Summary
One of the most fascinating challenges in neuroscience is the reconstruction of the connectivity map of the brain. Recent years have seen a rapid expansion in the field of connectomics, whose aim is to trace this map and understand its relationship with neural computation. Many different approaches, ranging from electron and optical microscopy to magnetic resonance imaging, have been proposed to address the connectomics challenge on various spatial scales and in different species. Here, we review the main technological advances in the microscopy techniques applied to connectomics, highlighting the potential and limitations of the different methods. Finally, we briefly discuss the role of connectomics in the Human Brain Project, the Future and Emerging Technologies (FET) Flagship recently approved by the European Commission.
Keywords: brain imaging , connectomics , electron microscopy , light microscopy , Human Brain Project
Introduction
The brain is probably the most complex structure in the known universe, complex enough to coordinate movements, gather and organize a vast amount of sensory data, perform abstract reasoning and develop new ideas. Understanding the mechanisms underlying brain function is therefore one of the biggest challenges of contemporary science.
Achieving a deeper understanding of the brain is a central issue not only for pure science; indeed, it would also have an enormous impact on society as a whole. In fact, in recent decades, the number of patients affected by central nervous system (CNS) disorders has increased dramatically, mainly because of the increase in life expectancy in developed countries and the direct/indirect effects of modern lifestyles on the brain. Although CNS disorders are not a major cause of death, they are a primary cause of disability for hundreds of millions of people worldwide ( Aarli et al. 2006 ), and a major cost for society in terms of health-care and their impact on the workforce, families and support groups.
The first insights into the structure of the brain were provided at the turn of the twentieth century by the Italian physician Camillo Golgi (1843–1926) , who discovered the reazione nera (black reaction), a staining method which randomly labels single neurons in their entirety ( Golgi,1885 ). The reazione nera was a true revolution for neuroanatomy, making it possible, for the first time, to observe single neurons inside the brain. Neurons appeared to be cells characterized by a long appendix (the axon) and a highly ramified ‘tree’ of dendrites. Golgi’s findings impressed a young Spanish pathologist, Santiago Ramón y Cajal (1852–1934) , who later refined the staining method and used it to provide the basis for what is now called the ‘neuron doctrine’ ( Ramon y Cajal, 1888). Although their respective visions of the function of the CNS were quite different, Golgi and Cajal shared the Nobel Prize in Physiology or Medicine in 1906.
Cajal’s ‘neuron doctrine’ formed the basis of the picture of the nervous system that we have inherited from the last century: the brain is an extremely complex network of neurons, each of which can transmit electrical signals over macroscopic distances along axons ( Kandel et al., 2000; Purves et al., 2004 ). This electrical activity regulates chemical communication between the axon and dendrites, belonging to other neurons, through specific subcellular structures called synapses. Chemical exchange at synapses can, in turn, elicit or inhibit electrical activity in downstream neurons, which subsequently activate or inhibit other neurons through synapses, and so on. In this way, the brain acts as a circuit in which elementary computational units (neurons) exchange information with other units in the network through unidirectional links (synapses). The unique features of the brain appear to emerge from its enormous number of units and links: a human brain consists of about 10 11 neurons connected by 10 14 –10 15 synapses ( Sporns et al., 2005). Even in the mouse, which is one of the most important animal models in brain research, we have to deal with about 10 8 neurons and 10 11 synapses ( Williams, 2000; Schüz and Palm, 1989). Adding to the complexity, connections between neural cells are not arranged in a lattice, but form a complex network lacking simple regularity ( Sporns et al., 2005). Finally, functional properties vary greatly between different neurons, which can be subdivided into an ever increasing number of neuronal types ( Kandel et al., 2000; Purves et al., 2004 ).
Our understanding of the functioning of neural cells, as well of the mechanisms underlying neuronal synapses, has advanced in recent years ( Sudhof, 2004; London and Hausser, 2005; Feldman, 2009). By comparison, our knowledge of neuronal connectivity in the brain lags a long way behind. Gross anatomical atlases of the brain are available, together with detailed descriptions, in terms of shape, functionality and genetic expression, of many types of neurons. However, a complete map of neuronal connections on a brain-wide scale, or ‘connectome’ ( Sporns et al., 2005), is still out of reach, because of the peculiar structure of the brain itself. In fact, although neurites are very small in diameter (sometimes 100 nm), they typically extend over large distances, even throughout the whole brain. For example, if we sum the length of all the branches of a single pyramidal neuron from the cerebral cortex, the result may exceed a centimeter in a mouse and a meter in a human brain ( Lichtman and Denk, 2011). Furthermore, neuronal processes are densely packed inside the brain, making it very difficult to distinguish between adjacent neurites and to discern between true synapses and random contacts ( Chklovskii et al., 2010; Mishchenko et al., 2010).
