Abstract
The multisubunit eukaryotic Mediator complex integrates diverse positive and negative gene regulatory signals and transmits them to the core transcription machinery. Mutations in individual subunits within the complex can lead to decreased or increased transcription of certain subsets of genes, which are highly specific to the mutated subunit. Recent studies suggest a role for Mediator in epigenetic silencing. Using white-opaque morphological switching in Candida albicans as a model, we have shown that Mediator is required for the stability of both the epigenetic silenced (white) and active (opaque) states of the bistable transcription circuit driven by the master regulator Wor1. Individual deletions of eight C. albicans Mediator subunits have shown that different Mediator subunits have dramatically diverse effects on the directionality, frequency, and environmental induction of epigenetic switching. Among the Mediator deletion mutants analyzed, only Med12 has a steady-state transcriptional effect on the components of the Wor1 circuit that clearly corresponds to its effect on switching. The MED16 and MED9 genes have been found to be among a small subset of genes that are required for the stability of both the white and opaque states. Deletion of the Med3 subunit completely destabilizes the opaque state, even though the Wor1 transcription circuit is intact and can be driven by ectopic expression of Wor1. The highly impaired ability of the med3 deletion mutant to mate, even when Wor1 expression is ectopically induced, reveals that the activation of the Wor1 circuit can be decoupled from the opaque state and one of its primary biological consequences.
INTRODUCTION
Epigenetic gene expression states require mechanisms that encode the state at specific loci, facilitate the stable inheritance of that state through cell division, and provide for a transcriptional readout. Feedback loops, DNA methylation, and chromatin modification and higher-order chromatin structure are among the features associated with epigenetic states (1–3). Bistable switches tend to be unidirectional in metazoan cells during development. In unicellular organisms, these switches are usually metastable (bidirectional) and are thought to provide an adaptive advantage in an ever-changing host environment (4–6). A combination of DNA-bound transcription factors, noncoding RNAs, local chromatin landscape, and coregulatory complexes specifies the gene expression patterns that define the physiology and morphology associated with an epigenetic state. Mediator is a central eukaryotic coregulatory complex that has also been implicated in the silencing of epigenetic states.
Transcription in purified and extract-based systems, in vivo studies, and cell-based assays have revealed that Mediator can be a critical functional/physical intermediary between DNA-bound activators and the core transcription machinery in all eukaryotes (7, 8). Saccharomyces cerevisiae Mediator (scMediator) has structurally distinct modules referred to as Tail, Middle, Head, and Cdk8 (9–12). Purification and characterization of Candida albicans Mediator (caMediator) (13) showed that the caMediator complex has a composition and architecture similar to that of scMediator. Certain scMediator subunits are critical for the activation of specific sets of genes (14, 15), and the Tail module has been shown to be a direct target of activation domains (16–19). Transcriptional profiling in vivo has shown that other subunits are essential for transcription of virtually all genes (20). Work in S. cerevisiae has also pointed to a role for several Mediator subunits in transcriptional repression and silencing that likely involves chromatin (21–27).
A specialized class of subunits, mainly in the Middle module and on the interface of the Middle and Tail modules, appears to have a primary role in regulating silencing and chromatin in S. cerevisiae. Although the Cdk8 module is clearly involved in transcription repression (15, 28), this functionality appears to be separable from those regulating chromatin, silencing, and certain forms of repression (24, 26, 29). Mutations in med16 (sin4) and med14 (rgr1), which lead to derepression of a subset of genes potentially through an epigenetic mechanism (30), are accompanied by gross alterations in chromatin structure in vivo (22, 31). Recently published work revealed that telomerically bound scMediator was required for the establishment of heterochromatin and the silencing of telomeric reporter genes (23, 26, 27, 29). This work showed that deletion of specific Mediator subunits, but not others, resulted in a decrease in Mediator occupancy at telomeres that was accompanied by a loss in heterochromatin and silencing (26). Mediator appears to be targeted to telomeric chromatin through direct interactions with the tails of histones H3 and H4 (27). This affinity of Mediator for H4 can be decreased by lysine acetylation of the H4 tail. Specifically, Mediator associates in vitro and in vivo with H4 tails lacking acetylation at K16, the hallmark of telomeric heterochromatin (27). This work suggests that Mediator-chromatin interactions are on the pathway leading to telomeric heterochromatin. Since telomeric silencing is a classical but highly specialized model for epigenetic silencing, we decided to adopt C. albicans white-opaque switching to study the mechanism of Mediator-facilitated stabilization of epigenetic states within a defined biological pathway.
Under certain conditions, genetically identical C. albicans cells can exist in two different morphological states, white and opaque (32). Phenotypic/morphological transitions in C. albicans are critical to the virulence and basic biology of this fungal human pathogen. The clearest biological impact of white-opaque switching in C. albicans is on its sexual cycle, as only opaque cells can mate (33). The white and opaque states can arise spontaneously and are inherited over ∼104 generations (32). Environmental conditions can regulate the switching frequency in a directional manner, but once established, the state can be inherited in the absence of the signal (34, 35). Hence, white-opaque switching can be defined as an epigenetic phenomenon. The white and opaque morphologies have distinct transcriptional states consisting of over 1,000 differentially regulated transcripts (36). The heart of the transcriptional circuit that governs these two states is a master regulator called Wor1. Wor1 is a fungus-specific DNA-bound transcription factor (37) whose expression is at a high level in the opaque state but silenced in the white state. Wor1 is required for the switch from the white state to the opaque state and is part of an interlocking feedback loop in which it binds and activates its own promoter (38–41). The white-opaque switching model has wide-ranging significance. Although the impact of white-opaque switching on C. albicans virulence is unclear (42, 43), Wor1 orthologs in other fungal pathogens of mammals (44) and plants (45–47) are clearly determinants of virulence. Meta-analysis has shown that the architecture and stability of the white-opaque transcriptional circuit share many properties with those of the Oct4 circuit, which governs the pluripotency and differentiation of mouse embryonic stem cells (36).
Wor1 drives white-opaque switching by binding at a wide variety of other loci throughout the genome—some that it activates and others that it represses (37, 40). Wor2, another transcription factor, is part of an interlocking positive-feedback loop with Wor1 and is required for the switch from white to opaque. On the other hand, the Efg1 transcription factor is part of an interlocking negative-feedback loop with Wor1, and the white state is highly unstable in its absence (41). A working model based on these studies posits that the epigenetic heritability of the opaque state derives from the Wor1 protein that is passed on to daughters during mitosis (42). In this model, switching originates from stochastic fluctuations in the amount of Wor1 above or below certain thresholds. In principle, the actual mechanistic Wor1 threshold could be anywhere along the pathway leading to Wor1 expression, from the status of chromatin in the promoters of key regulators of the circuit to their transcript or protein stability. Though many transcription networks utilize positive-feedback loops (48), few of them have documented heritably stable transcriptional states similar to those of the Wor1-driven circuit. Several recent studies have identified a key role for chromatin structure and dynamics in white-opaque switching (49–52). Given the connection between Mediator and telomeric heterochromatin and silencing in S. cerevisiae, we sought to determine whether C. albicans Mediator affects the bistability of gene expression in the white and opaque states.
Consistent with the idea that different parts of Mediator play highly specific roles in the stability of epigenetic transcription states, we have found that different subunits of C. albicans Mediator have diverse effects on the direction and frequency of white-opaque switching. For most of the mutants, the impact is likely happening through a mechanism that involves something other than a direct effect on the steady-state mRNA levels of the key components of the Wor1 transcriptional circuit in the final states. In addition, our analysis of the Med3 deletion mutant has shown that the Wor1 transcription circuit can be decoupled from its phenotypic output.
MATERIALS AND METHODS
Media and growth conditions.
C. albicans was routinely grown at 25°C in synthetic complete (SC) media (SC+Glu) (6.7 g/liter yeast nitrogen base [Difco/BD], 2 g/liter complete amino acid mix minus uracil [US Biological], 20 g/liter dextrose, 0.2 mM uridine, 0.1 μM ZnOAc [buffered with 20 mM K2HPO4/KH2PO4 to pH 6.8]) or at 30°C in YPD media (10 g/liter yeast extract, 20 g/liter peptone, 20 g/liter dextrose). To induce ectopic WOR1 expression, 20 g/liter maltose (Mal) was added instead of glucose. For cells growing on solid media, 2% agar plates were used.
Plasmid and strain construction.
The complete lists of C. albicans strains, primers, and plasmids used in this study are shown in Table S1, Table S2, and Table S3 in the supplemental material. Most strains, except where noted, were derived from the SN152 (53) MTLa/a strain. The specific open reading frames (ORFs) of Mediator subunit genes were previously identified (13). Mediator subunit single gene deletions (med5Δ/Δ, med9Δ/Δ, med1Δ/Δ, med16Δ/Δ, med3Δ/Δ, med15Δ/Δ, med20Δ/Δ, med12Δ/Δ) were created by using the C. maltosa LEU2 and C. dubliniensis HIS1 fusion PCR cassettes as described by Noble and Johnson (53). For each Mediator homozygous mutant, at least 4 independent strains derived from 2 independent heterozygous mutants were tested in switching assays. To complement the mutant strain, a region containing the upstream sequence and the complete ORF of a Mediator gene was amplified with the appropriate “complement 5′ F” and “complement 5′ R” primer pair, and the downstream region was amplified with the appropriate “complement 3′ F” and “complement 3′ R” primer pair. The upstream sequence and the complete ORF of MED12 were amplified in two parts due to its large size. The upstream/ORF PCR product and the downstream PCR product were double digested with enzymes listed together with the primers in Table S2. The two products were inserted sequentially into plasmid pSN69 (53), generating the complementation plasmid as shown in Table S3. This complementation cassette was then linearized and transformed into the mutant strain for complementation. At least 3 independent complemented strains for a single Mediator mutant were analyzed. The strains with ectopically expressed Wor1 were created as described by Stevenson and Liu (51) using a PMAL2-WOR1-3HA cassette. At least 3 independent Wor1-overexpression strains for each of the wild-type (WT) and Mediator mutant strains were analyzed.
White-opaque switching assay.
