Generation of antigen-loaded MR1 tetramers that specifically stain MAIT cells identifies heterogeneity in phenotypes and TCR repertoires in humans and mice.
Abstract
Mucosal-associated invariant T cells (MAIT cells) express a semi-invariant T cell receptor (TCR) α-chain, TRAV1-2–TRAJ33, and are activated by vitamin B metabolites bound by the major histocompatibility complex (MHC)–related class I–like molecule, MR1. Understanding MAIT cell biology has been restrained by the lack of reagents to specifically identify and characterize these cells. Furthermore, the use of surrogate markers may misrepresent the MAIT cell population. We show that modified human MR1 tetramers loaded with the potent MAIT cell ligand, reduced 6-hydroxymethyl-8-d-ribityllumazine (rRL-6-CH2OH), specifically detect all human MAIT cells. Tetramer+ MAIT subsets were predominantly CD8+ or CD4−CD8−, although a small subset of CD4+ MAIT cells was also detected. Notably, most human CD8+ MAIT cells were CD8α+CD8β−/lo, implying predominant expression of CD8αα homodimers. Tetramer-sorted MAIT cells displayed a TH1 cytokine phenotype upon antigen-specific activation. Similarly, mouse MR1–rRL-6-CH2OH tetramers detected CD4+, CD4−CD8− and CD8+ MAIT cells in Vα19 transgenic mice. Both human and mouse MAIT cells expressed a broad TCR-β repertoire, and although the majority of human MAIT cells expressed TRAV1-2–TRAJ33, some expressed TRAJ12 or TRAJ20 genes in conjunction with TRAV1-2. Accordingly, MR1 tetramers allow precise phenotypic characterization of human and mouse MAIT cells and revealed unanticipated TCR heterogeneity in this population.
Mucosal-associated invariant T cells (MAIT cells) are innate-like T cells, comprising up to 10% of the peripheral blood T cells in humans, and are present in high frequency in the gastrointestinal mucosa and liver (Treiner et al., 2003; Martin et al., 2009; Dusseaux et al., 2011). MAIT cells are also present in mice, although their frequencies are extremely rare in laboratory strains of mice tested to date (Tilloy et al., 1999; Treiner et al., 2003). MAIT cells may play a role in protective immunity and are implicated in several autoimmune disorders (Croxford et al., 2006; Gold et al., 2010; Le Bourhis et al., 2010, 2011, 2013; Miyazaki et al., 2011; Chiba et al., 2012; Chua et al., 2012; Cosgrove et al., 2013; Gold and Lewinsohn, 2013; Leeansyah et al., 2013; Meierovics et al., 2013). MAIT cells, when activated via the antigen (Ag)-specific αβ TCR, rapidly secrete cytokines, including IFN-γ, TNF, IL-17 in humans (Dusseaux et al., 2011) and IFN-γ, IL-4, IL-5, and IL-10 in Vα19i transgenic (Tg) mice (Kawachi et al., 2006). Consistent with their innate-like properties, MAIT cells express a very restricted T cell repertoire. Namely, in humans, MAIT cells express an invariant TCR α-chain, Vα7.2 (TRAV1-2), joined to Jα33 (TRAJ33), which is paired with a limited array of TCR β-chains (predominantly TRBV6 or TRBV20; Tilloy et al., 1999). In mice, the MAIT TCR repertoire comprises the orthologous TCR α-chain (Vα19Jα33) paired with Vβ6 or Vβ8 (TRBV19 or TRBV13). N-region additions are also found at the Vα-Jα junctions of MAIT TCRs, so the TCR α-chain is not completely invariant even though these residues are located at the base of the CDR3α loops rather than at the sites of direct Ag recognition (Reantragoon et al., 2012; Patel et al., 2013). The MAIT TCR is restricted to the ubiquitously expressed MHC class I (MHC-I)–related molecule MR1 (Treiner et al., 2003), which is only found in mammals and exhibits a very high level of sequence conservation between mice and humans, thereby underscoring the evolutionary importance of the MAIT–MR1 axis in immunity. Recently, we described a family of microbially derived vitamin B metabolites presented by MR1 that specifically activate MAIT cells and provided the molecular basis for MAIT TCR recognition of vitamin B metabolites (Kjer-Nielsen et al., 2012; Patel et al., 2013). These findings correlated with bacteria and yeast that stimulate MAIT cells possessing an intact riboflavin synthesis pathway, whereas this pathway is deficient in nonstimulatory microbes (Gold et al., 2010; Le Bourhis et al., 2010; Kjer-Nielsen et al., 2012). The definition of MR1-restricted ligands enables the function of MAIT cells to be probed in an Ag-dependent manner. However, a key to understanding MAIT cell physiology and pathology is the development of Ag-specific reagents, for example MR1-Ag tetramers, to characterize MAIT cells ex vivo. Tetramers of Ag-presenting molecules permit Ag-specific T cells to be isolated, quantified, tracked, and characterized from the milieu of T cells within the host (Altman et al., 1996; Davis et al., 2011). Indeed, the advent of tetramers and more elaborate multivalent technology has been of huge benefit in the characterization of MHC-I–, MHC-II–, and CD1d-restricted T cells and has also recently emerged for CD1b- and CD1c-restricted T cells (Benlagha et al., 2000; Matsuda et al., 2000; Kasmar et al., 2011; Ly et al., 2013).
Presently, human MAIT cells are phenotypically defined as CD3+, CD161hi, TRAV1-2+ T cells. There are clear limitations to this approach as their classification relies on reactivity with an anti–TRAV1-2 mAb, noting that TRAV1-2+ TCR usage is not limited to MAIT cells. For instance, public MHC-restricted T cells, as well as CD1b-restricted GEM T cells, also use this TRAV gene segment (Miles et al., 2005; Tynan et al., 2007; Van Rhijn et al., 2013), potentially leading to misleading classification of MAIT cells. Furthermore, there are risks associated with the use of this surrogate phenotype because of the potential down-regulation of CD161 after MAIT cell activation (Leeansyah et al., 2013) as exemplified in two recent studies of MAIT cells in HIV-infected subjects (Cosgrove et al., 2013; Leeansyah et al., 2013). Here, the loss of CD161hiTRAV1-2+ cells was interpreted as a loss of MAIT cells associated with HIV infection (Cosgrove et al., 2013); however, it was demonstrated that although many MAIT cells persisted, CD161 down-regulation rendered them undetectable using the CD161hi TRAV1-2 surrogate phenotype (Leeansyah et al., 2013). Moreover, the current characterization of MAIT cells in mice is even more challenging as Vα19-specific mAbs are unavailable for mouse MAIT cells. Recently, we identified a series of MR1-restricted ligands that originated from vitamin B, which includes the nonactivating folic acid metabolite, 6-formyl pterin (6-FP), and the highly potent riboflavin metabolite, reduced 6-hydroxymethyl-8-d-ribityllumazine (rRL-6-CH2OH), thereby providing the opportunity to develop Ag-specific multivalent reagents to track MAIT cells ex vivo (Kjer-Nielsen et al., 2012). Here, we show that MR1 tetramers bound to the potent MAIT cell–activating ligand rRL-6-CH2OH specifically detect MAIT cells in humans and mice, revealing heterogeneity in the human MAIT cell Ag receptor repertoire and defining phenotypic characteristics of these cells.
RESULTS
MR1 tetramer generation
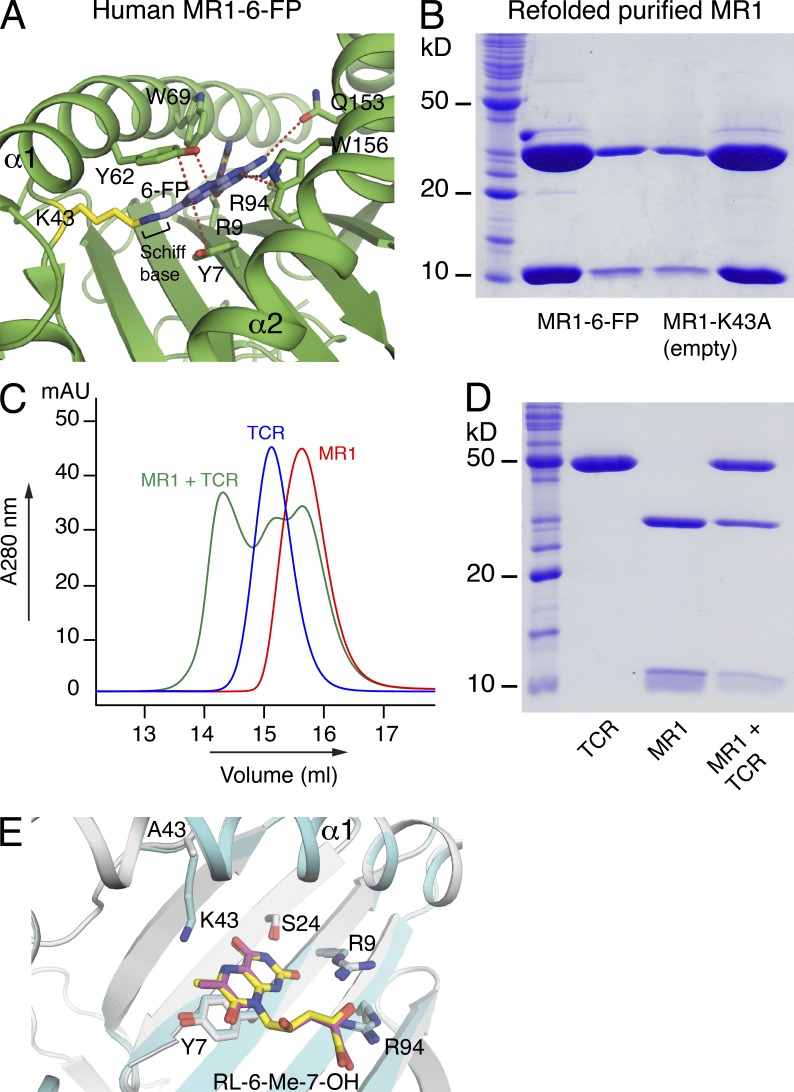
One potential hurdle in generating tetrameric reagents is that they generally require refolding and loading with a specific Ag such that the efficiency of this refolding/loading process can determine the broad applicability of the tetrameric reagent. Moreover, the associated stability of tetramers and potential Ag register shifts, as frequently observed with MHC-II tetramers, can impact their reliability (Davis et al., 2011). MR1 is distinct from other MHC-like Ag-presenting molecules, in that the Ag-binding cleft contains an aromatic cradle of residues that is ideally suited to bind small molecule metabolites (Kjer-Nielsen et al., 2012). Nevertheless, we have found the refolding efficiencies with the MR1 ligands to be very low (not depicted), creating a challenge to develop MR1-Ag tetramers as universal and readily applicable reagents. The MR1-binding cleft is mostly hydrophobic, but at the base of the cleft Lys43 forms a Schiff base with 6-FP (Fig. 1 A). Hence, we reasoned that the formation of a covalent bond with Lys43 might be a critical driving force in efficient refolding with MR1-restricted ligands. To formally test this, we engineered human and mouse MR1 in which Lys43 was mutated to Ala (K43A). Both the mouse and human MR1-K43A molecules refolded in the absence of any added ligand (Fig. 1 B). This was surprising as MHC and CD1 Ag-presenting molecules are typically only stable in the presence of bound Ag. The refolded “empty” MR1-K43A mutants (termed MR1-empty) exhibited similar biophysical properties to that of refolded wild-type MR1–6-FP complexes (Fig. 1 B and not depicted) and reacted with the anti-MR1 mAb 26.5 similarly to that of wild-type MR1–6-FP (not depicted). An empty and stable soluble MR1 suggested we could potentially load different MR1-restricted Ags of choice to generate MR1-Ag–specific reagents. Thus, we incubated the potent MAIT cell agonist rRL-6-CH2OH with MR1-empty, after which loading of MR1 with rRL-6-CH2OH was demonstrated by addition of soluble MAIT TCR and purification by gel-filtration chromatography of the ternary complex (Fig. 1, C and D). To further demonstrate that MR1-K43A binds ligands in the same manner as wild-type MR1, the ternary structure of refolded MR1-K43A, loaded with the MAIT cell agonist RL-6-Me-7-OH and complexed to a MAIT TCR, was determined (Fig. 1 E and Table 1). This revealed virtually identical modes of MAIT TCR binding to RL-6-Me-7-OH bound to MR1-K43A, when compared with the recently solved structure of the MAIT TCR–MR1–RL-6-Me-7-OH complex (Patel et al., 2013). Thus, MR1-K43A and wild-type MR1 present riboflavin metabolites to MAIT cells in a highly similar manner.
Figure 1.
Refolding and subsequent loading of mutant Lys43→Ala MR1 (MR1-K43A) with rRL-6-CH2OH. (A) Structure of wild-type human MR1 (green) with 6-FP (blue) forming a Schiff base with lysine 43 (K43; yellow). Van der Waal contacts (red) and hydrogen bonds (black) are shown (Kjer-Nielsen et al., 2012). (B) 15% SDS-PAGE of graded amounts of refolded and purified wild-type MR1 (MR1–6-FP; refolded with 6-FP), and empty MR1-K43A. Molecular mass markers are indicated in kilodaltons. This experiment was performed three times; shown is a representative experiment. (C) Gel filtration (S200 10/300 GL; GE Healthcare) purification of ternary complex: MR1-K43A (MR1) was loaded with rRL-6-CH2OH and subsequently complexed with soluble MAIT TCR (TCR). Ternary complex (MR1 + TCR) elutes at an earlier retention time than MR1 and MAIT TCR alone. Absorption at 280 nm and volume (ml) are shown on the y and x axis, respectively. (D) 15% SDS-PAGE of the ternary complex of MR1-K43A (loaded with rRL-6-CH2OH) and soluble MAIT TCR (purified by gel filtration as shown in C). Shown are molecular mass markers (in kilodaltons) and MAIT TCR (TCR), MR1-K43A (MR1), and ternary complex (MR1 + TCR) as indicated. (C and D) These experiments were performed twice; shown are representative experiments. (E) Overlay of ternary MR1-K43A–RL-6-Me-7-OH–MAIT TCR complex, with wild-type MR1–RL-6-Me-7-OH–MAIT TCR complex (Patel et al., 2013), viewed down into the Ag-binding cleft. RL-6-Me-7-OH in MR1-K43A–RL-6-Me-7-OH–MAIT TCR complex, yellow; RL-6-Me-7-OH in wild-type MR1–RL-6-Me-7-OH–MAIT TCR complex, magenta; MR1 in MR1-K43A–RL-6-Me-7-OH–MAIT TCR complex, gray; MR1 in wild-type MR1–RL-6-Me-7-OH–MAIT TCR complex, cyan; oxygen, red; nitrogen, blue.