The pursuit of the connectome therefore needs imaging techniques capable of nanometric resolution (to distinguish adjacent processes) in cm-wide samples (to follow long-projecting axons). One of the most popular techniques allowing nanometric resolution is electron microscopy (EM) which, however, is characterized by very slow data acquisition rates. EM is thus inappropriate for brain-wide studies; conversely, it has been used successfully to reconstruct local circuitry in small regions ( Briggman et al., 2011; Helmstaedter et al., 2011), and the whole nervous system in very small organisms such as the nematode Caenorhabditis elegans , which has only 302 neurons ( White et al., 1986). In addition to imaging-related problems, connectomics also poses great challenges in terms of data management and analysis. In fact, mapping the relatively small brain of a mouse (1 cm 3 ) with nanometric resolution will produce datasets exceeding tens of petabytes. To have an idea of the size of such datasets, one might consider that the information content of the printed collection of the Library of the Congress of the United States amounts to about one hundred terabytes ( Kasthuri and Lichtman, 2007). As current information technology is not ready to cope with such datasets, technological efforts to achieve full-resolution connectomes risk proving useless. A more realistic and useful goal is that of a mesoscale connectome, or ‘projectome’, i.e. the mapping of long-range projections of small clusters of neurons, such as the monoaminergic systems ( Sarter and Parikh, 2005; Popova, 2006; Glenthøj, 1995), without synaptic resolution ( Bohland et al., 2009; Kasthuri and Lichtman, 2007). Little or nothing is known about the trajectories of brain-wide projections, or about their variability between different individuals. There is, however, generally expected to be a high degree of structural invariance at the mesoscopic level of description in healthy subjects ( Bohland et al., 2009), while there is growing evidence that aberrant wiring plays a crucial role in schizophrenia ( Bullmore et al., 1997), autism ( Frith, 2001) and dyslexia ( Démonet et al., 2004).
As ‘meso-connectomics’ in mammals seems to be within reach from a technical point of view – it is indeed possible in smaller animals ( Tay et al., 2011) – and the datasets are expected to be of manageable size, the attainment of comprehensive (whole-brain) fine neuroanatomy could be the next breakthrough in neuroscience. The possibility of studying the correlation between external factors (behavior, stress, drug treatment) and brain wiring in healthy and diseased animal models, by means of any kind of high-throughput screening, is radically changing our view of the brain and of its function.
Optical methodologies could be the key to meso-resolution brain atlases as they provide micron-scale resolution at relatively fast acquisition rates ( Silvestri et al. 2013b). Furthermore, optical microscopy can be combined with cell-specific fluorescence labeling, allowing one to image only limited subsets of neurons. In fact, the advent of transgenic mice, in which selected neuronal populations are labeled with fluorescent proteins, opened up extensive prospects for mapping projections of neuronal subsystems ( Feng et al., 2000; Livet et al., 2007).
On the other hand, optical techniques are limited by other factors: for instance, large-volume imaging can be performed only in fixed specimens. Whole-brain structural connectivity can be inferred by diffusion imaging (DI), a method based on magnetic resonance which takes advantage of water diffusion to detect bundles of axonal fibers ( Mori and Zhang, 2006; Clayden, 2013; Craddock et al., 2013). Although the resolution achievable with DI is not suitable for following single processes, and the contrast mechanism is non-specific, this imaging method can be used to map brain connectivity in living animals and humans. For a more thorough discussion about DI and other MRI-based methods we refer the reader to the review by Clayden in this issue of Functional Neurology ( Clayden, 2013). In any case, cellular-level brain connectivity can be studied at present only in post-mortem tissue. In the following sections we discuss in some detail the state of the art of the various microscopy methods used in the pursuit of the connectome, to better ascertain what might be a worthwhile approach to adopt in order to obtain atlases of neuronal connections ( DeFelipe, 2010) or projections ( Kasthuri and Lichtman, 2007).
The techniques described are usually applied to animal brains, and their application to human tissue is not always straightforward. However, mapping the brain of selected animals could greatly help to further understanding of the human brain by, for example, clarifying the effect that pathology can have on neuronal wiring. Thus, while imaging technologies are being adapted to human tissue, the animal connectome might represent an intermediate step to the human one.
Electron microscopy
Electron microscopy has played a central role in neuroscience ever since the first experimental confirmation of the neuron doctrine in the 1950s ( Palade and Palay, 1954). The main advantage of EM is the outstanding resolution, down to the nm scale, that can be achieved thanks to the short de Broglie wavelength of electrons. EM techniques can be divided primarily into wide-field transmission electron microscopy (TEM) and scanning electron microscopy (SEM). In both cases, the sample is placed in a high vacuum chamber for imaging: biological specimens must therefore be completely dehydrated and embedded in a hard resin. With the correct sample preparation, EM can be applied to both animal and human brain tissue.
In TEM a collimated electron beam is projected onto an ultrathin (∼ 40 nm) tissue slice, and the transmitted electrons are used to form an image on a phosphor plate, which, in turn, is captured by a charge-coupled device. Since image acquisition is inherently parallel, i.e. with all the image pixels being acquired simultaneously, TEM allows relatively fast imaging. The overall process is, however, dramatically slowed down because each ultrathin slice needs to be manually cut and prepared for imaging. Furthermore, manual handling of slices unavoidably leads to mechanical distortions, with consequent problems of layer misalignment and surface mismatching. Neuroanatomical analysis with TEM thus required heroic reconstruction efforts, as witnessed by the 15-year-long mapping of the 302 neurons of Caenorhabditis elegans ( White et al., 1986). Nevertheless, of the different EM techniques, TEM still affords the best spatial resolution ( Briggman and Bock, 2012).