The white-opaque switching assay at 25°C was performed as previously described (38). Briefly, white cells from a frozen stock were streaked onto SC+Glu plates for 4 to 5 days of pregrowth at 25°C. Three or four pure white colonies were resuspended in water and plated onto SC+Glu+(5 μg/ml) phloxine B plates at a density of approximately 100 cells per plate. After approximately 6 to 7 days of growth at 25°C, the white-opaque switching frequency was scored as the percentage of single colonies that were opaque or had at least one opaque sector. The opaque-white switching assay was performed starting with pure opaque single colonies following a procedure similar to that described above. The induced opaque-white switching assay at 37°C was performed as described previously (54) with modifications. Freshly grown pure opaque colonies were inoculated into 2 ml SC+Glu liquid media for vigorous growth at 25°C for 4 h. The opaque cell culture at an optical density at 600 nm (OD600) of 1 was then diluted with prewarmed SC media to 8 ml at 37°C and grown on a roller drum for 9 h. Every 1.5 h, a 5-ml aliquot of the culture was pulled out for analysis and 5 ml fresh prewarmed SC media was added back to promote a constant growth rate (Matthew Lohse, personal communication). Cells were plated onto SC+Glu+phloxine B plates, and the total number of colonies and the number of white colonies were counted after 5 days of incubation at 25°C.
Purification of the med16Δ/Δ Mediator.
A 6×His-Flag tag was appended to the C terminus of the Med8 subunit in a WT strain (BWP17) and med16Δ/Δ strain (AZC34) and to the C terminus of Med15 in a med16Δ/Δ strain (AZC34) as previously described (13). To each of these strains, a hemagglutinin (HA) tag (55) was appended to the C terminus of Med3 to yield doubly tagged Med8-6×His-Flag/Med3-HA WT (cTTR03) and med16Δ/Δ (cTTR04) strains and a Med15-6×His-Flag/Med3-HA med16Δ/Δ (cTTR05) strain. Mediator was purified to homogeneity from each of these strains and analyzed as previously described (13).
RNA isolation and RT-qPCR.
Cultures were grown in SC+Glu media at 25°C starting from fresh and pure white or opaque colonies and harvested by centrifugation at an OD600 of approximately 1 to 2. For Mal induction, cells were grown in SC+Glu media for 3 h to an OD600 of 0.2, washed once with SC+Mal media, and harvested after 6.5 h growth in SC+Mal media. Pellets were washed once in double-distilled water (DDW) and flash-frozen in liquid nitrogen. Cells were lysed in 600 μl RTL buffer (RNeasy Minikit; Qiagen) by two rounds of bead beating (Mini-Beadbeater-16; BioSpec Products) for 1 min separated by 2 min cooling on ice. Following cell lysis, total RNA was extracted and purified following the manufacturer's protocols. About 1 μg of total RNA was treated with rDNase (Ambion) for 30 min at 37°C and then used in reverse transcription (RT) (iScript cDNA synthesis kit; Bio-Rad). cDNA amplification was performed by real-time PCR with SYBR green (Bio-Rad) on a Bio-Rad Q7500 cycler. The RT-quantitative PCR (RT-qPCR) primers for EFG1, CZF1, and WOR1-3HA (see Table S2 in the supplemental material) were designed using Beacon Designer 6 (Premier Biosoft).
Immunoblotting.
Cultures were grown in SC+Glu or SC+Mal media to an OD600 of 1 to 2 and harvested by centrifugation as described above. Pellets were washed once in DDW and lysed in ESB (1% SDS, 1% 2-mercaptoethanol, 15% glycerol, 0.01% bromophenol blue, 50 mM Tris-HCl, pH 6.8) with boiling followed by 5 min of bead beating. Western blotting was performed using the following primary antibodies: rabbit anti-Wor1 antibody (kindly provided by the Johnson laboratory) (40), mouse anti-HA.11 antibody (MMS-101R; Covance), rat anti-tubulin alpha antibody (MCA78G; AbD Serotec), mouse anti-Flag antibody (M2; Sigma), and rabbit anti-caMed1 antibody (13). Secondary antibodies were alkaline phosphatase conjugated and were used with a fluorescent substrate (ECF; GE Healthcare). Quantitative gel analysis was performed using ImageJ64 (NIH).
Quantitative mating assay.
Mating assays were performed as described previously (56) with modifications. Pure white and opaque colonies were inoculated into SC+Glu or SC+Mal liquid media for approximately 3 to 5 h of growth at 25°C to an OD600 of 2. Cells were then harvested, and 0.4 OD of MTL α/α cells and 0.6 OD of a/a cells were mixed in 300 μl DDW. The cell mixture was spotted onto YPD–55 μg/ml adenine or YPMal–55 μg/ml adenine plates for mating at 25°C. After 3 days, the lawn of cells was scraped from the plate and resuspended in DDW. Since the strains were not filamentous, filter discs were not necessary. After cell aggregates were broken up by vigorous vortexing and pipetting, cells were diluted and plated onto selective plates (SC-Arg plates for α + a/α cells and SC–Arg-His plates for a/α cells). After 3 days of growth at 25°C, colony formation on each plate was counted and the mating efficiency was calculated as (a/α)/(α + a/α).
RESULTS
Mediator subunits differentially regulate the direction and frequency of white-opaque switching.
Studies in C. albicans have recently shown that certain Mediator subunits play regulatory roles in cytokinesis, stress response, germ tube formation, filamentous growth, and virulence (13, 57). However, Mediator's impact on the regulation of white-opaque switching had never previously been investigated. Here we have constructed a series of C. albicans strains, in which genes encoding Mediator subunits (13) had been deleted, to determine how and if individual subunits regulated the directionality and frequency of white-opaque switching. To identify subunits encoded by nonessential genes in C. albicans, we drew upon the analysis of the orthologous Mediator genes in S. cerevisiae. All of the nonessential S. cerevisiae Mediator orthologs that we analyzed in C. albicans (see below) were also nonessential. The Mediator genes we chose to delete fell into four categories: subunits in the Tail module (MED5, MED16, MED3, MED15); subunits in the Middle module (MED1, MED9); a subunit in the Cdk8 module (MED12); and a subunit in the Head module (MED20) (13, 58, 59). We performed quantitative switching assays on these single mutants and compared both their white-opaque and opaque-white switching frequencies to those seen with a WT strain. Except for the med5Δ/Δ strain, all Mediator deletion mutants tested showed significant changes in switching frequency compared to the WT (Table 1 and Fig. 1). The med1Δ/Δ mutant and med20Δ/Δ mutant strains showed an increase in the white-opaque switching frequency, but a decrease in the opaque-white switching frequency, consistent with the idea that the opaque state is stabilized relative to the white state in these strains. The med12Δ/Δ mutant strain showed the opposite trend in switching frequency. This strain possessed a dramatically decreased white-opaque switching frequency accompanied by a very high (93%) opaque-white switching frequency. Opaque cells were never observed in the med3Δ/Δ or med15Δ/Δ strain, even under conditions of CO2 induction (see below), using either colony or single-cell morphology as a criterion. Previously, the inability to switch from the white state to the opaque state was observed only in the wor1Δ/Δ mutant (38–40), wor2Δ/Δ mutant (41), myo5Δ/Δ mutant (60), and sla2Δ/Δ mutant (60). In these mutants, the master regulation circuit was broken either by the absence of essential players or by the disruption of the cytoskeleton. The med9Δ/Δ and med16Δ/Δ strains belonged to a small class of mutants that displayed increased switching frequencies in both directions. Thus, Med9 and Med16 might play important roles in the stability of both the white state and opaque state. To confirm that the observed effect on switching was due solely to the Mediator mutation, we complemented each Mediator mutant at its original locus. Each complemented strain showed rescue of WT switching frequencies (Table 1). The observed differences in white-opaque switching between Mediator mutants suggest that Mediator can play both positive and negative roles in the regulation of white-opaque switching. To address the concern that stress induced by these mutants may have indirectly affected switching frequency (61), we analyzed the growth rates of both white and opaque cells of these strains under different growth conditions. The mutant and WT strains had comparable doubling times under all conditions used in our study (see Table S4 in the supplemental material) and had similar sensitivities to temperature stress (16°C and 37°C), oxidative stress (H2O2), salt stress (NaCl), and membrane integrity stress (SDS) (data not shown). Therefore, the switching phenotypes of Mediator mutants appear to be tightly related to the direct function of the complex in switching.
Table 1.
Differential effects of Mediator subunits on white-opaque switchinga
| C. albicans strain | White to opaque switching |
Opaque to white switching |
||
|---|---|---|---|---|
| Frequency (%) | n | Frequency (%) | n | |
| WT (a/a) | 4.5 ± 0.2 | 1,917 | 11.2 ± 1.9 | 1,059 |
| med5Δ/Δ | 6.9 ± 1.3 | 1,266 | 11.8 ± 4.0 | 1,299 |
| med9Δ/Δ | 20.3 ± 5.3* | 1,382 | 22.2 ± 4.8* | 1,091 |
| med1Δ/Δ | 17.3 ± 4.0* | 1,138 | 2.2 ± 0.8* | 1,103 |
| med16Δ/Δ | 19.3 ± 5.6* | 1,989 | 54.5 ± 6.9* | 1,106 |
| med3Δ/Δ | <10−4*** | 3,055 | ||
| med15Δ/Δ | <10−4*** | 2,000 | ||
| med20Δ/Δ | 73.4 ± 11.8** | 1,587 | 2.9 ± 1.0** | 1,112 |
| med12Δ/Δ | 0.6 ± 0.3*** | 2,780 | 93.3 ± 2.3*** | 1,191 |
| med9Δ/Δ-MED9 | 3.2 ± 0.8 | 1,215 | 8.7 ± 4.2 | 976 |
| med1Δ/Δ-MED1 | 4.2 ± 1.5 | 1,118 | 9.2 ± 5.1 | 1,077 |
| med16Δ/Δ-MED16 | 3.3 ± 1.8 | 1,457 | 8.0 ± 2.5 | 935 |
| med3Δ/Δ-MED3 | 4.7 ± 1.2 | 1,312 | 9.3 ± 1.8 | 1,067 |
| med15Δ/Δ-MED15 | 5.1 ± 1.3 | 1,057 | 8.5 ± 2.3 | 1,135 |
| med20Δ/Δ-MED20 | 6.3 ± 3.1 | 1,491 | 6.4 ± 2.9 | 1,048 |
| med12Δ/Δ-MED12 | 6.8 ± 2.6 | 959 | 4.5 ± 0.9 | 1,400 |
*, P < 0.05; **, P < 0.01; ***, P < 0.001 (relative to WT; Student's t test). Frequency, the percentage of the colonies that showed an alteration of the original phenotype, is displayed as means ± SD of the results of three independent experiments carried out with the same strain. n, total number of colonies counted for each strain. Independently derived mutants and complemented strains showed similar results (data not shown).
