Abstract
TNFα is a pleotropic cytokine that plays a central role in the inflammatory response by activating the NF-κB signaling pathway, and is targeted in a range of chronic inflammatory diseases, underscoring the therapeutic importance of understanding its underlying molecular mechanisms. Although K63-linked ubiquitination of RIP1 by TRAF2/5 and cIAP1/2 was thought to serve as a scaffold to activate the NF-κB pathway, the recent accumulation of conflicting results has challenged the necessity of these proteins in NF-κB activation. In addition, several serine/threonine kinases have been implicated in TNFα-induced IKK activation; however, the targeted disruption of these kinases had no effect on transient IKK activation. The recent discovery of RIP1-dependent and -independent activation of the early and delayed phases of IKK and TRAF2 phosphorylation-dependent activation of the prolonged phase of IKK offers a reconciliatory model for the interpretation of contradictory results in the field. Notably, the TNFα-induced inflammatory response is not exclusively controlled by the NF-κB pathway but is subject to regulatory crosstalk between NF-κB and other context-dependent pathways. Thus further elucidation of these spatiotemporally-coordinated signaling mechanisms has the potential to provide novel molecular targets and therapeutic strategies for NF-κB intervention.
Keywords: TNFα, RIP1, TRAF2, cIAP1, NF-κB
1. Introduction
Tumor necrosis factor-alpha (TNFα) regulates the host immune and inflammatory response by activating the nuclear factor-κB (NF-κB) pathway [1, 2]. While TNFα and NF-κB are required for protective inflammatory responses, excessive production of TNFα and resulting dysregulation of NF-κB are attributed to the pathologies of several human diseases, including chronic inflammatory disorders (CIDs) and cancer, making them ideal therapeutic targets for pharmacological inhibition. Importantly, TNFα blockade in some CIDs reduces the production of other pro-inflammatory cytokines, such as interleukin-1 (IL-1), IL-6 and IL-8, suggesting that TNFα plays a major role in the development of certain CIDs [3, 4]. Despite the therapeutic popularity of TNFα antagonists, the meta-analysis of long-term patient response data has exposed the unpredictable nature of these therapies, which can range from a low therapeutic responder to the onset of drug-induced autoimmune disease resulting from an increased production of autoantibodies [3]. Thus, a better understanding of the molecular mechanisms that control the strength and duration of TNFα signaling is necessary.
The NF-κB family of transcription factors consists of five members: p105/p50 (NF-κB1), p100/p52 (NF-κB2), p65 (RelA), c-Rel and RelB [5]. All NF-κB family members contain an N-terminal Rel homology domain (RHD) that mediates DNA binding and homo- and hetero-dimerization [5, 6]. Notably, only p65, c-Rel, and RelB harbor a transcription activation domain (TAD), whereas p50 and p52 lack this TAD, and thereby may repress transcription when they are not associated with p65, c-Rel or RelB. In most cell types, NF-κB dimers are retained in the cytoplasm by a family of inhibitory proteins known as inhibitors of NF-κB (IκBs) [5, 7]. Mammalian cells express three classical IκBs (IκBα, IκBβ and IκBε) and two novel IκBs (IκBζ and BCL-3). The IκB proteins are characterized by the presence of multiple ankyrin repeat domains that mediate IκB binding to NF-κB dimers and thereby interfere with the nuclear translocation of the NF-κB dimers. The C-terminal halves of the p105 and p100 proteins—the precursors of p50 and p52, respectively—also harbor multiple ankyrin repeats. This allows them to serve an IκB-like function, retaining the partners in the cytoplasm [5, 6].
2. TNFR1: a prototypical member of the TNF receptor superfamily
The TNF receptor (TNFR) superfamily is comprised of 29 receptors that serve to promote the innate immune defense, inflammatory response, and lymphoid organogenesis [2, 8]. These receptors are characterized by the presence of cysteine-rich subdomains in the extracellular ligand-binding domains. In contrast, their intracellular domains show no sequence homology, and are broadly divided into two groups based on the effectors recruited to their cytoplasmic domains [1, 2]. Receptors in the first group, such as TNFR1, Fas and death receptor 4 and 5 (DR4/5), contain an intracellular death domain (DD), whereas this domain is absent in the second group, such as TNFR2 and CD40 [1, 8].
2.1. Overview of TNFR family signaling
The TNFR superfamily members are devoid of intrinsic kinase activity, and thus initiate pro-inflammatory or pro-apoptotic pathways by recruiting one or more members of the TNFR-associated factor (TRAF) family or FAS-associated protein with a death-domain (FADD) [1, 8]. With the exception of TRAF1, TRAFs contain an N-terminal RING domain followed by several zinc finger (ZF) motifs, and are believed to function as E3 ubiquitin (Ub) ligases to induce the activation of the NF-κB and c-Jun N-terminal kinase (JNK) pathways [5, 9]. FADD contains both a DD and a death effector domain (DED), and thus can interact directly with DD domain containing proteins including Fas, DR4/5 and TNFR-associated death domain protein (TRADD) through homophilic DD interaction, as well as with DED domain containing proteins such as caspase-8 and cellular caspase-8 (FLICE)-like inhibitory protein (cFLIP) [10, 11]. Ligation of TNFR1 results in the recruitment of TRAFs and FADD indirectly through TRADD, and can lead to activation of both pro- and anti-apoptotic pathways depending on cellular context [5, 8]. On the other hand, ligand-bound TNFR2 and CD40 can directly recruit TRAFs to activate the pro-inflammatory JNK and NF-κB pathways, whereas ligated Fas and DR4/5 mainly activates the pro-apoptotic caspase-8 pathway by direct recruitment of FADD [10, 11].
NF-κB activation occurs mainly through two signaling pathways—the canonical and the non-canonical [6]. The canonical NF-κB pathway is activated by a variety of stimuli, including cytokines, growth factors, antigen receptors, pathogen associated molecular patterns (PAMPs), and cellular stresses. In contrast, the non-canonical NF-κB pathway is activated only by a subset of TNF family members, such as lymphotoxin α/β (LTα/β), B-cell activating factor (BAFF) and the CD40 ligand (CD40L) [5, 6]. Canonical NF-κB activation typically involves the dual phosphorylation of IκBα on serine 32/36 by the IKK complex, which leads to its ubiquitination-mediated degradation by the 26S proteasome, releasing NF-κB (most commonly the p65/p50 dimer) to translocate to the nucleus and activating the expression of genes involved in inflammation and the innate immune response. Non-canonical NF-κB activation requires TRAF2- and cellular inhibitors of apoptosis 1 and 2 (cIAP1/2)-dependent ubiquitination and degradation of TRAF3 and the subsequent accumulation of NF-κB-inducing kinase (NIK), which is habitually targeted for degradation by the TRAF2/TRAF3/cIAP1/2 E3 ligase complex in unstimulated cells [5, 6]. Accumulated NIK can then activate the IKKα homodimer, which in turn phosphorylates p100, triggering its ubiquitination-mediated proteasomal processing to p52. As the RHD domain of p100 is most commonly associated with RelB, activation of this pathway leads to nuclear translocation of the RelB/p52 dimer, resulting in expression of genes involved in adaptive immunity and lymphoid organogenesis [5, 6].
2.2. Pro- and anti-apoptotic TNFR1 signaling
TNFα can exert opposing biological effects – pro-inflammatory or pro-apoptotic – through activation of the NF-κB and capase-8 pathways, respectively [1, 12]. TNFα binds to two cognate receptors, TNFR1 and TNFR2 [1, 12]. Gene knockout studies revealed that both the pro-inflammatory and pro-apoptotic activities of TNFα are mediated by TNFR1 [7, 13]. In general, it is believed that ligation of TNFR1 by soluble TNFα elicits the sequential formation of two complexes with opposing effects on cell fate. Initially, TNFR1 interacts with TRADD through a homophilic DD interaction to initiate the recruitment of receptor-interacting protein 1 (RIP1) and TRAF2; TRAF2 then recruits cIAP1/2 to form the membrane-bound complex-I, leading to activation of the pro-survival NF-κB pathway. Thereafter, the TRADD/RIP1/TRAF2/cIAP1/2 complex dissociates from TNFR1 and binds FADD and caspase-8 in the cytoplasm to form proapoptotic complex-IIA [14].
2.3. RIP1 ubiquitination and cFLIP expression tip the balance between life and death
TNFα/TNFR1 signaling can result in cell survival, apoptosis or necrosis, depending on the cellular context [5, 7]. The balance between these rival pathways is usually tipped by two sequential events, cIAP1/2-mediated RIP1 ubiquitination and NF-κB -dependent expression of cFLIP. RIP1 is the best-characterized member of the RIP family and is highly conserved in vertebrates [15]. It contains an N-terminal kinase domain (KD), a C-terminal DD domain and an intermediate domain (ID) between the KD and DD domains [15]. Transient overexpression of RIP1 has long been known to be sufficient to induce both NF-κB activation and cell death [16]. Under a certain condition, RIP1 can directly interact with FADD and caspase-8 through its DD domain, and trigger both apoptotic and necrotic cell death [17]. In the case of TNFR1 signaling, TRAF2 and cIAP1/2, in conjunction with the E2 conjugating enzymes UbcH5 and/or Ubc13/Uev1a, conjugate RIP1 with K63-linked poly-ubiquitin chain [18]. This K63-linked poly-ubiquitination not only serves as a platform to recruit effectors and initiate NF-κB activation, but also prevents RIP1 from interacting with FADD and caspase-8 [16, 18]. As such, in the presence of TRAF2 and cIAP1/2 expression, pro-survival NF-κB activation dominates over pro-death caspase-8 activation. It is worth emphasizing that RIP1 ubiquitination-dependent inhibition of TNFα-induced cell death is independent of NF-κB activation and target gene expression [16].
The second event that tips the balance towards pro-survival is the NF-κB -dependent expression of cFLIP [14]. cFLIP contains two death effector domains (DEDs) at its N-terminus and a caspase-like domain at its C-terminus, and thus resembles caspase-8 structurally [10]. However, cFLIP lacks caspase activity owing to the substitution of critical amino acids in its caspase-like domain [10]. Following TNFα stimulation and NF-κB activation, the TRADD/RIP1/TRAF2/cIAP1/2 complex translocates from TNFR1 to the cytoplasm where it recruits FADD and caspase-8 to form the death-inducing signaling complex (DISC; also known as complex-IIA)[14]. However, under a physiological condition, NF-κB activation leads to inducible expression of cFLIP, which in turn competes with caspase-8 for incorporation into the DISC, and consequently limiting caspase-8 activation. Thus, if either NF-κB activation or protein synthesis is inhibited, DISC triggers caspase-8 activation and results in mitochondria-dependent and -independent apoptosis depending on cell types [5, 14].