In SEM a focused electron beam is scanned through the specimen, and back-scattered electrons are collected at each beam position, producing a raster map of the sample. Unlike what occurs with the transmission architecture, thick specimens can be observed, and axial resolution is afforded because of the limited penetration depth of the beam. The energy of the electron beam is typically much lower in SEM than in TEM, implying a lower signal-to-noise ratio and consequently a coarser spatial resolution. Nevertheless, SEM allows the use of automated strategies for sample cutting and handling. For example, an automated tape-collecting ultramicrotome ( Hayworth et al., 2006) has been devised which collects tissue sections on a tape well suited for automated SEM imaging. In other approaches ultrathin slices are removed from the specimen directly inside the microscope, in order to image its complete volume. Sections can be either mechanically cut away, as in serial block-face SEM ( Denk and Horstmann, 2004), or ablated by a focused ion beam (FIB-SEM) ( Knott et al., 2008).
Whatever the automated strategy used for data collection, EM is, in any case, inappropriate for whole-brain reconstructions because of its extremely slow frame rate, stemming from the raster scanning used for data collection (in SEM) and the manual handling of each single slice (in TEM). In fact, to our knowledge, the acquisition speed of EM never exceeds 10 μm 3 /s, implying that a cm 3 (the order of magnitude of a mouse brain) would require more than a thousand years to be imaged. Moreover, the enormous amount of data stemming from EM reconstructions needs to be analyzed manually [at least partially, given that no fully automated tools are available ( Helmstaedter et al., 2008)], thus further extending the time needed to extract quantitative information. However, EM is an invaluable technique for the study of dense local circuitry in 10–100 μm-sized volumes ( Fig. 1 ), as recently demonstrated by the mapping of neuronal wiring of the retina by Briggman et al. (2011) .
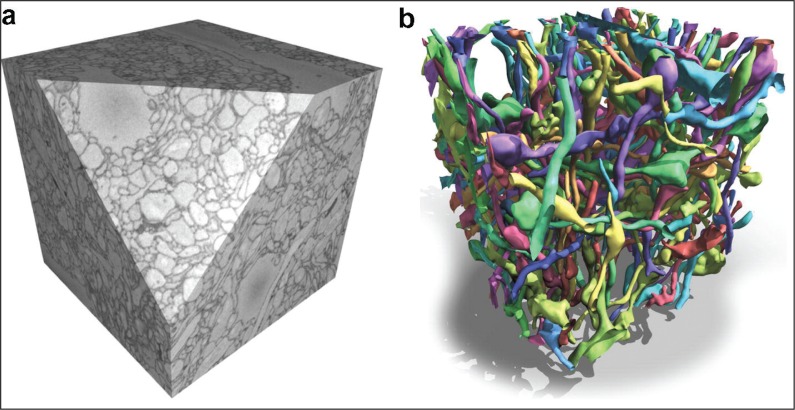
Figure 1 .
(a) A volume of about 12×12×15 nm in size from rabbit retina acquired with SBEM. (b) Reconstruction of randomly selected neuronal processes in the volume shown in (a). Reproduced with permission from Andres et al. (2012).
Optical methods
Integrating the ultra-resolution of EM with the in vivo imaging of DI is quite difficult, mainly because of the non-overlapping operative range (in terms of spatial resolution, acquisition time, field of view, sample preparation, etc.) of the two techniques. Optical approaches, which typically operate on the micron scale, can bridge the gap between high resolution in small volumes and low resolution in the whole brain. In practice, however, it is impossible to reconstruct the full neuronal network with light microscopy, since its spatial resolution is limited by diffraction to ∼ 200 nm, which is greater than the distance typically separating neurites. Super-resolution optical techniques, such as stimulated emission depletion microscopy ( Klar et al., 2000) or photo-activated localization microscopy ( Betzig et al., 2006), can break the diffraction barrier and increase the resolving power of optical microscopes to tens of nm. Anyway, a number of practical issues, above all the need for specialized chromophores and the long imaging time, still prevent the use of such approaches in large-scale neuroanatomical studies.
As regards diffraction-limited microscopy, the trick to get around the apparently difficult problem of neurite density is the same one that allowed Golgi and Ramon y Cajal to take the first steps into the realm of neuronal anatomy: if only a few neurons are visible, the resolution needed to observe them is lower, and can easily be afforded by optical techniques. Golgi’s reazione nera has recently been exploited to provide whole-brain mapping at subcellular level with micro-optical sectioning tomography (MOST) ( Li et al., 2010) and knife-edge scanning microscopy ( Mayerich et al., 2008). In these approaches the sample is embedded in a hard resin and sliced in ultrathin ribbons (∼ 1 μm) which are imaged just after slicing. The first demonstration of such techniques was in combination with Golgi staining, which randomly labels neurons with no distinctive feature. Recently, a new type of resin embedding, allowing good preservation of fluorescent proteins, has been exploited to image mice expressing either GFP or YFP in sparse neuronal subsets (fluorescence-MOST, fMOST) ( Gong et al., 2013). This improvement, thanks to the specificity of fluorescence labeling, opens up the possibility of studying the fine anatomy of selected neuronal populations within the whole brain, However, the time needed to image a single mouse brain with fMOST is about one month, limiting the practical applicability of the method.
