show the levels of cDNA products of LINE-1, IAP, and MusD. The same amounts of total cellular DNA were used as the template in PCR that was run for various cycles to show the linear range of the amplification reactions. The results shown represent those from three independent transfection experiments.
MOV10 Diminishes the Level of LINE-1 RNA
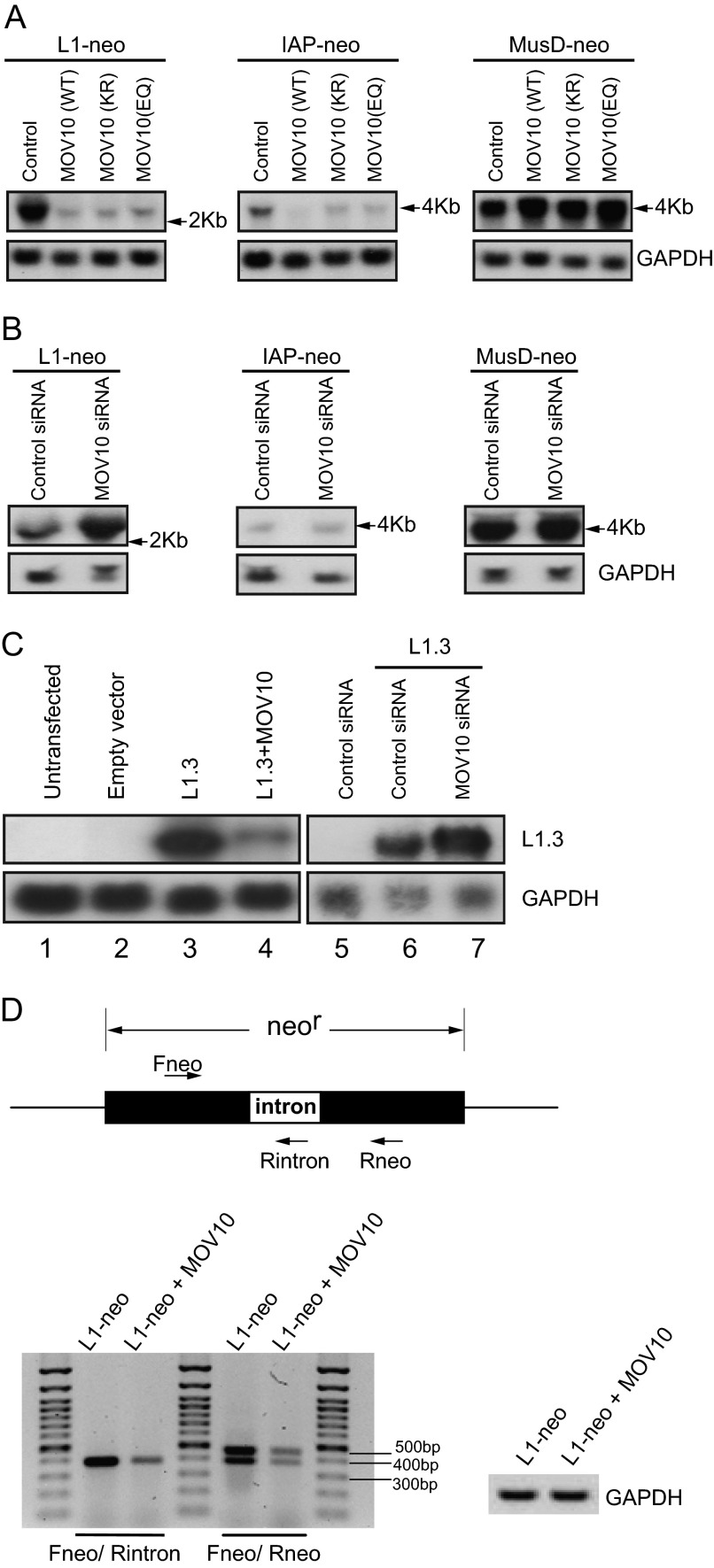
We next measured the effect of MOV10 on LINE-1 RNA production. MOV10 plasmid DNA was co-transfected into HEK293 cells together with CMV-L1-neoRT, IAP-neoTNF, or MusD-neoTNF. The levels of LINE-1, IAP, and MusD RNA were measured by Northern blot using 32P-labeled DNA probes that hybridize to the neomycin resistance gene sequences that are carried by the L1-neo, IAP-neo, and MusD-neo RNA. A drastic reduction in LINE-1 RNA and IAP RNA was seen when MOV10 was overexpressed, as opposed to a modest increase in MusD RNA (Fig. 5A). Depletion of endogenous MOV10 led to a significant increase in LINE-1 RNA (Fig. 5B). In contrast, no evident change was seen in IAP or MusD RNA signals (Fig. 5B). These observations are consistent with the impact of MOV10 on the transposition of LINE-1, IAP, or MusD that was measured in the colony assays either under conditions of MOV10 overexpression or knockdown (Figs. 1–3). We next performed similar experiments using a full-length human L1.3 DNA clone that does not bear any exogenous marker (47). Consistent with the observation that was made with the CMV-L1-neoRT reporter, overexpression of MOV10 profoundly diminished the level of L1.3 RNA, whereas MOV10 knockdown by siRNA oligonucleotides significantly increased L1.3 RNA expression (Fig. 5C). Notably, LINE-1 RNA transcript containing unspliced Neor, which should be generated from the transfected plasmid, was reduced to similar degree as spliced mRNA when MOV10 was expressed ectopically (Fig. 5D). This suggests that the reduction in LINE-1 RNA is most likely not due to inhibiting LINE-1 integration by MOV10, which would cause less copies of LINE-1 DNA integrated and consequently the decrease in LINE-1 RNA product. Therefore, we concluded that MOV10 is capable of diminishing LINE-1 RNA.
