Abstract
The nucleolus is implicated in sensing and responding to cellular stress by stabilizing p53. The pro-apoptotic effect of p53 is associated with several neurodegenerative disorders, including Huntington's disease (HD), which is characterized by the progressive loss of medium spiny neurons (MSNs) in the striatum. Here we show that disruption of nucleolar integrity and function causes nucleolar stress and is an early event in MSNs of R6/2 mice, a transgenic model of HD. Targeted perturbation of nucleolar function in MSNs by conditional knockout of the RNA polymerase I-specific transcription initiation factor IA (TIF-IA) leads to late progressive striatal degeneration, HD-like motor abnormalities and molecular signatures. Significantly, p53 prolongs neuronal survival in TIF-IA-deficient MSNs by transient upregulation of phosphatase and tensin homolog deleted on chromosome 10 (PTEN), a tumor suppressor that inhibits mammalian target of rapamycin signaling and induces autophagy. The results emphasize the initial role of nucleolar stress in neurodegeneration and uncover a p53/PTEN-dependent neuroprotective response.
Keywords: nucleolus, p53, Huntington's disease, PTEN, neuroprotection
Neurodegenerative diseases are associated with loss of effective defense mechanisms that protect neurons against stress conditions.1 It is known that cellular stress leads to downregulation of ribosomal RNA (rRNA) synthesis, a process that is associated with ageing, a prominent parameter for developing neurodegenerative diseases.2 Although neuronal vulnerability depends on the type of disease, impaired nucleolar activity and p53-mediated apoptosis is a common feature of neurodegenerative disorders.3, 4, 5, 6, 7, 8 Under normal conditions, cellular p53 levels are low, whereas both the amount and activity of p53 increase upon stress, thus regulating the balance between life and death.9
Cellular insults, such as oxidative stress or DNA damage, lead to nucleolar stress, namely disruption of nucleolar function and structure, stabilization of p53 and p53-dependent cell cycle arrest, senescence and apoptosis, reinforcing the central role of p53 in surveying cellular health.9, 10 This tight link between RNA-polymerase I (Pol I) activity, nucleolar integrity and p53-mediated damage control has also been observed after genetic inactivation of the Pol I-specific transcription initiation factor IA (TIF-IA), placing the Pol I transcription machinery in the centre of control pathways that are influenced by p53.11 Under normal conditions, the E3 ubiquitin-ligase MDM2/HDM2 controls the abundance of p53 by ubiquitinylation, marking p53 for proteasome-dependent proteolysis. Under conditions of nucleolar stress, for example, after genetic inactivation of TIF-IA, the p53-MDM2 complex is disrupted by the interaction with released ribosomal proteins, p53 levels are strongly enhanced and p53-dependent pathways are activated.12, 13 The finding that inhibition of Pol I transcription induces the p53 program raises the exciting possibility that conditional inactivation of TIF-IA may be a powerful approach to induce and investigate nucleolar and p53-dependent stress signaling in specific cells. Indeed, inhibition of rDNA transcription by Cre-mediated knock out of the TIF-IA gene in neural progenitors causes p53-dependent apoptosis.14 Moreover, perturbation of Pol I activity by TIF-IA ablation in dopaminergic neurons induces progressive neurodegeneration, leading to symptoms similar to those observed in Parkinson's disease.5
There is growing evidence that nucleolar stress is also involved in the pathogenesis of Huntington's disease (HD), a fatal neurodegenerative disorder caused by expansion of a CAG repeat in the gene encoding the protein huntingtin (Htt). HD results in specific degeneration of the striatum by yet poorly understood mechanisms. Abnormal nucleoli have been observed in HD15, 16 and Rrs1 (regulator of ribosome synthesis), a protein inhibiting transcription of both, rRNA and ribosomal protein genes, is upregulated in a HD mouse model.17 In cellular and animal models of HD, reduced levels of the Pol I transcription factor UBF1 have been observed,18 linking HD to impaired rDNA transcription. More recently, the expansion of CAG repeats has been shown to confer pathogenic property to the mutant RNA by impairing the activity of the nucleolar protein nucleolin and decreasing rRNA transcription.7 Interestingly, the activity of TIF-IA is regulated by brain-derived neurotrophic factor,19 a protein that is downregulated in HD and is essential to sustain striatal neurons.20 These findings underscore the functional link between nucleolar dysfunction and HD.
In this study, we have investigated the impact of nucleolar stress on striatal degeneration. We show that nucleolar structure and function are impaired in R6/2 transgenic mice that develop neurological symptoms similar to HD because of expression of a mutated human HTT fragment.21 Targeted ablation of TIF-IA in medium spiny neurons (MSNs), the major neuronal population of the striatum that is progressively lost in HD, causes disruption of nucleoli, accumulation of p53, progressive degeneration of the striatum and HD-like abnormalities. Interestingly, upregulation of p53 leads to a transient pro-survival response, involving p53-dependent induction of phosphatase and tensin homolog deleted on chromosome 10 (PTEN), inhibition of mammalian target of rapamycin (mTOR) signaling and induction of autophagy. The data reinforce the role of nucleolar stress in the development of HD and uncover a unique neuroprotective response exerted by the tumor suppressors p53 and PTEN.
Results
Targeted disruption of nucleoli in striatal neurons causes HD-like lesions
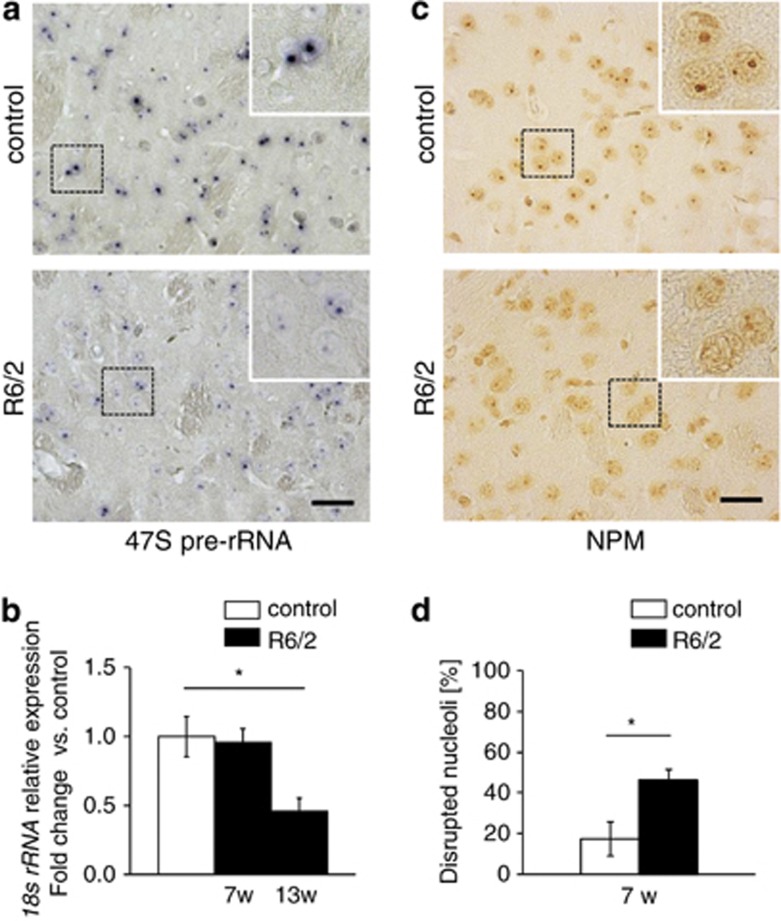
To test whether nucleolar structure and/or function is an early alteration in HD animal models, we analyzed pre-rRNA synthesis in striatal neurons of R6/2 transgenic mice, which express a mutant exon 1 of HTT and develop a progressive HD-like behavioral and neuropathological phenotype.21 At 7 weeks of age, a stage where most neurological symptoms are still absent and there is no loss of neurons (Supplementary Figures 1A, B and D) pre-rRNA levels in the striatum were decreased (Figure 1a). Consistently, the level of mature 18S rRNA monitored by qRT-PCR was significantly decreased in 13-week-old R6/2 mice (Figure 1b). Nucleolar stress and impaired ribosomal integrity in R6/2 mice were validated by decreased immunoreactivity of the rRNA-specific Y10b epitope22, 23 as well as diminished immunostaining of NPM/B23, a nucleolar marker protein, indicating that the nucleolar integrity and activity are impaired (Supplementary Figure 1C; Figures 1c and d).
Figure 1.