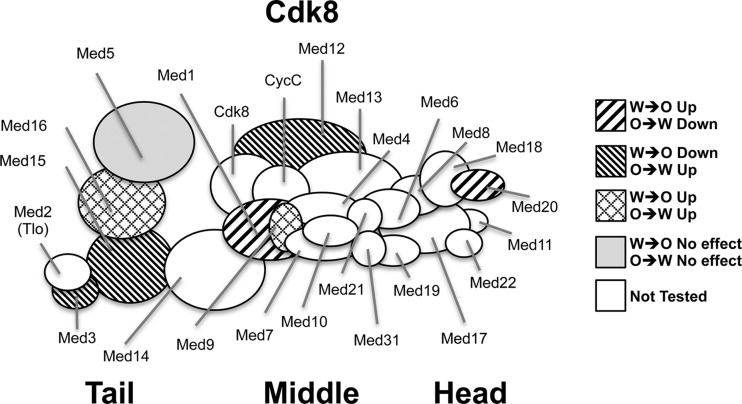
Fig 1.
Graphical summary of C. albicans Mediator subunit architecture and the impact of deletion mutants on white (W)-opaque (O) switching as described in detail in Table 1. The architecture of the C. albicans Tail module is based on published data (13) and data presented here, while the remainder is modeled on the structure of the S. cerevisiae complex (9).
Differences in switching among the med3Δ/Δ, med15Δ/Δ, and med16Δ/Δ strains are consistent with a subunit rearrangement within the C. albicans Tail module versus that of S. cerevisiae.
The architecture of the S. cerevisiae Mediator Tail module has the Med16 subunit serving as a bridge between the Med2, Med3, and Med15 submodule and the rest of the complex via Med14 (58). In Mediator purified from S. cerevisiae med16Δ strains, Med2, Med3, Med5, and Med15 become dissociated from the main complex (23). It was surprising that the med3Δ/Δ and med15Δ/Δ strains could not switch to opaque when Med3 and Med15 were dissociated from the complex in the med16Δ/Δ strain, which showed an increased white-opaque switching frequency. Hence, we asked whether the C. albicans Mediator Tail module had a different architecture from the S. cerevisiae complex. Purifying Mediator from a med16Δ/Δ C. albicans strain with a 6×His-Flag tag on Med8 and an HA tag to monitor the presence of the Tail module demonstrated that deletion of Med16 did not lead to dissociation of the Med2, Med3, and Med15 submodule (Fig. 2). Taking those results together with our previous purification of the C. albicans med3Δ/Δ Mediator, in which the entire Tail module (including Med2, Med3, Med5, Med15, and Med16) was lost (13), there appears to have been a “rewiring” of the protein-protein interactions in the C. albicans Tail module during evolution. The combination of the switching (Table 1) and structural (Fig. 2) data for med16Δ/Δ Mediator supports the idea that Med16 and Med5 represent a physically and functionally separable submodule of the Tail module in C. albicans (Fig. 1).
Fig 2.
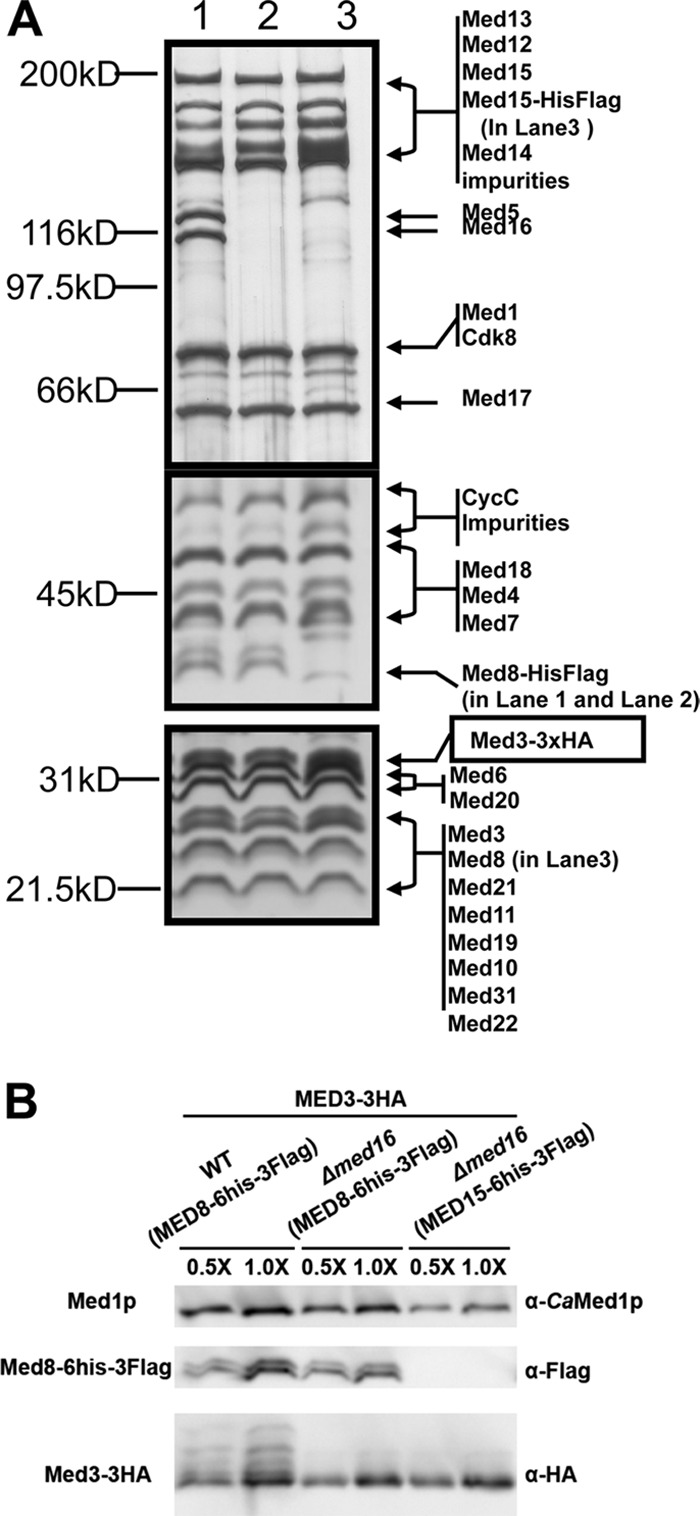
The Med2(Tlo)/Med3/Med15 Tail submodule of caMediator remains associated with the Middle and Head modules in a med16Δ/Δ strain. (A) Silver stain gels of Mediator purified from WT (Med8-6×His-Flag, Med3-3XHA) (lane 1), med16Δ/Δ (Med8-6×His-Flag, Med3-3XHA) (lane 2), and med16Δ/Δ (Med15-6×His-Flag, Med3-3XHA) (lane 3) strains. To obtain adequate resolution over the entire molecular mass range, we show panels from 6%, 7.5%, and 12.5% SDS-PAGE from top to bottom. (B) Western blot of the three samples analyzed as described for panel A, showing equal recoveries of Med3-HA (anti-HA) in WT and mutant Mediator using purification tags on Med8 or Med15. An antibody against caMed1 served as the recovery control.
Impact of Mediator mutants on environmentally induced white-opaque switching.
White-opaque switching frequency can be regulated by exposure to different environmental conditions. In principle, “spontaneous” switching could arise from “intrinsic” transcriptional noise in the Wor1 transcription circuit or from “extrinsic” noise in the signaling pathways that regulate switching, which would then lead to fluctuations in the Wor1 circuit. Hence, mutants that affect switching frequency could arise either from a direct effect on the transcriptional mechanisms regulating the Wor1 transcription circuit or by affecting the environmental signaling pathways that regulate the Wor1 circuit. Stimulation of white-opaque switching by CO2 and stimulation of opaque-white switching by growth at 37°C are among the best-documented conditions that affect switching (62, 63). To determine whether the signaling pathways leading to white-opaque switching were impacted in Mediator mutants, we measured opaque-white switching frequency at 37°C and white-opaque switching frequency in the presence of 5% CO2. Measuring switching frequency at 37°C affords the ability to monitor the kinetics of switching after the temperature of the liquid culture hosting the cells is raised from 25°C to 37°C (54). The med5Δ/Δ mutant strain, which did not affect opaque-white switching frequency (Table 1) compared to the WT, exhibited kinetics of switching identical to those seen with the WT at 37°C (Fig. 3). The med1Δ/Δ and med20Δ/Δ mutant strains, which both exhibited a decreased spontaneous opaque-white switching frequency (∼2%; Table 1), showed a delay in the kinetics of switching from opaque to white at 37°C (Fig. 3). The med1Δ/Δ mutant exhibited an ∼1.5-h delay in the T50 (time to reach 50% white cells in the population, termed the “commitment point”) (54) compared to the WT, while the med20Δ/Δ mutant exhibited a delay of greater than 4.5 h. The med1Δ/Δ mutant had a delay similar to that observed in a heterozygous knockout of the white state-specific transcription factor EFG1 (54). Despite having similar spontaneous opaque-white switching frequencies, the med20Δ/Δ mutant had a substantially longer delay in switching. This finding suggested that Med20 may positively regulate a step in the signaling pathway that leads from a shift to 37°C to shutting down the Wor1 transcription circuit. Among the mutants that led to an increased spontaneous opaque-white switching frequency, we observed a variety of effects on the kinetics of switching from opaque to white at 37°C. The med12Δ/Δ mutant had an almost 90% rate of spontaneous switching from opaque to white (Table 1). Consistent with this effect, the med12Δ/Δ mutant advanced the T50 ∼1.5 h versus that of the WT (Fig. 3). This number was similar to the advancement in the T50 observed in a heterozygous knockout of the opaque state-specific transcription factor WOR1 (54). This similarity was consistent with the finding that the amount of Wor1 and Wor2 expressed in the opaque phase in the med12Δ/Δ mutant was substantially smaller than the WT amount (Fig. 4A). In contrast to the med12Δ/Δ strain, the med16Δ/Δ and med9Δ/Δ mutant strains showed kinetic profiles of opaque-white switching at 37°C that differed from their impact on spontaneous opaque-white switching frequency. Despite a high spontaneous opaque-white switching frequency in the med16Δ/Δ strain (∼55%; Table 1), the kinetics of opaque-white switching at 37°C in the med16Δ/Δ and WT strains were nearly identical (Fig. 3). This result suggests that, at 25°C, Med16 may antagonize a step in the signaling pathway that leads to opaque-white switching at 37°C. The med9Δ/Δ mutant displayed a modest increase in the spontaneous opaque-white switching frequency (∼22%; Table 1) but surprisingly displayed a delayed T50 compared to the WT (Fig. 3). A potential explanation for this disparity may be that the med9Δ/Δ mutant Mediator became less stable at 37°C, which led to a differential impact on Mediator function in opaque-white switching at 37°C versus 25°C.