Notably, TNFα can also induce necrosis in certain experimental and pathological conditions. For example, if both caspase activity and protein synthesis are hindered by pharmacological inhibition, TNFα can cause necrosis in a manner dependent on RIP1 kinase activity and RIP3 expression [17]. Furthermore, depletion of cIAP1/2 by SMAC-mimetics (SM) followed by stimulation of the cells with TNFα induces the formation of a new cytoplasmic death-inducing complex, known as complex-IIB, which is different from complex-IIA, as it contains FADD, caspase-8, RIP1 and RIP3 rather than TRADD[19]. Moreover, depletion of cIAP1/2 and inhibition of basal RIP1 ubiquitination by stress-inducing agents such as etoposide induce the formation of yet another cytoplasmic death-inducing complex known as ripoptosome or necroptosome, which also contains FADD, caspase-8, RIP1 and potentially RIP3 [20, 21]. Although the composition of complex-IIB and the ripoptosome is similar, they are induced by different stimuli—the former by TNFα and the latter by cellular stress. In addition, cFLIP appear to suppress the formation and activation of the ripoptosome, but not complex-IIB. Collectively, these findings suggest that although different types of death stimuli may trigger necrosis through different pathways, the activity the necrosis-inducing core complex is tightly regulated by cIAP1/2-mediated RIP1 ubiquitination and NF-κB -dependent cFLIP expression.
3. RIP1-dependent and -independent NF-κB activation pathways
The current model for TNFα-induced NF-κB activation has been as follows: upon TNFα stimulation, TNFR1 recruits TRADD, RIP1, TRAF2/5 and cIAP1/2; the RING domains of cIAP1/2 work in conjunction with UbcH5 to catalyze K63-linked poly-ubiquitination of RIP1 (RIP1-K63-pUb) at K377 [18, 22, 23]; the RIP1-K63-pUb chain then recruits the TAK1 (consisting of TAK1, TAB1 andTAB2 subunits) and IKK (consisting of IKKα, IKKβ and IKKγ/NEMO subunits) complexes by interacting with the Ub-binding domains present on TAB2 and NEMO; once recruited by RIP1-K63-pUb chains, TAK1 directly activates IKKβ through proximity-mediated phosphorylation [18, 24, 25]. Recently, linear ubiquitination of NEMO and RIP1 by LUBAC (linear ubiquitin chain assembly complex consisting of HOIP, HOIL-1L and/or SHARPIN) has also been reported to be required for TNFα-induced NF-κB activation [26, 27].
3.1. Controversies and complexities of NF-κB activation mechanisms
Ironically, although TNFR1 signaling is among the most-studied pathways, our understanding of the molecular mechanisms underlying TNFR1-mediated activation of NF-κB has been complicated by the recent accumulation of apparently contradictory results [24, 25, 28-34]. Especially, the identities of the E2 enzymes (Ubc13 vs. UbcH5) and E3 ligases (TRAF2 vs. cIAP1), as well as the topology of the conjugated polyubiquitin chain remain controversial [35].
The RING domain of TRAF2 is believed to possess E3 ligase activity, but recent structural studies revealed that the distal N-terminus and nine amino acid residues between the RING and first zinc finger (ZF) domains sterically hinders interaction between the RING domain and E2 enzymes [30]. Conversely, a TRAF2-Ubc13 interaction can be observed by in vivo co-immunoprecipitation assays following TNFα stimulation, and this interaction appears to be destabilized by an Ub editing enzyme A20, as an alternative mechanism to limit RIP1 K63-linked ubiquitination and NF-κB activation [36]. It has also recently been reported that the bioactive lipid, sphingosine-1-phosphate (S1P), binds the TRAF2 RING domain to incite its E3 ligase activity [33]. Though TRAF2 and Ubc13 appear to interact in the presence of S1P or following TNFα stimulation, earlier studies demonstrated that disruption of Ubc13 by siRNA-mediated knockdown or genetic deletion has no effect on NF-κB activation in response to TNFα [29, 34]. A novel ubiquitin replacement strategy suggested that IKK activation by TNFα requires UbcH5 rather than Ubc13, whose poly-ubiquitination of RIP1 is not restricted to the K63-pUb chain [37]. In addition, K11-linked ubiquitination of RIP1 by cIAP1 and UbcH5 was also reported to occur and efficiently recruit NEMO to TNFR1 in response to TNFα stimulation [38].
Nevertheless, TNFα-induced RIP1 ubiquitination is impaired in TRAF2/5 double knockout (DKO) cells, because the recruitment of cIAP1/2 to TNFR1 is dependent on TRAF2 expression [7, 32]. Thus, TNFα-induced NF-κB activation in TRAF2/5 DKO cells was thought to be completely impaired [5, 7, 39]. Surprisingly, IKK kinase assays demonstrated that the classical IKK complex is constitutively activated to a certain degree in TRAF2 KO and TRAF2/5 DKO mouse embryonic fibroblasts (MEFs) due to the accumulation of NIK, and that stimulating these cells with TNFα further increases IKK activity, albeit weakly [32]. In line with this observation, stable expression of a TRAF2 mutant (in which the cIAP1/2-inetracting motif is deleted) in TRAF2/5 DKO cells attenuates, but does not completely abolish, TNFα-induced NF-κB activation [40]. Genetic deletion of both cIAP1 and cIAP2 also revealed that TNFα can still weakly activate NF-κB in cIAP1/2 DKO cells [41]. More surprisingly, a recent study showed that TNFα can activate NF-κB normally in various primary cells isolated from RIP1 KO mice [28]. In addition, although LUBAC has also been shown to be required for TNFα-induced NF-κB activation, Xu et al. reported that knockdown of HOIP (the catalytic component of LUBAC) has no effect on NF-κB activation [37].
3.2. Pitfalls of assessing NF-κB activation by analyzing IκBα degradation and target gene expression
One of the most common methods to assess stimulus-induced NF-κB activation is to examine IκBα degradation, as this event is essential for the release and nuclear translocation of canonical NF-κB dimers. In fact, the original conclusions that double knockout of TRAF2/5 or double knockdown (DKD) of cIAP1/2 completely impairs TNFα-induced NF-κB activation were based on an analysis of IκBα protein levels in these cells. However, IKK kinase assays revealed that the basal IKK activity is elevated and TNFα-induced IKK activation is attenuated, but not completely impaired, in TRAF2/5 DKO and cIAP1/2 DKD cells [32, 42]. Recent studies revealed the existence of extensive crosstalk between the canonical and non-canonical NF-κB pathways. For instance, accumulated NIK can activate both the classical IKK complex and the IKKα homodimer [32, 43]. NIK accumulates in TRAF2 KO, TRAF2/5 DKO, TRAF3 KO and cIAP1/2 DKD cells, and it is thus conceivable that basal IKK activity is elevated in these cells. In wild-type (WT) MEFs, TNFα-induced IκBα phosphorylation is robust and immediate, resulting in complete degradation of IκBα within 15 minutes post-stimulation. IκBα protein returns to a normal level within 60 minutes after TNFα stimulation, as the expression of IκBα itself is induced immediately by NF-κB [5, 7]. If, however, the IKK complex is constitutively activated to a certain degree, IκBα will likewise be constitutively phosphorylated, degraded and resynthesized, which will partially mask stimulation-induced IκBα degradation [32]. Thus, analysis of IκBα degradation in TRAF2/5 DKO and cIAP1/2 DKD cells does not clearly reflect the basal and inducible IKK activation [32, 42]. Collectively, these studies suggest that the IKK kinase assay is a more sensitive method to examine the basal and inducible activation of NF-κB.
Detection of target gene expression by RT-PCR is another common technique utilized to assess stimulus-induced NF-κB activation. NF-κB-dependent gene expression is regulated at multiple levels, including the post-translational modifications of NF-κB subunits (e.g. p65 phosphorylation by IKK, Akt, and PKCζ), the modification and accessibility of the target gene's promoter region, and the presence of additional cofactors resulting from cellular context or signal crosstalk [44-46]. For example, the expression of IP-10, ICAM-I and RANTES can be induced by both the canonical and non-canonical NF-κB pathways, whereas that of IL-6 through the regulated by cooperative actions of NF-κB and c-Jun [35, 47, 48]. As mentioned above, TNFα-induced expression of NF-κB target genes was thought to be impaired in TRAF2/5 DKO and cIAP1/2 DKD cells. However, real-time RT-PCR analysis of a set of NF-κB target genes in TRAF2/5 DKO MEFs revealed that while TNFα-induced IL-6 expression is completely impaired, the basal and inducible expression of IP-10, ICAM-I and RANTES are significantly increased compared to that observed in WT MEFs [32]. A possible explanation for these discrepancies is that the non-canonical NF-κB pathway is constitutively activated in TRAF2/5 DKO cells, while TNFα-induced activation of canonical NF-κB is attenuated but not impaired. As a result, these two NF-κB pathways synergistically induce the expression of IP-10, ICAM-I and RANTES in TRAF2/5 DKO MEFs, resulting in significantly elevated expression of these genes following TNFα stimulation. This could also partially explain an earlier observation in which TRAF2 KO macrophages were reported to overproduce TNFα and NO in response to TNFα stimulation [49]. In contrast, the absence of IL-6 induction in TRAF2/5 DKO cells is due impaired JNK activation, because the efficient induction of IL-6 depends on the activities of both c-Jun and NF-κB [32, 35, 47]. This is supported by the fact that TNFα-induced IL-6 expression is impaired in both JNK1/2 DKO and p65 KO MEFs [50, 51]. In addition, a subset of NF-κB target genes (e.g. IP-10 and IκBα) is particularly sensitive to NF-κB mediated transcription, as their promoters are immediately accessible to transcription factors. As a result, even transient NF-κB activation is sufficient to induce their transcription [35, 46]. In contrast, other promoters require stimulus-dependent modifications to chromatin structure in order to become accessible to NF-κB (e.g. RANTES and IL-6); thus they are activated only in the context of strong and prolonged NF-κB activation [48, 52]. Therefore, to better assess the role of TRAF2/5 and cIAP1/2 in TNFα-induced gene expression, it is necessary to examine the expression of a set of genes whose expression is regulated independently and/or cooperatively by canonical and non-canonical NF-κB, as well as by c-Jun.
