Abstract
Elucidating the molecular and biochemical details of bacterial infections can be challenging because of the many complex interactions that exist between a pathogen and its host. Consequently many tools have been developed to aid the study of bacterial pathogenesis. Small molecules are a valuable complement to traditional genetic techniques because they can be used to rapidly perturb genetically intractable systems and to monitor post-translationally regulated processes. Activity-based probes are a subset of small molecules that covalently label an enzyme of interest based on its catalytic mechanism. These tools allow monitoring of enzyme activation within the context of a native biological system and can be used to dissect the biochemical details of enzyme function. This review describes the development and application of activity-based probes to examine aspects of bacterial infection on both sides of the host-pathogen interface.
Keywords: Activity-based probes, bacterial pathogens, small molecule screening
Small molecules have long been applied to the study of bacteria and other microbes; however they have historically been used in the context of discovering antibiotics and other therapeutics capable of destroying the organism of interest. More recently, chemical tools have found use for creating and monitoring specific phenotypes in an effort to dissect the molecular events underlying pathogenesis. This is especially valuable for monitoring processes that are regulated post-translationally, such as enzymatic cascades that activate host immunity or remodel bacterial surfaces during infection. These events are difficult to study by easuring transcript or protein expression levels alone; and only sometimes can be monitored indirectly by traditional biochemical approaches such as western blotting. Methods employing small molecules that can rapidly and directly interact with enzymes of interest in a broadly applicable manner therefore provide a useful approach for dissecting host-pathogen interactions.
An overview of the general applications of activity-based probes
Activity-based probes (ABPs) are chemical tools that can be used to profile changes in enzyme activity levels (Figure 1).1 ABPs function by covalently modifying target enzymes via a specific chemical reaction that is dependent on the enzyme's catalytic mechanism. ABPs generally contain three components: an electrophilic trap or ‘warhead’, a specificity region, and a tag. The warhead portion of the probe is critical as it forms the covalent bond with the enzyme and, together with the specificity region, serves to direct the probe selectivity within the proteome. A wide variety of tags exist depending on the intended application, which can include direct visualization of enzyme activation or target isolation for further analysis. A number of comprehensive reviews have addressed the many possible components of ABPs and the benefits and applications of each type.1-3
Figure 1.
(a) Basic components of an activity-based probe (ABP). (b) General labeling mechanism of a target enzyme by an activity-based probe following post-translational enzyme activation. Green oval: zymogen pro-domain. Blue oval: enzyme catalytic domain. (c) Schematic of labeling target enzymes with clickable ABPs.
Because ABP labeling is dependent on an enzyme's catalytic mechanism and takes place within the active site, these tools can often be used to functionally dissect the biochemical details of enzyme structure and function. Although any enzyme target with a nucleophilic residue involved in catalysis is amenable to study using ABPs, and the diversity of ABPs being developed has dramatically increased over the past decade, the majority of examples where ABPs have been used to study protein function have focused on hydrolase targets. Proteases and other hydrolases often play critical roles in bacterial infection and host responses, making ABPs particularly valuable tools for studying these events.4-8
ABPs can be used to monitor enzymatic activation within a cellular setting, thus providing relevant information about target regulation in the context of its native environment. This also alleviates the need to purify or recombinantly express each protein of interest. Some ABP tags including fluorophores can be used to visualize localized enzymatic activity within a cell.9,10 The rapid readout of these probes makes them useful tools for studying the activation kinetics of an enzyme target, including real-time monitoring using ABPs containing quenched fluorophores.11,12
ABPs can also assess the biological targets and selectivity of small molecule inhibitors using a method known as competitive activity-based protein profiling (competitive ABPP) (Figure 2). In this strategy, a compound (or set of compounds) that produces a phenotype of interest are added to a sample followed by labeling with a broad-specificity ABP.13,14 Using this method it is possible to monitor the changes in activity of many related targets in response to inhibitor treatment. Furthermore, it is possible to begin to link a specific phenotype of a compound with inhibition of specific enzymatic targets. This approach also allows assessment of overall selectivity of an inhibitor within a related class of enzymes and can be used to assess the pharmacodynamic properties of a drug.13,14
Figure 2.
Scheme of the competitive activity-based protein profiling (competitive ABPP) method. A total proteome is either pre-incubated with a specific small molecule (top route) followed by labeling of residual enzyme activities using a broad-spectrum ABP and analysis by gel electrophoresis. This sample is compared to a sample from the same proteome in which the ABP is directly used to label all active targets (bottom route). This method allows direct assessment of target specificity across many related enzyme targets and can be used to develop and characterize enzyme inhibitors.
ABPs have broad utility in a number of areas of research. This review will give an overview of how ABPs have been applied to shed light on both bacterial infection mechanisms and host responses.
Applying activity-based probes to characterize bacterial factors associated with pathogenesis
Using ABPs to classify hydrolases involved in bacterial pathogenesis
The availability of genomic sequence information for many bacterial species has greatly improved the process of identifying new components of important biological process such as pathogenesis mechanisms. However, biochemical and functional characterization of these gene products has remained difficult, yet is critical to fully understand these processes. Because ABPs label enzymes based on their catalytic mode of action, they are ideal tools for biochemically classifying unknown proteins.14 This includes enzymes with activities associated with bacterial pathogenesis.
The Gram-positive bacterium Staphylococcus epidermidis is often thought of as a commensal organism associated with the skin and mucous membranes of the host. However, S. epidermidis also causes nosocomial infections at surgery sites and in immunocompromised patients. The broad spectrum papain family cysteine protease ABP DCG-04 was used monitor the activity of a putative protease with elastase activity termed Ecp in S. epidermidis cultures (Table 1).15,16 Probe labeling showed that Ecp activity was both cell wall associated and released into the growth media. By correlating labeling intensity with culture density, the authors were able to show that activity of the enzyme was growth-phase dependent and did not begin until mid-log phase. Because the protease was labeled by DCG-04 and inhibited by other small molecules specific for cysteine but not serine proteases, the authors classified Ecp as a cysteine protease. N-terminal sequencing identified Ecp as homologous to the staphopains A & B, which are known virulence factors in Staphylococcus aureus. It is therefore possible that S. epidermidis virulence mechanisms are similar to those of its relative S. aureus.
Table 1.
Activity-based probes used to profile bacterial factors associated with pathogenesis.
| Probe | Target Enzyme |
|---|---|

|
Staphylococcus epidermidis Cysteine protease Ecp16 |

|
Chlamydia trachomatis Peptide deformylase (PDF)17 |

|
Chlamydia trachomatis Serine protease CtHtrA18 |

|
Mycobacterium tuberculosis Serine protease Chp119 |
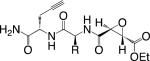
|
Clostridium difficile Cysteine protease |

|
Cwp8420 |

|
Staphylococcus aureus Serine protease ClpP, Various bacterial enzymes (hydrolases, ligases, transferases, oxidoreductases, etc.)27-30 |
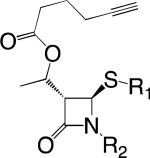
|
Staphylococcus aureus ClpP, β-Lactamase, MecR1, PBP232,33 |

|
Listeria monocytogenes Serine proteases ClpP1, ClpP234 |
|
|
Streptococcus pneumoniae Penicillin-binding proteins (PBPs)35 |

|
Escherichia coli Polyketide/non-ribosomal peptide synthase c245036 |

| |

|
Staphylococcus aureus Transferases MurA1, MurA2, PEP protein phosphotransferase (PtsI), Alkylhydroperoxide reductase C (AhpC)37 |

|
Staphylococcus aureus Protein tyrosine phosphotases PTPA and PTPB, Secretory antigen SsaA238 |