Nucleolar activity and integrity are decreased in R6/2 models of HD. (a) Visualization of 47S pre-rRNA by in situ hybridization on striatal sections of 7-week-old control (upper panel) and R6/2 mice (lower panel). (b) RT-qPCR analysis of 18S rRNA expression in striata from R6/2 mice (n=4) and controls (n=8) at 7 and 13 weeks. Values represent means±S.E.M. (c and d) Quantitative analysis of striatal cells with decreased nucleolar NPM/B23 (% of cells per mm2) in control and 7-week- old R6/2 mice. Values represent means±S.E.M. (n=3–4). Scale bars: 60 μm
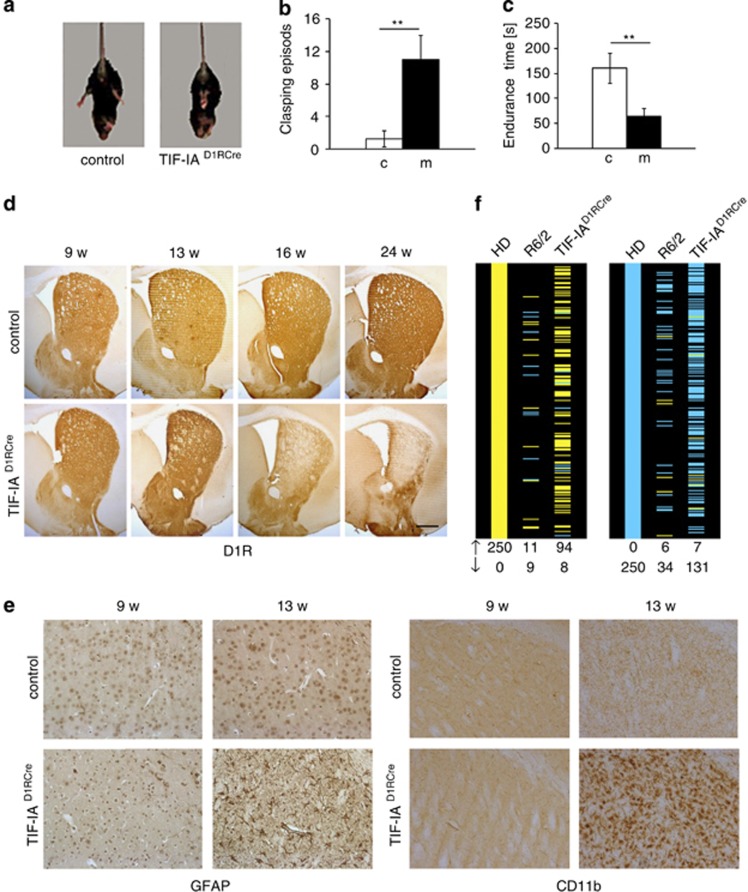
To investigate the link between impaired nucleolar function and striatal degeneration, we inhibited Pol I transcription in MSNs by specific ablation of the gene encoding the nucleolar transcription factor TIF-IA. Genetic inactivation of TIF-IA has been shown to lead to p53-dependent cell cycle arrest, senescence and apoptosis.11 To knockout TIF-IA in MSNs, we crossed TIF-IAflox/flox mice with transgenic mice expressing the Cre recombinase under the control of the promoter of the dopamine receptor D1 (D1R) gene.24 Mutant mice, termed TIF-IAD1RCre, were born at the expected ratio and showed no obvious phenotype after birth. After 40 weeks, TIF-IAD1RCre mice weighted less than control littermates (63%) and exhibited abnormal posture and gait (Movie S1). The onset of such abnormalities, monitored by feet clasping behavior and impaired motor coordination in rotarod performance tests was evident after 16 weeks (Figures 2a–c). Immunostaining of D1R-positive striatal neurons in brain sections demonstrated significant reduction of the striatum in TIF-IAD1RCre mice (Figure 2d), indicating that depletion of TIF-IA caused striatal neurodegeneration and motor deficits. Furthermore, staining of striatal sections with an antibody against the glial fibrillary acidic protein (GFAP) revealed reactive astrogliosis in 13-week-old TIF-IAD1RCre mice (Figure 2e, left panels). Signs of neuroinflammation and striatal lesions were also observed at the same stage by increased microgliosis (Figure 2e, right panels) and increased staining of degenerating neurons with FluoroJade C (Supplementary Figure 2A). TIF-IA-deficient MSNs suffered from enhanced oxidative damage, as shown by increased nitration of tyrosine residues, hydroxylation of deoxyguanosine-residues in DNA, formation of neuroketal (NK) adducts (Supplementary Figures 3A–C) and protein carbonylation (Supplementary Figure 4A).
Figure 2.
HD-like phenotypic alterations in TIF-IAD1RCre mice. (a) Representative images showing hind limb clasping in a tail-suspended TIF-IAD1RCre mutant, but not in a control mouse at 16 weeks. (b) Quantification of clasping frequency in control (c) and TIF-IAD1RCre (m) mice at 16 weeks. Values represent means±S.E.M. (n=4–7). (c) Analysis of motor coordination of control (c) and TIF-IAD1RCre (m) mice revealed by endurance in the rotarod test. Values for endurance time are means±S.E.M. (n=4–7). (d) Representative images showing staining of striata with anti-dopamine D1 receptor (D1R) antibodies in controls and TIF-IAD1RCre mutant mice at different stages. (e) Astro- and microgliosis detected by IHC with GFAP and CD11b-specific antibodies in TIF-IAD1RCre mutants and controls on striatal paraffin sections at 13 weeks of age but not at 9 weeks. Scale bars: D1R, 250 μm; GFAP and CD11b, 60 μm. (f) Comparison of striatal gene expression profiles in 13-week-old TIF-IAD1RCre mice, human HD and 9-week-old R6/2.25 Concordance between the 250 most significant genes upregulated (left panel) and downregulated (right panel) in human HD caudate and their orthologs in R6/2 and TIF-IAD1RCre mice (ranked by P-value). Yellow – statistically significant (P<0.001) upregulated genes, blue–statistically significant (P<0.001) downregulated genes. Data of HD and R6/2 profiling are derived from Strand et al.25
As knockout of TIF-IA in MSNs induced a HD-like phenotype, we profiled gene expression in 13-week-old control and TIF-IAD1RCre mice on microarrays and compared the data with published mRNA profiles from R6/2 mice midway through disease progression and from human HD patients of low pathological grade.25, 26 Of 250 significantly upregulated or downregulated genes in human HD brains, R6/2 mice exhibited an overlap of 11 upregulated and 34 downregulated gene orthologs (Figure 2f). Notably, in TIF-IAD1RCre mice, 94 upregulated and 131 downregulated genes matched to human genes that were differentially expressed in HD patients, indicating that inhibition of rRNA synthesis in MSNs triggers a transcriptional program that resembles human HD (Figure 2f).
p53 is required for survival of stress-exposed MSNs
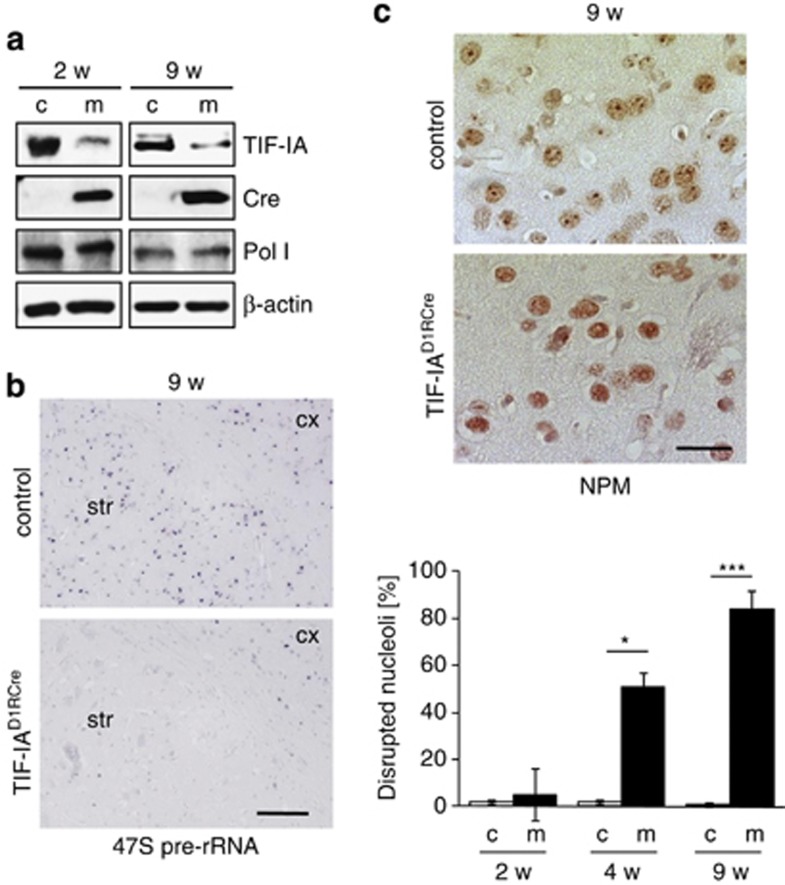
One of the most intriguing roles of the nucleolus in cellular homeostasis is its participation in sensing cellular stress signals and transmitting them to the p53 stabilization system.12, 27 Downstream effects of TIF-IA depletion in MSNs (Figure 3a) included inhibition of pre-rRNA synthesis (Figure 3b), loss of mature rRNA (Supplementary Figure 2B), perturbation of nucleolar structure (Figure 3c) and increased level of p53 protein (Figures 4a and b).
