Abstract
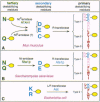
The N-end rule relates the in vivo half-life of a protein to the identity of its N-terminal residue. Similar but distinct versions of the N-end rule operate in all organisms examined, from mammals to fungi and bacteria. In eukaryotes, the N-end rule pathway is a part of the ubiquitin system. I discuss the mechanisms and functions of this pathway, and consider its applications.
Full text
PDF







Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aizenman E., Engelberg-Kulka H., Glaser G. An Escherichia coli chromosomal "addiction module" regulated by guanosine [corrected] 3',5'-bispyrophosphate: a model for programmed bacterial cell death. Proc Natl Acad Sci U S A. 1996 Jun 11;93(12):6059–6063. doi: 10.1073/pnas.93.12.6059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alagramam K., Naider F., Becker J. M. A recognition component of the ubiquitin system is required for peptide transport in Saccharomyces cerevisiae. Mol Microbiol. 1995 Jan;15(2):225–234. doi: 10.1111/j.1365-2958.1995.tb02237.x. [DOI] [PubMed] [Google Scholar]
- Arfin S. M., Bradshaw R. A. Cotranslational processing and protein turnover in eukaryotic cells. Biochemistry. 1988 Oct 18;27(21):7979–7984. doi: 10.1021/bi00421a001. [DOI] [PubMed] [Google Scholar]
- Arnason T., Ellison M. J. Stress resistance in Saccharomyces cerevisiae is strongly correlated with assembly of a novel type of multiubiquitin chain. Mol Cell Biol. 1994 Dec;14(12):7876–7883. doi: 10.1128/mcb.14.12.7876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baboshina O. V., Haas A. L. Novel multiubiquitin chain linkages catalyzed by the conjugating enzymes E2EPF and RAD6 are recognized by 26 S proteasome subunit 5. J Biol Chem. 1996 Feb 2;271(5):2823–2831. doi: 10.1074/jbc.271.5.2823. [DOI] [PubMed] [Google Scholar]
- Bachmair A., Finley D., Varshavsky A. In vivo half-life of a protein is a function of its amino-terminal residue. Science. 1986 Oct 10;234(4773):179–186. doi: 10.1126/science.3018930. [DOI] [PubMed] [Google Scholar]
- Bachmair A., Varshavsky A. The degradation signal in a short-lived protein. Cell. 1989 Mar 24;56(6):1019–1032. doi: 10.1016/0092-8674(89)90635-1. [DOI] [PubMed] [Google Scholar]
- Baker R. T., Varshavsky A. Inhibition of the N-end rule pathway in living cells. Proc Natl Acad Sci U S A. 1991 Feb 15;88(4):1090–1094. doi: 10.1073/pnas.88.4.1090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baker R. T., Varshavsky A. Yeast N-terminal amidase. A new enzyme and component of the N-end rule pathway. J Biol Chem. 1995 May 19;270(20):12065–12074. doi: 10.1074/jbc.270.20.12065. [DOI] [PubMed] [Google Scholar]
- Balzi E., Choder M., Chen W. N., Varshavsky A., Goffeau A. Cloning and functional analysis of the arginyl-tRNA-protein transferase gene ATE1 of Saccharomyces cerevisiae. J Biol Chem. 1990 May 5;265(13):7464–7471. [PubMed] [Google Scholar]
- Bartel B., Wünning I., Varshavsky A. The recognition component of the N-end rule pathway. EMBO J. 1990 Oct;9(10):3179–3189. doi: 10.1002/j.1460-2075.1990.tb07516.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chau V., Tobias J. W., Bachmair A., Marriott D., Ecker D. J., Gonda D. K., Varshavsky A. A multiubiquitin chain is confined to specific lysine in a targeted short-lived protein. Science. 1989 Mar 24;243(4898):1576–1583. doi: 10.1126/science.2538923. [DOI] [PubMed] [Google Scholar]
- Cormack B. P., Struhl K. The TATA-binding protein is required for transcription by all three nuclear RNA polymerases in yeast cells. Cell. 1992 May 15;69(4):685–696. doi: 10.1016/0092-8674(92)90232-2. [DOI] [PubMed] [Google Scholar]