Fig 3.
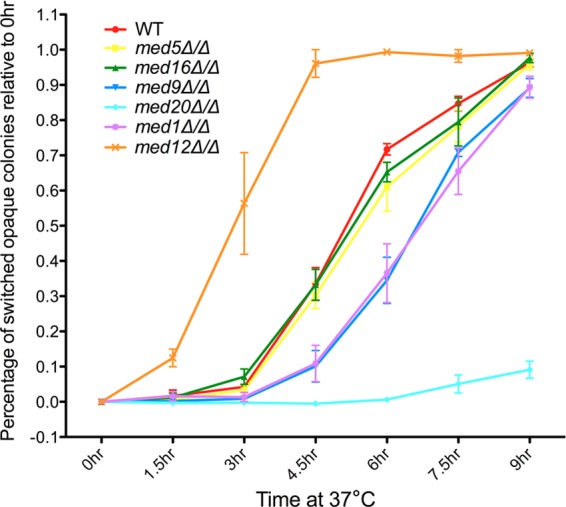
Kinetics of temperature-induced opaque-white switching in Mediator mutant strains. Opaque cells of WT and Mediator mutant strains growing in SC+Glu media, shifted from 25°C to 37°C, were removed every 1.5 h for analysis. The percentage of opaque cells that had switched to white (y axis) was calculated as the ratio of the fraction of opaque cells at each time point relative to the fraction of opaque cells at the starting time point (0 h), which was usually close to 100%. Data were plotted as means ± standard deviations (SD) of the results of three independent experiments with more than 1,500 colonies counted in total for each strain.
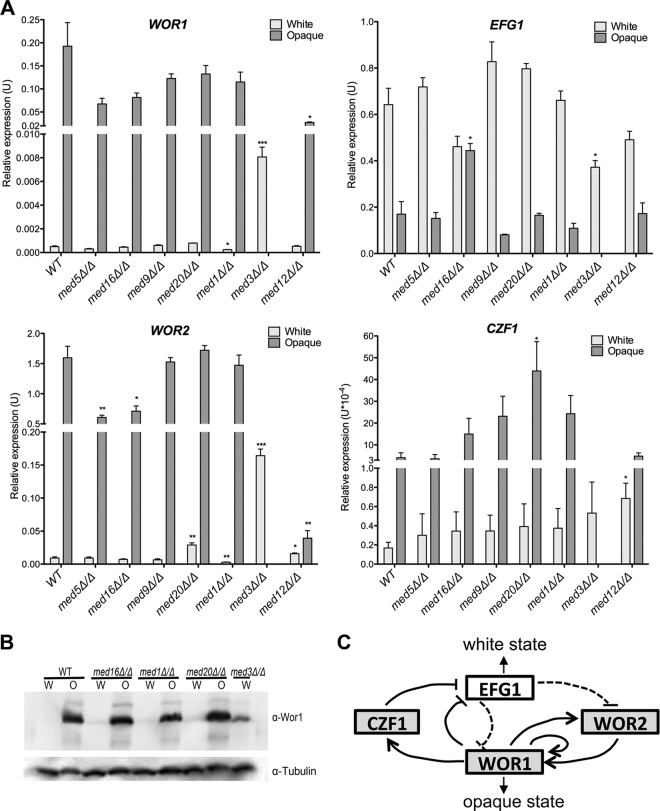
Fig 4.
Steady-state mRNA and protein levels of key transcriptional regulators of white-opaque switching. (A) RT-qPCR analysis of the mRNA levels of WOR1, EFG1, WOR2, and CZF1 in white and opaque cells growing in SC+Glu media. The qPCR was performed in triplicate. Results were normalized to ACT1. The amount of transcript in each mutant was compared to that in the WT white or opaque cells using a t test. Bars represent means ± SD. *, P < 0.05; **, P < 0.01; ***, P < 0.001. (B) Immunoblotting shows that the Wor1 protein level increased in med3Δ/Δ mutant strain white cells relative to the level in WT and other Mediator mutant cells. Tubulin was used as a loading control. (C) Diagram of the interlocking loop that regulates white-opaque switching (modified from PLOS Biology [41]). Dashed lines indicate regulatory controls active in the white state, and solid lines indicate regulatory controls active in the opaque state.
We determined the switching frequency on plates in the presence of 5% CO2 to investigate the effect of Mediator mutants under conditions that stimulate white-opaque switching. Similar to previous observations (62), exposure to CO2 led to an almost 40-fold increase in switching frequency in the WT strain (Table 2). The switching frequency under conditions of induction was close to 100% in the WT strain as well as in the mutants that led to increased spontaneous white-opaque switching. Hence, it was not possible to determine whether the increases in spontaneous switching frequency in these mutants originated from stimulating the CO2 signaling pathway. It is more useful to look at CO2-induced switching in the med3Δ/Δ, med15Δ/Δ and med12Δ/Δ mutant strains, which have decreased spontaneous white-opaque switching. CO2 induction did not lead to white-opaque switching in the med3Δ/Δ strain and med15Δ/Δ strain, consistent with the idea that the opaque state is unstable in these strains. However, CO2 induction was able to stimulate the impaired spontaneous white-opaque switching in the med12Δ/Δ strain and resulted in a fold increase similar to that seen with the WT (Table 2). This finding is consistent with the idea that the decrease in spontaneous white-opaque switching in the med12Δ/Δ strain is not a result of impaired CO2 signaling but is rather a direct effect on the transcriptional activation of the Wor1 circuit.
Table 2.
CO2-induced white to opaque switching of Mediator mutantsa
| Strain | Switching frequency (%) | n |
|---|---|---|
| WT | 94.9 ± 2.7 | 658 |
| med5Δ/Δ | 86.3 ± 6.2 | 741 |
| med9Δ/Δ | 97.6 ± 2.9 | 655 |
| med1Δ/Δ | 96.7 ± 1.1 | 707 |
| med16Δ/Δ | 78.8 ± 9.5 | 888 |
| med3Δ/Δ | <10−4*** | 779 |
| Med15Δ/Δ | <10−4*** | 785 |
| med20Δ/Δ | 100 | 660 |
| med12Δ/Δ | 19.7 ± 2.1*** | 813 |
Cells were preincubated on SC+Glu plates in air at 25°C and replated onto SC+Glu+phloxine B plates and incubated in 5% CO2 at 25°C for 5 days. ***, P < 0.001 (relative to WT; Student's t test). Switching frequency, the percentage of the colonies that showed an alteration of the original phenotype, is displayed as means ± SD of the results of three independent experiments carried out with the same strain. n, total number of colonies counted for each strain.
Steady-state mRNA expression levels of key regulators in white-opaque switching are affected only in select Mediator mutants.
White-opaque switching is under the tight regulation of an interlocking feedback loop composed of the four transcription factors Wor1, Efg1, Wor2, and Czf1 (Fig. 4C). Wor1 is the master regulator; Efg1 plays an inhibitory role in Wor1, while Wor2 and Czf1 play activating roles in this feedback circuit that maintains the metastable white and opaque states (41). A recent study indicated that an additional transcription factor, Wor3, plays a role in controlling this loop (64). Mediator is known to directly impact transcription, and modeling studies suggested that changes in the steady-state levels of the key regulators of the white-opaque switch are a straightforward way to explain changes in switching frequencies (65). Hence, we tested the steady-state mRNA levels of WOR1, EFG1, WOR2, and CZF1 by RT-qPCR in both white and opaque cells of the WT, med5Δ/Δ, med16Δ/Δ, med9Δ/Δ, med20Δ/Δ, and med12Δ/Δ strains and in white cells of the med3Δ/Δ strain since we could not obtain opaque cells of this mutant. Figure 4A shows that most Mediator mutant strains, in general, did not exhibit dramatic differences in the expression profiles of the four key players. However, we observed the following differences in mRNA levels compared to WT white or opaque cell levels: med16Δ/Δ opaque cells had higher EFG1 and lower WOR2 levels; med20Δ/Δ white cells had a higher WOR2 level that correlated with their increased white-opaque switching frequency; med1Δ/Δ white cells had lower WOR1 and WOR2 levels, despite the relative stability of the opaque state; med3Δ/Δ white cells had much higher levels of WOR1 and WOR2 and a lower level of EFG1 but still could not switch to opaque; and med12Δ/Δ white cells had higher WOR2 and CZF1 levels but opaque cells had lower WOR1 and WOR2 levels that are consistent with the compromised opaque state stability in this strain. The lower levels of WOR1 and WOR2 transcript in the opaque cell culture do not appear to be a result of “contaminating” white cells that arise from the high opaque-white switching frequency in the med12Δ/Δ strain. Under conditions of ectopic Wor1 expression that maintain the med12Δ/Δ cells in the opaque state, the endogenous levels of Wor1 are still decreased relative to the WT levels (see below).