3.3. Two parallel pathways mediate the early and delayed phases of IKK activation
Extensive analyses on the correlations amongst IKK activation, IκBα degradation and NF-κB dependent gene expression identified that the IKK kinase assay as a more sensitive means to assess the strength and duration of signaling output [31, 32]. A recent comprehensive analysis on the kinetics and amplitude of IKK activation in various KO, DKO and DKD cells revealed that while TNFα causes immediate and robust IKK activation in WT cells within 3 minutes of stimulation, this immediate IKK activation is delayed in RIP1 KO, TRAF2 KO, TRAF2/5 DKO and cIAP1/2 DKD cells; yet TNFα can still activate IKK in these KO, DKO and DKD cells 10 minutes after stimulation [42]. Interesting, knockdown of HOIP almost completely abolishes delayed IKK activation in RIP1 KO cells, but has no effect on immediate IKK activation in WT cells. Consistently, RIP1 ubiquitination is impaired in TRAF2/5 DKO and cIAP1/2 DKD but not in HOIP KD cells. Analysis of several NF-κB target genes revealed that while the induction of IP-10 and IκBα occurs normally, expression of IL-6 and RANTES is reduced significantly in RIP1 KO MEFs compared to that observed in WT MEFs [42]. These data provide evidence for the existence of two phases of transient IKK activation; the early phase induced by TRAF2/cIAP1-mediated ubiquitination of RIP1 which occurs within 3 minutes of TNFα stimulation, followed by the delayed phase activated by TRAF2/cIAP1-dependent recruitment of LUBAC 10 minutes post-stimulation. Thus, this study clarifies the details of TNFR1 signaling and further emphasizes the notion that an accurate assessment of TNFα-induced NF-κB activation requires evaluation of IKK activation kinetics and careful selection of NF-κB target genes for RT-PCR analysis.
Notably, in RIP1 KO cells, TNFα triggers increased auto-ubiquitination of TRAF2 and cIAP1/2 through K63- and K48-linkages, confirming that RIP1 as a bona fide substrate of TRAF2 and cIAP1/2, as E3 ligases undergo autoubiquitination in the absence of substrates [42, 53]. Interestingly, knockdown of HOIP in RIP1 KO cells has no effect on increased auto-ubiquitination of TRAF2 and cIAP1, though it completely impairs IKK activation [42]. Surprisingly, this increased K63-linked ubiquitination of TRAF2 is incapable of recruiting IKK to TNFR1 in RIP1 KO cells in the absence of HOIP, suggesting that the K63-pUb chain itself is not sufficient for immediate IKK recruitment in vivo. In addition, knockdown of HOIP in WT MEFs has no significant effect on TNFα-induced linear ubiquitination of RIP1 and immediate activation of IKK, suggesting that RIP1 linear ubiquitination can also be catalyzed by other E3 ligases in addition to LUBAC. Gerlach et al. reported by mass spectrometric analysis of the TNFR1 complex that RIP1 can be ubiquitinated through K48-, K63-, K11- and linear-linkages [54]. Furthermore, in vitro binding and ubiquitination assays with lysine-mutant forms of Ub demonstrated that cIAP1 can bind directly to and is capable of conjugating RIP1 with diverse types of ubiquitin chains, including K63- and linear-linkages [24, 25, 55]. Of note, when Ub acceptor sites are mutated, E3 ligases often conjugate Ub to alternative sites on substrates [23, 35]. Interestingly, when in vitro ubiquitination assays are performed with WT Ub, TRAF2 and cIAP1 together, but not either one alone, more efficiently and directly conjugates RIP1 with both K63- and linearly-linked pUb chains [42]. TRAF2 binds cIAP1/2 through its C-terminal TRAF-N domain. However, stable expression of the N-terminal RING domain-deleted TRAF2 in TRAF2 KO and TRAF2/5 DKO cells neither fully restores immediate RIP1 ubiquitination nor efficiently suppress constitutive p100 processing [31, 40]. These data suggest that TRAF2's function is not only to recruit cIAP1 but also to direct cIAP1-mediated RIP1 ubiquitination towards linear- and K63-linkages, and that linear-pUb chains recruit the IKK complex and K63-pUb chains recruit the TAK1 and LUBAC complexes (Fig. 1A). It is also likely that although NEMO binds to K63-pUb chains in vitro, it may be less competitive than TAB2 and HOIP for binding to K63-pUb chains in vivo.
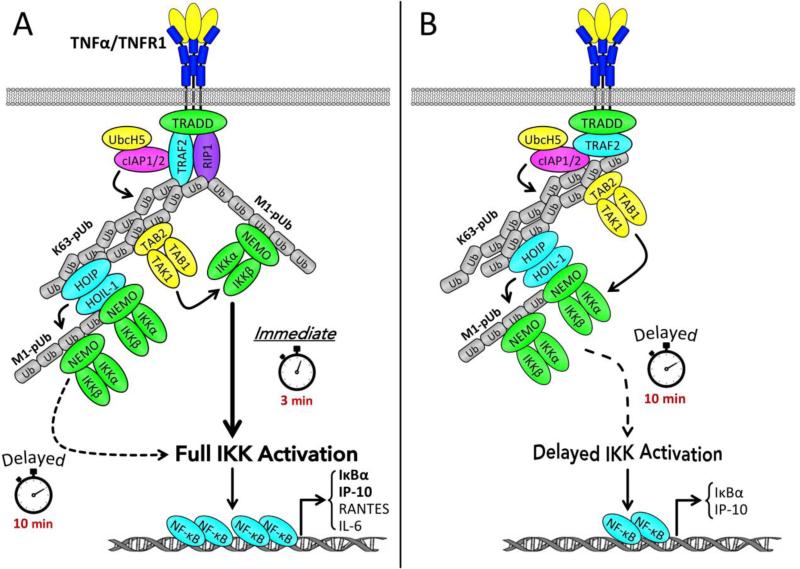
Fig. 1.
TNFα-mediated transient IKK activation pathways. (A) Signaling pathways to full IKK activation. Upon ligation, TNFR1 recruits TRADD, RIP1, TRAF2, and cIAP1/2. TRAF2 and cIAP1/2, in conjunction with UbcH5, catalyze RIP1 ubiquitination through linear (M1-pUb) and K63 linkages (K63-pUb), and also undergo autoubiquitination through K63-linkage. K63-pUb recruits the TAK1-TAB1-TAB2 complex, and M1-pUb recruits the IKKα-IKKβ-NEMO complex, leading to TAK1-mediated activation of the early phase of IKK. Thereafter, K63-pUb recruits the HOIL-1L–HOIP complex, which in turn recruits and catalyzes linear ubiquitination of NEMO, leading to more IKK recruitment and full activation of IKK. Such full IKK activation is required for the efficient expression of NF-κB target genes. (B) RIP1-independent pathway to delayed IKK activation. RIP1 protein is not detectable in embryonic hepatocytes. As a result, upon TNFR1 ligation, receptor-recruited TRAF2 and cIAP1/2 undergo increased autoubiquitination through K63 and K48 linkages. The K63-pUb chains then recruit the TAK1-TAB1-TAB2 and HOIP–HOIL-1 complexes, leading to activation of the delayed phase of IKK. Such delayed IKK activation is sufficient for induction of some NF-κB target genes (e.g., IκBα and IP-10).
3.4. Why are redundant pathways required for TNFα-induced NF-κB activation?
One of the perplexing questions in the field of TNFR1 study is the physiological justification for the abundance of E3 ligases and pUb chain topologies required for NF-κB activation [35]. As described above, recent studies demonstrated that RIP1 and LUBAC can independently mediate NF-κB activation in response to TNFα stimulation and inhibition of either pathway significantly reduces the expression of certain NF-κB target genes [42]. In fact, unlike p65 or IKKβ KO mice, RIP1 KO mice survive embryonic development but die perinatally due to multiple morphological defects, and further deletion of either TNFα or TNFR1 rescues RIP1 KO mice, allowing them to grow near normally [13, 56]. Interestingly, in embryonic hepatocytes, RIP1 protein is not detectable, due most likely to posttranslational degradation [28]. This suggests that during embryonic development TNFα can activate NF-κB in RIP1-deficient hepatocytes by the TRAF2-cIAP1/2-LUBAC pathway (Fig. 1B), and that during postnatal growth RIP1 is required for efficient activation of NF-κB and inhibition of TNFα-induced cell death. TNFα is a growth factor for hematopoietic cells, and its production is increased following physical trauma and pathogenic infection [2, 4]. It can be reasoned that the function of several E3 ligases and different types of pUb chains in TNFα signaling is to enhance the sensitivity of the cell's response to a slight increase in serum TNFα levels, so as to initiate an immediate and effective host response to infection and injury.
4. Regulation of the prolonged phase of IKK activation by TRAF2 phosphorylation
In addition to TAK1, several serine/threonine kinases, including Akt1, PKCζ, TBK1 and GSK3β, have been implicated in TNFR1-mediated NF-κB activation [5, 7]. However, individual knockout of these kinases has no effect on TNFα-induced transient IKK activation or IκBα degradation, though it does result in reduced NF-κB-dependent gene expression [7, 13]. For example, targeted disruption of PKCζ leads to reduced NF-κB activation without abolishing TNFα-induced transient IKK activation [57]. TBK1 knockout likewise imparts no effect on TNFα-induced immediate IKK activation, but it significantly curtails the expression of NF-κB target genes such as TLR2 and ICAM-1[58]. This suggests that though these serine/threonine kinases do not directly phosphorylate IKK in the activation loop, they may regulate NF-κB activity upstream or downstream of IKK.