Figure 3.
Early impairment of nucleolar activity and integrity precedes neurodegeneration in TIF-IAD1RCre mice. (a) Analysis of TIF-IA levels in striatal extracts from 2- to 9-week-old TIF-IAD1RCre mutants (m) and control (c) littermates by western blot. Detection of Cre-recombinase (Cre) confirmed the mutant phenotype, detection of Pol I and β-actin served as loading controls. (b) Visualization of 47S pre-rRNA by in situ hybridization of striatal sections from control mice (upper panel) and TIF-IAD1RCre mutants (lower panel) 9 weeks after birth. Scale bar, 120 μm. (c) Immunostaining showing decreased nucleolar NPM/B23 in striatal sections of 9-week-old control (left upper panel) and mutant (left lower panel) mice. Scale bar: 60 μm. Bottom panel: quantitative analysis of striatal cells with nucleoplasmatic NPM/B23 localization relative to the total number of cells counted per field. Values represent means±S.E.M. (n=3–4)
Figure 4.
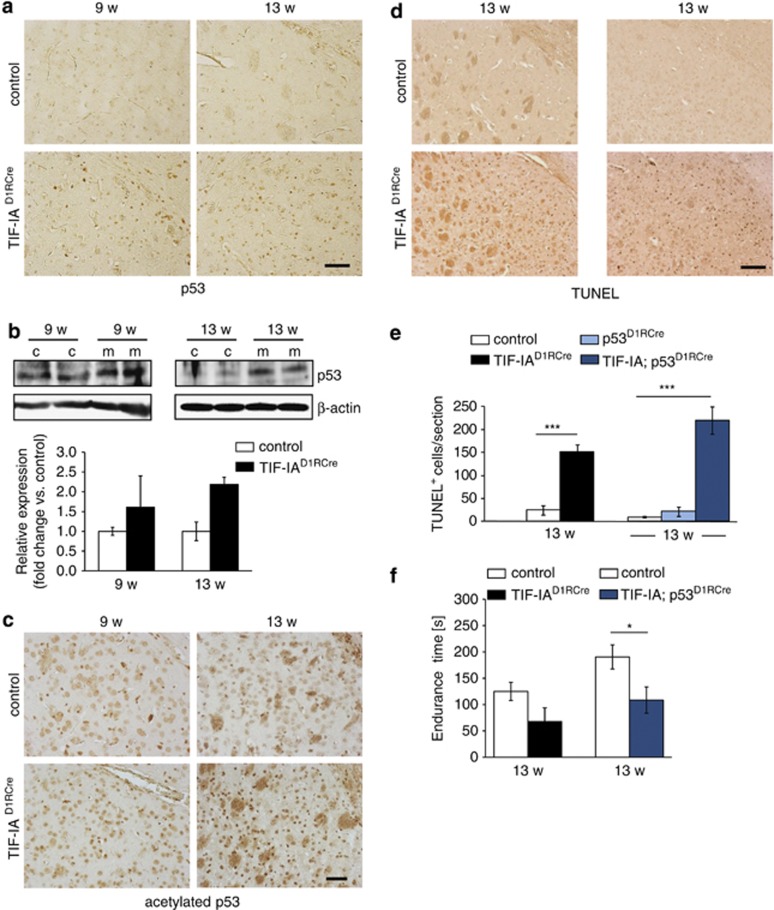
Nucleolar disruption in striatal neurons increases p53 levels and causes neuronal death. (a) Immunohistochemical analysis showing upregulation of p53 protein in 9- and 13-week-old TIF-IAD1RCre mutants. (b) Representative western blots detecting p53 in striatal extracts from control (c) and TIF-IAD1RCre mutants (m) at 9 and 13 weeks. Graphs show quantification of p53 levels relative to β-actin. Values represent means±S.E.M. (9 weeks, n=3; 13w, n=2). (c) Immunohistochemical analysis of striatal brain sections from 9- to 13-week-old mice stained with anti-acetyl-p53 (Lys373,382) antibodies. Scale bars: 60 μm. (d) Analysis of cell death by TUNEL assay in striata of control, TIF-IAD1RCre and TIF-IA;p53D1RCre mutant mice at the age of 13 weeks. Scale bar: 120 μm. (c) Immunohistochemical analysis on striatal brain sections from 9- to 13-week-old mice analyzed by anti-acetylp53 (Lys373,382) antibody in TIF-IAD1RCre mutants and controls. Scale bars: 60 μm. (d) Analysis of cell death by TUNEL assay in striata of control, TIF-IAD1RCre and TIF-IA;p53D1RCre mutants at the age of 13 weeks. Scale bar: 120 μm. (e) Quantification of TUNEL-positive cells within four striatal sections per mouse. Values represent means ±S.E.M. for TIF-IAD1RCre (n=4), TIF-IA;p53D1RCre (n=5), p53D1RCre (n=3) and the respective control mice (n=4–6). (f) Analysis of motor coordination of control, TIF-IAD1RCre and TIF-IA;p53D1RCre mice at 13 weeks analyzed by endurance in the rotarod test. Values for endurance time are means±S.E.M. (n=4–5, sm; n=6–10, dm)
It has been recently shown that p53 is stabilized by acetylation in TIF-IA-deficient cells.28 To investigate whether upregulation of p53 in TIF-IAD1RCre mice might be induced in a similar way, we analyzed p53 acetylation at Lys373 and Lys382. Staining of paraffin sections with anti-acetyl-p53 antibodies revealed sparse acetylation of p53 in 9-week-old TIF-IAD1RCre mice, but increased acetylation levels at 13 weeks (Figure 4c). This analysis revealed sparse acetylation of p53 in 9-week-old TIF-IAD1RCre mice, which was mostly increased at 13 weeks (Figure 4c). This result is consistent with data showing that acetylation of p53 is required for p53-dependent induction of apoptosis.29
To examine whether loss of p53 reduces neurodegeneration, we knocked out both TIF-IA and p53 in MSNs (TIF-IA; p53D1RCre mice) and monitored neuronal death in the striatum of TIF-IAD1RCre and TIF-IA;p53D1RCre mice. Terminal deoxynucleotidyl transferase dUTP nick end labeling (TUNEL) assays revealed increase of apoptotic cells in the striatum of 13-week-old TIF-IAD1RCre and TIF-IA;p53D1RCre mice compared with control animals, whereas loss of p53 alone did not affect MSN survival (Figures 4d and e). The number of MSNs with pyknotic nuclei was similar in 13-week-old TIF-IAD1RCre and TIF-IA;p53D1RCre mice (Supplementary Figures 5A and B), supporting that loss of p53 does not ameliorate the effects of nucleolar stress on neuronal survival. In addition, 13-week-old TIF-IAD1RCre and TIF-IA;p53D1RCre mice showed similar reduced endurance on the rotarod in comparison with control littermates (Figure 4f).
p53 induces expression of PTEN, inhibition of mTOR signaling and activation of autophagy
To investigate the molecular mechanisms regulating the balance between survival and death, we compared gene expression profiles in control and TIF-IAD1RCre mice at the early and late phase of p53 increase, that is, at 9 and 13 weeks after birth (Supplementary Figures 6A–C). At 9 weeks, the expression of 111 genes, mainly related to ribosomes and protein synthesis, was significantly changed in TIF-IAD1RCre mice (Supplementary Figure 6A and Supplementary Table 1). Interestingly, genes that are downregulated in human HD and R6/2 models, namely CaMK4, Cnr1, Homer1 and Ryr1,30, 31, 32, 33 were also less expressed in 13-week-old TIF-IAD1RCre mice (Supplementary Figure 6D), suggesting a common expression program upon striatal degeneration.