We used immunoblotting to determine whether Wor1 protein levels in selected mutants (Fig. 4B) exhibited a trend similar to that seen with respect to the steady-state mRNA levels revealed by RT-qPCR (Fig. 4A). Akin to the surprising upregulation of the Wor1 circuit in med3Δ/Δ white cells, we also detected an elevated level of Wor1 protein in this strain compared to the WT.
Ectopic Wor1 expression activates the Wor1 circuit in med3Δ/Δ and med12Δ/Δ mutants.
Spontaneous and induced white-opaque switching is highly impaired in the med12Δ/Δ mutant and undetectable in the med3Δ/Δ mutant. To determine how efficiently the Wor1 circuit could respond to the master regulator of the switch, we ectopically expressed Wor1 in these mutant strains. In mutant strains that have impaired white-opaque switching, overexpression of Wor1 was typically able to drive the formation of opaque cells in most previously tested mutant strains and of “opaque-like” cells in the wor1Δ/Δ and wor2Δ/Δ strains (38–41, 51, 52, 60). We hypothesized that the ability of Wor1 to stabilize the opaque state in the med3Δ/Δ strain was impaired, even when the self-sustaining feedback loop was activated. To test our hypothesis, we transformed the PMAL-WOR1-3HA transgene (51) into the med12Δ/Δ and med3Δ/Δ mutant strains. We assessed cell morphology and WOR1 expression levels in these cells grown under the noninduced (Glu) and induced (Mal) conditions (Table 3 and Fig. 5). Ectopic WOR1 expression led to 100% conversion to the opaque state in the med12Δ/Δ mutant with colony and cell morphology similar to those of the WT (data not shown). Although ectopic WOR1 expression in the med3Δ/Δ mutant drove the Wor1 circuit, the cells and colonies showed an altered morphology growing on SC+Mal+phloxine B plates that was distinct from that of typical opaque cells (see below). To show that the endogenous WOR1 autoactivation circuit was still functioning in the med3Δ/Δ mutant (and med12Δ/Δ mutant), we took advantage of the 3XHA tag fused with ectopic WOR1 to discriminate ectopically expressed WOR1 and endogenous WOR1. By using primers targeting the 3XHA of ectopic WOR1 and the 5′ untranslated region (5′UTR) of endogenous WOR1, we found that both the ectopic WOR1 and endogenous WOR1 were expressed at increased levels in the med3Δ/Δ mutant and med12Δ/Δ mutant under inducing conditions (Fig. 5A). These mutants showed a level of ectopic WOR1 expression similar to that seen with WT cells, both at the mRNA level and at the protein level. The ectopic WOR1 induced the expression of endogenous WOR1 in the mutants but at lower levels than those seen with the WT (Fig. 5A). We also tested ectopic and total Wor1 protein levels by using anti-HA antibodies and anti-Wor1 antibodies. As shown in Fig. 5B, the med3Δ/Δ and med12Δ/Δ mutants had a Wor1-3XHA level similar to that seen with the WT when being induced. However, they had less than half the concentration of total Wor1 protein compared to the WT. This difference could come only from a decrease in the endogenous Wor1 protein level. Under conditions of ectopic WOR1 expression, the med3Δ/Δ colonies showed wrinkling and reduced phloxine B staining (Fig. 5C) compared to the flat opaque WT colonies. Additionally, the cell dimension under liquid growth conditions was much less elongated in the mutant than in the WT cells (Fig. 5D). The wrinkled colony morphology of the med3Δ/Δ strain was a result of the ectopic expression of Wor1, as the med3Δ/Δ strain (without the Pmal-WOR1-3HA transgene) produced colonies on the inducing media (SC+Mal) (data not shown) that looked equivalent to those seen with the SC+Glu media. However, it is interesting that, even under noninducing conditions, the med3Δ/Δ cells adopted a somewhat intermediate phenotype between the white and opaque states (Fig. 5D; see also Fig. S2 in the supplemental material).
Table 3.
The induction of altered morphologies by ectopic expression of WOR1 and their stability in the med3Δ/Δ and med12Δ/Δ mutant strainsa
| C. albicans strain | Value for colonies |
|||
|---|---|---|---|---|
| After removal from Mal to Mal |
After removal from Mal to Glu |
|||
| Opaque (%) | n | Opaque (%) | n | |
| WT | <10−3 | 1,012 | <10−3 | 908 |
| WT; PMAL-WOR1-3HA | 100 | 1,119 | 79.0 ± 14.3 | 1,642 |
| med3Δ/Δ | <10−3 | 480 | <10−3 | 585 |
| med3Δ/Δ; PMAL-WOR1-3HA | 100* | 533 | <10−3 | 451 |
| med12Δ/Δ | <10−3 | 598 | <10−3 | 745 |
| med12Δ/Δ; PMAL-WOR1-3HA | 100 | 546 | 47.4 ± 18.3 | 1,090 |
Cells from the med3Δ/Δ strain with ectopically expressed WOR1-3HA under the control of the PMAL promoter formed an altered opaque colony morphology on SC+Mal+phloxine B plates. When opaque cells are removed from maltose plates to glucose plates, med3Δ/Δ grew as typical white colonies (Fig. 5C). At least 3 independent WOR1-overexpression strains for each WT and Mediator mutant were analyzed and showed similar results (data not shown).
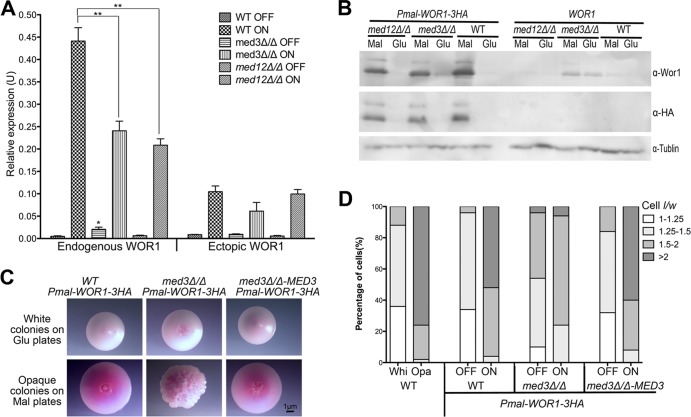
Fig 5.
Ectopic Wor1 expression activated the Wor1 circuit and altered morphologies in med3Δ/Δ and med12Δ/Δ mutants. (A) RT-qPCR analysis shows that ectopically expressed WOR1 induces endogenous WOR1 expression in both med3Δ/Δ and med12Δ/Δ mutants. Endogenous WOR1 was measured by primers targeting the 5′UTR of endogenous WOR1 (based on RNA-seq data from reference 63). Ectopic WOR1 was measured by primers targeting the end of the WOR1 ORF in junction with 3HA. The qPCR was performed in triplicate. Results were normalized to ACT1. Bars represent means ± SD. *, P < 0.05 (t test for the comparison between WT and med3Δ/Δ in SC+Glu); **, P < 0.01 (t test for comparisons between WT and med3Δ/Δ and med12Δ/Δ in SC+Mal). (B) Immunoblotting using an anti-HA antibody showed that ectopic Wor1-3HA protein expression was detected at similar levels in med3Δ/Δ and med12Δ/Δ mutants compared with the WT; however, densitometry of the Wor1 band (α-Wor1) band under conditions of ectopic Wor1 expression showed that the total amount of Wor1 was reduced by ∼50% in the med3Δ/Δ and med12Δ/Δ strains compared to the WT. An antibody against tubulin was used as a loading control. (C) A med3Δ/Δ mutant with ectopic Wor1 expression formed an altered opaque colony morphology. Cells were preincubated on either SC+Glu or SC+Mal plates for 3 days and replated onto phloxine B plates. Photos were taken with a Nikon SMZ1500 stereomicroscope (Nikon Instruments). (D) Measurement of cell dimensions showed that ectopic Wor1 expression induced altered cell morphology in a med3Δ/Δ mutant. The percentage of elongated cells (length/width > 2) was significantly less in the med3Δ/Δ mutant than in the WT and complemented strains. Cells were harvested from SC+Glucose (WT Whi and Opa, OFF) or SC+Mal (ON) liquid culture for microscopic analysis. A minimum of 50 cells was observed for each morphology evaluation. l, length; w, width (maximum diameter) (40). For representative figures of individual cell morphologies, see Fig. S2 in the supplemental material.
The self-sustaining property of the WOR1 feedback loop in WT cells ensures that once it is activated, it persists for many generations and maintains the opaque state (40). We had shown that this loop could be activated in the med3Δ/Δ and med12Δ/Δ mutants by ectopically expressed Wor1 to form cells of an altered morphology. To determine if this state was stable, we removed the opaque cells growing on SC+Mal to SC+Glu plates to shut down ectopic WOR1 expression. We then counted the percentage of opaque cells under this noninduced condition (Table 3). As expected, WT cells can maintain the opaque state, as the majority of colonies formed on SC+Glu were opaque. This ability was weakened in the med12Δ/Δ mutant, as half of colonies were opaque. This low percentage of opaque cells in the med12Δ/Δ mutant was likely the result of the high level of opaque-white switching in this mutant (Table 1). Consistent with the observation that the opaque state was unstable in the med3Δ/Δ mutant, we observed 100% reversion of the altered morphology to white cells upon removal from the induced condition to the noninduced condition. This feature distinguished the med12Δ/Δ mutant from the med3Δ/Δ mutant. Under conditions of ectopic Wor1 expression, both the med12Δ/Δ mutant and the med3Δ/Δ mutant exhibited activation of the Wor1 feedback loop at decreased levels, but only the med12Δ/Δ mutant cells could maintain the circuit once it was activated.
med3Δ/Δ mutant cells are incapable of mating.
