Significance
Current models propose that in eukaryotes the expression of a gene and its ability to recombine depend on whether regulatory factors or enzymes are present in the same subnuclear compartment as the gene. Nucleoprotein complexes might thereby modulate expression or recombination by conditionally tethering the regions they control to subnuclear compartments. Our research examines this type of spatial effect in fission yeast, showing how an ectopic ribosomal DNA repeat silences the chromosomal region in which it is integrated by relocalizing it to the perinucleolar space. Silencing of the relocalized region proceeds through heterochromatin formation and a redundant mechanism, plausibly antisense transcription. These observations establish fission yeast as a model system to investigate perinucleolar silencing, an evolutionarily conserved but poorly understood phenomenon.
Keywords: nuclear organization, gene silencing, chromatin, boundary elements, antisense transcription
Abstract
Our research addresses the relationship between subnuclear localization and gene expression in fission yeast. We observed the relocalization of a heterochromatic region, the mating-type region, from its natural location at the spindle-pole body to the immediate vicinity of the nucleolus. Relocalization occurred in response to a DNA rearrangement replacing a boundary element (IR-R) with a ribosomal DNA repeat (rDNA-R). Gene expression was strongly silenced in the relocalized mating-type region through mechanisms that differ from those operating in wild type. Also different from the wild-type situation, programmed recombination events failed to take place in the rDNA-R mutant. Increased silencing and perinucleolar localization depended on Reb1, a DNA-binding protein with cognate sites in the rDNA. Reb1 was recently shown to mediate long-range interchromosomal interactions in the nucleus through dimerization, providing a mechanism for the observed relocalization. Replacing the full rDNA repeat with Reb1-binding sites, and using mutants lacking the histone H3K9 methyltransferase Clr4, indicated that the relocalized region was silenced redundantly by heterochromatin and another mechanism, plausibly antisense transcription, achieving a high degree of repression in the rDNA-R strain.
Eukaryotic genomes are spatially organized within the nucleus (reviewed in refs. 1–5). In interphase, chromatin domains and entire chromosomes tend to occupy defined positions relative to each other or to nuclear landmarks. Chromosomal regions capable of associating with the nuclear periphery, with nuclear substructures, or with each other have been proposed to function as anchors underlying this organization (4–7). Coupled with a nonuniform distribution of proteins in the nucleus, the dynamic, directed localization of chromosomal regions to subnuclear compartments might influence and perhaps regulate gene expression and recombination. Both stable associations and transient contacts might impose lasting marks on the regions that participate in them (8, 9). Despite our increasingly detailed view of subnuclear organization, however, the two fundamental questions of how the organization is achieved and of its physiological significance remain largely open.
In the fission yeast Schizosaccharomyces pombe, extended silent heterochromatic domains are found at the three centromeres, at the telomeres of chromosome 1 and 2, and in the right arm of chromosome 2, in the mating-type region (10–16). Large clusters of rDNA repeats at both ends of chromosome 3 are also partially heterochromatic (13, 16). In addition to their distinctive chromatin structures, these regions display distinctive subnuclear localizations. The fission yeast nucleus is structured (17–23), and the nuclear proteome is not uniformally distributed (24). One pole of the interphase nucleus is defined by the microtubule organizing center or spindle-pole body (SPB) spanning the nuclear envelope. The three centromeres and mating-type region cluster at the SPB (17, 22). Telomeres are also at the nuclear periphery, although not associated with the SPB (17). The rDNA repeats merge opposite to the SPB to form the nucleolus, where rDNA genes are transcribed and ribosomes preassembled (25). Much has been learned regarding the biogenesis and composition of S. pombe chromatin (26), but with few exceptions (20–24, 27–29), the molecular processes discovered to date have not been studied in this spatial context.
We gained insight into effects exerted through the spatial organization of chromatin in S. pombe by searching for genomic elements capable of restoring silencing in a fragilized mating-type region. Heterochromatin in the mating-type region spans ∼20 kb. It encompasses two silent cassettes used for mating-type switching, mat2-P and mat3-M (Fig. 1A). Nucleosomes in the heterochromatic domain are methylated at histone H3 lysine 9 (H3K9me), and they are associated with chromodomain proteins and histone deacetylases, whereas nucleosomes in the flanking domains are predominantly methylated at histone H3 lysine 4 (H3K4me) and globally more acetylated (15, 30). Heterochromatin integrity depends on two boundary elements, the inverted repeats IR-L and IR-R (14). Although genes naturally or artificially placed between the two repeats are silenced in strains where the inverted repeats are intact, silencing is alleviated following deletion of one or the other repeat (14). Other reports on IR-L and IR-R (15, 21) drew the opposite conclusion, that the two elements function to block spreading of heterochromatin, but these studies were conducted in strains overexpressing Swi6 (triple gene dosage; swi6+-333 allele). In a wild-type background, IR-L and IR-R rather set a sharp transition between euchromatin and heterochromatin, and major effects of single or combined deletions are to alleviate silencing in the normally heterochromatic region (14). By searching for elements capable of restoring silencing in a strain lacking the IR-R boundary, we found that an rDNA repeat could strongly repress the mating-type region. The rDNA repeat caused a relocalization of the region to the perinucleolar space, indicating that distinct subnuclear environments can substitute for each other to silence gene expression. Relocalization of the region and its silencing depended on Reb1, a myb-domain DNA-binding protein capable of mediating physical interactions between chromosomal domains in other contexts (31). Mutational analysis, and the phenotype of cells in which the full rDNA repeat was replaced with binding sites for Reb1, lead us to propose that Reb1 tethers the rearranged mating-type region to the nucleolus, thereby causing its silencing through heterochromatin formation combined with a redundant mechanism that we suggest might be antisense transcription.
Fig. 1.
