Abstract
In the last decade, RNA interference (RNAi) advanced to one of the most widely applied techniques in the biomedical research field and several RNAi therapeutic clinical trials have been launched. We focus on RNAi-based inhibitors against the chronic infection with human immunodeficiency virus type 1 (HIV-1). A lentiviral gene therapy is proposed for HIV-infected patients that will protect and reconstitute the vital immune cell pool. The RNAi-based inhibitors that have been developed are short hairpin RNA molecules (shRNAs), of which multiple are needed to prevent viral escape. In ten distinct steps, we describe the selection process that started with 135 shRNA candidates, from the initial design criteria, via testing of the in vitro and in vivo antiviral activity and cytotoxicity to the final design of a combinatorial therapy with three shRNAs. These shRNAs satisfied all 10 selection criteria such as targeting conserved regions of the HIV-1 RNA genome, exhibiting robust inhibition of HIV-1 replication and having no impact on cell physiology. This combinatorial shRNA vector will soon move forward to the first clinical studies.
Keywords: Human immunodeficiency virus type 1, RNA interference, Gene therapy, “Human Immune System” mouse, Lentivirus
INTRODUCTION
After discovery of the mechanism of RNA interference (RNAi) in C. elegans in 1998[1], several RNAi approaches have been developed for use in therapeutic strategies, e.g., against inherited diseases or infectious pathogens[2]. The cellular RNAi pathway leads to the processing of small noncoding microRNAs (miRNAs) that regulate cellular gene expression at the post-transcriptional level to control cell differentiation and development[3]. This pathway may be primed by artificial short hairpin RNAs (shRNAs) that are produced in the cell from a transgene and processed into small interfering RNAs (siRNAs)[4]. Ready-to-use siRNAs can also be synthesized chemically and transfected into cells. Perfect base-pairing of the designed siRNA with the specific mRNA target results in cleavage of the latter by the RNA-induced silencing complex (RISC)[5]. Topical delivery of siRNAs in the lungs might be feasible for the treatment of acute infections with e.g., influenza A virus or the respiratory syncytial virus. However, chronic infections caused by pathogens such as the human immunodeficiency virus type 1 (HIV-1), hepatitis B virus and hepatitis C virus will require the continuous expression of RNAi inhibitors from a therapeutic transgene.
HIV-1 is replicating in cells of the human immune system, resulting in a constant depletion of the CD4+ T cells that contributes to the eventual progression to AIDS. Anti-HIV gene therapy aims to protect this indispensable cell pool from virus infection and destruction, which should lead to a (partial) reconstitution of the immune system. Due to the chronic nature of HIV-1 infection, cells must be protected life-long against HIV-1, which can be achieved by a stable RNAi gene therapy against the HIV-1 RNA genome. Apart from RNAi approaches, other antiviral strategies can be utilized such as ribozymes, antisense RNAs, dominant negative protein variants, decoy RNAs, or combinations of RNAi, ribozymes and RNA decoys[6]. However, the simultaneous use of multiple RNAi inhibitors seems one of the most promising approaches for a potent and durable therapy[7]. The therapeutic protocol that we have in mind starts with the isolation of blood mobilized CD34+ hematopoietic progenitor cells from HIV-infected patients, followed by ex vivo transduction with an shRNA-expressing lentiviral vector that stably integrates in the host cell DNA, and re-injection of the modified cells into the patients. Target cells for HIV-1 infection (CD4+ T cells, monocytes/macrophages and dendritic cells) that originate from these transduced pluripotent progenitor cells will express the antiviral RNAi constructs and thus prevent HIV-1 gene expression and virus replication. By that, HIV-1 infected CD4+ T cells evade also the destruction by CD8+ cytotoxic T cells as HIV-1 protein production can trigger viral peptide presentation via the MHC class I molecules to cytotoxic T cells.
We and others have previously demonstrated remarkably potent virus inhibition even with a single shRNA, but also observed that HIV-1 quickly escapes from RNAi pressure via the selection of mutations in the targeted sequence[8-13]. However, a combination of multiple potent shRNAs provided long-term suppression of HIV-1 replication[14-16]. For several years, we have designed and tested various alternative RNAi strategies against HIV-1. Extended shRNA designs and miRNA-like polycistron transcripts were optimized for the expression of multiple inhibitors, but the use of independent shRNA cassettes turned out to be most efficient[9,14,17-21]. Thus, the goal is to use a lentiviral vector with multiple shRNA cassettes that becomes stably incorporated in the human genome. We therefore designed a battery of shRNA inhibitors and tested these in a variety of in vitro and in vivo experimental settings to allow the selection of the most potent and safe RNAi antivirals. The top candidates were subsequently chosen for the development of a combinatorial RNAi gene therapy against HIV-1 that will be translated into a clinical trial[16]. Primary safety and efficacy studies were performed in the “Human Immune System” (HIS) mouse model[22,23]. Human CD34+ hematopoietic progenitor cells (hHPC) were transduced ex vivo with the lentiviral RNAi expression constructs and injected into immunocompromised newborn mice to monitor cell development and differentiation, shRNA expression, cytotoxicity and efficacy of the therapeutic regimen upon HIV-1 infection[24]. This pre-clinical animal model does closely mimic the anti-HIV gene therapy approach proposed for HIV-infected patients.
Here, we will discuss the numerous criteria and corresponding experimental tests that were used in selecting the optimal shRNA reagents for a combinatorial attack on the HIV-1 RNA genome. Ten distinct selection steps can be envisaged (Figure 1): (1) the basic design of the shRNA gene cassettes; (2 + 3) measurement of the antiviral activity in transiently transfected cells and stably transduced cells; (4) selection of the most conserved HIV-1 target sequences to maximize the number of sensitive viral isolates; (5) testing the viral escape possibilities as a measure of the durability of the therapeutic attack; (6) criteria imposed by the use of a lentiviral vector for delivery of the antiviral shRNA cassettes; (7 + 8) screens for possible adverse effects on cell physiology, both in vitro and in vivo; (9) target site alterations due to resistance mutations for clinically approved antiretroviral drugs; and (10) the assembly of multiple shRNAs to establish a combinatorial RNAi therapy. Along this selection pathway, which took over 7 years, we tested more than 135 shRNA candidates to end up with three potent and safe shRNAs that will be employed in a gene therapy trial (Table 1).
Figure 1.
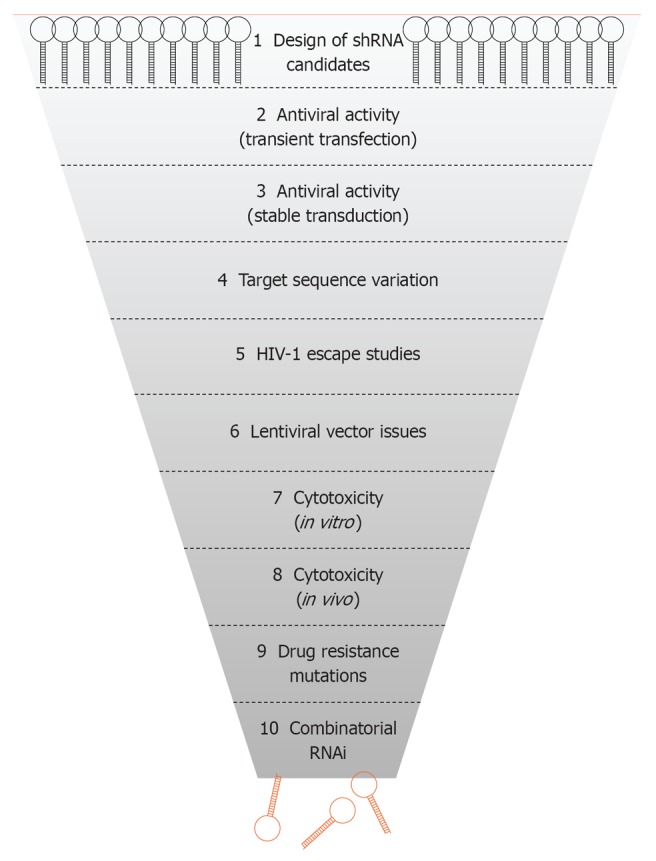
Selection of short hairpin RNAs against human immunodeficiency virus type 1. Scheme of the 10 steps used for the selection of the best shRNA inhibitors for development of an antiviral RNA interference (RNAi) gene therapy. HIV-1: Human immunodeficiency virus type 1.
Table 1.