C. albicans incorporates white-opaque switching into mating behavior—only MTL homozygous strains can switch and only opaque cells are capable of mating (66). We were interested in how mating was affected in the med3Δ/Δ mutant. This mutant exhibited several unique features: it did not have an opaque state that was stable enough for observation (Table 1); it upregulated opaque state-specific gene expression (WOR1, WOR2, OP4) and downregulated white state-specific gene expression (EFG1, WH11) in the white state (Fig. 4 and data not shown); and ectopic expression of Wor1 activated the Wor1 circuit but led to an altered morphology that was not stable when the ectopic Wor1 expression was shut down. In particular, we wanted to know if med3Δ/Δ white cells and the “enforced” altered-morphology cells could mate with cells of the opposite mating type. We compared the mating efficiency of these cells with WT MTL α/α cells (AZC19) to the mating efficiency of WT MTL a/a cells (AZC133) with WT MTL α/α cells on either YPD or YPMal plates. We never observed any successful mated cells from med3Δ/Δ white cells on media containing glucose or maltose (Table 4) or from med3Δ/Δ PMal-WOR1-3HA cells on glucose plates. However, for med3Δ/Δ PMal-WOR1-3HA cells growing on maltose plates, where ectopic Wor1 was expressed, we occasionally did observe one or two colonies on the doubly selective plates of the thousands tested. This result suggests that the deletion of the Med3 subunit severely impaired both the cell's ability to switch from white to opaque and its ability to mate efficiently. As controls, we also tested the mating efficiency of the med12Δ/Δ mutant either with or without Wor1 overexpression. In both cases, the cells showed mating efficiency similar to that of the WT strain when ectopic Wor1 was induced, largely due to the stabilization of the opaque state by Wor1 overexpression.
Table 4.
The med3Δ/Δ mutant shows highly impaired mating behaviora
| Strain | Mating efficiency |
|
|---|---|---|
| YPD | YPMal | |
| WT opaque | (3.6 ± 0.8) × 10−2 | (3.9 ± 1.5) × 10−2 |
| WT white | <10−5 | <10−5 |
| med3Δ/Δ | <10−5 | <10−5 |
| med3Δ/Δ; Pmal-WOR1-3HA | <10−5 | <10−4 |
| med12Δ/Δ opaque | (1.7 ± 0.9) × 10−3 | (4.2 ± 2.1) × 10−3 |
| med12Δ/Δ; Pmal-WOR1-3HA | (1.4 ± 0.2) × 10−3 | (1.5 ± 0.4) × 10−2 |
The quantitative mating assay was performed on YPD or YPMal plates supplemented with adenine for 3 days before cells were replated onto selective plates. All the strains listed in the table were MTLa/a and were mated with the same MTLα/α strain. The results shown in this table represent averages of the results of three independent experiments, with more than 1,000 colonies counted for each mating pair.
DISCUSSION
In all eukaryotes, Mediator has been shown to be a central hub of both positive and negative transcriptional regulation (7). Many mechanisms for Mediator have been invoked, including effects on initiation, reinitiation, elongation, and chromatin structure, via interactions with different factors (7, 8). The utilization of each of these mechanisms is likely to be dependent on different Mediator subunits and the identity of the protein, chromatin, and nucleic acids involved in the regulation of particular loci. Historically, the effect of particular Mediator mutations on transcription has been assessed by measuring differences in steady-state mRNA levels of specific genes. What emerges, in many cases, is a picture of Mediator as a molecular rheostat that facilitates certain levels of transcriptional activation or repression. Transcriptional switches, however, in which two or more stably heritable transcriptional states are adopted spontaneously or in response to an environmental cue, are of immense importance in microbes (67) and in many metazoan phenomena such as stem cell differentiation (68). Investigating the metastable white-opaque switch in C. albicans has allowed us to investigate the role of Mediator in the stability of epigenetic transcriptional states and how this relates to its effect on mRNA levels. Modeling studies have suggested that changes in expression levels of the key regulators of the switch have predictable effects on the directionality and frequency of switching (64). We have shown that different caMediator subunits can have dramatically different effects on the directionality, frequency, and environmental induction of epigenetic switching. These effects, however, are not always accompanied by the predicted changes in expression level of key regulators of the switch and suggest that Mediator can affect the stability of epigenetic states on multiple levels. The effect of each caMediator mutant on white-opaque switching has revealed important information about Mediator functionality that will be the basis for future mechanistic studies and provide important new information on the regulation of white-opaque switching.
Although components of the Cdk8 module, such as Med12, in yeast have been commonly associated with transcriptional repression (9), here the module appears to be necessary for high levels of activation of the WOR1 and WOR2 promoters. Of all the subunits tested, only the med12Δ/Δ mutant exhibited decreased levels of WOR1, and its required partner WOR2, in the opaque state (Fig. 4A). Consistent with a defect in the upregulation of WOR1, the med12Δ/Δ mutant showed a shift similar to that seen with a wor1Δ/+ heterozygote (34) in the kinetics of switching from opaque to white at 37°C (Fig. 3). It appears that this defect could be a result of inefficient activation of the WOR1 or WOR2 promoters by Wor1, as ectopic expression of Wor1 still leads to lower levels of the endogenous transcript. This finding is consistent with destabilization of the opaque state observed in the switching assays (Table 1). A role for the Cdk8 module in activated transcription is beginning to become more widely appreciated (69). Despite the lower activation levels of the Wor1 circuit in med12Δ/Δ opaque cells, the mutant is still capable of a mating efficiency that is comparable to that of the WT in the presence of ectopically expressed Wor1 (Table 4). We speculate that the decreased mating efficiency of med12Δ/Δ opaque cells in the absence of ectopically expressed Wor1 may result from opaque cells switching back to white during the course of the assay (Table 4). The med3Δ/Δ mutant, which also destabilizes the opaque state, provides an interesting contrast to the med12Δ/Δ strain.
As shown in studies based on work in S. cerevisiae, Med2, Med3, and Med15 are part of a submodule of the Tail that is primarily involved in activated transcription (70), although Med3 has been proposed to play a role in repression as well (71). Both the med3Δ/Δ and med15Δ/Δ strains are unable to switch to the opaque state (Table 1), even under environmental conditions that induce white-opaque switching up to 40-fold in WT cells (Table 2). In contrast to the med12Δ/Δ mutant, and despite its inability to switch, the med3Δ/Δ mutant appears to have partially derepressed the Wor1 circuit in white cells (Fig. 4). The med3Δ/Δ mutant, however, does not appear to have the potential to fully activate the Wor1 circuit, as the level of endogenous Wor1 driven by the ectopic expression of Wor1 is significantly lower than in the WT opaque cells (Fig. 5A and B). In addition to the lower levels of Wor1, the ectopic expression of Wor1 in the med3Δ/Δ mutant leads to a cell morphology that differs considerably from the traditional opaque morphology (Fig. 5C and D). Moreover, the med3Δ/Δ mutant, under conditions of ectopic Wor1 expression, has mating efficiency that is extremely reduced compared to that of the WT strain or even the med12Δ/Δ strain under conditions of ectopic Wor1 expression (Table 4). In combination, these results indicate that the Med2(Tlo)/Med3/Med15 Tail submodule has downstream effects from the Wor1 circuit that are necessary for true opaque cell morphology and mating behavior, beyond the direct effects on the Wor1 circuit itself. The finding that the med3Δ/Δ mutant cannot maintain the altered morphology after withdrawal of ectopic Wor1 expression (Table 3) indicates that these downstream effects likely provide feedback to the Wor1 circuit. Further analysis of the med3Δ/Δ mutant could help reveal the genes that define the opaque morphology and the mating behavior.
In contrast to the med3Δ/Δ and med12Δ/Δ mutants, the opaque state is stabilized relative to the white state in the med1Δ/Δ and med20Δ/Δ mutants (Table 1). Although both the med1Δ/Δ and med20Δ/Δ mutants exhibit decreased opaque-white switching, neither has an obvious change in the opaque mRNA levels of Wor1 that would explain the decrease (Fig. 4A). In the case of the med20Δ/Δ mutant, it is possible that increased levels of CZF1 could buffer fluctuations in Wor1 levels and lead to less frequent opaque-white switching. However, the increased CZF1 level in the opaque state does not appear to translate into decreased EFG1 levels such as the circuit would predict. Despite the med1Δ/Δ and med20Δ/Δ mutants having similar decreases in spontaneous opaque-white switching frequencies, the med20Δ/Δ mutant has a substantially longer delay in switching at 37°C (Fig. 3). It is possible that the signaling pathway that leads to switching at 37°C is partially functional at 25°C and that the decrease in spontaneous opaque-white switching at 25°C in the Med20 deletion results from a defect in this pathway in the med20Δ/Δ strain. It is still unclear how the signaling pathway from growth at 37°C to opaque-white switching occurs, and looking at the temporal pattern of gene expression changes upon switching at 37°C in the med20Δ/Δ mutant could shed new light on this issue. Given our recent studies of the importance of Mediator in silencing telomeres (26, 27), it is of interest that both the med1Δ/Δ and med20Δ/Δ mutants can be viewed as decreasing the stability of the epigenetically silenced (white) state of the Wor1 circuit. It is unclear what genes, if any, have been derepressed to achieve this result, although the increase in WOR2 expression in med20Δ/Δ white cells might be a clue (Fig. 4A). It is interesting that several histone deacetylase mutants also led to increased white-opaque switching combined with decreased (Hst1) (51) or unchanged (Hda1, Rpd31) (50) opaque-white switching. The med1Δ/Δ and med20Δ/Δ mutants, as well as the med9Δ/Δ and med16Δ/Δ mutants, may exert their effect through an impact on chromatin.