Generally speaking, sparse labeling in combination with fluorescence optical microscopy would in principle allow the reconstruction of neuronal subsystems throughout the whole brain on the micron scale. However, in practice no such map has hitherto been produced. This is due to a combination of various factors. First of all, confocal and two-photon microscopy, which are probably the most popular optical fluorescence techniques suitable for volume imaging, require the use of objective lenses with a high numerical aperture (NA), which have limited working distances. The brain must then be cut in thin slices (no thicker than ∼ 50 μm for confocal and ∼ 7–800 μm for two-photon imaging), which have to be matched after imaging. Surface distortion is more severe than in EM, since the tissue is not embedded in a hard resin, and volume reconstruction is thus more challenging. Furthermore, both confocal microscopy and two-photon microscopy are point-scanning techniques, with pixel dwell times of the order of μs at least ( Pawley, 2006; Conchello and Lichtman, 2005; Helmchen and Denk, 2005). The image acquisition rate is therefore still too low to cope with large volumes. Thanks to their high resolution and contrast, these methods are, however, very well suited to neuroanatomical studies of brain subregions ( Hama et al., 2011) or small animals ( Tay et al., 2011), or to whole-brain reconstruction with sparse axial sampling ( Ragan et al., 2012).
A promising technique to overcome the above-mentioned limitations of both confocal and two-photon microscopy, i.e. limited speed and need for slicing, is light sheet fluorescence microscopy ( Keller and Dodt, 2012). In this technique optical sectioning is achieved in a wide-field detection scheme through selective illumination of the focal plane by means of a sheet of light ( Huisken and Stainier, 2009). Thus, a single plane inside the specimen is imaged in one step and not sequentially as in the point-scanning technique. Furthermore, since optical sectioning is independent of the NA of detection optics ( Mertz, 2011), long working distance objective lenses can be used, preventing sample sectioning.
Coupled to tissue clearing protocols, based on refractive index matching ( Becker et al., 2012), light sheet microscopy has been exploited to image large biological samples, including whole mouse brains ( Dodt et al., 2007). To increase image contrast, light sheet illumination has recently been integrated with confocal detection ( Silvestri et al., 2012; Fahrbach and Rohrbach, 2012; Baumgart and Kubitscheck, 2012; Silvestri et al., 2013a). Confocal light sheet microscopy allows reconstruction of fluorescently-labeled entire mouse brains with a resolution of a few microns and with an imaging time of ∼ 72 hours per brain ( Silvestri et al., 2012). While neuronal soma can easily be detected ( Fig. 2 ), the signal-to-background ratio affordable with this technique is too low allow neuronal projections to be mapped in a reliable and repeatable manner.
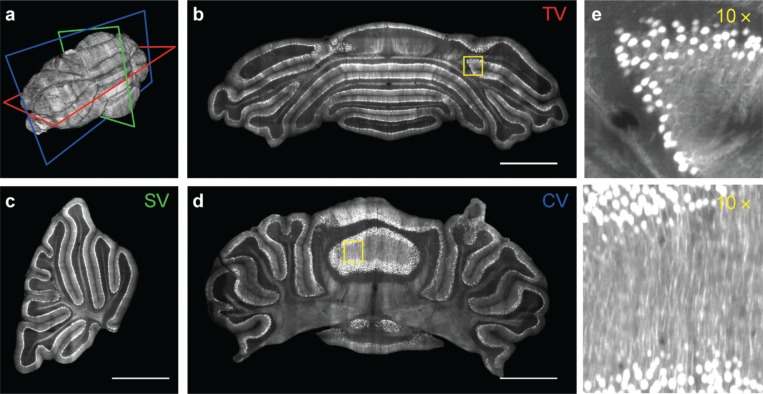
Figure 2 .
Neuroanatomy of Purkinje cells in the whole cerebellum imaged with confocal light sheet microscopy. (a) 3D volume rendering of a PND-10 L7-GFP mouse cerebellum. The superimposed planes refer to transverse (red), sagittal (green) and coronal (blue) digital sections shown in panels (b), (c) and (d) respectively. (b–d) Maximum intensity projections of 40 μm thick slabs. Scale bars, 1 mm. (e, f) 10 × magnification of the regions highlighted by the yellow boxes in panels (b) and (d). The lookup table saturates 2% of pixels for better visibility. Reproduced with permission from Silvestri et al. (2012).
The low levels of signal detectable in cleared samples with light sheet microscopy are due mainly to the clearing procedure itself. In fact, on the one hand, GFP fluorescence is impaired by the organic solvents used to render the tissue transparent. Novel clearing methods based on aqueous, fluorescence-friendly solutions ( Hama et al., 2011; Chung et al., 2013) may help to enhance the signal. On the other hand, there are no long working distance objective lenses corrected for the clearing solution used, and the refractive index mismatch between the design medium of the optics and the clearing agent introduces spherical aberrations which reduce image contrast by more than an order of magnitude ( Silvestri et al., 2013c). Correction of aberrations by means of adaptive optics ( Booth et al., 1998) may help to recover contrast and resolution, eventually allowing projection tracing in the whole mouse brain.
Concluding remarks
The reconstruction at cellular level of the neuronal wiring inside the brain is one of the greatest challenges facing contemporary neuroscience. However, to date it has been impossible to visualize entire neuronal networks across the whole brain. In fact, each of the methods that could potentially tackle the connectomics issue is, in practice, limited by critical drawbacks. For instance, EM can be used to reconstruct fine circuitry with nm resolution ( Briggman et al., 2011), but the acquisition rate is too low to allow the mapping of large volumes. On the other hand, large fiber bundles across the whole brain can be imaged in vivo with DI, but the resolution is too coarse to detect many single neuronal processes ( Clayden, 2013). Moreover, the non-specificity of DI contrast limits the classification of the observed fiber bundles.