Table 1.
Data collection and refinement statistics
| Parameter | MAIT–MR1-K43A–RL-6-Me-7-OH |
| Data collection | |
| Temperature | 100K |
| Space group | C2 |
| Cell dimensions | |
| a, b, c (Å) | 215.91, 69.36, 142.83 |
| α, β, γ (°) | 90, 104.30, 90 |
| Resolution (Å) | 75.01-2.40 (2.53-2.40) |
| Rpima | 7.9 (36.8) |
| I/σ1 | 7.4 (2.1) |
| Completeness (%) | 99.8 (99.9) |
| Total no. of observations | 344,942 (50,778) |
| No. of unique observations | 80,366 (11,654) |
| Multiplicity | 4.3 (4.4) |
| Refinement statistics | |
| Rfactor (%)b | 17.9 |
| Rfree (%)c | 22.7 |
| No. of atoms | |
| Protein | 12,417 |
| Ligand | 46 |
| Water | 334 |
| Ramachandran plot (%) | |
| Most favored | 91.0 |
| Allowed region | 9.0 |
| B factors (Å2) | |
| Protein | 35.4 |
| Ligand | 28.7 |
| rmsd bonds (Å) | 0.010 |
| rmsd angles (°) | 1.11 |
Values in parentheses refer to the highest resolution bin.
Rpim = Σhkl [1/(N − 1)]1/2 Σi|Ihkl, i − <Ihkl>|/Σhkl <Ihkl>.
Rfactor = (Σ||Fo| − |Fc||)/(Σ|Fo|) for all data except as indicated in footnote c.
5% of data was used for the Rfree calculation.
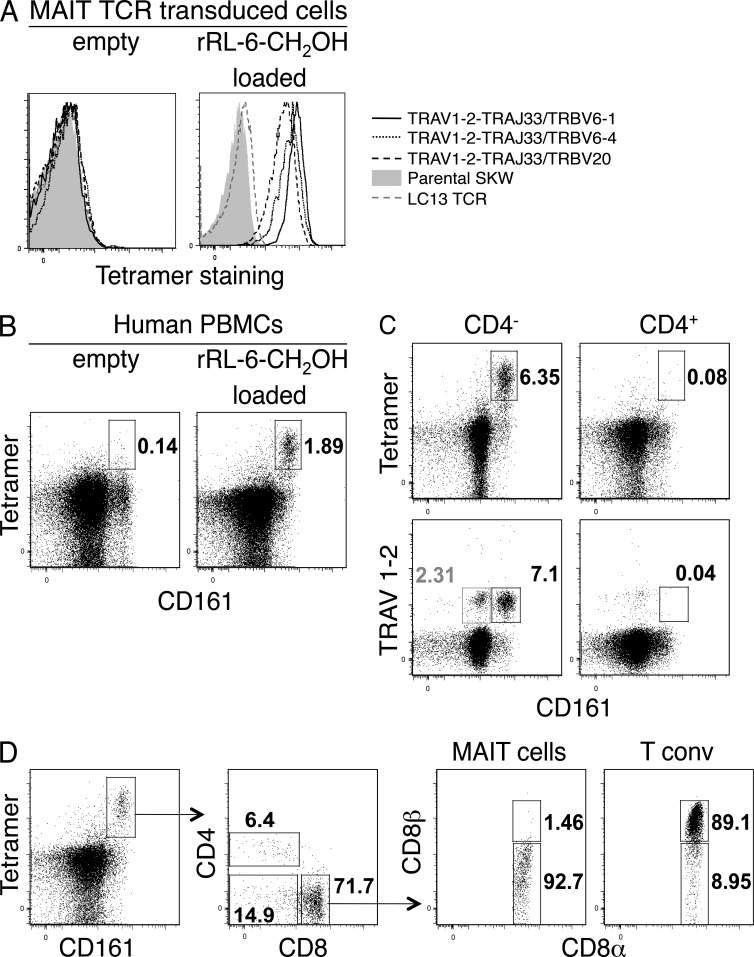
Tetrameric versions of human mutant K43A MR1-empty and MR1–rRL-6-CH2OH were formed by streptavidin cross-linking of biotin-labeled MR1, and these tetramers (MR1-Ag tetramers) were tested for their ability to stain the human T cell line SKW3 expressing MAIT TCRs (SKW.MAITs) with three different TCR β-chains (TRBV6-1, TRBV6-4, and TRBV20; Fig. 2 A). Empty MR1-tetramers did not bind to the SKW.MAIT cells (Fig. 2 A, left). MR1-Ag tetramers specifically bound all three SKW.MAIT cells but did not bind to SKW.LC13 cells expressing the MHC-I–restricted antiviral TCR LC13 (Fig. 2 A, right; Kjer-Nielsen et al., 2003). These data demonstrate that MR1-Ag tetramers could efficiently bind MAIT TCRs using a range of distinct TCR β-chains derived from human MAIT cells (Tilloy et al., 1999).
Figure 2.
Ag-specific identification of human MAIT cells. (A) Direct immunofluorescent staining of SKW3 cells expressing MAIT TCRs: TRAV1-2–TRAJ33 with TRBV6-1, TRBV6-4, or TRBV20, with empty (left) or rRL-6-CH2OH–loaded (right) human MR1 tetramer. Controls include nontransduced SKW3 (Parental SKW) and SKW3-transduced with an irrelevant TCR (LC13 TCR). (B) Human PBMCs were stained with CD3- and CD161-reactive mAbs and human MR1 tetramer. Shown are dot plots of CD3+ cells stained with empty (left) or rRL-6-CH2OH–loaded (right) human MR1 tetramer. Percentages of cells within boxed regions are indicated. Tetramer and CD161 staining are shown on the y and x axis, respectively. (C) Staining of human PBMCs comparing percentages of CD3+CD4− (left) and CD3+CD4+ (right) T cells detected by human MR1-Ag tetramer or TRAV1-2 mAb. Percentages of cells within (black) boxed regions are indicated. Also indicated is the percentage (gray) of CD3+CD4−TRAV1-2+CD161lo cells within the gray box (bottom left plot). Tetramer or TRAV1-2 and CD161 staining are shown on the y and x axis, respectively. Experiments in A–C were performed three times with similar results; shown are representative experiments. (D) Coreceptor expression on CD161+tetramer+ MAIT cells. CD3+CD161+tetramer+ cells were stained for cell surface expression of CD4, CD8α, and CD8β. CD8αα (CD8α+CD8β−) and CD8αβ (CD8α+CD8β+) expression on CD8+ MAIT cells was subsequently compared with expression on conventional CD8+ T cells (n = 6; shown is a representative staining experiment).
Detection of primary MAIT cell subsets
We next stained PBMCs from healthy donors with human MR1-Ag tetramers and compared the results with the established MAIT cell–staining protocol using anti-CD3, CD161, CD4, and TRAV1-2 markers. In agreement with MR1-Ag tetramer staining of the MAIT TCR–transduced SKW3 cells, a discrete population of PBMCs stained with the MR1-Ag tetramer in comparison with the MR1-empty tetramer, highlighting the specificity of the MR1-Ag tetramers (Fig. 2 B). Consistent with previous studies (Martin et al., 2009; Dusseaux et al., 2011), MR1-Ag tetramer+ cells were predominantly CD4− and CD161hi (Fig. 2 C). There was a close correlation between the percentage of MR1-Ag tetramer+ cells and the percentage of TRAV1-2+CD161hi cells (Fig. 2 C). Of note, from six donors, 10–24% of the TRAV1-2+, CD4− cells did not express high levels of CD161 (CD161hi), but an equivalent CD161lo tetramer-positive population was absent, presumably reflecting TRAV1-2 usage by non-MAIT cells (Fig. 2 C, bottom left). In addition, the TRAV1-2 mAb competes with MR1–rRL-6-CH2OH tetramer staining, indicating that the MR1-Ag tetramer was binding to TRAV1-2+ TCRs (not depicted).
As previously established for the current definition of MAIT cells within PBMCs, TRAV1-2+CD161hi T cells segregated mostly into the CD8+ and double-negative (DN) subsets, whereas the CD4+TRAV1-2+CD161hi subset was minor by comparison in five out of six donors (2–11% of MR1-Ag tetramer+ cells; Fig. 2, C and D). In the sixth donor, we observed low intensity of staining with MR1-Ag tetramer with a higher proportion (32%) of MR1-Ag tetramer+ cells that were CD4+ (not depicted). CD8 mediates adhesion and signal transduction, thus playing a crucial role in the development of T cells expressing MHC-I–restricted TCRs. CD8 comprises two subunits, CD8α and CD8β, encoded by two distinct genes, with CD8 existing in two isoforms, the αα homodimer and the αβ heterodimer. The CD8αβ heterodimer is by far the most predominant isoform (>90%) on MHC-restricted CD8+ T cells, with the CD8αα isoform being mostly restricted to gut intraepithelial lymphocytes (IELs; Das et al., 2000). Staining of PBMCs in six healthy donors using anti-CD8 mAbs demonstrated that the majority (>85%) of the tetramer+CD8+ subset was CD8α+ or CD8β−/lo, implying predominant expression of CD8αα homodimers, although some of these cells appear to coexpress CD8αβ heterodimers. In contrast, conventional MHC-restricted T cells mostly express high levels of the CD8αβ heterodimer (Fig. 2 D, right). Accordingly, these data indicate that the human MR1 tetramer+ cells differ in their CD8 expression from conventional T cells and are more like intestinal T cells, which express CD8αα homodimer.
Characterization of a noncanonical MAIT TCR repertoire in humans
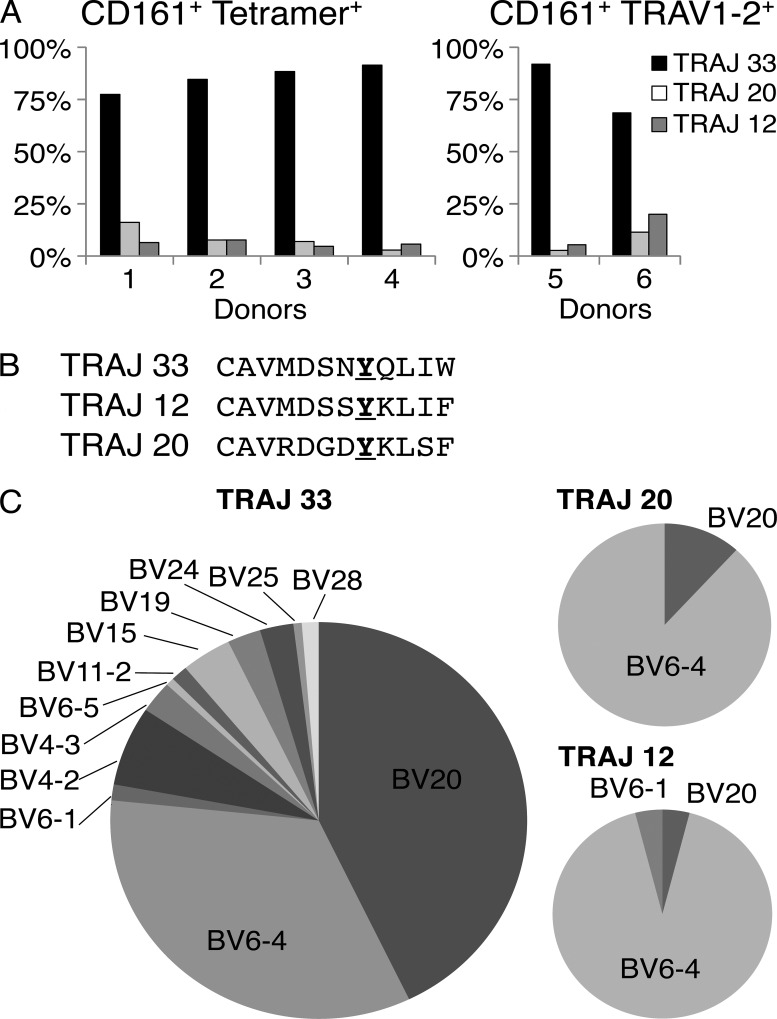
The canonical human MAIT TCR utilizes the invariant TRAV1-2–TRAJ33 α-chain paired predominantly with TRBV6 and TRBV20 β-chains (Tilloy et al., 1999). To investigate the MAIT TCR repertoire further in humans, the MR1-Ag tetramer and a TRAV1-2–reactive mAb (3C10; BioLegend) were used to stain PBMCs from four and two donors, respectively, after which single-cell sorted MR1-Ag tetramer+ or TRAV1-2+ cells were subjected to multiplex RT-PCR analysis (Wang et al., 2012) of their TCR genes, examining four T cell populations (CD3+CD4−CD161hiMR1-Ag tetramer+; CD3+CD4−CD161hiTRAV1-2+; CD3+CD4−CD161−TRAV1-2+; and CD3+CD4+CD161hiTRAV1-2+; Fig. 3). Most sorted MR1-Ag tetramer+ and TRAV1-2+ cells expressed the canonical TRAV1-2 gene segment joined to the TRAJ33 gene segment, thereby giving rise to the invariant TRAV1-2–TRAJ33 α-chain typically associated with MAIT TCRs (Tilloy et al., 1999). However, the TRAV1-2 gene did not exclusively join with the TRAJ33 gene segment (Fig. 3 A). Between 8 and 31% of sorted cells used TRAV1-2 joined with the TRAJ20 or TRAJ12 gene segments. This noncanonical TRAJ usage was observed using either MR1-Ag tetramers or mAb TRAV1-2+/CD161hi markers to identify MAIT cells. TRAJ20+ MAIT cells have previously been reported (Gold et al., 2010, 2013), and although there have been other studies suggestive of repertoire diversity in MAIT cells (Ebato et al., 1994; Maru et al., 2003; Hwang et al., 2006), these cannot be confirmed in the absence of definitive MAIT cell characterization. Our study demonstrates that in some individuals, a surprisingly high frequency (up to 31%) of MR1-Ag tetramer+ or mAb TRAV1-2+ cells can use alternative, non-TRAJ33 junctional genes. Although the sequences of TRAJ33, TRAJ20, and TRAJ12 differ slightly, they are relatively conserved overall, and notably, all three gene segments contained the conserved Tyr95α residue within the CDR3α loop (Fig. 3 B), which was previously shown to be crucial for MAIT cell activation (Reantragoon et al., 2012; Patel et al., 2013; Young and Gapin, 2013). A single TRAV1-2+TRAJ4+TRBV6-4+ clone was also identified within the TRAV1-2+CD161hi fraction, but this lacked Tyr95α, making it less likely to be a MAIT cell.