FIGURE 5.
MOV10 diminishes LINE-1 RNA level. A, effects of ectopic MOV10 on the RNA levels of LINE-1, IAP, and MusD. HEK293 cells were co-transfected with CMV-L1-neoRT, IAP-neoTNF, or MusD-neoTNF DNA together with wild type or mutant MOV10 DNA. Viral RNA levels were measured by Northern blotting using 32P-labeled DNA probes specific for the neomycin resistance gene. The bands indicated are the major splicing forms of L1-neoRT, IAP-neoTNF, and MusD-neoTNF. The cellular GAPDH mRNA served as internal controls. B, knockdown of MOV10 increases LINE-1 RNA level in HEK293 cells. HEK293 cells were transfected with control siRNA or MOV10-targeting siRNA prior to transfection with CMV-L1-neoRT, IAP-neoTNF, and MusD-neoTNF DNA. Northern blotting was performed to measure the RNA levels of L1-neoRT, IAP-neoTNF, and MusD-neoTNF. A representative result of three independent experiments is shown. C, MOV10 impairs L1.3 RNA expression. JM101/L1.3 DNA was either transfected into HEK293 cells together with wild type MOV10 DNA or transfected into HEK293 cells that had been treated with control or MOV10 siRNA oligonucleotides. Levels of L1.3 RNA were measured by Northern blotting. GAPDH mRNA was detected to serve as an internal control. The overexpression and knockdown of MOV10 in all transfection experiments were confirmed by Western blotting (data not shown). D, HEK293 cells were co-transfected with CMV-L1-neoRT and MOV10 expressing plasmid. Equal amount of total cellular RNA was treated with DNase to remove the potential contamination of plasmid DNA. The cDNA products were normalized by the GAPDH. The unspliced form of LINE-1 RNA was detected by PCR using the primers Fneo and Rintron, whereas the spliced form of LINE-1 RNA was detected by PCR using the primers Fneo and Rneo.
It has been reported that the first 670 nt of the LINE-1 5′-UTR display promoter activity (13), we therefore generated a reporter DNA construct by inserting LINE-1 promoter sequence upstream of the firefly luciferase reporter gene and then transfected this reporter DNA into HEK293 cells together with MOV10 DNA, so as to test whether MOV10 suppresses LINE-1 RNA expression by inhibiting the activity of LINE-1 promoter. Neither the wild type nor the mutant MOV10 proteins markedly affected the luciferase activity that was produced under the control of LINE-1 promoter (Fig. 6, A and B). Given no evidence showing, to our knowledge, that any cis-elements other than the 5′-UTR regulate LINE-1 transcription, our result suggests that MOV10 modulates LINE-1 RNA expression at the post-transcriptional stage such as through accelerating RNA degradation. To test this, we used actinomycin D (5 μg/ml) to block the activity of RNA polymerase II and wished to measure the decay kinetics of LINE-1 RNA with MOV10 overexpression. As shown in Fig. 6C, the amount of LINE-1 RNA in the cells expressing MOV10 was less than 10% of that in the control cells at time 0. This suggests a rather rapid degradation of LINE-1 mediated by MOV10. In the absence of MOV10, LINE-1 RNA was found really stable, i.e., only ∼45% reduction after an 8-h treatment with actinomycin D (Fig. 6C). Although it is still unclear how MOV10 causes this drastic decrease in LINE-1 RNA, this decrease may account for the subsequent reduction in LINE-1 cDNA production and LINE-1 transposition.
FIGURE 6.