|
Vibrio cholerae, Pseudomonas aeruginosa Family 3 β-D-glucosaminidase NagZ41 |

|
Porphyromonas gingivalis Arg-gingipain RgpB25,44 |

|
Porphyromonas gingivalis Lys-gingipain Kgp45 |

|
Clostridium difficile toxin TcdB47 |
An ABP was also used to help identify and classify a factor critical for the growth of Chlamydia trachomatis within host cells.17 Two matrix metalloprotease (MMP) inhibitors were found to block chlamydial growth, and subsequent gene sequencing of mutants resistant to the compounds revealed a point mutation in the promoter of peptide deformylase (PDF). PDF is an enzyme that uses zinc to remove an N-terminal formyl group from bacterial proteins following translation. A hydroxamate-based ABP termed AspR1 was used to confirm that these inhibitors were indeed targeting PDF, as incubation with the inhibitors competed with activity-based labeling of the enzyme. This study identifies a new target with clinical potential for battling C. trachomatis infection.
Recently, a small molecule protease inhibitor library was screened against a purified serine protease, CtHtrA, from Chlamydia trachomatis. This screen identified a peptide diphenyl phosphonate inhibitor JO146 that was then used to characterize the protease function in the bacteria.18 Inhibition of CtHtrA by JO146 resulted in destruction of Chlamydia morphology and elimination of bacteria-containing inclusion bodies from infected host cells. This compound was also efficacious in mouse models of Chlamydia infection. The original inhibitor hit was then converted into an ABP by addition of Cy5- and biotin-tags. These ABPs based on JO146 were used to confirm labeling of CtHtrA by both gel competition and proteomics analysis. This study defines an essential role for HtrA in Chlamydia pathogenesis, and highlights a possible new drug target for Chlamydia treatment.
Mycobacterium tuberculosis synthesizes a sulfatide known as sulfolipid-1 (SL-1) that is implicated in virulence mechanisms including manipulation of the host immune response. Biosynthesis of SL-1 by the pathogen is therefore of great interest, however the last steps of the synthetic pathway and subsequent transport have remained unknown. A recent study used bioinformatics to identify a putative membrane anchored glycolipid acyltransferase termed Chp1, and then showed that this enzyme catalyzes sequential steps at the end of SL-1 biosynthesis.19 The authors used the broad spectrum serine hydrolase probe ABP TAMRA-fluorophosphonate to show that Chp1 possesses serine hydrolase activity. Based on this information, the authors hypothesized that Chp1 could be a target of the FDA-approved lipase inhibitor tetrahydrolipstatin (THL), which has been shown to be bacteriocidal. Chp1 was identified as a target of THL in M. tuberculosis culture, making pharmacological inhibition of Chp1 and SL-1 biosynthesis a promising approach for future studies of Chp1.
Clostridium difficile is a Gram-positive anaerobe that is highly antibiotic resistant and the leading cause of hospital-acquired infections. The composition of C. difficile's surface includes a robust S-layer, which is critical for both the integrity of the bacterium and its interactions with the host. The S-layer is composed of a low and high molecular weight protein, both of which are expressed as a single pro-peptide called SlpA. SlpA is then processed by a previously unidentified protease. In order to identify this enzyme, researchers tested several different classes of protease inhibitors and found that the cysteine protease inhibitor E-64 weakly inhibited SlpA processing.20 Based on this information, the authors then made more potent, alkyne-tagged ABPs and used them to show that processing of SlpA most likely occurs following translocation into the S-layer, since incubation with the compounds led to an accumulation of unprocessed SlpA in the media. Because the labeled target appeared to be extracellular, the authors used biotin-labeled versions of the probes to label, affinity isolate, and identify the protease Cwp84. Subsequent co-expression experiments in E. coli confirmed that Cwp84 processes SlpA, and a knockout has confirmed this protease is solely responsible for SlpA cleavage.21 The authors have also recently improved upon the chemical tools initially used in their study by changing the reactive electrophilic warhead to a C-terminal Michael acceptor.22
Novel ABP Scaffolds for dissecting virulence and antibiotic resistance mechanisms
While antibiotics are critical for combating bacterial infection, they are also valuable resources for understanding the host-pathogen interface. Some of the first chemical tools that can be characterized as activity-based probes were radiolabeled and resin-linked versions of the β-lactam antibiotic penicillin G, which were used to identify serine hydrolases termed penicillin-binding proteins (PBPs) in bacterial membranes (Figure 3).23,24 This strategy has since been expanded upon and has proven valuable for understanding mechanisms of antibiotic resistance and also for creating new classes of ABPs. These compounds have also been used to dissect the regulation and activity of virulence mechanisms in several species of pathogens (Figure 4). It is important to understand both the targets of antibiotic classes and the mechanisms bacterial pathogens use to evade antimicrobial treatments in order to maximize the efficacy of these compounds.
Figure 3.
Labeling mechanism of the β-lactam antibiotic penicillin G. The red asterisk indicates the location of the 14C radiolabel on the probe used in the original identification studies.
Figure 4.
New antibiotic-based ABP scaffolds applied to study mechanisms of bacterial virulence and resistance. Top arrow: β-lactone probes label the widely conserved, virulence-associated protease ClpP. Probes of this type were used to identify inhibitors of ClpP using competitive activity-based protein profiling (ABPP). Bottom arrow: Probes containing antibiotic scaffolds are valuable for identifying differences in bacterial strains associated with antibiotic resistance, virulence, or other phenotypes of interest. MRSA: Multiresistant Staphylococcus aureus.
In addition to commonly used electrophilic warheads such as acyloxymethyl ketones (AOMKs),25 epoxides,15 and fluorophosphonates,26 the strained β-lactone ring has more recently been shown to be a viable electrophilic moiety for use in ABPs. Probes based on the β-lactone scaffold were successfully used to label a number of hydrolases in bacterial cultures, including the widely conserved serine protease ClpP (caseinolytic protein protease).27,28 ClpP has been associated with virulence in species such as the Gram-positive facultative anaerobes Staphylococcus aureus and Listeria monocytogenes. Competitive ABPP was subsequently used to create inhibitors for ClpP, and a refined inhibitor was shown to block activity of the enzyme in S. aureus culture (Figure 4). Using this inhibitor, invasive proteolytic and hemolytic activities were confirmed to be regulated directly by ClpP in wild-type and methicillin-resistant (MRSA) strains.29 In addition, these techniques were used to demonstrate that ClpP is upstream of several critical virulence factors in pyrogenic toxin superantigen (PTSA)-producing strains of S. aureus. These virulence factors included the enterotoxins SEB and SEC3 as well as toxic shock syndrome toxin (TSST-1). Similar experiments were also performed in L. monocytogenes cultures, and were able to attenuate virulence of this intracellular pathogen.30 The authors also showed that ClpP inhibition down-regulates the activity of the pore-forming toxin LLO as well as the virulence factor phosphatidylinositol-specific phospholipase C (PI-PLC).
‘Clickable’ ABPs contain a latent reactive group that allows conjugation of any desired tag to the probe using highly bio-orthogonal CLICK chemistry following target labeling (Figure 1c).3,31 ABPs of this type have been developed using common antibiotic and synthetic derivatives of the β-lactam scaffold as the primary recognition elements. These probes were used to label both lysates and cultures for several bacterial species.32 Probes with modifications found in natural antibiotics primarily labeled PBPs, while those with novel substituents labeled other enzymes including ClpP and the virulence-implicated β-lacatamase. These probes were also used to profile differences between antibiotic-sensitive and methicillin-resistant strains of S. aureus (Figure 4).33 This work identified several enzymes that were only active in MRSA cultures, including the known resistance-associated pair MecR1 and PBP2, as well as two novel hydrolases that could contribute to β-lactam resistance mechanisms in MRSA.
More recently, additional ABPs based on β-lactone and β-lactam scaffolds with improved specificity been developed. Some bacterial species including L. monocytogenes have two ClpP2 isoforms known as ClpP1 and 2. However, first-generation ABPs based on the β-lactone moiety were only able to label ClpP2. Therefore, a more strained, and therefore more reactive vibralactone-based probe was synthesized.34 This ABP was able to label both ClpP isoforms, and was used in E. coli co-expression studies to show that ClpP2 activates ClpP1 and that ClpP1 and Clp2 hetero-oligomerize with each other. A probe based on the β-lactam antibiotic cephalosporin C was also developed that exhibits enhanced selectivity towards a subset of PBPs.35 This probe was used to visualize active PBPs in a live uncapsulated derivative of the human pathogen Streptococcus pneumoniae.
In another study, a panel of α-alkylidene-γ-butyrolactones was synthesized and used to compare the activity profiles of enzymes in non-pathogenic and uropathogenic E. coli.36 The probes identified the enzyme c2450 as only active in the pathogenic strain. c2450 is a hybrid polyketide/non-ribosomal peptide synthase involved in synthesis of the virulence factor colibactin which activates the DNA checkpoint damage response in host cells, causing them to enter cell cycle arrest. This new insight into the structure and regulation of these important virulence factors highlights the power of activity-based protein profiling.
