Abstract
The Drosophila visual system has provided a model to study phototransduction and retinal degeneration. To identify new candidate proteins that contribute to these processes, we conducted a genome-wide screen for genes expressed predominately in the eye, using DNA microarrays. This screen appeared to be comprehensive as it led to the identification of all 22 eye-enriched genes previously shown to function in phototransduction or implicated in retinal degeneration. In addition, we identified 93 eye-enriched genes whose roles have not been previously defined. One of the eye-enriched genes encoded a member of a large family of transmembrane proteins, referred to as tetraspanins. We created a null mutation in the eye-enriched tetraspanin, Sunglasses (Sun), which resulted in light-induced retinal degeneration. We found that the Sun protein was distributed primarily in lysosomes, and functioned in a long-known but poorly understood phenomenon of light-induced degradation of rhodopsin. We propose that lysosomal tetraspanins in mammalian cells may also function in the downregulation of rhodopsin and other G-protein-coupled receptors, in response to intense or prolonged agonist stimulation.
Keywords: Drosophila, microarray, phototransduction, retinal degeneration, rhodopsin, tetraspanin
Introduction
The Drosophila visual system has emerged as a paradigm for characterizing the molecular mechanisms underlying phototransduction and retinal degeneration (reviewed in Pak, 1995; Montell, 1999). The common features between Drosophila and mammalian visual transduction include the employment of a number of key related molecules, such as rhodopsin, arrestin and heterotrimeric G-proteins. Both systems also display a high gain, rapid activation, sensitivity to light intensities that vary over many orders of magnitude and rapid adaptation (reviewed in Montell, 1999). However, in fly phototransduction, the effector of the G-protein is a phospholipase C rather than a phosphodiesterase (Bloomquist et al, 1988), and the cascade culminates with the opening of the light-sensitive cation channels, TRP, TRPL (reviewed in Montell, 1999) and TRPγ (Xu et al, 2000). Although the latter events in Drosophila visual transduction diverge from mammalian vision, they share important features with many other signaling cascades that utilize phosphoinositide signaling.
Many questions remain concerning Drosophila phototransduction, which are also relevant to mammalian phototransduction. These include the mechanisms mediating termination of signaling, adaptation and light-dependent targeting of proteins in photoreceptor cells. The molecular mechanisms underlying retinal degeneration are also incompletely understood, although Drosophila has provided important insights into the bases of certain types of retinal dystrophies. An additional phenomenon common between mammals and Drosophila is downregulation of rhodopsin in response to intense or continuous light (reviewed in Minke and Selinger, 1992; Penn and Anderson, 1992), although the proteins participating in this effect are unknown. Therefore, there remains considerable potential in using Drosophila genetics to identify additional genes that function in a variety of processes in photoreceptor cells.
Although many Drosophila loci have been characterized that function in phototransduction, or are associated with retinal degeneration, the forward genetic screens have fallen short of saturation. Therefore, in a few cases, mutations have been created in candidate retinal genes through a reverse genetics approach, which involved screening for protein nulls on Western blots (Dolph et al, 1993). During the last few years, there have been advances that have improved the ease of performing reverse genetics. These include the establishment of large collections of transposable element insertion lines, which provide the potential for generating mutations in flanking genes. Methods for homologous recombination have been developed (Rong and Golic, 2000) and an approach for effectively using RNAi in Drosophila photoreceptor cells has been described (Kalidas and Smith, 2002). Therefore, it is now feasible to perform reverse genetics more rapidly and on a larger scale than formerly possible.
A key problem with conducting large-scale reverse genetics is the establishment of reliable criteria for identifying genes likely to be associated with phototransduction or retinal degeneration without a bias toward genes already implicated to function in these processes. In an effort to do so, we have taken advantage of the observation that nearly all of the known mutations in the fruitfly that impair the visual response or cause age-dependent retinal degeneration (15 out of 16) disrupt genes whose expression is highly enriched in the eye (Hyde et al, 1990; Montell, 1999).
In the current work, we conducted a comprehensive microarray analysis to identify all of the eye-enriched genes among the 13 601 genes annotated by the Drosophila Genome Projects (Adams et al, 2000). This screen appeared to be effective as all 22 eye-enriched genes, known or strongly implicated to function in phototransduction or that are required to prevent retinal degeneration, emerged in this analysis. Furthermore, the screen led to the identification of 93 eye-enriched genes, which were not known or predicted to participate in retinal function. Among these genes was a locus encoding an eye-enriched tetraspanin, which we refer to as sunglasses (sun).
Tetraspanins consist of four transmembrane segments and comprise a large family in flies and humans, although the functions of most tetraspanins are unknown (Maecker et al, 1997; Todres et al, 2000). Among the 37 tetraspanins encoded in the Drosophila genome, a function has been unambiguously ascribed to only one, late-bloomer, which participates in synapse formation (Kopczynski et al, 1996). We found that a null mutation in sun caused light-dependent retinal degeneration. In contrast to all other proteins known to be associated with retinal degeneration, Sun was enriched in the lysosomes, suggesting that it functions in protein degradation. One protein, which could potentially undergo Sun-regulated degradation, is rhodopsin as exposure to intense or continuous light results in a decrease in rhodopsin concentration; however, the proteins mediating this effect have not been described. Here, we report that Sun is required for efficient light-induced downregulation of rhodopsin. Given that there are mammalian lysosomal associated tetraspanins (Fukuda, 1991), we propose that light-dependent downregulation of mammalian rhodopsin (Penn and Anderson, 1992) may be dependent on a tetraspanin related to Sun.
Results
Comprehensive screen for eye-enriched genes
To identify new candidate loci required for the physiology of photoreceptor cells, we conducted a comprehensive, genome-wide screen for eye-enriched genes in Drosophila using DNA microarrays. To perform this analysis, we isolated RNA from the heads of wild-type (wt) flies, from the heads of an eyeless mutant (sine oculis; so) and from wt bodies. Biotinylated cRNAs were probed with microarrays corresponding to the complete set of 13 601 genes cataloged by the Drosophila Genome Project.
A total of 128 genes were identified in the microarray analysis (wt head/so head ratios ⩾2.0), 93 of which were not previously known to be eye enriched. In all, 78 of these genes had not been characterized previously either at the molecular or genetic level (Table 1; Supplementary Table 1), and 14 corresponded to known genes that had not been analyzed in the context of retinal degeneration or phototransduction (Table 1, indicated by a). It was likely that most or all of the eye-enriched genes annotated in the Drosophila genome were identified since all 22 genes that were previously known to be eye enriched and which have been implicated in phototransduction/retinal degeneration were detected in this analysis (Table 2). Only one gene previously associated with retinal degeneration or phototransduction, rdgB, displayed a minor level of eye enrichment (wt head/so head ratio=1.3), consistent with prior studies (Vihtelic et al, 1991). In addition, 13 genes corresponded to loci known to function in processes in the eye other than phototransduction (Supplementary Table 2). Comparison of the chromosomal map positions of the 128 eye-enriched genes did not reveal significant clustering in the genome. The greatest concentration of eye-enriched genes consisted of 11, which spanned 92B2-94F14 on the third chromosome.
Table 1.