4.1. TRAF2 phosphorylation links serine/threonine kinases to IKK activation
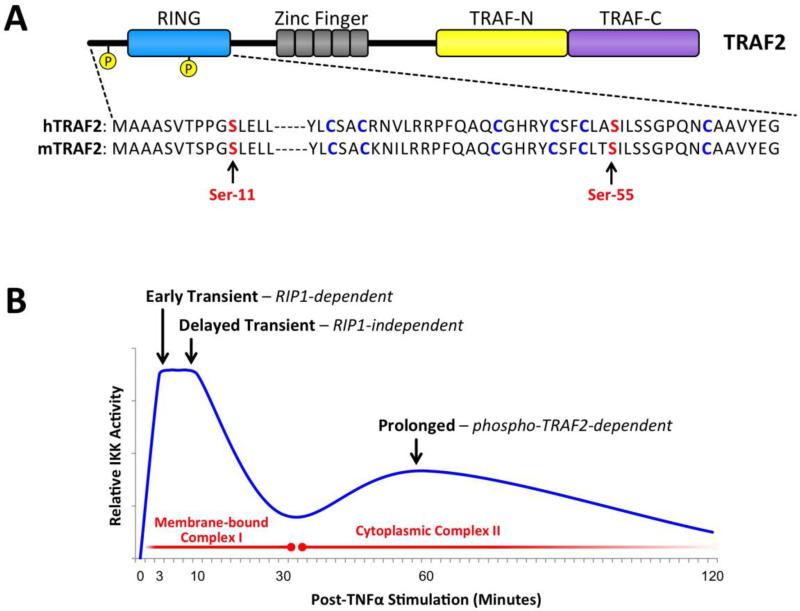
TRAF2 phosphorylation was first reported by Chaudhuri et al., though this study showed by in vivo 32P-orthophosphate labeling analysis that TRAF2 is phosphorylated at serine residue(s), it failed to identify any specific phosphosites [59]. In contrast, Li et al. reported by progressive site-directed mutagenesis complemented by functional assays that TNFα induces TRAF2 phosphorylation at the Thr-117 [60]. This Thr-117 phosphorylation appears to be mediated by PKC kinases to promote K63-linked ubiquitination of TRAF2 and subsequent recruitment of the TAK1 and IKK complexes [61]. Shortly thereafter, extensive phospho-mapping analyses in combination with in vivo 32P-orthophosphate labeling and two-dimensional separation of phospho-amino acids led to the identification of two serine residues, Ser-11 and Ser-55, phosphorylated in response to TNFα stimulation and PKC overexpression [62, 63]. These phosphorylation sites are conserved between mouse and human, and are located in the distal N-terminus and within the RING domain, respectively (Fig. 2A). Notably, these experiments were performed before and after TNFα stimulation, as well as after co-expression of TRAF2 with constitutively active forms of Akt and PKC in 293T and NIH 3T3 cells, yet they were unable to detect any phospho-threonine residues. In addition, further analyses revealed that Ser-11 is phosphorylated by IKKε, and Ser-55 is phosphorylated by PKCζ [62, 64].
Fig. 2.
TRAF2 phosphorylation regulates the prolonged phase of IKK activation. (A) Schematic illustration of the TRAF2 protein domains, phosphorylation sites, consensus C and H residues in the RING domain. (B) Upon TNFR1 ligation, the membrane bound complex-I activates the early and delayed phases of IKK through RIP1-dependent and -independent pathways. Thereafter, TRAF2 is phosphorylated at Ser-11 and Ser-55, which triggers the translocation of the TRADD/RIP1/TRAF2/cIAP1 complex from TNFR1 to the cytoplasm to form complex-IIA. This leads to activation of the prolonged phases of IKK in a TRAF2 phosphorylation-dependent manner, which is also required for the efficient expression of some NF-κB target genes.
TRAF2 phosphorylation peaks at 30 minutes after TNFα stimulation, kinetically lagging behind immediate IKK activation that occurs within 5 minutes post-stimulation [63]. Reconstitution of TRAF2 KO and TRAF2/5 DKO cells with a phospho-mutant TRAF2 in which either Ser-11 or Ser-55 is mutated to alanine (S11A or S55A) results in attenuated expression of NF-κB target genes following TNFα stimulation, but has no effect on immediate IκBα degradation, congruent with the results observed in TBK1 KO and PKCζ KO cells [62, 63]. Notably, IKK kinase assays revealed that TRAF2 phosphorylation promotes the prolonged phase of IKK activation that occurs between 45 and 75 minutes after TNFα stimulation, but not the immediate/transient phase—comprised of the previously described early and delayed phases— that occurs between 3 and 10 minutes post-stimulation [62, 63]. Mutation of both Ser-11 and Ser-55 to alanine (S11/55A) blunts the prolonged phase of IKK activation, but again has no effect on immediate IKK activation and IκBα degradation [65]. On the other hand, JNK kinase assays revealed that phosphorylation of Ser-11 and Ser-55 has the opposite effect on JNK activation. While Ser-11 phosphorylation is required for full activation of JNK, Ser-55 phosphorylation suppresses the prolonged phase of JNK activation that occurs 60 minutes after TNFα stimulation [62, 65]. This observation correlates well with the finding that genetic inactivation of Par-4, a PKCζ inhibitor, results in elevated NF-κB but decreased JNK activation in response to TNFα stimulation [66]. These data suggest that TRAF2 phosphorylation represents a new layer of regulation that links certain serine/threonine kinases to prolonged IKK activation and efficient NF-κB target gene expression in response to TNFα stimulation (Fig. 2B).
4.2. TRAF2 phosphorylation controls the fate of the TNFR1 signaling complex
Several independent studies have demonstrated that the TNFR1 complex also translocates to cholesterol- and sphingolipid-enriched lipid rafts, where it can then be internalized to promote prolonged JNK activation and production of reactive oxygen species (ROS) [11, 67, 68]. These studies demonstrated that the DISC remains associated with internalized TNFR1 in endosomal vesicles known as TNF receptosome, which does not correspond to either complex-IIA or -IIB[11]. These studies asserted that TNFR1 translocation to lipid rafts has no effect on NF-κB activation; however, others have shown that this event is required for TNFα-mediated NF-κB activation [69]. These findings suggest that the DISC can be formed in the cytoplasm as well as in the internalized TNF receptosome in a cell type- and context-dependent manner.
TRAF2 regulates almost all aspects of TNFR1 signaling [9, 35]. Several independent studies have clearly demonstrated that the intact RING domain of TRAF2 is essential for its activation of JNK signaling and translocation to lipid rafts in response to TNFR1 and CD40 ligation [34, 70, 71]. TRAF2 phosphorylation occurs within the TNFR1 complex 5-10 minutes after TNFα stimulation, and peaks at 30 minutes in the cytoplasm [63]. In TRAF2 KO cells reconstituted with TRAF2-S11/55A, complex-IIA is undetectable in the cytoplasm, indicating that TRAF2 phosphorylation mediates the translocation of the TRADD/RIP1/TRAF2/cIAP1 complex from TNFR1 to the cytoplasm [62, 63]. Although this complex-IIA is commonly regarded as an apoptosis-inducing complex, it does not cause cell death in normal cells unless protein synthesis or the NF-κB pathway is blocked [5, 14]. Thus, it is most likely that the physiological function of complex-IIA is to trigger the prolonged phase of IKK activation in a TRAF2 phosphorylation-dependent manner (Fig. 2B). Mechanistically, TRAF2 phosphorylation may directly or indirectly promote RIP1 ubiquitination within complex-IIA, retaining or recruiting more IKK. Based on these data, it is possible to hypothesize the following model: i) upon TNFα treatment, the membrane-bound complex-I induces transient IKK activation (including the early and delayed phases) independent of TRAF2 phosphorylation; ii) thereafter, TRAF2 is phosphorylated, and this phosphorylation induces the dissociation of the TRADD/RIP1/TRAF2/cIAP1/2/IKK complex from TNFR1 to form cytoplasmic complex-IIA, triggering the prolonged phase of IKK activation; and iii) in the absence of TRAF2 phosphorylation, the TRADD/RIP1/TRAF2/cIAP1/2/IKK complex remains associated with TNFR1, and thereby subsequently translocates to lipid rafts where it is internalized to activate the prolonged phase of JNK. While further studies are need to clarify the details of this model empirically, it is becoming apparent that TRAF2 phosphorylation regulates the spatial-temporal activation of the prolonged phase of IKK and JNK by controlling the subcellular translocation of the signaling complex.
4.3. TRAF2 dual phosphorylation promotes cell survival in response to oxidative stress
In addition to TNFα, a certain stress-inducing agents, such as menadione (oxidative stress inducer) and UV (DNA damage inducer), also strongly induce TRAF2 phosphorylation at both Ser-11 and Ser-55 [65]. In contrast to the increased susceptibility to TNFα-induced cell death, TRAF2 KO cells exhibit resistance to apoptosis induced by oxidative stress [72, 73]. Interestingly, expression of double mutant TRAF2-S11/55A in TRAF2 KO MEFs and MDA-MB-231 breast cancer cells significantly sensitizes them to oxidative stress-induced cell death [65]. Kinase assays revealed that this dual phosphorylation promotes cell survival by potentiating IKK activation while inhibiting prolonged JNK activation. ASK1 is a key kinase known to mediate sustained JNK activation in response to oxidative stress and is required for the onset of ROS-induced cell death [74]. An increase in intracellular ROS levels triggers the formation of a high-molecular mass complex consisting of ASK1, RIP1 and TRAF2, leading to sustained JNK activation [73, 75]. Co-immunoprecipitation assays revealed that TRAF2 phosphorylation has no effect on formation of the TRAF2/ASK1/RIP1 complex; rather, it appears that TRAF2 mediates the recruitment of IKK to this complex in a phosphorylation-dependent manner to elicit IKK activation and NF-κB target gene expression [65]. In addition, dual phosphorylation of TRAF2 also appears to suppress cell death under conditions of oxidative stress by preventing the degradation of Bcl-2 and cIAP1. The enhanced degradation of Bcl-2 in TRAF2-S11/55A-expressing cells is most likely caused by prolonged JNK activation, as it has been shown that JNK can directly phosphorylate Bcl-2 to instigate its degradation [76, 77]. On the other hand, cIAP1 degradation may be triggered by its auto-ubiquitination through K48-linkage [78]. These findings suggest that TRAF2 phosphorylation may promote cell survival under conditions of oxidative stress by directing the cIAP1 E3 ligase activity towards RIP1 ubiquitination to promote NF-κB activation and suppress prolonged JNK activation.
5. Crosstalk in TNFR-mediated signaling pathways
NF-κB activation is a key component of TNFR1 signal transduction; however, the NF-κB pathway does not exist in isolation, and there are many mechanisms that integrate its activity with other cell-signaling networks [79].
5.1. The canonical vs. non-canonical NF-κB pathway
Crosstalk exists at multiple levels between the canonical and non-canonical NF-κB pathways [79]. At the receptor level, ligation of a subset of the TNFR1 family members, such as LTβR, CD40 and BAFFR, leads to activation of both the canonical and non-canonical NF-κB pathways [6]. Of note, the expression of a dominant-negative NIK (DN-NIK) inhibits both p100 processing and IκBα degradation following ligation of these receptors [6, 80]. In line with these observations, NIK is overexpressed in a majority of primary myeloma patient samples and cell lines due to genetic or epigenetic alterations in genes (e.g. TRAF2, TRAF3, CYLD, BIR2 and BIR3) associated with the regulation of NIK protein stability [81, 82]. This accumulation of NIK results in constitutive activation of both NF-κB pathways, as siRNA-mediated knockdown of NIK in multiple myeloma cell lines with this genetic abnormality suppresses both the basal IκBα phosphorylation and abundance of nuclear p52 [81, 82]. These observations also indicate that NIK accumulation in TRAF2 KO, TRAF2/5 DKO and cIAP1/2 DKO cells can lead to constitutive activation of both NF-κB pathways. However, in these KO and DKO cells, p52 but not p65 accumulates in the nucleus [31, 32]. A possible explanation for this difference is that activation of canonical NF-κB is robust and transient, whereas activation of non-canonical NF-κB is slow and prolonged [6]. In addition, the canonical NF-κB pathway is subject to strong negative feedback regulation, which involves the recruitment of negative regulators, such as TRAF1 and deubiquitinases, to the TNFR1 complex, dephosphorylation of IKK at its T-loop, cleavage and degradation of p65 and rapid resynthesis of IκBα, among other [5, 6].