Inhibition of gene expression and recombination by an rDNA repeat. (A) Mating-type region. The mat2-P and mat3-M cassettes are in a ∼20-kb heterochromatic region flanked by the IR-L and IR-R boundary elements. They are silenced and used for gene conversions of mat1 leading to mating-type switching. The IR-R boundary was replaced with an rDNA repeat in rDNA-R strains. rDNA repeats consist of a transcribed region encoding the 18S, 5.8S, and 28S rRNA, and an untranscribed spacer containing an origin of replication (autonomously-replicating sequence, ars) and binding sites for the Reb1 and Sap1 proteins that block replication fork progression. (B) Tenfold serial dilutions of (EcoRV)::ade6+ cells with a wild-type boundary (IR-R+, MAM56), a deletion of the boundary (IR-RΔ, MAM46), or rDNA-R (MAM36) were spotted on selective media. Cells lacking (EcoRV)::ade6+ are shown for comparison [IR-R+(WT), MAM26]. Cells expressing ade6+ grow on AA-ade and form white colonies on yeast extract (YE) plates. (C) (EcoRV)::ade6+ expression measured by quantitative RT-PCR relative to the expression of ade6+ at its normal chromosomal location (IR-R+, PM7; IR-RΔ, PM3; rDNA-R, PM2). (D) Mating-type switching was assayed by iodine staining of colonies grown on minimum sporulation agar (MSA). Dark staining is indicative of efficient switching because S. pombe spores, but not vegetative cells, are stained brown by iodine vapors. Light staining of rDNA-R colonies reveals a mating-type switching defect. Strains as in B. (E) Southern blot of genomic DNA digested with HindIII and probed with a mat1 probe (10.5-kb HindIII fragment). The fragile site at mat1 gives rise to a double strand break (DSB). (Left) Strains as in B. (Right) swi1Δ strains (IR-R+, PM18; rDNA-R, PM20). (F) Quantification of mat1 content by PCR. From left to right: IR-R+, MAM56; IR-RΔ, MAM46; rDNA-R, MAM36; and IR-R+(WT), MAM26.
Results and Discussion
A Search for Boundary Elements Identifies an rDNA Repeat.
The mating-type region, depicted in Fig. 1A, contains one of the best characterized heterochromatic regions of fission yeast. Here, we used an ade6+ gene inserted near the mat3-M mating-type cassette [(EcoRV)::ade6+ in Fig. 1 A–C] (32). Cells with this reporter are predominantly Ade−, they produce red colonies when starved for adenine and barely detectable ade6+ transcripts due to heterochromatic silencing. Deleting the IR-R boundary (IR-RΔ) increased expression of (EcoRV)::ade6+ (Fig. 1 B and C). IR-RΔ cells formed white, Ade+ colonies, and their steady-state ade6+ transcript level was increased >20-fold compared with IR-R+ cells. This increased transcript level corresponds to approximately half the expression level of the endogenous ade6+ locus under the same conditions (Fig. 1C). Alleviated repression of (EcoRV)::ade6+ in IR-RΔ cells reflects the essential function of the boundary in protecting the repressed domain from encroachment by the flanking euchromatin (14).
We conducted a genomic screen for S. pombe DNA fragments capable of functionally replacing IR-R. Three libraries of S. pombe genomic DNA were introduced in the place of IR-R in the chromosome of (EcoRV)::ade6+ IR-RΔ cells. Most DNA integrations had no effect on ade6+ expression, producing white Ade+ transformants. A small proportion of red, Ade− transformants were also obtained, indicating that the fragment of genomic DNA inserted in these transformants repressed ade6+. Inserts were recovered from the latter class of transformants by PCR amplification and retested individually through a second round of chromosomal integration in the (EcoRV)::ade6+ IR-RΔ tester strain. Fragments reproducibly restoring ade6+ silencing were sequenced. One class of elements identified in this screen was rDNA repeats (Fig. 1).
The rDNA-R Boundary Inhibits both Gene Expression and Programmed DNA Rearrangments.
Fission yeast rDNA repeats consist of a 7.9-kb transcription unit and 3-kb intergenic spacer (Fig. 1A). The transcribed region is the template for the 28S, 5.8S, and 18S ribosomal RNAs synthesized as a long precursor by RNA Pol I. The spacer comprises a promoter region, an origin of replication, and replication fork barriers that ensure unidirectional replication of the repeats in the same direction as transcription (Fig. 1A) (33–35). A full 10.9-kb rDNA repeat, hereafter referred to as rDNA-R, repressed (EcoRV)::ade6+ tightly as judged both from a plating assay (Fig. 1B) and transcript quantification by RT-PCR (Fig. 1C). Curiously, more (EcoRV)::ade6+ transcript was detected in rDNA-R cells than in IR-R+ (Fig. 1C) cells even though repression by rDNA-R appeared tighter on plates than repression by IR-R+ (Fig. 1B). These different phenotypes of IR-R+ and rDNA-R might result from different transcript distributions in the cell populations, perhaps reflecting different modes of inheritance of expressed and repressed states in IR-R+ and rDNA-R cells. Alternatively, the (EcoRV)::ade6+ transcript detected by RT-PCR in rDNA-R cells might not be functional, as discussed further down.
Programmed recombination events in which genetic information is unidirectionally transferred from mat2-P or mat3-M to the expressed mat1 locus take place in wild-type cells, leading to mating-type switching. rDNA-R cells displayed a mating-type switching defect, causing reduced iodine staining on sporulation plates (Fig. 1D). This differs from IR-RΔ cells, where no defect was seen (Fig. 1D). Switching is initiated at a fragile site at mat1 that can be detected as a double-strand break (DSB) in chromosomal DNA preparations (36). The fragile site was detected by Southern blot in rDNA-R cells, indicating that its formation is not the primary defect leading to poor switching (Fig. 1E). Furthermore, the fragile site depended on the replication fork protection complex, as it does for wild-type and IR-RΔ cells (Swi1 and Swi3 proteins; Fig. 1E). In wild-type cells, the fragile site is converted to a DSB during DNA replication. It directs synthesis-dependent strand invasion of one or the other silent cassette cells. This process requires recombination proteins and directionality factors selecting the silent cassette that contains the information opposite to the information present at mat1 (37, 38). The viability of rDNA-R cells assayed by pedigree analysis was similar to wild type, showing that cells do not die as they attempt to switch. Donor choice, however, was strongly biased toward mat2-P in rDNA-R cells, whereas no bias is seen in IR-R+ or IR-RΔ cells (Fig. 1 D and F). Because use of mat3-M is favored by a recombination enhancer, the Swi2-dependent recombination enhancer SRE adjacent to mat3-M (38), this asymmetry suggests that rDNA-R blocks the association of cognate recombination factors with SRE, preventing use of mat3-M in P cells. This defect points to further differences between the action of rDNA-R and the action of IR-R in the mating-type region: rDNA-R inhibits both gene expression and recombination, whereas IR-R inhibits gene expression while permitting recombination.
The rDNA-R Boundary Causes a Relocalization of the Mating-Type Region to the Perinucleolar Space.