The antibiotic showdomycin highly resembles the nucleoside uridine, but with an electrophilic maleimide moiety conjugated to the pentose ring instead of a base. In order to analyze the mode of action of this potent class of compounds, one study described an alkyne-tagged ‘clickable’ version of the parent drug and used it to probe multiple strains of different bacterial species in culture (Figure 4).37 After lysis and analysis of the labeled proteins, the authors identified several families of enzymes critical for pathogen colonization and survival. This included the transferases MurA1 and MurA2 in S. aureus, which are essential for cell wall biosynthesis. Subsequently, the authors showed that while MurA2 is transcribed in four different S. aureus strains, it is only active in MRSA, highlighting the utility of a readout of activity levels. The MRSA strain was the only one resistant to higher concentrations of fosfomycin, a MurA inhibitor. These results provide a possible mechanism for the mode of action of showdomycin. The showdomycin-based ABP also more generally was able to probe pathogenesis mechanisms, by showing that phosphoenolpyruvate-protein phosphotransferase (PtsI) is only active in virulent toxin-producing S. aureus strains, including clinical isolates. In contrast, alkylhydroperoxide reductase C (AhpC) was only labeled in less pathogenic lab strain NCTC 8325. This work resulted in a highly novel ABP scaffold based on a naturally occurring maleimide electrophile that has already proven useful for gaining insight into differences between virulent and avirulent bacterial strains.
In theory, any compound containing an electrophilic moiety that has biological activity can be used to generate an activity-based probe. Cinnamic acid is a key flavor component that contains a Michael acceptor known to label cysteines. A library of peptides conjugated to cinnamic acid was used to make ABPs for detecting the activity of recombinantly expressed protein tyrosine phosphotases PTPA and PTPB from MRSA.38 These probes were then used to profile the MRSA membrane proteome. Although the authors were unable to detect endogenous levels of PTPs in the samples, the probes did label the staphylococcal secretory antigen SsaA2. The function of this enzyme is unknown, but has been implicated in S. aureus pathogenesis mechanisms including sepsis and endocarditis. This unique class of probes may be useful for further studying this potentially important virulence factor.
The glycoside hydrolase NagZ is involved in peptidoglycan recycling in Gram-negative bacteria and has also been shown to regulate AmpC, a β-lactamase that confers resistance to several types of β-lactam based antibiotics.39,40 NagZ is an exo-glycosidase that uses a catalytic nucleophile to hydrolyze N-acetylglucosamine residues during cell wall recycling. This nucleophile creates a covalent intermediate with its substrate, making it amenable to ABP profiling. An azide-labeled probe termed 2-azidoacetamido-2-deoxy-5-fluoro-β-D-glucopyranosyl fluoride (2AA5FGF) was created that stabilizes this covalent intermediate under acidic conditions and allows activity to be monitored after click chemistry attachment of a label.41 This tool was used to identify a presumed 3 β-D-glucosaminidase in P. aeruginosa lysate. Based on molecular weight, the authors hypothesized this enzyme was the putatively annotated PA3005. This was confirmed following cloning, expression, and biochemical characterization of PA3005. More recently, it has been shown that chemical or genetic inactivation of NagZ in P. aeruginosa results increased susceptibility to anti-pseudomonal β-lactam antibiotics,42 highlighting the importance of this target in resistance mechanisms.
Using ABPs to analyze mechanisms regulating enzymes involved in bacterial pathogenesis
Porphyromonas gingivalis is an anaerobic, Gram-negative pathogen that causes the widespread oral inflammatory disease, periodontitis. A major component of P. gingivalis virulence involves the secretion of cysteine endoproteases called gingipains. Gingipains include the Arg-gingipains RgpA and RgpB, and the Lys-gingipain Kgp. These proteases cleave their substrates after arginine or lysine residues, and are involved in all phases of pathogenesis, from colonization and nutrient acquisition to neutralization of host immune responses.43
Like most proteases, gingipains are synthesized as inactive zymogens. Their trafficking and activation is regulated by post-translational proteolytic processing. Gingipain activation has been studied using ABPs containing the cysteine protease-specific acyloxymethyl ketone (AOMK) warhead.25 In one study a biotin-labeled Arg-Lys-AOMK called BiRK was used to dissect the proteolytic activation sequence of pro-RgpB expressed in Saccharomyces cerevisiae.44 The authors used the probe at a sub-saturating concentration to determine the relative activity levels of different forms of the enzyme during processing. They then compared this with the total amounts of each form present as determined by saturated probe labeling. Using this information it was possible to demonstrate that RgpB gains activity during three discrete processing steps including truncation of both the N- and C-terminus, resulting in an overall 80-fold increase in activity. This finding was surprising because proteolytic activation of protease zymogens usually results in an activity increase of several orders of magnitude. This led the authors to conclude that the N-terminus of the zymogen may actually serve as a chaperone for folding of the catalytic domain, rather than as an inhibitor of the enzyme.
The gingipains Kgp and RgpA both function as large, non-covalent complexes on the surface of P. gingivalis. In a recent study, ABPs were used to examine the formation of these complexes by the Lys-gingipain Kgp.45 The authors identified an oligomerization sequence motif present in both the catalytic and hemagglutinin-adhesin domains of Kgp thought to be essential for complex processing and assembly. To test this hypothesis, they genetically disrupted this motif within the catalytic domain of Kgp. The authors used an ABP to show that the mutant Kgp was no longer properly processed and assembled on the surface of the bacteria, but instead the active catalytic domain was secreted into the growth media. The probes also did not label the full-length zymogen, indicating that the prodomain is inhibitory. These two studies are examples of how ABPs can provide insight into the processing and activation mechanisms of critical P. gingivalis virulence factors.
Clostridium difficile pathogenesis is also highly regulated by post-translational events. C. difficile secretes multidomain protein toxins, including the large glucosylating toxins TcdA and TcdB, that are responsible for a majority of the pathology associated with C. difficile infection (CDI).46 These toxins contain an internal cysteine protease domain (CPD) that autoproteolytically activates the toxin upon binding the eukaryote-specific small molecule inositol hexakisphosphate (InsP6) inside host epithelial cells. Our laboratory developed a set of covalent inhibitors and activity-based probes to study the substrate recognition and activation mechanisms of the TcdB CPD.47 As with other proteases, the library of peptidic inhibitors provided a way to rapidly gain biochemical insight into the specificity of the CPD active site that was not obvious from structural analysis alone.20,33
The probe AWP19 was designed from the inhibitor library screening information and has subsequently been used to biochemically dissect the details of an allosteric circuit in the TcdB CPD that regulates activation of this toxin protease domain upon infection of a host cell.48 The sensitivity of AWP19 enabled small differences in the activity levels of various CPD mutants to be quantified, allowing functional roles to be attributed to individual residues in the protein. Additionally, incubation of the CPD with saturating concentrations of AWP19 revealed that basal amounts of the enzyme are labeled even in the absence of the activator InsP6. This labeling was dependent on the presence of the CPD active site cysteine, and was not due to contaminating InsP6 from recombinant production of the CPD in E. coli. These data suggest that the CPD transiently adopts an active conformation without InsP6, and this active conformation is stabilized by InsP6.
AWP19 has also been used to compare the activation characteristics of the TcdB cysteine protease domain (CPD) from the historical C. difficile strain and the hypervirulent NAP1 variant within the context of the full-length holotoxin.49 This study revealed that there is significantly more processing of the TcdB toxin from the hypervirulent strain (TcdBHV) than the historical one. The authors also found that the active site of the full-length TcdBHV is less accessible to small molecule probes, leading them to hypothesize that the toxin CPD may have an increased affinity for its intramolecular substrate. These findings, together with studies that show TcdBHV is able to undergo conformational changes at a higher pH during endosome acidification supports the notion that the augmented activation characteristics of TcdBHV play a role in the hypervirulence of the NAP1 strain.50
Using activity-based probes to profile host responses to bacterial infection
The host response to bacterial infection involves a complex set of protective and destructive programs, many of which are regulated by proteolytic activity. For example, both the caspase and cathepsin protease families play key roles in cell death and inflammatory responses.51,52 Additionally, in many cases, the damaging effects of the host response contribute significantly to the disease pathology associated with bacterial infection. Because these processes involve hydrolases and are regulated post-translationally, ABPs are useful tools for studying both host defense mechanisms and contributions to bacterial pathogenesis. However, to date relatively few classes of ABPs have been used for this purpose.
Mice infected with the attaching-effacing enteropathogen Citrobacter rodentium serve as a model for disease caused by human enteropathogenic E. coli (EPEC) and enterohemorrhagic E. coli (EHEC). C. rodentium causes mucosal inflammation in the intestinal tract, and the proteinase-activated receptors (PARs) are known to play a role in controlling intestinal inflammatory responses. One study showed that host proteases capable of activating PAR2 are released into the intestinal lumen upon infection with C. rodentium.53 The authors used a diphenylphosphonate-based ABP called Bio-PK to characterize these host factors as serine proteases with trypsin-like activity (Table 2). Based on this result, the authors treated mice orally with soybean trypsin inhibitor (STI) and showed that it was possible to decrease the inflammation-based pathology associated with infection. These results provide a possible new target for reducing unproductive host inflammatory responses to enteropathogens.
Table 2.
Activity-based probes used to study host responses to bacterial infection.
| Probe | Target Enzyme |
|---|---|