- Dayal V. K., Chakraborty G., Sturman J. A., Ingoglia N. A. The site of amino acid addition to posttranslationally modified proteins of regenerating rat sciatic nerves. Biochim Biophys Acta. 1990 Apr 19;1038(2):172–177. doi: 10.1016/0167-4838(90)90201-p. [DOI] [PubMed] [Google Scholar]
- Deveraux Q., Ustrell V., Pickart C., Rechsteiner M. A 26 S protease subunit that binds ubiquitin conjugates. J Biol Chem. 1994 Mar 11;269(10):7059–7061. [PubMed] [Google Scholar]
- Dohmen R. J., Madura K., Bartel B., Varshavsky A. The N-end rule is mediated by the UBC2(RAD6) ubiquitin-conjugating enzyme. Proc Natl Acad Sci U S A. 1991 Aug 15;88(16):7351–7355. doi: 10.1073/pnas.88.16.7351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dohmen R. J., Wu P., Varshavsky A. Heat-inducible degron: a method for constructing temperature-sensitive mutants. Science. 1994 Mar 4;263(5151):1273–1276. doi: 10.1126/science.8122109. [DOI] [PubMed] [Google Scholar]
- Dougherty W. G., Semler B. L. Expression of virus-encoded proteinases: functional and structural similarities with cellular enzymes. Microbiol Rev. 1993 Dec;57(4):781–822. doi: 10.1128/mr.57.4.781-822.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ellis R. E., Yuan J. Y., Horvitz H. R. Mechanisms and functions of cell death. Annu Rev Cell Biol. 1991;7:663–698. doi: 10.1146/annurev.cb.07.110191.003311. [DOI] [PubMed] [Google Scholar]
- Emoto Y., Manome Y., Meinhardt G., Kisaki H., Kharbanda S., Robertson M., Ghayur T., Wong W. W., Kamen R., Weichselbaum R. Proteolytic activation of protein kinase C delta by an ICE-like protease in apoptotic cells. EMBO J. 1995 Dec 15;14(24):6148–6156. doi: 10.1002/j.1460-2075.1995.tb00305.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Finley D., Bartel B., Varshavsky A. The tails of ubiquitin precursors are ribosomal proteins whose fusion to ubiquitin facilitates ribosome biogenesis. Nature. 1989 Mar 30;338(6214):394–401. doi: 10.1038/338394a0. [DOI] [PubMed] [Google Scholar]
- Gonda D. K., Bachmair A., Wünning I., Tobias J. W., Lane W. S., Varshavsky A. Universality and structure of the N-end rule. J Biol Chem. 1989 Oct 5;264(28):16700–16712. [PubMed] [Google Scholar]
- Gottesman S., Maurizi M. R. Regulation by proteolysis: energy-dependent proteases and their targets. Microbiol Rev. 1992 Dec;56(4):592–621. doi: 10.1128/mr.56.4.592-621.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hershko A. The ubiquitin pathway for protein degradation. Trends Biochem Sci. 1991 Jul;16(7):265–268. doi: 10.1016/0968-0004(91)90101-z. [DOI] [PubMed] [Google Scholar]
- Hiller M. M., Finger A., Schweiger M., Wolf D. H. ER degradation of a misfolded luminal protein by the cytosolic ubiquitin-proteasome pathway. Science. 1996 Sep 20;273(5282):1725–1728. doi: 10.1126/science.273.5282.1725. [DOI] [PubMed] [Google Scholar]
- Hilt W., Wolf D. H. Proteasomes: destruction as a programme. Trends Biochem Sci. 1996 Mar;21(3):96–102. [PubMed] [Google Scholar]
- Hochstrasser M. Ubiquitin, proteasomes, and the regulation of intracellular protein degradation. Curr Opin Cell Biol. 1995 Apr;7(2):215–223. doi: 10.1016/0955-0674(95)80031-x. [DOI] [PubMed] [Google Scholar]
- Jentsch S. The ubiquitin-conjugation system. Annu Rev Genet. 1992;26:179–207. doi: 10.1146/annurev.ge.26.120192.001143. [DOI] [PubMed] [Google Scholar]
- Johnson E. S., Gonda D. K., Varshavsky A. cis-trans recognition and subunit-specific degradation of short-lived proteins. Nature. 1990 Jul 19;346(6281):287–291. doi: 10.1038/346287a0. [DOI] [PubMed] [Google Scholar]