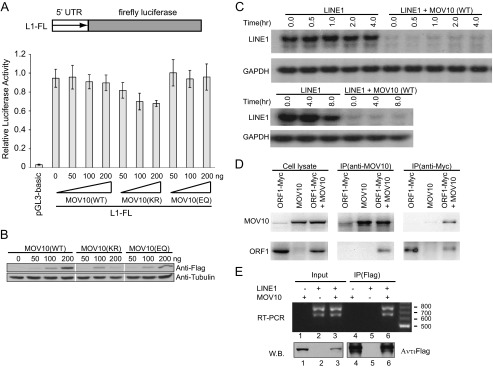
Effects of MOV10 overexpression on LINE-1 5′-UTR promoter activity and RNA stability. A, HEK293 cells were co-transfected with the wild type or mutant MOV10 DNA together with L1-FL. The firefly luciferase activity was measured to represent the activity of LINE-1 5′-UTR promoter. The pGL3-basic contains the firefly luciferase gene that lacks a promoter at its 5′ end. Firefly luciferase activity from this vector was measured to reflect the basal expression of firefly luciferase gene. The TK-RL plasmid DNA was included in each transfection to control the efficiency of different transfections. The results shown are the ratios of firefly luciferase activity versus Renilla luciferase activity. B, Western blots to show the expression of MOV10 and its mutants in the transiently transfected HEK293 cells. C, the same amounts of HEK293 cells were co-transfected with CMV-L1-neoRT and either MOV10 DNA or pcDNA3.1 and then treated with actinomycin D for various time periods as indicated. The RNA level of CMV-L1-neoRT was measured by Northern blotting as described previously. The cellular GAPDH mRNA served as internal controls. D, HEK293 cells were co-transfected with MOV10 DNA and plasmid encoding Myc-tagged ORF1. The cells were collected and lysed 48 h post-transfection. Equal amounts of supernatant were incubated with MOV10-specific antibody or anti-Myc antibody. The immunoprecipitated materials were analyzed by Western blots using anti-Myc or MOV10-specific antibody, respectively. E, HEK293 cells were transfected with FLAG/HA-tagged MOV10 and CMV-L1-neoRT DNA. The presence of LINE-1 RNA in the transfected cells as well as in the immunoprecipitated MOV10 complex was detected by RT-PCR using primers that amplify the neo coding sequence. W.B., Western blot.
The ORF1 and ORF2 proteins as well as LINE-1 RNA have been observed to form specific cytoplasmic foci in cultured cells (8–10). Although the role of these foci in LINE-1 replication is unclear, the viral RNP may protect LINE-1 RNA from the degradation by different kinds of host machinery. Therefore, it is possible that MOV10 targets LINE-1 RNP and then destabilizes LINE-1 RNA. To explore this possibility, we investigated whether MOV10 interacts with ORF1, the structural protein of LINE-1. The Myc-tagged ORF1 and/or MOV10 were expressed in 293 cells, and the interaction between the ORF1 and MOV10 was analyzed with a co-immunoprecipitation assay with either anti-MOV10 or anti-Myc antibody. As shown in Fig. 6D, MOV10 and ORF1 were co-immunoprecipitated, suggesting that MOV10 is able to interact with ORF1 either directly or indirectly. We went further to examine whether MOV10 associates with LINE-1 RNA by immunoprecipitation and RT-PCR. The results of Fig. 6E reveal the presence of LINE-1 RNA in the precipitated MOV10 complex. These data suggest a mechanism by which MOV10 causes degradation of LINE-1 RNA by association with LINE-1 RNA-protein complex.