Novel optical approaches, such as fMOST ( Gong et al., 2013) and confocal light sheet microscopy ( Silvestri et al., 2012), have the potential to bridge the gap between ultra-resolution and whole-brain imaging, opening up the possibility of investigating the projection patterns of specific neuronal subsystems. For example, axonal projections of localized nuclei [e.g. serotonergic ( Popova, 2006), cholinergic ( Sarter and Parikh, 2005), or dopaminergic systems ( Glenthøj, 1995)], corticothalamic circuits ( Briggs and Usrey, 2008), and interhemispheric connections ( Schulte and Muller-Oehring, 2010), etc., can be reconstructed using appropriate transgenic mouse models ( Feng et al.. 2000), retrograde labeling ( Marshel et al., 2010) or viral strategies ( Moriyoshi et al., 1996; Niedworok et al., 2012).
As we have shown in this article, all the imaging techniques tackling the connectomics challenge have their distinctive features in terms of imaging speed, contrast and resolution. In our opinion, the big puzzle of neuronal connectivity can be solved only if we put all these small pieces together. Data from EM, optical imaging and DI should be integrated in a multiscale/multiresolution model carrying different kinds of information: local circuitry, long-range projections, whole-brain cytoarchitecture. The goal of the Human Brain Project (HBP) is, precisely, to pool all our knowledge about the brain into unifying models, and to use these to simulate and even predict brain structure and function ( Markram, 2013; D’Angelo et al., 2013; Calimera et al., 2013; Redolfi et al., 2013). Within this ambitious project, strategic neuroanatomical data produced using various techniques will be integrated with data already present in the literature in order to feed brain models and obtain statistical predictions about those parts of the connectome map that are still unknown. Furthermore, brain simulations in the HBP will elucidate the coupling between structural and functional connectivity in order to provide a clearer view of how neuronal circuits process information in the brain ( Friston, 2011; Bargmann and Marder, 2013). Integration of structural and functional data will be a crucial step to gain insight into the big picture of the human brain.
The various types of information (anatomical, functional, molecular, etc.) gathered during the HBP will complete the data produced in other large-scale projects, such as the Human Connectome Project ( Van Essen et al., 2013) and the BRAIN initiative ( Insel et al., 2013). In addition, the integrative approach of the HBP will make it possible to exploit the collected data in brain simulations, inferring structural and functional brain rules from incomplete information. Thus, in the HBP some pieces of the big puzzle of the human brain will be inferred from the ones we already know, allowing neuroscience to move faster towards a more comprehensive and detailed view of the complex architecture of the brain.
Acknowledgments
This work has received funding from LASERLAB-EUROPE (grant agreements n° 228334 and 284464, EU’s Seventh Framework Programme) and has been supported by a Human Frontier Science Program research grant (RGP0027/2009), by the Italian Ministry for Education, University and Research in the framework of the Flagship Project NANOMAX, and by the Italian Ministry of Health in the framework of the ‘Stem Cells Call for Proposals’. This work has been carried out in the framework of the activities of the ICON foundation supported by “Ente Cassa di Risparmio di Firenze”. This work is part of the activities of the European Flagship Human Brain Project.
References
- Aarli JA , Dua T , Janca A , et al., editors. Neurological Disorders: Public Health Challenges . World Health Organization ; 2006 . [Google Scholar]
- Andres B , Koethe U , Kroeger T , et al. 3D segmentation of SBFSEM images of neuropil by a graphical model over supervoxel boundaries . Med Image Anal . 2012 ; 16 : 796 – 805 . doi: 10.1016/j.media.2011.11.004. [DOI] [PubMed] [Google Scholar]
- Bargmann CI , Marder E . From the connectome to brain function . Nat Methods . 2013 ; 10 : 483 – 490 . doi: 10.1038/nmeth.2451. [DOI] [PubMed] [Google Scholar]
- Baumgart E , Kubitscheck U . Scanned light sheet microscopy with confocal slit detection . Opt Express . 2012 ; 20 : 21805 – 21814 . doi: 10.1364/OE.20.021805. [DOI] [PubMed] [Google Scholar]
- Becker K , Jährling N , Saghafi S , et al. Chemical clearing and dehydration of GFP expressing mouse brains. PLoS One. 2012 ;7: e33916 . doi: 10.1371/journal.pone.0033916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Betzig E , Patterson GH , Sougrat R , et al. Imaging intra-cellular fluorescent proteins at nanometer resolution . Science . 2006 ; 313 : 1642 – 1645 . doi: 10.1126/science.1127344. [DOI] [PubMed] [Google Scholar]