Selection of short hairpin RNAs against human immunodeficiency virus type 1
| shRNA name |
HIV-1 target |
HIV-1 inhibition |
Target conservation (%)5 |
Viral escape6 |
Lentiviral vector |
Cell toxicity9 |
Drug resistance mutation10 | |||||
| Position1 | Gene | Transient3 | Stable4 | Subtype B | All subtypes | Vector targeting7 | Titer reduction8 | In vitro | In vivo | |||
| LDR2 | 327 | Leader | 95 | ++ | 66 | 61 | ND | + | ND | ND | ND | - |
| LDR3 | 3282 | Leader | 94 | ++ | 68 | 61 | ND | + | ND | ND | ND | - |
| LDR4 | 3292 | Leader | 99 | +++ | 68 | 61 | ND | + | ND | ND | ND | - |
| LDR5 | 3302 | Leader | 94 | ++ | 68 | 61 | ND | + | ND | ND | ND | - |
| LDR7 | 3322 | Leader | 84 | - | 69 | 61 | ND | + | ND | ND | ND | - |
| LDR8 | 3332 | Leader | 79 | ++ | 69 | 61 | ND | + | ND | ND | ND | - |
| LDR9 | 3342 | Leader | 91 | +++ | 70 | 63 | ND | + | ND | ND | ND | - |
| CA | 1032 | CA-p24 | 97 | - | 87 | 67 | ND | - | - | - | ND | - |
| Gag5 | 1365 | CA-p24 | 86 | ++ | 81 | 80 | + | - | + | + | + | - |
| Pol111 | 19102 | Prot11 | 9711 | +++11 | 8911 | 8511 | +11 | -11 | -11 | -11 | -11 | D30N, V32I11 |
| Pol2 | 19112 | Prot | 86 | - | 89 | 85 | ND | - | ND | ND | ND | D30N, V32I |
| Pro 1 | 19122 | Prot | 99 | +++ | 85 | 81 | ND | - | - | - | ND | D30N, V32I |
| Pro 2 | 19132 | Prot | 99 | ND | 80 | 79 | ND | - | ND | ND | ND | D30N, V32I |
| Pro 3 | 19142 | Prot | 99 | ND | 80 | 79 | ND | - | ND | ND | ND | D30N, V32I |
| Pro 4 | 19152 | Prot | 98 | ND | 77 | 76 | ND | - | ND | ND | ND | D30N, V32I, L33F |
| Pro 5 | 19162 | Prot | 97 | ND | 79 | 78 | ND | - | ND | ND | ND | D30N, V32I, L33F |
| Pro 6 | 19182 | Prot | 98 | ND | 78 | 77 | ND | - | ND | ND | ND | D30N, V32I, L33F |
| Pro 7 | 19192 | Prot | 98 | ND | 77 | 76 | ND | - | ND | ND | ND | D30N, V32I, L33F |
| Pro 8 | 2026 | RT | 87 | ND | 58 | 17 | ND | - | ND | ND | ND | D30N, V32I, L33F |
| Pol6 | 37552 | RT | 97 | - | 72 | 75 | ND | - | ND | ND | ND | - |
| RT 1 (A) | 37572 | RT | 95 | ++ | 71 | 74 | ND | - | - | - | ND | - |
| RT 2 (B) | 37582 | RT | 98 | ++ | 73 | 75 | ND | - | - | + | ND | - |
| RT 3 (C) | 37592 | RT | 95 | ++ | 73 | 75 | ND | - | - | - | ND | - |
| RT 4 (F) | 37602 | RT | 85 | ++ | 69 | 69 | ND | - | ND | ND | ND | - |
| RT 5 (G) | 37622 | RT | 94 | - | 72 | 72 | ND | - | - | - | ND | - |
| Int 1 | 4310 | Int | 91 | ND | 71 | 19 | ND | - | ND | ND | ND | - |
| Int 2 | 4344 | Int | 96 | +++ | 67 | 25 | ND | + | + | - | ND | - |
| Pol29 | 4393 | Int | 82 | - | 80 | 80 | ND | + | ND | ND | ND | - |
| Int 3 | 44202 | Int | 95 | +++ | 68 | 61 | ND | + | + | + | ND | - |
| Int 4 | 44222 | Int | 99 | ND | 71 | 64 | ND | + | ND | ND | ND | - |
| Int 5 | 4491 | Int | 81 | ND | 70 | 33 | ND | - | ND | ND | ND | - |
| Pol45 | 45432 | Int | 92 | ++ | 91 | 75 | ND | - | ND | ND | ND | - |
| Pol4711 | 45452 | Int11 | 9211 | +++11 | 9111 | 7311 | +11 | -11 | -11 | -11 | -11 | -11 |
| Vif | 4646 | Int/Vif | 96 | - | 82 | 31 | ND | - | - | - | ND | - |
| R/T511 | 5551 | Rev/Tat11 | 9311 | +++11 | 8711 | 7311 | +11 | -11 | -11 | -11 | -11 | -11 |
| Env 1 | 7250 | gp120 | 81 | ND | 83 | 76 | ND | + | ND | ND | ND | - |
| Env 2 | 78752 | gp41 | 83 | ND | 49 | 33 | ND | - | ND | ND | ND | - |
| Env 3 | 78812 | gp41 | 89 | ND | 19 | 6 | ND | + | ND | ND | ND | - |
| Env 4 | 78842 | gp41 | 85 | ND | 21 | 6 | ND | + | ND | ND | ND | - |
| Env 5 | 8026 | gp41 | 96 | +++ | 53 | 14 | ND | + | + | + | ND | - |
| Env 6 | 82772 | gp41 | 98 | ++ | 53 | 18 | ND | - | - | + | ND | - |
| Env 7 | 82782 | gp41 | 98 | ++ | 56 | 18 | ND | - | - | - | ND | - |
| Env 8 | 8359 | gp41 | 97 | +++ | 3 | 1 | ND | - | - | - | ND | - |
| LTR | 9072 | 3’LTR | 95 | ++ | 53 | 1 | ND | - | - | - | ND | - |
Position in human immunodeficiency virus type 1 (HIV-1) LAI mRNA;
Overlapping short hairpin RNA (shRNA) clusters;
Percentage of inhibition of HIV-1 production in co-transfected cells;
Inhibition of HIV-1 replication in stably transduced cells: +++ = strong, ++ = medium, + = low, - = no inhibition;
Percentage of sequences in Los Alamos database identical to shRNA target sequence;
Detection of escape mutations after prolonged culturing;
100% complementarity of the shRNA to JS1 lentiviral vector;
Titers compared to JS1 lentiviral vector; + = reduction > 1 log, - = reduction < 1 log;
Effects on cell growth; + = negative effect, - = no effect;
Drug resistance mutations in the shRNA target region;
Selected for the combinatorial gene therapy. ND: Not determined.
DESIGN OF shRNA MOLECULES
To identify new and potent shRNAs against HIV-1, different design criteria were applied. In general, the shRNA design was based on the prototype shRNA hairpin transcript published by Brummelkamp in 2002: complementary 19-nucleotide sense and antisense strands, a 9-nucleotide hairpin loop and 3’-UU overhang[4]. The antisense strand of this shRNA design will, upon Dicer processing, form the guide strand that instructs RISC for antiviral attack. The complete shRNA cassette consists of the RNA polymerase III H1 promoter, the shRNA sequence followed by the TTTTT termination signal. The H1 promoter, shRNA and termination signal were designed as synthetic DNA or as restriction fragments and cloned into the pSUPER vector (Figure 2A). This cassette can easily be transferred into the lentiviral vector JS1 (Figure 2B) for generation of stably transduced cells[25]. All shRNAs were checked in silico to avoid significant complementarity against cellular mRNAs to prevent putative off-target effects.
Figure 2.