Several studies have indicated that transcription and recombination obey special rules in the rDNA of eukaryotes, distinct from the bulk of the genome. Genes transcribed by PolI or PolII can be epigenetically silenced (13, 39–41), and some forms of recombination are also inhibited (42–47). In higher eukaryotes, the perinucleolar space is associated with heterochromatin (1, 4, 5, 48–50) and poorly expressed genes (7, 8). There is, furthermore, evidence that rDNA repeats are prone to clustering in S. pombe because the two rDNA arrays at the ends of chromosome 3 merge in the nucleolus (25) similar to nucleolar fusion in higher eukaryotes (50, 51). These observations taken together suggest that rDNA-R might exert its effects in the mating-type region by changing the subnuclear localization of the mating-type region from its natural location at the SPB to the nucleolus. We tested this hypothesis by using a chromosomal LacO array integrated at his2, a locus at 24 kb from the mating-type region, in strains expressing GFP-LacI (52). The mating-type region can be imaged in these cells as a fluorescent dot. We assayed the localization of the mating-type region relative to the nucleolus by coexpressing a nucleolar protein tagged with cyan fluorescent protein (CFP) (SPBC947.07) (24) and measuring the distance between the mating-type region (GFP dot) and the center of the nucleolus in 3D. Representative images and histograms of the measured distances are shown in Fig. 2 C–F. We found that the mating-type region in rDNA-R cells occupied a volume at the periphery of the nucleolus. This stands in contrast to IR-R+ or IR-RΔ cells in which the mating-type region was near the nuclear envelope diametrically opposed to the nucleolus (Fig. 2 C and D). Localization of the mating-type region at the nucleolus in rDNA-R cells indicates that factors normally facilitating the clustering of wild-type rDNA repeats attract rDNA-R to the nucleolus.
Fig. 2.
Relocalization of the mating-type region to the nucleolus in rDNA-R cells. (A) Dimensions of S. pombe cells. The distance d between the mating-type region (yellow) and the center of the nucleolus (blue) was measured in this experiment. (B) The S. pombe cell cycle. G2 cells (highlighted in blue) were used for distance measurements. (C−F) Fluorescence images (Left) and distance measurements (Right) for strains with the indicated genotypes. The distance d was determined in 3D for the reported number of cells. Mean values (d̅) and SDs are indicated. The IR-R+(WT) strain shown in C has a wild-type mating-type region; the strains shown in D−F have an (EcoRV)::ade6+ insertion. A Student’s t test showed that the mating-type region is significantly closer to the nucleolus in the rDNA-R mutant than in IR-R+ or IR-RΔ strains; P values comparing each distribution to C are indicated in D−F.
Relocalization of the Mating-Type Region to the Perinucleolar Space and Its Silencing Are Mediated by Reb1.
Reb1 is a myb-domain protein that binds at two sites in the fission yeast rDNA intergenic regions to facilitate transcription termination and to block replication fork progression (34, 35, 53, 54) (Fig. 1A). A recent study proposed that Reb1 can bring together unlinked chromosomal loci displaying cognate binding sites through dimerization (31). Here, deleting reb1 abrogated the tight association of the rDNA-R mating-type region with the nucleolus (Fig. 3A). In fluorescence images, the mating-type region was clearly dissociated from the nucleolus in a fraction of the cell population and remained close to the nucleolus in the remaining cells. Consistent with two populations, the mating-type region to nucleolus distance distribution appeared bimodal and could be fitted by a double Gaussian (Fig. 3A). The maxima of the two rDNA-R reb1Δ distributions have values close to the maximum for IR-R+ cells and rDNA-R cells, around 1.25 µm and 0.80 µm, respectively (Figs. 3A and 2E and F). These observations point to an important role for Reb1 in tethering the rDNA-R mating-type region to the nucleolus, and they indicate that additional factors facilitate perinucleolar association in the absence of Reb1, albeit inefficiently. The DNA-binding protein Sap1, with a binding site close to the Reb1 binding sites in the rDNA (Fig. 1A), might facilitate the association.
Fig. 3.
Reb1 mediates the relocalization of the rDNA-R mating-type region to the nucleolus and its silencing. (A) Distribution of the mating-type region to nucleolus distance d for (EcoRV)::ade6+ cells propagated in the presence or absence of adenine as indicated. The red dotted lines correspond to double Gaussian fits (mean values d̅1 and d̅2) indicating the presence of two subpopulations of cells differentially associated with the nucleolus. (B) Deletion of reb1 derepresses (EcoRV)::ade6+ in rDNA-R but not in IR-R+ cells. Spot tests as in Fig. 1B. From top to bottom: PM8, PM107, PM53, PM59, PM67, and PM51. (C) ade6+ expression measured by quantitative RT-PCR as in Fig. 1C. WT ade6+, 968; WT ade6+ reb1Δ, PG3772; IR-R+, PM8; IR-R+ reb1Δ, PM107 and PM108; rDNA-R, PM59; rDNA-R reb1Δ, PM67-69. (D) Relative localization of Reb1 (tagged with mCherry and expressed from endogenous locus; shown in red) and mating-type region (GFP; shown in green) in IR-R+ (PM131), IR-RΔ (PM132), and rDNA-R (PM127) cells. Reb1 and the mating-type region colocalize in a fraction of the rDNA-R cell population.
The second phenotype observed in reb1Δ cells was a derepression of (EcoRV)::ade6+. Cells grew in the absence of adenine (Fig. 3B), and increased ade6+ transcript levels were detected (Fig. 3C). As for the distance distributions, the growth assay indicated that only a fraction of the population was affected by the deletion of reb1 (Fig. 3B). We verified that Reb1 does not regulate ade6+ at its normal chromosomal location (Fig. 3C). We specifically assayed the localization of the mating-type region in cells expressing (EcoRV)::ade6+ by propagating reb1Δ cells in medium lacking adenine before microscopy. This selection enriched for cells in which the mating-type region was away from the nucleolus (Fig. 3A), and it was, as expected, accompanied by an increase in the expression level of ade6+ (Fig. 3C). This confirms that two cell populations exist in the rDNA-R reb1Δ mutant, cells in which the mating-type region is expressed and not associated with the nucleolus and cells in which the mating-type region remains repressed and associated with the nucleolus.
Antisense Transcription in the rDNA-R Mating-Type Region.
Our initial observations—that an rDNA repeat inserted at the edge of the mating-type region causes the relocalization of the mating-type region from the SPB to the vicinity of the nucleolus, that the rDNA repeat silences the mating-type region, and that the rDNA repeat inhibits directed recombination—do not on their own imply causal relationships between the three phenomena. Parameters other than localization might be affected by rDNA-R, which would in turn affect gene expression or recombination. Investigating this possibility, we detected an ade6+ antisense transcript in rDNA-R cells extending toward centromere 2 past (EcoRV)::ade6+ and mat3-M (Fig. 4). The levels of sense and antisense ade6+ transcript were not found to be anticorrelated (Fig. 4C). This indicates that antisense transcription is not directly responsible for shutting off ade6+ even though more complex mechanisms cannot be excluded (discussed below). One can also note that antisense transcripts have been detected at multiple loci in fission yeast genomes, but that anticorrelation with sense transcription was not generally observed (55). On the other hand, the fact that Reb1, a DNA-binding protein with sites in the rDNA, is required for both relocalization of the mating-type region to the nucleolus and its silencing indicates relocalization and silencing proceed through a common mechanism involving Reb1 bound to the rDNA repeat. In this context, the strict correlation between proximity of the mating-type region to the nucleolus and ade6+ repression is most simply explained by a model where proximity to the nucleolus causes ade6+ repression.