- Johnson E. S., Ma P. C., Ota I. M., Varshavsky A. A proteolytic pathway that recognizes ubiquitin as a degradation signal. J Biol Chem. 1995 Jul 21;270(29):17442–17456. doi: 10.1074/jbc.270.29.17442. [DOI] [PubMed] [Google Scholar]
- Johnston J. A., Johnson E. S., Waller P. R., Varshavsky A. Methotrexate inhibits proteolysis of dihydrofolate reductase by the N-end rule pathway. J Biol Chem. 1995 Apr 7;270(14):8172–8178. doi: 10.1074/jbc.270.14.8172. [DOI] [PubMed] [Google Scholar]
- Kaiser C. A., Preuss D., Grisafi P., Botstein D. Many random sequences functionally replace the secretion signal sequence of yeast invertase. Science. 1987 Jan 16;235(4786):312–317. doi: 10.1126/science.3541205. [DOI] [PubMed] [Google Scholar]
- Kayalar C., Ord T., Testa M. P., Zhong L. T., Bredesen D. E. Cleavage of actin by interleukin 1 beta-converting enzyme to reverse DNase I inhibition. Proc Natl Acad Sci U S A. 1996 Mar 5;93(5):2234–2238. doi: 10.1073/pnas.93.5.2234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knight S. A., Tamai K. T., Kosman D. J., Thiele D. J. Identification and analysis of a Saccharomyces cerevisiae copper homeostasis gene encoding a homeodomain protein. Mol Cell Biol. 1994 Dec;14(12):7792–7804. doi: 10.1128/mcb.14.12.7792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knighton D. R., Kan C. C., Howland E., Janson C. A., Hostomska Z., Welsh K. M., Matthews D. A. Structure of and kinetic channelling in bifunctional dihydrofolate reductase-thymidylate synthase. Nat Struct Biol. 1994 Mar;1(3):186–194. doi: 10.1038/nsb0394-186. [DOI] [PubMed] [Google Scholar]
- Levis M. J., Bourne H. R. Activation of the alpha subunit of Gs in intact cells alters its abundance, rate of degradation, and membrane avidity. J Cell Biol. 1992 Dec;119(5):1297–1307. doi: 10.1083/jcb.119.5.1297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li X., Chang Y. H. Amino-terminal protein processing in Saccharomyces cerevisiae is an essential function that requires two distinct methionine aminopeptidases. Proc Natl Acad Sci U S A. 1995 Dec 19;92(26):12357–12361. doi: 10.1073/pnas.92.26.12357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lévy F., Johnsson N., Rümenapf T., Varshavsky A. Using ubiquitin to follow the metabolic fate of a protein. Proc Natl Acad Sci U S A. 1996 May 14;93(10):4907–4912. doi: 10.1073/pnas.93.10.4907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Madura K., Dohmen R. J., Varshavsky A. N-recognin/Ubc2 interactions in the N-end rule pathway. J Biol Chem. 1993 Jun 5;268(16):12046–12054. [PubMed] [Google Scholar]
- Madura K., Varshavsky A. Degradation of G alpha by the N-end rule pathway. Science. 1994 Sep 2;265(5177):1454–1458. doi: 10.1126/science.8073290. [DOI] [PubMed] [Google Scholar]
- Maeda T., Wurgler-Murphy S. M., Saito H. A two-component system that regulates an osmosensing MAP kinase cascade in yeast. Nature. 1994 May 19;369(6477):242–245. doi: 10.1038/369242a0. [DOI] [PubMed] [Google Scholar]
- McGrath J. P., Jentsch S., Varshavsky A. UBA 1: an essential yeast gene encoding ubiquitin-activating enzyme. EMBO J. 1991 Jan;10(1):227–236. doi: 10.1002/j.1460-2075.1991.tb07940.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Negrutskii B. S., Deutscher M. P. Channeling of aminoacyl-tRNA for protein synthesis in vivo. Proc Natl Acad Sci U S A. 1991 Jun 1;88(11):4991–4995. doi: 10.1073/pnas.88.11.4991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishizawa M., Furuno N., Okazaki K., Tanaka H., Ogawa Y., Sagata N. Degradation of Mos by the N-terminal proline (Pro2)-dependent ubiquitin pathway on fertilization of Xenopus eggs: possible significance of natural selection for Pro2 in Mos. EMBO J. 1993 Oct;12(10):4021–4027. doi: 10.1002/j.1460-2075.1993.tb06080.