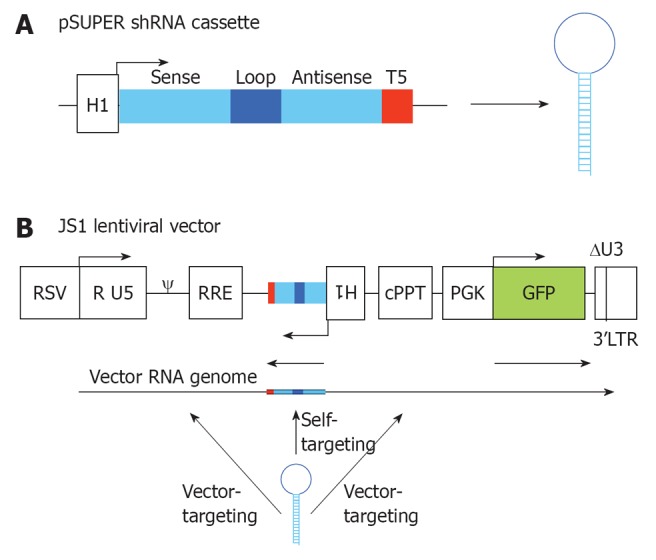
RNA interference vector constructs. A: Two complementary DNA oligonucleotides are annealed and cloned into pSUPER downstream of the H1 promoter that triggers short hairpin RNA (shRNA) expression. The shRNA cassette encodes a 19 nt sense strand, 9 nt loop, 19 nt antisense strand and a stretch of 5 T’s (T5), which is the termination signal; B: The shRNA cassette was cloned in the lentiviral vector JS1 for stable transduction of human T cells. The shRNA cassette is cloned in antisense direction to avoid promoter interference. During vector production three transcripts are produced from the lentiviral vector: the shRNA, the vector RNA genome and the GFP transcript. The shRNA will have a 100% target match with the shRNA-encoding sequence in the vector RNA genome (self-targeting), and potential targets in the human immunodeficiency virus type 1 (HIV-1) derived sequences of the lentiviral vector (vector-targeting). RSV: Respiratorial syncytial virus promoter; R U5: R and U5 element of HIV-1 promoter; ψ: Packaging signal; RRE: Rev responsive element; cPPT: Central polypurine tract; PGK GFP: Green fluorescent protein driven from a PGK promoter; 3’LTR: 3’ long terminal repeat of HIV-1, with deletion in the U3 region.
Over the years, several sets of shRNA inhibitors were tested in our laboratory. We initially described potent suppression of HIV-1 replication with an shRNA that targets nef gene sequences, but viral escape was apparent in prolonged cultures[9,13]. We next tested a first set of 86 antiviral shRNAs that were selected based solely on the conservedness of the target sequence among HIV-1 isolates[14]. Due to the high variability of HIV-1, this selection criterion has become very important for the development of a gene therapy that applies to a broad range of isolates. Our initial studies also revealed the importance of taking the target RNA structure into account for shRNA design as occluded targets are poorly recognized by the RNAi machinery[13,26]. Therefore, we generated a second set of shRNAs that targets particularly accessible regions of the HIV-1 RNA genome based on the SHAPE determined RNA structure model[27,28].
ANTIVIRAL ACTIVITY IN TRANSIENTLY TRANSFECTED CELLS
To evaluate the potency of the shRNAs in terms of anti-HIV activity, we developed a test to measure the inhibition of HIV-1 protein production[14]. For that reason, 293T cells were co-transfected with the HIV-1 molecular clone pLAI, the pSUPER-shRNA vector and the pRL Renilla vector to control for the transfection efficiency. These transfected 293T cells produce infectious virus but do not allow new rounds of infection due to the absence of relevant receptors for HIV-1 attachment and entry. At 48 h post-transfection, the HIV-1 capsid protein (CA-p24) production and Renilla production were quantified. CA-p24 can easily be measured via CA-p24 ELISA in the culture supernatant. Then, CA-p24 levels normalized for Renilla expression were compared to virus production with the empty pSUPER control plasmid obtained in co-transfections[14]. Figure 3 indicates the target sites for the most potent RNAi inhibitors plotted onto the HIV-1 genome. Of the 135 shRNA candidates, 44 exhibited at least 80% suppression of HIV-1 production. The upper panel depicts the first set, the lower panel marks the target sites for the second shRNA set[14,27]. Table 1 summarizes the characteristics of the 44 shRNAs that exhibit robust inhibition.
Figure 3.
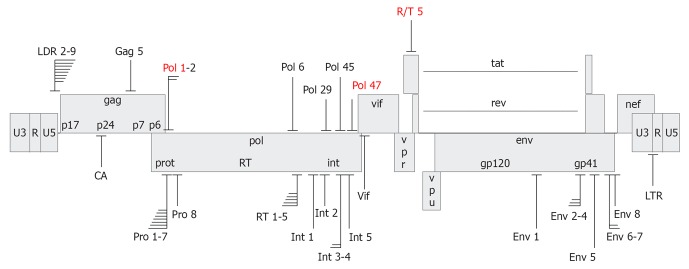
Target sites of anti-human immunodeficiency virus short hairpin RNAs. Depicted is the human immunodeficiency virus type 1 (HIV-1) LAI proviral genome with short hairpin RNA (shRNA) target sites that yielded > 80% suppression of HIV-1 production. Several clusters of overlapping shRNA targets are indicated. The shRNAs have been designed based on conservedness of the target sequence (upper panel) or accessibility in the structured HIV-1 RNA genome (lower panel). shRNAs selected for the R3 combinatorial RNA interference vector are marked in red.
ANTIVIRAL ACTIVITY IN STABLY TRANSDUCED T CELLS
Several of the shRNAs that exhibited significant antiviral activity in the transient transfection assay were subsequently tested in stably transduced CD4+ T cells. To do so, the shRNA expression cassettes were cloned into the lentiviral vector JS1 to allow stable transduction of SupT1 T cells (Figure 2B)[11,14,25]. SupT1 is a commonly used CD4+ T cell line that is permissive for HIV-1 infection and that shows clear cytopathic effects (syncytia) upon virus replication. The cells were transduced at a low multiplicity of infection of 0.15 to assure that maximally a single copy of the lentiviral vector integrates per cell. The expression of a GFP reporter gene by the JS1 lentiviral vector allows the easy separation of transduced from non-transduced cells by fluorescence activated cell sorting. The cells are usually sorted 2 d after transduction to obtain a pure GFP-positive population. The transduced cells can subsequently be challenged with HIV-1 and virus replication can be monitored. Infected cultures were inspected on a daily basis under the microscope to monitor cytopathic effects and supernatants were collected to measure CA-p24 production (Figure 4A). SupT1 cells transduced with the empty JS1 vector served as control cells to measure uninhibited viral spread. For future gene therapy applications, shRNAs were only considered if they conferred strong HIV-1 inhibition in transient co-transfections and stably transduced T cells. In Table 1, this is indicated as “+++” in the respective column.
Figure 4.
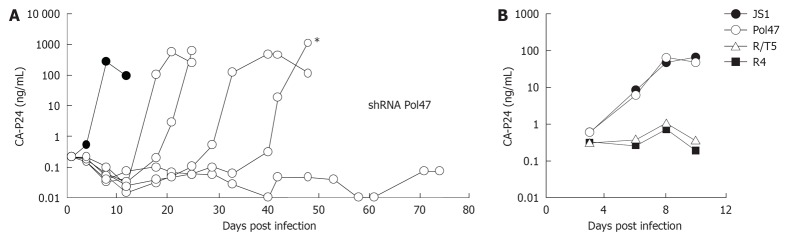
Long-term short hairpin RNA activity in human immunodeficiency virus type 1 replication studies. A: SupT1 T cells expressing short hairpin RNA (shRNA) Pol47 were infected with HIV-1 and virus replication was monitored in five cultures by measuring CA-p24 production for up to 75 d. Cells transduced with the empty lentiviral vector JS1 served as control (dark circles); B: Supernatant from the indicated culture (asterisk in panel A) was passaged on new cells to test the escape phenotype. The virus replicated on control and shRNA Pol47 cells, but not on cells that express another antiviral shRNA (R/T5) or the R4 construct. Adapted from[16].
HIV-1 TARGET SEQUENCE VARIABILITY
Due to the high variability of HIV-1, it is especially important for an anti-HIV gene therapy to target sequences that are relatively conserved among different virus isolates and several HIV-1 subtypes. For most shRNAs, this was an important selection criterion and Table 1 provides an overview of the conservedness of the shRNA targets, both for subtype B that is most prevalent in the Western world and the other subtypes that belong to the HIV-1 group M. To determine the degree of conservedness of the siRNA target sequences, all HIV-1 genome sequences present in the Los Alamos National Laboratory database (http://www.hiv.lanl.gov/) were aligned. The alignment provides the percentage of sequences that are fully complementary to the siRNA for the entire group M and also per subtype. To ensure high sequence conservation, minimally 70% of the viral sequences of each target region have to form a perfect match with the siRNA[14]. This was important for the group M sequences that comprise all subtypes, and specifically the subtype B sequences that are most prevalent in the Western world (Table 1). This standard ensures targeting of a broad spectrum of viral isolates and also decreases the risk of rapid viral escape because mutations of well-conserved HIV-1 sequences are more likely to cause a loss of viral replication efficiency[9-11,13,29,30]. As a current standard diagnostic procedure, the patient-derived HIV-1 sequences of the pol gene are genotyped, including the target sequences for the shRNA inhibitors Pol1 and Pol47. Thus, one will be able to confirm the conservation, such that a full match with the shRNA is guaranteed. Genotyping will also reveal the presence of non-B subtypes, exotic HIV-1 strains and even super-infections that may complicate the RNAi gene therapy[31].