Fig. 4.
Antisense transcription in the mating-type region of rDNA-R cells. (A) The positions of seven primer pairs used in B and C are indicated above the mating-type region. (B) Antisense transcription (i.e., polarity opposite to ade6+) was detected by real-time RT-PCR at the seven positions assayed in rDNA-R (PM59) but not in IR-R+ (PM8) cells. Transcript levels were estimated relative to act1+. (C) Sense and antisense (EcoRV)::ade6+ transcripts were measured with primer pair 1 in independent biological isolates. Three independent isolates were examined for rDNA-R reb1+, five for rDNA-R reb1Δ cells propagated in the presence of adenine, and three for rDNA-R reb1Δ cells propagated in the absence of adenine. Each RT-PCR was set up in triplicate. Transcript levels were normalized to ade6+ expressed from its normal chromosomal location. No anticorrelation between sense and antisense transcription was observed, indicating that antisense transcription is not, or not solely, responsible for changes in sense transcript abundance.
Dependency of Reb1- and rDNA-R−Mediated Silencing on Heterochromatic Factors.
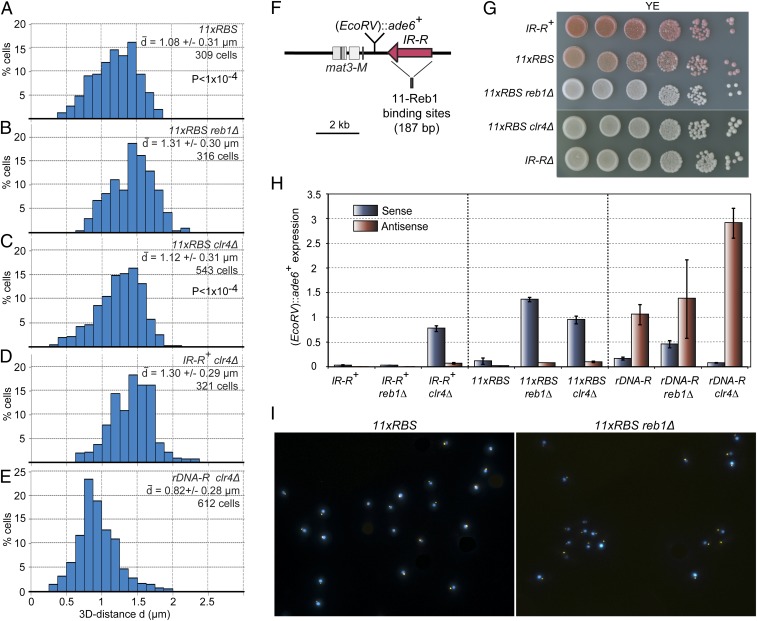
We investigated the extent to which Reb1 can relocalize and silence the mating-type region by introducing 11 Reb1-binding sites instead of rDNA-R in an IR-RΔ strain (referred to as 11xRBS below). The 11xRBS recapitulated some, but not all, effects of rDNA-R. Like rDNA-R, 11xRBS both relocalized the mating-type region to the nucleolus and repressed (EcoRV)::ade6+ (Fig. 5 and Figs. S1 and S2). Both effects required Reb1 (Fig. 5 and Fig. S1). Relocalization (compare Fig. 2E with Fig. 5A) and silencing (see Fig. S2 for growth on selective plates) by 11xRBS were not as tight as for rDNA-R, consistent with other factors acting at rDNA-R in addition to Reb1. Two major differences between 11xRBS and rDNA-R were, first, that no (EcoRV)::ade6+ antisense transcript was detected in the 11xRBS strain (Fig. 5H and Fig. S1) and, second, that silencing by 11xRBS required the histone H3K9 methyltransferase Clr4 (Fig. 5 G and H and Figs. S1 and S2) whereas silencing by rDNA-R did not (Figs. 3B and 5H and Fig. S2). For both 11xRBS and rDNA-R, the association of the mating-type region with the nucleolus remained unchanged in the absence of Clr4 (Fig. 5 C and E). This is consistent with relocalization preceding silencing. The requirement for other factors important for heterochromatin formation at the major heterochromatic loci was assayed (Fig. S2). None of the mutations tested reduced repression by rDNA-R (clr3Δ; sir2Δ; ago1Δ; rdp1Δ) whereas deletion of genes coding for histone deacetylases (sir2Δ; clr3Δ) abrogated silencing by 11xRBS as it does for IR-R+ cells (Fig. S2).
Fig. 5.
Relocalization and repression of the mating-type region by Reb1-binding sites. (A−E) Distribution of the mating-type region to nucleolus distance d for strains with the indicated genotypes. The mating-type region was associated with the nucleolus in a fraction of the 11xRBS (A) and 11xRBS clr4Δ (C) populations, leading to shorter mean distances than for 11xRBS reb1Δ cells (B) where the association was lost. The nucleolar association of rDNA-R remained in clr4Δ (E). Student t tests showed that the distributions in A and B were statistically different from each other (P < 1 × 10−4), as were the distributions in B and C (P < 1 × 10−4), whereas A and C were not (P = 0.05). The strains were, from A to E, TP360, TP361, TP362, PM16, and PM14. (F) Schema depicting the replacement of the IR-R+ boundary with 11 Reb1-binding sites (11xRBS). (G) The 11xRBS represses (EcoRV)::ade6+ in a Reb1- and Clr4-dependent manner. Spot tests as in Fig. 1B with, from top to bottom, PM8, TP360, TP361, TP362, and PM3. (H) (EcoRV)::ade6+ sense and antisense transcripts measured by quantitative RT-PCR normalized to euchromatic ade6+ sense expression as in Fig. 1C. The 11xRBS, but not rDNA-R, required the methyltransferase Clr4 for its silencing effects, and an antisense (EcoRV)::ade6+ transcript was abundant in rDNA-R cells, but not in 11xRBS cells. The means of several biological isolates are presented for each genotype. Quantification for each individual isolate is presented in Fig. S1. Strains, from left to right: PM8, PM107-108, PM31, TP360, TP361, TP362, PM59, PM67-69, and PM27. (I) Example of fluorescence images used for quantification in A and B.