Classes of genes identified by the microarray analysis that were not previously known to be eye enriched
| Gene | Map | Description | Ratio | REL |
|---|---|---|---|---|
| Redox control/detoxification | ||||
| CG6781 | 66D4 | Glutathione transferase | 12.1±2.8 | 209.6±63.3 |
| Eip71CDa | 71C4 | Protein methionine reductase | 3.7±0.1 | 916.2±33.4 |
| CG11765 | 47A6 | Glutathione peroxidase | 2.6±0.3 | 1553.9±356.2 |
| Intermediary metabolism | ||||
| CG7077 | 94A14 | Short-chain dehydrogenase | 42.8±4.9 | 873.1±309.4 |
| CG13356 | 20B1 | Short-chain dehydrogenase | 23.5±4.9 | 473.4±86.4 |
| CG9150 | 26B3 | Short-chain dehydrogenase | 6.9±1.3 | 121.5±39.8 |
| CG6870 | 36C7 | Heme-binding protein | 9.5±0.8 | 2650.9±925.6 |
| RPE65a | 87F10 | B-carotene 15,15′-dioxygenase | 2.6±0.3 | 154.4±11.6 |
| CG7059 | 94B1 | Phosphoglycerate mutase | 2.0±0.2 | 83.0±21.5 |
| Post translational modifications | ||||
| Kinase/phosphatase | ||||
| CG8889 | 48E2 | Ser/Thr phosphatase | 10.7±1.4 | 192.3±162.8 |
| CG2049b | 45C1 | Ser/Thr kinase | 5.0±1.3 | 84.9±14.0 |
| Fps85Da | 85D11 | Protein tyrosine kinase | 2.3±0.4 | 102.0±10.4 |
| Ubiquitination | ||||
| CG1839 | 12E1 | F-box containing protein | 5.6±0.4 | 149.1±37.1 |
| CG5798 | 93C1 | Ubiquitin carboxyl hydrolase | 4.3±0.6 | 285.5±43.1 |
| CG7220 | 47B7 | Ubiquitin conjugating enzyme | 3.1±0.3 | 1399.7±113.6 |
| Other modifications | ||||
| CG9171 | 26A2 | Acetylglucosaminyltransferase | 5.3±0.7 | 72.6±5.8 |
| Unda | 30D1 | Methionyl aminopeptidase | 3.2±0.9 | 245.9±18.8 |
| CG5027 | 72D10 | Disulfide isomerase | 2.0±0.2 | 146.3±11.2 |
| Lipid metabolism | ||||
| CG12120 | 8D4 | Weakly related to acyl-trans. | 47.3±5.6 | 783.8±26.3 |
| CG17352 | 66C8 | LDL receptor | 38.0±17.3 | 926.6±318.3 |
| CG11440 | 79E4 | PA-phosphatase | 15.7±3.8 | 241.3±47.5 |
| CG15151 | 36D3 | Insulin like PLA2 inhibitor | 3.3±0.8 | 75.5±11.0 |
| CG9245 | 13E14 | CDP-DAG inositol-3.P. trans. | 2.4±0.1 | 1001.9±49.9 |
| Protease and protease inhibitors | ||||
| CG12955 | 51E9 | Serine protease inhibitor | 5.3±0.8 | 92.0±3.5 |
| CG17739 | 48F8 | Trypsin inhibitor | 3.2±0.2 | 775.9±45.1 |
| CG4948 | 66F4 | Serine-type endopeptidase | 2.3±0.1 | 662.1±62.4 |
| Synaptic function | ||||
| Sloba,b | 28B1 | Slowpoke-binding protein | 8.8±0.8 | 776.8±129.0 |
| CG6656 | 93F9 | Histidine acid phosphatase | 7.8±0.5 | 152.8±9.8 |
| CG14723 | 86F9 | Histamine-gated Cl- channel | 4.4±1.5 | 86.2±9.2 |
| CG9935 | 102D | Glutamate receptor | 5.1±0.5 | 70.6±2.1 |
| Hka | 9B7 | Voltage-gated K+ channel | 2.3±0.4 | 313.5±46.0 |
| Eaat2a | 21D1 | Na+/glutamate cotransporter | 2.2±0.1 | 649.6±121.8 |
| CG7411 | 92B1 | Histamine-gated Cl- channel | 2.0±0.6 | 68.7±20.7 |
| Vesicle transport | ||||
| RabRP1a,b | 45B3 | Small GTPase | 4.3±0.1 | 1119.7±39.4 |
| Rab14a | 35A1 | Small GTPase | 2.1±0.1 | 805.7±140.5 |
| Signaling | ||||
| CG10233 | 83A1 | Related to junctophilin | 146.7±12.0 | 2506.2±221.0 |
| Cbp53Ea | 53E4 | Ca2+-binding protein | 2.8±1.2 | 345.2±79.6 |
| CG7823 | 76D5 | Rho GDI | 2.1±0.4 | 470.3±139.4 |
| Apoptosis | ||||
| CG10257 | 51D7 | Anti-FAS apoptosis inhibitor | 3.7±1.4 | 61.4±8.2 |
| Transporters | ||||
| CG15891 | 5E1 | Related H+/cation antiporter | 9.0±2.5 | 201.3±51.5 |
| CG3790 | 49D3 | Organic cation transporter | 8.0±1.3 | 152.8±25.7 |
| Structural/morphological maintenance | ||||
| CG18066 | 57A8 | Insect cuticle protein | 25.1±0.5 | 419.8±51.4 |
| CG12143 | 42E1 | Tetraspanin | 13.2±2.4 | 1143.7±108.4 |
| CG4784b | 72F1 | Cuticle protein | 8.0±1.8 | 401.4±35.6 |
| GH11322 | 24E4 | C-type immunoglobulin | 2.8±0.1 | 646.4±63.3 |
| CG8511 | 49A3 | Cuticle protein | 4.5±0.6 | 64.2±19.5 |
| CG7106 | 28D2 | C-type lectin | 2.1±0.3 | 73.6±10.9 |
| Transcription factors/nucleic acid-binding proteins | ||||
| CG2016 | 82E7 | Takeout like protein | 25.7±1.6 | 432.0±68.1 |
| CG11820 | 96B16 | Transcription factor | 11.2±1.8 | 176.4±33.2 |
| Mstaa | 2E2 | BOP zinc finger protein | 10.6±1.5 | 896.6±27.4 |
| Pph13 | 21C6 | Pol. II transcription factor | 9.8±1.5 | 148.0±11.7 |
| CG10446 | 37B10 | Transcription factor | 7.3±3.0 | 122.5±21.4 |
| CG2650 | 3B4 | Circadian clock gene | 7.8±2.3 | 131.2±59.7 |
| SP2523 | 66A8 | Related to RNA helicase | 4.6±0.6 | 541.5±151.5 |
| mre11a | 32E2 | DNA repair protein | 3.7±0.5 | 130.4±17.1 |
| Rbp9a | 23C1 | RNA-binding protein | 3.3±0.6 | 365.0±20.2 |
| CG6866 | 34B9 | RNA-binding protein | 2.3±0.3 | 156.6±22.2 |
| Others with predictable functions | ||||
| CG13409 | 94A1 | Biotin synthetase | ||
| CG8483 | 87E2 | Related to venom allergen5 | 3.4±0.1 | 195.8±22.8 |
| CHIPa | 31E2 | Tetratricopeptide repeat | 2.1±0.2 | 206.1±38.6 |
| CG12020 | 62B4 | Heat-shock protein | 2.0±0.2 | 48.7±10.7 |
| No recognizable domains (homologs in a diversity of organisms) | ||||
| CG13057 | 72E4 | D, M | 491.5±67.5 | 8834.3±2357.0 |
| CG9427 | 85E1 | C, D, M | 9.9±2.9 | 300.1±53.4 |
| CG6983 | 66D1 | D, M, H | 7.3±1.4 | 107.0±21.5 |
| CG4686 | 92B6 | A, C, D, M, H | 2.9±0.3 | 734.9±85.3 |
| CG14464 | 41A1 | H | 2.3±0.4 | 243.9±94.5 |
| CG4306 | 75C6 | M, H | 2.2±0.2 | 53.4±8.2 |
| No recognizable domains (invertebrate homologs only) | ||||
| CG1561 | 10C5 | C, D | 93.1±30.6 | 1596.7±63.0 |
| CG7135 |
16F7 |
C, D |
3.6±1.0 |
100.9±13.1 |
| Minimum wt head/so head ratio is 2.0. The relative expression levels (RELs) were obtained by subtracting the expression level in so heads from that in wt (Canton S) heads. The standard errors (SEs) are shown (n=3). | ||||
| a Genes that have been previously described, but not previously reported to be eye enriched or known to function in visual transduction. | ||||
| b Genes that are regulated during the circadian cycle (Claridge-Chang et al, 2001; McDonald and Rosbash, 2001). | ||||
| A: A. thaliana; C: C. elegans; D: D. melanogaster; H: H. sapiens; M: M. musculus. | ||||
| No recognizable domains and no homologs—24 genes (Supplementary Table 1). | ||||
Table 2.
Ratios of genes known or implicated in retinal degeneration or phototransduction
| Gene | Affymetrix ratio |
REL | |
|---|---|---|---|
| wt head/so head | wt head/wt body | ||
| trp | 172.2±31.3 | 232.0±67.6 | 2941.4±463.8 |
| Gβe | 137.6±9.9 | 182.5±33.6 | 2346.0±198.9 |
| Rh4 | 135.2±18.5 | 140.0±10.5 | 2261.9±86.3 |
| trpl | 116.3±20.2 | 209.7±39.5 | 2996.0±280.4 |
| ninaC (myosin III) | 86.5±2.9 | 131.6±32.7 | 1712.5±87.0 |
| arrestin1 | 84.9±28.7 | 112.5±9.6 | 11269.8±2103.9 |
| inaD (PDZ protein) | 80.6±12.4 | 104.4±26.5 | 1335.5±31.2 |
| arrestin2 | 68.7±22.8 | 130.0±24.3 | 7823.6±1568.7 |
| ninaE (Rh1) | 67.2±31.3 | 42.3±3.17 | 14592.7±3191.9 |
| Rh3 | 54.7±11.7 | 53.5±11.0 | 3125.1±389.0 |
| Rh2 | 52.3±4.4 | 65.7±10.7 | 880.6±59.3 |
| Rh5 | 49.7±11.5 | 23.6±2.3 | 1935.4±36.7 |
| inaF | 46.0±1.8 | 57.6±8.8 | 819.0±106.8 |
| Gqα | 27.5±5.7 | 41.4±9.6 | 577.6±47.9 |
| ninaA (Rh chaperone) | 24.9±8.4 | 26.9±3.2 | 754.4±126.0 |
| inaC (PKC) | 23.6±3.9 | 33.0±11.1 | 393.3±32.8 |
| Rh6 | 23.2±7.3 | 78.6±33.5 | 2120.2±152.8 |
| norpA (PLC) | 12.1±2.3 | 36.4±10.0 | 426.0±46.7 |
| Gγ30A | 12.0±2.7 | 48.3±15.1 | 2041.8±941.2 |
| rdgC (Rh phosphatase) | 6.2±1.7 | 14.1±6.4 | 134.5±27.9 |
| rdgA (DAG kinase) | 2.8±0.5 | 3.9±0.5 | 107.7±46.2 |
|
cds |
2.6±0.2 |
7.5±0.9 |
250.0±42.5 |
| The expression ratios and RELs are presented along with the SEs. | |||
To provide additional evidence to support the GeneChip data, we performed Northern blots and in situ hybridizations to adult heads using probes corresponding to selected genes. We found that there was strong agreement between the Northern blot, in situ hybridizations and microarray data (Figures 1A and B; Supplementary Table 3). Thus, the current microarray analysis appears to have identified all of the eye-enriched genes annotated by the genome project. This was in contrast to a microarray analysis in which a subset of Drosophila genes (29.6%) was surveyed for targets of the transcription factor Eyes-Absent (EYA) (Arbeitman et al, 2002). Among those known eye-enriched genes, which were included on their GeneChips, over half were missed in their analysis (for more details, see the legend to Supplementary Table 2). Although this study purports to identify eye-enriched genes, EYA is also abundantly expressed in the ovary and affects female fertility (Bonini et al, 1998). As RNA was prepared from whole flies, rather than isolated heads, many of the identified genes (Arbeitman et al, 2002) were likely to be regulated by EYA in the ovary rather than in the eye. In fact, four of the EYA targets are bodyenriched and only five appear on the list of novel eye-enriched genes identified in the current analysis (see the legend to Supplementary Table 2).