HIV-1 ESCAPE STUDIES
Viral escape from the shRNA pressure can occur similar to what is observed under antiviral therapy with antiviral drugs when the HIV-1 target sequence accumulates one or multiple mutations. Extensive viral escape studies have been performed for some shRNAs[8-11,13,32-34]. Transduced and GFP-sorted SupT1 cells were challenged with a high amount of virus and passaged over time. When viral outgrowth was observed, the cell-free supernatant was transferred to a new culture of shRNA-expressing cells to confirm the resistance phenotype (Figure 4A and B). Cellular DNA with the integrated proviral genome can subsequently be isolated, the siRNA target region can be PCR-amplified and cloned into a plasmid for sequence analysis[11,13,16,35]. At this point, it is important to filter out ‘pseudo-escape’ events that are due to breakthrough virus replication when a high virus input is tested, often in combination with a sub-optimal inhibitory shRNA regimen. In this scenario, the pseudo-escape virus can be recognized because it will not carry any resistance mutation and will obviously lack the resistance phenotype[9,11,16].
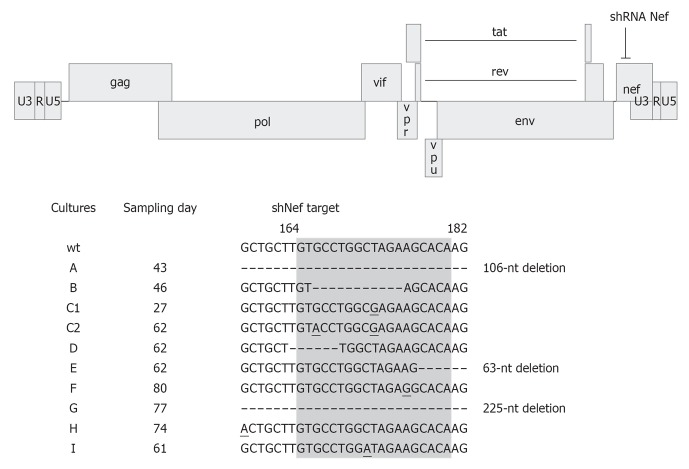
We routinely test multiple HIV-1 evolution cultures in parallel because viral evolution is a chance process driven by randomly occurring mutations, some of which are beneficial and thus subsequently selected under RNAi pressure. For example, the diverse viral escape routes observed in independent cultures of shRNA Nef expressing cells are depicted in Figure 5[13]. We reported three types of HIV-1 escape: (1) Mutation(s) in the siRNA target sequence; (2) a mutation in the flanking region that influences the local RNA structure; and (3) partial or even complete deletion of the target sequence. The latter escape route seems possible only in case non-essential viral sequences are targeted. Indeed, deletion-based viral escape was never witnessed when essential HIV-1 sequences encoding the Protease and Integrase enzymes were targeted[10,32]. This observation supports the notion to target well-conserved viral sequences that usually encode the more essential viral functions. It must be pointed out that viral escape studies are extremely time and labor-intensive. Therefore, such investigations should only be conducted for candidate shRNAs that fulfill multiple criteria, e.g., potent inhibition in transient transfections and stably transduced T cell infections.
Figure 5.
Human immunodeficiency virus type 1 escapes from short hairpin RNA Nef. The human immunodeficiency virus type 1 (HIV-1) LAI proviral genome and the short hairpin RNA (shRNA) Nef target site are indicated. SupT1 cells expressing shRNA Nef were infected with HIV-1 and passaged twice a week until viral escape occurred. Nine different cultures were examined in parallel and the day of sampling is indicated. Part of the nef gene is shown with the shRNA Nef target site highlighted in gray. Numbers refer to the nucleotide position in the nef gene. Escape was apparent by (1) one or more escape mutations in the target sequence; (2) mutations outside the target region; and (3) complete or partial deletions of the target region. Mutations are underlined. Adapted from[13].
LENTIVIRAL VECTOR CONSIDERATIONS
HIV-1 causes a persistent infection in humans, which requires durable expression of the inhibitory shRNAs. Therefore, the use of a lentiviral vector seems ideal because of its property to stably integrate into the host cell genome, which allows a constant supply of antiviral shRNAs. The third generation lentiviral vectors have proven to be safe for use in humans and no insertional oncogenesis has been reported thus far[36-38]. These vectors transduce dividing and non-dividing cells and can thus be applied, e.g., in hematopoietic progenitor cells[23,39-43]. For clinical application, it is important that the vector can be produced to high titers. We and others previously reported titer problems with lentiviral vectors encoding antiviral shRNAs that may obstruct clinical application[44-48]. Lentiviral vectors are produced by co-transfection of the lentiviral vector construct JS1 (Figure 2B), Gag-Pol and Rev-expression plasmids, and a VSV-G envelope construct. During lentiviral vector production, all the different mRNAs are expressed in the producer cell together with the shRNA transcript. The vector transcript does in fact include the shRNA sequence and will thus have a perfect target for siRNA-mediated degradation. However, such self-targeting does not easily occur because the target is occluded in a stable shRNA hairpin structure and therefore protected from RNAi attack. Further complications arise when the shRNA targets HIV-1 derived sequences in one of the lentiviral vector constructs. This is referred to as vector targeting. We previously discussed in detail all possible routes by which shRNAs could impede lentiviral vector production and how to prevent or overcome these specific problems[45,48]. Of course, acute cytotoxicity of the expressed shRNA can also cause a serious titer reduction due to effects on the producer cell viability and this may eventually also affect the viability of transduced cells, i.e., the gene therapy target cells.
CYTOTOXICITY IN IN VITRO CELL CULTURE
The antiviral shRNA may exhibit adverse effects on cell growth through silencing of cellular mRNAs (off-target effects) or saturation of the RNAi machinery, in particular when the shRNA is overexpressed[49]. These effects cannot easily be predicted and should thus be tested experimentally. There are several ways to score the impact of shRNA expression on cell viability and physiology. One could for instance determine the cellular doubling time by frequent cell counting. We recently developed a very user-friendly and ultra-sensitive assay that follows over time the ratio of shRNA-expressing GFP-positive cells vs untransduced GFP-negative cells in a co-culture assay[50]. This competitive cell growth or CCG assay has some clear advantages over other well-established cell proliferation assays: (1) After cell transduction, only a small aliquot of the culture is needed to launch the CCG assay, without any extra steps; (2) The CCG assay is internally controlled as it starts with a mixture of transduced and untransduced cells; and (3) Even minor effects on the cellular proliferation rate caused by shRNA expression can be detected. We screened all promising shRNAs in this assay (Figure 6). Besides single shRNA-expressing vectors, we also investigated combinatorial vectors such as R4 (Gag5, Pol1, Pol47, and R/T5) and R3 (Pol1, Pol47, and R/T5). shRNAs that exhibit negative effects on cell growth such as Gag5 should be excluded from combinatorial RNAi vectors (Figure 6, Table 1). Removal of Gag5 from the R4 vector that exhibited impaired cell proliferation led to the design of the R3 lentiviral vector that scored no negative cell growth effects. Cytotoxicity by saturation of the cellular RNAi pathway is especially critical for the combinatorial shRNA vectors and might have contributed to the adverse R4 effects.
Figure 6.
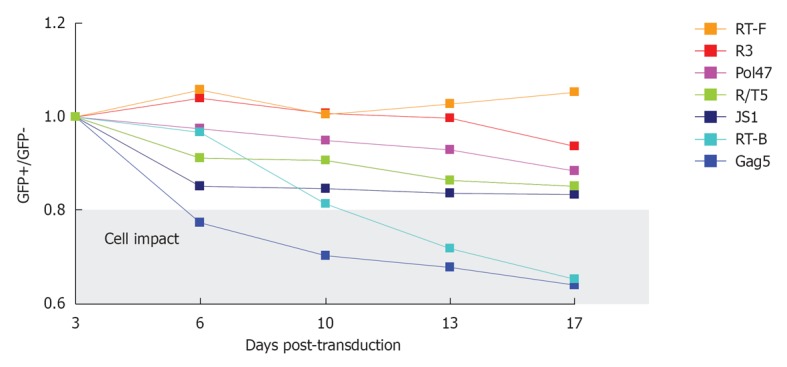
Competitive cell growth assay. SupT1 T cells were transduced with a short hairpin RNA (shRNA)-expressing lentiviral vector, yielding a cell population with approximately 40% GFP-positive cells. The ratio of GFP-positive cells at day 3 after transduction was set at 1 and measured longitudinally. The cells were passaged and analyzed via FACS measurement twice a week. JS1 represents the empty lentiviral vector without shRNA expression. The gray window highlights shRNAs that have a significant adverse effect on cell growth.