Collectively, these phenotypes strongly support our hypothesis that Reb1 bound to cognate sites triggers the association of the mating-type region with the nucleolus and its silencing (Fig. 6). This would occur in both 11xRBS and rDNA-R cells. We previously found that Clr4 and other chromodomain proteins impose a variegated repression on PolII-transcribed genes inserted into the native rDNA arrays (13). These factors might strengthen heterochromatin formation when the 11xRBS or rDNA-R mating-type region pairs with the rDNA. Other nucleolar proteins such as Grc3, which both catalyzes rRNA processing in the nucleolus and heterochromatin formation in the major heterochromatic regions (27), or proximity to the nuclear exosome abundant in the nucleolus (56), might also facilitate silencing in this specific spatial context.
Fig. 6.
Model. Insertion of an rDNA repeat in the mating-type region (rDNA-R) in the place of a boundary element (IR-R+) leads to a relocalization of the region away from its natural location at the spindle-pole body to the perinucleolar compartment. Perinucleolar association is stabilized by the dimerization of the myb-domain protein Reb1. This brings the mating-type region under the influence of macromolecules predominantly abundant or active in the nucleolus resulting in gene silencing.
rDNA-R would exert a redundant repression due to runaway transcription initiated by RNA PolI within rDNA-R (Fig. 4 and Fig. S1). That antisense transcription has a role in silencing is supported by the observation that only one insert orientation of rDNA-R was recovered in the screen, the orientation in which PolI transcription is most likely to run across (EcoRV)::ade6+ (see Materials and Methods). Alleviated repression of (EcoRV)::ade6+ in rDNA-R reb1Δ cells whose mating-type region is far from the nucleolus, even in cases where these cells contain high levels of antisense ade6+ transcript (Fig. 4C), would indicate that antisense transcription, or the antisense transcript, silences most effectively near the nucleolus. Runaway transcription across mat3-M might furthermore impair switching in rDNA-R cells by disrupting interactions between the SRE element and recombination factors (38). No switching defect was observed in the 11xRBS strain (Fig. S2), where runaway transcription does not occur (Fig. 5H and Fig. S1).
Several mechanisms can be envisioned for how antisense transcription or an antisense transcript would inhibit gene expression. Collisions between RNA polymerases transcribing in opposite directions might cause premature termination; transcription by RNA PolI might be accompanied by chromatin modifications preventing initiation by RNA PolII or in other ways prevent RNA PolII transcription (57, 58); some types of RNA-DNA hybrids might promote gene silencing (59). Fig. 1 showed that more ade6+ sense transcript was detected in rDNA-R cells than in IR-R+ cells, yet rDNA-R cells grew more poorly than IR-R+ cells in the absence of adenine. The transcripts detected by RT-PCR might not be full length, or interactions with the antisense might affect their processing and prevent export of functional transcripts from the perinucleolar space to the cytoplasm.
Evolutionary Conservation of Perinucleolar Silencing.
Several lines of evidence suggest that the perinucleolar compartment functions as a repressive environment in other cell types. Nucleoli in higher eukaryotes are surrounded by a layer of heterochromatin (1, 4, 5, 48–50). In mammalian ES cell cultures, the inactive X chromosome (Xi) or autosomes expressing the Xist transcript visit the perinucleolar space in S phase (9). This localization, which requires Xist, has been proposed to facilitate the inheritance of the Xi heterochromatic state during replication by allowing the action of chromatin remodeling factors particularly abundant at the nucleolar periphery (9). Similarly, a DNA element essential for the silencing of the imprinted Kcnq1ot1 locus targets DNA to the perinucleolar space in S phase, and this localization has been suggested to participate in epigenetically silencing the locus (60, 61). Recent genomics studies have identified chromatin domains preferentially associated with the nucleolus in human cell lines (7, 8). These suggest that the periphery of the nucleolus is a transcription-poor environment that might mediate gene repression by sequestering genes away from more active environments. Consistently, artificially targeting a locus to the nucleolus of human cells with a 5S RNA sequence leads to reduced expression (62). Our observations provide an example of such spatial regulation in fission yeast.
Materials and Methods
S.pombe Strains.
Strain genotypes are listed in Table S1.
Screen for Boundary Elements.
Three libraries of S. pombe genomic DNA were created in the plasmid pGT299 and used in a genetic screen for boundary and silencing elements. pGT299 is a pBluescript SKII(-) (Agilent) derivative obtained by cloning the 2.8-kb XmnI-PstI fragment of S. pombe genomic DNA that is normally centromere-distal to IR-R+ into the filled-in XbaI -PstI sites of the polylinker, and inserting the S. cerevisiae LEU2 gene at the BlpI site of this fragment (14). S. pombe genomic DNA from the wild-type 968 strain (63) digested with SpeI, NheI, or XbaI was cloned into the SpeI site of pGT299, creating three libraries of genomic DNA. The SpeI library contained ∼7,000 clones, 50% of which had an insert with an average size of 3.2 kb. The NheI and XbaI libraries each contained ∼4,000 clones, 90% of which had inserts with respective average sizes of 7.5 kb and 3.9 kb. Double digests were set up for each library, with NotI and KpnI, and with NotI and XhoI, to release inserts suitable for chromosomal integration at the edge of the mating-type region. The digests were transformed into strain PG2897 using the lithium acetate method. PG2897 contains (EcoRV)::ade6+ (32) (Fig. 1), a deletion of the IR-R+ boundary element (IR-RΔ), and an insertion of the S. pombe ura4+ gene at the first SpeI centromere-distal to IR-RΔ (SpeI::ura4+) (14). This SpeI site is the same used to generate the three libraries in pGT299. PG2897 also contains the leu1-32 mutation, which can be complemented by S. cerevisiae LEU2. Chromosomal integration of a library insert by homologous recombination in PG2897 [mat3-M (EcoRV)::ade6+ IR-RΔ SpeI::ura4+] results in a Leu+ 5-fluoroorotic acid (FOA)-resistant transformant [mat3-M (EcoRV)::ade6+ IR-RΔ (SpeI)::insert BlpI::LEU2]. Leu+ transformants were selected in a first step on AA-leu drop-out medium and subsequently replicated onto FOA plates containing 12 mg/L adenine. Approximately 500,000 Leu+ transformants were obtained in total, of which ∼50,000 were also FOA resistant. Southern blotting revealed that integration had occurred as expected in all Leu+ FOA-resistant transformants tested (>30). Transformants in which (EcoRV)::ade6+ was expressed formed white colonies on the 12 mg/L adenine plates used, as does the parental PG2897 strain. A small fraction of transformants formed red colonies, indicating that the fragment of genomic DNA integrated in these transformants repressed (EcoRV)::ade6+. The integrated DNA and flanking regions were amplified from these transformants using the Expand long-template PCR system (Roche) with GTO-216 and GTO-217. The PCR products were cloned into PCR2.1-TOPO (Invitrogen), partially sequenced, and reintroduced into PG2897 to retest their ability to repress (EcoRV)::ade6+. These PCR products do not contain LEU2, but their integration in PG2897 displaces SpeI::ura4+, resulting in FOA resistance. A 10.9-kb SpeI fragment containing a full rDNA repeat was repeatedly obtained in the screen. Other fragments will be described elsewhere.