MOV10 May Inhibit LINE-1 Mobility by an Ago2-independent Mechanism
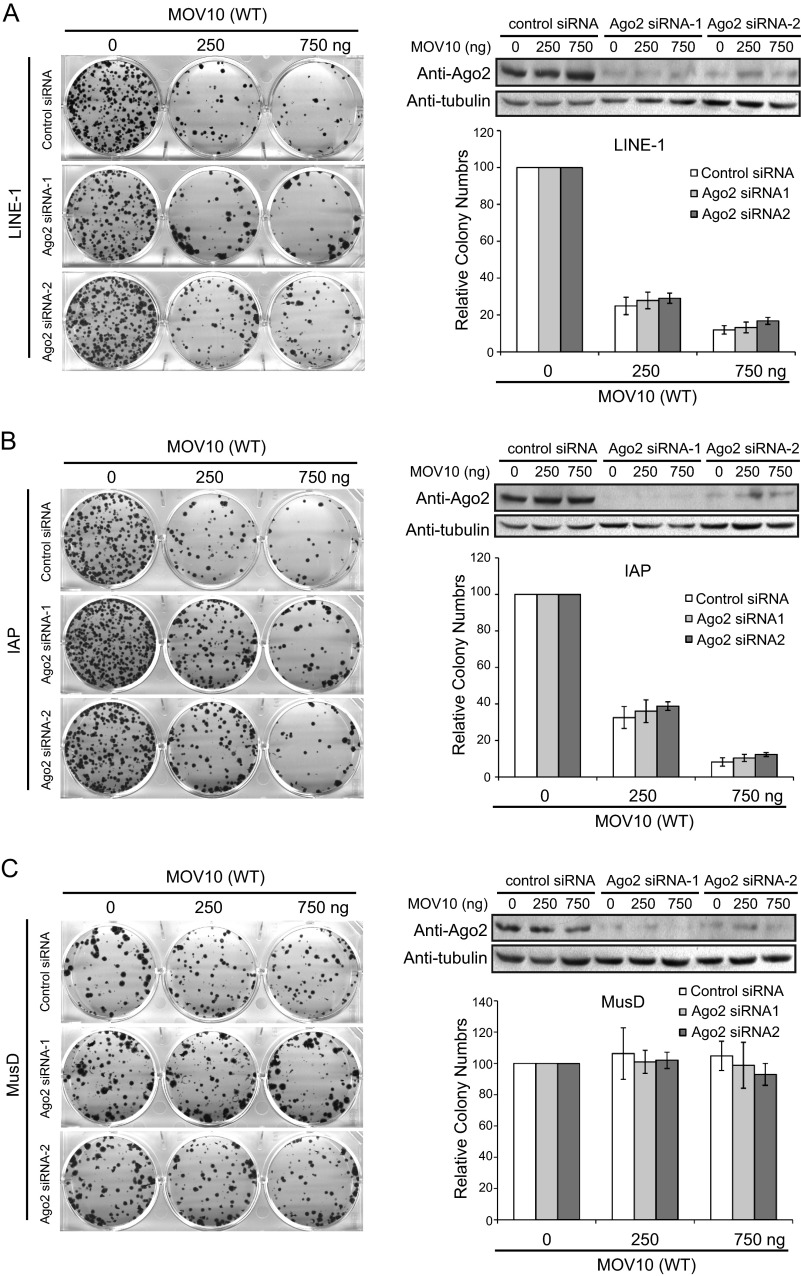
MOV10 has been reported to associate with Ago2 and has a role in microRNA-mediated gene silencing (38). We therefore asked whether MOV10 inhibits LINE-1 retrotransposition via the microRNA pathway. To answer this question, we first knocked down the endogenous Ago2 in HeLa cells using siRNA and then co-transfected MOV10 and CMV-L1-neoRT DNA. The results showed that MOV10 diminished the number of G418-resistant cell colonies to the same degree in cells that were either treated with control siRNA or with siRNA that targeted Ago2 (Fig. 7). Although we cannot rule out the possibility that the residual amounts of Ago2 following siRNA treatment are sufficient to support MOV10 to inhibit LINE-1 activity, one interpretation of these data is that MOV10 may not require Ago2 to restrict LINE-1 retrotransposition.
FIGURE 7.
MOV10 inhibits LINE-1 retrotransposition when Ago2 is knocked down. Western blots were performed to determine the efficiency of Ago2 knockdown in each transfection experiment. Colony assays were conducted to assess the retrotransposition efficiency of LINE-1 (A), IAP (B), or MusD (C) in HeLa cells that had been transfected with either control siRNA or Ago2-targeting siRNA oligonucleotides. The results of three independent transfection experiments are summarized in each bar graph.
DISCUSSION
Our data show that MOV10 restricts LINE-1 replication through suppressing LINE-1 RNA expression, which further supports the results recently reported in three independent studies (55–57). Because the transcription activity of LINE-1 promoter is not evidently affected by MOV10 (Fig. 6), we speculate that MOV10 may diminish LINE-1 RNA level by accelerating its decay. Indeed, the RNA stability analysis suggests a rapid degradation of LINE-1 RNA induced by MOV10. This mechanism of inhibition at the LINE-1 RNA level is also observed by Goodier et al. (55).
The ORF1 and ORF2 proteins, as well as LINE-1 RNA, have been observed to form specific cytoplasmic foci in cultured cells (8–10). These LINE-1 foci are often associated with stress granules that store mRNA molecules that are translation dormant and may also be subject to degradation. Indeed, our data herein showed that MOV10 interacts with ORF1. A most recent work also showed that MOV10 associates with the LINE-1 RNP and co-localize with ORF1 in cytoplasmic granules, accompanied by a decrease in cytoplasmic LINE-1RNPs (55). These observations raise the possibility that MOV10 sequesters LINE-1 RNA in these foci and engages RNA degradation by cellular machinery, thereby controlling the activity of LINE-1. In support of the hypothesis, we found that two mutated MOV10 unable to form cytoplasmic foci exhibited impaired activity against LINE-1.