Figure 3.
Characterization of MR1-Ag tetramer-reactive MAIT cells. (A) Bar graphs showing TRAJ usage of CD3+CD4−CD161hitetramer+ cells (all of which expressed TRAV1-2; left, donors 1–4) or CD3+CD4−CD161hiTRAV1-2+ cells (stained with the TRAV1-2–specific mAb 3C10; right, donors 5 and 6). Bars represent percentage of total sequences (n = 32 [donor 1], 13 [donor 2], 43 [donor 3], 35 [donor 4], 37 [donor 5], and 35 [donor 6]) obtained from PCR amplification. (B) Alignment of CDR3α regions of TRAV1-2 TCR α-chains containing TRAJ33, TRAJ12, and TRAJ20 segments from CD161+tetramer+ cells. The conserved Tyr95α residue is highlighted in bold and underlined. (C) Pie charts comparing TRBV usage of MAIT cells with TRAV1-2 α-chains containing either TRAJ33 (left), TRAJ20 (top right), or TRAJ12 segments (bottom right; n, total number of paired sequences pooled from donors 1–6, = 154 [TRAJ33], 17 [TRAJ20], and 25 [TRAJ12]).
In the MAIT cell population where TRAV1-2 was joined with TRAJ33, the paired TCR-β segments were dominated by TRBV6-4 and TRBV20 genes, as previously reported (Tilloy et al., 1999), but TRAV1-2–TRAJ33 α-chains also paired with TRBV28, TRBV25, TRBV24, TRBV19, TRBV15, TRBV11-2, TRBV6-5, TRBV6-1, TRBV4-3, and TRBV4-2 (Fig. 3, A and C). This is consistent with mutagenesis and structural studies of the human MAIT TCR showing that the residues critical to MR1-mediated recognition were largely confined to the MAIT TCR α-chain and CDR3β loop of the MAIT TCR β-chain (Reantragoon et al., 2012; Patel et al., 2013; Young and Gapin, 2013). Interestingly, although sampled from a more limited number of cells, the TRBV usage in the noncanonical TRAJ20+ and TRAJ12+ MAIT cells was dominated by TRBV6-4 genes (P < 0.0001 for TRAJ12 and P < 0.0001 for TRAJ20, Fisher’s exact test), suggesting preferential pairing of these TCRs may be associated with noncanonical MAIT cell development (Fig. 3 C). These findings collectively reveal unexpected heterogeneity in the MAIT TCR repertoire.
Identical Ag dependence of MAIT cells expressing noncanonical TCRs
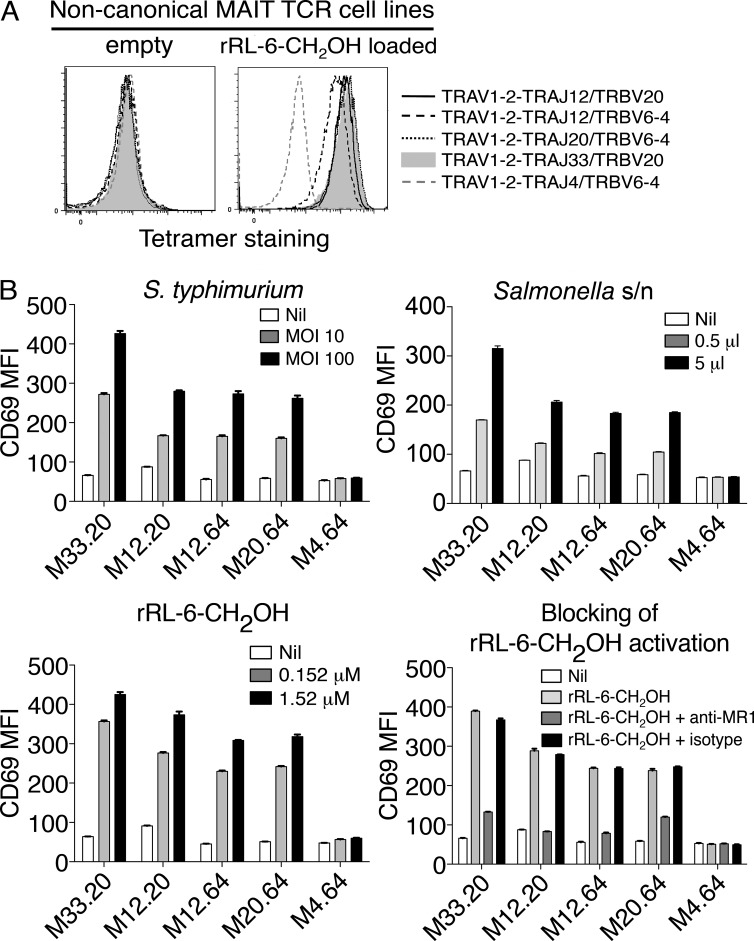
To formally investigate whether the noncanonical, cell surface phenotype–defined MAIT TCRs were MR1 restricted and Ag specific, we transduced genes encoding TRAV1-2–TRAJ20 and TRAV1-2–TRAJ12 TCRs into SKW3 cells, along with their cognate TRBV genes. These SKW.MAIT TCR transductants bound to MR1-Ag tetramers but not to MR1-empty tetramers (Fig. 4 A). The single TRAV1-2–TRAJ4 TRBV6-4 TCR was also included in this panel of transductants, but, as anticipated by the lack of Tyr95α, it did not stain with MR1-Ag tetramers (Fig. 4 A). Furthermore, the TRAV1-2–TRAJ4+ SKW transductant was not activated by Salmonella typhimurium infection, S. typhimurium supernatant, or synthetic rRL-6-CH2OH Ag (Fig. 4 B) but was activated upon stimulation with CD3/CD28 mAbs (not depicted). Thus, this TCR did not appear to represent a MAIT TCR but served as a valuable negative control for the other noncanonical MAIT TCRs. However, the remaining SKW.MAIT transductants were specifically activated by S. typhimurium–infected C1R cells and by C1R cells to which MAIT-activating ligands, either furnished from bacterial supernatant or as synthetic rRL-6-CH2OH, had been added (Fig. 4 B; Kjer-Nielsen et al., 2012). Importantly, the activation of the SKW transductants expressing these noncanonical MAIT TCRs was blocked by coincubation with an anti-MR1 mAb, but not an isotype-matched HLA-ABC–reactive mAb, confirming the MR1 restriction of these cells (Fig. 4 B). Thus, noncanonical MAIT TCRs share ligand specificity and functional properties of canonical MAIT cells expressing TRAV1-2–TRAJ33.
Figure 4.
T cells expressing TRAV1-2–TRAJ12 and TRAV1-2–TRAJ20 α-chains reconstitute features of canonical TRAV1-2–TRAJ33 MAIT cells. (A) Direct immunofluorescent staining of SKW3 cells expressing noncanonical MAIT TCRs: TRAV1-2–TRAJ12 with TRBV20 or TRBV6-4 and TRAV1-2–TRAJ20 with TRBV6-4, with empty (left) or human MR1-Ag tetramer (rRL-6-CH2OH loaded; right). Controls include a canonical MAIT TCR comprising TRAV1-2–TRAJ33 and TRBV20 α- and β-chains and a non-MAIT TCR comprising TRAV1-2–TRAJ4 and TRBV6-4 α- and β-chains. This experiment was performed twice with similar results. (B) SKW3 cells expressing noncanonical MAIT TCRs using TRAV1-2 joined with different TRAJs (M33.20, TRAV1-2–TRAJ33 plus TRBV20; M12.20, TRAV1-2–TRAJ12 plus TRBV20; M12.64, TRAV1-2–TRAJ12 plus TRBV6-4; M20.64, TRAV1-2–TRAJ20 plus TRBV6-4; and control non-MAIT TCR M4.64, TRAV1-2–TRAJ4 plus TRBV6-4) were activated by either infection with S. typhimurium (top left) or by addition of S. typhimurium supernatant (s/n; top right), rRL-6-CH2OH (bottom left), or rRL-6-CH2OH in the absence or presence of the anti-MR1 mAb (26.5) or an isotype control mAb (W6/32; bottom right). CD69 mean fluorescence intensity (MFI) is shown on the y axis. The tetramer-staining experiment was performed twice; the S. typhimurium infection, addition of S. typhimurium supernatant, and addition of rRL-6-CH2-OH experiments were performed once, three times, and three times, respectively, with similar results. Error bars show SEM of triplicates. Representative results are shown.
MAIT cells in the intestinal mucosa
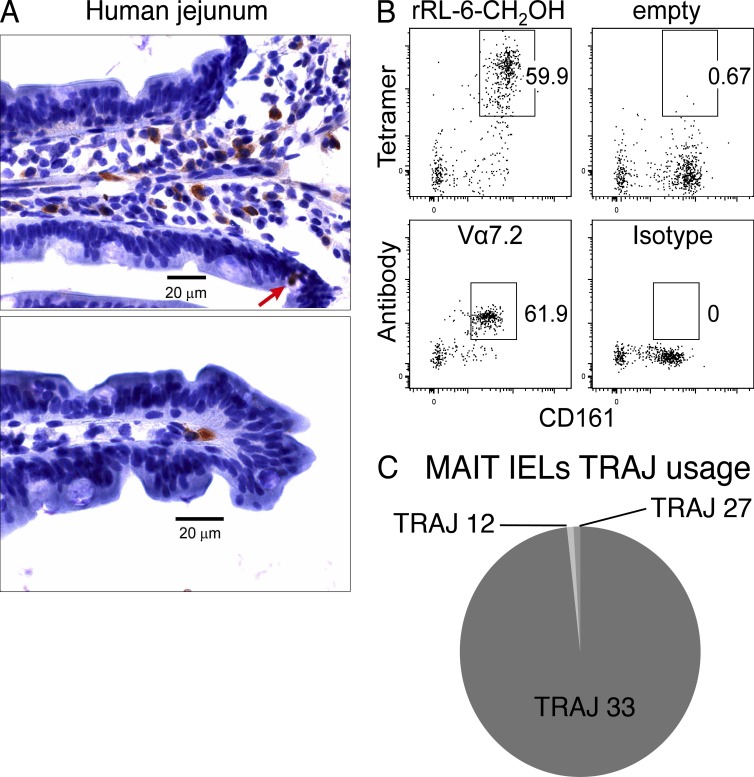
As MAIT cells are enriched at mucosal sites, we determined whether TRAV1-2+ T cells could be detected in tissue sections from intestinal mucosa and whether these cells expressed the canonical MAIT TCR α-chain sequences. Healthy human jejunal tissue was collected into cold PBS, after a Whipple’s procedure, and then fixed in cold 4% formaldehyde. Fixation buffer was washed out with PBS, and cryosections were processed for immunohistochemistry using the TRAV1-2–specific mAb D5 (Kjer-Nielsen et al., 2012). TRAV1-2+ T cells were observed in the lamina propria, predominantly at the bases of the villi (Fig. 5 A, top). Occasionally, cells were observed at the villus tip (Fig. 5 A, bottom), and there were rare IELs (Fig. 5 A, top, arrow). For detection of MAIT cells by flow cytometry, single-cell suspensions were prepared from jejunum tissue and were stained for CD3, CD4, CD161, and either MR1-Ag tetramer or anti–TRAV1-2 mAb (D5), or empty MR1 tetramer or isotype control mAb (Fig. 5 B). The proportion of MR1-Ag tetramer+CD161hi cells and TRAV1-2+CD161hi was similar (59.9–61.9% of CD3+CD4− lymphocytes, respectively), revealing a significant enrichment of MAIT cells in the jejunal mucosa. Moreover, the majority of the TRAV1-2+ cells expressed TRAJ33 (Fig. 5 C) and paired with one or other of the TRBV6-4, TRBV20, TRBV19, and TRBV3-1 TCR β-chains (not depicted). Accordingly, MR1-Ag tetramer+ MAIT cells are highly enriched in healthy jejunal tissue and express a TCR repertoire largely typical of blood MAIT cells.
Figure 5.