Figure 1.
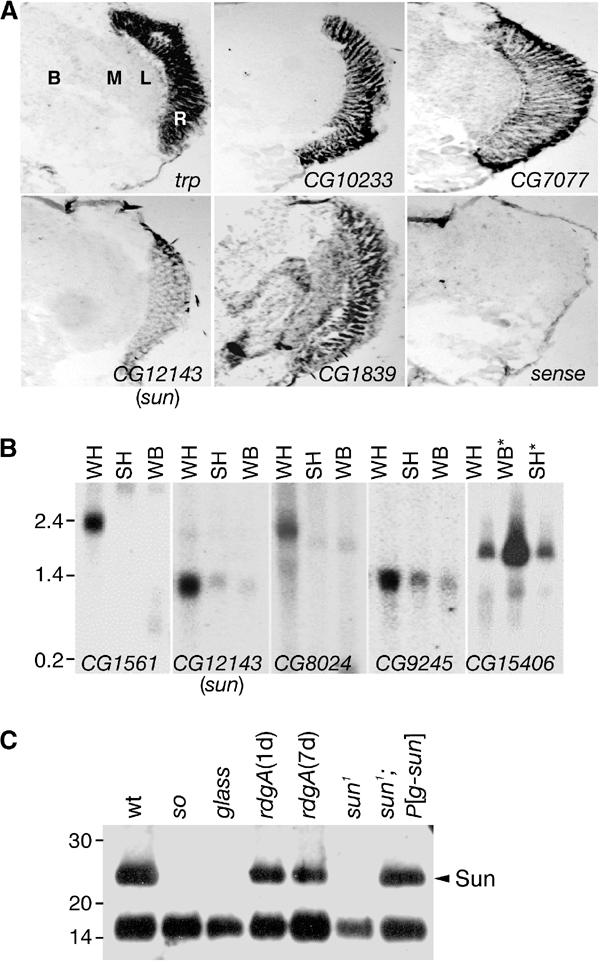
Expression patterns of sun and novel eye-enriched genes. (A) Spatial distributions of selected eye-enriched mRNAs in adult heads. In situ hybridizations to frontal sections were performed with antisense RNAs, except for the trp sense control (sense). The Flybase ID numbers are indicated. B, brain; L, lamina; M, medulla; R, retina. (B) Expression of sun and selected eye-enriched mRNAs by Northern blot analysis. Lanes contained mRNAs prepared from wt fly heads (WH), so heads (SH) and wt bodies (WB). CG15406 was predicted to be body enriched, based on the GeneChip data (Supplementary Table 3). *The order of the SH and WB samples probed with the CG15406 DNA was reversed relative to the other samples. The filters were reprobed with rp49, demonstrating that the RNA loading in each lane was similar (data not shown). (C) The Sun protein was enriched in photoreceptor cells. Anti-Sun antibodies were used to probe a Western blot containing Triton X-100 soluble proteins extracted from the indicated 1-day-old flies (except for 7-day-old rdgA). The lower 15 kDa protein of unknown identity cross-reacted with the anti-Sun antibodies. This band provided an internal loading control.
Classifications of eye-enriched genes
The 93 eye-enriched genes identified here encode multiple classes of proteins whose roles in visual physiology were either poorly understood or completely unknown (Table 1). Examples include proteins, which may have roles in detoxification, such as a glutathione transferase-related protein and a methionine sulfoxide reductase. In addition, there were three short-chain dehydrogenase-like proteins, which might function either in detoxification or synthesis of the retinal. At least nine genes encode proteins that appear to function in post-translational modifications ranging from phosphorylation to ubiquitination, removal of N-terminal methionines, glycosylation and lipid modification. Three eye-enriched genes were related to proteins that regulate proteolysis. These included two protease inhibitor-like proteins and one protein related to serine-type proteases. Five novel eye-enriched proteins may function in lipid metabolism, several of which may regulate the production of lipid second messengers. This latter group includes a phosphatidic acid phosphatase, a CDP-DAG-inositol 3-phosphatidyltransferase and a protein related to a phospholipase A2 inhibitor. In addition, there was a protein related to the LDL receptor. One of the most highly eye-enriched genes encoded a protein related to junctophilin, a protein that has been implicated in Ca2+ signaling (Takeshima et al, 2000). Other examples of novel eye-enriched genes include two uncharacterized Rab-related proteins, which may function in protein or vesicle transport. There were at least seven genes, which were not formerly known to be eye enriched, that encoded proteins likely to have roles in synaptic function. In addition, there were two eye-enriched genes encoding cation transporters.
Retinal degeneration in sun1 flies
Included among the eye-enriched genes was a member of a large family of tetraspanins (Todres et al, 2000). Given that the functions of many such proteins are poorly characterized, we set out to determine the phenotype of disrupting the eye-enriched tetraspanin, which we refer to as sunglasses (sun). As with other tetraspanins, Sun is characterized by four predicted transmembrane segments, a large extracellular domain between the third and fourth transmembrane segments and key residues conserved among most tetraspanins (Figure 2A) (Maecker et al, 1997). The tetraspanin most similar to Sun was a mammalian lysosomal protein, CD63/LAMP3/LIMP1 (Fukuda, 1991) (Figure 2A). Consistent with the microarray data, we found, on the basis of Northern blots (Figure 1B; Supplementary Table 3; CG12143) and in situ hybridizations (Figure 1A; CG12143), that the sun RNA was concentrated in the retina.
Figure 2.
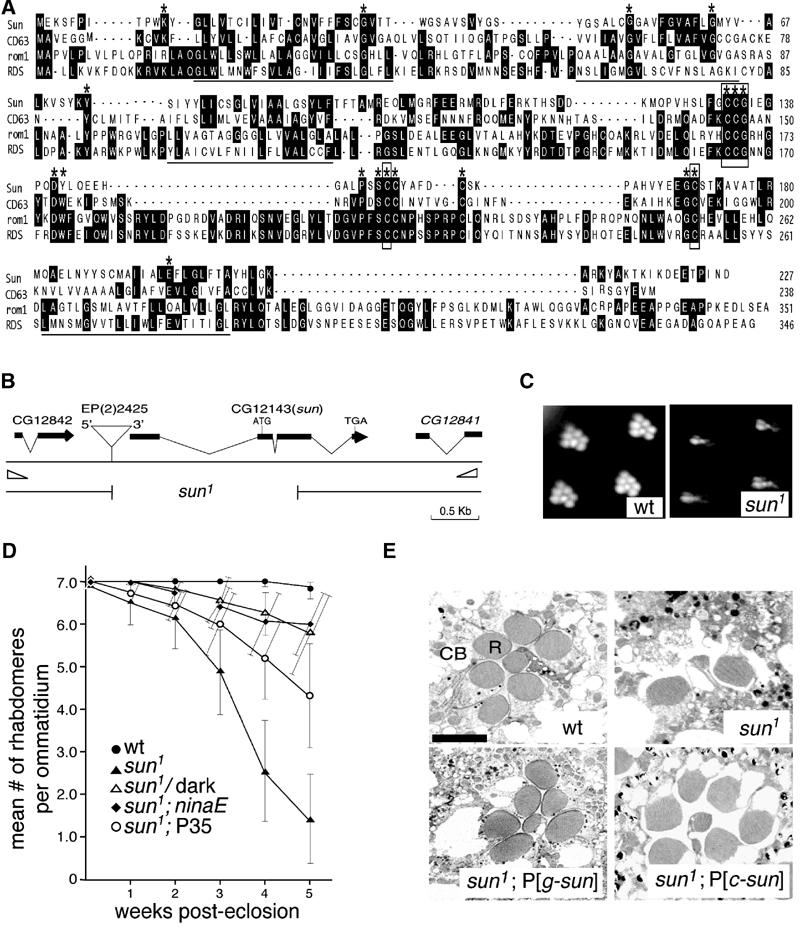
sun gene and sun1 retinal degeneration phenotype. (A) Alignment of Sun with other tetraspanins. The putative transmembrane segments are underlined. *Residues that are ⩾80% identical among all tetraspanins. Invariant residues are boxed. (B) sun gene. The two open triangles at the left and right ends indicate the locations of the primers used to screen for deletions generated by imprecise excision of the P-element, EP(2)2425. The 1.8 kb deletion in sun1 is indicated. (C) Rhabdomeres from 4-week-old wt and sun1 flies observed using the optica neutralization technique (Franceschini and Kirschfeld, 1971). The flies were reared at 25°C under a 12 h light/12 h dark cycle. (D) Mean number of rhabdomeres/ommatidium as a function of age. The flies were reared at 25°C under a 12 h light/12 h dark cycle, except for the dark-reared sun1 flies. The number of rhabdomeres was determined by the optical neutralization technique. Each data point was based on examination of ⩾90 ommatidia from ⩾8 flies. Error bars represent the SDs. (E) Retinal morphology of 4-week-old flies examined by transmission EM. Tangential sections were obtained from the distal region of the compound eye. Sections from wt, sun1, the genomic transgene in a sun1 background (sun1;P[g-sun]) and the myc-sun cDNA transgene in a sun1 background (sun1;P[c-sun]) are shown.