At the transcriptional level, p100 expression can be induced by canonical NF-κB to form a negative feedback loop [79, 83]. RelB expression is also dependent on the basal activity of RelA/p50, as knockout of RelA impairs RelB/p52 activation induced by LTβR [83, 84]. RelA KO mice show an impaired development of Peyer's Patch and lymph nodes, supporting the cross-regulatory mechanism between canonical and non-canonical NF-κB pathways [85]. Conversely, canonical NF-κB -mediated expression of cIAP1, cIAP2, TRAF2 and TRAF3 plays a negative role in non-canonical signaling by degrading NIK [35, 84, 86]. Moreover, p50 and p52 appear to have largely overlapping functions in dimer formation and target gene expression, although p50 and p52 display preferences for RelA and RelB, respectively [79, 83]. For instance, in p105 KO cells, RelA can form a dimer with p52; likewise, knockout of p100/p52 results in formation of the RelB/p50 dimer [79, 83]. These observations imply that stimulus-responsive dynamic regulation of the canonical and non-canonical NF-κB pathways is achieved through an extensive network of biochemical reactions that govern the accumulation and activation of upstream kinases, the synthesis and degradation of NF-κB and IκB proteins and their interactions with each other.
5.2. The JNK and NF-κB pathways
Almost all type of cytokines, growth factors and cellular stresses that induce NF-κB activation can also trigger JNK activation [47]. In general, proinflammatory cytokines such as TNFα and IL-1 induce transient JNK activation to promote cell proliferation through activation of the AP-1 family transcription factors, such as c-Jun and ATF2. In contrast, stress-inducing agents, like ROS, trigger strong and prolonged JNK activation to promote the induction of apoptosis [47, 74]. The effects of ROS can extend to JNK activation through three distinct mechanisms. First, ROS can inactivate MAPK phosphatases (MPKs) by oxidizing their catalytic cysteine residues, potentiating prolonged JNK activation [87]. Second, ROS elicits the release of JNK from its cytoplasmic inhibitor glutathione-S-transferase pi (GSTpi), facilitating redox-dependent activation [88]. Lastly, ROS directly oxidize the redox-sensitive thiols on Thioredoxin (Thx), a cytoplasmic ASK inhibitor, triggering the dissociation of Thx from ASK1, allowing its auto-phosphorylation, and subsequent activation of JNK [89].
In most cell types, strong and prolonged JNK activation induces apoptosis through several mechanisms. JNK can directly phosphorylate Bcl-2 to suppress its anti-apoptotic activity [77], while JNK-mediated phosphorylation of Itch triggers its E3 ligase activity towards cFLIP, resulting in cFLIP degradation and release of caspase-8 inhibition [90]. In addition, JNK-mediated phosphorylation of certain BCL-2 family members (e.g. Bax, Bmf and Bim) promotes their pro-apoptotic activity through as yet ill-defined mechanisms [91-94]. Moreover, prolonged JNK activation can elicit the cleavage of Bid to jBid, promoting its localization to the mitochondria and the release of pro-apoptotic Smac/DIABLO [95].
As described above, ligation of TNFR1 can lead to activation of the NF-κB, JNK and caspase-8 pathways. However, activation of the JNK and caspase-8 pathways is tightly regulated by NF-κB target gene products at several levels: TRAF2 and cIAP1/2 suppress RIP1-mediated caspase-8 activation by ubiquitinating RIP1; cFLIP limits caspase-8 activity by competing for FADD interaction; X-linked IAP (XIAP) and GADD45β inhibits prolonged JNK activation by suppressing MEKK1-mediated MKK7 activation (Fig.3) [5, 7]. As such, inhibition of the NF-κB pathway by pharmacological inhibitors or knockout of TRAF2, cIAP1/2, IKKβ, NEMO or RelA, all result in caspase-8 and prolonged JNK activation, resulting in concomitant cell death in response to TNFα stimulation [5, 7, 13]. Notably, NF-κB-mediated inhibition of JNK provides a tumor-suppressing function in some cell types, including epidermal cells. For instance, keratinocyte-specific ablation of IKKβ, NEMO, TAK1 and RelA, or expression of a degradation-resistant form of IκBα, all lead to skin inflammation and hyperplasia in puppies after birth through the JNK/c-Jun pathway [5, 7, 13]. Thus, prolonged JNK activation can either be pro-apoptotic or pro-proliferative in a cell type- and context-dependent manner.
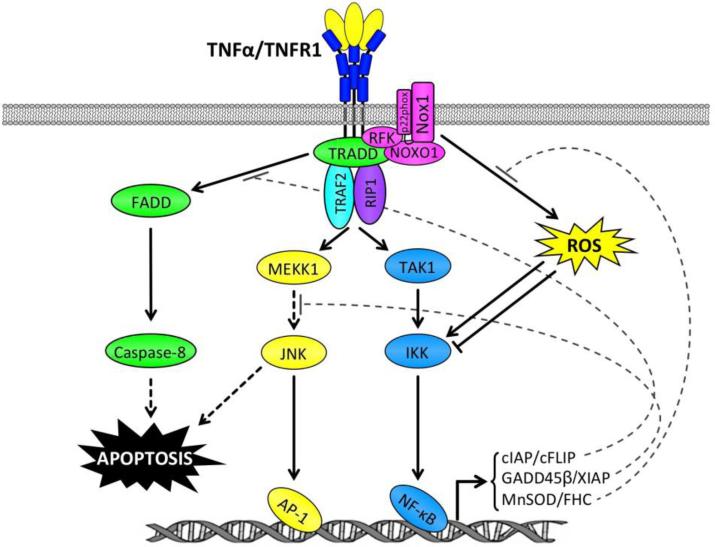
Fig.3.
Crosstalk in TNFR1 signaling. Ligation of TNFR1 leads to activation of the pro-inflammatory NF-κB and JNK, and pro-apoptotic caspase-8 and ROS pathways. However, in normal cells, NF-κB target gene products suppress pro-apoptotic pathways. For instance, cIAP1/2 and cFLIP inhibit caspase-8 activation, XIAP and GADD45β suppress prolonged JNK activation, and FHC and Mn-SOD inhibit ROS accumulation. Therefore, unless NF-κB pathway is blocked, TNFα is insufficient to induce cell death.
5.3. The NF-κB pathway and reactive oxygen species
TNFR1 ligation can also instigate ROS production through mitochondria and NADPH oxidases [96, 97]. While prolonged JNK activation can lead to increased ROS production in mitochondria, the NOX family of ROS-generating NADPH oxidases utilize NADPH to reduce oxygen to superoxide [98]. Following TNF stimulation, the TNFR1/TRADD/RIP1 complex recruits Nox1 or Nox2 through riboflavin kinase (RFK) and flavin adenine dinucleotide (FAD) to potentiate ROS production[97]. Intracellular ROS can react non-specifically with biomolecules, instigating protein oxidation, DNA damage, lipid peroxidation. However, in a physiological condition, TNFα-induced and NF-κB-dependent gene expression serves to actively suppresses ROS accumulation in most cell types [96]. Several antioxidant proteins are NF-κB target genes, including Manganese Superoxide Dismutase (MnSOD) and Ferritin Heavy Chain (FHC). MnSOD is a mitochondrial enzyme that catalyzes the dismutation of to oxygen and hydrogen peroxide (H2O2). H2O2 is then neutralized into water by catalases and peroxidases[96]. Mice lacking Mn-SOD die perinatally due to massive oxidative stress [99]. FHC protects the cell from oxidative damage by preventing the iron-dependent generation of •OH radicals from H2O2, known as the Fenton Reaction [98].
ROS can exert a dual function on TNFα-induced NF-κB activation through direct or indirect post-translational modifications of IKK, IκBα and NF-κB dimers. For instance, H2O2-mediated formation of disulfide bridges between Cys-54 and -347 in NEMO can promote its homo-dimerization to potentiate NF-κB activation [100], while ROS-induced and PKAc-mediated phosphorylation of RelA on Ser-276 promotes its interaction with transcriptional coactivators, CBP/p300 and P-TEFb, promoting the expression of a subset of NF-κB target genes [101, 102]. Conversely, S-glutathionation of IKKβ at Cys-179 inactivates its kinase activity [103], and the S-glutathionation of IκBα at Cys-189 is sufficient to attenuate its conventional phosphorylation at Ser-32/36 by IKK [104]. In addition, H2O2 can trigger casein kinase II-mediated phosphorylation of IκBα on Tyr-42, facilitating its association with the SH2 domain of PI3K and release of NF-κB, independent of IκBα degradation [105-107]. Moreover, the direct oxidation of p50 at Cys-62 within its RHD inhibits its ability to bind DNA. Collectively, these findings suggest that while TNFα-induced NF-κB activation suppresses ROS accumulation and plays an integral role in maintaining cellular homeostasis, NF-κB activation can also be positively or negatively regulated by ROS, depending on the extend of ROS accumulation and type of cells (Fi. 3).
6. Conclusions and future perspectives
Excessive TNFα production is the major cause of several immune-mediated inflammatory diseases. Three functional antagonists of TNFα are currently approved for the treatment of RA and IBD [3, 4]. Although the development of these therapies has undoubtedly improved the quality of life for many patients, numerous issues have come to light with the continued studies on the efficacy and long-term effects of TNF blockade, which include initial or acquired desensitization, an increased risk of infection, and the development of drug-induced autoimmune disorders [108-117]. In addition, increased TNFα production is not only an important risk factor in tumor development and progression, but also an integral mediator of cancer-associated chronic inflammation [3, 4]. However, several studies have reported that TNFα antagonism can lead to an increased risk of malignancy as a result of diminished TNFα-mediated cancer immunosurveillance [110, 118].