In situ localization and TCR repertoire analysis of jejunal MAIT cells. (A) Immunohistochemical staining of MAIT cells in the lamina propria in a human jejunal tissue section. Brown indicates TRAV1-2 (D5 mAb) staining detected with peroxidase-conjugated secondary antibody. The red arrow shows an example of a TRAV1-2+ intraepithelial MAIT cell stained with the D5 mAb. (B) Flow analysis of human IELs prepared from jejunal sections with mAbs against CD3, CD4, and CD161 and either human MR1-Ag tetramer (rRL-6-CH2OH; top left) or empty MR1-tetramer (top right) or TRAV1-2–reactive mAb (Vα7.2; bottom left) or isotype control (bottom right). Tetramer or TRAV1-2 and CD161 staining are shown on the y and x axis, respectively. This staining experiment was performed three times with similar results. (C) Pie chart showing TRAJ usage of MAIT IELs (CD161hiTRAV1-2+). Proportions are calculated from a total of 113 sequences.
Functional activity of MR1-tetramer–reactive MAIT cells
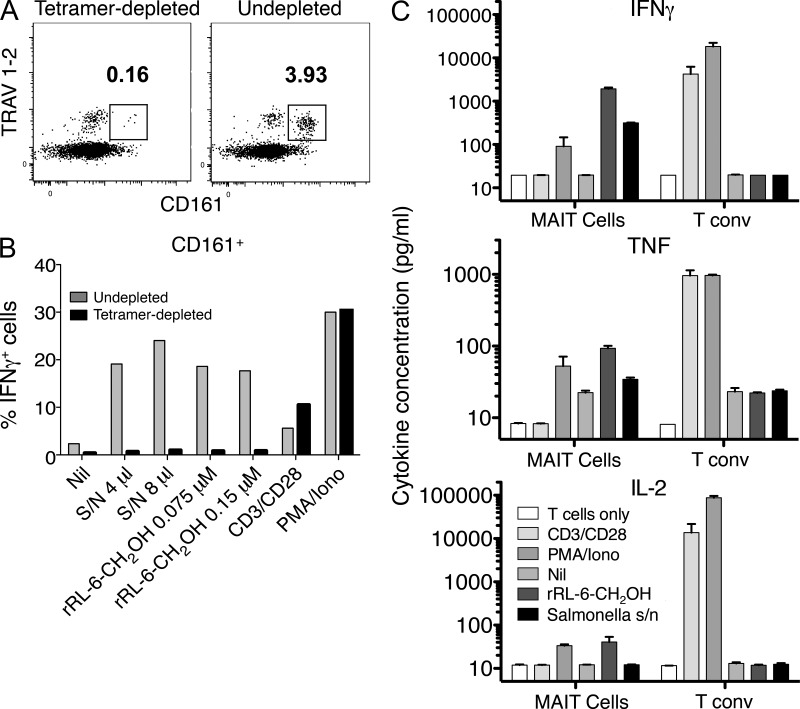
To determine whether the MR1-Ag tetramers identified all detectable human MAIT cells, we performed depletion of MR1-tetramer+ cells by flow sorting and then tested the functional activity of the remaining T cells. Depletion of cells with MR1-Ag tetramers removed T cell reactivity to both synthetic rRL-6-CH2OH and S. typhimurium supernatant, despite the presence of residual T cells expressing the TRAV1-2 TCR (Fig. 6, A and B). Both depleted and nondepleted CD161+ T cells produced IFN-γ in response to anti-CD3 and CD28 and PMA/ionomycin stimulation, demonstrating that CD161+ cells were capable of non-TCR–mediated activation (Fig. 6 B).
Figure 6.
MR1-Ag tetramer detects all MAIT cells. (A and B) Human PBMCs were depleted of MAIT cells by staining with rRL-6-CH2OH–loaded human MR1 tetramer and subsequent sorting by flow cytometry. Two-dimensional dot plots show analytical staining of a fraction of cells after sorting with TRAV1-2 staining (y axis; using the anti–TRAV1-2–specific mAb D5) versus CD161 staining (x axis) after tetramer depletion (left) or with control undepleted cells (right). Cells subsequently used in the experiment depicted in B were not subjected to prior treatment with mAbs. Undepleted and tetramer-depleted PBMCs were treated with rRL-6-CH2OH or S. typhimurium supernatant (S/N) or had no treatment (Nil) in the presence of C1R.MR1 cells or were treated with anti-CD3/anti-CD28 beads or PMA/ionomycin (Iono). Bar graphs shown are gated on CD3+CD4−CD161+ T cells. Bars represent percentage of IFN-γ–producing cells (% IFNγ+ cells). This MAIT tetramer depletion experiment was performed three times; a representative experiment is shown. The experiment was performed in triplicate, and shown are means. (C) Human PBMCs were stained with CD3- and CD161-reactive mAbs, and rRL-6-CH2OH–loaded MR1 tetramer and CD3+CD161hi MR1-Ag tetramer+ cells were then sorted before either no treatment (T cells only) or treatment with anti-CD3/anti-CD28 beads (CD3/CD28) or PMA/ionomycin or were coincubated with C1R.MR1 cells in the absence of treatment (Nil) or with the addition of rRL-6-CH2OH or S. typhimurium supernatant. Production (pg/ml) of IFN-γ, TNF, and IL-2 by MAIT cells or control CD3+CD161− MR1-Ag tetramer− T cells (T conv) was measured using a CBA assay. The mean of duplicate samples is shown, with error bars indicating SEM.
We next examined the cytokine profile of MR1-Ag tetramer+ cells activated by S. typhimurium supernatant or synthetic rRL-6-CH2OH Ag using a cytometric bead array (CBA) assay. Human MR1-Ag tetramer+ CD3+ cells were purified by flow cytometric sorting and stimulated in the presence of different activation agents for 60 h in culture. Conventional (MR1-Ag tetramer−, CD161−) T cells produced IFN-γ, TNF, and IL-2 upon incubation with anti-CD3/anti-CD28 beads and by PMA/ionomycin; however, as expected, they were not significantly activated by synthetic rRL-6-CH2OH Ag or S. typhimurium supernatant in the presence of C1R-MR1 APCs (Fig. 6 C). In contrast, CD161hi MR1-Ag tetramer+ T cells produced IFN-γ, TNF, and IL-2 upon incubation with synthetic rRL-6-CH2OH Ag or S. typhimurium supernatant in the presence of C1R.MR1 APCs. Surprisingly, activation of MR1-Ag tetramer+ cells by MAIT ligands did not induce production of IL-17, in contrast with an earlier study showing that human blood MAIT cells possess a TH17-like functional phenotype (Dusseaux et al., 2011). This may reflect differences in the stimulation conditions and/or donor differences between the two studies. Collectively, these data indicate that the MR1-Ag tetramers recognize all MAIT cells and can be used for their functional characterization without tetramer-mediated preactivation through the MAIT TCR.
Identification of mouse MAIT cells with mouse MR1-Ag tetramers
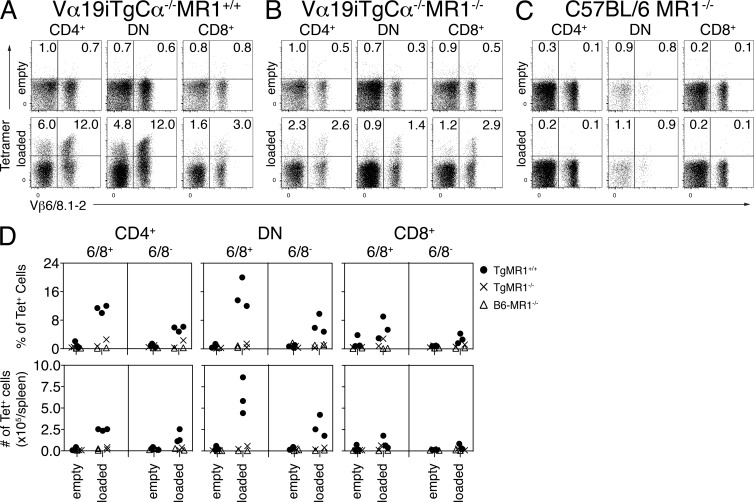
MAIT cells are extremely rare in standard laboratory strains of mice compared with humans (Tilloy et al., 1999; Treiner et al., 2003). Hence, most studies of mouse MAIT cells have used Tg lines expressing Vα19i on a Cα−/− background such that the Tg MAIT TCR α-chains pair with endogenous TCR β-chains (predominantly Vβ6 and Vβ8), thus enriching the number of MAIT cells (Okamoto et al., 2005; Croxford et al., 2006; Kawachi et al., 2006; Martin et al., 2009; Le Bourhis et al., 2010; Chua et al., 2012). To confirm the shared ligand specificity of human and mouse MAIT cells and to characterize the phenotype of mouse Tg MAIT cells, mouse MR1-Ag tetramers were used to stain splenic T cells from Vα19iTg-Cα−/−MR1+/+ mice as well as control Vα19iTg-Cα−/−MR1−/−, C57BL/6-MR1−/− (Fig.7, A–C), and C57BL/6 mice (not depicted). Empty MR1 tetramers did not stain a significant proportion of CD3+ cells from the spleens of any of these mice (Fig. 7, A–C, top), whereas the mouse MR1-Ag tetramers identified tetramer+ T cells in the CD4+, DN, and CD8+ T cell populations of Vα19iTg-Cα−/−MR1+/+ mice (Fig. 7, A, bottom).
Figure 7.
Detection of mouse MAIT cells in Vα19iMR1+/+ Tg mice. (A–C) Staining of splenic pan T cells from Vα19iTg-Cα−/−MR1+/+ (A), Vα19iTg-Cα−/−MR1−/− (B), and C57BL/6-MR1−/− (C) mice with empty (top) or rRL-6-CH2OH–loaded (bottom) mouse MR1 tetramer in conjunction with mAbs reactive to Vβ6, Vβ8.1, and Vβ8.2. Representative dot plots from one of three experiments show staining of CD4+ (left), CD4−CD8− DN (middle), or CD8+ (right) cell populations. Tetramer and Vβ6, Vβ8.1, Vβ8.2 staining is shown on the y and x axis, respectively. (D) Composite data from three separate experiments showing percentages (top row) and absolute numbers (bottom row) of splenocytes from Vα19iTg-Cα−/−MR1+/+ (TgMR1+/+), Vα19iTg-Cα−/−MR1−/− (TgMR1−/−), or C57BL/6-MR1−/− (B6-MR1−/−) mice stained with empty or rRL-6-CH2OH–loaded mouse MR1 tetramer. Shown are staining of CD4+ (left), CD4−CD8− DN (middle), or CD8+ (right) splenocytes that were either MR1-Ag+ and Vβ6/Vβ8.1/Vβ8.2+ (6/8+) or Vβ6/Vβ8.1/Vβ8.2− (6/8−).
In contrast to human MAIT cells in which only a small proportion of MAIT cells expressed CD4 (mean 6%; Fig. 2 D), >40% of MR1-Ag tetramer+ cells from a Vα19iTg-Cα−/−MR1+/+ mouse were CD4+, with the remainder containing more DN than CD8+ cells (ratio 3–4:1; Fig. 7, A, bottom). Thus the phenotype of splenic mouse MR1-Ag tetramer+ cells created in the Tg model differs from the most frequent phenotype of natural human MAIT cells in blood. Interestingly, we found that MR1-Ag tetramer+, CD4+ splenocytes were present in greater numbers in Vα19iTg-Cα−/−MR1+/+ mice when compared with Vα19iTg-Cα−/−MR1−/− mice (Fig. 7, A, B, and D). In contrast, Kawachi et al. (2006) did not find significantly greater numbers of CD4+ splenocytes in Vα19iTg-Cα−/−MR1+/+ in comparison with Vα19iTg-Cα−/−MR1−/− mice. Notably, about two thirds of mouse MR1-Ag tetramer+ cells expressed the predominantly used Vβ6 or Vβ8 genes, and about one third of the MR1-Ag tetramer+ cells were Vβ6− and Vβ8− (Fig. 7, A, bottom). It was not surprising that the T cells not expressing the preferentially used Vβ6 and Vβ8.1/8.2 segments in the Vα19iTg-Cα−/−MR1+/+ mice were reactive to MR1-Ag tetramer as other Vβ segments can be used (Tilloy et al., 1999; Kawachi et al., 2006). Thus, it appears that in the same way that there is predominant but nonexclusive TRBV6 and TRBV20 chain selection by human MAIT TCRs, mouse MAIT TCRs can also use other β-chains in addition to the predominantly used Vβ6 and Vβ8. We confirmed this finding by RT-PCR analysis of TCR genes from BW5147 hybridomas generated from fusion with sorted MR1-Ag tetramer+ Vβ6−Vβ8− and Vβ6+Vβ8+ T cells (Fig. 8 A and not depicted). Interestingly, tetramer-reactive cells were not completely absent from Vα19iTg-Cα−/−MR1−/− mice lacking the MR1 restriction element (Fig. 7, B [bottom] and D). However, the proportion of tetramer-reactive cells, both as a percentage and as absolute numbers, was significantly less in splenocytes from Vα19iTg-Cα−/−MR1−/− mice when compared with Vα19iTg-Cα−/−MR1+/+ mice (Fig. 7, A, B, and D). This result is similar to what has been found for NKT cells, where CD1d–α-GalCer tetramers stained a proportion of T cells from CD1d−/− mice (Wei et al., 2006). Importantly, no clear population of splenocytes from both wild-type and C57BL/6-MR1−/− mice was detected with MR1-Ag tetramer when compared with staining by control empty MR1 tetramer (Fig. 7, C and D; and not depicted). Because the numbers of MAIT cells in wild-type C57BL/6 mice are extremely low when compared with humans (Tilloy et al., 1999; Treiner et al., 2003), this observation is consistent with the notion that the presence of both Vβ6+Vβ8+ and Vβ6−Vβ8− MR1-Ag tetramer+ cells from Vα19iTg-Cα−/− mice is not caused by nonspecific staining by MR1-Ag tetramers. Accordingly, mouse MR1-Ag tetramers specifically detect Vα19i+ T cells in the Tg mouse.