Although all experimental evidence thus far supports the action of RNA degradation, the mechanism underlying is still unclear. MOV10 has been reported to associate with Ago2 and regulate the activity of holo-RISC (RNA-induced silencing complex) (38), which may have enabled MOV10 to regulate local mRNA translation at neuron synapses (39). It is thus possible that MOV10 diminishes LINE-1 RNA stability through engaging the activity of LINE-1-targeting microRNA molecules that already exist in cells. However, MOV10 effectively inhibited LINE-1 replication when Ago2 was knocked down with siRNA oligonucleotides (Fig. 7), suggesting that a fully functional microRNA pathway is likely unessential for MOV10 to suppress LINE-1, although we cannot rule out that the residual amount of Ago2 after siRNA treatment suffices for this inhibitory function of MOV10. Consistent with our conclusion, a recent work showed that MOV10 is not necessary for microRNA or siRNA-mediated mRNA silencing (57). In a similar vein, overexpression of MOV10, but not Dicer or Ago2, diminishes the production of infectious HIV-1 particles (43). These observations collectively support that MOV10 may also directly associate with LINE-1 RNA and target it for degradation independent of the microRNA or siRNA pathway.
The work herein shows that MOV10 mutants bearing mutated Walker A and B motifs dramatically reduced their ability to restrict LINE-1 transposition (Figs. 1–4) in agreement with the previous report (55). These data suggest that helicase activity might be important to inhibitory activity of LINE-1. However, it is worthy note that the MOV10 (EQ), a mutant form containing a substitution of glutamic to glutamine in Walker B motif, still exhibited considerable restriction activity against LINE-1. Because the glutamic acid in Walker B motif is suggested to act as a catalytic base in ATP hydrolysis and to be essential for the helicase activity (37, 58), this observation raises a possibility that other function of MOV10 may play a role in its inhibitory effect on retrotransposon. Consistent with our finding, Goodier et al. (55) reported that mutated motifs Ia and V, the other two motifs conserved in RNA helicase, had no effect anti-LINE-1 activity of MOV10. Furthermore, MOV10 was shown to inhibit IAP RTP and decrease significantly the products of reverse transcriptase (57). Two other reports also provided evidence of MOV10 mediated inhibition of HIV-1 reverse transcription (43, 44). Although our work suggests that the reduction in LINE-1 RNA is most likely not due to inhibiting LINE-1 integration by MOV10 (Fig. 5E), it is worth exploring the possibility that MOV10 blocks the reverse transcription and/or integration step of LINE-1 replication as an extra mechanism.
LINE-1 activity is controlled by host cells via a number of mechanisms that target distinct stages of LINE-1 replication (12). One layer of control involves methylation of LINE-1 DNA, which silences LINE-1 RNA synthesis. Loss of LINE-1 DNA methylation, as seen in the early stage of mouse embryogenesis and primordial germ cell development, results in active LINE-1 gene expression (22, 23). It is known that mouse primordial germ cells are guarded by piRNAs that control the activated LINE-1 by restoring LINE-1 DNA methylation (25, 26). MOV10L1, the homologue of MOV10, has been shown to associate with Piwi proteins and is required for controlling the expression of retrotransposons in germ cells (59, 60). In analogy to MOV10L1, MOV10 may play a similar role in suppressing retrotransposons in somatic cells. Understanding of the molecular mechanism behind this function of MOV10 will require identifying the associated cellular factors and elucidating the involved RNA decay pathways.
Acknowledgments
We thank Daniel Stetson, Thierry Heidmann, Thomas Tuschl, Rongtuan Lin, Brian Cullen, Jose Luis Garcia-Perez, and John Moran for providing valuable reagents.
This work was supported by Canadian Institutes of Health Research 973 Program Grant 2012CB911102 and National Natural Science Foundation of China Grant 31270210.
- LTR
- long terminal repeat.
REFERENCES
- 1. Brouha B., Schustak J., Badge R. M., Lutz-Prigge S., Farley A. H., Moran J. V., Kazazian H. H., Jr. (2003) Hot L1s account for the bulk of retrotransposition in the human population. Proc. Natl. Acad. Sci. U.S.A. 100, 5280–5285 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Belancio V. P., Hedges D. J., Deininger P. (2008) Mammalian non-LTR retrotransposons. For better or worse, in sickness and in health. Genome Res 18, 343–358 [DOI] [PubMed] [Google Scholar]