Taking into account the features of the SpeI library (50% inserts; 3.2-kb mean insert size), the rDNA repeat size (10.9 kb), the number of rDNA repeats in each genome (100−150), and the genome size (∼14 Mb), we estimate that rDNA repeats ought to have replaced IR-R+ in 1–2% of the SpeI library transformants. Based on the distinctive dark red, stably repressed phenotype of the transformants from which rDNA inserts were recovered, we estimate that 0.25–0.5% of the transformants contained a repressive rDNA insert. This frequency being slightly lower than calculated might reflect, first, an underrepresentation of pGT299-rDNA clones in the library due to large plasmid size (>18 kb) and, second, the fact that only one rDNA insert orientation resulted in repression. In six out of six transformants from which rDNA boundaries were recovered by long-range PCR, the rDNA insert was in the same orientation (shown in Fig. 1). Repressive rDNA inserts were not recovered from the XbaI or NheI libraries.
Construction of Strains with 11 Reb1-Binding Sites (11xRBS).
Long complementary oligonucleotides containing 10 Reb1 binding sites and protruding SpeI compatible ends (10xReb1-For and 10xReb1-Rev, Table S2) were annealed and cloned into the SpeI site of plasmid pGT299 (described above). Clones with various numbers of Reb1-binding sites were obtained. A plasmid with 11 sites (11xRBS; rDNA1 plasmid) was digested with NotI and PstI, and its insert was integrated into the chromosome of strain PG3737. PCR on chromosomal DNA of Leu+, FOA-resistant transformants, and sequencing of the PCR products confirmed that 11 sites had been integrated. One of the transformants was saved and used in genetic crosses to produce TP360, TP361, TP362, TP380, and TP381.
Quantitative Multiplex PCR.
PCR was performed as in ref. 38 to determine the ratio of mat1-M/mat1-P alleles in strains of interest. PCR products were run on 1.5% agarose gel, and ethidium bromide staining was quantified using ImageJ.
Fluorescent Labeling.
The mating-type region was labeled by a lac-Op array inserted at the tightly linked his2 locus (52). LacI-GFP was expressed in the same strains from the his7 locus under control of the dis1 promoter (52). The nucleolus was labeled as in ref. 24 by expressing a fluorescently tagged version of the ribosome biogenesis protein Rrp14-C (SPBC947.07; 08/08C03). For the purpose of our experiment, the YFP tag used by ref. 24 was replaced with CFP. The Rrp14-C-CFP fusion protein was expressed from the leu1 locus, under control of the thiamine-repressible promoter nmt1. Before microscopy, cells were propagated in liquid medium (AA with 0.25 µM thiamine or EMM2 with 0.1 µM thiamine). Reb1 was imaged by expressing an mCherry-Reb1 fusion protein from the endogenous reb1 locus using a previously made construct (34) in which we replaced GFP with mCherry.
Data Acquisition.
An Imager.Z1 microscope from Zeiss equipped with a cooled Orca-ER CCD camera (Hamamatsu) was used for the fluorescence microscopy. The illumination source was an HXP 120C lamp from Leica. All images were captured at 100-fold magnification using a Plan-Apochromat 100×, 1.4 NA objective and the acquisition program Volocity. For each field of cells, 11 fluorescent images were obtained at each of the relevant wavelengths at 0.4-µm intervals along the z axis. The fluorophores were visualized using band-pass CFP (31044 v2) and YFP (41028) filter sets from Chroma (Brattleboro, VT). Image acquisition times were 500 ms to 1 s for the GFP signal visualized through the YFP band-pass filter and 200 ms to 600 ms for the CFP signal.
Data Analysis.
Three-dimensional distances between the mating-type region and the center of the nucleolus were measured using a custom-made Matlab program. Each field imaged in the microscopy sessions contained a set of cells to analyze, for which n stacked images had been acquired in each YFP and CFP channel (here, n = 11). Following contrast enhancement, the YFP and CFP image stacks were imported into Matlab, and all images were filtered using a Difference of Gaussians filter and converted to binary images. The two stacks were processed in parallel as follows. Each mating-type region or nucleolus (object) corresponds to clustered pixels in the binary images. Clusters were labeled, at first in 2D. Each cluster was assigned an integer number, and the x and y coordinates of its centroid were determined and coupled to the intensity of the centroid pixel in the original image. Consecutive images were then compared, and a common number was assigned to all clusters representing the same object. This analysis resulted in a 3D matrix containing n−1 2D matrices, each containing information about the comparison between two consecutive images, i.e., centroid coordinates, centroid intensity, slice number, and cluster number. All clusters with the same number were compared, and the vector from the cluster with the highest intensity was saved in a new 2D matrix. Finally, the mating-type region (YFP) and nucleolus (CFP) matrices were compared. If the x and y coordinates of a YFP and CFP cluster were within a certain confidence interval, they were coupled as belonging to the same cell and the mating-type region to nucleolus distance d was calculated.
Statistical Analyses.
A Student t test assuming near-Gaussian distributions, independent samples, and a common variance was performed to determine whether the three distance distributions shown in Fig. 2 D–F (IR-RΔ, rDNA-R, and IR-R+) are significantly different from the distribution in Fig. 2C [IR-R+(WT)]. The null hypothesis tested was that the samples were drawn from populations having the same mean mating-type region to nucleolus distance. The three strains were tested one by one against IR-R+(WT) with a confidence limit of 95%, producing the P values reported near the histograms. Similarly, the distributions in Fig. 5 A–C were compared pairwise. The double Gaussian fits in Fig. 3A were obtained using Matlab’s maximum likelihood estimation function.
RNA Extraction and Real-Time PCR.