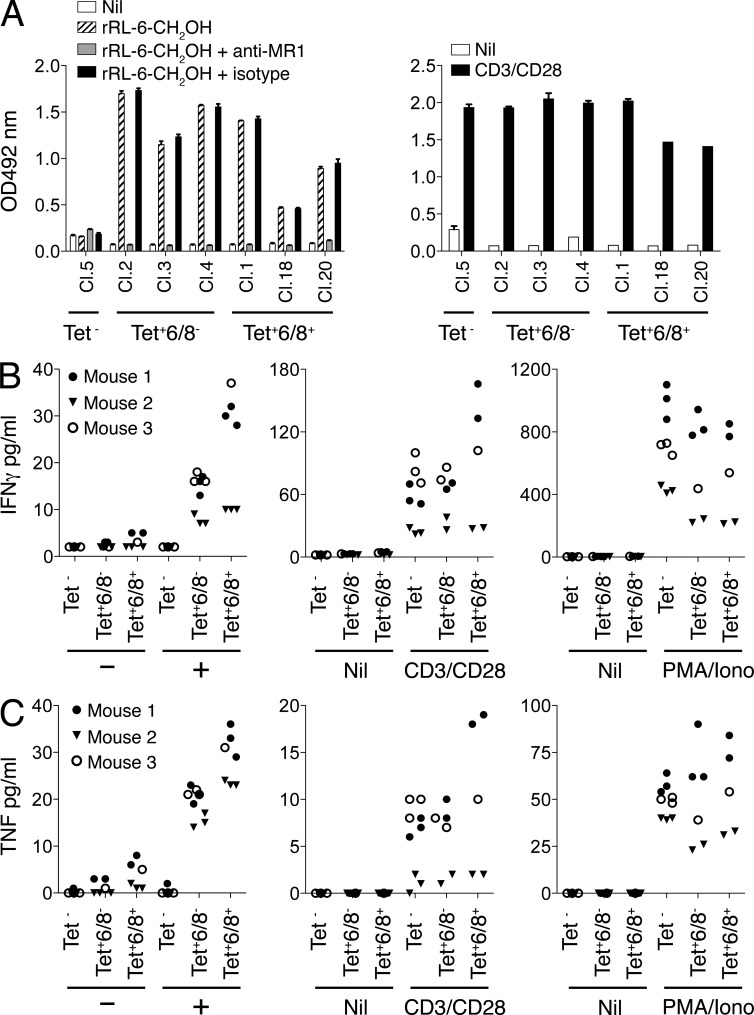
Figure 8.
Characterization of MR1-Ag tetramer–reactive mouse MAIT cells. (A) Activation of both Vβ6/Vβ8− and Vβ6/Vβ8+ BW5147-MAIT hybridomas by rRL-6-CH2OH. BW5147-MAIT hybridomas, either nonreactive with rRL-6-CH2OH–loaded MR1 tetramer (Tet−; Clone 5: Cl.5) or either MR1-Ag tetramer+Vβ6/Vβ8− (Tet+6/8−; Cl.2,3,4) or MR1-Ag tetramer+Vβ6/Vβ8+ (Tet+6/8+; Cl.1,18,20) were then (left) coincubated with M12.C3.MR1 cells in the absence (Nil) or presence of 15.2 µM rRL-6-CH2OH or presence of 15.2 µM rRL-6-CH2OH with prior addition of either the MR1-reactive mAb 26.5 (+anti-MR1) or an isotype control mAb, W6/32 (+isotype). Alternatively (right), BW5147-MAIT hybridomas were incubated alone (Nil) or in the presence of CD3- and CD28-reactive mAbs. IL-2 release by BW5147-MAIT hybridomas was measured by the conversion of o-phenylenediamine dihydrochloride by horseradish peroxidase in an indirect ELISA assay, and absorption at 492 nm is shown on the y axis. Results are shown as mean and SEM of triplicates. This experiment was performed three times; shown is a representative experiment. (B and C) Activation of both Vα19iTg+ Vβ6/Vβ8− and Vβ6/Vβ8+ MAIT cells by rRL-6-CH2OH. Mouse splenocytes from three separate Vα19iTg-Cα−/−MR1+ littermates were stained with CD3- and Vβ6/Vβ8-reactive mAbs and rRL-6-CH2OH–loaded MR1 tetramer. Sorted CD3+MR1-Ag tetramer− (Tet−), CD3+MR1-Ag tetramer+Vβ6/Vβ8− (Tet+6/8−), or CD3+MR1-Ag tetramer+Vβ6/Vβ8+ (Tet+6/8+) cells were then coincubated with M12.C3.MR1 cells in the absence (−) or presence (+) of 15.2 µM rRL-6-CH2OH. Sorted T cells were also treated with either CD3- and CD28-reactive mAbs (CD3/CD28) or PMA and ionomycin (PMA/iono) or left untreated (Nil). Production (pg/ml) of IFN-γ (B) and TNF (C) by MAIT cells or control MR1-Ag tetramer–negative T cells was measured using a CBA assay after 24 h. These data are representative of two independent experiments. In this experiment, MAIT cells and T cells were separately sorted from three Vα19iTg-Cα−/−MR1+ littermates (mouse 1, MR1+/+; mouse 2, MR1+/−; mouse 3, MR1+/−), with each symbol representing an individual data point. Depending on numbers of available sorted cells, one to three in vitro replicates per parameter were tested. A similar result was obtained when this experiment was repeated with MAIT and T cells separately sorted from two Vα19iTg-Cα−/−MR1+ littermates.
Functional activity of both Vα19iTg+ Vβ6/Vβ8− and Vβ6/Vβ8+ MR1-Ag tetramer–reactive mouse MAIT cells
There is currently no equivalent Vα19-specific mAb for the detection of MAIT cells in mice, such as the TRAV1-2–specific mAbs available for human MAIT cells. To verify that both Vβ6+/Vβ8+ and Vβ6−/Vβ8− MR1-Ag tetramer+ T cells from Vα19iTg mice were capable of stimulation by rRL-6-CH2OH in an MR1-restricted manner, we tested Vβ6+/Vβ8+ and Vβ6−/Vβ8− BW5147-MAIT cell hybridomas (derived from the fusion of MR1-Ag tetramer sorted splenocytes from a Vα19iTg-Cα−/−MR1+ mouse) for their capacity to be activated by rRL-6-CH2OH (Fig. 8 A). Both Vβ6+/Vβ8+ and Vβ6−/Vβ8− BW5147-MAIT hybridomas (which were MR1-Ag tetramer+) were stimulated by rRL-6-CH2OH to produce IL-2, and this activation was blocked by the MR1-specific mAb 26.5 but not the isotype control mAb W6/32 (Fig. 8 A, left). In contrast, a control BW5147 hybridoma, which did not stain with MR1-Ag tetramer, was not activated by rRL-6-CH2OH. (Fig. 8 A, left). As expected, RT-PCR analysis of RNA from the BW5147-MAIT hybridomas confirmed the presence of non-Vβ6/Vβ8 and Vβ6/Vβ8 TCR chains in Vβ6−/Vβ8− and Vβ6+/Vβ8+ hybridomas, respectively (not depicted). All BW5147 hybridomas were activated upon anti-CD3/CD28 mAb stimulation (Fig. 8 A, right). To further verify that MR1-Ag tetramer–reactive mouse splenocytes recognize rRL-6-CH2OH, CD3+MR1-Ag tetramer+Vβ6+/Vβ8+, CD3+MR1-Ag tetramer+Vβ6−/Vβ8−, or CD3+MR1-Ag tetramer− splenocytes from Vα19iTg+ mice were purified by flow cytometric sorting. Sorted cells were then coincubated with M12.C3.MR1 cells (expressing high levels of transduced mouse MR1) in the presence or absence of rRL-6-CH2OH, and production of IFN-γ and TNF by MR1-Ag tetramer+ cells was compared with that of control MR1-Ag tetramer− T cells using a CBA assay. MR1-Ag tetramer–positive cells, when stimulated by rRL-6-CH2OH in the presence of M12.C3.MR1 Ag-presenting cells, produced both IFN-γ and TNF, whether or not they were Vβ6+/Vβ8+ or Vβ6−/Vβ8− (Fig. 8, B and C). In contrast, MR1-Ag tetramer–negative cells were not stimulated by this Ag, although they could be stimulated by anti-CD3/CD28 or PMA/ionomycin (Fig. 8, B and C). Collectively, the mouse MR1-Ag tetramers offer a highly specific tool for tracking MAIT cells and verify the shared ligand specificity of human and mouse MAIT cells.
DISCUSSION
The advent of both classical and nonclassical MHC tetramer technology has revolutionized investigations into T cell–mediated immunity (Davis et al., 2011). Within the MHC axis, specific MHC-laden peptides need to be bound in a defined register to enable epitope-specific tracking of T cells. For CD1d-restricted NKT cells, which are usually characterized by invariant TRAV10-TRAJ18 TCR usage (which defines type I NKT cells), a universal CD1d-restricted Ag, α-galactosylceramide, is routinely used to track virtually the entire population of these cells (Benlagha et al., 2000; Matsuda et al., 2000). Moreover, tetrameric versions of less potent CD1d-restricted Ags can help delineate how variations within the type I NKT cell TCR β-chain repertoire can impact Ag recognition (Pellicci et al., 2009; Matulis et al., 2010). More recent studies have also demonstrated the potential for use of CD1b and CD1c tetramers loaded with microbial Ags to detect CD1b- and CD1c-restricted T cells, respectively (Kasmar et al., 2011; Ly et al., 2013). In this study, we have generated MR1-Ag–loaded tetramers and demonstrate the power of tetramer technology for the study of MAIT cells.
Despite the abundance of MAIT cells in humans (up to 10% of PBMC T cells), they have largely remained unexplored, and clinical practice is essentially oblivious to the presence and therapeutic use of MAIT cells. Furthermore, the utility of the surrogate TRAV1-2+, CD3+, CD161hi phenotype for MAIT cells has recently been called into question with a study showing that CD161 can be down-regulated on MAIT cells in HIV patients (Leeansyah et al., 2013). In mice, the problem of detection of MAIT cells is further confounded by the lack of a Vα19-specific mAb. Our limited understanding of MAIT cell function is principally attributable to lack of knowledge of the nature of the MR1-restricted MAIT cell Ag, and consequently not having Ag-specific reagents to track and characterize MAIT cells ex vivo. The recent description of riboflavin metabolites as MAIT cell Ags (Kjer-Nielsen et al., 2012) has afforded the opportunity to develop an MR1-Ag tetrameric reagent that can essentially monitor the entire population of MAIT cells.
As the efficiency of MR1 refolding with the vitamin B metabolites was found to be very low, thereby potentially limiting widespread use of a tetrameric reagent, we engineered MR1 so that it could be refolded in the absence of an added ligand. This empty MR1 exhibited essentially identical chromatographic and structural properties when compared with wild-type MR1, yet possessed the potential of being loaded with any putative MR1-restricted Ag of choice. To test the ability of this reagent to detect MAIT cells, we generated MR1 tetramers loaded with the potent rRL-6-CH2OH Ag. We demonstrate that MR1-Ag tetramers specifically detect human and mouse MAIT cells, thereby obviating the need for relying on the restricted TRAV1-2+CD161+CD3+ cell surface phenotype that is currently used to study human MAIT cells (Martin et al., 2009). Moreover, in the absence of a Vα19-reactive mAb, the mouse MR1-Ag tetramers represent the first reagent that can be used to reliably track MAIT cells from the Vα19iTg mouse. Comparison of staining of human and mouse lymphocytes with MR1-Ag tetramers revealed that there was a difference in the coreceptor-dependent subsets between mouse (as defined by the Vα19iTg mouse) and human MAIT cells, with the CD4+ subset representing ∼40–50% in the mouse, in contrast with the presence of 2–11% of CD4+ MAIT cells in humans, thereby suggesting differing coreceptor requirements between these species. Furthermore, it was notable that in humans the CD8+ MAIT cell subset was mostly CD8α+ CD8β−/lo, suggesting they predominantly express the CD8αα coreceptor, a CD8 coreceptor form which is also found on IELs located within the gut and other epithelial surfaces, in parallel with the predominant location of MAIT cells. The presence of low levels of CD8β on CD8α-high MAIT cells has also been previously observed (Martin et al., 2009; Walker et al., 2012) and suggests that some MAIT cells may coexpress CD8αβ heterodimer along with CD8αα homodimer. CD8 binds predominantly to the α3 domain of MHC-I, and a comparison of the MHC-I–CD8αα footprint reveals that the equivalent residues and structural features are mostly conserved/conservatively substituted between HLA-I and MR1 (Kjer-Nielsen et al., 2012). This suggests that MR1 may have the ability to bind to CD8αα, and accordingly may play a role in augmenting the MAIT cell immune response (Gao et al., 1997).