- Bohland JW , Wu C , Barbas H , et al. A proposal for a coordinated effort for the determination of brainwide neuroanatomical connectivity in model organisms at a mesoscopic scale . PLoS Comput Biol. 2009 ;5: e1000334 . doi: 10.1371/journal.pcbi.1000334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Booth MJ , Neil MAA , Wilson T . Aberration correction for confocal imaging in refractive-index-mismatched media . Journal of Microscopy . 1998 ; 192 : 90 – 98 . [Google Scholar]
- Briggman KL , Bock DD . Volume electron microscopy for neuronal circuit reconstruction . Curr Opin Neurobiol . 2012 ; 22 : 154 – 161 . doi: 10.1016/j.conb.2011.10.022. [DOI] [PubMed] [Google Scholar]
- Briggman KL , Helmstaedter M , Denk W . Wiring specificity in the direction-selectivity circuit of the retina . Nature . 2011; 471 : 183 – 188 . doi: 10.1038/nature09818. [DOI] [PubMed] [Google Scholar]
- Briggs F , Usrey WM . Emerging views of corticothalamic function . Curr Opin Neurobiol . 2008 ; 18 : 403 – 407 . doi: 10.1016/j.conb.2008.09.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bullmore ET , Frangou S , Murray RM . The dysplastic net hypothesis: an integration of developmental and dysconnectivity theories of schizophrenia . Schizophr Res . 1997 ; 28 : 143 – 156 . doi: 10.1016/s0920-9964(97)00114-x. [DOI] [PubMed] [Google Scholar]
- Calimera A , Poncino M , Macii E . Human brain project and neuromorphic computing . Funct Neurol . 2013 ; 28 : 191 – 196 . [PMC free article] [PubMed] [Google Scholar]
- Chklovskii DB , Vitaladevuni S , Scheffer LK . Semi-automated reconstruction of neural circuits using electron microscopy . Curr Opin Neurobiol . 2010 ; 20 : 667 – 675 . doi: 10.1016/j.conb.2010.08.002. [DOI] [PubMed] [Google Scholar]
- Chung K , Wallace J , Kim SY , et al. Structural and molecular interrogation of intact biological systems . Nature . 2013 ; 497 : 332 – 337 . doi: 10.1038/nature12107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clayden JD . Imaging connectivity: MRI and the structural networks of the brain . Funct Neurol . 2013 ; 28 : 197 – 203 . [PMC free article] [PubMed] [Google Scholar]
- Conchello JA , Lichtman JW . Optical sectioning microscopy . Nat Methods . 2005 ; 2 : 920 – 931 . doi: 10.1038/nmeth815. [DOI] [PubMed] [Google Scholar]
- Craddock RC , Jbabdi S , Yan CG , et al. Imaging human connectomes at the macroscale . Nat Methods . 2013 ; 10 : 524 – 539 . doi: 10.1038/nmeth.2482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- D’Angelo E , Solinas S , Garrido J . Realistic modeling of neurons and networks: towards brain simulation . Funct Neurol . 2013 ; 28 : 153 – 166 . [PMC free article] [PubMed] [Google Scholar]
- DeFelipe J . From the connectome to the synaptome: an epic love story . Science . 2010 ; 330 : 1198 – 1201 . doi: 10.1126/science.1193378. [DOI] [PubMed] [Google Scholar]
- Démonet JF , Taylor MJ , Chaix Y . Developmental dyslexia . Lancet . 2004 ; 363 : 1451 – 1460 . doi: 10.1016/S0140-6736(04)16106-0. [DOI] [PubMed] [Google Scholar]
- Denk W , Horstmann H . Serial block-face scanning electron microscopy to reconstruct three-dimensional tissue anostructure. PLoS Biol. 2004 ;2: e329 . doi: 10.1371/journal.pbio.0020329. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dodt HU , Leischner U , Schierloh A , et al. Ultramicroscopy: three-dimensional visualization of neuronal networks in the whole mouse brain . Nat Methods . 2007 ; 4 : 331 – 336 . doi: 10.1038/nmeth1036. [DOI] [PubMed] [Google Scholar]
- Fahrbach FO , Rohrbach A . Propagation stability of self-reconstructing Bessel beams enables contrast-enhanced imaging in thick media . Nat Commun . 2012 ; 3 : 632 . doi: 10.1038/ncomms1646. [DOI] [PubMed] [Google Scholar]
- Feldman DE . Synaptic mechanisms for plasticity in neo-cortex . Annu Rev Neurosci . 2009 ; 32 : 33 – 55 . doi: 10.1146/annurev.neuro.051508.135516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feng G , Mellor RH , Bernstein M , et al. Imaging neuronal subsets in transgenic mice expressing multiple spectral variants of GFP . Neuron . 2000 ; 28 : 41 – 51 . doi: 10.1016/s0896-6273(00)00084-2. [DOI] [PubMed] [Google Scholar]
- Friston KJ . Functional and effective connectivity: a review . Brain Connect . 2011 ; 1 : 13 – 36 . doi: 10.1089/brain.2011.0008. [DOI] [PubMed] [Google Scholar]
- Frith U . Mind blindness and the brain in autism . Neuron . 2001 ; 32 : 969 – 979 . doi: 10.1016/s0896-6273(01)00552-9. [DOI] [PubMed] [Google Scholar]
- Glenthøj BY . The brain dopaminergic system. Pharmacological, behavioural and electrophysiological studies . Dan Med Bull . 1995 ; 42 : 1 – 21 . [PubMed] [Google Scholar]
- Golgi C . Sulla fina anatomia degli organi centrali del sistema nervoso. 1885 Reggio Emilia, Calderini. [Google Scholar]
- Gong H , Zeng S , Yan C , et al. Continuously tracing brain-wide long-distance axonal projections in mice at a one-micron voxel resolution . Neuroimage . 2013 ; 74 : 87 – 98 . doi: 10.1016/j.neuroimage.2013.02.005. [DOI] [PubMed] [Google Scholar]
- Hama H , Kurokawa H , Kawano H , et al. Scale: a chemical approach for fluorescence imaging and reconstruction of transparent mouse brain . Nat Neurosci . 2011 ; 14 : 1481 – 1488 . doi: 10.1038/nn.2928. [DOI] [PubMed] [Google Scholar]
- Hayworth KJ , Kasthuri N , Schalek R , et al. Automating the collection of ultrathin serial sections for large volume TEM reconstructions . Microscopy and Microanalysis. 2006 ;12(Suppl 2): 86 – 87 . [Google Scholar]
- Helmchen F , Denk W . Deep tissue two-photon microscopy . Nat Methods . 2005 ; 2 : 932 – 940 . doi: 10.1038/nmeth818. [DOI] [PubMed] [Google Scholar]
- Helmstaedter M , Briggman KL , Denk W . 3D structural imaging of the brain with photons and electrons . Curr Opin Neurobiol . 2008 ; 18 : 633 – 641 . doi: 10.1016/j.conb.2009.03.005. [DOI] [PubMed] [Google Scholar]
- Helmstaedter M , Briggman KL , Denk W . High-accuracy neurite reconstruction for high-throughput neuroanatomy . Nat Neurosci . 2011 ; 14 : 1081 – 1088 . doi: 10.1038/nn.2868. [DOI] [PubMed] [Google Scholar]
- Huisken J , Stainier DY . Selective plane illumination microscopy techniques in developmental biology . Development . 2009 ; 136 : 1963 – 1975 . doi: 10.1242/dev.022426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Insel TR , Landis SC , Collins FS . Research priorities. The NIH BRAIN Initiative . Science. 2013 ; 340 : 687 – 688 . doi: 10.1126/science.1239276. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kandel ER , Schwartz JH , Jessell TM . Principles of Neural Science. 4th ed . New York : McGraw-Hill, Health Professions Division ; 2000 . [Google Scholar]
- Kasthuri N , Lichtman JW . The rise of the ‘projectome’ . Nat Methods . 2007 ; 4 : 307 – 308 . doi: 10.1038/nmeth0407-307. [DOI] [PubMed] [Google Scholar]
- Keller PJ , Dodt HU . Light sheet microscopy of living or cleared specimens . Curr Opin Neurobiol . 2012 ; 22 : 138 – 143 . doi: 10.1016/j.conb.2011.08.003. [DOI] [PubMed] [Google Scholar]
- Klar TA , Jakobs S , Dyba M , et al. Fluorescence microscopy with diffraction resolution barrier broken by stimulated emission . Proc Natl Acad Sci U S A . 2000 ; 97 : 8206 – 8210 . doi: 10.1073/pnas.97.15.8206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knott G , Marchman H , Wall D , et al. Serial section scanning electron microscopy of adult brain tissue using focused ion beam milling . J Neurosci . 2008 ; 28 : 2959 – 2964 . doi: 10.1523/JNEUROSCI.3189-07.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li A , Gong H , Zhang B , et al. Micro-optical sectioning tomography to obtain a high-resolution atlas of the mouse brain . Science . 2010 ; 330 : 1404 – 1408 . doi: 10.1126/science.1191776. [DOI] [PubMed] [Google Scholar]
- Lichtman JW , Denk W . The big and the small: challenges of imaging the brain’s circuits . Science . 2011 ; 334 : 618 – 623 . doi: 10.1126/science.1209168. [DOI] [PubMed] [Google Scholar]
- Livet J , Weissman TA , Kang H , et al. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system . Nature . 2007 ; 450 : 56 – 62 . doi: 10.1038/nature06293. [DOI] [PubMed] [Google Scholar]
- London M , Hausser M . Dendritic computation . Annu Rev Neurosci . 2005 ; 28 : 503 – 532 . doi: 10.1146/annurev.neuro.28.061604.135703. [DOI] [PubMed] [Google Scholar]
- Markram H . Seven challanges for neuroscience . Funct Neurol . 2013 ; 28 : 145 – 151 . [PMC free article] [PubMed] [Google Scholar]
- Marshel JH , Mori T , Nielsen KJ , et al. Targeting single neuronal networks for gene expression and cell labeling in vivo . Neuron . 2010 ; 67 : 562 – 574 . doi: 10.1016/j.neuron.2010.08.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mayerich D , Abbott L , McCormick B . Knife-edge scanning microscopy for imaging and reconstruction of three-dimensional anatomical structures of the mouse brain . J Microsc . 2008 ; 231 : 134 – 143 . doi: 10.1111/j.1365-2818.2008.02024.x. [DOI] [PubMed] [Google Scholar]
- Mertz J . Optical sectioning microscopy with planar or structured illumination . Nat Methods . 2011 ; 8 : 811 – 819 . doi: 10.1038/nmeth.1709. [DOI] [PubMed] [Google Scholar]
- Mishchenko Y , Hu T , Spacek J , et al. Ultrastructural analysis of hippocampal neuropil from the connectomics perspective . Neuron . 2010 ; 67 : 1009 – 1020 . doi: 10.1016/j.neuron.2010.08.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mori S , Zhang J . Principles of diffusion tensor imaging and its applications to basic neuroscience research . Neuron . 2006 ; 51 : 527 – 539 . doi: 10.1016/j.neuron.2006.08.012. [DOI] [PubMed] [Google Scholar]
- Moriyoshi K , Richards LJ , Akazawa C , et al. Labeling neural cells using adenoviral gene transfer of membrane-targeted GFP . Neuron . 1996 ; 16 : 255 – 260 . doi: 10.1016/s0896-6273(00)80044-6. [DOI] [PubMed] [Google Scholar]
- Niedworok CJ , Schwarz I , Ledderose J , et al. Charting monosynaptic connectivity maps by two-color light-sheet fluorescence microscopy . Cell Rep . 2012 ; 2 : 1375 – 1386 . doi: 10.1016/j.celrep.2012.10.008. [DOI] [PubMed] [Google Scholar]
- Palade GE , Palay SL . Electron microscope observations of interneuronal and neuromuscular synapses . Anatomical Record . 1954 ; 118 : 335 – 336 . [Google Scholar]
- Pawley JB . Handbook of Biological Confocal Microscopy. 3rd ed . New York, NY : Springer ; 2006 . [Google Scholar]
- Popova NK . From genes to aggressive behavior: the role of serotonergic system . Bioessays . 2006 ; 28 : 495 – 503 . doi: 10.1002/bies.20412. [DOI] [PubMed] [Google Scholar]
- Purves D , Augustine GJ , Fitzpatrick D , et al., editors. Neuroscience. Third Edition . Sunderland, Mass: Sinauer Associates Inc ; 2004 . [Google Scholar]
- Ragan T , Kadiri LR , Venkataraju KU , et al. Serial two-photon tomography for automated ex vivo mouse brain imaging . Nat Methods . 2012 ; 9 : 255 – 258 . doi: 10.1038/nmeth.1854. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramon y Cajal S . Estructura de los centros nerviosos de la aves . Rev Trim Histol Norm Pat . 1888 ; 1 : 1 – 10 . [Google Scholar]
- Redolfi A , Frisoni GB . Brain investigation and brain conceptualization . Funct Neurol . 2013 ; 28 : 175 – 190 . [PMC free article] [PubMed] [Google Scholar]
- Sarter M , Parikh V . Choline transporters, cholinergic transmission and cognition . Nat Rev Neurosci . 2005 ; 6 : 48 – 56 . doi: 10.1038/nrn1588. [DOI] [PubMed] [Google Scholar]
- Schulte T , Muller-Oehring EM . Contribution of callosal connections to the interhemispheric integration of visuomotor and cognitive processes . Neuropsychol Rev . 2010 ; 20 : 174 – 190 . doi: 10.1007/s11065-010-9130-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schüz A , Palm G . Density of neurons and synapses in the cerebral cortex of the mouse . J Comp Neurol . 1989 ; 286 : 442 – 455 . doi: 10.1002/cne.902860404. [DOI] [PubMed] [Google Scholar]
- Silvestri L , Allegra Mascaro AL , Costantini I , et al. Correlative two-photon and light sheet microscopy. Methods. 2013a ;pii doi: 10.1016/j.ymeth.2013.06.013. S1046-2023(13)00214-4. [DOI] [PubMed] [Google Scholar]
- Silvestri L , Allegra Mascaro AL , Lotti J , et al. Advanced optical techniques to explore brain structure and function . Journal of Innovative Optical Health Sciences . 2013b ; 06 ( 01 ): 1230002 . [Google Scholar]
- Silvestri L , Bria A , Sacconi L , et al. Confocal light sheet microscopy: micron-scale neuroanatomy of the entire mouse brain . Opt Express . 2012 ; 20 : 20582 – 20598 . doi: 10.1364/OE.20.020582. [DOI] [PubMed] [Google Scholar]
- Silvestri L , Sacconi L , Pavone FS . Light sheet microscopy of cleared mouse brains: aberrations effects caused by refractive index mismatch . Proceedings of the SPIE. 2013c ; 8804 : 880405 . [Google Scholar]
- Sporns O , Tononi G , Kötter R . The human connectome: a structural description of the human brain. PLoS Comput Biol. 2005 ;1: e42 . doi: 10.1371/journal.pcbi.0010042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sudhof TC . The synaptic vesicle cycle . Annu Rev Neurosci . 2004 ; 27 : 509 – 547 . doi: 10.1146/annurev.neuro.26.041002.131412. [DOI] [PubMed] [Google Scholar]
- Tay TL , Ronneberger O , Ryu S , et al. Comprehensive catecholaminergic projectome analysis reveals single-neuron integration of zebrafish ascending and descending dopaminergic systems . Nat Commun . 2011 ; 2 : 171 . doi: 10.1038/ncomms1171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Essen DC , Smith SM , Barch DM , et al. The WUMinn Human Connectome Project: an overview . Neuroimage . 2013 ; 80 : 62 – 79 . doi: 10.1016/j.neuroimage.2013.05.041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- White JG , Southgate E , Thomson JN , et al. The structure of the nervous system of the nematode Caenorhabditis elegans . Philos Trans R Soc Lond B Biol Sci . 1986 ; 314 : 1 – 340 . doi: 10.1098/rstb.1986.0056. [DOI] [PubMed] [Google Scholar]
- Williams RW . Mapping genes that modulate mouse brain development: a quantitative genetic approach . Results Probl Cell Differ . 2000 ; 30 : 21 – 49 . doi: 10.1007/978-3-540-48002-0_2. [DOI] [PubMed] [Google Scholar]