Among the 22 genes that are differentially expressed in 9- and 13-week-old TIF-IAD1RCre mutants (Supplementary Figure 6B), the tumor-suppressor gene Pten was upregulated after 9 weeks but not after 13 weeks. This transient increase of PTEN was validated at the mRNA and protein level (Figures 5a and b, upper panel). Consistent with Pten being regulated by p53,34 its expression was not increased in striata of TIF-IA;p53D1RCre mice (double mutants, dm) lacking both TIF-IA and p53 (Figure 5b, lower panel and Figure 5c). As PTEN sustains cell survival by suppressing the phosphatidylinositol-3-kinase (PI3K)/AKT/mTOR pathway,35 we examined whether PTEN exerts this function also in TIF-IA-deficient MSNs. Indeed, phosphorylation of the ribosomal protein S6, a downstream target of mTOR, was significantly decreased in 9-week-old TIF-IAD1RCre mice (Figure 5d and Supplementary Figure 7A). Similarly, phosphorylation of the eukaryotic initiation factor 4E binding protein, a direct mTOR target, was reduced in TIF-IAD1RCre mice, supporting that upregulation of PTEN impairs mTOR function in MSNs (Figure 5e).
Figure 5.
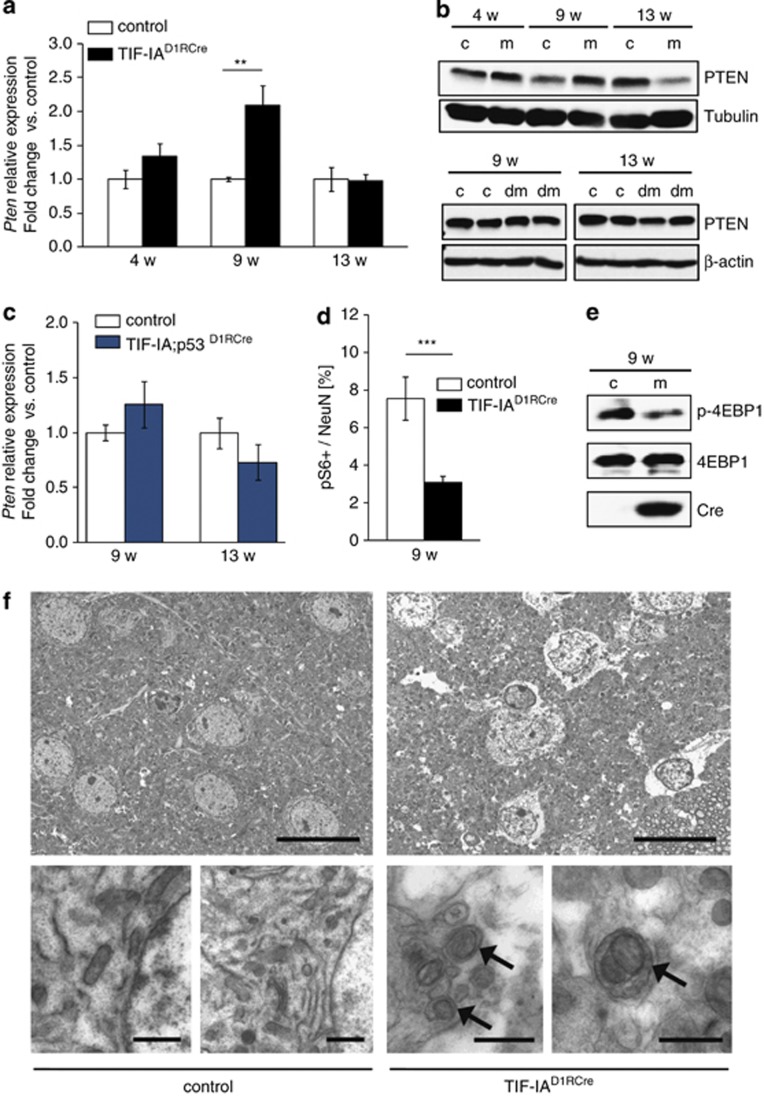
p53-dependent upregulation of PTEN inhibits mTOR and activates autophagy in MSNs. (a) RT-qPCR analysis of Pten expression in striata from TIF-IAD1RCre mutants (m) and control littermates (c) at different stages. Values represent means±S.E.M. (n=4–5). (b) Representative western blots monitoring striatal PTEN protein levels in control (c), TIF-IAD1RCre (m) and TIF-IA;p53D1RCre (dm) mice at different age. Tubulin or β-actin were used as loading controls. (c) Pten expression analyzed by RT-qPCR using RNA from striata of TIF-IA;p53D1RCre mutants or control mice at 9 and 13 weeks. Values represent means ±S.E.M. (n=3–4). (d) Quantification of the phosphoS6 (pS6)-positive cells relative to the total number of NeuN-positive cells detected by immunostaining of striatal sections (5–8 serial sections per mouse) from controls and TIF-IAD1RCre mutants at 9 weeks. Values represent means ±S.E.M. (n= 3, control; n=4, mutant). (e) Representative western blot showing downregulation of phosphorylated 4EBP1 (p-4EBP1) in TIF-IAD1RCre mutants at 9 weeks of age. (f) Ultrastructural analysis of the striatal tissue in control and mutant mice (9 weeks) by electron microscopy. Left upper and lower panels: cytology of striatal neurons from control animals. Right upper panel: striatal neurons of TIF-IAD1RCre mutants show less electrondense cytoplasm than in the control, indicating reduced density of the endoplasm granular mass as a sign of general cellular impairment. Right lower panel: higher magnification reveals double membrane autophagosomes (arrows) engulfing entire organelles in the mutant. Scale bar in upper panel: 1.8 μm. Scale bar in lower panel: 0.8 μm
Inhibition of mTOR activity is known to trigger autophagy, a catabolic process that is beneficial for cell survival by removal and recycling of damaged cellular components.36 To examine whether autophagy is induced in TIF-IA-deficient MSNs, we monitored formation of autophagosomes in striata of 9-week-old control and mutant mice by electron microscopy. In TIF-IAD1RCre mutants, but not in control animals, autophagosomes partly engulfing entire organelles, such as mitochondria, were readily detectable and pathological alterations in the cellular structure, including reduced density of the endoplasmic granular mass and cytoplasmic swelling, were evident (Figure 5f). In addition, visualization of autophagosomes by staining with LC3B antibodies supported autophagy activation in the striatum of TIF-IAD1RCre mutants (Supplementary Figures 7B and C). Thus, mTOR signaling and subsequent activation of autophagy appears to be a protective mechanism that promotes the survival of MSNs upon nucleolar stress.
p53 and PTEN prolong neuronal survival upon nucleolar stress
To confirm that PTEN serves a neuroprotective function in TIF-IAD1RCre mice, we generated TIF-IA;PTEND1RCre mice, having both TIF-IA and Pten genes deleted in MSNs. In contrast to TIF-IAD1RCre mice, phosphorylation of S6 was not reduced in the striata of the double mutants (Figure 6a), indicating that nucleolar stress-induced PTEN acts in the downregulation of mTOR signaling pathway induced by nucleolar stress. In agreement with this, abrogation of TIF-IA in combination with p53 or PTEN prevented the decrease of autophagy-related proteins LC3-I and p62 (also known as sequestosome 1/SQSTM1 or A170),37, 38 which was observed in TIF-IAD1RCre mice (Figure 6b), indicating that proper induction of autophagy requires functional p53 and PTEN proteins. Consistent with autophagy being a mean to prolong survival, the amount of apoptotic cells in striata from 9-week-old TIF-IA;PTEND1RCre and TIF-IA;p53D1RCre mice was significantly increased, whereas in TIF-IAD1RCre, p53D1RCre and PTEND1RCre mutant mice no adverse effect on MSN survival was observed (Figure 6c). Thus, p53-dependent upregulation of PTEN may cause inactivation of mTOR with a temporary neuroprotective effect. The crucial role of PTEN in transmitting such a survival response was confirmed by immunostaining of the neuronal marker protein NeuN showing loss of striatal neurons in 9-week-old TIF-IA;PTEND1RCre compared with TIF-IAD1RCre mice (Figure 6d). These data imply a neuroprotective circuitry in MSNs that is initiated by disruption of nucleoli leading to increased levels of p53 and PTEN. PTEN in turn downregulates mTOR and induces autophagy, which may alleviate metabolic impairment caused by abrogation of ribosome biogenesis.