The MAIT cell repertoires are characterized in humans and mice by varied TCR β-chain usage, with particular β-chains being predominantly used by both humans and mice. We show that the MR1-Ag tetramer binds equally well to human MAIT TCRs expressing the TRBV6-1,TRBV 6-4, and TRBV20 genes; and mouse MAIT TCRs using Vβ6, Vβ8, and other Vβ-chains are capable of binding to MR1-Ag tetramer. In agreement with previous studies (Tilloy et al., 1999; Walker et al., 2012), TCR sequence analysis from human MR1-Ag tetramer–sorted PBMCs shows that as well as the frequently used TRBV6 and TRBV20 TCR β-chains, within the MAIT cell repertoire there is diversification through the (less frequent) usage of TRBV28, TRBV25, TRBV24, TRBV19, TRBV15, TRBV11-2, TRBV6-5, TRBV4-3, and TRBV4-2 β-chains. Given that the TCR-β repertoire is far more diverse in cord blood compared with adult blood (Walker et al., 2012), this suggests that TCR β-chain usage is less of a factor in MAIT TCR development but perhaps reflects peripheral expansion of MAIT cells in response to microbial challenge. However, it is equally clear that human MAIT cells expressing TCR β-chains other than TRBV6-4 and TRBV20 can be identified by the MR1-Ag tetramer. The less stringent requirement for particular TCR β-chains is consistent with recent MAIT TCR structural and mutagenesis data (Reantragoon et al., 2012; Patel et al., 2013; Young et al., 2013). Alternatively, varied MAIT TCR β-chain usage may modulate the affinity threshold to which MAIT cells respond to MR1-restricted Ags, in a manner analogous to that observed in type I NKT cells (Mallevaey et al., 2009; Pellicci et al., 2009; Patel et al., 2011; Rossjohn et al., 2012). Indeed, although the majority of MAIT cells were MR1-Ag tetramerhi, a smaller population was tetramerintermediate, suggesting that the MAIT TCR repertoire can impact avidity toward MR1-Ag. However, we did not observe a difference in TCR repertoire between MR1-Ag tetramerhi and tetramerintermediate cells (not depicted). Surprisingly, in addition to the varied MAIT TCR β-chain repertoire, we show that the human MAIT TCR α-chain repertoire can also be heterogeneous. Namely, although the TRAV1-2 gene usage appears to be fixed, the TRAJ33 gene can be replaced by TRAJ20 and TRAJ12 gene segments, which in some individuals appear to represent a considerable subset of MAIT cells. Although these alternate MAIT TCR α-chains display similar functionality in comparison with the TRAV1-2–TRAJ33+ MAIT TCRs, it was interesting to note that they showed a bias in TRBV usage, with the vast majority expressing TRBV6-4, suggesting preferential pairing of these TCR β-chains with the noncanonical TCR α-chains. This alternate use of the TRAJ20/TRAJ12 and TRAJ33 gene segments seem to be underpinned by a conserved Tyr95α residue within the J region of the CDR3α loop, whereupon this residue was shown to be a key requirement for MAIT TCR recognition (Reantragoon et al., 2012).
Despite the extensive heterogeneity of both the human and mouse MAIT cell populations, extending toward varied coreceptor usage, TCR α-chain TRAJ usage (for human MAIT cells), and TCR β-chain usage, the MR1-Ag tetramer effectively stains the majority of the human and Vα19iTg mouse MAIT cell population ex vivo. Moreover, coupled with the TRAV1-2 mAb, MR1-Ag tetramers can be used to verify MAIT cells in tissue sections. We demonstrate that human as well as Vα19iTg mouse MAIT cells, when activated by the potent rRL-6-CH2OH Ag, secrete a narrow range of TH1-biasing cytokines, which is in stark contrast to the TH1 and TH2 range of cytokines produced by the type I NKT cells upon stimulation by CD1d–α-GalCer (Kawano et al., 1997). Accordingly, the phenotypic heterogeneity of MAIT cells converges to functional homogeneity toward the potent MAIT cell Ag rRL-6-CH2OH. Our findings describe a highly specific tetrameric reagent to enable the tracking of MAIT cells ex vivo, while simultaneously providing fundamental insight into MAIT cell biology and MAIT TCR diversity. This reagent will greatly enhance studies of MAIT cells in disease as it will enable investigation of these cells in mice, and it will overcome concerns about limitations of surrogate phenotypes in humans. Collectively, these advances should see the development and use of MAIT cell reagents and therapeutics in the clinic.
MATERIALS AND METHODS
Generation of genes encoding mutant MR1-K43A.
Mutation of the genes encoding soluble human and mouse MR1 at position K43A was generated by the QuikChange Site-Directed Mutagenesis method (Agilent Technologies). A codon-encoding cysteine was next placed at the 3′ end the human and mouse MR1-K43A genes. Genes encoding the C-terminal cysteine–tagged human and mouse MR1-K43A heavy chains and human and mouse β2m were expressed separately in BL21 Escherichia coli, and inclusion body protein was prepared and solubilized in 8 M urea, 20 mM Tris-HCl, pH 8.0, 0.5 mM Na-EDTA, and 1 mM DTT as previously described (Kjer-Nielsen et al., 2012). MR1 and β2m were refolded in a buffer containing either no urea (human MR1) or containing 5 M urea (mouse MR1), 100 mM Tris, pH 8.0, 2 mM Na-EDTA, 400 mM l-arginine–HCl, 0.5 mM oxidized glutathione, 5 mM reduced glutathione, PMSF, and pepstatin A and dialyzed in 10 mM Tris before FPLC purification by sequential DEAE anion exchange, gel filtration, and Mono-Q anion exchange chromatography as described previously (Kjer-Nielsen et al., 2012).
Generation of human and mouse MR1-K43A tetramers.
Purified C-terminal cysteine–tagged MR1-K43A was “loaded” with rRL-6-CH2OH by incubating with a 100 molar excess of rRL-6-CH2OH for 4 h at room temperature in the dark. rRL-6-CH2OH–loaded cysteine-tagged MR1-K43A was then reduced with 5 mM DTT for 20 min before buffer exchange into PBS using a PD-10 column (GE Healthcare). The cysteine-tagged MR1-K43A was biotinylated with Maleimide-PEG2 biotin (Thermo Fisher Scientific) with a 30:1 molar ratio of biotin/protein at 4°C for 16 h in the dark. Biotinylated MR1 was subjected to S200 10/300 GL (GE Healthcare) chromatography to remove excess biotin. Biotinylated, rRL-6-CH2OH–loaded MR1-K43A monomers were tetramerized with streptavidin conjugated to PE (SA-PE; BD) or APC (SA-APC; Invitrogen).
Isolation of PBMCs.
Whole blood from healthy donors was collected (authorized by the Australian Red Blood Cross Service Material Supply Agreement with the University of Melbourne), and PBMCs were separated using Ficoll-Paque Premium (GE Healthcare). PBMCs were harvested and resuspended in fresh RPMI medium. Cells were then washed twice before resuspension in 10% DMSO in FCS. Before use, PBMCS were stored in liquid nitrogen.
MR1–rRL-6-CH2OH tetramer staining of human MAIT SKW3 cell lines, human PBMCs, and human IELs.
Approximately 2 × 105 of MAIT SKW3 cell lines, or human IELs, or 106 human PBMCs were stained with the empty or rRL-6-CH2OH–loaded versions of human MR1-K43A tetramer at 20 µg/ml for 40 min at room temperature in the dark. MAIT SKW3 cell lines were then stained with CD3-PE, and human PBMCs were stained with CD3-Pe-Cy7 (eBioscience), CD4-APC-Cy7 (BioLegend), CD8α-PerCP (BD), and CD161-APC (Miltenyi Biotec) or, alternatively, CD3–Alexa Fluor 700 (EBioscience), CD8β-PE (BD), or CD161-PeCy7 (BioLegend) for 30 min at 4°C. IELs were stained with CD3-Pe-Cy7 (eBioscience), CD4-APC-Cy7 (BioLegend), CD161-APC (Miltenyi Biotec), and anti–TRAV1-2 D5-FITC or isotype control (anti–pre-TCR) 8A5-FITC. Cells were then washed once with 2 ml FACS wash (2% FBS in PBS) and resuspended in 90 µl FACS fix (2.1% glucose and 1% paraformaldehyde in PBS).
Intestinal lymphocyte cell preparation and tissue immunohistochemistry.
Human jejunum mucosa was obtained from a patient undergoing a Whipple’s procedure and processed by the Australian Phenomics Histopathology Facility at the University of Melbourne. IELs and lamina propria lymphocytes were prepared essentially as described previously (Van Damme et al., 2001). In brief, mucosa was dissected away from underlying cell layers and washed briefly in PBS to remove blood and debris. Mucosal tissue was then incubated in PBS for 1 h at 37° with gentle agitation to allow dispersion of IELs. Tissue was allowed to settle and cells were then filtered (100 µM). The lamina propria lymphocyte cell fraction was obtained through further processing of tissue by manual cutting to 1–5 mm, followed by digestion with 50 U/ml Collagenase IV (Worthington Biochemical Corporation) for 2 h at 37°C with gentle agitation. Tissue was allowed to settle and cells were then filtered (100 µM) before analysis. For immunohistochemistry experiments, jejunal tissue was collected into cold PBS in the operating theater and then fixed in cold 4% formaldehyde. Fixative was washed out with PBS, and cryosections were made and processed for immunohistochemistry using the anti–TRAV1-2 mAb D5 (Kjer-Nielsen et al., 2012). mAb reactivity was detected using a peroxidase-coupled second antibody, and standard immunohistochemistry was conducted at the Australian Phenomics Histopathology and Organ Pathology service at the Department of Anatomy and Neuroscience at the University of Melbourne. This work was conducted under Ethics Approval H2011/04231.
CBA with human PBMCs.
Human PBMCs were stained with the rRL-6-CH2OH–loaded human MR1 tetramer–PE at 20 µg/ml for 40 min at room temperature before staining with CD3-PeCy7 (eBioscience), CD4–Pacific blue (eBioscience), γδ-TCR-FITC (BD), CD161-APC (Miltenyi Biotec), and CD8α-APCH7 (BD) for 30 min at 4°C in the dark. CD3+CD161+tetramer+ cells were sorted. Approximately 5,000 cells were incubated with anti-CD3/anti-CD28 beads (4 × 105 beads/well), 10 ng/ml PMA/1 µg/ml ionomycin, or with 10,000 C1R.MR1 cells in the presence of 1.52 µM rRL-6-CH2OH or 8 µl S. typhimurium supernatant in a final volume of 80 µl. At 24 and 60 h, 20 µl of supernatant was collected and replaced with fresh media (RPMI-1640 [Gibco] in supplement and 10% FCS). Supernatants were analyzed using CBA human flex sets for IFN-γ, TNF, and IL-2 (BD). In brief, 10 µl of culture supernatant was incubated with 10 µl of detection bead mix at room temperature for 1 h before addition of 10 µl of detection antibody mix and incubated for a further 1 h. Beads were then washed twice before data acquisition.
Tetramer-depleted intracellular cytokine staining assay.
Human PBMCs were stained with rRL-6-CH2OH–loaded human MR1 tetramer–PE at 20 µg/ml for 40 min at room temperature and bulk sorted for the tetramer-negative population (tetramer depleted). Undepleted control cells were PBMCs not stained with tetramer, but otherwise similarly bulk sorted for lymphocytes (using SSC versus FSC gates for both tetramer-depleted and undepleted cells). 2 × 105 of undepleted and tetramer-depleted cells were cultured with 2 × 105 C1R.MR1 cells (C1R cells transduced with and expressing high levels of human MR1) in the presence of 0.15 or 0.075 µM rRL-6-CH2OH or 4 or 8 µl S. typhimurium culture supernatant. Additionally, undepleted and tetramer-depleted cells were cultured without C1R.MR1 cells, in the absence or presence of either anti-CD3/anti-CD28 mAbs or 10 ng/ml PMA/1 µg/ml ionomycin. After overnight culture, cells were surface stained with CD3-Pe-Cy7 (eBioscience), CD4-APC-Cy7 (BioLegend), CD8α-PerCP (BD), and CD161-APC (Miltenyi Biotec) and TRAV1-2–FITC (D5; Kjer-Nielsen et al., 2012) mAbs for 30 min at 4°C in the dark and then fixed with 1% paraformaldehyde (ProSciTech), after which intracellular cytokine staining was performed with IFN-γ–Pacific blue (BioLegend) and granzyme-B–Alexa Fluor 700 (BD) in 0.3% Saponin (Sigma-Aldrich) for 40 min at 4°C in the dark.
Activation assay of human MAIT TCR SKW3 cell lines.
105 human MAIT TCR SKW3 cells were cultured with 105 C1R cells in the presence of 0.152 or 1.52 µM rRL-6-CH2OH, 0.5 or 5 µl S. typhimurium supernatant, or S. typhimurium (multiplicity of 10 or 100) in a total volume of 200 µl. For the MR1 blocking assay, anti-MR1 mAb (26.5; Huang et al., 2005) or isotype control mAb (W6/32; Maziarz et al., 1986) was added to C1R cells (20 µg/ml final concentration) for 1 h before culturing with SKW3 cells. After overnight culture, cells were stained with CD3-PE (BD) and CD69-APC (BD) for 30 min at 4°C, before analysis of CD69 surface expression by flow cytometry.
Data acquisition on flow cytometers.
Data were acquired on FACSCanto II, LSR II, and LSRFortessa flow cytometers (BD). Data were analyzed using FlowJo analysis software (Tree Star).
Mice.
MR1−/−, Vα19iTg-Cα−/−MR1+/+, and Vα19iTg-Cα−/−MR1−/− mice (all on C57BL/6 genetic background) were a gift from S. Gilfillan (Washington University in St. Louis School of Medicine, St. Louis, MO) to T.H. Hansen and Z. Chen. The Vα19iTg mice were crossed with TCRα−/− (Cα−/−) mice to eliminate expression of endogenous TCRα chains and then to MR1−/− mice (Vα19iTg-Cα−/−MR1−/−; Kawachi et al., 2006). Therefore, Vα19iTg-Cα−/−MR1+/+ and Vα19iTg-Cα−/−MR1−/− mice exclusively express a Vα19i transgene that is the canonical TCR Vα of mouse MAIT cells. As previously reported, MAIT cells isolated from these Tg mice use endogenous TCR Vβ chains and have surface marker characteristics of polyclonal MAIT cells (Kawachi et al., 2006). Mice were bred and maintained under specific pathogen–free conditions, and protocols used were approved by the Animal Studies Committee of Washington University in St. Louis and the University of Melbourne.
