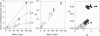Abstract
We have investigated the evolution of glycerol-3-phosphate dehydrogenase (Gpdh). The rate of amino acid replacements is 1 x 10(-10)/site/year when Drosophila species are compared. The rate is 2.7 times greater when Drosophila and Chymomyza species are compared; and about 5 times greater when any of those species are compared with the medfly Ceratitis capitata. This rate of 5 x 10(-10)/site/year is also the rate observed in comparisons between mammals, or between different animal phyla, or between the three multicellular kingdoms. We have also studied the evolution of Cu,Zn superoxide dismutase (Sod). The rate of amino acid replacements is about 17 x 10(-10)/site/year when comparisons are made between dipterans or between mammals, but only 5 x 10(-10) when animal phyla are compared, and only 3 x 10(-10) when the multicellular kingdoms are compared. The apparent decrease by about a factor of 5 in the rate of SOD evolution as the divergence between species increases can be consistent with the molecular clock hypothesis by assuming the covarion hypothesis (namely, that the number of amino acids that can change is constant, but the set of such amino acids changes from time to time and from lineage to lineage). However, we know of no model consistent with the molecular clock hypothesis that would account for the increase in the rate of GPDH evolution as the divergence between species increases.
Full text
PDF


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Ayala F. J. On the virtues and pitfalls of the molecular evolutionary clock. J Hered. 1986 Jul-Aug;77(4):226–235. doi: 10.1093/oxfordjournals.jhered.a110227. [DOI] [PubMed] [Google Scholar]
- Bewley G. C., Cook J. L., Kusakabe S., Mukai T., Rigby D. L., Chambers G. K. Sequence, structure and evolution of the gene coding for sn-glycerol-3-phosphate dehydrogenase in Drosophila melanogaster. Nucleic Acids Res. 1989 Nov 11;17(21):8553–8567. doi: 10.1093/nar/17.21.8553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cook J. L., Bewley G. C., Shaffer J. B. Drosophila sn-glycerol-3-phosphate dehydrogenase isozymes are generated by alternate pathways of RNA processing resulting in different carboxyl-terminal amino acid sequences. J Biol Chem. 1988 Aug 5;263(22):10858–10864. [PubMed] [Google Scholar]
- Fitch W. M., Ayala F. J. The superoxide dismutase molecular clock revisited. Proc Natl Acad Sci U S A. 1994 Jul 19;91(15):6802–6807. doi: 10.1073/pnas.91.15.6802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kimura M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol. 1980 Dec;16(2):111–120. doi: 10.1007/BF01731581. [DOI] [PubMed] [Google Scholar]
- Kimura M. Evolutionary rate at the molecular level. Nature. 1968 Feb 17;217(5129):624–626. doi: 10.1038/217624a0. [DOI] [PubMed] [Google Scholar]
- Kwiatowski J., Skarecky D., Ayala F. J. Structure and sequence of the Cu,Zn Sod gene in the Mediterranean fruit fly, Ceratitis capitata: intron insertion/deletion and evolution of the gene. Mol Phylogenet Evol. 1992 Mar;1(1):72–82. doi: 10.1016/1055-7903(92)90037-h. [DOI] [PubMed] [Google Scholar]
- Kwiatowski J., Skarecky D., Bailey K., Ayala F. J. Phylogeny of Drosophila and related genera inferred from the nucleotide sequence of the Cu,Zn Sod gene. J Mol Evol. 1994 May;38(5):443–454. doi: 10.1007/BF00178844. [DOI] [PubMed] [Google Scholar]
- Lee Y. M., Friedman D. J., Ayala F. J. Superoxide dismutase: an evolutionary puzzle. Proc Natl Acad Sci U S A. 1985 Feb;82(3):824–828. doi: 10.1073/pnas.82.3.824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li W. H., Ellsworth D. L., Krushkal J., Chang B. H., Hewett-Emmett D. Rates of nucleotide substitution in primates and rodents and the generation-time effect hypothesis. Mol Phylogenet Evol. 1996 Feb;5(1):182–187. doi: 10.1006/mpev.1996.0012. [DOI] [PubMed] [Google Scholar]
- Li W. H. Unbiased estimation of the rates of synonymous and nonsynonymous substitution. J Mol Evol. 1993 Jan;36(1):96–99. doi: 10.1007/BF02407308. [DOI] [PubMed] [Google Scholar]
- Ohta T. Role of very slightly deleterious mutations in molecular evolution and polymorphism. Theor Popul Biol. 1976 Dec;10(3):254–275. doi: 10.1016/0040-5809(76)90019-8. [DOI] [PubMed] [Google Scholar]
- Wilson A. C., Carlson S. S., White T. J. Biochemical evolution. Annu Rev Biochem. 1977;46:573–639. doi: 10.1146/annurev.bi.46.070177.003041. [DOI] [PubMed] [Google Scholar]
- von Kalm L., Weaver J., DeMarco J., MacIntyre R. J., Sullivan D. T. Structural characterization of the alpha-glycerol-3-phosphate dehydrogenase-encoding gene of Drosophila melanogaster. Proc Natl Acad Sci U S A. 1989 Jul;86(13):5020–5024. doi: 10.1073/pnas.86.13.5020. [DOI] [PMC free article] [PubMed] [Google Scholar]