Cells were propagated in EMM2 minimal medium supplemented as needed, to OD600 ≈ 0.4, and RNA was extracted according to a previously described procedure (64). The following primers were used for transcript detection by RT-PCR: ade6+, GTO-218 and GTO-219; act1+, TJO-55 and TJO-58; and antisense transcript shown in Fig. S1: position 1, GTO-218 and GTO-219; position 2, CHIP-23 and CHIP-24; position 3, CHIP-19 and CHIP-20; position 4, CHIP-35 and CHIP-36; position 5, CHIP-44 and CHIP-45; position 6, CHIP-5 and CHIP-6; position 7, CHIP-31 and CHIP-32. The sequences of these primers are shown in Table S2. Real-time PCR was performed on a BioRad CFX96 system, using a QuantiTect SYBR Green PCR Kit from Qiagen according to the manufacturer instructions. The reverse-transcription step was performed at 50 °C for 20 min using single primers to generate strand-specific cDNAs. The following primers were used to detect the antisense transcript shown in Fig. 4: GTO-218, CHIP-23, CHIP-19, CHIP-35, CHIP-44, CHIP-5, and CHIP-31. Following reverse transcription, the samples were heated at 95 °C for 15 min, and subjected to 40 cycles of (94 °C for 15 s, 55 °C for 30 s, and 72 °C for 30 s). All reactions were set up in triplicate, and the melting curve of all PCR products was determined after amplification. Fourfold dilution series of genomic DNA were used to determine primer efficiencies and the exponential range of amplification for each primer pairs. All experiments were performed in this range. Mean normalized expression (MNE) values for transcript levels were calculated according to Eq. 1 and SEs on the mean (SEMNE) according to Eq. 2 (65).
where Eref is the efficiency of act1 amplification and CTref,mean is its mean cycle threshold; Etarget is the efficiency of amplification of the transcript of interest and CTtarget,mean is its mean cycle threshold. To compare the expression of (EcoRV)::ade6+ to ade6+ at its native chromosomal location, an additional normalization was performed by estimating the ade6+ to act1+ expression ratio in a wild-type strain (968; Fig. 3C).
Supplementary Material
Acknowledgments
We thank Pablo Hernandez, the Yeast Genetic Resource Center (Osaka University), and the RIKEN Bioresource Center for materials, and Stanley Brown, Laerke Holm, Sandeep Krishna, and Julie Køhler for their comments on the manuscript. Our research was supported by the Lundbeck Foundation, the Danish Research Council, the Copenhagen University Center of Excellence MolPhysX, and the Danish National Research Foundation (Center for Models of Life).
Footnotes
The authors declare no conflict of interest.
This article is a PNAS Direct Submission.
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1315581110/-/DCSupplemental.
References
- 1.Comings DE. Arrangement of chromatin in the nucleus. Hum Genet. 1980;53(2):131–143. doi: 10.1007/BF00273484. [DOI] [PubMed] [Google Scholar]
- 2.Marshall WF, Fung JC, Sedat JW. Deconstructing the nucleus: Global architecture from local interactions. Curr Opin Genet Dev. 1997;7(2):259–263. doi: 10.1016/s0959-437x(97)80136-0. [DOI] [PubMed] [Google Scholar]
- 3.Mekhail K, Moazed D. The nuclear envelope in genome organization, expression and stability. Nat Rev Mol Cell Biol. 2010;11(5):317–328. doi: 10.1038/nrm2894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Ritland Politz JC, Scalzo D, Groudine M. Something silent this way forms: The functional organization of the nuclear repressive compartment. Annu Rev Cell Dev Biol. 2013;29:241−270. doi: 10.1146/annurev-cellbio-101512-122317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Németh A, Längst G. Genome organization in and around the nucleolus. Trends Genet. 2011;27(4):149–156. doi: 10.1016/j.tig.2011.01.002. [DOI] [PubMed] [Google Scholar]
- 6.van Koningsbruggen S, et al. High-resolution whole-genome sequencing reveals that specific chromatin domains from most human chromosomes associate with nucleoli. Mol Biol Cell. 2010;21(21):3735–3748. doi: 10.1091/mbc.E10-06-0508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Németh A, et al. Initial genomics of the human nucleolus. PLoS Genet. 2010;6(3):e1000889. doi: 10.1371/journal.pgen.1000889. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Csink AK, Henikoff S. Large-scale chromosomal movements during interphase progression in Drosophila. J Cell Biol. 1998;143(1):13–22. doi: 10.1083/jcb.143.1.13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Zhang LF, Huynh KD, Lee JT. Perinucleolar targeting of the inactive X during S phase: Evidence for a role in the maintenance of silencing. Cell. 2007;129(4):693–706. doi: 10.1016/j.cell.2007.03.036. [DOI] [PubMed] [Google Scholar]
- 10.Thon G, Klar AJS. The clr1 locus regulates the expression of the cryptic mating-type loci of fission yeast. Genetics. 1992;131(2):287–296. doi: 10.1093/genetics/131.2.287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Allshire RC, Javerzat JP, Redhead NJ, Cranston G. Position effect variegation at fission yeast centromeres. Cell. 1994;76(1):157–169. doi: 10.1016/0092-8674(94)90180-5. [DOI] [PubMed] [Google Scholar]
- 12.Nimmo ER, Cranston G, Allshire RC. Telomere-associated chromosome breakage in fission yeast results in variegated expression of adjacent genes. EMBO J. 1994;13(16):3801–3811. doi: 10.1002/j.1460-2075.1994.tb06691.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Thon G, Verhein-Hansen J. Four chromo-domain proteins of Schizosaccharomyces pombe differentially repress transcription at various chromosomal locations. Genetics. 2000;155(2):551–568. doi: 10.1093/genetics/155.2.551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Thon G, Bjerling P, Bünner CM, Verhein-Hansen J. Expression-state boundaries in the mating-type region of fission yeast. Genetics. 2002;161(2):611–622. doi: 10.1093/genetics/161.2.611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Noma K, Allis CD, Grewal SI. Transitions in distinct histone H3 methylation patterns at the heterochromatin domain boundaries. Science. 2001;293(5532):1150–1155. doi: 10.1126/science.1064150. [DOI] [PubMed] [Google Scholar]
- 16.Cam HP, et al. Comprehensive analysis of heterochromatin- and RNAi-mediated epigenetic control of the fission yeast genome. Nat Genet. 2005;37(8):809–819. doi: 10.1038/ng1602. [DOI] [PubMed] [Google Scholar]