|
Trypsin-like serine protease activators of proteinase-activated receptor PAR2 in response to Citrobacter rodentium infection.53 |

|
Lysosomal cysteine protease cathepsin B in response to Salmonella enterica serovar Typhimurium infection.9 |

|
Pro-inflammatory cysteine protease caspase-1 activation in response to several bacterial pathogens.55-60,65-67 |

|
Pro-apoptotic caspases-3, -7 and pro-inflammatory caspase-1 in response to bacterial infection.10 |
Salmonella enterica serovar Typhimurium (S. typhimurium) is a Gram-negative intracellular pathogen that invades host cells by triggering phagocytosis. One study used an ABP to monitor activation of the lysosomal cysteine protease cathepsin B in endocytic compartments containing S. typhimurium.9 The authors synthesized an ABP with an epoxide warhead and a click tag, and used it to image cathepsin B activity by microscopy (Figure 5). The authors discovered that active CatB was absent from vacuoles containing Salmonella, suggesting that the bacterium is capable of inhibiting CatB activation. Because the cathepsins are involved in inflammatory responses, the authors surmised that this could be part of Salmonella's method for evading host defense mechanisms.
Figure 5.
Applications of small molecule probes to the study of pathogen induced inflammation pathways. (a) Inflammasome-mediated caspase-1 activation by the intracellular pathogen Salmonella. Caspase-1 activation leads to pyrogenic cytokine processing and release and also triggers the pro-inflammatory cell death program pyroptosis. The tan oval highlights where the activity-based probes Azido-E-64 has been used to study cathepsin B activity and the grey oval highlights where FLICA and AWP28 have been used to monitor caspase-1 activation during infection. Caspase-3 and -7 activation and apoptosis is triggered in the absence of inflammasome-mediated caspase-1 activation. (b) Protein domain structures of selected inflammasome components involved in Salmonella recognition. These proteins assemble via homotypic domain interactions.
Using FLICA to study inflammasome-mediated caspase-1 activation and pyroptosis in response to bacterial infection
The cysteine protease caspase-1 plays a critical role in controlling host inflammatory response to infection. A cytosolic, multi-protein complex called the inflammasome activates the procaspase-1 zymogen upon detection of multiple pro-inflammatory signals, including pathogen-associated molecular patterns (PAMPs) (Figure 5).4,7 In phagocytes including macrophages, caspase-1 catalyzes the processing and subsequent release of pyrogenic cytokines including interleukin-1β and interleukin-18. These signals mount an innate immune response to the offending stimulus. Caspase-1 also triggers the pro-inflammatory cell death program termed pyroptosis, which leads to osmotic lysis of the cell in an effort to control infection by destroying the replicative niche of the pathogen.51,54 Because caspase-1 is activated post-translationally by the inflammasome in response to bacterial pathogens, it is an ideal target to study using ABPs. The fluorescent inhibitor of caspases (FLICA) probes are commercially available fluoromethyl ketone (FMK)-based ABPs that have been used extensively to study the host response to a variety of bacterial pathogens. This includes species of Shigella,55Chlamydia,56Mycobacterium,57Helicobacter,58 and others including some discussed here in more detail. FLICA is primarily used with host cell lines lacking specific response genes in order to determine whether these genes play a role in caspase-1 activation using microscopy or flow cytometry. These studies are useful for elucidating intracellular pathways involved in pathogen detection.
FLICA has been used to study to the spatial organization of active caspase-1 in macrophages that have been infected by intracellular pathogens including S. typhimurium (Figure 5). One study showed that the cytosolic receptors NLRP3 and NLRC4 respond to distinct Salmonella PAMPs upon infection and they integrate these signals into a single, defined macromolecular focus that contains the adapter protein ASC.59 Formation of this structure was independent of caspase-1, which is subsequently recruited to the focus. Microscopy with FLICA showed that while approximately 80% of the foci contained caspase-1 in wild-type cells, only about 10% contained active caspase-1. The authors used a caspase-1 inhibitor to determine that this focus is the site of interleukin processing. Subsequent work revealed that receptors containing a caspase-recruitment domain (CARD), including NLRC4, trigger pyroptosis by directly associating with caspase-1 in the absence of ASC (Figure 5).60 In these cases, the basal activity of the procaspase-1 zymogen is sufficient to cause pyroptosis. Microscopy with FLICA showed that no large focus containing active caspase-1 is formed in pyroptotic ASC knockout cells. This work spatially and biochemically decouples the cell death and cytokine processing roles of caspase-1 in response to bacterial infection.
Some bacterial pathogens actively manipulate host responses during infection. FLICA has been used with microscopy to dissect how species of the Gram-negative Yersinia genus trigger and manipulate cell death programs in infected macrophages. The Yersinia effector protein YopJ is secreted into host cells via the bacteria's type III secretion system (T3SS), and inhibits host survival signals including mitogen-activated protein (MAP) kinase pathways, which can lead to apoptosis.61 Other Yersinia T3SS effectors have also been implicated in caspase-1 inhibition.62,63 FLICA was used to show that pre-stimulation of macrophages with toll-like receptor (TLR) agonists including LPS causes macrophages to activate caspase-1 and undergo pyroptosis in response to infection with Y. pseudotuberculosis in a YopJ-independent manner.64 FLICA was also used to show that infecting macrophages with the KIM strain of Y. pestis, the causative agent of plague, triggers caspase-1 activation.65 These cells release processed IL-1β and undergo cell death, but the death was found to be independent of caspase-1. Subsequent work showed that this strain harbors a form of YopJ that inhibits the host protein inhibitor of nuclear factor kappa-B kinase beta (IKKβ), and FLICA experiments in IKKβ knockout macrophages showed that infection of cells lacking IKKβ have increased caspase-1 activation.66 Together these studies confirm that the interaction between Yersinia and its host consists of a balance of both apoptotic and pyroptotic cell death programs.
FLICA has also been used to show that active caspase-1 has a pro-survival role in response to some bacterial virulence factors. Many bacteria including Aeromonas and Listeria species use pore-forming bacterial toxins during infection, leading to loss of intracellular potassium. One study used FLICA and flow cytometry to show that caspase-1 is activated by pore-forming proteins such as the Aeromonas toxin aerolysin.67 Knockdown experiments confirmed that full caspase-1 activation involved contributions from NLRC4, NLRP3, and ASC. Furthermore, caspase-1 activation was shown to promote pro-survival membrane biogenesis pathways, and abrogation of these pathways results in increased cell death. This study provides an example for another, non-canonical role for caspase-1 activity in response to bacterial pathogenesis.
FLICA as a commercial probe for caspase-1