- 3. Chen J. M., Stenson P. D., Cooper D. N., Férec C. (2005) A systematic analysis of LINE-1 endonuclease-dependent retrotranspositional events causing human genetic disease. Hum. Genet. 117, 411–427 [DOI] [PubMed] [Google Scholar]
- 4. Babushok D. V., Kazazian H. H., Jr. (2007) Progress in understanding the biology of the human mutagen LINE-1. Hum. Mutat. 28, 527–539 [DOI] [PubMed] [Google Scholar]
- 5. Kulpa D. A., Moran J. V. (2006) Cis-preferential LINE-1 reverse transcriptase activity in ribonucleoprotein particles. Nat. Struct. Mol. Biol. 13, 655–660 [DOI] [PubMed] [Google Scholar]
- 6. Feng Q., Moran J. V., Kazazian H. H., Jr., Boeke J. D. (1996) Human L1 retrotransposon encodes a conserved endonuclease required for retrotransposition. Cell 87, 905–916 [DOI] [PubMed] [Google Scholar]
- 7. Moran J. V., Holmes S. E., Naas T. P., DeBerardinis R. J., Boeke J. D., Kazazian H. H., Jr. (1996) High frequency retrotransposition in cultured mammalian cells. Cell 87, 917–927 [DOI] [PubMed] [Google Scholar]
- 8. Goodier J. L., Zhang L., Vetter M. R., Kazazian H. H., Jr. (2007) LINE-1 ORF1 protein localizes in stress granules with other RNA-binding proteins, including components of RNA interference RNA-induced silencing complex. Mol. Cell. Biol. 27, 6469–6483 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Goodier J. L., Mandal P. K., Zhang L., Kazazian H. H., Jr. (2010) Discrete subcellular partitioning of human retrotransposon RNAs despite a common mechanism of genome insertion. Hum. Mol. Genet. 19, 1712–1725 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Doucet A. J., Hulme A. E., Sahinovic E., Kulpa D. A., Moldovan J. B., Kopera H. C., Athanikar J. N., Hasnaoui M., Bucheton A., Moran J. V., Gilbert N. (2010) Characterization of LINE-1 ribonucleoprotein particles. PLoS Genet. 6, e1001150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Cordaux R., Batzer M. A. (2009) The impact of retrotransposons on human genome evolution. Nat Rev Genet 10, 691–703 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Zamudio N., Bourc'his D. (2010) Transposable elements in the mammalian germline. A comfortable niche or a deadly trap? Heredity 105, 92–104 [DOI] [PubMed] [Google Scholar]
- 13. Swergold G. D. (1990) Identification, characterization, and cell specificity of a human LINE-1 promoter. Mol. Cell. Biol. 10, 6718–6729 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Yoder J. A., Walsh C. P., Bestor T. H. (1997) Cytosine methylation and the ecology of intragenomic parasites. Trends Genet. 13, 335–340 [DOI] [PubMed] [Google Scholar]
- 15. Kato Y., Kaneda M., Hata K., Kumaki K., Hisano M., Kohara Y., Okano M., Li E., Nozaki M., Sasaki H. (2007) Role of the Dnmt3 family in de novo methylation of imprinted and repetitive sequences during male germ cell development in the mouse. Hum. Mol. Genet. 16, 2272–2280 [DOI] [PubMed] [Google Scholar]
- 16. Bourc'his D., Bestor T. H. (2004) Meiotic catastrophe and retrotransposon reactivation in male germ cells lacking Dnmt3L. Nature 431, 96–99 [DOI] [PubMed] [Google Scholar]
- 17. Walsh C. P., Chaillet J. R., Bestor T. H. (1998) Transcription of IAP endogenous retroviruses is constrained by cytosine methylation. Nat. Genet. 20, 116–117 [DOI] [PubMed] [Google Scholar]
- 18. Branciforte D., Martin S. L. (1994) Developmental and cell type specificity of LINE-1 expression in mouse testis. Implications for transposition. Mol. Cell. Biol. 14, 2584–2592 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Kano H., Godoy I., Courtney C., Vetter M. R., Gerton G. L., Ostertag E. M., Kazazian H. H., Jr. (2009) L1 retrotransposition occurs mainly in embryogenesis and creates somatic mosaicism. Genes Dev. 23, 1303–1312 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Garcia-Perez J. L., Marchetto M. C., Muotri A. R., Coufal N. G., Gage F. H., O'Shea K. S., Moran J. V. (2007) LINE-1 retrotransposition in human embryonic stem cells. Hum. Mol. Genet. 16, 1569–1577 [DOI] [PubMed] [Google Scholar]
- 21. van den Hurk J. A., Meij I. C., Seleme M. C., Kano H., Nikopoulos K., Hoefsloot L. H., Sistermans E. A., de Wijs I. J., Mukhopadhyay A., Plomp A. S., de Jong P. T., Kazazian H. H., Cremers F. P. (2007) L1 retrotransposition can occur early in human embryonic development. Hum. Mol. Genet. 16, 1587–1592 [DOI] [PubMed] [Google Scholar]
- 22. Hajkova P., Erhardt S., Lane N., Haaf T., El-Maarri O., Reik W., Walter J., Surani M. A. (2002) Epigenetic reprogramming in mouse primordial germ cells. Mech. Dev. 117, 15–23 [DOI] [PubMed] [Google Scholar]