The med9Δ/Δ and med16Δ/Δ mutants belong to an unusual class of white-opaque switching mutants that increase switching frequencies in both directions (Table 1). The only other two mutations found to have a similar impact are deletion of CAC2, which is part of the CAF-1 complex that functions mainly in replication-coupled nucleosome assembly (52), and deletion of RPD3, a histone deacetylase associated with transcriptional repression (49). With the exception of the increased EFG1 expression in med16Δ/Δ opaque cells, there appear to be no significant changes in the expression levels of the key components of the Wor1 circuit (Fig. 4A). This finding is consistent with modeling studies (65) of the interlocking feedback loop that predict that simple changes in the expression level of a component(s) of the circuit would lead to unidirectional changes in switching frequency (i.e., an increase in Wor1 production leading to an increase in white-opaque switching frequency accompanied by a decreased opaque-white switching frequency). The med9Δ/Δ and med16Δ/Δ mutants appear to have roles in the stability of both of the epigenetic states. Two parameters in the modeling studies that can decrease the stability of both states are an increase in noise or a decrease in the cooperativity of promoter binding by the DNA-bound regulators. Assessing the noise in Wor1 expression in single cells by green fluorescent protein (GFP) tagging of Wor1 indicates that the med16Δ/Δ mutant has cellular Wor1-GFP noise in opaque cells similar to that seen with the WT or other mutants (see Fig. S1 in the supplemental material). An alternative explanation supported by previous studies is that there are specific chromatin structures associated with both epigenetic transcriptional states and that mutations which impair the formation of these structures can impair the stability of one or both states (50, 52). Previous studies of the med16Δ mutation in S. cerevisiae support the idea that Med16 could impact the stability of chromatin structures and epigenetic states (22, 26), although the mechanism is still unclear. It is interesting that deletion of the Med5 subunit, which is anchored through the Med16 subunit (Fig. 1 and 2) and affects telomeric silencing in S. cerevisiae (26), had no effect on white-opaque switching. Lastly, our studies of purified med16Δ/Δ Mediator have revealed an interesting change in protein-protein interactions that has occurred within the Mediator complex during fungal evolution. Since Med16 is required for the stable incorporation of the Med2/Med3/Med15 Tail submodule into scMediator (72), we were surprised by the dramatic differences in switching between the med16Δ/Δ strain and the med3Δ/Δ and med15Δ/Δ strains. Our purification of a med16Δ/Δ caMediator that retained the Med2(Tlo)/Med3/Med15 Tail submodule not only reconciled the switching data but revealed that, although there are orthologs for all the scMediator subunits in C. albicans, rewiring of interactions among the subunits can occur. These changes in the network of Mediator subunit interactions are reminiscent of the changes in the wiring of transcription factor networks among fungi (73) and could also potentially occur also through the exchange of weak protein-protein interactions during evolution.
Supplementary Material
ACKNOWLEDGMENTS
We thank Richard Bennett (Brown University), Haoping Liu (UC Irvine), Matt Lohse (UCSF), and Malcolm Whiteway (NRC/BRI) for strains and/or plasmids. We are very grateful to Deborah Hogan, Paula Sundstrom for advice and assistance during the course of experiments. We thank Zhong Li for assistance in writing the MATLAB code for single-cell GFP analysis.
This work was supported by National Institute of General Medical Sciences grant R01 GM62483 to L.C.M. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Footnotes
Published ahead of print 19 July 2013
Supplemental material for this article may be found at http://dx.doi.org/10.1128/EC.00137-13.
REFERENCES
- 1.Bonasio R, Tu S, Reinberg D. 2010. Molecular signals of epigenetic states. Science 330:612–616 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Margueron R, Reinberg D. 2010. Chromatin structure and the inheritance of epigenetic information. Nat. Rev. Genet. 11:285–296 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Moazed D. 2011. Mechanisms for the inheritance of chromatin states. Cell 146:510–518 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Acar M, Mettetal JT, van Oudenaarden A. 2008. Stochastic switching as a survival strategy in fluctuating environments. Nat. Genet. 40:471–475 [DOI] [PubMed] [Google Scholar]
- 5.Levy SF, Ziv N, Siegal ML. 2012. Bet hedging in yeast by heterogeneous, age-correlated expression of a stress protectant. PLoS Biol. 10:e1001325. 10.1371/journal.pbio.1001325 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Veening JW, Smits WK, Kuipers OP. 2008. Bistability, epigenetics, and bet-hedging in bacteria. Annu. Rev. Microbiol. 62:193–210 [DOI] [PubMed] [Google Scholar]
- 7.Conaway RC, Conaway JW. 2011. Function and regulation of the Mediator complex. Curr. Opin. Genet. Dev. 21:225–230 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Ansari SA, Morse RH. 2013. Mechanisms of Mediator complex action in transcriptional activation. Cell. Mol. Life Sci. 70:2743–2756 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Björklund S, Gustafsson CM. 2005. The yeast Mediator complex and its regulation. Trends Biochem. Sci. 30:240–244 [DOI] [PubMed] [Google Scholar]
- 10.Borggrefe T, Davis R, Erdjument-Bromage H, Tempst P, Kornberg RD. 2002. A complex of the Srb8, -9, -10 and -11 transcriptional regulatory proteins from yeast. J. Biol. Chem. 277:44202–44207 [DOI] [PubMed] [Google Scholar]
- 11.Chadick JZ, Asturias FJ. 2005. Structure of eukaryotic Mediator complexes. Trends Biochem. Sci. 30:264–271 [DOI] [PubMed] [Google Scholar]
- 12.Samuelsen CO, Baraznenok V, Khorosjutina O, Spahr H, Kieselbach T, Holmberg S, Gustafsson CM. 2003. TRAP230/ARC240 and TRAP240/ARC250 Mediator subunits are functionally conserved through evolution. Proc. Natl. Acad. Sci. U. S. A. 100:6422–6427 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Zhang A, Petrov KO, Hyun ER, Liu Z, Gerber SA, Myers LC. 2012. The Tlo proteins are stoichiometric components of C. albicans Mediator anchored via the Med3 subunit. Eukaryot. Cell 11:874–884 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Myers LC, Gustafsson CM, Hayashibara KC, Brown PO, Kornberg RD. 1999. Mediator protein mutations that selectively abolish activated transcription. Proc. Natl. Acad. Sci. U. S. A. 96:67–72 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.van de Peppel J, Kettelarij N, van Bakel H, Kockelkorn TT, van Leenen D, Holstege FC. 2005. Mediator expression profiling epistasis reveals a signal transduction pathway with antagonistic submodules and highly specific downstream targets. Mol. Cell 19:511–522 [DOI] [PubMed] [Google Scholar]
- 16.Brzovic PS, Heikaus CC, Kisselev L, Vernon R, Herbig E, Pacheco D, Warfield L, Littlefield P, Baker D, Klevit RE, Hahn S. 2011. The acidic transcription activator Gcn4 binds the mediator subunit Gal11/Med15 using a simple protein interface forming a fuzzy complex. Mol. Cell 44:942–953 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Jedidi I, Zhang F, Qiu H, Stahl SJ, Palmer I, Kaufman JD, Nadaud PS, Mukherjee S, Wingfield PT, Jaroniec CP, Hinnebusch AG. 2010. Activator Gcn4 employs multiple segments of Med15/Gal11, including the KIX domain, to recruit mediator to target genes in vivo. J. Biol. Chem. 285:2438–2455 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Herbig E, Warfield L, Fish L, Fishburn J, Knutson BA, Moorefield B, Pacheco D, Hahn S. 2010. Mechanism of Mediator recruitment by tandem Gcn4 activation domains and three Gal11 activator-binding domains. Mol. Cell. Biol. 30:2376–2390 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Reeves WM, Hahn S. 2005. Targets of the Gal4 transcription activator in functional transcription complexes. Mol. Cell. Biol. 25:9092–9102 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Holstege FC, Jennings EG, Wyrick JJ, Lee TI, Hengartner CJ, Green MR, Golub TR, Lander ES, Young RA. 1998. Dissecting the regulatory circuitry of a eukaryotic genome. Cell 95:717–728 [DOI] [PubMed] [Google Scholar]
- 21.Howard SC, Chang YW, Budovskaya YV, Herman PK. 2001. The Ras/PKA signaling pathway of Saccharomyces cerevisiae exhibits a functional interaction with the Sin4p complex of the RNA polymerase II holoenzyme. Genetics 159:77–89 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Jiang YW, Dohrmann PR, Stillman DJ. 1995. Genetic and physical interactions between yeast RGR1 and SIN4 in chromatin organization and transcriptional regulation. Genetics 140:47–54 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Liu Z, Myers LC. 2012. Med5 (Nut1) and Med17 (Srb4) are direct targets of Mediator histone H4 Tail interactions. PLoS One 7:e38416. 10.1371/journal.pone.0038416 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Singh H, Erkine AM, Kremer SB, Duttweiler HM, Davis DA, Iqbal J, Gross RR, Gross DS. 2006. A functional module of yeast mediator that governs the dynamic range of heat-shock gene expression. Genetics 172:2169–2184 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Song W, Treich I, Qian N, Kuchin S, Carlson M. 1996. SSN genes that affect transcriptional repression in Saccharomyces cerevisiae encode SIN4, ROX3, and SRB proteins associated with RNA polymerase II. Mol. Cell. Biol. 16:115–120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Zhu X, Liu B, Carlsten JO, Beve J, Nyström T, Myers LC, Gustafsson CM. 2011. Mediator influences telomeric silencing and cellular life span. Mol. Cell. Biol. 31:2413–2421 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zhu X, Zhang Y, Bjornsdottir G, Liu Z, Quan A, Costanzo M, Dávila López M, Westholm JO, Ronne H, Boone C, Gustafsson CM, Myers LC. 2011. Histone modifications influence mediator interactions with chromatin. Nucleic Acids Res. 39:8342–8354 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Elmlund H, Baraznenok V, Lindahl M, Samuelsen CO, Koeck PJ, Holmberg S, Hebert H, Gustafsson CM. 2006. The cyclin-dependent kinase 8 module sterically blocks Mediator interactions with RNA polymerase II. Proc. Natl. Acad. Sci. U. S. A. 103:15788–15793 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Peng J, Zhou JQ. 2012. The Tail-module of yeast Mediator complex is required for telomere heterochromatin maintenance. Nucleic Acids Res. 40:581–593 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Jiang YW, Stillman DJ. 1996. Epigenetic effects on yeast transcription caused by mutations in an actin-related protein present in the nucleus. Genes Dev. 10:604–619 [DOI] [PubMed] [Google Scholar]