While the ABP FLICA provides a useful readout of caspase-1 activation in the study of host-pathogen interactions, caution must be taken when using these tools for imaging purposes. Several studies have shown that FMK-based probes are not selective for their caspase target, and instead label additional active caspases and other classes of proteases such as cathepsins.10,68-70 This has led to some confusion in how to interpret results generated using the FLICA probes.
For example, studies used FLICA with flow cytometry to show that caspase-1 activation by Legionella requires the cytosolic receptor NAIP5.71 However, a six-fold increase in FLICA labeling was also observed in infected Casp1−/− macrophages when compared to uninfected cells, which the authors attributed to cross-reactivity of the probe with other caspases. The use of these unselective probes to detect caspase-1 activity was subsequently questioned in a conflicting report.72 This particular issue was finally resolved when biochemical reconstitution and mouse knockout experiments definitively proved a role for both NAIP5 and NLRC4 in the activation of caspase-1 upon detection of cytosolic flagellin from bacteria including Legionella.73,74
FLICA labeling also produced confusing results in the previously highlighted work examining the role of caspase-1 autoproteolytic activation in cytokine processing and pyroptosis.60 This study showed that when FLICA-labeled macrophages were examined by microscopy, labeling was found in areas where caspase-1 was not detected by a selective caspase-1 antibody. This led the authors to question the specificity of FLICA. Despite these reports and the known lack of selectivity for FMK-based probes, the scope of targets labeled by FLICA upon bacterial infection have not been analyzed biochemically.
Development and use of AWP28 to study pyroptosis
Our laboratory has attempted to address the limitations of FLICA by creating an optimized activity-based probe for selective visualization of caspase activation in response to bacterial infection. This recently developed probe called AWP28 makes use of the AOMK electrophile that has greater overall selectivity for cysteine protease targets coupled with a more caspase-1 targeted peptide backbone.10 Due to overlapping substrate specificities of many proteases including caspases, it remains difficult to target individual caspases.75 While AWP28 labels multiple active caspases upon infection, the optimized properties of this probe allow the labeled caspases to be analyzed and identified by gel electrophoresis. Therefore it was possible to use AWP28 to show that apoptosis is robustly triggered by Salmonella infection in macrophages lacking the ability to activate pyroptosis through the caspase-1 pathway. Furthermore, these studies confirmed that caspase-1 activity is required to bypass the apoptotic response; identifying a possible feedback mechanism between two critical cell death programs. This work may provide insight into a potential intracellular arms race between the host and pathogens capable of suppressing caspase-1 activation, and also highlights the balance that exists between pro- and anti-inflammatory cellular responses.
Conclusions and future uses of chemical probes for studying bacterial pathogenesis
Using small molecule tools to study biological processes can be complicated by difficulties accessing the intracellular environment. This is especially true in the case of bacteria, which possess many layers of resistance to chemical warfare. The outer membrane of Gram-negative bacteria is notoriously difficult to penetrate using small molecules, and many bacterial species possess arrays of efflux pumps.76,77 It therefore can be difficult to use chemical probes to study the internal physiology of bacterial pathogens. Fortunately, there have been some successes developing cell permeable small molecule tools to build upon. Activity-based probes (ABPs) tagged with a bio-orthogonal moiety such as an azide or alkyne have been used to label intracellular enzymes in bacterial culture.37
Broadening the enzymatic targets of ABPs will also be critical to obtaining a holistic view of the molecular mechanisms involved in pathogenesis. The development of new ABP scaffolds requires an intimate understanding of the target enzyme family's mechanism of catalysis, since these probes harness this mechanism to label in an activity-dependent manner. A recently created probe targeting histidine kinases is promising, as these enzymes are involved in bacterial two-component signaling which can be critical for regulating bacterial virulence.78,79
The application of ABPs has diverged into two vastly different directions that both take advantage of their ability to report enzyme activity. One approach focuses on the use of general structures and electrophiles to globally profile the activity of a specific type of nucleophilic mechanism. For example, iodoacetamide has been used in reactive cysteine profiling to determine how the activity of nucleophilic cysteine residues varies within and between enzymes in complex proteomes.80 These general techniques take advantage of powerful proteomics methods and can provide insight into how whole families of enzymes are regulated under distinct biological conditions. General ABPs can be used to determine the targets of compounds that create phenotypes of interest using competitive activity-based protein profiling.13,29,81 Such probes are also capable of rapidly profiling differences between virulent and avirulent bacterial strains,36 and may therefore provide a means to functionally assess different isolates. These techniques have a great deal of untapped potential for understanding pathogenic mechanisms in diverse strains.
A second application involves using ABPs for imaging purposes. In these cases, highly selective compounds are needed in order to assure that the observed readout can be attributed to the activity of a specific target enzyme. This can be a difficult task because many families of related enzymes have overlapping substrate profiles.75 This means the selectivity of an ABP can often not be made absolute by modifying the specificity region alone. One way to improve the selectivity of chemical probes is to make use of complementary genetic techniques. For example, “bump-hole” approaches can be used to modify an enzyme of interest so it is a unique target for a complementary chemical probe.82 Recently, this approach has been applied to protease targets as well.83,84 This technique is viable for studying bacterial virulence mechanisms in genetically tractable pathogens. Imaging enzyme activation is invaluable for dynamic host-pathogen interactions that are regulated post-translationally, and it will be exciting to see how this is accomplished in the future.
Many chemical biology tools languish following the proof of concept stage because they lack the simplicity and robustness necessary to answer biological questions. In contrast, activity-based probes are an example of a powerful technology that can be used to rapidly gain biological and biochemical insight into a variety enzymes and systems including bacterial infection and host responses. The studies discussed here highlight the power of using these tools to dissect mechanisms of bacterial pathogenesis.
Acknowledgments
Funding
A.W.P. was supported by an N.S.F. Graduate Research Fellowship. This work was funded by a grant from the National Institutes of Health R01 EB005011 (to M.B.).