CYTOTOXICITY IN VIVO
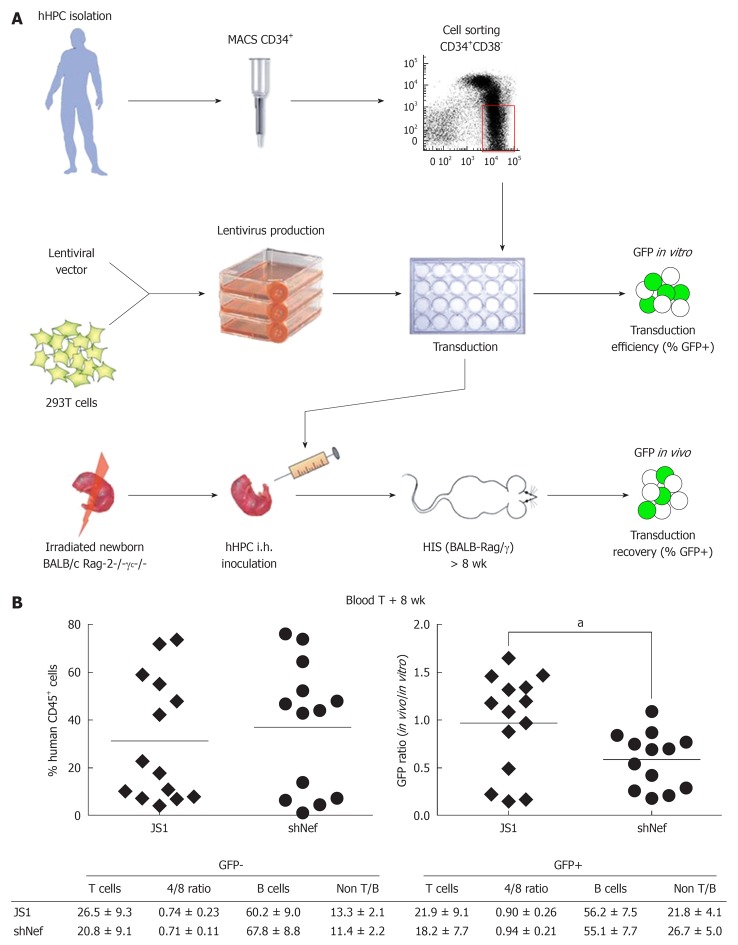
Before one can proceed to a gene therapy trial in humans, safety should ideally be demonstrated in a preclinical animal model. As the RNAi mechanism is based on perfect sequence complementarities between siRNA and the viral target, the simian immunodeficiency virus/macaque model cannot be used for such studies. However, mice with the complete human immune system were created by injection of hHPCs into immunodeficient (BALB/c Rag2-/- IL2Rγc-/-) newborn mice. All major subsets of the human innate and adaptive immune system are found in the reconstituted HIS mice[22,23]. This HIS mouse model is ideally suited to test our gene therapy for several reasons. First, the hHPCs that are engrafted in the Rag-2-/-γc-/- newborn mice (Figure 7A) are similar to the ones that we propose to modify in our ex vivo gene therapy of HIV-1 seropositive patients. Second, as the hHPCs transplanted in the mice consist of a mixture of transduced (GFP+) and non-transduced hHPCs (GFP-), the human immune system in the animals will thus be constituted by transduced shRNA-expressing cells (GFP+) and non-transduced cells (GFP-). This provides an internal control to test for adverse effects of shRNA expression on hematopoiesis. Finally, as the HIS mouse can be infected by HIV-1, both the safety and efficacy of the shRNA therapy can be evaluated in the HIS mouse model.
Figure 7.
In vivo safety studies in the HIS mouse model. A: Cell suspensions enriched for human hematopoietic progenitor cells (hHPC) are prepared from fetal liver tissue. Live nucleated CD34+ cells are magnetically sorted and further enriched for the hHPC (CD34+CD38- fraction) using fluorescence activated cell sorting. Lentiviral supernatants are produced on 293T cells. hHPC are transduced ex vivo with the shRNA-expressing lentiviral vector and injected intrahepatically into sublethally irradiated newborn BALB/c Rag-2-/-γc-/- mice. The transduction efficiency is evaluated based on GFP expression after 3.5 d in culture (GFP in vitro); B: The HIS (BALB-Rag/γ) mice are analyzed in the blood and the organs after at least 8 wk post-transplantation for the presence of human cells (%CD45+ cells) (left graph), which were analyzed for GFP recovery (right graph). The GFP recovery is the ratio between the frequency of human GFP+ cells measured in the animals (GFP in vivo) and the frequency of GFP+ hHPC injected in the newborn mice (GFP in vitro, transduction efficiency). The major subsets of the human immune system in the blood are also analyzed for their frequency and absolute number in the human GFP+ and GFP- population. Adapted from[24,70]. aP < 0.05.
The safety of an shRNA is assessed in the blood and the organs of the animals by multiple factors: (1) The presence of the human hematopoiesis-derived CD45+ GFP+ and GFP- cells; (2) the ratio between the frequency of human GFP+ cells measured in the animals and the frequency of human GFP+CD34+ cells injected in the newborn mice; and (3) the frequency and absolute number of different cell subsets of the human immune system, such as CD4+ and CD8+ T cells, B cells, monocytes and dentritic cells (Figure 7B). We previously tested the feasibility of an shRNA-based gene therapy in HIS mice reconstituted with hHPC transduced with a lentiviral vector expressing an shRNA against the HIV-1 nef gene[24]. In this model hHPC expressing anti-HIV shRNAs give rise to multi-lineage reconstitution of the human immune system in vivo and generate CD4+ T cells with the ability to resist HIV-1 replication in a sequence-specific manner. We tested our four candidate shRNAs and observed normal development of the human immune system in the animals for three of these shRNAs (Centlivre et al manuscript in preparation). A negative impact of Gag5 on the hematopoiesis of the HIS mouse was scored, confirming the in vitro findings in the CCG assay. These combined results led to the exclusion of Gag5 from the combination gene therapy (Table 1). The three “safe” shRNAs will now be combined into a single lentiviral vector for further in vivo safety tests.
PRE-EXISTING DRUG RESISTANCE MUTATIONS
Our proposed anti-HIV gene therapy will be developed for therapy-experienced HIV-1 infected individuals who have failed on regular antiretroviral drug regimens. As drug resistance mutations may affect the viral genome sequences targeted by RNAi, we investigated whether the target sequences of the top shRNA candidates are likely to acquire drug-resistance causing mutations. For this, we screened the Stanford HIV-1 drug resistance database[51,52]. The relevant drug resistance substitutions in the inhibited viral proteins are plotted in Table 1. In particular, the Protease gene sequence targeted by the set of overlapping shRNAs (Pol1-2, Pro1-7) has been implicated in the acquisition of resistance against Protease inhibitors like Nelfinavir, Aprenavir, Ritonavir and Indinavir at codons 30, 32 and 33. A treatment history that includes one of these Protease inhibitors and genotyping results that demonstrate the presence of at least one of these mutations will be an exclusion criterion for gene therapy participants.
COMBINATORIAL RNAi
The stable expression of anti-HIV shRNAs in T cells results in potent virus inhibition[16,53,54]. However, the application of a single shRNA inhibitor is not sufficient to maintain inhibition. Virus escape variants can emerge after extensive culturing[8-11,13,34]. Therefore, multiple antiviral shRNAs should be expressed simultaneously to achieve durable inhibition by raising the genetic threshold for viral escape[48,55,56]. This combinatorial strategy is analogous to current antiretroviral therapy regimens with multiple drugs that have led to significant clinical success in HIV-1 infected patients[57].