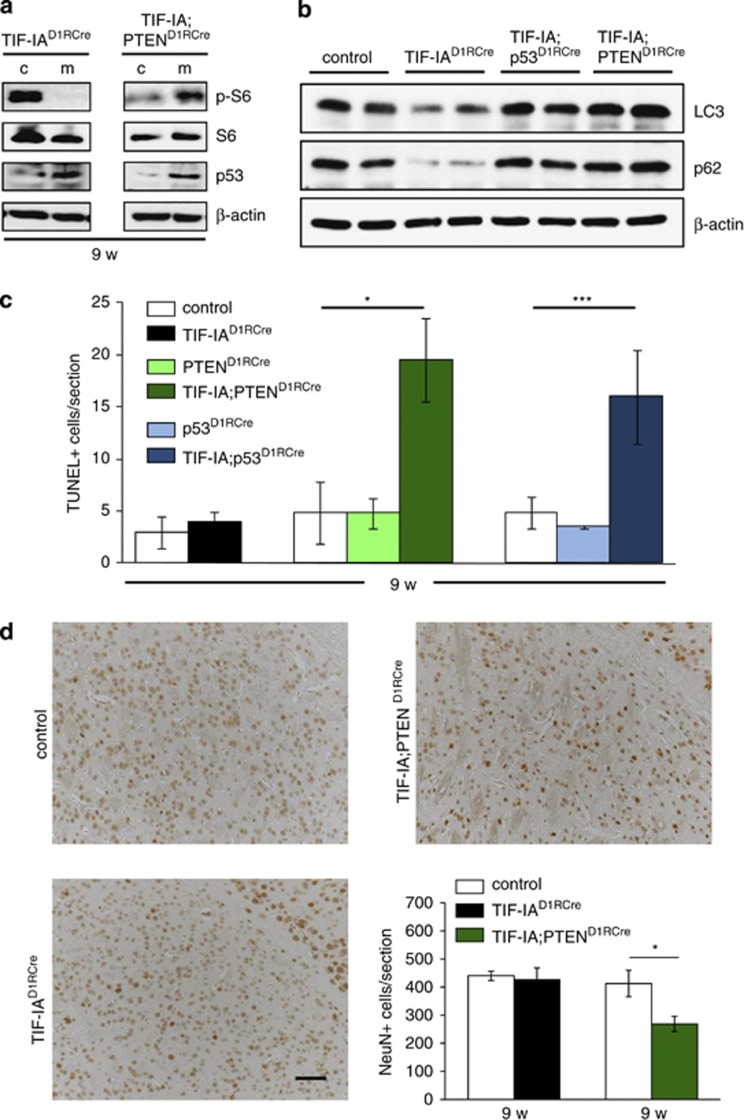
Figure 6.
p53-dependent upregulation of PTEN sustains survival of stress-exposed MSNs. (a) Representative western blots showing the levels of pS6 in striatal extracts from 9-week-old controls, TIF-IAD1RCre (left panel) or TIF-IA;PTEND1RCre (right panel) mutants. For normalization, the total levels of S6 were detected. Levels of p53 and β-actin were also monitored. (b) Representative western blots showing the levels of LC3-I, p62 and β-actin in striatal extracts from 9-week-old control, TIF-IAD1RCre, TIF-IA;p53D1RCre and TIF-IA;PTEND1RCre mice. Note that the LC3-I antibody has no reactivity toward the autophagosome-associated, phosphatidylethanolamine-conjugated LC3-II isoform. (c) Quantification of TUNEL-positive cells monitoring death of MSNs in response to TIF-IA depletion in the presence or absence of p53 or PTEN. The diagram depicts number of TUNEL-positive cells per section (mm2). Bars represent mean values±S.E.M. for TIF-IAD1RCre (n=4), PTEND1RCre (n=4) and p53D1RCre (n=5) single mutants, TIF-IA; PTEND1RCre (n=4) and TIF-IA; p53D1RCre (n=6) double mutants and for the respective control littermates (n=4–6). Striata were analyzed from all animals at 9 weeks. (d) Analysis of NeuN-positive cells by immunohistochemistry in the striatum of 9-week-old control, TIF-IAD1RCre and TIF-IA;PTEND1RCre mice. Scale bar: 60 μm. The diagram on the lower right panel shows the quantification of NeuN-positive cells per section (mm2). Values represent means±S.E.M. (n=4)
Discussion
Downregulation of rRNA synthesis is a major event to preserve cellular homeostasis in response to changes in environmental conditions.12 An increasing number of reports highlight the role of dysregulated nucleolar activity in neurodegenerative diseases.3, 4, 5, 6, 7, 16, 39, 40 Here we provide in vivo evidence that nucleolar stress leads to an initial neuroprotective phase dependent on upregulation of PTEN and p53, downregulation of mTOR and induction of autophagy. The initial survival phase of MSNs is followed by a progressive loss of striatal neurons and HD-like symptoms (Supplementary Figure 7D).
The p53-PTEN axis is known to be relevant for tumor suppression keeping PI3K/AKT/mTOR signaling in check to avoid uncontrolled cell proliferation.34 PTEN antagonizes tumor progression, latency and invasiveness, thus functioning as a pro-apoptotic factor.41 In support of PTEN protecting neural cells from stress, loss of PTEN in dopaminergic neurons appears to suppress to a certain extent behavioral abnormalities.42, 43 Moreover, silencing of the PTEN gene prevents HIV-1 gp120(IIIB)-induced degeneration of striatal neurons.44 In contrast, PTEN-dependent downregulation of mTOR signaling and subsequent activation of autophagy might be pro-survival responses that enable MSNs to survive for a certain time in stress conditions. A recent study shows that autophagy levels are reduced in the forebrain-specific conditional Pten knock-out mice.45 Accordingly, autophagy mediated by inhibition of mTOR has neuroprotective effects, indicating that autophagy is a cellular surveillance mechanism that counteracts metabolic impairment.46, 47, 48 Interestingly, autophagy is induced in solid tumor cells by inhibition of Pol I-dependent transcription.49 Apparently, the proper balance between the pro-apoptotic or neuroprotective role of PTEN depends on the type of neurons and on specific stress signals.
Consistent with the intrinsic self-defense mechanism and the HD-like phenotype in TIF-IA-deficient mice, aggregation of mutated Htt in fly and mouse models of HD has been shown to reduce neurotoxicity by inhibiting mTOR and enhancing autophagy.46 The molecular mechanism underlying neuronal degeneration in TIF-IAD1RCre mice is obviously different from those caused by mutant Htt. However, the gene expression profiles of 13-week-old TIF-IAD1RCre mice and HD patients display a striking similarity, supporting the involvement of nucleolar stress in HD. Mutant Htt led to inhibition of rRNA synthesis and nucleolar disruption at a stage at which R6/2 mice did not show motor abnormalities, significant loss of body weight or neuronal loss (Supplementary Figures 1A, B and D). Similarly, changes in striatal volume measured by NMR are not significant before 16 weeks.21 Interestingly, R6/2 mice show higher resistance to various brain insults, including ischemia and excitotoxicity50, 51 and showed dopamine-induced autophagy in striatal neurons.52 A recent report has demonstrated that mutation of Htt induces DNA damage, thereby activating p53 long before neurodegeneration is obvious, implying that striatal cells can counteract stress for long periods.53 It has been recently shown that loss of TIF-IA induces acetylation of p53, which in turn leads to stabilization and increased levels of p53.28 In accord with these data, we found increased levels of p53 acetylation in TIF-IAD1RCre mice, especially at later stages. This suggests that post-translational modifications of p53 may have a role in neuronal survival upon nucleolar stress and that by increased acetylation p53 may switch from a neuroprotective to a pro-apoptotic function. Apparently, p53 mediates an initial pro-survival response lasting until the cellular resources are used up and cells inevitably die.
Taken together, these results support the development of therapeutical strategies to control nucleolar activity at an appropriate time window, thus sustaining survival of neurons in HD and possibly also in other neurological disorders.
Materials and Methods
Mice and behavioral tests
Homozygous TIF-IAflox/flox; D1RCre (TIF-IAD1RCre) mutant mice were generated by crossing mice carrying the TIF-IA floxed allele (TIF-IAflox/flox)11 to the transgenic line D1RCre.24 TIF-IA+/flox; D1RCre-positive mice were again crossed with TIF-IAflox/flox mice. A similar strategy was used to generate TIF-IAflox/flox; PTENflox/flox; D1RCre (TIF-IA; PTEND1RCre) and TIF-IAflox/flox; p53flox/flox; D1RCre (TIF-IA; p53D1RCre) mice. Heterozygous R6/2 mice (originally from Dr. Jia-Yi Li, Lund University) were periodically analyzed for CAG triplet content (Laragen, Culver City, CA, USA) and mice with >180 repeats were excluded from breeding. To test motor performance of mice, hind limb clasping episodes after suspension by the tail were recorded and counted within 5-s intervals. Coordination tests were done on a TSE rotarod system (model 7650) and the time spent on the accelerating rod (4 to 40 r.p.m. in 60 s) was measured.5 Procedures involving animal care were approved by the Committee on Animal Care and Use (Regierungspräsidium Karlsruhe) in accordance with the local Animal Welfare Act and the European Communities Council Directive of 24 November 1986 (86/609/EEC).