References
- 1.Cravatt BF, Wright AT, Kozarich JW. Activity-based protein profiling: from enzyme chemistry to proteomic chemistry. Annu Rev Biochem. 2008;77:383–414. doi: 10.1146/annurev.biochem.75.101304.124125. [DOI] [PubMed] [Google Scholar]
- 2.Edgington LE, Verdoes M, Bogyo M. Functional imaging of proteases: recent advances in the design and application of substrate-based and activity-based probes. Curr Opin Chem Biol. 2012;15:798–805. doi: 10.1016/j.cbpa.2011.10.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Sadaghiani AM, Verhelst SH, Bogyo M. Tagging and detection strategies for activity-based proteomics. Curr Opin Chem Biol. 2007;11:20–28. doi: 10.1016/j.cbpa.2006.11.030. [DOI] [PubMed] [Google Scholar]
- 4.Broz P, Monack DM. Molecular mechanisms of inflammasome activation during microbial infections. Immunol Rev. 2011;243:174–190. doi: 10.1111/j.1600-065X.2011.01041.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Butler SM, Festa RA, Pearce MJ, Darwin KH. Self-compartmentalized bacterial proteases and pathogenesis. Mol Microbiol. 2006;60:553–562. doi: 10.1111/j.1365-2958.2006.05128.x. [DOI] [PubMed] [Google Scholar]
- 6.Gur E, Biran D, Ron EZ. Regulated proteolysis in Gram-negative bacteria--how and when? Nat Rev Microbiol. 2011;9:839–848. doi: 10.1038/nrmicro2669. [DOI] [PubMed] [Google Scholar]
- 7.Schroder K, Tschopp J. The inflammasomes. Cell. 2010;140:821–832. doi: 10.1016/j.cell.2010.01.040. [DOI] [PubMed] [Google Scholar]
- 8.Shen A. Autoproteolytic activation of bacterial toxins. Toxins (Basel) 2010;2:963–977. doi: 10.3390/toxins2050963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Hang HC, Loureiro J, Spooner E, van der Velden AW, Kim YM, Pollington AM, Maehr R, Starnbach MN, Ploegh HL. Mechanism-based probe for the analysis of cathepsin cysteine proteases in living cells. ACS Chem Biol. 2006;1:713–723. doi: 10.1021/cb600431a. [DOI] [PubMed] [Google Scholar]
- 10.Puri AW, Broz P, Shen A, Monack DM, Bogyo M. Caspase-1 activity is required to bypass macrophage apoptosis upon Salmonella infection. Nature chemical biology. 2012;8:745–747. doi: 10.1038/nchembio.1023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Blum G, Mullins SR, Keren K, Fonovic M, Jedeszko C, Rice MJ, Sloane BF, Bogyo M. Dynamic imaging of protease activity with fluorescently quenched activity-based probes. Nat Chem Biol. 2005;1:203–209. doi: 10.1038/nchembio728. [DOI] [PubMed] [Google Scholar]
- 12.Edgington LE, Berger AB, Blum G, Albrow VE, Paulick MG, Lineberry N, Bogyo M. Noninvasive optical imaging of apoptosis by caspase-targeted activity-based probes. Nat Med. 2009;15:967–973. doi: 10.1038/nm.1938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Barglow KT, Cravatt BF. Activity-based protein profiling for the functional annotation of enzymes. Nat Methods. 2007;4:822–827. doi: 10.1038/nmeth1092. [DOI] [PubMed] [Google Scholar]
- 14.Greenbaum DC, Arnold WD, Lu F, Hayrapetian L, Baruch A, Krumrine J, Toba S, Chehade K, Bromme D, Kuntz ID, Bogyo M. Small molecule affinity fingerprinting. A tool for enzyme family subclassification, target identification, and inhibitor design. Chem Biol. 2002;9:1085–1094. doi: 10.1016/s1074-5521(02)00238-7. [DOI] [PubMed] [Google Scholar]
- 15.Greenbaum D, Medzihradszky KF, Burlingame A, Bogyo M. Epoxide electrophiles as activity-dependent cysteine protease profiling and discovery tools. Chem Biol. 2000;7:569–581. doi: 10.1016/s1074-5521(00)00014-4. [DOI] [PubMed] [Google Scholar]
- 16.Oleksy A, Golonka E, Banbula A, Szmyd G, Moon J, Kubica M, Greenbaum D, Bogyo M, Foster TJ, Travis J, Potempa J. Growth phase-dependent production of a cell wall-associated elastinolytic cysteine proteinase by Staphylococcus epidermidis. Biol Chem. 2004;385:525–535. doi: 10.1515/BC.2004.062. [DOI] [PubMed] [Google Scholar]
- 17.Balakrishnan A, Patel B, Sieber SA, Chen D, Pachikara N, Zhong G, Cravatt BF, Fan H. Metalloprotease inhibitors GM6001 and TAPI-0 inhibit the obligate intracellular human pathogen Chlamydia trachomatis by targeting peptide deformylase of the bacterium. J Biol Chem. 2006;281:16691–16699. doi: 10.1074/jbc.M513648200. [DOI] [PubMed] [Google Scholar]
- 18.Gloeckl S, Ong VA, Patel P, Tyndall JD, Timms P, Beagley KW, Allan JA, Armitage CW, Turnbull L, Whitchurch CB, Merdanovic M, Ehrmann M, Powers JC, Oleksyszyn J, Verdoes M, Bogyo M, Huston WM. Identification of a serine protease inhibitor which causes inclusion vacuole reduction and is lethal to Chlamydia trachomatis. Mol Microbiol. 2013 doi: 10.1111/mmi.12306. doi: 10.1111/mmi.12306. [DOI] [PubMed] [Google Scholar]
- 19.Seeliger JC, Holsclaw CM, Schelle MW, Botyanszki Z, Gilmore SA, Tully SE, Niederweis M, Cravatt BF, Leary JA, Bertozzi CR. Elucidation and chemical modulation of sulfolipid-1 biosynthesis in Mycobacterium tuberculosis. J Biol Chem. 2012;287:7990–8000. doi: 10.1074/jbc.M111.315473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Dang TH, de la Riva L, Fagan RP, Storck EM, Heal WP, Janoir C, Fairweather NF, Tate EW. Chemical probes of surface layer biogenesis in Clostridium difficile. ACS Chem Biol. 2010;5:279–285. doi: 10.1021/cb9002859. [DOI] [PubMed] [Google Scholar]
- 21.Kirby JM, Ahern H, Roberts AK, Kumar V, Freeman Z, Acharya KR, Shone CC. Cwp84, a surface-associated cysteine protease, plays a role in the maturation of the surface layer of Clostridium difficile. J Biol Chem. 2009;284:34666–34673. doi: 10.1074/jbc.M109.051177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Tam Dang TH, Fagan RP, Fairweather NF, Tate EW. Novel inhibitors of surface layer processing in Clostridium difficile. Bioorg Med Chem. 2012;20:614–621. doi: 10.1016/j.bmc.2011.06.042. [DOI] [PubMed] [Google Scholar]
- 23.Suginaka H, Blumberg PM, Strominger JL. Multiple penicillin-binding components in Bacillus subtilis, Bacillus cereus, Staphylococcus aureus, and Escherichia coli. J Biol Chem. 1972;247:5279–5288. [PubMed] [Google Scholar]
- 24.Blumberg PM, Strominger JL. Covalent affinity chromatography of penicillin-binding components from bacterial membranes. Methods Enzymol. 1974;34:401–405. doi: 10.1016/s0076-6879(74)34046-3. [DOI] [PubMed] [Google Scholar]
- 25.Kato D, Boatright KM, Berger AB, Nazif T, Blum G, Ryan C, Chehade KA, Salvesen GS, Bogyo M. Activity-based probes that target diverse cysteine protease families. Nat Chem Biol. 2005;1:33–38. doi: 10.1038/nchembio707. [DOI] [PubMed] [Google Scholar]
- 26.Liu Y, Patricelli MP, Cravatt BF. Activity-based protein profiling: the serine hydrolases. Proc Natl Acad Sci U S A. 1999;96:14694–14699. doi: 10.1073/pnas.96.26.14694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Bottcher T, Sieber SA. Beta-lactones as specific inhibitors of ClpP attenuate the production of extracellular virulence factors of Staphylococcus aureus. J Am Chem Soc. 2008;130:14400–14401. doi: 10.1021/ja8051365. [DOI] [PubMed] [Google Scholar]
- 28.Bottcher T, Sieber SA. Beta-lactones as privileged structures for the active-site labeling of versatile bacterial enzyme classes. Angew Chem Int Ed Engl. 2008;47:4600–4603. doi: 10.1002/anie.200705768. [DOI] [PubMed] [Google Scholar]
- 29.Bottcher T, Sieber SA. Structurally refined beta-lactones as potent inhibitors of devastating bacterial virulence factors. Chembiochem. 2009;10:663–666. doi: 10.1002/cbic.200800743. [DOI] [PubMed] [Google Scholar]