There are several ways to express multiple RNAi inhibitors against HIV-1, ranging from polycistronic miRNAs to extended multimeric shRNA transcripts[17-21]. We achieved most promising results with multiple shRNA cassettes[14,16]. To express multiple shRNAs, we initially inserted several H1-driven shRNA expression cassettes into the lentiviral vector. However, these vectors are extremely unstable as shRNA cassettes were deleted during transduction due to slippage of the Reverse Transcriptase enzyme on the repeated H1 promoter sequences[16,58]. To prevent recombination-mediated deletion of shRNA cassettes, we designed shRNA cassettes with unique promoter elements. The RNA polymerase III promoters H1, U6 and 7SK and the RNA polymerase II promoter U1 were used[4,59-61]. All these promoters have precise transcription start and termination sites and the shRNA expression levels are similar. The combination of four promoter-shRNA cassettes in R4 (U1-R/T5, U6-Pol1, 7SK-Gag5, and H1-Pol47) leads to durable virus inhibition in stably transduced T cells[16]. At present, the R4 combinatorial RNAi vector has been modified to R3 without Gag5 due to adverse cellular effects of this shRNA. The R3 lentiviral vector that confers the same potent and durable inhibition is proposed for the future clinical anti-HIV gene therapy trial.
CONCLUSION
We describe here the course that was taken to select the most potent and safe shRNA inhibitors against HIV-1, which will contribute to the development of an exclusively shRNA-based gene therapy against HIV-1. Currently, the combinatorial RNAi approach comprises three shRNAs targeting three distinct and highly conserved regions of the HIV-1 RNA genome.
The proposed RNAi gene therapy against HIV-1 will be developed for therapy-experienced HIV-1 infected individuals. During antiretroviral therapy, mutations can be selected in the genes that encode the drug-targeted viral proteins. Such mutations can interfere with RNAi attack when siRNA target sites are altered. Indeed, one shRNA inhibitor of the combinatorial shRNA vector targets a viral sequence that frequently acquires mutations to escape from Protease inhibitors. Patients who failed on Protease inhibitor containing regimens or that harbor viruses with such resistance mutations have to be excluded from the current combinatorial RNAi gene therapy. To overcome this issue, alternative shRNA regimens could be established that do not target viral genome regions known to acquire prominent drug resistance mutations. Alternatively, one could attack the resistant HIV-1 strains with modified shRNA inhibitors. We previously showed that a combination shRNA strategy directed against the wildtype and drug-escape variants was able to efficiently and durably suppress virus replication[35,48].
The selection of HIV-1 escape variants must be prevented to durably suppress the chronic virus infection. To achieve this, targeting of conserved HIV-1 RNA regions is important, as well as the simultaneous application of multiple shRNAs. Potent virus inhibition will reduce the chance of virus escape by limiting the occurrence of mutations and the genetic threshold for resistance development is increased when multiple viral sequences are targeted. We continuously work on improvement of the shRNA design and recently identified loop sequences such as miRNA-derived loops that improve the siRNA processing and yield more potent gene knockdown and HIV-1 inhibition[62]. Alternatively, targeting of cellular cofactors that are essential for HIV-1 replication represents a promising anti-escape approach. The mutation rate of the cellular DNA replication machinery is significantly lower than that of the lentiviral Reverse Transcriptase enzyme. Thus, the chance that resistance mutations are selected in host mRNAs is negligible compared to HIV-1 target sequences. Recent screens revealed several cofactor-encoding mRNAs whose knock-down resulted in diminished HIV-1 replication[63-67]. However, knock-down of cellular proteins at the mRNA level might have negative effects on cell viability and anti-host shRNAs must be carefully designed. An ideal candidate cofactor is the CCR5 co-receptor. A natural deletion in the CCR5 gene (CCR5-Δ32) has been found at 1% frequency in the Caucasian population. These individuals are resistant to HIV-1 infection and do not appear to suffer from major biological effects or health issues due to the absence of this receptor protein[68].
Within a couple of years, RNAi has moved from the laboratory to clinical trials as novel therapeutic against a variety of diseases. In 2008, the first antiviral shRNA was used in combination with a TAR decoy and CCR5-ribozyme as an RNA-based gene therapy for HIV-1 infected individuals. The transfused cells were successfully engrafted and the anti-tat/rev siRNA was detected in peripheral blood mononuclear cells (PBMCs) up to 24 mo[69]. This initial clinical result provides encouragement for the anti-HIV gene therapy that we develop based exclusively on multiple shRNAs. The extensive preclinical assays in the humanized mouse model demonstrated the safety and efficacy of this combinatorial RNAi approach, which will soon move towards clinical testing.
Footnotes
Supported by The NWO-CW (Chemical Sciences), ZonMw (Medical Sciences), and the Dutch AIDS Fund (project 2006006); the DAAD (German Academic Exchange Service); the FRM (Fondation pour la Recherche Medicale)
Peer reviewer: Eiichi N Kodama, MD, PhD, Assistant Professor, Department of Internal Medicine, Division of Emerging Infectious Diseases, Tohoku University School of Medicine, Building 1 Room 515, 2-1 Seiryo, Aoba-ku, Sendai 980-8575, Japan
S- Editor Zhang SS L- Editor A E- Editor Zheng XM
References
- 1.Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature. 1998;391:806–811. doi: 10.1038/35888. [DOI] [PubMed] [Google Scholar]
- 2.Angaji SA, Hedayati SS, Poor RH, Madani S, Poor SS, Panahi S. Application of RNA interference in treating human diseases. J Genet. 2010;89:527–537. doi: 10.1007/s12041-010-0073-3. [DOI] [PubMed] [Google Scholar]
- 3.He L, Hannon GJ. MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet. 2004;5:522–531. doi: 10.1038/nrg1379. [DOI] [PubMed] [Google Scholar]
- 4.Brummelkamp TR, Bernards R, Agami R. A system for stable expression of short interfering RNAs in mammalian cells. Science. 2002;296:550–553. doi: 10.1126/science.1068999. [DOI] [PubMed] [Google Scholar]
- 5.Hannon GJ. RNA interference. Nature. 2002;418:244–251. doi: 10.1038/418244a. [DOI] [PubMed] [Google Scholar]
- 6.Haasnoot J, Berkhout B. Nucleic acids-based therapeutics in the battle against pathogenic viruses. Handb Exp Pharmacol. 2009;189:243–263. doi: 10.1007/978-3-540-79086-0_9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Berkhout B, Sanders RW. Molecular strategies to design an escape-proof antiviral therapy. Antiviral Res. 2011;92:7–14. doi: 10.1016/j.antiviral.2011.04.002. [DOI] [PubMed] [Google Scholar]
- 8.Boden D, Pusch O, Lee F, Tucker L, Ramratnam B. Human immunodeficiency virus type 1 escape from RNA interference. J Virol. 2003;77:11531–11535. doi: 10.1128/JVI.77.21.11531-11535.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Das AT, Brummelkamp TR, Westerhout EM, Vink M, Madiredjo M, Bernards R, Berkhout B. Human immunodeficiency virus type 1 escapes from RNA interference-mediated inhibition. J Virol. 2004;78:2601–2605. doi: 10.1128/JVI.78.5.2601-2605.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.von Eije KJ, ter Brake O, Berkhout B. Human immunodeficiency virus type 1 escape is restricted when conserved genome sequences are targeted by RNA interference. J Virol. 2008;82:2895–2903. doi: 10.1128/JVI.02035-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.von Eije KJ, ter Brake O, Berkhout B. Stringent testing identifies highly potent and escape-proof anti-HIV short hairpin RNAs. J Gene Med. 2009;11:459–467. doi: 10.1002/jgm.1329. [DOI] [PubMed] [Google Scholar]