To obtain a mutation in sun, we mobilized a P-element, which inserted near the 5′ end of sun (Figure 2B), since the original element did not exhibit any visual physiology defect (data not shown). To identify a deletion of sun, we screened pools of genomic DNA by PCR (see Materials and methods and Figure 2B). One excision line (sun1) contained a 1.8 kb deletion, which removed nearly half of the region encoding Sun (Figure 2B). We raised anti-Sun antibodies and found that expression of the Sun protein was eliminated in sun1 and restored in mutant flies containing the sun genomic DNA transgene, P[g-sun] (Figure 1C).
We examined whether sun1 flies exhibited retinal degeneration using an assay on intact fly heads, the optical neutralization technique (Figure 2C) (Franceschini and Kirschfeld, 1971). The fly compound eye consists of ∼800 ommatidial units, each of which contains eight photoreceptor cells, although only seven are present in any given plane of section. The photoreceptor cells include a microvillar structure, the rhabdomere, which is the site of phototransduction. Flies were either maintained under a normal light/dark cycle or kept in the dark, and the average number of rhabdomeres per ommatidium was assessed. In wt ommatidia, seven rhabdomeres were detected regardless of the age of the flies or light conditions (Figures 2C and D).
We found that sun1 flies exhibited light- and age-dependent retinal degeneration. In mutant flies that were incubated in the dark, there was relatively little reduction in rhabdomere number as the flies aged (Figure 2D). However, sun1 flies maintained under a light/dark cycle exhibited a gradual loss of rhabdomeres (Figure 2D). To confirm these data, we used transmission EM. While sections of 4-week-old wt ommatidia displayed seven rhabdomeres, few (0–4 rhabdomeres/ommatidia) were detected in sun1 exposed to a light/dark cycle (Figure 2E). In contrast to most known retinal degeneration mutants in Drosophila, which display the initial defects in the rhabdomeres, the degeneration in sun photoreceptor cells was first evident in the cell bodies (data not shown). The full complement of rhabdomeres was restored in sun1 flies containing the sun genomic or cDNA transgenes (Figure 2E). The retinal degeneration was significantly suppressed either in a genetic background (ninaEP332), which greatly reduced the concentration of the major rhodopsin, or upon introduction of a transgene that expressed Baculovirus P35, a broad-spectrum caspase inhibitor (Figure 2D).
Sun is a lysosomal-enriched protein
To address the function of Sun, we examined the expression pattern of the protein, initially using a biochemical approach. The Sun protein appeared to be expressed predominantly in photoreceptor cells, as it was eliminated in a mutant, glass, which does not form photoreceptor cells (Figure 1C). Sun was present in rdgA flies, which were devoid of rhabdomeres due to degeneration (Figure 1C), indicating that a significant proportion of the Sun pool was present in the cell bodies. However, Sun did not appear to be specific to the cell bodies as there was a small but reproducible decrease in Sun levels in rdgA. The anti-Sun antibodies did not recognize Sun in sections of compound eyes. Therefore, to examine the subcellular distribution of Sun, we generated transgenic flies expressing a Myc-tagged version of Sun (P[w+;c-sun]). The Myc-Sun protein was functional in vivo as it largely rescued the Sun mutant phenotype (Figure 2E). Consistent with the Western blot data, the anti-Myc staining was detected primarily in a punctate pattern in the cell bodies (Figures 3A–C). A small amount of labeling was detected near the base of some rhabdomeres (Figure 3C). Only a low level of background staining resulted from using anti-Myc antibodies on sections of nontransgenic flies and this staining was primarily associated with the extracellular central matrix (Figures 3D–F).
Figure 3.
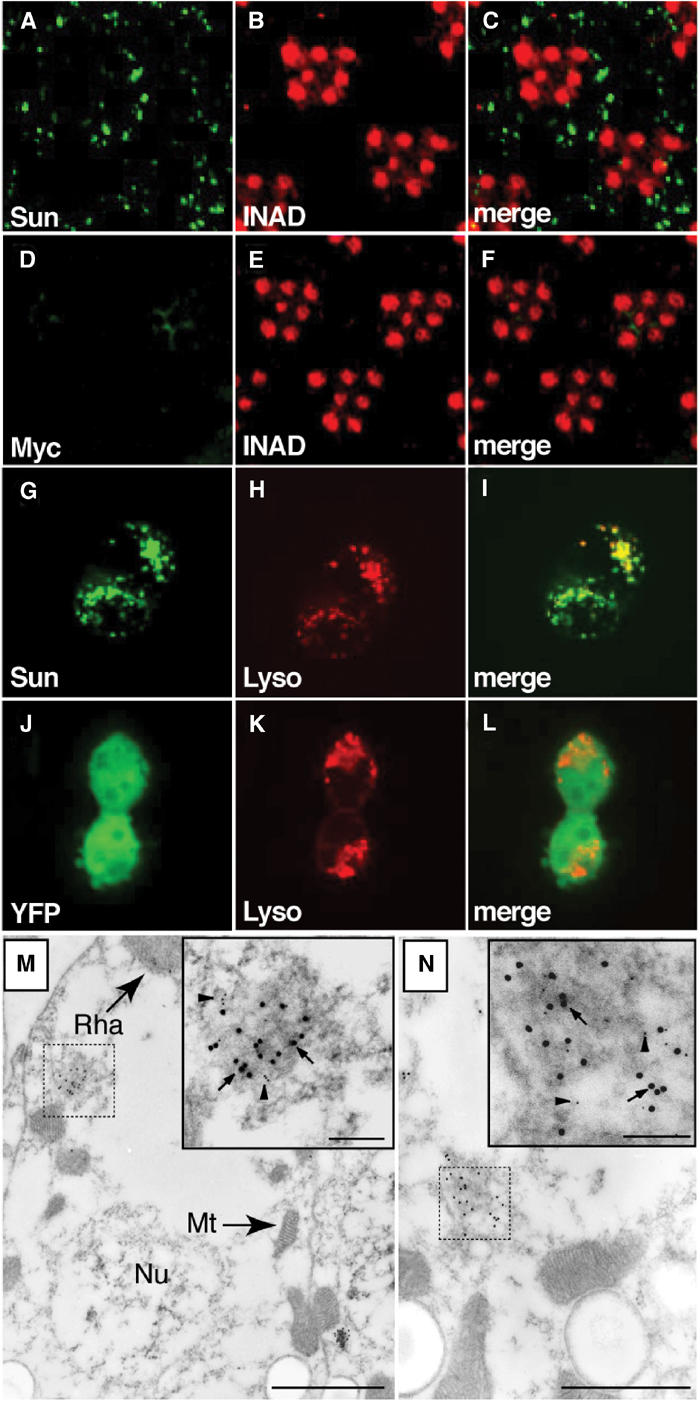
Immunolocalization of Sun. (A–F) Fly heads were embedded in LR White and 0.5 μm cross-sections of compound eyes were stained with rabbit anti-Myc and a rhabdomeral marker, rat anti-INAD. (A) Section from myc-sun transgenic fly, P[w+;c-sun];cn bw, stained with rabbit anti-Myc polyclonal antibodies. (B) Same section as in (A) stained with rat anti-INAD antibodies. (C) Merged images from (A, B). (D) Section from a white-eyed (cn bw) nontransgenic fly stained with anti-Myc antibodies. (E) Same section as in (D) stained with anti-INAD. (F) Merged images from (D, E). (G–I) 293 cells transfected with a plasmid encoding a YFP-Sun fusion protein. (G) Localization of YFP. (H) Same cells as in (J) stained with LysoTracker. (I) Merged images from (G, H). (J–L) 293 cells transfected with a plasmid encoding YFP. (J) Localization of YFP. (K) Same cells as (G) stained with LysoTracker. (L) Merged images from (J, K). (M, N) Immuno-EM localization of Myc-Sun and Rh1 in the cell bodies of P[c-sun];cn bw photoreceptor cells. Sections were stained with anti-Myc rabbit polyclonal antibodies (15 nm gold particles; several are indicated by arrows) and anti-Rh1 mouse monoclonal antibodies (5 nm gold particles, several are indicated by arrowheads). The dashed and solid boxes show lower and higher magnification views of Myc-Sun and Rh1 positive lysosomal-like vesicles (M) and MVB-like vesicles (N). The small (inset) and large-scale bars represent 0.1 and 1.0 μm, respectively. Mt, mitochondrion; Nu, nucleus; Rha, rhabdomere.