Isolation of mouse splenic cells.
Spleen cells from naive Vα19iTg-Cα−/−MR1+/+ mice, Vα19iTg-Cα−/−MR1−/−, C57BL/6-MR1−/−, and wild-type C57BL/6 mice were aseptically removed and used as the source for T cell isolation. A single-cell suspension was prepared, and red blood cells were lysed with ammonium chloride buffer. Cells were washed, counted, and subjected to magnetic labeling and separation according to the instructions of the pan-T cell isolation kit II, negative selection (Miltenyi Biotec).
MR1–rRL-6-CH2OH tetramer staining of mouse splenic T cells.
Approximately 106 purified T cells were washed with cold FACS staining buffer (1× PBS with 2% FBS), pelleted, and stained with 20 µg/ml PE-conjugated negative control mouse MR1-K43A tetramer (empty) or PE-conjugated rRL-6-CH2OH–loaded mouse MR1 tetramer in ∼10 µl (final volume) FACS staining buffer (in FACS tubes; BD) for 45 min at room temperature in the dark. Immediately thereafter (no washing step), surface Ags, including CD3ε, CD4, CD8α, Vβ6 TCR, and Vβ8.1/8.2 TCR were stained for 20 min on ice with anti–mouse CD3–Pacific blue (BD), anti–mouse CD4-PerCP (BioLegend), anti–mouse CD8α-APC-H7 (BD), anti–mouse Vβ6 TCR-FITC (BD), and anti–mouse Vβ8.1/8.2 TCR-FITC (BD), respectively. Cells were then washed once with 2 ml FACS staining buffer and resuspended in 200 µl FACS staining buffer.
CBA with mouse splenic T cells.
Splenocytes from a Vα19iTg-Cα−/−MR1+/+ mouse were stained with 20 µg/ml PE-conjugated rRL-6-CH2OH–loaded mouse MR1-K43A tetramer in ∼10 µl (final volume) FACS staining buffer before staining with anti–mouse CD3-APC, anti–mouse Vβ6 TCR-FITC (BD), and anti–mouse Vβ8.1/8.2 TCR-FITC (BD). Vβ6−/Vβ8.1/8.2−CD3+tetramer+, Vβ6+/Vβ8.1/8.2+CD3+tetramer+, or control CD3+tetramer− cells were sorted, and ∼20,000 cells were incubated with anti-CD3/anti-CD28 mAbs or 10 ng/ml PMA/1 µg/ml ionomycin or with 20,000 M12.C3.murineMR1 cells (Griffith et al., 1988; expressing high levels of transduced mouse MR1) in the absence or presence of 15.2 µM rRL-6-CH2OH in a final volume of 200 µl. At 24 h, 5 µl of supernatant was collected and analyzed using CBA mouse flex sets for IFN-γ and TNF (BD). In brief, 5 µl of culture supernatant (diluted to 10 µl) was incubated with 10 µl of detection bead mix at room temperature for 1 h before addition of 10 µl of detection antibody mix and incubated for a further 1 h. Beads were then washed twice before data acquisition.
Activation assay of BW5147-mouse MAIT TCR cell lines.
Splenocytes from a Vα19iTg-Cα−/−MR1+/+ mouse were stained with CD3- and Vβ6/Vβ8-reactive mAbs, and rRL-6-CH2OH–loaded MR1-K43A tetramer and CD3+MR1 tetramer+Vβ6/Vβ8− or CD3+MR1 tetramer+Vβ6/Vβ8+ cells were then sorted by flow cytometry before fusion with the BW5147 cell line as previously described (de Kauwe et al., 2009). BW5147-MAIT hybridomas, either nonreactive with rRL-6-CH2OH–loaded mouse MR1-K43A tetramer or either MR1-Ag tetramer+Vβ6/Vβ8− or MR1-Ag tetramer+Vβ6/Vβ8+, were then coincubated with M12.C3.MR1 cells in the absence or presence of 15.2 µM rRL-6-CH2OH. Alternatively BW5147-MAIT hybridomas were incubated alone or in the presence of plate-bound CD3- and CD28-reactive mAbs (BD). IL-2 release by BW5147-MAIT hybridomas was assayed by indirect ELISA with the conversion of o-phenylenediamine dihydrochloride by horseradish peroxidase and measurement of absorption at 492 nm.
PCR amplification of TCR sequences.
Human PBMCs stained with MR1-K43A tetramer loaded with rRL-6-CH2OH or IELs stained with TRAV1-2 (D5)–FITC and costained with CD3-PE-Cy7, CD4-APC-Cy7, and CD161-APC were single-cell sorted on a FACSAria flow cytometer into 96-well twin.tec skirted PCR plates (Eppendorf). Reverse transcription was performed with a Superscript VILO cDNA Synthesis kit (Invitrogen). Plates were incubated at 25°C for 5 min, 42°C for 30 min, and 80°C for 5 min and held at 16°C, as previously described (Wang et al., 2012). cDNA was then amplified with two rounds of nested PCR with a panel of multiple primers (external primers and internal primers, respectively) targeting α- and β-chains of the TCR. Plates were incubated at 95°C for 2 min, then followed by 35 cycles of 95°C for 20 s, 52°C for 20 s, and 72°C for 45 s, and lastly by 1 cycle of 72°C for 7 min. Samples were fractionated on 2% agarose gels to confirm for the presence of amplified products. Only samples with visible paired α- and β-chains were selected for sequencing.
Sequencing of TCR repertoire.
Samples were treated with 2 µl ExoSTAR (GE Healthcare), according to the manufacturer’s protocol, and incubated at 37°C for 15 min and 80°C for 15 min. Samples were then sequenced with internal primers with Big Dye 3.1 (Walter and Eliza Hall Institute of Medical Research) and incubated at 95°C for 5 min, followed next by 35 cycles of 96°C for 10 s, 50°C for 5 s, and 60°C for 4 min. The DyeEx 96 kit (QIAGEN) was used to remove dye terminators, according to the manufacturer’s protocol. Samples were sent to the Sequencing and Genotyping Facility, Department of Pathology, the University of Melbourne for electrophoresis and sequencing. Sequences were determined using IMGT/V-QUEST software.
Crystallization, structure determination, and refinement.
Ternary MR1-K43A–RL-6-Me-7-OH plus MAIT TCR complex (6–10 mg/ml) crystallized at 294 K in 0.2 M sodium acetate, 0.1 M Bis-Tris-Propane, pH 6.5, and 20% PEG 3350. Crystals were flash frozen before data collection using 10% glycerol as the cryoprotectant. The data were collected at 100 K on the 031D1 beamline at the Australian Synchrotron, Melbourne. The crystals diffracted to 2.4 Å and belong to the space group C2, with two molecules within the asymmetric unit. The data were processed using Mosflm version 7.0.5 (Leslie, 2006) and scaled using SCALA from the CCP4 Suite (CCP4, 1994). The data for ternary MR1-K43A–RL-6-Me-7-OH plus MAIT TCR complex was solved using MR1–RL-6-Me-7-OH plus MAIT TCR complex (Protein Data Bank accession no. 4L4V) without the ligand (Patel et al., 2013). To prevent model bias, the Rfree set of the MAIT TCR-MR1–6-FP (Protein Data Bank accession no. 4L4T) data was used in the experimental intensities scaling using SCALA as well as the implementation of the simulated annealing protocol in Phenix (Zwart et al., 2008). Refinement was performed using BUSTER 2.10. Model building was performed using COOT (Crystallographic Object-Oriented Toolkit). The quality of structure was validated at the Research Collaboratory for Structural Bioinformatics Protein Data Bank Data Validation and Deposition Services. All molecular graphics representations were created using PyMOL. The coordinates were deposited in the Protein Data Bank under the accession number 4LCW.
Acknowledgments
We thank Susan Gilfillan for generously making available Vα19iTg mice and Daniel Pellicci for help with generating TCR-transduced cell lines. We thank the Australian Synchrotron for assistance with data collection and the Australian Phenomics Network Histopathology Service for use of facilities.
This research was supported by the National Health and Medical Research Council of Australia (NHMRC). T.H. Hansen was supported by National Institutes of Health grant AI046553. R. Reantragoon was supported by the Faculty of Medicine, Chulalongkorn University and Chulalongkorn Hospital, Thai Red Cross Society scholarships. N.A. Gherardin was supported by a Leukemia Foundation postgraduate scholarship. D.P. Fairlie and D.I. Godfrey were supported by NHMRC Senior Principal Research Fellowships. O. Patel was supported by an Australian Research Council Future Fellowship. J. Rossjohn was supported by an NHMRC Australia Fellowship.
The authors declare that they have no competing financial interests.
Footnotes
Abbreviations used:
- 6-FP
- 6-formyl pterin
- Ag
- antigen
- CBA
- cytometric bead array
- DN
- double negative
- IEL
- intraepithelial lymphocyte
- MAIT cell
- mucosal-associated invariant T cell
- rRL-6-CH2OH
- reduced 6-hydroxymethyl-8-d-ribityllumazine
- Tg
- transgenic
References
- Altman J.D., Moss P.A., Goulder P.J., Barouch D.H., McHeyzer-Williams M.G., Bell J.I., McMichael A.J., Davis M.M. 1996. Phenotypic analysis of antigen-specific T lymphocytes. Science. 274:94–96 10.1126/science.274.5284.94 [DOI] [PubMed] [Google Scholar]
- Benlagha K., Weiss A., Beavis A., Teyton L., Bendelac A. 2000. In vivo identification of glycolipid antigen-specific T cells using fluorescent CD1d tetramers. J. Exp. Med. 191:1895–1903 10.1084/jem.191.11.1895 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chiba A., Tajima R., Tomi C., Miyazaki Y., Yamamura T., Miyake S. 2012. Mucosal-associated invariant T cells promote inflammation and exacerbate disease in murine models of arthritis. Arthritis Rheum. 64:153–161 10.1002/art.33314 [DOI] [PubMed] [Google Scholar]
- Chua W.J., Truscott S.M., Eickhoff C.S., Blazevic A., Hoft D.F., Hansen T.H. 2012. Polyclonal mucosa-associated invariant T cells have unique innate functions in bacterial infection. Infect. Immun. 80:3256–3267 10.1128/IAI.00279-12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cosgrove C., Ussher J.E., Rauch A., Gärtner K., Kurioka A., Hühn M.H., Adelmann K., Kang Y.H., Fergusson J.R., Simmonds P., et al. 2013. Early and nonreversible decrease of CD161++ /MAIT cells in HIV infection. Blood. 121:951–961 10.1182/blood-2012-06-436436 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Croxford J.L., Miyake S., Huang Y.Y., Shimamura M., Yamamura T. 2006. Invariant V(alpha)19i T cells regulate autoimmune inflammation. Nat. Immunol. 7:987–994 10.1038/ni1370 [DOI] [PubMed] [Google Scholar]
- Das G., Gould D.S., Augustine M.M., Fragoso G., Sciutto E., Stroynowski I., Van Kaer L., Schust D.J., Ploegh H., Janeway C.A., Jr 2000. Qa-2–dependent selection of CD8α/α T cell receptor α/β+ cells in murine intestinal intraepithelial lymphocytes. J. Exp. Med. 192:1521–1528 (published erratum appears in J. Exp. Med. 2001. 193:413) 10.1084/jem.192.10.1521 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davis M.M., Altman J.D., Newell E.W. 2011. Interrogating the repertoire: broadening the scope of peptide-MHC multimer analysis. Nat. Rev. Immunol. 11:551–558 10.1038/nri3020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Kauwe A.L., Chen Z., Anderson R.P., Keech C.L., Price J.D., Wijburg O., Jackson D.C., Ladhams J., Allison J., McCluskey J. 2009. Resistance to celiac disease in humanized HLA-DR3-DQ2-transgenic mice expressing specific anti-gliadin CD4+ T cells. J. Immunol. 182:7440–7450 10.4049/jimmunol.0900233 [DOI] [PubMed] [Google Scholar]
- Dusseaux M., Martin E., Serriari N., Péguillet I., Premel V., Louis D., Milder M., Le Bourhis L., Soudais C., Treiner E., Lantz O. 2011. Human MAIT cells are xenobiotic-resistant, tissue-targeted, CD161hi IL-17-secreting T cells. Blood. 117:1250–1259 10.1182/blood-2010-08-303339 [DOI] [PubMed] [Google Scholar]
- Ebato M., Nitta T., Yagita H., Sato K., Okumura K. 1994. Shared amino acid sequences in the ND beta N and N alpha regions of the T cell receptors of tumor-infiltrating lymphocytes within malignant glioma. Eur. J. Immunol. 24:2987–2992 10.1002/eji.1830241210 [DOI] [PubMed] [Google Scholar]
- Gao G.F., Tormo J., Gerth U.C., Wyer J.R., McMichael A.J., Stuart D.I., Bell J.I., Jones E.Y., Jakobsen B.K. 1997. Crystal structure of the complex between human CD8alpha(alpha) and HLA-A2. Nature. 387:630–634 10.1038/42523 [DOI] [PubMed] [Google Scholar]
- Gold M.C., Lewinsohn D.M. 2013. Co-dependents: MR1-restricted MAIT cells and their antimicrobial function. Nat. Rev. Microbiol. 11:14–19 10.1038/nrmicro2918 [DOI] [PubMed] [Google Scholar]