- 23. Rougier N., Bourc'his D., Gomes D. M., Niveleau A., Plachot M., Pàldi A., Viegas-Péquignot E. (1998) Chromosome methylation patterns during mammalian preimplantation development. Genes Dev. 12, 2108–2113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Smith Z. D., Chan M. M., Mikkelsen T. S., Gu H., Gnirke A., Regev A., Meissner A. (2012) A unique regulatory phase of DNA methylation in the early mammalian embryo. Nature 484, 339–344 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Carmell M. A., Girard A., van de Kant H. J., Bourc'his D., Bestor T. H., de Rooij D. G., Hannon G. J. (2007) MIWI2 is essential for spermatogenesis and repression of transposons in the mouse male germline. Dev. Cell 12, 503–514 [DOI] [PubMed] [Google Scholar]
- 26. Aravin A. A., Hannon G. J., Brennecke J. (2007) The Piwi-piRNA pathway provides an adaptive defense in the transposon arms race. Science 318, 761–764 [DOI] [PubMed] [Google Scholar]
- 27. Suspène R., Aynaud M. M., Koch S., Pasdeloup D., Labetoulle M., Gaertner B., Vartanian J. P., Meyerhans A., Wain-Hobson S. (2011) Genetic editing of herpes simplex virus 1 and Epstein-Barr herpesvirus genomes by human APOBEC3 cytidine deaminases in culture and in vivo. J. Virol. 85, 7594–7602 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Chiu Y. L., Greene W. C. (2008) The APOBEC3 cytidine deaminases. An innate defensive network opposing exogenous retroviruses and endogenous retroelements. Annu Rev Immunol 26, 317–353 [DOI] [PubMed] [Google Scholar]
- 29. Narvaiza I., Linfesty D. C., Greener B. N., Hakata Y., Pintel D. J., Logue E., Landau N. R., Weitzman M. D. (2009) Deaminase-independent inhibition of parvoviruses by the APOBEC3A cytidine deaminase. PLoS Pathog. 5, e1000439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Vartanian J. P., Henry M., Marchio A., Suspène R., Aynaud M. M., Guétard D., Cervantes-Gonzalez M., Battiston C., Mazzaferro V., Pineau P., Dejean A., Wain-Hobson S. (2010) Massive APOBEC3 editing of hepatitis B viral DNA in cirrhosis. PLoS Pathog. 6, e1000928. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Renard M., Henry M., Guétard D., Vartanian J. P., Wain-Hobson S. (2010) APOBEC1 and APOBEC3 cytidine deaminases as restriction factors for hepadnaviral genomes in non-humans in vivo. J. Mol. Biol. 400, 323–334 [DOI] [PubMed] [Google Scholar]
- 32. Esnault C., Priet S., Ribet D., Heidmann O., Heidmann T. (2008) Restriction by APOBEC3 proteins of endogenous retroviruses with an extracellular life cycle. Ex vivo effects and in vivo “traces” on the murine IAPE and human HERV-K elements. Retrovirology 5, 75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Wissing S., Montano M., Garcia-Perez J. L., Moran J. V., Greene W. C. (2011) Endogenous APOBEC3B restricts LINE-1 retrotransposition in transformed cells and human embryonic stem cells. J. Biol. Chem. 286, 36427–36437 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Goila-Gaur R., Strebel K. (2008) HIV-1 Vif, APOBEC, and intrinsic immunity. Retrovirology 5, 51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Mooslehner K., Müller U., Karls U., Hamann L., Harbers K. (1991) Structure and expression of a gene encoding a putative GTP-binding protein identified by provirus integration in a transgenic mouse strain. Mol. Cell. Biol. 11, 886–893 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Fairman-Williams M. E., Guenther U. P., Jankowsky E. (2010) SF1 and SF2 helicases. Family matters. Curr. Opin. Struct. Biol. 20, 313–324 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Walker J. E., Saraste M., Runswick M. J., Gay N. J. (1982) Distantly related sequences in the α- and β-subunits of ATP synthase, myosin, kinases and other ATP-requiring enzymes and a common nucleotide binding fold. EMBO J. 1, 945–951 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Meister G., Landthaler M., Peters L., Chen P. Y., Urlaub H., Lührmann R., Tuschl T. (2005) Identification of novel argonaute-associated proteins. Curr. Biol. 15, 2149–2155 [DOI] [PubMed] [Google Scholar]
- 39. Banerjee S., Neveu P., Kosik K. S. (2009) A coordinated local translational control point at the synapse involving relief from silencing and MOV10 degradation. Neuron 64, 871–884 [DOI] [PubMed] [Google Scholar]