Mice were killed with CO2 and striata were dissected from removed brains. For immunohistochemistry, striata were fixed in 4% paraformaldehyde overnight and paraffin embedded or sectioned on a vibratome. Sections were incubated with primary antibodies overnight at 4 °C. Visualization of antigen-bound primary antibodies was carried out using either a biotinylated secondary antibody together with the avidin–biotin system and the VECTOR peroxidase kit (Vector Laboratories, Burlingame, CA, USA), or an anti-rabbit Alexa-488 coupled secondary antibody (Molecular Probes, Eugene, OR, USA). For western blotting, striata were lysed in modified RIPA buffer: 20 mM Tris-HCl, pH 7.5, 200 mM NaCl, 1% Triton X-100, 1 mM EDTA, 1 mM EGTA, 50 mM NaF, 10 mM β-glycerolphosphate, 10 mM K2HPO4, 1 mM sodium-vanadate, 1 × protease-inhibitor-mix (Roche, Applied Science, Mannheim, Germany) and homogenized by passing through a 23-gauge needle. For detection of autophagy-associated proteins striata were lysed and homogenized in buffer containing 20 mM HEPES [pH 7.5], 10 mM KCl, 1.5 mM MgCl2, 1 mM EDTA, 1 mM EGTA, 1 mM DTT, 0.5% CHAPS and protease inhibitor cocktail (Roche).54 Lysates were cleared by centrifugation, resolved on SDS–PAGE gels, transferred to nitrocellulose membranes and probed with primary antibodies. Blots were developed by incubation with horseradish peroxidase-conjugated secondary antibodies followed by ECL plus reagent (Amersham, GE Healthcare, Freiburg, Germany). Chemiluminescence was digitally recorded using a CCD camera (LAS 3000, Fujifilm, Tokyo, Japan). Quantification of the western blots was performed by the MultiGauge Version 3.0 software (Fujifilm). Primary antibodies for immunostaining and western blotting were: anti-nucleophosmin (NPM/B23), anti-nitrosylated tyrosine, anti-NKs, anti-8-hydroxydeoxyguanosine anti-GFAP (Millipore, Billerica, MA, USA), anti-S6, anti-phosphoS6, anti-4EBP1, anti-phospho4EBP1, anti-GAPDH (Cell Signaling, Danvers, MA, USA), anti-D1R, anti-β-actin anti-p62 (Sigma, St. Louis, MO, USA), anti-p53 (Novocastra, /Leica, Wetzlar, Germany), anti-CD11b (Serotec, Oxfordshire, UK), anti-LC3B for immunofluorescence (Abcam, Cambridge, UK), anti-LC3 for western blot (Sigma), anti-acetyl-p53(Lys373, Lys382) for IHC (Millipore).55 Anti-TIF-IA antibodies were a kind gift of Brian McStay and anti-Cre and anti-Pol I (RPA116) antibodies have been described elsewhere.11
In situ hybridization
Non-radioactive in situ hybridization was performed on paraffin sections as previously described,14 using specific riboprobes hybridizing to regions in the mature 28S rRNA or in the 5′ external transcribed spacer of the pre-rRNA.
Analysis of cell death
Cell death-induced fragmentation of genomic DNA was detected by in situ staining of DNA ends with terminal TUNEL, according to the manufacturer's instructions (Roche).
Electron microscopy
For electron microscopy, animals were perfused intracardially by heparinized saline solution followed by incubation in 2% glutaraldehyde/1% paraformaldehyde in 0.1 M sodium phosphate buffer pH 7.4. Brains were post-fixed for 2 h, then coronal vibratome sections (250 μm) were cut, fixed in osmium-ferrocyanide for 1 h, stained en bloc with 1% uranyl acetate, dehydrated and embedded in araldite.56 Ultrathin sections (0.07 μm) were examined with a Philips CM10 transmission electron microscope Hillsboro, OR, USA.
Gene expression profiling
Gene expression analysis was performed in male TIF-IAD1RCre mutant mice and controls at age of 9 and 13 weeks. Total RNA was isolated from dissected striatum and nucleus accumbens using the RNeasy Mini Kit (Qiagen, Hilden, Germany). RNA integrity from all samples was controlled using LabChips and Bioanalyzer 2100 (Agilent, Santa Clara, CA, USA). RNA was reverse transcribed and cDNA representing a single animal was hybridized to a GeneChip Mouse Genome 430A 2.0 array (Affymetrix, Santa Clara, CA, USA), for a total of 12 arrays. Analysis of array data was performed as described before.57 Briefly, raw data were normalized and expression values were computed using affy and gcrma and limma packages from R/Bioconductor.58 Statistical analysis was performed using a linear model (with limma from Bioconductor) and the Benjamini and Hochberg method to assess false discovery rate.59 Ontology analyses were carried out using GSEA. MultiExperiment Viewer (MeV ver. 4.0; Boston, MA, USA) was used for identification of patterns of gene expression and data visualization (heat-map). The data are stored in the GEO database (http://www.ncbi.nlm.nih.gov/geo/), record number GSE29647.
RT-qPCR
Reverse transcription was performed using the Oligo d(T)16 primers and MultiScribe Reverse Transcriptase (Applied Biosystems, Foster City, CA, USA) following the manufacturer's instructions. TaqMan quantitative PCR (real-time PCR) (qPCR) was performed using the Chromo4 Platform (Bio-Rad, Hercules, CA, USA). The following TaqMan inventoried gene expression assays were used: CamK4 (Mm01135329_m1), Cnr1 (Mm01212171_s1), Eif2c2 (Ago2) (Mm00838341_m1), Gprk6 (Mm00442425_m1), Hip1 (Mm00524503_m1), Homer1 (Mm00516275_m1), Hprt (Mm00446968_m1), Pde10a (Mm00449329_m1), Pten (Mm00477210_m1), Smpd3 (Mm00491359_m1), Ryr1 (Mm01175211_m1) and Vim (Mm00449201_m1) (Applied Biosystems/Life Technologies, Carlsbad, CA, USA). Hypoxanthine-phophoribosyltransferase (Hprt) was chosen as a housekeeping gene. Expression changes were calculated as a fold change versus mean of control samples.
Statistical analysis
Statistical analysis was performed with GraphPad Prism 4.0 software, La Jolla, CA, USA. Two-tailed P-values using Student's t-test or one-way analysis of variance followed by the Fisher's LSD (least significant difference) test for significance assessment. Quantification of NPM/B23 in R6/2 mice was performed scoring in average 100 cells per sections (microscopic field of 1 mm2; eight sections per animal). Percentage of phosphorylated RPS6 neurons co-stained for NeuN was determined from 5 to 8 serial sections of each animal (n=3, control; n=4, mutant). Quantification of the number of NeuN+ cells was done in a microscopic field of 1 mm2 from 4 to 8 sections per each genotype (n=4). Quantification of autophagosomes was carried on using ImageJ (NIH, Bethesda, MA, USA) after thresholding.60 Confocal images were acquired at 63 × and on average eight cells per microscopic field were used for the quantification. The relatively low number of cells was due to reduced LC3B staining in the mutants (confirmed by the western blot in Figure 6b); three sections per sample were counted (n=4, ctrl; n=5, mut). The ImageJ function ‘Analyze particles' was used to define the outlines of the puncta. The size field was set to include the puncta area by manual review of the outlines and given our staining conditions it was fixed at 20 pixel.2 Mean values and S.E.M. are displayed in the figures. P-values are expressed in comparison with respective control. The results of significance test are reported as follows: (*P<0.05; **P<0.01; ***P<0.001).
Acknowledgments
We thank Professor Robert Burke for critical reading of the manuscript and Drs. Antonio Piras and Andrii Domanskyi for discussion and advices. We thank Dr. Oliver Kretz for the Affymetrix platform and Dr. Witold Konopka for help with mouse perfusion for electron microscopy. The work was supported by the German Federal Ministry for Education and Research (BMBF) through NGFNplus grant FZK 01GS08142, the DKFZ Intramural Funding Program to RP, the CIBERNED Intramural Funding to JRN, the Alfried Krupp Prize to BL, Polish National Science Center grant NCN 2011/03/13/N27/05949 to GK. The project based on this report was also funded by the program for medical genome research with financial support from BMBF under the support code 01GS08141. The author is responsible for the content of this publication.
Glossary
- D1R
D1 receptor
- GFAP
glial fibrillary acidic protein
- HD
Huntington's disease
- Hprt
hypoxanthine-phophoribosyltransferase
- MSNs
medium spiny neurons
- mTOR
mammalian target of rapamycin
- NK
neuroketal
- NPM
nucleophosmin
- PI3K
phosphatidylinositol-3-kinase
- Pol I
polymerase I
- PTEN
phosphatase and tensin homolog deleted on chromosome 10
- qPCR
quantitative PCR (real-time PCR)
- TIF-IA
transcription initiation factor IA
- TUNEL
terminal deoxynucleotidyl transferase dUTP nick end labeling
The authors declare no conflict of interest.