Immunolocalization at the EM level confirmed that the anti-Myc staining was within vesicles in the cell bodies. No immunoreactivity was detected in the nucleus or mitochondria (Figures 3M and N). Given that the tetraspanin most related to Sun, CD63, is a lysosomal protein, we considered whether Sun is also present in lysosomes. However, it is not possible to unequivocally identify lysosomes in these preparations due to an absence of multilamellar structures after the fixation and staining protocol. Nevertheless, the Myc-Sun positive vesicles appeared as dense bodies, reminiscent of structures previously described as lysosome-like and multivesicular body (MVB)-like (Suzuki and Hirosawa, 1991). To address whether Sun was localized in lysosomes, we expressed a YFP-tagged version of Sun in tissue culture cells. In contrast to the wide distribution of YFP alone (Figures 3J–L), YFP-Sun displayed a punctate pattern that colocalized with a lysosomal/late endosomal marker, LysoTracker (Figures 3G–I). There was also significant colocalization of Sun with the lysosomal protein LAMP1 (Supplementary Figures 1A–C). A small proportion of the YFP-Sun pool colocalized with an endosomal marker, RhoB (Supplementary Figures 1D–F). Additional evidence that Sun is primarily a lysosomal protein is presented below.
Defects in light-dependent turnover of rhodopsin in sun1
The spatial distribution of Sun suggested that it might be involved in the turnover of a photoreceptor cell protein. Therefore, we compared the concentrations of a variety of proteins in wt and sun1 heads. Young flies were used in these experiments, which did not exhibit any significant age- and light-dependent degeneration due to the sun1 mutation. The flies were either maintained in ambient light or exposed for 6 h to bright blue light, which stably converts rhodopsin to the light-activated metarhodopsin. We found that the levels of the photoreceptor cell enriched proteins, TRP, INAD and the rhodopsin binding protein Arrestin2, were similar in wt and sun1 flies, regardless of the light conditions (Figure 4A).
Figure 4.
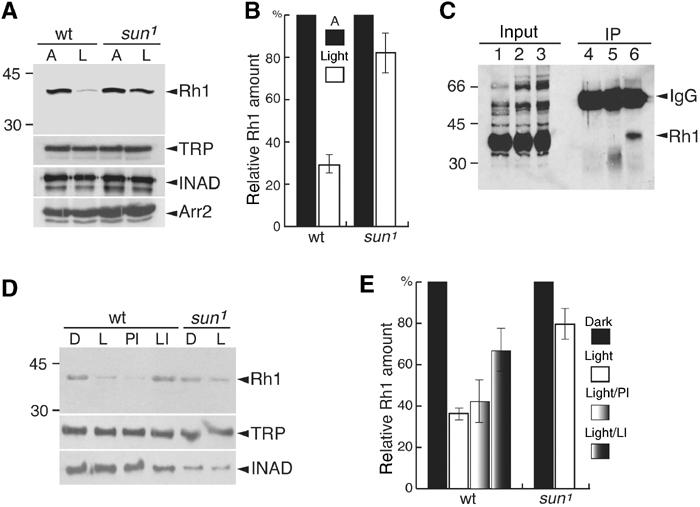
Defect in blue light-induced lysosomal degradation of Rh1 in sun1. (A) Rh1 levels in wt and sun1 flies. Three-day-old flies were either kept under ambient light (A) or stimulated for 6 h with blue light (L). Head extracts were prepared and a Western blot was probed with anti-Rh1 antibodies and 125I-labeled-anti-mouse IgG. Parallel Western blots, using the same samples, were probed with anti-TRP, anti-INAD and anti-Arr2 antibodies. (B) Quantitation of relative Rh1 levels from (A). Band intensities were determined using a phosphoimager. (C) Rh1 co-immunoprecipitated (co-IPed) with Sun in vivo. Head extracts were prepared from wt or sun1. IPs were performed with anti-Sun antibodies or preimmune serum and a Western blot was probed with anti-Rh1 antibodies. A 1% input in lanes 1–3 corresponds to lanes 4–6, respectively. Lanes: 4, IP with anti-Sun and sun1 extracts; 5, IP with preimmune serum and wt extracts; 6, IP with anti-Sun and wt extracts. In some but not all experiments with nonimmune serum, an ∼35 kDa band was detected. This band was not detected with anti-Sun antibodies. (D) A lysosomal inhibitor suppressed blue light-induced degradation of Rh1. Isolated wt or sun1 ommatidia were maintained in the dark (D) or exposed to blue light for 4 h (L). Prior to the light treatment, some of the ommatidia were transferred to media containing either a lysosomal inhibitor (LI; 10 μg/μl leupeptin) or a proteasomal inhibitor (PI; 10 μM lactacystin). The Western blot was probed with anti-Rh1 antibodies and 125I-labeled-anti-mouse IgG. Parallel Western blots containing the same samples were probed with antibodies that recognize the retinal-enriched proteins INAD and TRP. (E) Quantitation of the relative Rh1 levels from (D). The relative intensities of the bands were determined using a phosphoimager.
The major rhodopsin (Rh1) was an excellent candidate for degradation through a Sun-dependent pathway since it is the only photoreceptor cell protein reported to undergo downregulation in response to continuous bright light (reviewed in Minke and Selinger, 1992). In wt photoreceptor cells challenged for 6 h with blue light, the level of rhodopsin was greatly reduced (Figures 4A and B; 31±5% of wt levels). This reduction in Rh1 levels was similar in the arrestin2 mutant (data not shown), which was unexpected given that downregulation of G-protein-coupled receptors (GPCRs) typically occurs through an arrestin-dependent mechanism (Claing et al, 2002). This suggested that there was a mechanism for bright light-dependent downregulation of Rh1, in addition to clathrin-mediated endocytosis of Rh1/arrestin complexes (Alloway et al, 2000; Kiselev et al, 2000). Of primary significance here, the reduction in Rh1 concentration resulting from exposure to continuous bright blue light was suppressed to a significant extent in sun1 (Figures 4A and B; 82±18%). The combination of these data suggests that Sun participates in the turnover of Rh1 through promoting its degradation in lysosomes.
We found that most of the Rh1 pool in the cell bodies colocalized with the Sun-containing vesicles in photoreceptor cells (Figures 3M and N). Evidence that the Rh1-containing vesicles in photoreceptor cells are lysosomes is presented below. Rh1 appeared to associate with Sun in vivo as Rh1 was detected in immune complexes after performing immunoprecipitations (IPs) with anti-Sun antibodies (Figure 4C). Another abundant membrane protein, TRP, did not co-IP with Sun (data not shown). We did not detect Rh1 after performing IPs with anti-Sun from sun1 extracts or after using preimmune serum and wt extracts (Figure 4C).
Light-dependent lysosomal localization of Rh1 in isolated ommatidia
To characterize the mechanism through which Sun contributes to light-dependent degradation of Rh1 in greater detail, we attempted to recapitulate the phenomenon in isolated ommatidia. To facilitate this analysis, we examined the relative concentrations and localization of GFP-Rh1 in young wt and sun1 photoreceptor cells, before and after exposure of the isolated ommatidia to bright blue light. After exposing isolated ommatidia to blue light for 4 h, the concentration of rhodopsin was decreased (Figures 4D and E).
To examine the effects of light on the distribution of GFP-Rh1, we examined GFP staining in ommatidia after a 1.5 h exposure to blue light, prior to when there is a dramatic decrease in the overall concentration of GFP-Rh1. In ommatidia incubated in the dark, the GFP-Rh1 was present almost exclusively in the rhabdomeres (Figure 5A). After the blue light exposure, GFP-Rh1 was also detected in many puncta in the cytoplasm (Figure 5B). Many of these puncta corresponded to lysosomes as there was significant overlap between GFP-Rh1 and LysoTracker (Figures 5B–E). Since after exposure to bright light GFP-Rh1 is detected in the cell bodies, primarily in lysosomes, and since Sun and Rh1 colocalize in vesicles (Figures 3M and N), these data further support the conclusion that Sun is a lysosomal protein. Additional evidence that lysosomes participated in the bright light-induced degradation of Rh1 was that the light-dependent degradation of Rh1 was significantly suppressed by a lysosomal inhibitor, but unaffected by a proteasomal inhibitor (Figures 4D and E).
Figure 5.
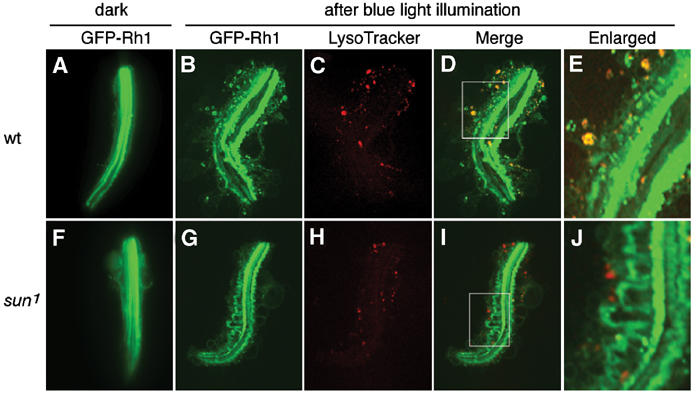
Light-dependent translocation of Rh1 in wt but not sun1 isolated ommatidia. (A–E) Ommatidia from wt expressing a GFP-Rh1 transgene. (F–J) Ommatidia from sun1 expressing a GFP-Rh1 transgene. (A, F) GFP-Rh1 staining (green) in ommatidia maintained in the dark. (B, G) Ommatidia incubated in media containing LysoTracker and exposed to blue light for 1.5 h. The GFP-Rh1 expression is shown (green). (C, H) LysoTracker staining (red) in the same ommatidia shown in (B) and (G), respectively. (D, I) Merge of staining shown in (B, C) and (G, H), respectively. (E, J) Higher magnifications of regions indicated by boxes in (D, I), respectively.