- 12.ter Brake O, von Eije KJ, Berkhout B. Probing the sequence space available for HIV-1 evolution. AIDS. 2008;22:1875–1877. doi: 10.1097/QAD.0b013e328309efe3. [DOI] [PubMed] [Google Scholar]
- 13.Westerhout EM, Ooms M, Vink M, Das AT, Berkhout B. HIV-1 can escape from RNA interference by evolving an alternative structure in its RNA genome. Nucleic Acids Res. 2005;33:796–804. doi: 10.1093/nar/gki220. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.ter Brake O, Konstantinova P, Ceylan M, Berkhout B. Silencing of HIV-1 with RNA interference: a multiple shRNA approach. Mol Ther. 2006;14:883–892. doi: 10.1016/j.ymthe.2006.07.007. [DOI] [PubMed] [Google Scholar]
- 15.ter Brake O, Berkhout B. A novel approach for inhibition of HIV-1 by RNA interference: counteracting viral escape with a second generation of siRNAs. J RNAi Gene Silencing. 2005;1:56–65. [PMC free article] [PubMed] [Google Scholar]
- 16.ter Brake O, 't Hooft K, Liu YP, Centlivre M, von Eije KJ, Berkhout B. Lentiviral vector design for multiple shRNA expression and durable HIV-1 inhibition. Mol Ther. 2008;16:557–564. doi: 10.1038/sj.mt.6300382. [DOI] [PubMed] [Google Scholar]
- 17.Liu YP, Haasnoot J, Berkhout B. Design of extended short hairpin RNAs for HIV-1 inhibition. Nucleic Acids Res. 2007;35:5683–5693. doi: 10.1093/nar/gkm596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Liu YP, Haasnoot J, ter Brake O, Berkhout B, Konstantinova P. Inhibition of HIV-1 by multiple siRNAs expressed from a single microRNA polycistron. Nucleic Acids Res. 2008;36:2811–2824. doi: 10.1093/nar/gkn109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Liu YP, Gruber J, Haasnoot J, Konstantinova P, Berkhout B. RNAi-mediated inhibition of HIV-1 by targeting partially complementary viral sequences. Nucleic Acids Res. 2009;37:6194–6204. doi: 10.1093/nar/gkp644. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Liu YP, von Eije KJ, Schopman NC, Westerink JT, ter Brake O, Haasnoot J, Berkhout B. Combinatorial RNAi against HIV-1 using extended short hairpin RNAs. Mol Ther. 2009;17:1712–1723. doi: 10.1038/mt.2009.176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Konstantinova P, de Vries W, Haasnoot J, ter Brake O, de Haan P, Berkhout B. Inhibition of human immunodeficiency virus type 1 by RNA interference using long-hairpin RNA. Gene Ther. 2006;13:1403–1413. doi: 10.1038/sj.gt.3302786. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Traggiai E, Chicha L, Mazzucchelli L, Bronz L, Piffaretti JC, Lanzavecchia A, Manz MG. Development of a human adaptive immune system in cord blood cell-transplanted mice. Science. 2004;304:104–107. doi: 10.1126/science.1093933. [DOI] [PubMed] [Google Scholar]
- 23.Gimeno R, Weijer K, Voordouw A, Uittenbogaart CH, Legrand N, Alves NL, Wijnands E, Blom B, Spits H. Monitoring the effect of gene silencing by RNA interference in human CD34+ cells injected into newborn RAG2-/- gammac-/- mice: functional inactivation of p53 in developing T cells. Blood. 2004;104:3886–3893. doi: 10.1182/blood-2004-02-0656. [DOI] [PubMed] [Google Scholar]
- 24.ter Brake O, Legrand N, von Eije KJ, Centlivre M, Spits H, Weijer K, Blom B, Berkhout B. Evaluation of safety and efficacy of RNAi against HIV-1 in the human immune system (Rag-2(-/-)gammac(-/-)) mouse model. Gene Ther. 2009;16:148–153. doi: 10.1038/gt.2008.124. [DOI] [PubMed] [Google Scholar]
- 25.Seppen J, Rijnberg M, Cooreman MP, Oude Elferink RP. Lentiviral vectors for efficient transduction of isolated primary quiescent hepatocytes. J Hepatol. 2002;36:459–465. doi: 10.1016/s0168-8278(01)00308-7. [DOI] [PubMed] [Google Scholar]
- 26.Westerhout EM, Berkhout B. A systematic analysis of the effect of target RNA structure on RNA interference. Nucleic Acids Res. 2007;35:4322–4330. doi: 10.1093/nar/gkm437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Low JT, Knoepfel SA, Watts JM, ter Brake O, Berkhout B, Weeks KM. SHAPE-directed discovery of potent shRNA inhibitors of HIV-1. Mol Ther. 2012;20:820–828. doi: 10.1038/mt.2011.299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Watts JM, Dang KK, Gorelick RJ, Leonard CW, Bess JW, Swanstrom R, Burch CL, Weeks KM. Architecture and secondary structure of an entire HIV-1 RNA genome. Nature. 2009;460:711–716. doi: 10.1038/nature08237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Haasnoot J, Westerhout EM, Berkhout B. RNA interference against viruses: strike and counterstrike. Nat Biotechnol. 2007;25:1435–1443. doi: 10.1038/nbt1369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.McIntyre GJ, Groneman JL, Yu YH, Jaramillo A, Shen S, Applegate TL. 96 shRNAs designed for maximal coverage of HIV-1 variants. Retrovirology. 2009;6:55. doi: 10.1186/1742-4690-6-55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.van der Kuyl AC, Jurriaans S, Back NK, Sprenger HG, van der Werf TS, Zorgdrager F, Berkhout B, Cornelissen M. Unusual cluster of HIV type 1 dual infections in Groningen, The Netherlands. AIDS Res Hum Retroviruses. 2011;27:429–433. doi: 10.1089/aid.2010.0175. [DOI] [PubMed] [Google Scholar]
- 32.Nishitsuji H, Kohara M, Kannagi M, Masuda T. Effective suppression of human immunodeficiency virus type 1 through a combination of short- or long-hairpin RNAs targeting essential sequences for retroviral integration. J Virol. 2006;80:7658–7666. doi: 10.1128/JVI.00078-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Lee SK, Dykxhoorn DM, Kumar P, Ranjbar S, Song E, Maliszewski LE, François-Bongarçon V, Goldfeld A, Swamy NM, Lieberman J, Shankar P. Lentiviral delivery of short hairpin RNAs protects CD4 T cells from multiple clades and primary isolates of HIV. Blood. 2005;106:818–826. doi: 10.1182/blood-2004-10-3959. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Unwalla HJ, Li HT, Bahner I, Li MJ, Kohn D, Rossi JJ. Novel Pol II fusion promoter directs human immunodeficiency virus type 1-inducible coexpression of a short hairpin RNA and protein. J Virol. 2006;80:1863–1873. doi: 10.1128/JVI.80.4.1863-1873.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Schopman NC, ter Brake O, Berkhout B. Anticipating and blocking HIV-1 escape by second generation antiviral shRNAs. Retrovirology. 2010;7:52. doi: 10.1186/1742-4690-7-52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Biffi A, Bartolomae CC, Cesana D, Cartier N, Aubourg P, Ranzani M, Cesani M, Benedicenti F, Plati T, Rubagotti E, et al. Lentiviral vector common integration sites in preclinical models and a clinical trial reflect a benign integration bias and not oncogenic selection. Blood. 2011;117:5332–5339. doi: 10.1182/blood-2010-09-306761. [DOI] [PubMed] [Google Scholar]
- 37.Dropulic B. Lentivirus in the clinic. Mol Ther. 2001;4:511–512. doi: 10.1006/mthe.2001.0501. [DOI] [PubMed] [Google Scholar]
- 38.Cartier N, Hacein-Bey-Abina S, Bartholomae CC, Veres G, Schmidt M, Kutschera I, Vidaud M, Abel U, Dal-Cortivo L, Caccavelli L, et al. Hematopoietic stem cell gene therapy with a lentiviral vector in X-linked adrenoleukodystrophy. Science. 2009;326:818–823. doi: 10.1126/science.1171242. [DOI] [PubMed] [Google Scholar]
- 39.Van den Haute C, Eggermont K, Nuttin B, Debyser Z, Baekelandt V. Lentiviral vector-mediated delivery of short hairpin RNA results in persistent knockdown of gene expression in mouse brain. Hum Gene Ther. 2003;14:1799–1807. doi: 10.1089/104303403322611809. [DOI] [PubMed] [Google Scholar]