- 30.Bottcher T, Sieber SA. Beta-lactones decrease the intracellular virulence of Listeria monocytogenes in macrophages. ChemMedChem. 2009;4:1260–1263. doi: 10.1002/cmdc.200900157. [DOI] [PubMed] [Google Scholar]
- 31.Agard NJ, Baskin JM, Prescher JA, Lo A, Bertozzi CR. A comparative study of bioorthogonal reactions with azides. ACS Chem Biol. 2006;1:644–648. doi: 10.1021/cb6003228. [DOI] [PubMed] [Google Scholar]
- 32.Staub I, Sieber SA. Beta-lactams as selective chemical probes for the in vivo labeling of bacterial enzymes involved in cell wall biosynthesis, antibiotic resistance, and virulence. J Am Chem Soc. 2008;130:13400–13409. doi: 10.1021/ja803349j. [DOI] [PubMed] [Google Scholar]
- 33.Staub I, Sieber SA. Beta-lactam probes as selective chemical-proteomic tools for the identification and functional characterization of resistance associated enzymes in MRSA. J Am Chem Soc. 2009;131:6271–6276. doi: 10.1021/ja901304n. [DOI] [PubMed] [Google Scholar]
- 34.Zeiler E, Braun N, Bottcher T, Kastenmuller A, Weinkauf S, Sieber SA. Vibralactone as a tool to study the activity and structure of the ClpP1P2 complex from Listeria monocytogenes. Angew Chem Int Ed Engl. 2011;50:11001–11004. doi: 10.1002/anie.201104391. [DOI] [PubMed] [Google Scholar]
- 35.Kocaoglu O, Calvo RA, Sham LT, Cozy LM, Lanning BR, Francis S, Winkler ME, Kearns DB, Carlson EE. Selective penicillin-binding protein imaging probes reveal substructure in bacterial cell division. ACS Chem Biol. 2012;7:1746–1753. doi: 10.1021/cb300329r. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Kunzmann MH, Sieber SA. Target analysis of alpha-alkylidene-gamma-butyrolactones in uropathogenic E. coli. Mol Biosyst. 2012;8:3061–3067. doi: 10.1039/c2mb25313e. [DOI] [PubMed] [Google Scholar]
- 37.Bottcher T, Sieber SA. Showdomycin as a versatile chemical tool for the detection of pathogenesis-associated enzymes in bacteria. J Am Chem Soc. 2010;132:6964–6972. doi: 10.1021/ja909150y. [DOI] [PubMed] [Google Scholar]
- 38.Pitscheider M, Sieber SA. Cinnamic aldehyde derived probes for the active site labelling of pathogenesis associated enzymes. Chem Commun (Camb) 2009:3741–3743. doi: 10.1039/b905527d. [DOI] [PubMed] [Google Scholar]
- 39.Cheng Q, Li H, Merdek K, Park JT. Molecular characterization of the beta-N-acetylglucosaminidase of Escherichia coli and its role in cell wall recycling. J Bacteriol. 2000;182:4836–4840. doi: 10.1128/jb.182.17.4836-4840.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Votsch W, Templin MF. Characterization of a beta -N-acetylglucosaminidase of Escherichia coli and elucidation of its role in muropeptide recycling and beta -lactamase induction. J Biol Chem. 2000;275:39032–39038. doi: 10.1074/jbc.M004797200. [DOI] [PubMed] [Google Scholar]
- 41.Stubbs KA, Scaffidi A, Debowski AW, Mark BL, Stick RV, Vocadlo DJ. Synthesis and use of mechanism-based protein-profiling probes for retaining beta-D-glucosaminidases facilitate identification of Pseudomonas aeruginosa NagZ. J Am Chem Soc. 2008;130:327–335. doi: 10.1021/ja0763605. [DOI] [PubMed] [Google Scholar]
- 42.Asgarali A, Stubbs KA, Oliver A, Vocadlo DJ, Mark BL. Inactivation of the glycoside hydrolase NagZ attenuates antipseudomonal beta-lactam resistance in Pseudomonas aeruginosa. Antimicrob Agents Chemother. 2009;53:2274–2282. doi: 10.1128/AAC.01617-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Guo Y, Nguyen KA, Potempa J. Dichotomy of gingipains action as virulence factors: from cleaving substrates with the precision of a surgeon's knife to a meat chopper-like brutal degradation of proteins. Periodontol 2000. 2010;54:15–44. doi: 10.1111/j.1600-0757.2010.00377.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Mikolajczyk J, Boatright KM, Stennicke HR, Nazif T, Potempa J, Bogyo M, Salvesen GS. Sequential autolytic processing activates the zymogen of Arg-gingipain. J Biol Chem. 2003;278:10458–10464. doi: 10.1074/jbc.M210564200. [DOI] [PubMed] [Google Scholar]
- 45.Sztukowska M, Veillard F, Potempa B, Bogyo M, Enghild JJ, Thogersen IB, Nguyen KA, Potempa J. Disruption of gingipain oligomerization into non-covalent cell-surface attached complexes. Biol Chem. 2012;393:971–977. doi: 10.1515/hsz-2012-0175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Shen A. Clostridium difficile toxins: mediators of inflammation. J Innate Immun. 2012;4:149–158. doi: 10.1159/000332946. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Puri AW, Lupardus PJ, Deu E, Albrow VE, Garcia KC, Bogyo M, Shen A. Rational design of inhibitors and activity-based probes targeting Clostridium difficile virulence factor TcdB. Chem Biol. 2010;17:1201–1211. doi: 10.1016/j.chembiol.2010.09.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Shen A, Lupardus PJ, Gersch MM, Puri AW, Albrow VE, Garcia KC, Bogyo M. Defining an allosteric circuit in the cysteine protease domain of Clostridium difficile toxins. Nat Struct Mol Biol. 2011;18:364–371. doi: 10.1038/nsmb.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Lanis JM, Hightower LD, Shen A, Ballard JD. TcdB from hypervirulent Clostridium difficile exhibits increased efficiency of autoprocessing. Mol Microbiol. 2012;84:66–76. doi: 10.1111/j.1365-2958.2012.08009.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Lanis JM, Barua S, Ballard JD. Variations in TcdB activity and the hypervirulence of emerging strains of Clostridium difficile. PLoS Pathog. 2010;6 doi: 10.1371/journal.ppat.1001061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Lamkanfi M. Emerging inflammasome effector mechanisms. Nat Rev Immunol. 2011;11:213–220. doi: 10.1038/nri2936. [DOI] [PubMed] [Google Scholar]
- 52.Turk B, Turk V. Lysosomes as “suicide bags” in cell death: myth or reality? J Biol Chem. 2009;284:21783–21787. doi: 10.1074/jbc.R109.023820. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Hansen KK, Sherman PM, Cellars L, Andrade-Gordon P, Pan Z, Baruch A, Wallace JL, Hollenberg MD, Vergnolle N. A major role for proteolytic activity and proteinase-activated receptor-2 in the pathogenesis of infectious colitis. Proc Natl Acad Sci U S A. 2005;102:8363–8368. doi: 10.1073/pnas.0409535102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Miao EA, Leaf IA, Treuting PM, Mao DP, Dors M, Sarkar A, Warren SE, Wewers MD, Aderem A. Caspase-1-induced pyroptosis is an innate immune effector mechanism against intracellular bacteria. Nat Immunol. 2010;11:1136–1142. doi: 10.1038/ni.1960. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Dupont N, Lacas-Gervais S, Bertout J, Paz I, Freche B, Van Nhieu GT, van der Goot FG, Sansonetti PJ, Lafont F. Shigella phagocytic vacuolar membrane remnants participate in the cellular response to pathogen invasion and are regulated by autophagy. Cell Host Microbe. 2009;6:137–149. doi: 10.1016/j.chom.2009.07.005. [DOI] [PubMed] [Google Scholar]
- 56.Olivares-Zavaleta N, Carmody A, Messer R, Whitmire WM, Caldwell HD. Chlamydia pneumoniae inhibits activated human T lymphocyte proliferation by the induction of apoptotic and pyroptotic pathways. J Immunol. 2011;186:7120–7126. doi: 10.4049/jimmunol.1100393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Danelishvili L, Everman JL, McNamara MJ, Bermudez LE. Inhibition of the Plasma-Membrane-Associated Serine Protease Cathepsin G by Mycobacterium tuberculosis Rv3364c Suppresses Caspase-1 and Pyroptosis in Macrophages. Front Microbiol. 2012;2:281. doi: 10.3389/fmicb.2011.00281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Hitzler I, Sayi A, Kohler E, Engler DB, Koch KN, Hardt WD, Muller A. Caspase-1 has both proinflammatory and regulatory properties in Helicobacter infections, which are differentially mediated by its substrates IL-1beta and IL-18. J Immunol. 2012;188:3594–3602. doi: 10.4049/jimmunol.1103212. [DOI] [PubMed] [Google Scholar]