NF-κB is ubiquitously expressed and can be activated by almost all types of stimuli that cause inflammation and cellular stress [5]. NF-κB activation induces the expression of proinflammatory cytokines, chemokines, angiogenic factors, and adhesion molecules, and these factors in turn further propagate and amplify the inflammatory response, thereby promoting tumorigenesis, progression and metastasis [35, 119]. In addition, NF-κB activation induces the expression of cIAP1/2, Bcl-XL, survivin and cFLIP, and these pro-survival proteins then confer upon cancer cells the ability to survive and adapt to stressful tumor microenvironments characterized by hypoxia, hypoglycemia, and the increased production of free radicals – conditions that potentiate chronic inflammation and oxidative stress [119, 120]. In line with these findings, numerous studies have demonstrated that NF-κB is constitutively activated in various types of human cancers. These findings also made NF-κB an ideal target for pharmacological inhibition. However, although over 700 different inhibitors of NF-κB have been reported, none of them reached an approvable level for human use, due to severe immune side effects [121]. It is now becoming clear that NF-κB activation induced by PAMP is not only essential for the physiological immune and inflammatory responses, but also for epithelial cell survival, and proliferation [46, 122]. As a consequence, inhibition of the core elements of the NF-κB pathway (e.g. IKK, IκBα and p65) could result in disruption of epithelial homeostasis, and subsequently in infection and inflammation [122]. These observations suggest that both hyper- and hypo-activation of NF-κB can lead to inflammation; therefore, the major challenges facing the researchers is to develop NF-κB inhibitors aimed at disease-specific signaling pathways, thereby avoiding the risk of undesired side effects. As such, further studies also should apply live-imaging and proteomic approaches to study the dynamic interactions of Ub and Ub-binding proteins, the molecular events that determine the topology of ubiquitination, regulatory crosstalk between the NF-κB and other pathways, and the mechanisms by which serine/threonine kinases regulate the basal and inducible IKK activity.
Acknowledgements
We would like to apologize to all scientists whose contributions to the field are not cited owing to space restriction. We thank the members of Habelhah laboratory for critical reading of the manuscript. Support by NCI grant CA138475 is gratefully acknowledged.
Abbreviations
- ASK1
apoptosis signal-regulating kinase 1
- cFLIP
cellular caspase-8 (FLICE)-like inhibitory protein
- cIAP1
cellular inhibitor of apoptosis 1
- DICS
death-inducing signaling complex
- DKO
double knockout
- FADD
Fas-associated death domain
- IκB
inhibitor of κB
- IKK
IκB kinase
- IL-1
interleukin-1
- IL-6
interleukin-6
- JNK
c-Jun N-terminal kinase
- KO
knockout
- LUBAC
linear ubiquitin chain assembly complex
- MEFs
mouse embryonic fibroblasts
- MEKK1
mitogen-activated protein kinase kinase kinase 1
- NF-κB
nuclear factor κB
- NIK
NF-κB, inducing kinase
- RA
rheumatoid arthritis
- RIP1
receptor interacting protein 1
- ROS
reactive oxygen species
- SM
SMAC mimetic
- S1P
sphingosine-1-phosphate
- TAK1
transforming growth factor β-activated kinase 1
- TAB1
TAK1 binding protein 1
- TBK1
TANK-binding kinase 1
- TNF
tumor necrosis factor
- TNFR
TNF receptor
- TRADD
TNF receptor associated death domain
- TRAF
TNF receptor associated factor
- UBD
ubiquitin-binding domain
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Conflict of interest
The authors declared no potential conflict of interest with respect to the authorship and publication of this article.
References
- 1.Bodmer JL, Schneider P, Tschopp J. Trends in Biochemical Sciences. 2002;27:19–26. doi: 10.1016/s0968-0004(01)01995-8. [DOI] [PubMed] [Google Scholar]
- 2.Hehlgans T, Pfeffer K. Immunology. 2005;115:1–20. doi: 10.1111/j.1365-2567.2005.02143.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Lin J, Ziring D, Desai S, Kim S, Wong M, Korin Y, Braun J, Reed E, Gjertson D, Singh RR. Clinical Immunology. 2008;126:13–30. doi: 10.1016/j.clim.2007.08.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Valesini G, Iannuccelli C, Marocchi E, Pascoli L, Scalzi V, Di Franco M. Autoimmunity Reviews. 2007;7:35–41. doi: 10.1016/j.autrev.2007.03.003. [DOI] [PubMed] [Google Scholar]
- 5.Hayden MS, Ghosh S. Cell. 2008;132:344–362. doi: 10.1016/j.cell.2008.01.020. [DOI] [PubMed] [Google Scholar]
- 6.Bonizzi G, Karin M. Trends in Immunology. 2004;25:280–288. doi: 10.1016/j.it.2004.03.008. [DOI] [PubMed] [Google Scholar]
- 7.Vallabhapurapu S, Karin M. Annual Review of Immunology. 2009;27:693–733. doi: 10.1146/annurev.immunol.021908.132641. [DOI] [PubMed] [Google Scholar]
- 8.Wallach D, Varfolomeev EE, Malinin NL, Goltsev YV, Kovalenko AV, Boldin MP. Annual Review of Immunology. 1999;17:331–367. doi: 10.1146/annurev.immunol.17.1.331. [DOI] [PubMed] [Google Scholar]
- 9.Wajant H, Henkler F, Scheurich P. Cellular Signalling. 2001;13:389–400. doi: 10.1016/s0898-6568(01)00160-7. [DOI] [PubMed] [Google Scholar]
- 10.Hyer ML, Samuel T, Reed JC. Clinical Cancer Research. 2006;12:5929–5931. doi: 10.1158/1078-0432.CCR-06-2098. [DOI] [PubMed] [Google Scholar]
- 11.Schutze S, Tchikov V, Schneider-Brachert W. Nature Reviews. Molecular Cell Biology. 2008;9:655–662. doi: 10.1038/nrm2430. [DOI] [PubMed] [Google Scholar]
- 12.Hehlgans T, Mannel DN. Biological Chemistry. 2002;383:1581–1585. doi: 10.1515/BC.2002.178. [DOI] [PubMed] [Google Scholar]
- 13.Gerondakis S, Grumont R, Gugasyan R, Wong L, Isomura I, Ho W, Banerjee A. Oncogene. 2006;25:6781–6799. doi: 10.1038/sj.onc.1209944. [DOI] [PubMed] [Google Scholar]
- 14.Micheau O, Tschopp J. Cell. 2003;114:181–190. doi: 10.1016/s0092-8674(03)00521-x. [DOI] [PubMed] [Google Scholar]
- 15.Festjens N, Vanden Berghe T, Cornelis S, Vandenabeele P. Cell Death and Differentiation. 2007;14:400–410. doi: 10.1038/sj.cdd.4402085. [DOI] [PubMed] [Google Scholar]
- 16.O'Donnell MA, Legarda-Addison D, Skountzos P, Yeh WC, Ting AT. Current Biology. 2007;17:418–424. doi: 10.1016/j.cub.2007.01.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Vandenabeele P, Declercq W, Van Herreweghe F, Vanden Berghe T. Sci Signal. 2010;3:re4. doi: 10.1126/scisignal.3115re4. [DOI] [PubMed] [Google Scholar]
- 18.Chen ZJ, Sun LJ. Molecular Cell. 2009;33:275–286. doi: 10.1016/j.molcel.2009.01.014. [DOI] [PubMed] [Google Scholar]
- 19.Wang L, Du F, Wang X. Cell. 2008;133:693–703. doi: 10.1016/j.cell.2008.03.036. [DOI] [PubMed] [Google Scholar]
- 20.Tenev T, Bianchi K, Darding M, Broemer M, Langlais C, Wallberg F, Zachariou A, Lopez J, MacFarlane M, Cain K, Meier P. Molecular Cell. 2011;43:432–448. doi: 10.1016/j.molcel.2011.06.006. [DOI] [PubMed] [Google Scholar]
- 21.Feoktistova M, Geserick P, Kellert B, Dimitrova DP, Langlais C, Hupe M, Cain K, MacFarlane M, Hacker G, Leverkus M. Molecular Cell. 2011;43:449–463. doi: 10.1016/j.molcel.2011.06.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Wu CJ, Conze DB, Li T, Srinivasula SM, Ashwell JD. Nature Cell Biology. 2006;8:398–406. doi: 10.1038/ncb1384. [DOI] [PubMed] [Google Scholar]
- 23.Ea CK, Deng L, Xia ZP, Pineda G, Chen ZJ. Molecular Cell. 2006;22:245–257. doi: 10.1016/j.molcel.2006.03.026. [DOI] [PubMed] [Google Scholar]
- 24.Mahoney DJ, Cheung HH, Mrad RL, Plenchette S, Simard C, Enwere E, Arora V, Mak TW, Lacasse EC, Waring J, Korneluk RG. Proceedings of the National Academy of Sciences of the United States of America. 2008;105:11778–11783. doi: 10.1073/pnas.0711122105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Varfolomeev E, Goncharov T, Fedorova AV, Dynek JN, Zobel K, Deshayes K, Fairbrother WJ, Vucic D. Journal of Biological Chemistry. 2008;283:24295–24299. doi: 10.1074/jbc.C800128200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Haas TL, Emmerich CH, Gerlach B, Schmukle AC, Cordier SM, Rieser E, Feltham R, Vince J, Warnken U, Wenger T, Koschny R, Komander D, Silke J, Walczak H. Molecular Cell. 2009;36:831–844. doi: 10.1016/j.molcel.2009.10.013. [DOI] [PubMed] [Google Scholar]
- 27.Tokunaga F, Sakata S, Saeki Y, Satomi Y, Kirisako T, Kamei K, Nakagawa T, Kato M, Murata S, Yamaoka S, Yamamoto M, Akira S, Takao T, Tanaka K, Iwai K. Nature Cell Biology. 2009;11:123–132. doi: 10.1038/ncb1821. [DOI] [PubMed] [Google Scholar]
- 28.Wong WW, Gentle IE, Nachbur U, Anderton H, Vaux DL, Silke J. Cell Death and Differentiation. 2010;17:482–487. doi: 10.1038/cdd.2009.178. [DOI] [PubMed] [Google Scholar]
- 29.Yamamoto M, Okamoto T, Takeda K, Sato S, Sanjo H, Uematsu S, Saitoh T, Yamamoto N, Sakurai H, Ishii KJ, Yamaoka S, Kawai T, Matsuura Y, Takeuchi O, Akira S. Nature Immunology. 2006;7:962–970. doi: 10.1038/ni1367. [DOI] [PubMed] [Google Scholar]