- 40.Harper SQ, Staber PD, He X, Eliason SL, Martins IH, Mao Q, Yang L, Kotin RM, Paulson HL, Davidson BL. RNA interference improves motor and neuropathological abnormalities in a Huntington's disease mouse model. Proc Natl Acad Sci USA. 2005;102:5820–5825. doi: 10.1073/pnas.0501507102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Blömer U, Naldini L, Verma IM, Trono D, Gage FH. Applications of gene therapy to the CNS. Hum Mol Genet. 1996;5 Spec No:1397–1404. doi: 10.1093/hmg/5.supplement_1.1397. [DOI] [PubMed] [Google Scholar]
- 42.Naldini L. Lentiviruses as gene transfer agents for delivery to non-dividing cells. Curr Opin Biotechnol. 1998;9:457–463. doi: 10.1016/s0958-1669(98)80029-3. [DOI] [PubMed] [Google Scholar]
- 43.Ralph GS, Radcliffe PA, Day DM, Carthy JM, Leroux MA, Lee DC, Wong LF, Bilsland LG, Greensmith L, Kingsman SM, et al. Silencing mutant SOD1 using RNAi protects against neurodegeneration and extends survival in an ALS model. Nat Med. 2005;11:429–433. doi: 10.1038/nm1205. [DOI] [PubMed] [Google Scholar]
- 44.Liu YP, Berkhout B. Lentiviral delivery of RNAi effectors against HIV-1. Curr Top Med Chem. 2009;9:1130–1143. doi: 10.2174/156802609789630866. [DOI] [PubMed] [Google Scholar]
- 45.Liu YP, Vink MA, Westerink JT, Ramirez de Arellano E, Konstantinova P, Ter Brake O, Berkhout B. Titers of lentiviral vectors encoding shRNAs and miRNAs are reduced by different mechanisms that require distinct repair strategies. RNA. 2010;16:1328–1339. doi: 10.1261/rna.1887910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Liu YP, Westerink JT, ter Brake O, Berkhout B. RNAi-inducing lentiviral vectors for anti-HIV-1 gene therapy. Methods Mol Biol. 2011;721:293–311. doi: 10.1007/978-1-61779-037-9_18. [DOI] [PubMed] [Google Scholar]
- 47.Poluri A, Sutton RE. Titers of HIV-based vectors encoding shRNAs are reduced by a dicer-dependent mechanism. Mol Ther. 2008;16:378–386. doi: 10.1038/sj.mt.6300370. [DOI] [PubMed] [Google Scholar]
- 48.ter Brake O, Berkhout B. Lentiviral vectors that carry anti-HIV shRNAs: problems and solutions. J Gene Med. 2007;9:743–750. doi: 10.1002/jgm.1078. [DOI] [PubMed] [Google Scholar]
- 49.Grimm D, Streetz KL, Jopling CL, Storm TA, Pandey K, Davis CR, Marion P, Salazar F, Kay MA. Fatality in mice due to oversaturation of cellular microRNA/short hairpin RNA pathways. Nature. 2006;441:537–541. doi: 10.1038/nature04791. [DOI] [PubMed] [Google Scholar]
- 50.Eekels JJ, Pasternak AO, Schut AM, Geerts D, Jeeninga RE, Berkhout B. A competitive cell growth assay for the detection of subtle effects of gene transduction on cell proliferation. Gene Ther. 2011:Epub ahead of print. doi: 10.1038/gt.2011.191. [DOI] [PubMed] [Google Scholar]
- 51.Rhee SY, Gonzales MJ, Kantor R, Betts BJ, Ravela J, Shafer RW. Human immunodeficiency virus reverse transcriptase and protease sequence database. Nucleic Acids Res. 2003;31:298–303. doi: 10.1093/nar/gkg100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Shafer RW. Rationale and uses of a public HIV drug-resistance database. J Infect Dis. 2006;194 Suppl 1:S51–S58. doi: 10.1086/505356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Yamamoto T, Miyoshi H, Yamamoto N, Yamamoto N, Inoue J, Tsunetsugu-Yokota Y. Lentivirus vectors expressing short hairpin RNAs against the U3-overlapping region of HIV nef inhibit HIV replication and infectivity in primary macrophages. Blood. 2006;108:3305–3312. doi: 10.1182/blood-2006-04-014829. [DOI] [PubMed] [Google Scholar]
- 54.Li MJ, Bauer G, Michienzi A, Yee JK, Lee NS, Kim J, Li S, Castanotto D, Zaia J, Rossi JJ. Inhibition of HIV-1 infection by lentiviral vectors expressing Pol III-promoted anti-HIV RNAs. Mol Ther. 2003;8:196–206. doi: 10.1016/s1525-0016(03)00165-5. [DOI] [PubMed] [Google Scholar]
- 55.Berkhout B. RNA interference as an antiviral approach: targeting HIV-1. Curr Opin Mol Ther. 2004;6:141–145. [PubMed] [Google Scholar]
- 56.Leonard JN, Schaffer DV. Computational design of antiviral RNA interference strategies that resist human immunodeficiency virus escape. J Virol. 2005;79:1645–1654. doi: 10.1128/JVI.79.3.1645-1654.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Deeks SG. Antiretroviral treatment of HIV infected adults. BMJ. 2006;332:1489. doi: 10.1136/bmj.332.7556.1489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Mcintyre GJ, Yu YH, Tran A, Jaramillo AB, Arndt AJ, Millington ML, Boyd MP, Elliott FA, Shen SW, Murray JM, et al. Cassette deletion in multiple shRNA lentiviral vectors for HIV-1 and its impact on treatment success. Virol J. 2009;6:184. doi: 10.1186/1743-422X-6-184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Yu JY, DeRuiter SL, Turner DL. RNA interference by expression of short-interfering RNAs and hairpin RNAs in mammalian cells. Proc Natl Acad Sci USA. 2002;99:6047–6052. doi: 10.1073/pnas.092143499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Koper-Emde D, Herrmann L, Sandrock B, Benecke BJ. RNA interference by small hairpin RNAs synthesised under control of the human 7S K RNA promoter. Biol Chem. 2004;385:791–794. doi: 10.1515/BC.2004.103. [DOI] [PubMed] [Google Scholar]
- 61.Denti MA, Rosa A, Sthandier O, De Angelis FG, Bozzoni I. A new vector, based on the PolII promoter of the U1 snRNA gene, for the expression of siRNAs in mammalian cells. Mol Ther. 2004;10:191–199. doi: 10.1016/j.ymthe.2004.04.008. [DOI] [PubMed] [Google Scholar]
- 62.Schopman NC, Liu YP, Konstantinova P, ter Brake O, Berkhout B. Optimization of shRNA inhibitors by variation of the terminal loop sequence. Antiviral Res. 2010;86:204–211. doi: 10.1016/j.antiviral.2010.02.320. [DOI] [PubMed] [Google Scholar]
- 63.Yeung ML, Houzet L, Yedavalli VS, Jeang KT. A genome-wide short hairpin RNA screening of jurkat T-cells for human proteins contributing to productive HIV-1 replication. J Biol Chem. 2009;284:19463–19473. doi: 10.1074/jbc.M109.010033. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Eekels JJ, Geerts D, Jeeninga RE, Berkhout B. Long-term inhibition of HIV-1 replication with RNA interference against cellular co-factors. Antiviral Res. 2011;89:43–53. doi: 10.1016/j.antiviral.2010.11.005. [DOI] [PubMed] [Google Scholar]
- 65.Brass AL, Dykxhoorn DM, Benita Y, Yan N, Engelman A, Xavier RJ, Lieberman J, Elledge SJ. Identification of host proteins required for HIV infection through a functional genomic screen. Science. 2008;319:921–926. doi: 10.1126/science.1152725. [DOI] [PubMed] [Google Scholar]
- 66.König R, Zhou Y, Elleder D, Diamond TL, Bonamy GM, Irelan JT, Chiang CY, Tu BP, De Jesus PD, Lilley CE, et al. Global analysis of host-pathogen interactions that regulate early-stage HIV-1 replication. Cell. 2008;135:49–60. doi: 10.1016/j.cell.2008.07.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Zhou H, Xu M, Huang Q, Gates AT, Zhang XD, Castle JC, Stec E, Ferrer M, Strulovici B, Hazuda DJ, et al. Genome-scale RNAi screen for host factors required for HIV replication. Cell Host Microbe. 2008;4:495–504. doi: 10.1016/j.chom.2008.10.004. [DOI] [PubMed] [Google Scholar]
- 68.Huang Y, Paxton WA, Wolinsky SM, Neumann AU, Zhang L, He T, Kang S, Ceradini D, Jin Z, Yazdanbakhsh K, et al. The role of a mutant CCR5 allele in HIV-1 transmission and disease progression. Nat Med. 1996;2:1240–1243. doi: 10.1038/nm1196-1240. [DOI] [PubMed] [Google Scholar]
- 69.DiGiusto DL, Krishnan A, Li L, Li H, Li S, Rao A, Mi S, Yam P, Stinson S, Kalos M, et al. RNA-based gene therapy for HIV with lentiviral vector-modified CD34(+) cells in patients undergoing transplantation for AIDS-related lymphoma. Sci Transl Med. 2010;2:36ra43. doi: 10.1126/scitranslmed.3000931. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Centlivre M, Zhou X, Pouw SM, Weijer K, Kleibeuker W, Das AT, Blom B, Seppen J, Berkhout B, Legrand N. Autoregulatory lentiviral vectors allow multiple cycles of doxycycline-inducible gene expression in human hematopoietic cells in vivo. Gene Ther. 2010;17:14–25. doi: 10.1038/gt.2009.109. [DOI] [PubMed] [Google Scholar]