To determine whether there was a decrease in the localization of Rh1 in sun1 lysosomes, we examined the distribution of LysoTracker and GFP-Rh1 in ommatidia isolated from young sun1 flies, prior to any apparent retinal degeneration. The sun1 and wt photoreceptor cells contained a similar number of lysosomes, indicating that Sun was not required for lysosomal biogenesis (Supplementary Figures 2). Of significance here, we found that the overlap between LysoTracker and GFP-Rh1 was significantly reduced in sun1 (Figures 5G–J), consistent with the conclusion that Sun facilitates targeting or entry of Rh1 into lysosomes. In sun1 photoreceptor cells, the GFP-Rh1 staining was also detected in an extensive array of large loops emanating from the rhabdomeres (Figures 5G, I and J). Such loops, which were fewer in number and much smaller in wt (Figure 5E), appeared to correspond to invaginations of the rhabdomeral membrane, which are dynamically produced in wt in a light-dependent manner (Kosloff et al, 2003).
Discussion
Comparison of eye-enriched genes in flies and mammals
The prime motivation for performing the microarray study was to identify candidate loci that participate in phototransduction or which would provide new insights into retinal dystrophies. In support of the efficacy of the microarray analysis, all 22 of the known eye-enriched genes, which function or have been implicated to function in retinal degeneration or phototransduction, emerged in this study. Furthermore, there was excellent concordance between the GeneChip, Northern blot and in situ hybridization data.
In addition to the known eye-enriched genes, 93 genes were identified that encode multiple classes of proteins whose roles in visual physiology were either poorly understood or completely unknown. Many of these genes fall into the same classes as those reported in an analysis of rod-enriched genes in the mouse (Blackshaw et al, 2001). A large class encodes proteins that function in post-translational modifications, which includes related ubiquitin hydrolases and similar glycosyltransferases. Another group consists of eye-enriched genes that encode proteins that may be involved in redox control and/or detoxification. These proteins may be present to reduce the deleterious effects resulting from oxidative stress in photoreceptor cells. An eye-enriched DNA repair protein, Mre11, may also be present to limit chromosomal damage induced by intense illumination. Similar mechanisms for controlling oxidative stress and light-induced damage may exist in flies and mammals as functionally related glutathione peroxidase and DNA repair proteins are enriched in rod cells (Blackshaw et al, 2001).
Several genes encoding proteins that function in proteolysis were eye enriched in both Drosophila and the mouse. In addition to light-dependent downregulation of rhodopsin, these proteins might control the levels of photoreceptor cell proteins that oscillate during the circadian cycle. Ten eye-enriched RNAs oscillate during the circadian cycle (Claridge-Chang et al, 2001; McDonald and Rosbash, 2001), including the ubiquitin hydrolase, which is similar to the rod-enriched protein (Blackshaw et al, 2001). In addition, protein degradation may be essential for the elimination of damaged proteins generated during metabolic stress (reviewed in Penn and Anderson, 1992). Regulated removal of such damaged proteins could provide a mechanism to limit the accumulation of inactive proteins, which might lead to photoreceptor cell death.
Several other classes of formerly unrecognized eye-enriched genes in flies were also represented among the mammalian rod-enriched genes. These included genes encoding proteins involved in synaptic function, the regulation of cell death and lipid metabolism. This latter category was expected to be included among the Drosophila eye-enriched genes as fly phototransduction utilizes phosphoinositide signaling. However, the second messenger in mammalian phototransduction is cGMP. A regulatory role for phosphoinositides, which may be common between mammalian and Drosophila photoreceptors, may be the light-dependent movement of proteins, as occurs for Drosophila visual arrestin (Lee et al, 2003). Thus, despite the striking differences in morphology and second-messenger systems, the similarities in classes of proteins between the fly and mammalian visual systems may reflect similar mechanisms for dealing with light-dependent and oxidative damage, rapid signaling, light-dependent protein translocation and circadian rhythm.
Requirement for Sun for downregulation of rhodopsin and to prevent light-dependent retinal degeneration
In the current work, we focused on sun, since it was the only eye-enriched member of a particularly large family of transmembrane proteins, many of which are poorly characterized. We found that mutation of sun resulted in light- and age-dependent retinal degeneration. To address the normal function of Sun in wt photoreceptor cells, we first examined the spatial distribution of the protein. We found that Sun was expressed in lysosomes, suggesting that it might play a role in the degradation of one or more photoreceptor proteins. In support of this possibility, we found that the light-induced degradation of Rh1 was reduced in a sun1 background. Moreover, Rh1 and Sun appeared to colocalize in lysosomes and formed a complex in vivo. Therefore, an attractive model is that Sun is required for light-dependent downregulation of rhodopsin by promoting its degradation in lysosomes.
The model presented here is consistent with findings that other GPCRs are internalized and then targeted either to lysosomes or recycling endosomes in response to strong or prolonged exposure to their agonists (reviewed in Ferguson, 2001). Rh1 may normally be internalized and degraded in lysosomes to downregulate the concentration of the receptor in the presence of intense or constant light, in a manner similar to agonist-induced downregulation of most GPCRs. Upon exposure to bright continuous light, there were large invaginations of the rhabdomeral membrane, which were detected in sun but not wt ommatidia. These large invaginations might accumulate in sun photoreceptor cells due to a requirement for Sun for efficient transfer of Rh1 from these rhabdomeral invaginations to MVBs and subsequently to lysosomes. In the absence of transfer of Rh1-containing membranes to the MVB/lysosomal pathway, the invaginations may increase in size and accumulate. These data raise the possibility that a defect in the downregulation of activated receptors and accumulation of membrane invaginations may underlie the light-dependent retinal degeneration that occurs in sun1 mutants.
The requirement for Sun is distinct from that of two other tetraspanins that are expressed in the disk rims of mammalian photoreceptor cells, peripherin/rds and rom-1, and appear to be structural proteins (Travis et al, 1991; Bascom et al, 1992). Additional tetraspanins may be situated in intracellular compartments of a variety of cell types, including mammalian photoreceptor cells, and function in agonist-dependent protein degradation. Given that the tetraspanin that is among the most related to Sun (CD63) is present in late endosomal and lysosomal membranes (Fukuda, 1991), we propose that other tetraspanins may interact with and participate in the downregulation of other GPCRs.
Materials and methods
RNA preparation, hybridization to Drosophila genome arrays, scanning and staining
Detailed descriptions of the methods that we used to prepare the RNA samples (P-MEXP-2143) and labeling of the cRNAs (P-MEXP-2178) have been deposited in The European Bioinformatics Institute ArrayExpress database (http://www.ebi.ac.uk/arrayexpress). Hybridization, scanning and analysis of Affymetrix Drosophila genome arrays were performed according to the manufacturer's instructions. The annotation for the probe set is available at http://www.affymetrix.com/products/arrays/specific/fly.affx. Biotinylated oligonucleotide (B2) and bioB, bioC, bioD and cre control cRNAs, which were supplied by Affymetrix, were spiked into each sample. The hybridization cocktail consisted of 20 μg of fragmented biotinylated cRNA (final concentration: 0.07 μg/μl), control oligonucleotide B2 (50 pM), control cRNAs bioB, C, D and cre (1.5, 5, 25 and 100 pM, respectively), herring sperm DNA (0.1 mg/ml), acetylated BSA (0.5 mg/ml) and hybridization buffer (100 mM MES, 1 M sodium, 20 mM EDTA, 0.01% Tween 20 in a total volume of 300 μl). The cocktail was heated to 99°C for 5 min before hybridization. The Affymetrix Fluidics Station and Scanner were both controlled by the Affymetrix Microarray Suite version 4 software (MAS4.0) using the default protocols and parameters throughout. The arrays were stained with streptavidin phycoerthyrin (SAPE) solution and scanned once. A second stain was then performed with a biotinylated antibody and visualized with SAPE. The image resulting from this second scan was further analyzed with MAS4.0. The samples were then hybridized to the array for 16 h at 45°C in a rotating oven. Three independent RNA samples were analyzed for each sample group.
Analysis of microarray data
A slight variation of the MAS4.0 algorithm was used to calculate expression values for each probe set as described below. The background subtraction was as described for MAS4.0. The expression values reported are essentially the same as the MAS4.0 ‘average difference' expression metric, except that any probe pairs in a probe set excluded as outliers in a sample (using the MAS4.0 criteria of 3 s.d. from the mean of the probe set) were excluded uniformly from all of the arrays when calculating the fold changes. This uniform masking of outliers is the only way in which the analysis algorithm varies from the standard MAS4.0 software. Each array was scaled (normalized) as in the MAS4.0 algorithm so that the trimmed mean (mean of the middle 96%) expression value for each array was 150. The expression levels for each of three pairs of samples in each group were used to compute the ratio and expression level changes for each probe set. The standard errors for the absolute values and ratios are presented in Tables 1 and 2 and Supplementary Tables 1 and 2. The complete set of microarray data has been deposited in the EBI ArrayExpress database.
Verification of microarray results
Selected eye-enriched genes with different enrichment ratios were used to probe Northern blots. The band intensities were quantitated using a Fuji PhosphoImager (MacBACV2.5 software). In situ hybridizations on sections of adult fly heads were performed as described (Schaeren-Wiemers and Gerfin-Moser, 1993) using digoxigenin-labeled RNA probes.