- 40. Haussecker D., Cao D., Huang Y., Parameswaran P., Fire A. Z., Kay M. A. (2008) Capped small RNAs and MOV10 in human hepatitis delta virus replication. Nat. Struct. Mol. Biol. 15, 714–721 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41. Schoggins J. W., Wilson S. J., Panis M., Murphy M. Y., Jones C. T., Bieniasz P., Rice C. M. (2011) A diverse range of gene products are effectors of the type I interferon antiviral response. Nature 472, 481–485 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Furtak V., Mulky A., Rawlings S. A., Kozhaya L., Lee K., Kewalramani V. N., Unutmaz D. (2010) Perturbation of the P-body component Mov10 inhibits HIV-1 infectivity. PLoS One 5, e9081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Wang X., Han Y., Dang Y., Fu W., Zhou T., Ptak R. G., Zheng Y. H. (2010) Moloney leukemia virus 10 (MOV10) protein inhibits retrovirus replication. J. Biol. Chem. 285, 14346–14355 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Burdick R., Smith J. L., Chaipan C., Friew Y., Chen J., Venkatachari N. J., Delviks-Frankenberry K. A., Hu W. S., Pathak V. K. (2010) P body-associated protein Mov10 inhibits HIV-1 replication at multiple stages. J. Virol. 84, 10241–10253 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Esnault C., Maestre J., Heidmann T. (2000) Human LINE retrotransposons generate processed pseudogenes. Nat. Genet. 24, 363–367 [DOI] [PubMed] [Google Scholar]
- 46. Dewannieux M., Dupressoir A., Harper F., Pierron G., Heidmann T. (2004) Identification of autonomous IAP LTR retrotransposons mobile in mammalian cells. Nat. Genet. 36, 534–539 [DOI] [PubMed] [Google Scholar]
- 47. Dombroski B. A., Scott A. F., Kazazian H. H., Jr. (1993) Two additional potential retrotransposons isolated from a human L1 subfamily that contains an active retrotransposable element. Proc. Natl. Acad. Sci. U.S.A. 90, 6513–6517 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Stetson D. B., Ko J. S., Heidmann T., Medzhitov R. (2008) Trex1 prevents cell-intrinsic initiation of autoimmunity. Cell 134, 587–598 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Esnault C., Millet J., Schwartz O., Heidmann T. (2006) Dual inhibitory effects of APOBEC family proteins on retrotransposition of mammalian endogenous retroviruses. Nucleic Acids Res. 34, 1522–1531 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50. Zhou Y., Ma J., Bushan Roy B., Wu J. Y., Pan Q., Rong L., Liang C. (2008) The packaging of human immunodeficiency virus type 1 RNA is restricted by overexpression of an RNA helicase DHX30. Virology 372, 97–106 [DOI] [PubMed] [Google Scholar]
- 51. Belancio V. P., Roy-Engel A. M., Pochampally R. R., Deininger P. (2010) Somatic expression of LINE-1 elements in human tissues. Nucleic Acids Res. 38, 3909–3922 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Belancio V. P., Hedges D. J., Deininger P. (2006) LINE-1 RNA splicing and influences on mammalian gene expression. Nucleic Acids Res. 34, 1512–1521 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Meister G., Landthaler M., Patkaniowska A., Dorsett Y., Teng G., Tuschl T. (2004) Human Argonaute2 mediates RNA cleavage targeted by miRNAs and siRNAs. Mol. Cell 15, 185–197 [DOI] [PubMed] [Google Scholar]
- 54. Singleton M. R., Dillingham M. S., Wigley D. B. (2007) Structure and mechanism of helicases and nucleic acid translocases. Annu. Rev. Biochem. 76, 23–50 [DOI] [PubMed] [Google Scholar]
- 55. Goodier J. L., Cheung L. E., Kazazian H. H., Jr. (2012) MOV10 RNA helicase is a potent inhibitor of retrotransposition in cells. PLoS Genet. 8, e1002941. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56. Arjan-Odedra S., Swanson C. M., Sherer N. M., Wolinsky S. M., Malim M. H. (2012) Endogenous MOV10 inhibits the retrotransposition of endogenous retroelements but not the replication of exogenous retroviruses. Retrovirology 9, 53. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Lu C., Luo Z., Jäger S., Krogan N. J., Peterlin B. M. (2012) Moloney leukemia virus type 10 inhibits reverse transcription and retrotransposition of intracisternal a particles. J. Virol. 86, 10517–10523 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58. Subramanya H. S., Bird L. E., Brannigan J. A., Wigley D. B. (1996) Crystal structure of a DExx box DNA helicase. Nature 384, 379–383 [DOI] [PubMed] [Google Scholar]
- 59. Frost R. J., Hamra F. K., Richardson J. A., Qi X., Bassel-Duby R., Olson E. N. (2010) MOV10L1 is necessary for protection of spermatocytes against retrotransposons by Piwi-interacting RNAs. Proc. Natl. Acad. Sci. U.S.A. 107, 11847–11852 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Zheng K., Xiol J., Reuter M., Eckardt S., Leu N. A., McLaughlin K. J., Stark A., Sachidanandam R., Pillai R. S., Wang P. J. (2010) Mouse MOV10L1 associates with Piwi proteins and is an essential component of the Piwi-interacting RNA (piRNA) pathway. Proc. Natl. Acad. Sci. U.S.A. 107, 11841–11846 [DOI] [PMC free article] [PubMed] [Google Scholar]