- 31.Macatee T, Jiang YW, Stillman DJ, Roth SY. 1997. Global alterations in chromatin accessibility associated with loss of SIN4 function. Nucleic Acids Res. 25:1240–1247 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Lohse MB, Johnson AD. 2009. White-opaque switching in Candida albicans. Curr. Opin. Microbiol. 12:650–654 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Miller MG, Johnson AD. 2002. White-opaque switching in Candida albicans is controlled by mating-type locus homeodomain proteins and allows efficient mating. Cell 110:293–302 [DOI] [PubMed] [Google Scholar]
- 34.Huang G. 2012. Regulation of phenotypic transitions in the fungal pathogen Candida albicans. Virulence 3:251–261 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Morschhäuser J. 2010. Regulation of white-opaque switching in Candida albicans. Med. Microbiol. Immunol. 199:165–172 [DOI] [PubMed] [Google Scholar]
- 36.Tuch BB, Mitrovich QM, Homann OR, Hernday AD, Monighetti CK, De La Vega FM, Johnson AD. 2010. The transcriptomes of two heritable cell types illuminate the circuit governing their differentiation. PLoS Genet. 6:e1001070. 10.1371/journal.pgen.1001070 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Lohse MB, Zordan RE, Cain CW, Johnson AD. 2010. Distinct class of DNA-binding domains is exemplified by a master regulator of phenotypic switching in Candida albicans. Proc. Natl. Acad. Sci. U. S. A. 107:14105–14110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Huang G, Wang H, Chou S, Nie X, Chen J, Liu H. 2006. Bistable expression of WOR1, a master regulator of white-opaque switching in Candida albicans. Proc. Natl. Acad. Sci. U. S. A. 103:12813–12818 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Srikantha T, Borneman AR, Daniels KJ, Pujol C, Wu W, Seringhaus MR, Gerstein M, Yi S, Snyder M, Soll DR. 2006. TOS9 regulates white-opaque switching in Candida albicans. Eukaryot. Cell 5:1674–1687 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Zordan RE, Galgoczy DJ, Johnson AD. 2006. Epigenetic properties of white-opaque switching in Candida albicans are based on a self-sustaining transcriptional feedback loop. Proc. Natl. Acad. Sci. U. S. A. 103:12807–12812 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Zordan RE, Miller MG, Galgoczy DJ, Tuch BB, Johnson AD. 2007. Interlocking transcriptional feedback loops control white-opaque switching in Candida albicans. PLoS Biol. 5:e256. 10.1371/journal.pbio.0050256 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Soll DR. 2009. Why does Candida albicans switch? FEMS Yeast Res. 9:973–989 [DOI] [PubMed] [Google Scholar]
- 43.Yi S, Sahni N, Daniels KJ, Lu KL, Srikantha T, Huang G, Garnaas AM, Soll DR. 2011. Alternative mating type configurations (a/α versus a/a or α/α) of Candida albicans result in alternative biofilms regulated by different pathways. PLoS Biol. 9:e1001117. 10.1371/journal.pbio.1001117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Nguyen VQ, Sil A. 2008. Temperature-induced switch to the pathogenic yeast form of Histoplasma capsulatum requires Ryp1, a conserved transcriptional regulator. Proc. Natl. Acad. Sci. U. S. A. 105:4880–4885 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Jonkers W, Dong Y, Broz K, Kistler HC. 2012. The Wor1-like protein Fgp1 regulates pathogenicity, toxin synthesis and reproduction in the phytopathogenic fungus Fusarium graminearum. PLoS Pathog. 8:e1002724. 10.1371/journal.ppat.1002724 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Michielse CB, Becker M, Heller J, Moraga J, Collado IG, Tudzynski P. 2011. The Botrytis cinerea Reg1 protein, a putative transcriptional regulator, is required for pathogenicity, conidiogenesis, and the production of secondary metabolites. Mol. Plant Microbe Interact. 24:1074–1085 [DOI] [PubMed] [Google Scholar]
- 47.Michielse CB, van Wijk R, Reijnen L, Manders EM, Boas S, Olivain C, Alabouvette C, Rep M. 2009. The nuclear protein Sge1 of Fusarium oxysporum is required for parasitic growth. PLoS Pathog. 5:e1000637. 10.1371/journal.ppat.1000637 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Lee TI, Rinaldi NJ, Robert F, Odom DT, Bar-Joseph Z, Gerber GK, Hannett NM, Harbison CT, Thompson CM, Simon I, Zeitlinger J, Jennings EG, Murray HL, Gordon DB, Ren B, Wyrick JJ, Tagne JB, Volkert TL, Fraenkel E, Gifford DK, Young RA. 2002. Transcriptional regulatory networks in Saccharomyces cerevisiae. Science 298:799–804 [DOI] [PubMed] [Google Scholar]
- 49.Srikantha T, Tsai L, Daniels K, Klar AJ, Soll DR. 2001. The histone deacetylase genes HDA1 and RPD3 play distinct roles in regulation of high-frequency phenotypic switching in Candida albicans. J. Bacteriol. 183:4614–4625 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Hnisz D, Schwarzmüller T, Kuchler K. 2009. Transcriptional loops meet chromatin: a dual-layer network controls white-opaque switching in Candida albicans. Mol. Microbiol. 74:1–15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Stevenson JS, Liu H. 2011. Regulation of white and opaque cell-type formation in Candida albicans by Rtt109 and Hst3. Mol. Microbiol. 81:1078–1091 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Stevenson JS, Liu H. 2013. Nucleosome assembly factors CAF-1 and HIR modulate epigenetic switching frequencies in an H3K56 acetylation-associated manner in Candida albicans. Eukaryot. Cell 12:591–603 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Noble SM, Johnson AD. 2005. Strains and strategies for large-scale gene deletion studies of the diploid human fungal pathogen Candida albicans. Eukaryot. Cell 4:298–309 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Lohse MB, Johnson AD. 2010. Temporal anatomy of an epigenetic switch in cell programming: the white-opaque transition of C. albicans. Mol. Microbiol. 78:331–343 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Lavoie H, Sellam A, Askew C, Nantel A, Whiteway M. 2008. A toolbox for epitope-tagging and genome-wide location analysis in Candida albicans. BMC Genomics 9:578. 10.1186/1471-2164-9-578 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Booth LN, Tuch BB, Johnson AD. 2010. Intercalation of a new tier of transcription regulation into an ancient circuit. Nature 468:959–963 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Uwamahoro N, Qu Y, Jelicic B, Lo TL, Beaurepaire C, Bantun F, Quenault T, Boag PR, Ramm G, Callaghan J, Beilharz TH, Nantel A, Peleg AY, Traven A. 2012. The functions of Mediator in Candida albicans support a role in shaping species-specific gene expression. PLoS Genet. 8:e1002613. 10.1371/journal.pgen.1002613 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Guglielmi B, van Berkum N, Klapholz B, Bijma T, Boube M, Boschiero C, Bourbon H, Holstege FC, Werner M. 2004. A high resolution protein interaction map of the yeast Mediator complex. Nucleic Acids Res. 32:5379–5391 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Kang JS, Kim SH, Hwang MS, Han SJ, Lee YC, Kim YJ. 2001. The structural and functional organization of the yeast mediator complex. J. Biol. Chem. 276:42003–42010 [DOI] [PubMed] [Google Scholar]
- 60.Kachurina N, Turcotte B, Whiteway M. 2012. Motor protein Myo5p is required to maintain the regulatory circuit controlling WOR1 expression in Candida albicans. Eukaryot. Cell 11:626–637 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Alby K, Bennett RJ. 2009. Stress-induced phenotypic switching in Candida albicans. Mol. Biol. Cell 20:3178–3191 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Huang G, Srikantha T, Sahni N, Yi S, Soll DR. 2009. CO2 regulates white-to-opaque switching in Candida albicans. Curr. Biol. 19:330–334 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Slutsky B, Staebell M, Anderson J, Risen L, Pfaller M, Soll DR. 1987. “White-opaque transition”: a second high-frequency switching system in Candida albicans. J. Bacteriol. 169:189–197 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Lohse MB, Hernday AD, Fordyce PM, Noiman L, Sorrells TR, Hanson-Smith V, Nobile CJ, DeRisi JL, Johnson AD. 2013. Identification and characterization of a previously undescribed family of sequence-specific DNA-binding domains. Proc. Natl. Acad. Sci. U. S. A. 110:7660–7665 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Sriram K, Soliman S, Fages F. 2009. Dynamics of the interlocked positive feedback loops explaining the robust epigenetic switching in Candida albicans. J. Theor. Biol. 258:71–88 [DOI] [PubMed] [Google Scholar]
- 66.Bennett RJ, Johnson AD. 2005. Mating in Candida albicans and the search for a sexual cycle. Annu. Rev. Microbiol. 59:233–255 [DOI] [PubMed] [Google Scholar]
- 67.Avery SV. 2006. Microbial cell individuality and the underlying sources of heterogeneity. Nat. Rev. Microbiol. 4:577–587 [DOI] [PubMed] [Google Scholar]
- 68.Young RA. 2011. Control of the embryonic stem cell state. Cell 144:940–954 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Galbraith MD, Donner AJ, Espinosa JM. 2010. CDK8: a positive regulator of transcription. Transcription 1:4–12 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Ansari SA, Morse RH. 2012. Selective role of Mediator Tail module in the transcription of highly regulated genes in yeast. Transcription 3:110–114 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Papamichos-Chronakis M, Conlan RS, Gounalaki N, Copf T, Tzamarias D. 2000. Hrs1/Med3 is a Cyc8-Tup1 corepressor target in the RNA polymerase II holoenzyme. J. Biol. Chem. 275:8397–8403 [DOI] [PubMed] [Google Scholar]
- 72.Myers LC, Kornberg RD. 2000. Mediator of transcriptional regulation. Annu. Rev. Biochem. 69:729–749 [DOI] [PubMed] [Google Scholar]
- 73.Baker CR, Booth LN, Sorrells TR, Johnson AD. 2012. Protein modularity, cooperative binding, and hybrid regulatory states underlie transcriptional network diversification. Cell 151:80–95 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.