- 59.Broz P, Newton K, Lamkanfi M, Mariathasan S, Dixit VM, Monack DM. Redundant roles for inflammasome receptors NLRP3 and NLRC4 in host defense against Salmonella. J Exp Med. 2010;207:1745–1755. doi: 10.1084/jem.20100257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Broz P, von Moltke J, Jones JW, Vance RE, Monack DM. Differential requirement for Caspase-1 autoproteolysis in pathogen-induced cell death and cytokine processing. Cell Host Microbe. 2010;8:471–483. doi: 10.1016/j.chom.2010.11.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Zhang Y, Bliska JB. Role of Toll-like receptor signaling in the apoptotic response of macrophages to Yersinia infection. Infect Immun. 2003;71:1513–1519. doi: 10.1128/IAI.71.3.1513-1519.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Brodsky IE, Palm NW, Sadanand S, Ryndak MB, Sutterwala FS, Flavell RA, Bliska JB, Medzhitov R. A Yersinia effector protein promotes virulence by preventing inflammasome recognition of the type III secretion system. Cell Host Microbe. 2010;7:376–387. doi: 10.1016/j.chom.2010.04.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Schotte P, Denecker G, Van Den Broeke A, Vandenabeele P, Cornelis GR, Beyaert R. Targeting Rac1 by the Yersinia effector protein YopE inhibits caspase-1-mediated maturation and release of interleukin-1beta. J Biol Chem. 2004;279:25134–25142. doi: 10.1074/jbc.M401245200. [DOI] [PubMed] [Google Scholar]
- 64.Bergsbaken T, Cookson BT. Macrophage activation redirects Yersinia-infected host cell death from apoptosis to caspase-1-dependent pyroptosis. PLoS Pathog. 2007;3:e161. doi: 10.1371/journal.ppat.0030161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Lilo S, Zheng Y, Bliska JB. Caspase-1 activation in macrophages infected with Yersinia pestis KIM requires the type III secretion system effector YopJ. Infect Immun. 2008;76:3911–3923. doi: 10.1128/IAI.01695-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Zheng Y, Lilo S, Brodsky IE, Zhang Y, Medzhitov R, Marcu KB, Bliska JB. A Yersinia effector with enhanced inhibitory activity on the NF-kappaB pathway activates the NLRP3/ASC/caspase-1 inflammasome in macrophages. PLoS Pathog. 2011;7:e1002026. doi: 10.1371/journal.ppat.1002026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Gurcel L, Abrami L, Girardin S, Tschopp J, van der Goot FG. Caspase-1 activation of lipid metabolic pathways in response to bacterial pore-forming toxins promotes cell survival. Cell. 2006;126:1135–1145. doi: 10.1016/j.cell.2006.07.033. [DOI] [PubMed] [Google Scholar]
- 68.Berger AB, Sexton KB, Bogyo M. Commonly used caspase inhibitors designed based on substrate specificity profiles lack selectivity. Cell Res. 2006;16:961–963. doi: 10.1038/sj.cr.7310112. [DOI] [PubMed] [Google Scholar]
- 69.Darzynkiewicz Z, Pozarowski P. All that glitters is not gold: all that FLICA binds is not caspase. A caution in data interpretation--and new opportunities. Cytometry A. 2007;71:536–537. doi: 10.1002/cyto.a.20425. [DOI] [PubMed] [Google Scholar]
- 70.Schotte P, Declercq W, Van Huffel S, Vandenabeele P, Beyaert R. Non-specific effects of methyl ketone peptide inhibitors of caspases. FEBS Lett. 1999;442:117–121. doi: 10.1016/s0014-5793(98)01640-8. [DOI] [PubMed] [Google Scholar]
- 71.Zamboni DS, Kobayashi KS, Kohlsdorf T, Ogura Y, Long EM, Vance RE, Kuida K, Mariathasan S, Dixit VM, Flavell RA, Dietrich WF, Roy CR. The Birc1e cytosolic pattern-recognition receptor contributes to the detection and control of Legionella pneumophila infection. Nat Immunol. 2006;7:318–325. doi: 10.1038/ni1305. [DOI] [PubMed] [Google Scholar]
- 72.Lamkanfi M, Amer A, Kanneganti TD, Munoz-Planillo R, Chen G, Vandenabeele P, Fortier A, Gros P, Nunez G. The Nod-like receptor family member Naip5/Birc1e restricts Legionella pneumophila growth independently of caspase-1 activation. J Immunol. 2007;178:8022–8027. doi: 10.4049/jimmunol.178.12.8022. [DOI] [PubMed] [Google Scholar]
- 73.Lightfield KL, Persson J, Brubaker SW, Witte CE, von Moltke J, Dunipace EA, Henry T, Sun YH, Cado D, Dietrich WF, Monack DM, Tsolis RM, Vance RE. Critical function for Naip5 in inflammasome activation by a conserved carboxy-terminal domain of flagellin. Nat Immunol. 2008;9:1171–1178. doi: 10.1038/ni.1646. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Kofoed EM, Vance RE. Innate immune recognition of bacterial ligands by NAIPs determines inflammasome specificity. Nature. 2011;477:592–595. doi: 10.1038/nature10394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Thornberry NA, Rano TA, Peterson EP, Rasper DM, Timkey T, Garcia-Calvo M, Houtzager VM, Nordstrom PA, Roy S, Vaillancourt JP, Chapman KT, Nicholson DW. A combinatorial approach defines specificities of members of the caspase family and granzyme B. Functional relationships established for key mediators of apoptosis. J Biol Chem. 1997;272:17907–17911. doi: 10.1074/jbc.272.29.17907. [DOI] [PubMed] [Google Scholar]
- 76.Nikaido H, Pages JM. Broad-specificity efflux pumps and their role in multidrug resistance of Gram-negative bacteria. FEMS microbiology reviews. 2012;36:340–363. doi: 10.1111/j.1574-6976.2011.00290.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Nikaido H. Multidrug resistance in bacteria. Annual review of biochemistry. 2009;78:119–146. doi: 10.1146/annurev.biochem.78.082907.145923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Wilke KE, Francis S, Carlson EE. Activity-based probe for histidine kinase signaling. J Am Chem Soc. 2012;134:9150–9153. doi: 10.1021/ja3041702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Wu HJ, Wang AH, Jennings MP. Discovery of virulence factors of pathogenic bacteria. Curr Opin Chem Biol. 2008;12:93–101. doi: 10.1016/j.cbpa.2008.01.023. [DOI] [PubMed] [Google Scholar]
- 80.Weerapana E, Wang C, Simon GM, Richter F, Khare S, Dillon MB, Bachovchin DA, Mowen K, Baker D, Cravatt BF. Quantitative reactivity profiling predicts functional cysteines in proteomes. Nature. 2010;468:790–795. doi: 10.1038/nature09472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Puri AW, Bogyo M. Using small molecules to dissect mechanisms of microbial pathogenesis. ACS Chem Biol. 2009;4:603–616. doi: 10.1021/cb9001409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 82.Bishop AC, Ubersax JA, Petsch DT, Matheos DP, Gray NS, Blethrow J, Shimizu E, Tsien JZ, Schultz PG, Rose MD, Wood JL, Morgan DO, Shokat KM. A chemical switch for inhibitor-sensitive alleles of any protein kinase. Nature. 2000;407:395–401. doi: 10.1038/35030148. [DOI] [PubMed] [Google Scholar]
- 83.Morell M, Nguyen Duc T, Willis AL, Syed S, Lee J, Deu E, Deng Y, Xiao J, Turk BE, Jessen JR, Weiss SJ, Bogyo M. Coupling protein engineering with probe design to inhibit and image matrix metalloproteinases with controlled specificity. J Am Chem Soc. 2013;135:9139–9148. doi: 10.1021/ja403523p. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 84.Xiao J, Broz P, Puri AW, Deu E, Morell M, Monack DM, Bogyo M. A coupled protein and probe engineering approach for selective inhibition and activity-based probe labeling of the caspases. J Am Chem Soc. 2013;135:9130–9138. doi: 10.1021/ja403521u. [DOI] [PMC free article] [PubMed] [Google Scholar]