Footnotes
Supplementary Information accompanies this paper on Cell Death and Differentiation website (http://www.nature.com/cdd)
Edited by L Greene
Supplementary Material
References
- Mattson MP, Magnus T. Ageing and neuronal vulnerability. Nat Rev Neurosci. 2006;7:278–294. doi: 10.1038/nrn1886. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson FB, Marciniak RA, Guarente L. Telomeres, the nucleolus and aging. Curr Opin Cell Biol. 1998;10:332–338. doi: 10.1016/s0955-0674(98)80008-2. [DOI] [PubMed] [Google Scholar]
- Hetman M, Pietrzak M. Emerging roles of the neuronal nucleolus. Trends Neurosci. 2012;35:305–314. doi: 10.1016/j.tins.2012.01.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pietrzak M, Rempala G, Nelson PT, Zheng JJ, Hetman M. Epigenetic silencing of nucleolar rRNA genes in Alzheimer's disease. PLoS One. 2011;6:e22585. doi: 10.1371/journal.pone.0022585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rieker C, Engblom D, Kreiner G, Domanskyi A, Schober A, Stotz S, et al. Nucleolar disruption in dopaminergic neurons leads to oxidative damage and parkinsonism through repression of mammalian target of rapamycin signaling. J Neurosci. 2011;31:453–460. doi: 10.1523/JNEUROSCI.0590-10.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vilotti S, Codrich M, Dal Ferro M, Pinto M, Ferrer I, Collavin L, et al. Parkinson's disease DJ-1 L166P alters rRNA biogenesis by exclusion of TTRAP from the nucleolus and sequestration into cytoplasmic aggregates via TRAF6. PLoS One. 2012;7:e35051. doi: 10.1371/journal.pone.0035051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tsoi H, Lau TC, Tsang SY, Lau KF, Chan HY. CAG expansion induces nucleolar stress in polyglutamine diseases. Proc Natl Acad Sci USA. 2012;109:13428–13433. doi: 10.1073/pnas.1204089109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lanni C, Racchi M, Memo M, Govoni S, Uberti D. p53 at the crossroads between cancer and neurodegeneration. Free Radic Biol Med. 2012;52:1727–1733. doi: 10.1016/j.freeradbiomed.2012.02.034. [DOI] [PubMed] [Google Scholar]
- Vousden KH, Prives C. Blinded by the light: the growing complexity of p53. Cell. 2009;137:413–431. doi: 10.1016/j.cell.2009.04.037. [DOI] [PubMed] [Google Scholar]
- Mayer C, Grummt I. Cellular stress and nucleolar function. Cell Cycle. 2005;4:1036–1038. doi: 10.4161/cc.4.8.1925. [DOI] [PubMed] [Google Scholar]
- Yuan X, Zhou Y, Casanova E, Chai M, Kiss E, Grone HJ, et al. Genetic inactivation of the transcription factor TIF-IA leads to nucleolar disruption, cell cycle arrest, and p53-mediated apoptosis. Mol Cell. 2005;19:77–87. doi: 10.1016/j.molcel.2005.05.023. [DOI] [PubMed] [Google Scholar]
- Boulon S, Westman BJ, Hutten S, Boisvert FM, Lamond AI. The nucleolus under stress. Mol Cell. 2010;40:216–227. doi: 10.1016/j.molcel.2010.09.024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deisenroth C, Zhang Y. Ribosome biogenesis surveillance: probing the ribosomal protein-Mdm2-p53 pathway. Oncogene. 2010;29:4253–4260. doi: 10.1038/onc.2010.189. [DOI] [PubMed] [Google Scholar]
- Parlato R, Kreiner G, Erdmann G, Rieker C, Stotz S, Savenkova E, et al. Activation of an endogenous suicide response after perturbation of rRNA synthesis leads to neurodegeneration in mice. J Neurosci. 2008;28:12759–12764. doi: 10.1523/JNEUROSCI.2439-08.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hilditch-Maguire P, Trettel F, Passani LA, Auerbach A, Persichetti F, MacDonald ME. Huntingtin: an iron-regulated protein essential for normal nuclear and perinuclear organelles. Hum Mol Genet. 2000;9:2789–2797. doi: 10.1093/hmg/9.19.2789. [DOI] [PubMed] [Google Scholar]
- Latonen L. Nucleolar aggresomes as counterparts of cytoplasmic aggresomes in proteotoxic stress. Proteasome inhibitors induce nuclear ribonucleoprotein inclusions that accumulate several key factors of neurodegenerative diseases and cancer. Bioessays. 2011;33:386–395. doi: 10.1002/bies.201100008. [DOI] [PubMed] [Google Scholar]
- Carnemolla A, Fossale E, Agostoni E, Michelazzi S, Calligaris R, De Maso L, et al. Rrs1 is involved in endoplasmic reticulum stress response in Huntington disease. J Biol Chem. 2009;284:18167–18173. doi: 10.1074/jbc.M109.018325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee J, Hwang YJ, Boo JH, Han D, Kwon OK, Todorova K, et al. Dysregulation of upstream binding factor-1 acetylation at K352 is linked to impaired ribosomal DNA transcription in Huntington's disease. Cell Death Differ. 2011;18:1726–1735. doi: 10.1038/cdd.2011.38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gomes C, Smith SC, Youssef MN, Zheng JJ, Hagg T, Hetman M. RNA polymerase 1-driven transcription as a mediator of BDNF-induced neurite outgrowth. J Biol Chem. 2011;286:4357–4363. doi: 10.1074/jbc.M110.170134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zuccato C, Ciammola A, Rigamonti D, Leavitt BR, Goffredo D, Conti L, et al. Loss of huntingtin-mediated BDNF gene transcription in Huntington's disease. Science. 2001;293:493–498. doi: 10.1126/science.1059581. [DOI] [PubMed] [Google Scholar]
- Ferrante RJ. Mouse models of Huntington's disease and methodological considerations for therapeutic trials. Biochim Biophys Acta. 2009;1792:506–520. doi: 10.1016/j.bbadis.2009.04.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garden GA, Canady KS, Lurie DI, Bothwell M, Rubel EW. A biphasic change in ribosomal conformation during transneuronal degeneration is altered by inhibition of mitochondrial, but not cytoplasmic protein synthesis. J Neurosci. 1994;14:1994–2008. doi: 10.1523/JNEUROSCI.14-04-01994.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dixon J, Jones NC, Sandell LL, Jayasinghe SM, Crane J, Rey JP, et al. Tcof1/Treacle is required for neural crest cell formation and proliferation deficiencies that cause craniofacial abnormalities. Proc Natl Acad Sci USA. 2006;103:13403–13408. doi: 10.1073/pnas.0603730103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lemberger T, Parlato R, Dassesse D, Westphal M, Casanova E, Turiault M, et al. Expression of Cre recombinase in dopaminoceptive neurons. BMC Neurosci. 2007;8:4. doi: 10.1186/1471-2202-8-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strand AD, Baquet ZC, Aragaki AK, Holmans P, Yang L, Cleren C, et al. Expression profiling of Huntington's disease models suggests that brain-derived neurotrophic factor depletion plays a major role in striatal degeneration. J Neurosci. 2007;27:11758–11768. doi: 10.1523/JNEUROSCI.2461-07.2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ju TC, Chen HM, Lin JT, Chang CP, Chang WC, Kang JJ, et al. Nuclear translocation of AMPK-alpha1 potentiates striatal neurodegeneration in Huntington's disease. J Cell Biol. 2011;194:209–227. doi: 10.1083/jcb.201105010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rubbi CP, Milner J. Disruption of the nucleolus mediates stabilization of p53 in response to DNA damage and other stresses. EMBO J. 2003;22:6068–6077. doi: 10.1093/emboj/cdg579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuroda T, Murayama A, Katagiri N, Ohta YM, Fujita E, Masumoto H, et al. RNA content in the nucleolus alters p53 acetylation via MYBBP1A. EMBO J. 2011;30:1054–1066. doi: 10.1038/emboj.2011.23. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tang Y, Zhao W, Chen Y, Zhao Y, Gu W. Acetylation is indispensable for p53 activation. Cell. 2008;133:612–626. doi: 10.1016/j.cell.2008.03.025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deckel AW, Elder R, Fuhrer G. Biphasic developmental changes in Ca2+/calmodulin-dependent proteins in R6/2 Huntington's disease mice. Neuroreport. 2002;13:707–711. doi: 10.1097/00001756-200204160-00034. [DOI] [PubMed] [Google Scholar]