- 30.Yin Q, Lamothe B, Darnay BG, Wu H. Biochemistry. 2009;48:10558–10567. doi: 10.1021/bi901462e. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Zhang L, Blackwell K, Shi Z, Habelhah H. Journal of Molecular Biology. 2010;396:528–539. doi: 10.1016/j.jmb.2010.01.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Zhang L, Blackwell K, Thomas GS, Sun S, Yeh WC, Habelhah H. Journal of Molecular Biology. 2009;389:495–510. doi: 10.1016/j.jmb.2009.04.054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Alvarez SE, Harikumar KB, Hait NC, Allegood J, Strub GM, Kim EY, Maceyka M, Jiang H, Luo C, Kordula T, Milstien S, Spiegel S. Nature. 2010;465:1084–1088. doi: 10.1038/nature09128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Habelhah H, Takahashi S, Cho SG, Kadoya T, Watanabe T, Ronai Z. EMBO Journal. 2004;23:322–332. doi: 10.1038/sj.emboj.7600044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Habelhah H. Genes Cancer. 2010;1:735–747. doi: 10.1177/1947601910382900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Shembade N, Ma A, Harhaj EW. Science. 2010;327:1135–1139. doi: 10.1126/science.1182364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Xu M, Skaug B, Zeng W, Chen ZJ. Molecular Cell. 2009;36:302–314. doi: 10.1016/j.molcel.2009.10.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Dynek JN, Goncharov T, Dueber EC, Fedorova AV, Izrael-Tomasevic A, Phu L, Helgason E, Fairbrother WJ, Deshayes K, Kirkpatrick DS, Vucic D. EMBO Journal. 2010;29:4198–4209. doi: 10.1038/emboj.2010.300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Tada K, Okazaki T, Sakon S, Kobarai T, Kurosawa K, Yamaoka S, Hashimoto H, Mak TW, Yagita H, Okumura K, Yeh WC, Nakano H. Journal of Biological Chemistry. 2001;276:36530–36534. doi: 10.1074/jbc.M104837200. [DOI] [PubMed] [Google Scholar]
- 40.Vince JE, Pantaki D, Feltham R, Mace PD, Cordier SM, Schmukle AC, Davidson AJ, Callus BA, Wong WW, Gentle IE, Carter H, Lee EF, Walczak H, Day CL, Vaux DL, Silke J. Journal of Biological Chemistry. 2009;284:35906–35915. doi: 10.1074/jbc.M109.072256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Feltham R, Moulin M, Vince JE, Mace PD, Wong WW, Anderton H, Day CL, Vaux DL, Silke J. Journal of Biological Chemistry. 2010;285:17525–17536. doi: 10.1074/jbc.M109.087635. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Blackwell K, Zhang L, Workman LM, Ting AT, Iwai K, Habelhah H. Molecular and Cellular Biology. 2013 doi: 10.1128/MCB.01416-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Zarnegar B, Yamazaki S, He JQ, Cheng G. Proceedings of the National Academy of Sciences of the United States of America. 2008;105:3503–3508. doi: 10.1073/pnas.0707959105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Hoffmann A, Levchenko A, Scott ML, Baltimore D. Science. 2002;298:1241–1245. doi: 10.1126/science.1071914. [DOI] [PubMed] [Google Scholar]
- 45.Werner SL, Barken D, Hoffmann A. Science. 2005;309:1857–1861. doi: 10.1126/science.1113319. [DOI] [PubMed] [Google Scholar]
- 46.Smale ST. Nature Immunol. 2011;12:689–694. doi: 10.1038/ni.2070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Davis RJ. Cell. 2000;103:239–252. doi: 10.1016/s0092-8674(00)00116-1. [DOI] [PubMed] [Google Scholar]
- 48.Hoffmann A, Leung TH, Baltimore D. EMBO Journal. 2003;22:5530–5539. doi: 10.1093/emboj/cdg534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Nguyen LT, Duncan GS, Mirtsos C, Ng M, Speiser DE, Shahinian A, Marino MW, Mak TW, Ohashi PS, Yeh WC. Immunity. 1999;11:379–389. doi: 10.1016/s1074-7613(00)80113-2. [DOI] [PubMed] [Google Scholar]
- 50.Ventura JJ, Kennedy NJ, Lamb JA, Flavell RA, Davis RJ. Molecular and Cellular Biology. 2003;23:2871–2882. doi: 10.1128/MCB.23.8.2871-2882.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Okazaki T, Sakon S, Sasazuki T, Sakurai H, Doi T, Yagita H, Okumura K, Nakano H. Biochemical and Biophysical Research Communications. 2003;300:807–812. doi: 10.1016/s0006-291x(02)02932-7. [DOI] [PubMed] [Google Scholar]
- 52.Saccani S, Pantano S, Natoli G. Journal of Experimental Medicine. 2001;193:1351–1359. doi: 10.1084/jem.193.12.1351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Gentle IE, Wong WW, Evans JM, Bankovacki A, Cook WD, Khan NR, Nachbur U, Rickard J, Anderton H, Moulin M, Lluis JM, Moujalled DM, Silke J, Vaux DL. Journal of Biological Chemistry. 2011;286:13282–13291. doi: 10.1074/jbc.M110.216226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Gerlach B, Cordier SM, Schmukle AC, Emmerich CH, Rieser E, Haas TL, Webb AI, Rickard JA, Anderton H, Wong WW, Nachbur U, Gangoda L, Warnken U, Purcell AW, Silke J, Walczak H. Nature. 2011;471:591–596. doi: 10.1038/nature09816. [DOI] [PubMed] [Google Scholar]
- 55.Bertrand MJ, Lippens S, Staes A, Gilbert B, Roelandt R, De Medts J, Gevaert K, Declercq W, Vandenabeele P. PLoS One. 2011;6:e22356. doi: 10.1371/journal.pone.0022356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Kelliher MA, Grimm S, Ishida Y, Kuo F, Stanger BZ, Leder P. Immunity. 1998;8:297–303. doi: 10.1016/s1074-7613(00)80535-x. [DOI] [PubMed] [Google Scholar]
- 57.Leitges M, Sanz L, Martin P, Duran A, Braun U, Garcia JF, Camacho F, Diaz-Meco MT, Rennert PD, Moscat J. Molecular Cell. 2001;8:771–780. doi: 10.1016/s1097-2765(01)00361-6. [DOI] [PubMed] [Google Scholar]
- 58.Bonnard M, Mirtsos C, Suzuki S, Graham K, Huang J, Ng M, Itie A, Wakeham A, Shahinian A, Henzel WJ, Elia AJ, Shillinglaw W, Mak TW, Cao Z, Yeh WC. EMBO Journal. 2000;19:4976–4985. doi: 10.1093/emboj/19.18.4976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Chaudhuri A, Orme S, Vo T, Wang W, Cherayil BJ. Biochemical and Biophysical Research Communications. 1999;256:620–625. doi: 10.1006/bbrc.1999.0385. [DOI] [PubMed] [Google Scholar]
- 60.Li S, Wang L, Berman MA, Zhang Y, Dorf ME. Molecular Cell. 2006;24:497–509. doi: 10.1016/j.molcel.2006.10.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Li S, Wang L, Dorf ME. Molecular Cell. 2009;33:30–42. doi: 10.1016/j.molcel.2008.11.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Thomas GS, Zhang L, Blackwell K, Habelhah H. Cancer Research. 2009;69:3665–3672. doi: 10.1158/0008-5472.CAN-08-4867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Blackwell K, Zhang L, Thomas GS, Sun S, Nakano H, Habelhah H. Molecular and Cellular Biology. 2009;29:303–314. doi: 10.1128/MCB.00699-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Shen RR, Zhou AY, Kim E, Lim E, Habelhah H, Hahn WC. Molecular and Cellular Biology. 2012;32:4756–4768. doi: 10.1128/MCB.00468-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Zhang L, Blackwell K, Altaeva A, Shi Z, Habelhah H. Molecular Biology of the Cell. 2011;22:128–140. doi: 10.1091/mbc.E10-06-0556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Moscat J, Rennert P, Diaz-Meco MT. Cell Death and Differentiation. 2006;13:702–711. doi: 10.1038/sj.cdd.4401823. [DOI] [PubMed] [Google Scholar]
- 67.Schneider-Brachert W, Tchikov V, Neumeyer J, Jakob M, Winoto-Morbach S, Held-Feindt J, Heinrich M, Merkel O, Ehrenschwender M, Adam D, Mentlein R, Kabelitz D, Schutze S. Immunity. 2004;21:415–428. doi: 10.1016/j.immuni.2004.08.017. [DOI] [PubMed] [Google Scholar]
- 68.Woo CH, Kim TH, Choi JA, Ryu HC, Lee JE, You HJ, Bae YS, Kim JH. Biochemical and Biophysical Research Communications. 2006;351:972–978. doi: 10.1016/j.bbrc.2006.10.154. [DOI] [PubMed] [Google Scholar]
- 69.Legler DF, Micheau O, Doucey MA, Tschopp J, Bron C. Immunity. 2003;18:655–664. doi: 10.1016/s1074-7613(03)00092-x. [DOI] [PubMed] [Google Scholar]
- 70.Hostager BS, Catlett IM, Bishop GA. Journal of Biological Chemistry. 2000;275:15392–15398. doi: 10.1074/jbc.M909520199. [DOI] [PubMed] [Google Scholar]
- 71.Arron JR, Pewzner-Jung Y, Walsh MC, Kobayashi T, Choi Y. Journal of Experimental Medicine. 2002;196:923–934. doi: 10.1084/jem.20020774. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Lin Y, Choksi S, Shen HM, Yang QF, Hur GM, Kim YS, Tran JH, Nedospasov SA, Liu ZG. Journal of Biological Chemistry. 2004;279:10822–10828. doi: 10.1074/jbc.M313141200. [DOI] [PubMed] [Google Scholar]
- 73.Noguchi T, Takeda K, Matsuzawa A, Saegusa K, Nakano H, Gohda J, Inoue J, Ichijo H. Journal of Biological Chemistry. 2005;280:37033–37040. doi: 10.1074/jbc.M506771200. [DOI] [PubMed] [Google Scholar]
- 74.Tobiume K, Matsuzawa A, Takahashi T, Nishitoh H, Morita K, Takeda K, Minowa O, Miyazono K, Noda T, Ichijo H. EMBO Report. 2001;2:222–228. doi: 10.1093/embo-reports/kve046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Shen HM, Lin Y, Choksi S, Tran J, Jin T, Chang L, Karin M, Zhang J, Liu ZG. Molecular and Cellular Biology. 2004;24:5914–5922. doi: 10.1128/MCB.24.13.5914-5922.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Fan M, Goodwin M, Vu T, Brantley-Finley C, Gaarde WA, Chambers TC. Journal of Biological Chemistry. 2000;275:29980–29985. doi: 10.1074/jbc.M003776200. [DOI] [PubMed] [Google Scholar]