Generation of a sun mutant
The w+ P-element insertion, EP(2)2425, which was used to generate the deletion in sun by imprecise excision of the P-element, inserted 128 base pairs 5′ from the transcription start site of CG12143 (sun) predicted by the BDGP. EP(2)2425 was mobilized by genetically introducing transposase using the Δ2–3 line (Robertson et al, 1988). In all, 93 individual w− excision chromosomes were placed over a deficiency that spanned sun (Df(2R)Drl[rv25]). Genomic DNAs were combined into 18 pools and screened for deletions that removed sun by PCR. Individual DNAs in the positive pools were rescreened by PCR and the deletions were confirmed by DNA sequencing.
Transgenic flies
A 5.0 kb sun genomic DNA fragment (nucleotides 37586–42771 in AE003842) was generated by PCR and subcloned into the pCaSpeR4 vector (Thummel and Pirrotta, 1992). This clone, which included 1.5 and 1.0 kb 5′ and 3′ of the sun gene, was sequenced to confirm that there were no mutations in the sun coding region. The sun cDNA, which was fused to an N-terminal Myc tag, was linked at the 5′ end to the ninaE (Rh1) promoter and the ninaE 3′ UTR and subcloned into pCaSpeR4. The constructs were injected into w1118 embryos and the resulting transgenes were genetically introduced into w1118;sun flies.
Production of anti-Sun antibodies
A GST fusion protein was generated (pGEX5X-1 vector, Pharmacia), which included the region corresponding to the loop between transmembrane segments 3 and 4 (residues 99–187) and the C-terminus (residues 206–227) of Sun. The fusion protein was fractionated by SDS–PAGE, electroeluted as described (Montell and Rubin, 1988) and injected into rabbits (Covance).
Localization of Sun
Immunofluorescence staining of 0.5 μm sections of P[w+;Rh1::myc::sun];cn bw compound eyes was performed as described (Li and Montell, 2000) using rabbit polyclonal anti-Myc antibodies. To perform the immunostaining at the EM level, compound eyes from 6-day-old w;P[w+;Rh1::myc::sun] flies were fixed, embedded in LR White, thin-sectioned and immunolabeled as described (Tsunoda et al, 2001). The double labeling was performed using mouse anti-Rh1 monoclonal antibodies and rabbit anti-Myc primary antibodies and 15 nm colloidal gold-conjugated anti-rabbit IgG and 5 nm colloidal gold-conjugated anti-mouse IgG secondary antibodies. The immunolabeled sections were stained with 1% aqueous uranyl acetate for 30 s and examined using a JEM 1200EX electron microscope. To determine the localization of Sun in tissue culture cells, HEK cells were transfected with pcDNA3-yfp or pcDNA3-yfp::sun. In some experiments the cells were subsequently loaded with LysoTracker Red DND-99 (Molecular Probes). Confocal images were acquired using an UltraView LCI System (Perkin-Elmer).
Analyses of retinal degeneration
The flies used for morphological analyses were reared at 25°C under a 12 h light/12 h dark cycle. To assay the time course of photoreceptor cell degeneration, the mean number of rhabdomeres per ommatidium was determined using the optical neutralization technique (Franceschini and Kirschfeld, 1971). Each data point was based on examination of ⩾90 ommatidia. Transmission EM was performed as described (Li and Montell, 2000) using hemisected fly heads embedded in LR White.
Blue light-induced changes in Rh1 concentration in flies
w1118 and w1118;sun flies were exposed to 10 mW blue light for 6 h, and the fly heads were homogenized in Tris-buffered saline (20 mM Tris (pH 7.5), 150 mM NaCl) containing 0.5% Triton X-100 and protease inhibitors, and then mixed with an equal volume of 2 × SDS sample buffer. The extracts equal to 0.2 fly heads were fractionated by SDS–PAGE and the Western blots were probed with anti-Rh1 antibodies. Separate blots containing the same extracts were probed with anti-Arr2, anti-INAD and anti-TRP antibodies.
Co-immunoprecipitations
The Sun and Rh1 co-IPs were performed as described (Chevesich et al, 1997) with minor modifications. In all, 10 mg fly heads were homogenized in 1 ml buffer A (20 mM Tris–HCl (pH 7.5), 100 mM NaCl, 5 mM MgCl2, 10% sucrose, 1% glycerol, 1 mM EDTA and 1 × c[I]mplete protease inhibitors containing 1% CHAPS) on ice. The extracts were centrifuged at 16 000 g for 5 min at 4°C, 200 μl aliquots of the supernatant were diluted with 800 μl buffer A without CHAPS, and 1 μl primary antibody or preimmune serum was added and incubated for 3 h at 4°C. The protein A beads were blocked with buffer A containing 0.2% CHAPS and 1% BSA for 30 min at 4°C and 50 μl of blocked protein A beads were added to the tubes containing the immune complexes and incubated for 1 h at 4°C. After washing 2 × with buffer A containing 0.2% CHAPS, the immune complexes were eluted with 2 × SDS sample buffer and a Western blot was probed with anti-Rh1 monoclonal antibodies.
Reconstitution of blue light-induced degradation of Rh1 in isolated ommatidia
A transgene expressing Rh1 tagged at the N-terminus with GPF (gift from Dr J O'Tousa) was introduced into a sun1 background. Ommatidia were dissected from adult flies as described (Hardie, 1991) with minor modifications. Ommatidia were dissected in DMEM medium, transferred to the Schneider's Drosophila medium and dark adapted for 1 h at room temperature. The ommatidia were either kept in the dark or illuminated with blue light for 1.5 h in Schneider's medium containing 50 nM LysoTracker-Blue DND-22 (Molecular Probes), and then confocal images were acquired.
To assay for light-induced degradation of Rh1, dark-adapted ommatidia were either stimulated with blue light for 4 h or kept in the dark. Western blots containing ommatidial extracts were probed with anti-Rh1 monoclonal antibodies. For experiments using lysosomal and proteasomal inhibitors, the ommatidia were preincubated with either 10 μg/μl leupeptin or 10 μM lactacystin before initiation of the light treatment. To determine the relative concentrations of Rh1 on Westerns, the blots were probed with anti-Rh1 primary antibodies, washed and probed with 125I-labeled anti-mouse IgG (ICN). As a control for equal loading, we used rabbit anti-INAD and 125I-labeled protein A. The band intensities were quantified using a Fuji Phosphoimager and MacBACV2.5 software.
Supplementary Material
Legends to Supplementary Tables and Figures
Acknowledgments
We thank Y Pan and M Sepanski for preparing sections of compound eyes, B Hu for help with the microinjections and Dr J O'Tousa for the transgenic flies expressing GFP-Rh1. This work was supported by a grant to CM from the NEI (EY08117), by grants from the NCI (R21 CA94393 and RO1 CA92190) to LAC and by a Grant-in-Aid for Scientific Research (C) from the Japan Society for the Promotion of Science to ES.