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishizawa M., Okazaki K., Furuno N., Watanabe N., Sagata N. The 'second-codon rule' and autophosphorylation govern the stability and activity of Mos during the meiotic cell cycle in Xenopus oocytes. EMBO J. 1992 Jul;11(7):2433–2446. doi: 10.1002/j.1460-2075.1992.tb05308.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ota I. M., Varshavsky A. A gene encoding a putative tyrosine phosphatase suppresses lethality of an N-end rule-dependent mutant. Proc Natl Acad Sci U S A. 1992 Mar 15;89(6):2355–2359. doi: 10.1073/pnas.89.6.2355. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ota I. M., Varshavsky A. A yeast protein similar to bacterial two-component regulators. Science. 1993 Oct 22;262(5133):566–569. doi: 10.1126/science.8211183. [DOI] [PubMed] [Google Scholar]
- Ozkaynak E., Finley D., Solomon M. J., Varshavsky A. The yeast ubiquitin genes: a family of natural gene fusions. EMBO J. 1987 May;6(5):1429–1439. doi: 10.1002/j.1460-2075.1987.tb02384.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pahl H. L., Baeuerle P. A. Control of gene expression by proteolysis. Curr Opin Cell Biol. 1996 Jun;8(3):340–347. doi: 10.1016/s0955-0674(96)80007-x. [DOI] [PubMed] [Google Scholar]
- Park E. C., Finley D., Szostak J. W. A strategy for the generation of conditional mutations by protein destabilization. Proc Natl Acad Sci U S A. 1992 Feb 15;89(4):1249–1252. doi: 10.1073/pnas.89.4.1249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rechsteiner M., Hoffman L., Dubiel W. The multicatalytic and 26 S proteases. J Biol Chem. 1993 Mar 25;268(9):6065–6068. [PubMed] [Google Scholar]
- Reiss Y., Kaim D., Hershko A. Specificity of binding of NH2-terminal residue of proteins to ubiquitin-protein ligase. Use of amino acid derivatives to characterize specific binding sites. J Biol Chem. 1988 Feb 25;263(6):2693–2698. [PubMed] [Google Scholar]
- Sadis S., Atienza C., Jr, Finley D. Synthetic signals for ubiquitin-dependent proteolysis. Mol Cell Biol. 1995 Aug;15(8):4086–4094. doi: 10.1128/mcb.15.8.4086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schatz G., Dobberstein B. Common principles of protein translocation across membranes. Science. 1996 Mar 15;271(5255):1519–1526. doi: 10.1126/science.271.5255.1519. [DOI] [PubMed] [Google Scholar]
- Scheffner M., Nuber U., Huibregtse J. M. Protein ubiquitination involving an E1-E2-E3 enzyme ubiquitin thioester cascade. Nature. 1995 Jan 5;373(6509):81–83. doi: 10.1038/373081a0. [DOI] [PubMed] [Google Scholar]
- Scheffner M., Werness B. A., Huibregtse J. M., Levine A. J., Howley P. M. The E6 oncoprotein encoded by human papillomavirus types 16 and 18 promotes the degradation of p53. Cell. 1990 Dec 21;63(6):1129–1136. doi: 10.1016/0092-8674(90)90409-8. [DOI] [PubMed] [Google Scholar]
- Scherer D. C., Brockman J. A., Chen Z., Maniatis T., Ballard D. W. Signal-induced degradation of I kappa B alpha requires site-specific ubiquitination. Proc Natl Acad Sci U S A. 1995 Nov 21;92(24):11259–11263. doi: 10.1073/pnas.92.24.11259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sherman F., Stewart J. W., Tsunasawa S. Methionine or not methionine at the beginning of a protein. Bioessays. 1985 Jul;3(1):27–31. doi: 10.1002/bies.950030108. [DOI] [PubMed] [Google Scholar]