- Gold M.C., Cerri S., Smyk-Pearson S., Cansler M.E., Vogt T.M., Delepine J., Winata E., Swarbrick G.M., Chua W.J., Yu Y.Y., et al. 2010. Human mucosal associated invariant T cells detect bacterially infected cells. PLoS Biol. 8:e1000407 10.1371/journal.pbio.1000407 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gold M.C., Eid T., Smyk-Pearson S., Eberling Y., Swarbrick G.M., Langley S.M., Streeter P.R., Lewinsohn D.A., Lewinsohn D.M. 2013. Human thymic MR1-restricted MAIT cells are innate pathogen-reactive effectors that adapt following thymic egress. Mucosal Immunol. 6:35–44 10.1038/mi.2012.45 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Griffith I.J., Nabavi N., Ghogawala Z., Chase C.G., Rodriguez M., McKean D.J., Glimcher L.H. 1988. Structural mutation affecting intracellular transport and cell surface expression of murine class II molecules. J. Exp. Med. 167:541–555 10.1084/jem.167.2.541 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang S., Gilfillan S., Cella M., Miley M.J., Lantz O., Lybarger L., Fremont D.H., Hansen T.H. 2005. Evidence for MR1 antigen presentation to mucosal-associated invariant T cells. J. Biol. Chem. 280:21183–21193 10.1074/jbc.M501087200 [DOI] [PubMed] [Google Scholar]
- Hwang H.Y., Kim T.G., Kim T.Y. 2006. Analysis of T cell receptor alpha-chain variable region (Valpha) usage and CDR3alpha of T cells infiltrated into lesions of psoriasis patients. Mol. Immunol. 43:420–425 10.1016/j.molimm.2005.03.004 [DOI] [PubMed] [Google Scholar]
- Kasmar A.G., van Rhijn I., Cheng T.Y., Turner M., Seshadri C., Schiefner A., Kalathur R.C., Annand J.W., de Jong A., Shires J., et al. 2011. CD1b tetramers bind αβ T cell receptors to identify a mycobacterial glycolipid-reactive T cell repertoire in humans. J. Exp. Med. 208:1741–1747 10.1084/jem.20110665 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kawachi I., Maldonado J., Strader C., Gilfillan S. 2006. MR1-restricted V alpha 19i mucosal-associated invariant T cells are innate T cells in the gut lamina propria that provide a rapid and diverse cytokine response. J. Immunol. 176:1618–1627 [DOI] [PubMed] [Google Scholar]
- Kawano T., Cui J., Koezuka Y., Toura I., Kaneko Y., Motoki K., Ueno H., Nakagawa R., Sato H., Kondo E., et al. 1997. CD1d-restricted and TCR-mediated activation of valpha14 NKT cells by glycosylceramides. Science. 278:1626–1629 10.1126/science.278.5343.1626 [DOI] [PubMed] [Google Scholar]
- Kjer-Nielsen L., Clements C.S., Purcell A.W., Brooks A.G., Whisstock J.C., Burrows S.R., McCluskey J., Rossjohn J. 2003. A structural basis for the selection of dominant alphabeta T cell receptors in antiviral immunity. Immunity. 18:53–64 10.1016/S1074-7613(02)00513-7 [DOI] [PubMed] [Google Scholar]
- Kjer-Nielsen L., Patel O., Corbett A.J., Le Nours J., Meehan B., Liu L., Bhati M., Chen Z., Kostenko L., Reantragoon R., et al. 2012. MR1 presents microbial vitamin B metabolites to MAIT cells. Nature. 491:717–723 [DOI] [PubMed] [Google Scholar]
- Le Bourhis L., Martin E., Péguillet I., Guihot A., Froux N., Coré M., Lévy E., Dusseaux M., Meyssonnier V., Premel V., et al. 2010. Antimicrobial activity of mucosal-associated invariant T cells. Nat. Immunol. 11:701–708 (published erratum appears in Nat. Immunol. 2010. 11:969) 10.1038/ni.1890 [DOI] [PubMed] [Google Scholar]
- Le Bourhis L., Guerri L., Dusseaux M., Martin E., Soudais C., Lantz O. 2011. Mucosal-associated invariant T cells: unconventional development and function. Trends Immunol. 32:212–218 10.1016/j.it.2011.02.005 [DOI] [PubMed] [Google Scholar]
- Le Bourhis L., Mburu Y.K., Lantz O. 2013. MAIT cells, surveyors of a new class of antigen: development and functions. Curr. Opin. Immunol. 25:174–180 10.1016/j.coi.2013.01.005 [DOI] [PubMed] [Google Scholar]
- Leeansyah E., Ganesh A., Quigley M.F., Sönnerborg A., Andersson J., Hunt P.W., Somsouk M., Deeks S.G., Martin J.N., Moll M., et al. 2013. Activation, exhaustion, and persistent decline of the antimicrobial MR1-restricted MAIT-cell population in chronic HIV-1 infection. Blood. 121:1124–1135 10.1182/blood-2012-07-445429 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leslie A.G. 2006. The integration of macromolecular diffraction data. Acta Crystallogr. D Biol. Crystallogr. 62:48–57 10.1107/S0907444905039107 [DOI] [PubMed] [Google Scholar]
- Ly D., Kasmar A.G., Cheng T.Y., de Jong A., Huang S., Roy S., Bhatt A., van Summeren R.P., Altman J.D., Jacobs W.R., Jr, et al. 2013. CD1c tetramers detect ex vivo T cell responses to processed phosphomycoketide antigens. J. Exp. Med. 210:729–741 10.1084/jem.20120624 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mallevaey T., Scott-Browne J.P., Matsuda J.L., Young M.H., Pellicci D.G., Patel O., Thakur M., Kjer-Nielsen L., Richardson S.K., Cerundolo V., et al. 2009. T cell receptor CDR2 beta and CDR3 beta loops collaborate functionally to shape the iNKT cell repertoire. Immunity. 31:60–71 10.1016/j.immuni.2009.05.010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin E., Treiner E., Duban L., Guerri L., Laude H., Toly C., Premel V., Devys A., Moura I.C., Tilloy F., et al. 2009. Stepwise development of MAIT cells in mouse and human. PLoS Biol. 7:e54 10.1371/journal.pbio.1000054 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maru Y., Yokosuka O., Imazeki F., Saisho H., Omata M. 2003. Analysis of T cell receptor variable regions and complementarity determining region 3 of infiltrating T lymphocytes in the liver of patients with chronic type B hepatitis. Intervirology. 46:277–288 10.1159/000073207 [DOI] [PubMed] [Google Scholar]
- Matsuda J.L., Naidenko O.V., Gapin L., Nakayama T., Taniguchi M., Wang C.R., Koezuka Y., Kronenberg M. 2000. Tracking the response of natural killer T cells to a glycolipid antigen using CD1d tetramers. J. Exp. Med. 192:741–754 10.1084/jem.192.5.741 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matulis G., Sanderson J.P., Lissin N.M., Asparuhova M.B., Bommineni G.R., Schümperli D., Schmidt R.R., Villiger P.M., Jakobsen B.K., Gadola S.D. 2010. Innate-like control of human iNKT cell autoreactivity via the hypervariable CDR3beta loop. PLoS Biol. 8:e1000402 10.1371/journal.pbio.1000402 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maziarz R.T., Fraser J., Strominger J.L., Burakoff S.J. 1986. The human HLA-specific monoclonal antibody W6/32 recognizes a discontinuous epitope within the alpha 2 domain of murine H-2Db. Immunogenetics. 24:206–208 [DOI] [PubMed] [Google Scholar]
- Meierovics A., Yankelevich W.J., Cowley S.C. 2013. MAIT cells are critical for optimal mucosal immune responses during in vivo pulmonary bacterial infection. Proc. Natl. Acad. Sci. USA. 110:E3119–E3128 10.1073/pnas.1302799110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miles J.J., Elhassen D., Borg N.A., Silins S.L., Tynan F.E., Burrows J.M., Purcell A.W., Kjer-Nielsen L., Rossjohn J., Burrows S.R., McCluskey J. 2005. CTL recognition of a bulged viral peptide involves biased TCR selection. J. Immunol. 175:3826–3834 [DOI] [PubMed] [Google Scholar]
- Miyazaki Y., Miyake S., Chiba A., Lantz O., Yamamura T. 2011. Mucosal-associated invariant T cells regulate Th1 response in multiple sclerosis. Int. Immunol. 23:529–535 10.1093/intimm/dxr047 [DOI] [PubMed] [Google Scholar]
- Okamoto N., Kanie O., Huang Y.Y., Fujii R., Watanabe H., Shimamura M. 2005. Synthetic alpha-mannosyl ceramide as a potent stimulant for an NKT cell repertoire bearing the invariant Valpha19-Jalpha26 TCR alpha chain. Chem. Biol. 12:677–683 10.1016/j.chembiol.2005.04.014 [DOI] [PubMed] [Google Scholar]
- Patel O., Pellicci D.G., Uldrich A.P., Sullivan L.C., Bhati M., McKnight M., Richardson S.K., Howell A.R., Mallevaey T., Zhang J., et al. 2011. Vβ2 natural killer T cell antigen receptor-mediated recognition of CD1d-glycolipid antigen. Proc. Natl. Acad. Sci. USA. 108:19007–19012 10.1073/pnas.1109066108 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patel O., Kjer-Nielsen L., Le Nours J., Eckle S.B.G., Birkinshaw R., Beddoe T., Corbett A.J., Liu L., Miles J.J., Meehan B., et al. 2013. Recognition of vitamin B metabolites by mucosal-associated invariant T cells. Nat. Commun. 4:2142. [DOI] [PubMed] [Google Scholar]
- Pellicci D.G., Patel O., Kjer-Nielsen L., Pang S.S., Sullivan L.C., Kyparissoudis K., Brooks A.G., Reid H.H., Gras S., Lucet I.S., et al. 2009. Differential recognition of CD1d-alpha-galactosyl ceramide by the V beta 8.2 and V beta 7 semi-invariant NKT T cell receptors. Immunity. 31:47–59 10.1016/j.immuni.2009.04.018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reantragoon R., Kjer-Nielsen L., Patel O., Chen Z., Illing P.T., Bhati M., Kostenko L., Bharadwaj M., Meehan B., Hansen T.H., et al. 2012. Structural insight into MR1-mediated recognition of the mucosal associated invariant T cell receptor. J. Exp. Med. 209:761–774 10.1084/jem.20112095 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rossjohn J., Pellicci D.G., Patel O., Gapin L., Godfrey D.I. 2012. Recognition of CD1d-restricted antigens by natural killer T cells. Nat. Rev. Immunol. 12:845–857 10.1038/nri3328 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tilloy F., Treiner E., Park S.H., Garcia C., Lemonnier F., de la Salle H., Bendelac A., Bonneville M., Lantz O. 1999. An invariant T cell receptor α chain defines a novel TAP-independent major histocompatibility complex class Ib–restricted α/β T cell subpopulation in mammals. J. Exp. Med. 189:1907–1921 10.1084/jem.189.12.1907 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Treiner E., Duban L., Bahram S., Radosavljevic M., Wanner V., Tilloy F., Affaticati P., Gilfillan S., Lantz O. 2003. Selection of evolutionarily conserved mucosal-associated invariant T cells by MR1. Nature. 422:164–169 10.1038/nature01433 [DOI] [PubMed] [Google Scholar]
- Tynan F.E., Reid H.H., Kjer-Nielsen L., Miles J.J., Wilce M.C., Kostenko L., Borg N.A., Williamson N.A., Beddoe T., Purcell A.W., et al. 2007. A T cell receptor flattens a bulged antigenic peptide presented by a major histocompatibility complex class I molecule. Nat. Immunol. 8:268–276 10.1038/ni1432 [DOI] [PubMed] [Google Scholar]
- Van Damme N., De Vos M., Baeten D., Demetter P., Mielants H., Verbruggen G., Cuvelier C., Veys E.M., De Keyser F. 2001. Flow cytometric analysis of gut mucosal lymphocytes supports an impaired Th1 cytokine profile in spondyloarthropathy. Ann. Rheum. Dis. 60:495–499 10.1136/ard.60.5.495 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Rhijn I., Kasmar A., de Jong A., Gras S., Bhati M., Doorenspleet M.E., de Vries N., Godfrey D.I., Altman J.D., de Jager W., et al. 2013. A conserved human T cell population targets mycobacterial antigens presented by CD1b. Nat. Immunol. 14:706–713 10.1038/ni.2630 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walker L.J., Kang Y.H., Smith M.O., Tharmalingham H., Ramamurthy N., Fleming V.M., Sahgal N., Leslie A., Oo Y., Geremia A., et al. 2012. Human MAIT and CD8αα cells develop from a pool of type-17 precommitted CD8+ T cells. Blood. 119:422–433 10.1182/blood-2011-05-353789 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang G.C., Dash P., McCullers J.A., Doherty P.C., Thomas P.G. 2012. T cell receptor αβ diversity inversely correlates with pathogen-specific antibody levels in human cytomegalovirus infection. Sci. Transl. Med. 4:128ra42. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wei D.G., Curran S.A., Savage P.B., Teyton L., Bendelac A. 2006. Mechanisms imposing the Vβ bias of Vα14 natural killer T cells and consequences for microbial glycolipid recognition. J. Exp. Med. 203:1197–1207 10.1084/jem.20060418 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young M.H., Gapin L. 2013. Mucosal associated invariant T cells: don’t forget your vitamins. Cell Res. 23:460–462 10.1038/cr.2012.168 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young M.H., U’Ren L., Huang S., Mallevaey T., Scott-Browne J., Crawford F., Lantz O., Hansen T.H., Kappler J., Marrack P., Gapin L. 2013. MAIT cell recognition of MR1 on bacterially infected and uninfected cells. PLoS ONE. 8:e53789 10.1371/journal.pone.0053789 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zwart P.H., Afonine P.V., Grosse-Kunstleve R.W., Hung L.W., Ioerger T.R., McCoy A.J., McKee E., Moriarty N.W., Read R.J., Sacchettini J.C., et al. 2008. Automated structure solution with the PHENIX suite. Methods Mol. Biol. 426:419–435 10.1007/978-1-60327-058-8_28 [DOI] [PubMed] [Google Scholar]