References
- Adams MD, Celniker SE, Holt RA, Evans CA, Gocayne JD, Amanatides PG, Scherer SE, Li PW, Hoskins RA, Galle RF, George RA, Lewis SE, Richards S, Ashburner M, Henderson SN, Sutton GG, Wortman JR, Yandell MD, Zhang Q, Chen LX, Brandon RC, Rogers YH, Blazej RG, Champe M, Pfeiffer BD, Wan KH, Doyle C, Baxter EG, Helt G, Nelson CR, Gabor Miklos GL, Abril JF, Agbayani A, An HJ, Andrews-Pfannkoch C, Baldwin D, Ballew RM, Basu A, Baxendale J, Bayraktaroglu L, Beasley EM, Beeson KY, Benos PV, Berman BP, Bhandari D, Bolshakov S, Borkova D, Botchan MR, Bouck J, Brokstein P, Brottier P, Burtis KC, Busam DA, Butler H, Cadieu E, Center A, Chandra I, Cherry JM, Cawley S, Dahlke C, Davenport LB, Davies P, de Pablos B, Delcher A, Deng Z, Mays AD, Dew I, Deitz SM, Dodson K, Doup LE, Downes M, Dugan-Rocha S, Dunkov BC, Dunn P, Durbin KJ, Evangelista CC, Ferraz C, Ferriera S, Fleischmann W, Fosler C, Gabrielian AE, Garg NS, Gelbart WM, Glasser K, Glodek A, Gong F, Gorrell JH, Gu Z, Guan P, Harris M, Harris NL, Harvey D, Heiman TJ, Hernandez JR, Houck J, Hostin D, Houston KA, Howland TJ, Wei MH, Ibegwam C, Jalali M, Kalush F, Karpen GH, Ke Z, Kennison JA, Ketchum KA, Kimmel BE, Kodira CD, Kraft C, Kravitz S, Kulp D, Lai Z, Lasko P, Lei Y, Levitsky AA, Li J, Li Z, Liang Y, Lin X, Liu X, Mattei B, McIntosh TC, McLeod MP, McPherson D, Merkulov G, Milshina NV, Mobarry C, Morris J, Moshrefi A, Mount SM, Moy M, Murphy B, Murphy L, Muzny DM, Nelson DL, Nelson DR, Nelson KA, Nixon K, Nusskern DR, Pacleb JM, Palazzolo M, Pittman GS, Pan S, Pollard J, Puri V, Reese MG, Reinert K, Remington K, Saunders RD, Scheeler F, Shen H, Shue BC, Siden-Kiamos I, Simpson M, Skupski MP, Smith T, Spier E, Spradling AC, Stapleton M, Strong R, Sun E, Svirskas R, Tector C, Turner R, Venter E, Wang AH, Wang X, Wang ZY, Wassarman DA, Weinstock GM, Weissenbach J, Williams SM, Woodage T, Worley KC, Wu D, Yang S, Yao QA, Ye J, Yeh RF, Zaveri JS, Zhan M, Zhang G, Zhao Q, Zheng L, Zheng XH, Zhong FN, Zhong W, Zhou X, Zhu S, Zhu X, Smith HO, Gibbs RA, Myers EW, Rubin GM, Venter JC (2000) The genome sequence of Drosophila melanogaster. Science 287: 2185–2195 [DOI] [PubMed] [Google Scholar]
- Alloway PG, Howard L, Dolph PJ (2000) The formation of stable rhodopsin-arrestin complexes induces apoptosis and photoreceptor cell degeneration. Neuron 28: 129–138 [DOI] [PubMed] [Google Scholar]
- Arbeitman MN, Furlong EE, Imam F, Johnson E, Null BH, Baker BS, Krasnow MA, Scott MP, Davis RW, White KP (2002) Gene expression during the life cycle of Drosophila melanogaster. Science 297: 2270–2275 [DOI] [PubMed] [Google Scholar]
- Bascom RA, Manara S, Collins L, Molday RS, Kalnins VI, McInnes RR (1992) Cloning of the cDNA for a novel photoreceptor membrane protein (rom-1) identifies a disk rim protein family implicated in human retinopathies. Neuron 8: 1171–1184 [DOI] [PubMed] [Google Scholar]
- Blackshaw S, Fraioli RE, Furukawa T, Cepko CL (2001) Comprehensive analysis of photoreceptor gene expression and the identification of candidate retinal disease genes. Cell 107: 579–589 [DOI] [PubMed] [Google Scholar]
- Bloomquist BT, Shortridge RD, Schneuwly S, Perdew M, Montell C, Steller H, Rubin G, Pak WL (1988) Isolation of a putative phospholipase C gene of Drosophila, norpA, and its role in phototransduction. Cell 54: 723–733 [DOI] [PubMed] [Google Scholar]
- Bonini NM, Leiserson WM, Benzer S (1998) Multiple roles of the eyes absent gene in Drosophila. Dev Biol 196: 42–57 [DOI] [PubMed] [Google Scholar]
- Chevesich J, Kreuz AJ, Montell C (1997) Requirement for the PDZ domain protein, INAD, for localization of the TRP store-operated channel to a signaling complex. Neuron 18: 95–105 [DOI] [PubMed] [Google Scholar]
- Claing A, Laporte SA, Caron MG, Lefkowitz RJ (2002) Endocytosis of G protein-coupled receptors: roles of G protein-coupled receptor kinases and β-arrestin proteins. Prog Neurobiol 66: 61–79 [DOI] [PubMed] [Google Scholar]
- Claridge-Chang A, Wijnen H, Naef F, Boothroyd C, Rajewsky N, Young MW (2001) Circadian regulation of gene expression systems in the Drosophila head. Neuron 32: 657–671 [DOI] [PubMed] [Google Scholar]
- Dolph PJ, Ranganathan R, Colley NJ, Hardy RW, Socolich M, Zuker CS (1993) Arrestin function in inactivation of G protein-coupled receptor rhodopsin in vivo. Science 260: 1910–1916 [DOI] [PubMed] [Google Scholar]
- Ferguson SS (2001) Evolving concepts in G protein-coupled receptor endocytosis: the role in receptor desensitization and signaling. Pharmacol Rev 53: 1–24 [PubMed] [Google Scholar]
- Franceschini N, Kirschfeld K (1971) Les phénomènes de pseudopupille dans l'oeil composé de Drosophila. Kybertik 9: 159–182 [DOI] [PubMed] [Google Scholar]
- Fukuda M (1991) Lysosomal membrane glycoproteins. Structure, biosynthesis, and intracellular trafficking. J Biol Chem 266: 21327–21330 [PubMed] [Google Scholar]
- Hardie RC (1991) Whole-cell recordings of the light induced current in dissociated Drosophila photoreceptors: evidence for feedback by calcium permeating the light-sensitive channels. Proc R Soc Lond (B) 245: 203–210 [Google Scholar]
- Hyde DR, Mecklenburg KL, Pollock JA, Vihtelic TS, Benzer S (1990) Twenty Drosophila visual system cDNA clones: one is a homolog of human arrestin. Proc Natl Acad Sci USA 87: 1008–1012 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kalidas S, Smith DP (2002) Novel genomic cDNA hybrids produce effective RNA interference in adult Drosophila. Neuron 33: 177–184 [DOI] [PubMed] [Google Scholar]
- Kiselev A, Socolich M, Vinos J, Hardy RW, Zuker CS, Ranganathan R (2000) A molecular pathway for light-dependent photoreceptor apoptosis in Drosophila. Neuron 28: 139–152 [DOI] [PubMed] [Google Scholar]
- Kopczynski CC, Davis GW, Goodman CS (1996) A neural tetraspanin, encoded by late bloomer, that facilitates synapse formation. Science 271: 1867–1870 [DOI] [PubMed] [Google Scholar]
- Kosloff M, Elia N, Joel-Almagor T, Timberg R, Zars TD, Hyde DR, Minke B, Selinger Z (2003) Regulation of light-dependent Gqα translocation and morphological changes in fly photoreceptors. EMBO J 22: 459–468 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee SJ, Xu H, Kang LW, Amzel LM, Montell C (2003) Light adaptation through phosphoinositide-regulated translocation of Drosophila visual arrestin. Neuron 39: 121–132 [DOI] [PubMed] [Google Scholar]
- Li HS, Montell C (2000) TRP and the PDZ protein, INAD, form the core complex required for retention of the signalplex in Drosophila photoreceptor cells. J Cell Biol 150: 1411–1422 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maecker HT, Todd SC, Levy S (1997) The tetraspanin superfamily: molecular facilitators. FASEB J 11: 428–442 [PubMed] [Google Scholar]
- McDonald MJ, Rosbash M (2001) Microarray analysis and organization of circadian gene expression in Drosophila. Cell 107: 567–578 [DOI] [PubMed] [Google Scholar]
- Minke B, Selinger Z (1992) Intracellular messengers in invertebrate photoreceptors studied in mutant flies. In Neuromethods, Boulton A, Baker G, Taylor C (eds), pp 517–563. Clifton, NJ: The Humana Press Inc. [Google Scholar]
- Montell C (1999) Drosophila visual transduction. Ann Rev Cell Dev Biol 15: 231–268 [DOI] [PubMed] [Google Scholar]
- Montell C, Rubin GM (1988) The Drosophila ninaC locus encodes two photoreceptor cell specific proteins with domains homologous to protein kinases and the myosin heavy chain head. Cell 52: 757–772 [DOI] [PubMed] [Google Scholar]
- Pak WL (1995) Drosophila in vision research. Invest Ophthalmol Vis Sci 36: 2340–2357 [PubMed] [Google Scholar]
- Penn JS, Anderson RE (1992) Effects of light history on the rat retina. In Progress in Retinal Research, Osborne N, Chader G (eds), Vol. 11, pp 75–98. New York: Pergamon Press [Google Scholar]
- Robertson HM, Preston CR, Phillis RW, Johnson-Schlitz DM, Benz WK, Engels WR (1988) A stable genomic source of P element transposase in Drosophila melanogaster. Genetics 118: 461–470 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rong YS, Golic KG (2000) Gene targeting by homologous recombination in Drosophila. Science 288: 2013–2018 [DOI] [PubMed] [Google Scholar]
- Schaeren-Wiemers N, Gerfin-Moser A (1993) A single protocol to detect transcripts of various types and expression levels in neural tissue and cultured cells: in situ hybridization using digoxigenin-labelled cRNA probes. Histochemistry 100: 431–440 [DOI] [PubMed] [Google Scholar]
- Suzuki E, Hirosawa K (1991) Immunoelectron microscopic study of the opsin distribution in the photoreceptor cells of Drosophila melanogaster. J Electron Microsc (Tokyo) 40: 187–192 [PubMed] [Google Scholar]
- Takeshima H, Komazaki S, Nishi M, Iino M, Kangawa K (2000) Junctophilins: a novel family of junctional membrane complex proteins. Mol Cell 6: 11–22 [DOI] [PubMed] [Google Scholar]
- Thummel CS, Pirrotta V (1992) New pCaSpeR P element vectors. Dros Info Serv 71: 150 [Google Scholar]
- Todres E, Nardi JB, Robertson HM (2000) The tetraspanin superfamily in insects. Insect Mol Biol 9: 581–590 [DOI] [PubMed] [Google Scholar]
- Travis GH, Sutcliffe JG, Bok D (1991) The retinal degeneration slow (rds) gene product is a photoreceptor disc membrane-associated glycoprotein. Neuron 6: 61–70 [DOI] [PubMed] [Google Scholar]
- Tsunoda S, Sun Y, Suzuki E, Zuker C (2001) Independent anchoring and assembly mechanisms of INAD signaling complexes in Drosophila photoreceptors. J Neurosci 21: 150–158 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vihtelic TS, Hyde DR, O'Tousa JE (1991) Isolation and characterization of the Drosophila retinal degeneration B (rdgB) gene. Genetics 127: 761–768 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu XZ, Chien F, Butler A, Salkoff L, Montell C (2000) TRPγ, a Drosophila TRP-related subunit, forms a regulated cation channel with TRPL. Neuron 26: 647–657 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Legends to Supplementary Tables and Figures