- 17.Funabiki H, Hagan I, Uzawa S, Yanagida M. Cell cycle-dependent specific positioning and clustering of centromeres and telomeres in fission yeast. J Cell Biol. 1993;121(5):961–976. doi: 10.1083/jcb.121.5.961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Tanizawa H, et al. Mapping of long-range associations throughout the fission yeast genome reveals global genome organization linked to transcriptional regulation. Nucleic Acids Res. 2010;38(22):8164–8177. doi: 10.1093/nar/gkq955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Molnar M, Kleckner N. Examination of interchromosomal interactions in vegetatively growing diploid Schizosaccharomyces pombe cells by Cre/loxP site-specific recombination. Genetics. 2008;178(1):99–112. doi: 10.1534/genetics.107.082826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Steglich B, Filion GJ, van Steensel B, Ekwall K. The inner nuclear membrane proteins Man1 and Ima1 link to two different types of chromatin at the nuclear periphery in S. pombe. Nucleus. 2012;3(1):77–87. doi: 10.4161/nucl.18825. [DOI] [PubMed] [Google Scholar]
- 21.Noma K, Cam HP, Maraia RJ, Grewal SIS. A role for TFIIIC transcription factor complex in genome organization. Cell. 2006;125(5):859–872. doi: 10.1016/j.cell.2006.04.028. [DOI] [PubMed] [Google Scholar]
- 22.Alfredsson-Timmins J, Henningson F, Bjerling P. The Clr4 methyltransferase determines the subnuclear localization of the mating-type region in fission yeast. J Cell Sci. 2007;120(Pt 11):1935–1943. doi: 10.1242/jcs.03457. [DOI] [PubMed] [Google Scholar]
- 23.Alfredsson-Timmins J, Kristell C, Henningson F, Lyckman S, Bjerling P. Reorganization of chromatin is an early response to nitrogen starvation in Schizosaccharomyces pombe. Chromosoma. 2009;118(1):99–112. doi: 10.1007/s00412-008-0180-6. [DOI] [PubMed] [Google Scholar]
- 24.Matsuyama A, et al. ORFeome cloning and global analysis of protein localization in the fission yeast Schizosaccharomyces pombe. Nat Biotechnol. 2006;24(7):841–847. doi: 10.1038/nbt1222. [DOI] [PubMed] [Google Scholar]
- 25.Umesono K, Hiraoka Y, Toda T, Yanagida M. Visualization of chromosomes in mitotically arrested cells of the fission yeast Schizosaccharomyces pombe. Curr Genet. 1983;7:123–128. doi: 10.1007/BF00365637. [DOI] [PubMed] [Google Scholar]
- 26.Creamer KM, Partridge JF. RITS-connecting transcription, RNA interference, and heterochromatin assembly in fission yeast. Wiley Interdiscip Rev RNA. 2011;2(5):632–646. doi: 10.1002/wrna.80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Kitano E, Hayashi A, Kanai D, Shinmyozu K, Nakayama J. Roles of fission yeast Grc3 protein in ribosomal RNA processing and heterochromatic gene silencing. J Biol Chem. 2011;286(17):15391–15402. doi: 10.1074/jbc.M110.201343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Woolcock KJ, et al. RNAi keeps Atf1-bound stress response genes in check at nuclear pores. Genes Dev. 2012;26(7):683–692. doi: 10.1101/gad.186866.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Kawakami K, Hayashi A, Nakayama J, Murakami Y. A novel RNAi protein, Dsh1, assembles RNAi machinery on chromatin to amplify heterochromatic siRNA. Genes Dev. 2012;26(16):1811–1824. doi: 10.1101/gad.190272.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Hansen KR, et al. H3K9me-independent gene silencing in fission yeast heterochromatin by Clr5 and histone deacetylases. PLoS Genet. 2011;7(1):e1001268. doi: 10.1371/journal.pgen.1001268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Singh SK, Sabatinos S, Forsburg S, Bastia D. Regulation of replication termination by Reb1 protein-mediated action at a distance. Cell. 2010;142(6):868–878. doi: 10.1016/j.cell.2010.08.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Thon G, Bjerling KP, Nielsen IS. Localization and properties of a silencing element near the mat3-M mating-type cassette of Schizosaccharomyces pombe. Genetics. 1999;151(3):945–963. doi: 10.1093/genetics/151.3.945. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Sanchez JA, Kim SM, Huberman JA. Ribosomal DNA replication in the fission yeast, Schizosaccharomyces pombe. Exp Cell Res. 1998;238(1):220–230. doi: 10.1006/excr.1997.3835. [DOI] [PubMed] [Google Scholar]
- 34.Sánchez-Gorostiaga A, López-Estraño C, Krimer DB, Schvartzman JB, Hernández P. Transcription termination factor reb1p causes two replication fork barriers at its cognate sites in fission yeast ribosomal DNA in vivo. Mol Cell Biol. 2004;24(1):398–406. doi: 10.1128/MCB.24.1.398-406.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Krings G, Bastia D. swi1- and swi3-dependent and independent replication fork arrest at the ribosomal DNA of Schizosaccharomyces pombe. Proc Natl Acad Sci USA. 2004;101(39):14085–14090. doi: 10.1073/pnas.0406037101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Arcangioli B. A site- and strand-specific DNA break confers asymmetric switching potential in fission yeast. EMBO J. 1998;17(15):4503–4510. doi: 10.1093/emboj/17.15.4503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Thon G, Klar AJS. Directionality of fission yeast mating-type interconversion is controlled by the location of the donor loci. Genetics. 1993;134(4):1045–1054. doi: 10.1093/genetics/134.4.1045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Jia S, Yamada T, Grewal SIS. Heterochromatin regulates cell type-specific long-range chromatin interactions essential for directed recombination. Cell. 2004;119(4):469–480. doi: 10.1016/j.cell.2004.10.020. [DOI] [PubMed] [Google Scholar]
- 39.McStay B, Grummt I. The epigenetics of rRNA genes: From molecular to chromosome biology. Annu Rev Cell Dev Biol. 2008;24:131–157. doi: 10.1146/annurev.cellbio.24.110707.175259. [DOI] [PubMed] [Google Scholar]
- 40.Bryk M, et al. Transcriptional silencing of Ty1 elements in the RDN1 locus of yeast. Genes Dev. 1997;11(2):255–269. doi: 10.1101/gad.11.2.255. [DOI] [PubMed] [Google Scholar]
- 41.Smith JS, Boeke JD. An unusual form of transcriptional silencing in yeast ribosomal DNA. Genes Dev. 1997;11(2):241–254. doi: 10.1101/gad.11.2.241. [DOI] [PubMed] [Google Scholar]
- 42.Gottlieb S, Esposito RE. A new role for a yeast transcriptional silencer gene, SIR2, in regulation of recombination in ribosomal DNA. Cell. 1989;56(5):771–776. doi: 10.1016/0092-8674(89)90681-8. [DOI] [PubMed] [Google Scholar]
- 43.Straight AF, et al. Net1, a Sir2-associated nucleolar protein required for rDNA silencing and nucleolar integrity. Cell. 1999;97(2):245–256. doi: 10.1016/s0092-8674(00)80734-5. [DOI] [PubMed] [Google Scholar]
- 44.Huang J, et al. Inhibition of homologous recombination by a cohesin-associated clamp complex recruited to the rDNA recombination enhancer. Genes Dev. 2006;20(20):2887–2901. doi: 10.1101/gad.1472706. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Torres-Rosell J, et al. The Smc5-Smc6 complex and SUMO modification of Rad52 regulates recombinational repair at the ribosomal gene locus. Nat Cell Biol. 2007;9(8):923–931. doi: 10.1038/ncb1619. [DOI] [PubMed] [Google Scholar]
- 46.Mekhail K, Seebacher J, Gygi SP, