Summary
Benzene is a widespread and toxic contaminant. The fate of benzene in contaminated aquifers seems to be primarily controlled by the abundance of oxygen: benzene is aerobically degraded at high rates by ubiquitous microorganisms, and the oxygen‐dependent pathways for its breakdown were elucidated more than 50 years ago. In contrast, benzene was thought to be persistent under anoxic conditions until 25 years ago. Nevertheless, within the last 15 years, several benzene‐degrading cultures have been enriched under varying electron acceptor conditions in laboratories around the world, and organisms involved in anaerobic benzene degradation have been identified, indicating that anaerobic benzene degradation is a relevant environmental process. However, only a few benzene degraders have been isolated in pure culture so far, and they all use nitrate as an electron acceptor. In some highly enriched strictly anaerobic cultures, benzene has been described to be mineralized cooperatively by two or more different organisms. Despite great efforts, the biochemical mechanism by which the aromatic ring of benzene is activated in the absence of oxygen is still not fully elucidated; methylation, hydroxylation and carboxylation are discussed as likely reactions. This review summarizes the current knowledge about the ‘key players’ of anaerobic benzene degradation under different electron acceptor conditions and the possible pathway(s) of anaerobic benzene degradation.
Introduction
Benzene is a common component of fuels, particularly gasoline, and an important raw chemical used as solvent or chemical intermediate. Due to its extensive use, benzene is a widespread anthropogenic contaminant in aqueous environments. Compared with other hydrocarbons, benzene is highly water‐soluble (saturation: 24 mM at 25°C) and toxic; the US Environmental Protection Agency has classified benzene as a Group A human carcinogen. Chemically, benzene is stable under typical environmental conditions, as the compound is stabilized by the aromatic ring system (π‐electron system) without any potentially reactive substituent.
The persistence of benzene in aqueous environments seems to be primarily controlled by the abundance of oxygen as benzene is often persistent under anoxic conditions. Aerobic benzene‐degrading microorganisms are ubiquitous and have been known for a long time – the first report in regard to aerobic benzene‐degrading microorganisms dates almost 100 years back (Söhngen, 1913). Benzene‐degrading organisms contain mono‐ or dioxygenases which activate the aromatic nucleus by introducing molecular oxygen to yield phenol or cis‐benzene dihydrodiol, compounds that are further oxidized to catechol (Gibson and Parales, 2000; Tao et al., 2004). The aromatic ring of catechol is finally cleaved by further dioxygenases in ortho‐ or meta‐position (Vaillancourt et al., 2006).
Contaminant plumes in aquifers usually become anoxic due to the low solubility and rapid microbial consumption of oxygen. Therefore, knowledge about anaerobic benzene degradation is highly relevant to understand the fate of benzene in the environment. Until the beginning of 1980, aromatic hydrocarbons were thought to be generally recalcitrant under anoxic conditions (Atlas, 1981). In the last 25 years, it was realized that many hydrocarbons including aromatics are biodegradable under several electron‐accepting conditions (for a review see Foght, 2008). However, anaerobic benzene degradation is usually slow and associated with long lag‐times. Benzene is considered to be more persistent under anoxic conditions than its alkylated derivatives toluene, ethylbenzene and xylene isomers and the reasons for the recalcitrance of benzene are not yet clear. Co‐contaminants have been shown to inhibit anaerobic benzene degradation (Edwards et al., 1992; Cunningham et al., 2001; Ruiz‐Aguilar et al., 2003; Da Silva and Alvarez, 2007) – and other studies suggest that anaerobic benzene degraders are not ubiquitous in subsurface sediments (Kazumi et al., 1997; Nales et al., 1998; Weiner and Lovley, 1998a; Phelps and Young, 1999). As we outline in this review, benzene can be degraded by syntrophic consortia which may require narrow and specific environmental conditions for optimal functioning, possibly explaining why benzene is not or only slowly degraded at some field sites or in laboratory enrichment cultures.
The number of laboratory enrichment cultures capable to degrade benzene under anoxic conditions has increased in the last 10 years, but is still low compared with the number of cultures described for anoxic degradation of other hydrocarbons, e.g. toluene. Notably, only four benzene‐degrading pure cultures were described, and all of them use nitrate as electron acceptor (Coates et al., 2001; Kasai et al., 2006). Under strictly anoxic conditions, pure benzene‐degrading strains have been not yet isolated, although some enrichment cultures seem to be dominated by specific organisms. The lack of capable laboratory cultures and the slow growth of the available cultures might be the main reasons why the pathway for anaerobic benzene degradation is still not elucidated.
This review summarizes the current knowledge about the organisms involved in anaerobic benzene degradation and the possible mechanisms by which the benzene ring is activated and transformed in the absence of oxygen, including innovative experimental results published by different research groups in the last 5 years. The reader may find additional information regarding anaerobic benzene degradation also in previously released excellent reviews (Lovley, 2000; Coates et al., 2002; Foght, 2008; Weelink et al., 2010).
Benzene degradation in laboratory microcosms – electron acceptors and ‘key players’
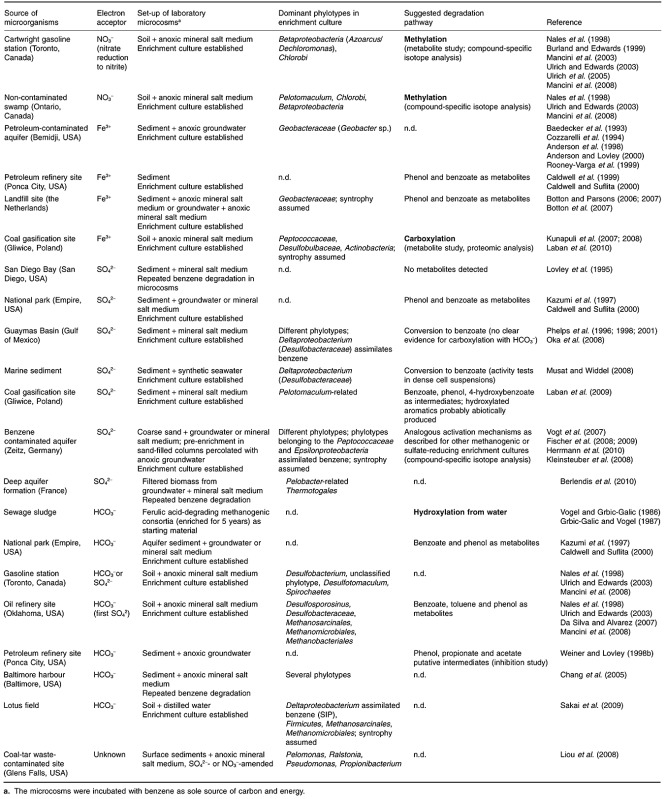
Benzene‐mineralizing laboratory microcosms have been established under several electron‐acceptor conditions; in most cases, successful enrichments of anaerobic benzene degraders were established by using anoxic sediment or soil taken from petroleum contaminated sites (for an overview see Table 1). The major part of microorganisms living in the subsurface are attached to sediment or soil particles (Harvey et al., 1984; Kölbel‐Boelke et al., 1988; Hazen et al., 1991; Alfreider et al., 1997; Griebler et al., 2002; Anneser et al., 2010). Correspondingly, microcosms prepared from sediment usually show higher degradation rates and shorter lag‐phases compared with microcosms prepared from groundwater (Holm et al., 1992). For most of the enrichment cultures, however, long lag‐phases (sometimes more than 100 days) were observed before anaerobic benzene degradation was detectable. Initial degradation rates were low, indicating that the specific rate of anaerobic benzene degradation is generally rather low. Nevertheless, long lag‐phases and low degradation rates seem to be not the only reasons for the limited number of anaerobic benzene‐degrading cultures successfully enriched so far. On the one hand, several authors observed that anaerobic benzene degraders could not be detected at all at distinct sites (e.g. Langenhoff et al., 1996; Kazumi et al., 1997; Nales et al., 1998; Weiner and Lovley, 1998a; Phelps and Young, 1999), indicating that anaerobic benzene degraders may not be ubiquitous. On the other hand, it has been shown that the majority of bacteria cannot be cultured in the laboratory yet (Rappe and Giovannoni, 2003; Keller and Zengler, 2004). These yet uncultured organisms might include anaerobic benzene degraders. Possibly, ‘not detected’ means ‘beyond the detection limit’ in some cases. In our laboratory, we have set up anaerobic enrichment cultures from different contaminated sites with 13C‐labelled benzene as substrate, and observed in some (but not all) cultures a small but continuous release of 13C‐CO2, demonstrating that benzene is mineralized in those cultures, but for long incubation times only detectable for highly sensitive gas chromatographic isotope ratio mass spectrometers (Carsten Vogt, unpubl. results). Similar results were obtained by Morasch and colleagues (2007).
Table 1.
Overview about anaerobic benzene‐degrading microcosms described so far: sources, microcosm preparation, dominant organisms and putative degradation pathway.
Interestingly, the growth behaviour of benzene‐degrading cultures seems to be not correlated to the amount of potential energy available by the electron acceptor. The biomass yields of benzene‐degrading nitrate‐reducing pure and enrichment cultures were reported to be low and comparable to the yield of methanogenic benzene‐degrading enrichment cultures (Coates et al., 2001; Ulrich and Edwards, 2003). This is surprising as the standard free energy for benzene mineralization with nitrate or ferric iron as electron acceptor is more than an order of magnitude higher compared with the acceptors sulfate and carbon dioxide (Table 2).
Table 2.
Stoichiometric equations and standard free energy changes (ΔG0′) for benzene oxidation with different electron acceptors.
| Electron acceptors (oxidized/reduced) | Stoichiometric equation | ΔG0′ (kJ mol−1) |
|---|---|---|
| CO2/CH4 | C6H6 + 6.75 H2O → 2.25 HCO3‐ + 3.75 CH4 + 2.25 H+ | −116a |
| SO42−/H2S | C6H6 + 3 H2O + 3.75 SO42− → 6 HCO3‐ + 1.875 H2S + 1.875 HS‐ + 0.375 H+ | −185b |
| Fe3+/Fe2+ | C6H6 + 18 H2O + 30 Fe3+ → 6 HCO3‐ + 30 Fe2+ + 36 H+ | −3070a |
| NO3‐/N2 | C6H6 + 6 NO3‐ → 6 HCO3‐ + 3 N2 | −2978c |
| NO3‐/NO2‐ | C6H6+ 15 NO3‐ + 3 H2O → 6 HCO3‐ + 15 NO2‐ + 6 H+ | −2061c |
| ClO3‐/Cl‐ | C6H6 + 5 ClO3‐ + 3 H2O → 6 HCO3‐ + 5 Cl‐ + 6 H+ | −3813c |
| O2/H2O | C6H6 + 7.5 O2 + 3 H2O → 6 HCO3‐ + 6 H+ | −3173c |
At some sites, enrichment cultures could be established using different electron acceptors [e.g. Ponca City, USA: ferric iron (Caldwell et al., 1999; Caldwell and Suflita, 2000), sulfate (Anderson and Lovley, 2000), carbon dioxide (Weiner and Lovley, 1998a); Toronto Gas Station: nitrate, sulfate, carbon dioxide (Nales et al., 1998; Ulrich and Edwards, 2003)]. A few cultures were also described to switch from sulfate to carbon dioxide as electron acceptor and vice versa (Ulrich and Edwards, 2003). The latter examples support the hypothesis of syntrophic processes governing anaerobic benzene degradation, which are discussed in more detail below.
Benzene degradation under methanogenic conditions
Benzene degradation under methanogenic conditions was occasionally observed in laboratory microcosms (Wilson et al., 1986; Grbic‐Galic and Vogel, 1987; Kazumi et al., 1997; Nales et al., 1998; Weiner and Lovley, 1998b; Ulrich and Edwards, 2003; Chang et al., 2005; Sakai et al., 2009), but also verified in situ by means of push‐pull experiments (Reinhard et al., 2005). The first demonstration of anaerobic benzene mineralization was reported by Grbic‐Galic and Vogel (1987) who detected a small production (less than 10%) of 14C‐CO2 from ring‐labelled 14C‐benzene in a methanogenic culture which was originally prepared from sewage sludge and pre‐enriched with ferulic acid as substrate. Complete mineralization of benzene to equal amounts of carbon dioxide and methane was shown later in microcosms set up with sediment samples taken from a benzene‐contaminated aquifer and using 14C‐labelled benzene as substrate (Weiner and Lovley, 1998b). Added acetate (1 mM) or propionate (100 µM) strongly inhibited benzene mineralization, indicating that these compounds were fermentatively produced during benzene degradation. Two cultures initially enriched with sulfate as electron acceptor could switch to carbon dioxide; notably, degradation rates were significantly higher under methanogenic conditions (Ulrich and Edwards, 2003). In these cultures, phylotypes affiliated to the genera Desulfobacterium and Desulfosporosinus as well as aceticlastic methanogens were identified as dominant members of the community after prolonged incubation with benzene (Ulrich and Edwards, 2003; Da Silva and Alvarez, 2007; Mancini et al., 2008). Recently, a deltaproteobacterium distantly related to the Syntrophaceae was identified by stable isotope probing of DNA (DNA‐SIP) and real‐time PCR as a key player of anaerobic benzene degradation in a methanogenic enrichment culture set up from non‐contaminated soil (Sakai et al., 2009). The archaeal clone library constructed from this enrichment culture was dominated by phylotypes belonging to the Methanosarcinales and Methanomicrobiales. The authors concluded that benzene was sequentially degraded by a consortium of fermenters, aceticlastic methanogens and hydrogenotrophic methanogens. Such syntrophic relationships likely exist in all benzene‐degrading cultures under methanogenic conditions, as methanogens have a narrow organic substrate spectrum restricted to simple low‐molecular‐weight organic compounds, and are not known for degrading aromatic compounds. Thus, methanogens are assumed to consume hydrogen, acetate or other small molecules released by fermenting organisms. Such syntrophic relationships were also described for the degradation of long‐chain alkanes under methanogenic conditions (Zengler et al., 1999).
Benzene degradation under sulfate‐reducing conditions
Sulfate‐dependent benzene mineralization was demonstrated for the first time in sediment microcosms from freshwater and marine origin (Lovley et al., 1995; Phelps et al., 1996). These studies verified a previous observation of benzene mineralization where the electron acceptor was assumed to be sulfate (Edwards et al., 1992); actually, this study was the first stating ‘complete’ anaerobic mineralization of benzene. In situ anaerobic benzene degradation at the Ponca City site (USA) could be stimulated by adding sulfate (Anderson and Lovley, 2000), indicating its preferred use as electron acceptor for anaerobic benzene degradation at this site. Sediment laboratory microcosms from this site degraded benzene also with carbon dioxide or ferric iron as electron acceptor (Weiner and Lovley, 1998a; Caldwell et al., 1999).
In the last 15 years, several cultures degrading benzene under sulfate‐reducing conditions were enriched and the dominant organisms were described. The marine culture enriched by Phelps and colleagues (1996) contained 12 different phylotypes after incubation for 3 years with benzene as sole source of carbon and energy (Phelps et al., 1998). Four clones belonged to the Desulfobacteraceae, the other phylotypes were affiliated to Thiomicrospira (Gammaproteobacteria), Sulfurovum (Epsilonproteobacteria), Bellilinea (Chloroflexi), Exiguobacterium (Bacilli) as well as several members of the Clostridia and Bacteroidetes (according to the RDP Classifier; Wang et al., 2007). After more than 10 years of incubation, a phylotype belonging to the Desulfobacteraceae (‘clone SB‐21’) was identified by DNA‐SIP for assimilating benzene in this culture (Oka et al., 2008). Notably, a phylotype closely related to clone SB‐21 was found to be the dominant organism in a benzene‐mineralizing sulfate‐reducing culture enriched also from marine sediment (Musat and Widdel, 2008). Furthermore, phylotypes related to the Desulfobacteraceae were detected in higher abundances in two benzene‐degrading freshwater enrichments able to use sulfate or carbon dioxide as electron acceptor (Ulrich and Edwards, 2003; Mancini et al., 2008). These observations suggest that members of the Desulfobacteraceae are key players of benzene degradation under sulfate‐reducing conditions. However, despite enrichment with benzene for several years, all of these cultures contained still other phylotypes, and a single benzene‐degrading Desulfobacteraceae‐like organism could not be isolated from any of the described cultures. Therefore, it is possible that benzene is mineralized in these cultures by synergetic or syntrophic relationships of Desulfobacteraceae and other organisms; this can be at least assumed for the enrichment cultures which can use both sulfate and carbon dioxide as electron acceptors.
In addition to phylotypes belonging to the Gram‐negative Desulfobacteraceae, Gram‐positive members of the family Peptococcaceae seem to be key players of sulfate‐dependent anaerobic benzene degradation. The enrichment culture of Laban and colleagues (2009) was dominated by a phylotype related to the genus Pelotomaculum; the authors assumed that this organism may assimilate benzene solely using sulfate as electron acceptor. In our own culture enriched from groundwater‐percolated sand columns at the field site Zeitz (Vogt et al., 2007), a phylotype affiliated to the Cryptanaerobacter/Pelotomaculum group within the Desulfotomaculum subcluster Ih of the Peptococcaceae (Imachi et al., 2006) was shown to be crucial for sulfate‐dependent benzene mineralization, as T‐RFLP peak abundances of this organism increased after prolonged incubation with benzene and decreased during growth on other substrates (Kleinsteuber et al., 2008). Moreover, this phylotype and also an Epsilonproteobacterium distantly related to the genus Sulfurovum assimilated carbon from 13C‐labelled benzene in a subsequent DNA‐SIP experiment (Herrmann et al., 2010). In this experiment, small amounts of 13C‐labelled benzene were converted to 13C‐labelled methane, and members of the aceticlastic Methanosaetaceae were detected in an archaeal 16S rRNA gene clone library. In addition, several sulfate‐reducing Deltaproteobacteria were detected but not found to be 13C‐labelled in spite of prolonged incubation with benzene (Kleinsteuber et al., 2008; Herrmann et al., 2010). It was further shown that benzene mineralization could be reversibly inhibited by addition of hydrogen or low amounts of acetate (0.3 mM), indicating that both compounds are intermediates during anaerobic benzene fermentation (Rakoczy et al., unpubl. data). It was therefore suggested that benzene is mineralized by a consortium consisting of syntrophic fermenters, hydrogenotrophic sulfate reducers, aceticlastic methanogens and other acetate‐consuming bacteria (Fig. 1). Contaminated aquifers might be a specific ecological niche for syntrophic benzene degradation, as syntrophs are versatile regarding the terminal electron acceptor and specifically adapted to a life at the thermodynamic limit. At the Zeitz site, anaerobic benzene degradation, likely with sulfate as electron acceptor, has been verified also in the flow path of the benzene plume by compound‐specific stable isotope analysis (CSIA) (Fischer et al., 2007; 2009). This method, which is explained in more detail below, has generally great potential to verify anaerobic benzene degradation in contaminated aquifers due to the strong hydrogen isotope fractionation linked to anaerobic benzene degradation.
Figure 1.
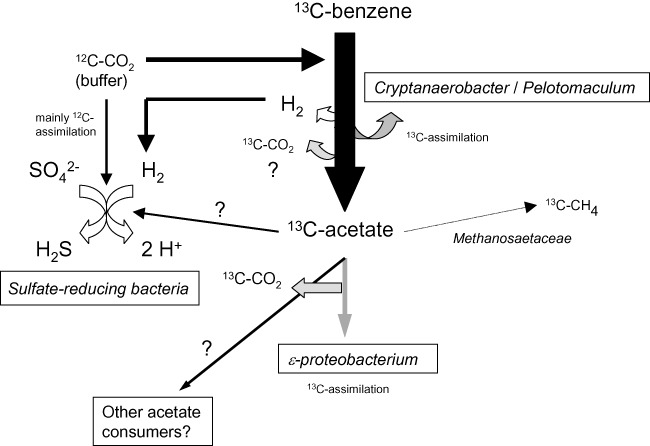
Proposed degradation pathway for sulfate‐dependent syntrophic benzene mineralization, based on previous observations (Vogt et al., 2007; Kleinsteuber et al., 2008; Herrmann et al., 2010; J. Rakoczy, K.M. Schleinitz, N. Müller, H.‐H. Richnow, unpubl. data). T‐RFLP and SIP data suggest that the Cryptanaerobacter/Pelotomaculum (CP) phylotype assimilates the majority of benzene within the consortium. During benzene oxidation to acetate by the CP phylotype, hydrogen is formed and consumed by various sulfate reducers, driving the syntrophic process. Benzene‐derived acetate might be consumed by several ecophysiologically different organisms, but especially by an epsilonproteobacterium. The majority of the produced carbon dioxide is thought to be formed from the acetate released by the CP phylotype. Small amounts of acetate are aceticlastically converted to methane.
Recently, a sulfate‐reducing benzene‐degrading culture was enriched from groundwater sampled from an underground gas storage area (Berlendis et al., 2010). The abundant phylotypes in this enrichment culture were distantly related to Pelobacter, Thermotogales and Methanolobus, indicating that benzene is syntrophically degraded.
Benzene degradation under iron‐reducing conditions
Several benzene‐mineralizing iron‐reducing cultures were successfully established using amorphous iron hydroxide [Fe(OH)3] as electron acceptor (Kazumi et al., 1997; Villatoro‐Monzon et al., 2003; Jahn et al., 2005; Botton and Parsons, 2006; Kunapuli et al., 2007). Iron‐dependent benzene mineralization could also be stimulated by adding nitriloacetic acid (NTA) or other iron‐chelating compounds (Lovley et al., 1994; 1996; Caldwell et al., 1999). Presumably one of the best‐investigated aquifers in which iron‐driven benzene degradation takes place is the Bemidji site in Minnesota (USA), where a crude oil pipeline ruptured in 1979 and contaminated the adjacent aquifer (Essaid et al., 2011). Benzene degradation or mineralization in anaerobic microcosms prepared with sediment and groundwater from the iron‐reducing zone of this aquifer was repeatedly reported (Baedecker et al., 1993; Cozzarelli et al., 1994; Anderson et al., 1998). Recently, benzene degradation was also verified directly in the anoxic iron‐reducing zone of the plume by an in situ microcosms approach (Cozzarelli et al., 2010). Rooney‐Varga and colleagues (1999) investigated the community structure of iron‐reducing benzene‐mineralizing enrichment cultures and sediment samples of the Bemidji site. MPN‐PCR revealed an increase of Geobacter‐related 16S rRNA gene copies in benzene‐mineralizing sediments and enrichment cultures, indicating an involvement of Geobacteraceae in benzene mineralization under iron‐reducing conditions.
Geobacteraceae were also dominant in iron‐reducing benzene‐degrading enrichment cultures set up from a landfill site in the Netherlands (Botton and Parsons, 2007). In contrast, no Geobacteraceae were identified in a highly enriched iron‐reducing culture originating from a contaminated site in Poland (Kunapuli et al., 2007; Laban et al., 2010). Here, a phylotype affiliated to the Peptococcaceae was most abundant and assimilated 13C‐benzene in a DNA‐SIP experiment (Kunapuli et al., 2007). Phylotypes affiliated to the Desulfobulbaceae and members of the Actinobacteria were also prominent. The authors suggested that benzene is syntrophically mineralized in this culture, with the Peptococcaceae phylotype as the primary benzene oxidizer.
Benzene degradation under nitrate‐reducing conditions
Major and colleagues (1988) demonstrated first benzene degradation dependent on nitrate as electron acceptor, using microcosms set up from hydrocarbon‐contaminated sediments and anoxic groundwater. Nitrate was shown to be reduced to dinitrogen during benzene degradation; the electron balance of reduced nitrate and degraded benzene indicated that benzene was mineralized, although carbon dioxide production from benzene was not experimentally confirmed. Benzene degradation under nitrate‐reducing conditions was later observed in microcosms made of soil and anoxic groundwater taken from three different sites (Nales et al., 1998). Afterwards, stable benzene‐degrading enrichment cultures were established and further examined. As demonstrated for one culture in experiments with 14C‐labelled benzene, more than 90% of the benzene's carbon was released as CO2; benzene degradation appeared to be coupled to nitrate reduction to nitrite (Burland and Edwards, 1999). Notably, the biomass yield of the nitrate‐reducing cultures was in the range observed for sulfate‐reducing or methanogenic benzene‐degrading enrichments (Ulrich and Edwards, 2003), despite the high redox potential of the nitrate/nitrite couple (Table 2). Dominant phylotypes in the cultures, determined after being incubated for several years in the laboratory, belonged to the Betaproteobacteria (genera Azoarcus and Dechloromonas), the Peptococcaceae (genus Pelotomaculum) and Chlorobi (Ulrich and Edwards, 2003; Mancini et al., 2008). Analogously to cultures enriched with ferric iron, sulfate or carbon dioxide as electron acceptor, it was apparently not possible to isolate a single benzene degrader out of these cultures, despite the long laboratory incubation time.
Benzene degradation with electron acceptors other than nitrate, sulfate, ferric iron or carbon dioxide
Manganese(IV) was shown to be used as electron acceptor for anaerobic benzene degradation in sediment microcosms and columns. The benzene degradation rates were significantly higher for manganese(IV) compared with ferric iron, indicating that this process is relevant in the environment (Villatoro‐Monzon et al., 2003; 2008). Recently, Zhang and colleagues (2010) showed that benzene was mineralized by sediment microorganisms using a graphite anode as electron acceptor.
Pure cultures of anaerobic benzene degraders
Notably, the only pure cultures described to mineralize benzene using nitrate as electron acceptor were not isolated by the ‘classical approach’ from benzene‐degrading enrichment cultures. The strains JJ and RCB, belonging to the genus Dechloromonas, were originally isolated using humic substances and nitrate (JJ), or chlorobenzoate and chlorate (RCB) as electron donors and acceptors respectively (Coates et al., 2001). Kasai and colleagues (2006) identified a phylotype affiliated to the genus Azoarcus for assimilation of benzene with nitrate as electron acceptor in an RNA‐SIP experiment targeting gasoline‐contaminated groundwater. The authors than isolated several pure strains out of the groundwater using a non‐selective medium and were able to identify two Azoarcus strains as nitrate‐reducing benzene degraders in subsequent experiments with 14C‐labelled benzene (Kasai et al., 2006).
Tan and colleagues (2006) and Weelink and colleagues (2007) enriched a chlorate‐reducing benzene‐degrading culture. Later, a strain belonging to Alicycliphilus denitrificans was isolated that was able to mineralize benzene with chlorate as electron acceptor (Weelink et al., 2008). However, in the last step of the chlorate reduction pathway, chlorite dismutates to oxygen and chloride. Biochemical and physiological data suggest that this bacterium uses the oxygen released during chlorate reduction as a co‐substrate for the initial attack and ring cleavage of benzene; thus it cannot be regarded as a true anaerobic benzene degrader. Benzene degradation by chlorate reduction is excellently and extensively described in the recent review of Weelink and colleagues (2010).
Mechanisms of anaerobic benzene activation
The question how benzene is activated in the absence of oxygen is still not convincingly answered, although several attempts have been made since the end of the 1980s to elucidate the activation mechanism. In principle, the benzene molecule is thermodynamically very stable due to the symmetric π‐electron system of the aromatic ring and the lack of potentially destabilizing or reactive substituents.
Many alkylated aromatic hydrocarbons, e.g. toluene and xylenes, are activated under anoxic conditions by a reaction sequence in which fumarate is added to the alkyl side‐chain of the aromatic ring (for a review see Heider, 2007). In the first step of this reaction, a relatively stable benzyl radical is thought to be formed. One might suggest that benzene is activated by a similar mechanism; however, the formation of a phenyl radical as a reactive intermediate for a subsequent methylation or other reaction is rather unlikely for energetic reasons. The abstraction of a hydrogen atom from benzene would need an activation energy of more than 460 kJ mol−1 which is roughly 100 kJ mol−1 more compared with the formation of a benzyl radical from alkylated benzene derivatives (Widdel and Rabus, 2001; Musat and Widdel, 2008). Thus, three other activation mechanisms have been intensively discussed: (i) an anaerobic hydroxylation of benzene yielding phenol, (ii) a Friedel‐Crafts‐type methylation of benzene yielding toluene and (iii) a carboxylation of benzene yielding benzoate (Fig. 2). We will discuss the pros and cons for all three possible activation reactions. Generally, isotope‐based methods have been used to elucidate the reaction mechanism, either by CSIA, or by detection of 13C‐ or 14C‐labelled metabolites which are formed during transformation of 13C‐ or 14C‐labelled benzene. Due to the lack of pure cultures, most studies aiming to elucidate the reaction mechanism were performed with enrichment cultures.
Figure 2.
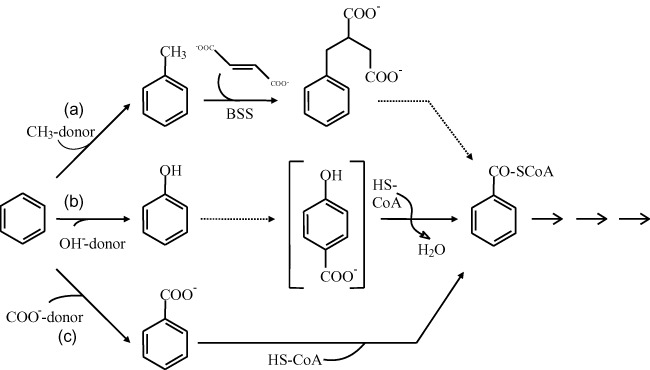
Possible anaerobic benzene activation steps and further transformation reactions to benzoyl‐CoA as central metabolite (modified from Foght, 2008 and Laban et al., 2010). (a) Methylation; (b) hydroxylation; (c) carboxylation. Benzoyl‐CoA can be further reduced by ATP‐dependent or ‐independent benzoyl‐CoA reductases (Kung et al., 2009).
Benzene hydroxylation
Benzene hydroxylation by hydroxyl ions stemming from water is an attractive model for benzene activation as the redox potential for this reaction is comparably low (E°′ = −0.09 V; Musat and Widdel, 2008) and hence likely possible even under sulfate‐reducing or methanogenic conditions. Phenol itself has been shown to be mineralized by facultative and strictly anaerobic bacteria; the degradation pathway proceeds via a carbon dioxide‐dependent carboxylation of the aromatic ring to 4‐hydroxybenzoate and further transformation to benzoyl‐CoA (Bak and Widdel, 1986; Knoll and Winter, 1987; Tschech and Fuchs, 1987; Lovley and Lonergan, 1990; Zhang and Wiegel, 1994; Qiu et al., 2008; Schleinitz et al., 2009).
Hydroxylation of benzene had been already suggested as an initial reaction mechanism for benzene activation in one of the first reports regarding anaerobic benzene degradation (Vogel and Grbic‐Galic, 1986). Addition of 18O‐labelled water resulted in the formation of 18O‐labelled phenol, indicating that the introduced hydroxyl group originated from water. In subsequent reports, phenol was often detected in enrichments as metabolite of benzene degradation under different electron acceptor conditions: in iron‐reducing cultures (Caldwell and Suflita, 2000; Botton and Parsons, 2007; Kunapuli et al., 2008), sulfate‐reducing cultures (Caldwell and Suflita, 2000; Laban et al., 2009) and methanogenic cultures (Weiner and Lovley, 1998b; Caldwell and Suflita, 2000; Ulrich et al., 2005). In some studies, benzoate was concomitantly detected with phenol (Caldwell and Suflita, 2000; Ulrich et al., 2005; Kunapuli et al., 2008).
It is noteworthy that it has recently been observed that phenol can be abiotically formed from benzene in culture media from iron and sulfate reducers by contact with air after sampling (Kunapuli et al., 2008). Hydroxyl radicals were likely generated by oxidation of iron in the sample during work up, which then reacted rapidly with benzene producing small amounts of phenol before sample analysis. The formation of 2‐hydroxybenzoate and 4‐hydroxybenzoate from benzoate was also explained by these mechanisms (Laban et al., 2009). These results indicate that it is generally problematic to distinguish between biotic and abiotic phenol formation in strongly reduced culture samples, probably preventing any clear evidence for benzene hydroxylation under strictly anoxic conditions by means of metabolite analysis. Hence, other methodological approaches are needed in addition. Substrate consumption tests with two highly enriched sulfate‐reducing benzene‐degrading cultures revealed that phenol is either not consumed (Laban et al., 2009) or only consumed after a certain lag‐phase (Musat and Widdel, 2008), strongly indicating that phenol is unlikely to be an intermediate during benzene degradation in these cultures.
Chakraborty and Coates (2005) suggested that Dechloromonas strain RCB hydroxylated benzene to phenol, which was subsequently transformed to benzoate; the reaction was dependent on the presence of nitrate as electron acceptor. Here, the authors did not report phenol formation or benzene degradation in anoxic, nitrate‐free control cultures amended with benzene. The origin of the introduced hydroxyl group could not be identified. When cells degraded benzene in H218O‐enriched mineral salt medium, the formed phenol was only slightly enriched with 18O, suggesting that the hydroxyl group did not originate from water. On the other hand, hydroxyl free radical scavengers strongly inhibited benzene degradation and phenol formation, indicating that hydroxyl radicals were the source of the hydroxyl group in phenol. In an additional study, Chakraborty and colleagues (2005) showed that strain RCB could degrade benzene and several other aromatic hydrocarbons with nitrate, chlorate or oxygen as electron acceptor. Surprisingly, none of the known genes for anaerobic degradation of aromatic compounds could be found in the genome of strain RCB, which was recently sequenced (Salinero et al., 2009). Due to the lack of these genes, anaerobic benzene degradation in strain RCB ‘remains enigmatic’ (Salinero et al., 2009). On the other hand, strain RCB encodes several aerobic pathways for aromatics degradation. In the presence of chlorate, this organism releases oxygen during chlorate respiration allowing aromatics degradation by means of oxygenases even in the initial absence of oxygen in the culture medium, as shown also for the chlorate‐reducing benzene degrader Alicycliphilus denitrificans (Weelink et al., 2008). Interestingly, it has been recently demonstrated that oxygen can be released during reduction of nitric oxide (NO) (Ettwig et al., 2010), an intermediate of the classical nitrate reduction pathway to dinitrogen. As also suggested by Weelink and colleagues (2010), strain RCB may contain this enzyme, allowing the use of oxygen for the initial attack of benzene and other aromatics even under nitrate‐reducing conditions, explaining the apparently inconsistent physiological and genetic data. Unfortunately, the oxygen‐releasing enzyme has not been characterized and can therefore not yet be identified in the genome of strain RCB. The genome of strain RCB contains the genes for the classical nitrate reduction to dinitrogen pathway, including those for nitric oxide reductase which catalyses nitric oxide reduction to nitrous oxide (N2O) (Salinero et al., 2009).
Benzene methylation
Benzene methylation via Friedel‐Crafts‐type reaction is exergonic using the unique biological methyl donors S‐adenosyl‐methionine or methyl‐tetrahydrofolate (Coates et al., 2002), which open the doors for another hypothesis for anaerobic benzene activation. Actually, S‐adenosyl‐methione‐dependent alkylation of benzene (and substituted aromatics) has been observed in bone marrow (Flesher and Myers, 1991). Methylation has been also proposed for the anaerobic activation of the non‐substituted aromatic hydrocarbon naphthalene (Safinowski and Meckenstock, 2006). If benzene is methylated by anaerobes, the reaction product toluene could be further activated by addition of fumarate to the methyl group of toluene catalysed by the enzyme benzylsuccinate synthase (BSS), leading to the characteristic compound benzylsuccinate as intermediate. BSS has been detected in several anaerobic toluene‐degrading pure and mixed cultures, and fumarate addition seems to be a unique activation mechanism for anaerobic toluene degradation (for an overview see Heider, 2007). PCR primers for the gene encoding the protein subunit which contains the reactive centre, bssA, have been also developed (Winderl et al., 2007). Thus, reasonable strategies for verifying benzene activation by methylation are detecting the intermediates toluene and benzylsuccinate or detecting the presence or expression of bssA‐like genes or the induction or activity of BSS.
Ulrich and colleagues (2005) detected [ring‐13C]‐labelled toluene and [ring‐13C]‐labelled benzoate as intermediates in [13C6]‐benzene‐spiked nitrate‐reducing and methanogenic enrichment cultures. The formation of [13C6]‐phenol was observed only in the methanogenic culture. The nitrate‐reducing culture degraded toluene rapidly and at higher rates than benzene, supporting the hypothesis that toluene might be an intermediate during benzene degradation in this culture. In contrast, toluene was only slightly degraded by the methanogenic culture. The authors concluded that two degradation pathways exist: (i) a methylation pathway leading to toluene with subsequent transformation to benzoate operating in the nitrate‐reducing and methanogenic culture, and (ii) a hydroxylation pathway leading to phenol with subsequent formation of benzoate operating only in the methanogenic culture. This hypothesis was supported by studies in which CSIA was used for characterizing the initial step of benzene activation in different cultures (Mancini et al., 2003; 2008; Fischer et al., 2008). Mancini and colleagues (2008) showed that the ratio of hydrogen isotope fractionation (Δδ2H) versus carbon isotope fractionation (Δδ13C) – a value defined as lambda: Λ = Δδ2H/Δδ13C (Fischer et al., 2008) – for anaerobic benzene degradation was significantly higher for the methanogenic culture (Λ = 39 ± 5) compared with the nitrate‐reducing enrichment culture (Λ = 16 ± 2) investigated by Ulrich and colleagues (2005). Simplified, the lambda value can be seen as a biochemical fingerprint of a given biochemical reaction. Concordantly, other nitrate‐reducing cultures showed lambda values in the range between 8 and 19, whereas for other methanogenic or sulfate‐reducing cultures lambda values between 28 and 31 were determined (Mancini et al., 2003; 2008; Fischer et al., 2008). Thus, the CSIA data indicate that benzene activation under nitrate‐reducing conditions is different from benzene activation under sulfate‐reducing and methanogenic conditions. Nevertheless, ‘different reaction mechanism’ means that either the reactions are truly different (different products are formed), or the reactions are similar on paper (same products are formed) but proceed via different reaction mechanisms catalysed by different enzymes or cofactors leading to different fractionation patterns; the latter has been recently shown for toluene activation by benzylsuccinate synthase (Vogt et al., 2008; Herrmann et al., 2009). Hence, further research is needed to conclusively demonstrate that benzene can be methylated under nitrate‐reducing conditions. Some highly enriched strictly anaerobic benzene‐degrading cultures cannot degrade toluene (Kunapuli et al., 2008; Musat and Widdel, 2008; Laban et al., 2009), probably excluding biomethylation of benzene to form toluene as activation mechanism.
Benzene carboxylation
Similar to hydroxylation or methylation of benzene, benzene carboxylation is slightly exergonic or close to the thermodynamic equilibrium depending on the carboxyl donor and thus feasible even in sulfate‐reducing or methanogenic cultures (Musat and Widdel, 2008). Additionally, for some non‐substituted aromatic compounds, e.g. naphthalene (Zhang and Young, 1997; Musat et al., 2009; DiDonato et al., 2010) or phenanthrene (Zhang and Young, 1997; Davidova et al., 2007), a carboxylation reaction was suggested for ring activation, indicating that carboxylation might be an important activation principle for the degradation of non‐substituted aromatic compounds in nature. Indeed, benzoate has been detected as intermediate of anaerobic benzene degradation in sulfate‐reducing (Caldwell and Suflita, 2000; Phelps et al., 2001; Laban et al., 2009), iron‐reducing (Caldwell and Suflita, 2000; Kunapuli et al., 2008), nitrate‐reducing (Ulrich et al., 2005) or methanogenic (Caldwell and Suflita, 2000; Ulrich et al., 2005) enrichment cultures, concomitantly with the intermediates phenol (Caldwell and Suflita, 2000; Ulrich et al., 2005; Kunapuli et al., 2008; Laban et al., 2009) or toluene (Ulrich et al., 2005). Phelps and colleagues (2001) found deuterated benzoate (D5) as sole intermediate of deuterated benzene (D6) degradation in their highly enriched marine sulfate‐reducing culture. Interestingly, incubation in the presence of a 13C‐labelled bicarbonate buffer system did not lead to 13C incorporation in the carboxyl group of the benzoate intermediate, indicating that the introduced carboxyl group did not originate from carbon dioxide. This was in accordance with results presented by Caldwell and Suflita (2000) for their benzene‐degrading sulfate‐reducing freshwater culture. Here, fully [13C7]‐labelled benzoate was formed when the culture was spiked with fully [13C6]‐labelled benzene, showing that the carboxyl group of benzoate was stemming from transformation products of [13C6]‐benzene itself, but not from the non‐labelled bicarbonate buffer system. In contrast, Kunapuli and colleagues (2008) found both [13C6]‐benzoate and [13C7]‐benzoate in their iron‐reducing enrichment culture during incubation with fully labelled [13C6]‐benzene. Furthermore, the authors detected 13C‐carboxy group‐labelled benzoate if cells were incubated in medium prepared with non‐labelled benzene and 13C‐labelled bicarbonate buffer, favouring the hypothesis that the bicarbonate buffer was the carboxyl group donor for benzoate formation.
However, it is generally difficult to interpret all these observations since benzoyl‐CoA, the activated form of benzoate, is a common intermediate within the anaerobic degradation pathways of several aromatic compounds including toluene and phenol (for an overview see Carmona et al., 2009; see also Fig. 2); in addition, benzoate has been reported as an excreted intermediate during phenol degradation under methanogenic conditions (Knoll and Winter, 1987; Kobayashi et al., 1989; Bechard et al., 1990; Karlsson et al., 2000). Hence, the metabolite benzoate might be formed directly from benzene by a carboxylation step, but could be formed as well in later steps during anaerobic benzene degradation pathways starting, e.g. with a methylation or hydroxylation step.
Lately, Laban and colleagues (2010) investigated their highly enriched iron‐reducing benzene‐degrading culture using an approach combining metagenomics and metaproteomics. Subcultures were grown with benzene, phenol or benzoate as sole substrates, and peptide sequences were subsequently identified based on the metagenome which had been sequenced before. Proteins similar to the phenylphosphate carboxylase subunits PpcA and PpcD of Azoarcus sp. strain EbN1 and to the benzoate‐CoA ligase of Geobacter metallireducens were specifically expressed during anaerobic benzene degradation. Based on these results, the authors suggested that benzene is directly carboxylated by a putative anaerobic benzene carboxylase. The formed benzoate might be further activated by a benzoate‐CoA ligase to benzoyl‐CoA. However, an enzyme activity test for the putative anaerobic benzene carboxylase, ultimately proving this hypothesis, could not be established yet.
Outlook – what is needed for a better understanding of the mechanisms of anaerobic benzene degradation?
Isolating strictly anaerobic benzene‐degrading bacteria is still important for a better understanding of anaerobic benzene degradation. Sequencing the genome of such an organism may allow revealing the upper and lower degradation pathway; also more controlled physiological experiments would be possible for elucidating the anaerobic benzene activation mechanism. However, the gene encoding the responsible enzyme might be not yet detectable within the genome due to the unknown enzyme properties. Establishing any in vitro or in vivo assay undoubtedly proving a specific activation mechanism is clearly needed.
For highly enriched benzene‐degrading cultures, sequencing of the metagenome might be very helpful to clarify syntrophic or cooperative relationships, as, for example, recently shown for anaerobic methane oxidation (Meyerdierks et al., 2010). If a metagenome is available, mRNA expression or proteomic studies may follow for further functional characterization (Ram et al., 2005; Meyerdierks et al., 2010).
Acknowledgments
This work is integrated into the internal research and development programme of the UFZ as well as the SAFIRA project. We acknowledge funding also from the German Research Foundation, Priority Program 1319.
References
- Alfreider A., Krossbacher M., Psenner R. Groundwater samples do not reflect bacterial densities and activity in subsurface systems. Water Res. 1997;31:832–840. [Google Scholar]
- Anderson R.T., Lovley D.R. Anaerobic bioremediation of benzene under sulfate‐reducing conditions in a petroleum‐contaminated aquifer. Environ Sci Technol. 2000;34:2261–2266. [Google Scholar]
- Anderson R.T., Rooney‐Varga J.N., Gaw C.V., Lovley D.R. Anaerobic benzene oxidation in the Fe(III) reduction zone of petroleum contaminated aquifers. Environ Sci Technol. 1998;32:1222–1229. [Google Scholar]
- Anneser B., Pilloni G., Bayer A., Lueders T., Griebler C., Einsiedl F., Richters L. High resolution analysis of contaminated aquifer sediments and groundwater: what can be learned in terms of Natural Attenuation? Geomicrobiol J. 2010;27:130–142. [Google Scholar]
- Atlas R.M. Microbial degradation of petroleum hydrocarbons – an environmental perspective. Microbiol Rev. 1981;45:180–209. doi: 10.1128/mr.45.1.180-209.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baedecker M.J., Cozzarelli I.M., Eganhouse R.P., Siegel D.I., Bennett P.C. Crude oil in a shallow sand and gravel aquifer‐III. Biogeochemical reactions and mass balance modelling in anoxic groundwater. Appl Geochem. 1993;8:569–586. [Google Scholar]
- Bak F., Widdel F. Anaerobic degradation of phenol and phenol derivates by Desulfobacterium phenolicum sp. nov. Arch Microbiol. 1986;146:177–180. [Google Scholar]
- Bechard G., Bisaillon J.G., Beaudet R., Sylvestre M. Degradation of phenol by a bacterial consortium under methanogenic conditions. Can J Microbiol. 1990;36:573–578. [Google Scholar]
- Berlendis S., Lascourreges J.F., Schraauwers B., Sivadon P., Magot M. Anaerobic biodegradation of BTEX by original bacterial communities from an underground gas storage aquifer. Environ Sci Technol. 2010;44:3621–3628. doi: 10.1021/es100123b. [DOI] [PubMed] [Google Scholar]
- Botton S., Parsons J.R. Degradation of BTEX compounds under iron‐reducing conditions in contaminated aquifer microcosms. Environ Toxicol Chem. 2006;25:2630–2638. doi: 10.1897/06-004r.1. [DOI] [PubMed] [Google Scholar]
- Botton S., van Harmelen M., Braster M., Parsons J.R., Roling W.F.M. Dominance of Geobacteraceae in BTX‐degrading enrichments from an iron‐reducing aquifer. FEMS Microbiol Ecol. 2007;62:118–130. doi: 10.1111/j.1574-6941.2007.00371.x. [DOI] [PubMed] [Google Scholar]
- Botton S., Parsons J.R. Degradation of BTX by dissimilatory iron‐reducing cultures. Biodegradation. 2007;18:371–381. doi: 10.1007/s10532-006-9071-9. [DOI] [PubMed] [Google Scholar]
- Burland S.M., Edwards E.A. Anaerobic benzene biodegradation linked to nitrate reduction. Appl Environ Microbiol. 1999;65:529–533. doi: 10.1128/aem.65.2.529-533.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Caldwell M.E., Suflita J.M. Detection of phenol and benzoate as intermediates of anaerobic benzene biodegradation under different terminal electron‐accepting conditions. Environ Sci Technol. 2000;34:1216–1220. [Google Scholar]
- Caldwell M.E., Tanner R.S., Suflita J.M. Microbial metabolism of benzene and the oxidation of ferrous iron under anaerobic conditions: implications for bioremediation. Anaerobe. 1999;5:595–603. [Google Scholar]
- Carmona M., Zamarro M.T., Blazquez B., Durante‐Rodriguez G., Juarez J.F., Valderrama J.A. Anaerobic catabolism of aromatic compounds: a genetic and genomic view. Microbiol Mol Biol Rev. 2009;73:71–133. doi: 10.1128/MMBR.00021-08. et al. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chakraborty R., Coates J.D. Hydroxylation and carboxylation – two crucial steps of anaerobic benzene degradation by Dechloromonas strain RCB. Appl Environ Microbiol. 2005;71:5427–5432. doi: 10.1128/AEM.71.9.5427-5432.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chakraborty R., O'Connor S.M., Chan E., Coates J.D. Anaerobic degradation of benzene, toluene, ethylbenzene, and xylene compounds by Dechloromonas strain RCB. Appl Environ Microbiol. 2005;71:8649–8655. doi: 10.1128/AEM.71.12.8649-8655.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang W., Um Y., Holoman T.R.P. Molecular characterization of anaerobic microbial communities from benzene‐degrading sediments under methanogenic conditions. Biotechnol Prog. 2005;21:1789–1794. doi: 10.1021/bp050250p. [DOI] [PubMed] [Google Scholar]
- Coates J.D., Chakraborty R., Lack J.G., O'Connor S.M., Cole K.A., Bender K.S., Achenbach L.A. Anaerobic benzene oxidation coupled to nitrate reduction in pure culture by two strains of Dechloromonas. Nature. 2001;411:1039–1043. doi: 10.1038/35082545. [DOI] [PubMed] [Google Scholar]
- Coates J.D., Chakraborty R., McInerney M.J. Anaerobic benzene biodegradation – a new era. Res Microbiol. 2002;153:621–628. doi: 10.1016/s0923-2508(02)01378-5. [DOI] [PubMed] [Google Scholar]
- Cozzarelli I.M., Baedecker M.J., Eganhouse R.P., Goerlitz D.F. The geochemical evolution of low molecular weight organic acids derived from the degradation of petroleum contaminants in groundwater. Geochim Cosmochim Acta. 1994;58:863–877. [Google Scholar]
- Cozzarelli I.M., Bekins B.A., Eganhouse R.P., Warren E., Essaid H.I. In situ measurements of volatile aromatic hydrocarbon biodegradation rates in groundwater. J Contam Hydrol. 2010;111:48–64. doi: 10.1016/j.jconhyd.2009.12.001. [DOI] [PubMed] [Google Scholar]
- Cunningham J.A., Rahme H., Hopkins G.D., Lebron C., Reinhard M. Enhanced in situ bioremediation of BTEX contaminated groundwater by combined injection of nitrate and sulfate. Environ Sci Technol. 2001;35:1663–1670. doi: 10.1021/es001722t. [DOI] [PubMed] [Google Scholar]
- Da Silva M.L.B., Alvarez P.J.J. Assessment of anaerobic benzene degradation potential using 16S rRNA gene‐targeted real‐time PCR. Environ Microbiol. 2007;9:72–80. doi: 10.1111/j.1462-2920.2005.01116.x. [DOI] [PubMed] [Google Scholar]
- Davidova I.A., Gieg L.M., Duncan K.E., Suflita J.M. Anaerobic phenanthrene mineralization by a carboxylating sulfate‐reducing bacterial enrichment. ISME J. 2007;1:436–442. doi: 10.1038/ismej.2007.48. [DOI] [PubMed] [Google Scholar]
- DiDonato R.J., Young N.D., Butler J.E., Chin K.J., Hixson K.K., Mouser P. Genome sequence of the Deltaproteobacterial strain NaphS2 and analysis of differential gene expression during anaerobic growth on naphthalene. PLoS ONE. 2010;5:11. doi: 10.1371/journal.pone.0014072. et al. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Edwards E.A., Wills L.E., Reinhard M., Grbicgalic D. Anaerobic degradation of toluene and xylene by aquifer microorganisms under sulfate‐reducing conditions. Appl Environ Microbiol. 1992;58:794–800. doi: 10.1128/aem.58.3.794-800.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Essaid H.I., Bekins B.A., Herkelrath W.N., Delin G.N. Crude oil at the Bemidji site: 25 years of monitoring, modeling, and understanding. Ground Water. 2011 doi: 10.1111/j.1745-6584.2009.00654.x. [DOI] [PubMed] [Google Scholar]
- Ettwig K.F., Butler M.K., Le Paslier D., Pelletier E., Mangenot S., Kuypers M.M.M. Nitrite‐driven anaerobic methane oxidation by oxygenic bacteria. Nature. 2010;464:543–548. doi: 10.1038/nature08883. et al. [DOI] [PubMed] [Google Scholar]
- Fischer A., Theuerkorn K., Stelzer N., Gehre M., Thullner M., Richnow H.H. Applicability of stable isotope fractionation analysis for the characterization of benzene biodegradation in a BTEX‐contaminated aquifer. Environ Sci Technol. 2007;41:3689–3696. doi: 10.1021/es061514m. [DOI] [PubMed] [Google Scholar]
- Fischer A., Herklotz I., Herrmann S., Thullner M., Weelink S.A.B., Stams A.J.M. Combined carbon and hydrogen isotope fractionation investigations for elucidating benzene biodegradation pathways. Environ Sci Technol. 2008;42:4356–4363. doi: 10.1021/es702468f. et al. [DOI] [PubMed] [Google Scholar]
- Fischer A., Gehre M., Breitfeld J., Richnow H.H., Vogt C. Carbon and hydrogen isotope fractionation of benzene during biodegradation under sulfate‐reducing conditions: a laboratory to field site approach. Rapid Commun Mass Spectrom. 2009;23:2439–2447. doi: 10.1002/rcm.4049. [DOI] [PubMed] [Google Scholar]
- Flesher J.W., Myers S.R. Methyl‐substitution of benzene and toluene in preparations of human bone marrow. Life Sci. 1991;48:843–850. doi: 10.1016/0024-3205(91)90100-p. [DOI] [PubMed] [Google Scholar]
- Foght J. Anaerobic biodegradation of aromatic hydrocarbons: pathways and prospects. J Mol Microbiol Biotechnol. 2008;15:93–120. doi: 10.1159/000121324. [DOI] [PubMed] [Google Scholar]
- Gibson D.T., Parales R.E. Aromatic hydrocarbon dioxygenases in environmental biotechnology. Curr Opin Biotechnol. 2000;11:236–243. doi: 10.1016/s0958-1669(00)00090-2. [DOI] [PubMed] [Google Scholar]
- Grbic‐Galic D., Vogel T.M. Transformation of toluene and benzene by mixed methanogenic cultures. Appl Environ Microbiol. 1987;53:254–260. doi: 10.1128/aem.53.2.254-260.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Griebler C., Mindl B., Slezak D., Geiger‐Kaiser M. Distribution patterns of attached and suspended bacteria in pristine and contaminated shallow aquifers studied with an in situ sediment exposure microcosm. Aquat Microb Ecol. 2002;28:117–129. [Google Scholar]
- Harvey R.W., Smith R.L., George L. Effect of organic contamination upon microbial distributions and heterotrophic uptake in a cape cod, mass, aquifer. Appl Environ Microbiol. 1984;48:1197–1202. doi: 10.1128/aem.48.6.1197-1202.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hazen T.C., Jimenez L., Devictoria G.L., Fliermans C.B. Comparison of bacteria from deep subsurface sediment and adjacent groundwater. Microb Ecol. 1991;22:293–304. doi: 10.1007/BF02540231. [DOI] [PubMed] [Google Scholar]
- Heider J. Adding handles to unhandy substrates: anaerobic hydrocarbon activation mechanisms. Curr Opin Chem Biol. 2007;11:188–194. doi: 10.1016/j.cbpa.2007.02.027. [DOI] [PubMed] [Google Scholar]
- Herrmann S., Vogt C., Fischer A., Kuppardt A., Richnow H.H. Characterization of anaerobic xylene biodegradation by two‐dimensional isotope fractionation analysis. Environ Microbiol Rep. 2009;1:535–544. doi: 10.1111/j.1758-2229.2009.00076.x. [DOI] [PubMed] [Google Scholar]
- Herrmann S., Kleinsteuber S., Chatzinotas A., Kuppardt S., Lueders T., Richnow H.H., Vogt C. Functional characterization of an anaerobic benzene‐degrading enrichment culture by DNA stable isotope probing. Environ Microbiol. 2010;12:401–411. doi: 10.1111/j.1462-2920.2009.02077.x. [DOI] [PubMed] [Google Scholar]
- Holm P.E., Nielsen P.H., Albrechtsen H.J., Christensen T.H. Importance of unattached bacteria and bacteria attached to sediment in determining potentials for degradation of xenobiotic organic contaminants in an aerobic aquifer. Appl Environ Microbiol. 1992;58:3020–3026. doi: 10.1128/aem.58.9.3020-3026.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Imachi H., Sekiguchi Y., Kamagata Y., Loy A., Qiu Y.L., Hugenholtz P. Non‐sulfate‐reducing, syntrophic bacteria affiliated with Desulfotomaculum cluster I are widely distributed in methanogenic environments. Appl Environ Microbiol. 2006;72:2080–2091. doi: 10.1128/AEM.72.3.2080-2091.2006. et al. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jahn M.K., Haderlein S.B., Meckenstock R.U. Anaerobic degradation of benzene, toluene, ethylbenzene, and o‐xylene in sediment‐free iron‐reducing enrichment cultures. Appl Environ Microbiol. 2005;71:3355–3358. doi: 10.1128/AEM.71.6.3355-3358.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karlsson A., Ejlertsson J., Svensson B.H. CO2‐dependent fermentation of phenol to acetate, butyrate and benzoate by an anaerobic, pasteurised culture. Arch Microbiol. 2000;173:398–402. doi: 10.1007/s002030000160. [DOI] [PubMed] [Google Scholar]
- Kasai Y., Takahata Y., Manefield M., Watanabe K. RNA‐based stable isotope probing and isolation of anaerobic benzene‐degrading bacteria from gasoline‐contaminated groundwater. Appl Environ Microbiol. 2006;72:3586–3592. doi: 10.1128/AEM.72.5.3586-3592.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kazumi J., Caldwell M.E., Suflita J.M., Lovley D.R., Young L.Y. Anaerobic degradation of benzene in diverse anoxic environments. Environ Sci Technol. 1997;31:813–818. [Google Scholar]
- Keller M., Zengler K. Tapping into microbial diversity. Nat Rev Microbiol. 2004;2:141–150. doi: 10.1038/nrmicro819. [DOI] [PubMed] [Google Scholar]
- Kleinsteuber S., Schleinitz K.M., Breitfeld J., Harms H., Richnow H.H., Vogt C. Molecular characterization of bacterial communities mineralizing benzene under sulfate‐reducing conditions. FEMS Microbiol Ecol. 2008;66:143–157. doi: 10.1111/j.1574-6941.2008.00536.x. [DOI] [PubMed] [Google Scholar]
- Knoll G., Winter J. Anaerobic degradation of phenol in sewage sludge – benzoate formation from phenol and CO2 in the presence of hydrogen. Appl Microbiol Biotechnol. 1987;25:384–391. [Google Scholar]
- Kobayashi T., Hashinaga T., Mikami E., Suzuki T. Methanogenic degradation of phenol and benzoate in acclimated sludges. Water Sci Technol. 1989;21:55–65. [Google Scholar]
- Kölbel‐Boelke J., Anders E.M., Nehrkorn A. Microbial communities in the saturated groundwater environment. 2: diversity of bacterial communities in a pleistocene sand aquifer and their in vitro activities. Microb Ecol. 1988;16:31–48. doi: 10.1007/BF02097403. [DOI] [PubMed] [Google Scholar]
- Kunapuli U., Lueders T., Meckenstock R.U. The use of stable isotope probing to identify key iron‐reducing microorganisms involved in anaerobic benzene degradation. ISME J. 2007;1:643. doi: 10.1038/ismej.2007.73. [DOI] [PubMed] [Google Scholar]
- Kunapuli U., Griebler C., Beller H.R., Meckenstock R.U. Identification of intermediates formed during anaerobic benzene degradation by an iron‐reducing enrichment culture. Environ Microbiol. 2008;10:1703–1712. doi: 10.1111/j.1462-2920.2008.01588.x. [DOI] [PubMed] [Google Scholar]
- Kung J.W., Loffler C., Dorner K., Heintz D., Gallien S., Van Dorsselaer A. Identification and characterization of the tungsten‐containing class of benzoyl‐coenzyme A reductases. Proc Natl Acad Sci USA. 2009;106:17687–17692. doi: 10.1073/pnas.0905073106. et al. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laban N.A., Selesi D., Jobelius C., Meckenstock R.U. Anaerobic benzene degradation by Gram‐positive sulfate‐reducing bacteria. FEMS Microbiol Ecol. 2009;68:300–311. doi: 10.1111/j.1574-6941.2009.00672.x. [DOI] [PubMed] [Google Scholar]
- Laban N.A., Selesi D., Rattei T., Tischler P., Meckenstock R.U. Identification of enzymes involved in anaerobic benzene degradation by a strictly anaerobic iron‐reducing enrichment culture. Environ Microbiol. 2010;12:2783–2796. doi: 10.1111/j.1462-2920.2010.02248.x. [DOI] [PubMed] [Google Scholar]
- Langenhoff A.A.M., Zehnder A.J.B., Schraa G. Behaviour of toluene, benzene and naphthalene under anaerobic conditions in sediment columns. Biodegradation. 1996;7:267–274. [Google Scholar]
- Liou J.S.C., DeRito C.M., Madsen E.L. Field‐based and laboratory stable isotope probing surveys of the identities of both aerobic and anaerobic benzene‐metabolizing microorganisms in freshwater sediment. Environ Microbiol. 2008;10:1964–1977. doi: 10.1111/j.1462-2920.2008.01612.x. [DOI] [PubMed] [Google Scholar]
- Lovley D.R. Anaerobic benzene degradation. Biodegradation. 2000;11:107–116. doi: 10.1023/a:1011191220463. [DOI] [PubMed] [Google Scholar]
- Lovley D.R., Lonergan D.J. Anaerobic oxidation of toluene, phenol, and para‐cresol by the dissimilatory iron‐reducing organism, GS 15. Appl Environ Microbiol. 1990;56:1858–1864. doi: 10.1128/aem.56.6.1858-1864.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lovley D.R., Woodward J.C., Chapelle F.H. Stimulated anoxic biodegradation of aromatic hydrocarbons using Fe(III) ligands. Nature. 1994;370:128–131. doi: 10.1038/370128a0. [DOI] [PubMed] [Google Scholar]
- Lovley D.R., Coates J.D., Woodward J.C., Phillips E.J.P. Benzene oxidation coupled to sulfate reduction. Appl Environ Microbiol. 1995;61:953–958. doi: 10.1128/aem.61.3.953-958.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lovley D.R., Woodward J.C., Chapelle F.H. Rapid anaerobic benzene oxidation with a variety of chelated Fe(III) forms. Appl Environ Microbiol. 1996;62:288–291. doi: 10.1128/aem.62.1.288-291.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Major D.W., Mayfield C.I., Barker J.F. Biotransformation of benzene by denitrification in aquifer sand. Ground Water. 1988;26:8–14. [Google Scholar]
- Mancini S.A., Ulrich A.C., Lacrampe‐Couloume G., Sleep B., Edwards E.A., Lollar B.S. Carbon and hydrogen isotopic fractionation during anaerobic biodegradation of benzene. Appl Environ Microbiol. 2003;69:191–198. doi: 10.1128/AEM.69.1.191-198.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mancini S.A., Devine C.E., Elsner M., Nandi M.E., Ulrich A.C., Edwards E.A., Sherwood Lollar B. Isotopic evidence suggests different initial reaction mechanisms for anaerobic benzene biodegradation. Environ Sci Technol. 2008;42:8290–8296. doi: 10.1021/es801107g. [DOI] [PubMed] [Google Scholar]
- Meyerdierks A., Kube M., Kostadinov I., Teeling H., Glockner F.O., Reinhardt R., Amann R. Metagenome and mRNA expression analyses of anaerobic methanotrophic archaea of the ANME‐1 group. Environ Microbiol. 2010;12:422–439. doi: 10.1111/j.1462-2920.2009.02083.x. [DOI] [PubMed] [Google Scholar]
- Morasch B., Hohener P., Hunkeler D. Evidence for in situ degradation of mono‐and polyaromatic hydrocarbons in alluvial sediments based on microcosm experiments with C‐13‐labeled contaminants. Environ Pollut. 2007;148:739–748. doi: 10.1016/j.envpol.2007.01.029. [DOI] [PubMed] [Google Scholar]
- Musat F., Widdel F. Anaerobic degradation of benzene by a marine sulfate‐reducing enrichment culture, and cell hybridization of the dominant phylotype. Environ Microbiol. 2008;10:10–19. doi: 10.1111/j.1462-2920.2007.01425.x. [DOI] [PubMed] [Google Scholar]
- Musat F., Galushko A., Jacob J., Widdel F., Kube M., Reinhardt R. Anaerobic degradation of naphthalene and 2‐methylnaphthalene by strains of marine sulfate‐reducing bacteria. Environ Microbiol. 2009;11:209–219. doi: 10.1111/j.1462-2920.2008.01756.x. et al. [DOI] [PubMed] [Google Scholar]
- Nales M., Butler B.J., Edwards E.A. Anaerobic benzene biodegradation: a microcosm survey. Bioremediat J. 1998;2:125–144. [Google Scholar]
- Oka A.R., Phelps C.D., McGuinness L.M., Mumford A., Young L.Y., Kerkhof L.J. Identification of critical members in a sulfidogenic benzene‐degrading consortium by DNA stable isotope probing. Appl Environ Microbiol. 2008;74:6476–6480. doi: 10.1128/AEM.01082-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Phelps C.D., Young L.Y. Anaerobic biodegradation of BTEX and gasoline in various aquatic sediments. Biodegradation. 1999;10:15–25. doi: 10.1023/a:1008303729431. [DOI] [PubMed] [Google Scholar]
- Phelps C.D., Kazumi J., Young L.Y. Anaerobic degradation of benzene in BTX mixtures dependent on sulfate reduction. FEMS Microbiol Lett. 1996;145:433–437. doi: 10.1111/j.1574-6968.1996.tb08612.x. [DOI] [PubMed] [Google Scholar]
- Phelps C.D., Kerkhof L.J., Young L.Y. Molecular characterization of a sulfate‐reducing consortium which mineralizes benzene. FEMS Microbiol Ecol. 1998;27:269–279. [Google Scholar]
- Phelps C.D., Zhang Z.M., Young L.Y. Use of stable isotopes to identify benzoate as a metabolite of benzene degradation in a sulphidogenic consortium. Environ Microbiol. 2001;3:600–603. doi: 10.1046/j.1462-2920.2001.00228.x. [DOI] [PubMed] [Google Scholar]
- Qiu Y.L., Hanada S., Ohashi A., Harada H., Kamagata Y., Sekiguchi Y. Syntrophorhabdus aromaticivorans gen. nov., sp nov., the first cultured anaerobe capable of degrading phenol to acetate in obligate syntrophic associations with a hydrogenotrophic methanogen. Appl Environ Microbiol. 2008;74:2051–2058. doi: 10.1128/AEM.02378-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ram R.J., VerBerkmoes N.C., Thelen M.P., Tyson G.W., Baker B.J., Blake R.C. Community proteomics of a natural microbial biofilm. Science. 2005;308:1915–1920. et al. [PubMed] [Google Scholar]
- Rappe M.S., Giovannoni S.J. The uncultured microbial majority. Annu Rev Microbiol. 2003;57:369–394. doi: 10.1146/annurev.micro.57.030502.090759. [DOI] [PubMed] [Google Scholar]
- Reinhard M., Hopkins G.D., Steinle‐Darling E., LeBron C.A. In situ biotransformation of BTEX compounds under methanogenic conditions. Ground Water Monit Remediat. 2005;25:50–59. [Google Scholar]
- Rooney‐Varga J.N., Anderson R.T., Fraga J.L., Ringelberg D., Lovley D.R. Microbial communities associated with anaerobic benzene degradation in a petroleum‐contaminated aquifer. Appl Environ Microbiol. 1999;65:3056–3063. doi: 10.1128/aem.65.7.3056-3063.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruiz‐Aguilar G.M.L., O'Reilly K., Alvarez P.J.J. A comparison of benzene and toluene plume lengths for sites contaminated with regular vs. ethanol‐amended gasoline. Ground Water Monit Remediat. 2003;23:48–53. [Google Scholar]
- Safinowski M., Meckenstock R.U. Methylation is the initial reaction in anaerobic naphthalene degradation by a sulfate‐reducing enrichment culture. Environ Microbiol. 2006;8:347–352. doi: 10.1111/j.1462-2920.2005.00900.x. [DOI] [PubMed] [Google Scholar]
- Sakai N., Kurisu F., Yagi O., Nakajima F., Yamamoto K. Identification of putative benzene‐degrading bacteria in methanogenic enrichment cultures. J Biosci Bioeng. 2009;108:501–507. doi: 10.1016/j.jbiosc.2009.06.005. [DOI] [PubMed] [Google Scholar]
- Salinero K.K., Keller K., Feil W.S., Feil H., Trong S., Di Bartolo G., Lapidus A. Metabolic analysis of the soil microbe Dechloromonas aromatica str. RCB: indications of a surprisingly complex life‐style and cryptic anaerobic pathways for aromatic degradation. BMC Genomics. 2009;10:23. doi: 10.1186/1471-2164-10-351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schleinitz K.M., Schmeling S., Jehmlich N., von Bergen M., Harms H., Kleinsteuber S. Phenol degradation in the strictly anaerobic iron‐reducing bacterium Geobacter metallireducens GS‐15. Appl Environ Microbiol. 2009;75:3912–3919. doi: 10.1128/AEM.01525-08. et al. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Söhngen N.L. Benzin, Petroleum, Paraffinol und Paraffin als Kohlenstoff‐ und Energiequellen für Mikroben. Centr Bakteriol Parasitenk Abt II. 1913;37:595–609. [Google Scholar]
- Tan N.C.G., van Doesburg W., Langenhoff A.A.M., Stams A.J.M. Benzene degradation coupled with chlorate reduction in a soil column study. Biodegradation. 2006;17:11–17. doi: 10.1007/s10532-005-5335-z. [DOI] [PubMed] [Google Scholar]
- Tao Y., Fishman A., Bentley W.E., Wood T.K. Oxidation of benzene to phenol, catechol, and 1,2,3‐trihydroxybenzene by toluene 4‐monooxygenase of Pseudomonas mendocina KR1 and toluene 3‐monooxygenase of Ralstonia pickettii PKO1. Appl Environ Microbiol. 2004;70:3814–3820. doi: 10.1128/AEM.70.7.3814-3820.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tschech A., Fuchs G. Anaerobic degradation of phenol by pure cultures of newly isolated denitrifying Pseudomonads. Arch Microbiol. 1987;148:213–217. doi: 10.1007/BF00414814. [DOI] [PubMed] [Google Scholar]
- Ulrich A.C., Edwards E.A. Physiological and molecular characterization of anaerobic benzene‐degrading mixed cultures. Environ Microbiol. 2003;5:92–102. doi: 10.1046/j.1462-2920.2003.00390.x. [DOI] [PubMed] [Google Scholar]
- Ulrich A.C., Beller H.R., Edwards E.A. Metabolites detected during biodegradation of C‐13(6)‐benzene in nitrate‐reducing and methanogenic enrichment cultures. Environ Sci Technol. 2005;39:6681–6691. doi: 10.1021/es050294u. [DOI] [PubMed] [Google Scholar]
- Vaillancourt F.H., Bolin J.T., Eltis L.D. The ins and outs of ring‐cleaving dioxygenases. Crit Rev Biochem Mol Biol. 2006;41:241–267. doi: 10.1080/10409230600817422. [DOI] [PubMed] [Google Scholar]
- Villatoro‐Monzon W.R., Mesta‐Howard A.M., Razo‐Flores E. Anaerobic biodegradation of BTEX using Mn(IV) and Fe(III) as alternative electron acceptors. Water Sci Technol. 2003;48:125–131. [PubMed] [Google Scholar]
- Villatoro‐Monzon W.R., Morales‐Ibarria M.G., Velazquez E.K., Ramirez‐Saad H., Razo‐Flores E. Benzene biodegradation under anaerobic conditions coupled with metal oxides reduction. Water Air Soil Pollut. 2008;192:165–172. [Google Scholar]
- Vogel T.M., Grbic‐Galic D. Incorporation of oxygen from water into toluene and benzene during anaerobic fermentative transformation. Appl Environ Microbiol. 1986;52:200–202. doi: 10.1128/aem.52.1.200-202.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vogt C., Godeke S., Treutler H.C., Weiss H., Schirmer M., Richnow H.H. Benzene oxidation under sulfate‐reducing conditions in columns simulating in situ conditions. Biodegradation. 2007;18:625–636. doi: 10.1007/s10532-006-9095-1. [DOI] [PubMed] [Google Scholar]
- Vogt C., Cyrus E., Herklotz I., Schlosser D., Bahr A., Herrmann S. Evaluation of toluene degradation pathways by two‐dimensional stable isotope fractionation. Environ Sci Technol. 2008;42:7793–7800. doi: 10.1021/es8003415. et al. [DOI] [PubMed] [Google Scholar]
- Wang Q., Garrity G.M., Tiedje J.M., Cole J.R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol. 2007;73:5261–5267. doi: 10.1128/AEM.00062-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weelink S.A.B., Tan N.C.G., ten Broeke H., van Doesburg W., Langenhoff A.A.M., Gerritse J., Stams A.J.M. Physiological and phylogenetic characterization of a stable benzene‐degrading, chlorate‐reducing microbial community. FEMS Microbiol Ecol. 2007;60:312–321. doi: 10.1111/j.1574-6941.2007.00289.x. [DOI] [PubMed] [Google Scholar]
- Weelink S.A.B., Tan N.C.G., ten Broeke H., van den Kieboom C., van Doesburg W., Langenhoff A.A.M. Isolation and characterization of Alicycliphilus denitrificans strain BC, which grows on benzene with chlorate as the electron acceptor. Appl Environ Microbiol. 2008;74:6672–6681. doi: 10.1128/AEM.00835-08. et al. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weelink S.A.B., van Eekert M.H.A., Stams A.J.M. Degradation of BTEX by anaerobic bacteria: physiology and application. Rev Environ Sci Bio/Technol. 2010;9:359–385. [Google Scholar]
- Weiner J.M., Lovley D.R. Anaerobic benzene degradation in petroleum‐contaminated aquifer sediments after inoculation with a benzene‐oxidizing enrichment. Appl Environ Microbiol. 1998a;64:775–778. doi: 10.1128/aem.64.2.775-778.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiner J.M., Lovley D.R. Rapid benzene degradation in methanogenic sediments from a petroleum‐contaminated aquifer. Appl Environ Microbiol. 1998b;64:1937–1939. doi: 10.1128/aem.64.5.1937-1939.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Widdel F., Rabus R. Anaerobic biodegradation of saturated and aromatic hydrocarbons. Curr Opin Biotechnol. 2001;12:259–276. doi: 10.1016/s0958-1669(00)00209-3. [DOI] [PubMed] [Google Scholar]
- Wilson B.H., Smith G.B., Rees J.F. Biotransformations of selected alkylbenzenes and halogenated aliphatic hydrocarbons in methanogenic aquifer material – a microcosm study. Environ Sci Technol. 1986;20:997–1002. doi: 10.1021/es00152a005. [DOI] [PubMed] [Google Scholar]
- Winderl C., Schaefer S., Lueders T. Detection of anaerobic toluene and hydrocarbon degraders in contaminated aquifers using benzylsuccinate synthase (bssA) genes as a functional marker. Environ Microbiol. 2007;9:1035–1046. doi: 10.1111/j.1462-2920.2006.01230.x. [DOI] [PubMed] [Google Scholar]
- Zengler K., Richnow H.H., Rossello‐Mora R., Michaelis W., Widdel F. Methane formation from long‐chain alkanes by anaerobic microorganisms. Nature. 1999;401:266–269. doi: 10.1038/45777. [DOI] [PubMed] [Google Scholar]
- Zhang X.M., Wiegel J. Reversible conversion of 4‐hydroxybenzoate and phenol by Clostridium hydroxybenzoicum. Appl Environ Microbiol. 1994;60:4182–4185. doi: 10.1128/aem.60.11.4182-4185.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang X.M., Young L.Y. Carboxylation as an initial reaction in the anaerobic metabolism of naphthalene and phenanthrene by sulfidogenic consortia. Appl Environ Microbiol. 1997;63:4759–4764. doi: 10.1128/aem.63.12.4759-4764.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang T., Gannon S.M., Nevin K.P., Franks A.E., Lovley D.R. Stimulating the anaerobic degradation of aromatic hydrocarbons in contaminated sediments by providing an electrode as the electron acceptor. Environ Microbiol. 2010;12:1011–1020. doi: 10.1111/j.1462-2920.2009.02145.x. [DOI] [PubMed] [Google Scholar]