- 77.Yamamoto K, Ichijo H, Korsmeyer SJ. Molecular and Cellular Biology. 1999;19:8469–8478. doi: 10.1128/mcb.19.12.8469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Csomos RA, Brady GF, Duckett CS. Journal of Biological Chemistry. 2009;284:20531–20539. doi: 10.1074/jbc.M109.029983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Oeckinghaus A, Hayden MS, Ghosh S. Nature Immunol. 2011;12:695–708. doi: 10.1038/ni.2065. [DOI] [PubMed] [Google Scholar]
- 80.Ramakrishnan P, Wang W, Wallach D. Immunity. 2004;21:477–489. doi: 10.1016/j.immuni.2004.08.009. [DOI] [PubMed] [Google Scholar]
- 81.Annunziata CM, Davis RE, Demchenko Y, Bellamy W, Gabrea A, Zhan F, Lenz G, Hanamura I, Wright G, Xiao W, Dave S, Hurt EM, Tan B, Zhao H, Stephens O, Santra M, Williams DR, Dang L, Barlogie B, Shaughnessy JD, Jr., Kuehl WM, Staudt LM. Cancer Cell. 2007;12:115–130. doi: 10.1016/j.ccr.2007.07.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Keats JJ, Fonseca R, Chesi M, Schop R, Baker A, Chng WJ, Van Wier S, Tiedemann R, Shi CX, Sebag M, Braggio E, Henry T, Zhu YX, Fogle H, Price-Troska T, Ahmann G, Mancini C, Brents LA, Kumar S, Greipp P, Dispenzieri A, Bryant B, Mulligan G, Bruhn L, Barrett M, Valdez R, Trent J, Stewart AK, Carpten J, Bergsagel PL. Cancer Cell. 2007;12:131–144. doi: 10.1016/j.ccr.2007.07.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 83.Shih VF, Tsui R, Caldwell A, Hoffmann A. Cell Research. 2011;21:86–102. doi: 10.1038/cr.2010.161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Basak S, Shih VF, Hoffmann A. Molecular and Cellular Biology. 2008;28:3139–3150. doi: 10.1128/MCB.01469-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 85.Alcamo E, Hacohen N, Schulte LC, Rennert PD, Hynes RO, Baltimore D. Journal of Experimental Medicine. 2002;195:233–244. doi: 10.1084/jem.20011885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 86.Wang CY, Mayo MW, Korneluk RG, Goeddel DV, Baldwin AS., Jr. Science. 1998;281:1680–1683. doi: 10.1126/science.281.5383.1680. [DOI] [PubMed] [Google Scholar]
- 87.Kamata H, Honda S, Maeda S, Chang L, Hirata H, Karin M. Cell. 2005;120:649–661. doi: 10.1016/j.cell.2004.12.041. [DOI] [PubMed] [Google Scholar]
- 88.Adler V, Yin Z, Fuchs SY, Benezra M, Rosario L, Tew KD, Pincus MR, Sardana M, Henderson CJ, Wolf CR, Davis RJ, Ronai Z. EMBO Journal. 1999;18:1321–1334. doi: 10.1093/emboj/18.5.1321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Saitoh M, Nishitoh H, Fujii M, Takeda K, Tobiume K, Sawada Y, Kawabata M, Miyazono K, Ichijo H. EMBO Journal. 1998;17:2596–2606. doi: 10.1093/emboj/17.9.2596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 90.Chang L, Kamata H, Solinas G, Luo JL, Maeda S, Venuprasad K, Liu YC, Karin M. Cell. 2006;124:601–613. doi: 10.1016/j.cell.2006.01.021. [DOI] [PubMed] [Google Scholar]
- 91.Kim BJ, Ryu SW, Song BJ. Journal of Biological Chemistry. 2006;281:21256–21265. doi: 10.1074/jbc.M510644200. [DOI] [PubMed] [Google Scholar]
- 92.Lei K, Davis RJ. Proceedings of the National Academy of Sciences of the United States of America. 2003;100:2432–2437. doi: 10.1073/pnas.0438011100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Putcha GV, Le S, Frank S, Besirli CG, Clark K, Chu B, Alix S, Youle RJ, LaMarche A, Maroney AC, Johnson EM., Jr. Neuron. 2003;38:899–914. doi: 10.1016/s0896-6273(03)00355-6. [DOI] [PubMed] [Google Scholar]
- 94.Becker EB, Howell J, Kodama Y, Barker PA, Bonni A. Journal of Neuroscience. 2004;24:8762–8770. doi: 10.1523/JNEUROSCI.2953-04.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 95.Deng Y, Ren X, Yang L, Lin Y, Wu X. Cell. 2003;115:61–70. doi: 10.1016/s0092-8674(03)00757-8. [DOI] [PubMed] [Google Scholar]
- 96.Shen HM, Pervaiz S. FASEB Journal. 2006;20:1589–1598. doi: 10.1096/fj.05-5603rev. [DOI] [PubMed] [Google Scholar]
- 97.Yazdanpanah B, Wiegmann K, Tchikov V, Krut O, Pongratz C, Schramm M, Kleinridders A, Wunderlich T, Kashkar H, Utermohlen O, Bruning JC, Schutze S, Kronke M. Nature. 2009;460:1159–1163. doi: 10.1038/nature08206. [DOI] [PubMed] [Google Scholar]
- 98.Lambeth JD. Nature Review Immunology. 2004;4:181–189. doi: 10.1038/nri1312. [DOI] [PubMed] [Google Scholar]
- 99.Li Y, Huang TT, Carlson EJ, Melov S, Ursell PC, Olson JL, Noble LJ, Yoshimura MP, Berger C, Chan PH, Wallace DC, Epstein CJ. Nature Genetics. 1995;11:376–381. doi: 10.1038/ng1295-376. [DOI] [PubMed] [Google Scholar]
- 100.Herscovitch M, Comb W, Ennis T, Coleman K, Yong S, Armstead B, Kalaitzidis D, Chandani S, Gilmore TD. Biochemical and Biophysical Research Communications. 2008;367:103–108. doi: 10.1016/j.bbrc.2007.12.123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Jamaluddin M, Wang S, Boldogh I, Tian B, Brasier AR. Cellular Signalling. 2007;19:1419–1433. doi: 10.1016/j.cellsig.2007.01.020. [DOI] [PubMed] [Google Scholar]
- 102.Zhong H, Voll RE, Ghosh S. Molecular Cell. 1998;1:661–671. doi: 10.1016/s1097-2765(00)80066-0. [DOI] [PubMed] [Google Scholar]
- 103.Reynaert NL, van der Vliet A, Guala AS, McGovern T, Hristova M, Pantano C, Heintz NH, Heim J, Ho YS, Matthews DE, Wouters EF, Janssen-Heininger YM. Proceedings of the National Academy of Sciences of the United States of America. 2006;103:13086–13091. doi: 10.1073/pnas.0603290103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Kil IS, Kim SY, Park JW. Biochemical and Biophysical Research Communications. 2008;373:169–173. doi: 10.1016/j.bbrc.2008.06.007. [DOI] [PubMed] [Google Scholar]
- 105.Takada Y, Mukhopadhyay A, Kundu GC, Mahabeleshwar GH, Singh S, Aggarwal BB. Journal of Biological Chemistry. 2003;278:24233–24241. doi: 10.1074/jbc.M212389200. [DOI] [PubMed] [Google Scholar]
- 106.Canty TG, Jr., Boyle EM, Jr., Farr A, Morgan EN, Verrier ED, Pohlman TH. Circulation. 1999;100:II361–364. doi: 10.1161/01.cir.100.suppl_2.ii-361. [DOI] [PubMed] [Google Scholar]
- 107.Beraud C, Henzel WJ, Baeuerle PA. Proceedings of the National Academy of Sciences of the United States of America. 1999;96:429–434. doi: 10.1073/pnas.96.2.429. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 108.Sidiropoulos PI, Boumpas DT. Annals of the Rheumatic Diseases. 2006;65:701–703. doi: 10.1136/ard.2005.049890. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 109.Gardam MA, Keystone EC, Menzies R, Manners S, Skamene E, Long R, Vinh DC. The Lancet Infectious Diseases. 2003;3:148–155. doi: 10.1016/s1473-3099(03)00545-0. [DOI] [PubMed] [Google Scholar]
- 110.Bongartz T, Sutton AJ, Sweeting MJ, Buchan I, Matteson EL, Montori V. JAMA: The journal of American Medical Association. 2006;295:2275–2285. doi: 10.1001/jama.295.19.2275. [DOI] [PubMed] [Google Scholar]
- 111.Day R. Lancet. 2002;359:540–541. doi: 10.1016/S0140-6736(02)07718-8. [DOI] [PubMed] [Google Scholar]
- 112.Hyrich KL, Silman AJ, Watson KD, Symmons DP. Annals of the Rheumatic Diseases. 2004;63:1538–1543. doi: 10.1136/ard.2004.024737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 113.Ramos-Casals M, Brito-Zeron P, Munoz S, Soria N, Galiana D, Bertolaccini L, Cuadrado MJ, Khamashta MA. Medicine. 2007;86:242–251. doi: 10.1097/MD.0b013e3181441a68. [DOI] [PubMed] [Google Scholar]
- 114.Jarrett SJ, Cunnane G, Conaghan PG, Bingham SJ, Buch MH, Quinn MA, Emery P. Journal of Rheumatology. 2003;30:2287–2291. [PubMed] [Google Scholar]
- 115.Tack CJ, Kleijwegt FS, Van Riel PL, Roep BO. Diabtologia. 2009;52:1442–1444. doi: 10.1007/s00125-009-1381-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 116.Sicotte NL, Voskuhl RR. Neurology. 2001;57:1885–1888. doi: 10.1212/wnl.57.10.1885. [DOI] [PubMed] [Google Scholar]
- 117.Sari I, Akar S, Birlik M, Sis B, Onen F, Akkoc N. Journal of Rheumatology. 2006;33:1411–1414. [PubMed] [Google Scholar]
- 118.Diak P, Siegel J, La Grenade L, Choi L, Lemery S, McMahon A. Arthritis and Rheumatism. 2010;62:2517–2524. doi: 10.1002/art.27511. [DOI] [PubMed] [Google Scholar]
- 119.Luo JL, Kamata H, Karin M. Journal of Clinical Investigation. 2005;115:2625–2632. doi: 10.1172/JCI26322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 120.Ben-Neriah Y, Karin M. Nature Immunology. 2011;12:715–723. doi: 10.1038/ni.2060. [DOI] [PubMed] [Google Scholar]
- 121.Gupta SC, Sundaram C, Reuter S, Aggarwal BB. Biochimica et Biophysica Acta. 2010;1799:775–787. doi: 10.1016/j.bbagrm.2010.05.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 122.Pasparakis M. Nature Reviews. Immunology. 2009;9:778–788. doi: 10.1038/nri2655. [DOI] [PubMed] [Google Scholar]