- Blazquez C, Chiarlone A, Sagredo O, Aguado T, Pazos MR, Resel E, et al. Loss of striatal type 1 cannabinoid receptors is a key pathogenic factor in Huntington's disease. Brain. 2011;134 (Pt 1:119–136. doi: 10.1093/brain/awq278. [DOI] [PubMed] [Google Scholar]
- Hebb AL, Robertson HA, Denovan-Wright EM. Striatal phosphodiesterase mRNA and protein levels are reduced in Huntington's disease transgenic mice prior to the onset of motor symptoms. Neuroscience. 2004;123:967–981. doi: 10.1016/j.neuroscience.2003.11.009. [DOI] [PubMed] [Google Scholar]
- Luthi-Carter R, Hanson SA, Strand AD, Bergstrom DA, Chun W, Peters NL, et al. Dysregulation of gene expression in the R6/2 model of polyglutamine disease: parallel changes in muscle and brain. Hum Mol Genet. 2002;11:1911–1926. doi: 10.1093/hmg/11.17.1911. [DOI] [PubMed] [Google Scholar]
- Stambolic V, MacPherson D, Sas D, Lin Y, Snow B, Jang Y, et al. Regulation of PTEN transcription by p53. Mol Cell. 2001;8:317–325. doi: 10.1016/s1097-2765(01)00323-9. [DOI] [PubMed] [Google Scholar]
- Song MS, Salmena L, Pandolfi PP. The functions and regulation of the PTEN tumour suppressor. Nat Rev Mol Cell Biol. 2012;13:283–296. doi: 10.1038/nrm3330. [DOI] [PubMed] [Google Scholar]
- Mizushima N, Levine B, Cuervo AM, Klionsky DJ. Autophagy fights disease through cellular self-digestion. Nature. 2008;451:1069–1075. doi: 10.1038/nature06639. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mizushima N, Yoshimori T. How to interpret LC3 immunoblotting. Autophagy. 2007;3:542–545. doi: 10.4161/auto.4600. [DOI] [PubMed] [Google Scholar]
- Johansen T, Lamark T. Selective autophagy mediated by autophagic adapter proteins. Autophagy. 2011;7:279–296. doi: 10.4161/auto.7.3.14487. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Caudle WM, Kitsou E, Li J, Bradner J, Zhang J. A role for a novel protein, nucleolin, in Parkinson's disease. Neurosci Lett. 2009;459:11–15. doi: 10.1016/j.neulet.2009.04.060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baltanas FC, Casafont I, Weruaga E, Alonso JR, Berciano MT, Lafarga M. Nucleolar disruption and cajal body disassembly are nuclear hallmarks of DNA damage-induced neurodegeneration in purkinje cells. Brain Pathol. 2011;21:374–388. doi: 10.1111/j.1750-3639.2010.00461.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alimonti A, Carracedo A, Clohessy JG, Trotman LC, Nardella C, Egia A, et al. Subtle variations in Pten dose determine cancer susceptibility. Nat Genet. 2010;42:454–458. doi: 10.1038/ng.556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diaz-Ruiz O, Zapata A, Shan L, Zhang Y, Tomac AC, Malik N, et al. Selective deletion of PTEN in dopamine neurons leads to trophic effects and adaptation of striatal medium spiny projecting neurons. PLoS One. 2009;4:e7027. doi: 10.1371/journal.pone.0007027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Domanskyi A, Geissler C, Vinnikov IA, Alter H, Schober A, Vogt MA, et al. Pten ablation in adult dopaminergic neurons is neuroprotective in Parkinson's disease models. FASEB J. 2011;25:2898–2910. doi: 10.1096/fj.11-181958. [DOI] [PubMed] [Google Scholar]
- Zou S, El-Hage N, Podhaizer EM, Knapp PE, Hauser KF. PTEN gene silencing prevents HIV-1 gp120(IIIB)-induced degeneration of striatal neurons. J Neurovirol. 2011;17:41–49. doi: 10.1007/s13365-010-0016-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McMahon J, Huang X, Yang J, Komatsu M, Yue Z, Qian J, et al. Impaired autophagy in neurons after disinhibition of mammalian target of rapamycin and its contribution to epileptogenesis. J Neurosci. 2012;32:15704–15714. doi: 10.1523/JNEUROSCI.2392-12.2012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ravikumar B, Vacher C, Berger Z, Davies JE, Luo S, Oroz LG, et al. Inhibition of mTOR induces autophagy and reduces toxicity of polyglutamine expansions in fly and mouse models of Huntington disease. Nat Genet. 2004;36:585–595. doi: 10.1038/ng1362. [DOI] [PubMed] [Google Scholar]
- Komatsu M, Waguri S, Chiba T, Murata S, Iwata J, Tanida I, et al. Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature. 2006;441:880–884. doi: 10.1038/nature04723. [DOI] [PubMed] [Google Scholar]
- Hara T, Nakamura K, Matsui M, Yamamoto A, Nakahara Y, Suzuki-Migishima R, et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature. 2006;441:885–889. doi: 10.1038/nature04724. [DOI] [PubMed] [Google Scholar]
- Drygin D, Lin A, Bliesath J, Ho CB, O'Brien SE, Proffitt C, et al. Targeting RNA polymerase I with an oral small molecule CX-5461 inhibits ribosomal RNA synthesis and solid tumor growth. Cancer Res. 2011;71:1418–1430. doi: 10.1158/0008-5472.CAN-10-1728. [DOI] [PubMed] [Google Scholar]
- Klapstein GJ, Levine MS. Age-dependent biphasic changes in ischemic sensitivity in the striatum of Huntington's disease R6/2 transgenic mice. J Neurophysiol. 2005;93:758–765. doi: 10.1152/jn.00483.2004. [DOI] [PubMed] [Google Scholar]
- Hansson O, Guatteo E, Mercuri NB, Bernardi G, Li XJ, Castilho RF, et al. Resistance to NMDA toxicity correlates with appearance of nuclear inclusions, behavioural deficits and changes in calcium homeostasis in mice transgenic for exon 1 of the huntington gene. Eur J Neurosci. 2001;14:1492–1504. doi: 10.1046/j.0953-816x.2001.01767.x. [DOI] [PubMed] [Google Scholar]
- Petersen A, Larsen KE, Behr GG, Romero N, Przedborski S, Brundin P, et al. Expanded CAG repeats in exon 1 of the Huntington's disease gene stimulate dopamine-mediated striatal neuron autophagy and degeneration. Hum Mol Genet. 2001;10:1243–1254. doi: 10.1093/hmg/10.12.1243. [DOI] [PubMed] [Google Scholar]
- Grison A, Mantovani F, Comel A, Agostoni E, Gustincich S, Persichetti F, et al. Ser46 phosphorylation and prolyl-isomerase Pin1-mediated isomerization of p53 are key events in p53-dependent apoptosis induced by mutant huntingtin. Proc Natl Acad Sci USA. 2011;108:17979–17984. doi: 10.1073/pnas.1106198108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piras A, Gianetto D, Conte D, Bosone A, Vercelli A. Activation of autophagy in a rat model of retinal ischemia following high intraocular pressure. PLoS One. 2011;6:e22514. doi: 10.1371/journal.pone.0022514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li X, Khanna A, Li N, Wang E. Circulatory miR34a as an RNAbased, noninvasive biomarker for brain aging. Aging (Albany NY) 2011;3:985–1002. doi: 10.18632/aging.100371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ponti G, Aimar P, Bonfanti L. Cellular composition and cytoarchitecture of the rabbit subventricular zone and its extensions in the forebrain. J Comp Neurol. 2006;498:491–507. doi: 10.1002/cne.21043. [DOI] [PubMed] [Google Scholar]
- Bilbao A, Parkitna JR, Engblom D, Perreau-Lenz S, Sanchis-Segura C, Schneider M, et al. Loss of the Ca2+/calmodulin-dependent protein kinase type IV in dopaminoceptive neurons enhances behavioral effects of cocaine. Proc Natl Acad Sci USA. 2008;105:17549–17554. doi: 10.1073/pnas.0803959105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 2004;5:R80. doi: 10.1186/gb-2004-5-10-r80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benjamini Y, Hochberg Y. Controlling the false discovery rate - a practical and powerful approach to multiple testing. J Roy Stat Soc Ser B Methodol. 1995;57:289–300. [Google Scholar]
- Koga H, Martinez-Vicente M, Arias E, Kaushik S, Sulzer D, Cuervo AM. Constitutive upregulation of chaperone-mediated autophagy in Huntington's disease. J Neurosci. 2011;31:18492–18505. doi: 10.1523/JNEUROSCI.3219-11.2011. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.