- Shrader T. E., Tobias J. W., Varshavsky A. The N-end rule in Escherichia coli: cloning and analysis of the leucyl, phenylalanyl-tRNA-protein transferase gene aat. J Bacteriol. 1993 Jul;175(14):4364–4374. doi: 10.1128/jb.175.14.4364-4374.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spence J., Sadis S., Haas A. L., Finley D. A ubiquitin mutant with specific defects in DNA repair and multiubiquitination. Mol Cell Biol. 1995 Mar;15(3):1265–1273. doi: 10.1128/mcb.15.3.1265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stewart A. E., Arfin S. M., Bradshaw R. A. The sequence of porcine protein NH2-terminal asparagine amidohydrolase. A new component of the N-end Rule pathway. J Biol Chem. 1995 Jan 6;270(1):25–28. doi: 10.1074/jbc.270.1.25. [DOI] [PubMed] [Google Scholar]
- Strauss J. H., Strauss E. G. The alphaviruses: gene expression, replication, and evolution. Microbiol Rev. 1994 Sep;58(3):491–562. doi: 10.1128/mr.58.3.491-562.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thornberry N. A., Bull H. G., Calaycay J. R., Chapman K. T., Howard A. D., Kostura M. J., Miller D. K., Molineaux S. M., Weidner J. R., Aunins J. A novel heterodimeric cysteine protease is required for interleukin-1 beta processing in monocytes. Nature. 1992 Apr 30;356(6372):768–774. doi: 10.1038/356768a0. [DOI] [PubMed] [Google Scholar]
- Tobias J. W., Shrader T. E., Rocap G., Varshavsky A. The N-end rule in bacteria. Science. 1991 Nov 29;254(5036):1374–1377. doi: 10.1126/science.1962196. [DOI] [PubMed] [Google Scholar]
- Varshavsky A. Codominance and toxins: a path to drugs of nearly unlimited selectivity. Proc Natl Acad Sci U S A. 1995 Apr 25;92(9):3663–3667. doi: 10.1073/pnas.92.9.3663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Varshavsky A. Naming a targeting signal. Cell. 1991 Jan 11;64(1):13–15. doi: 10.1016/0092-8674(91)90202-a. [DOI] [PubMed] [Google Scholar]
- Varshavsky A. The N-end rule. Cold Spring Harb Symp Quant Biol. 1995;60:461–478. doi: 10.1101/sqb.1995.060.01.051. [DOI] [PubMed] [Google Scholar]
- Varshavsky A. The N-end rule. Cell. 1992 May 29;69(5):725–735. doi: 10.1016/0092-8674(92)90285-k. [DOI] [PubMed] [Google Scholar]
- Watanabe N., Hunt T., Ikawa Y., Sagata N. Independent inactivation of MPF and cytostatic factor (Mos) upon fertilization of Xenopus eggs. Nature. 1991 Jul 18;352(6332):247–248. doi: 10.1038/352247a0. [DOI] [PubMed] [Google Scholar]
- Watkins J. F., Sung P., Prakash L., Prakash S. The Saccharomyces cerevisiae DNA repair gene RAD23 encodes a nuclear protein containing a ubiquitin-like domain required for biological function. Mol Cell Biol. 1993 Dec;13(12):7757–7765. doi: 10.1128/mcb.13.12.7757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wickner S., Gottesman S., Skowyra D., Hoskins J., McKenney K., Maurizi M. R. A molecular chaperone, ClpA, functions like DnaK and DnaJ. Proc Natl Acad Sci U S A. 1994 Dec 6;91(25):12218–12222. doi: 10.1073/pnas.91.25.12218. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wiertz E. J., Jones T. R., Sun L., Bogyo M., Geuze H. J., Ploegh H. L. The human cytomegalovirus US11 gene product dislocates MHC class I heavy chains from the endoplasmic reticulum to the cytosol. Cell. 1996 Mar 8;84(5):769–779. doi: 10.1016/s0092-8674(00)81054-5. [DOI] [PubMed] [Google Scholar]
- Wolf S., Lottspeich F., Baumeister W. Ubiquitin found in the archaebacterium Thermoplasma acidophilum. FEBS Lett. 1993 Jul 12;326(1-3):42–44. doi: 10.1016/0014-5793(93)81757-q. [DOI] [PubMed] [Google Scholar]
- de Groot R. J., Rümenapf T., Kuhn R. J., Strauss E. G., Strauss J. H. Sindbis virus RNA polymerase is degraded by the N-end rule pathway. Proc Natl Acad Sci U S A. 1991 Oct 15;88(20):8967–8971. doi: 10.1073/pnas.88.20.8967. [DOI] [PMC free article] [PubMed] [Google Scholar]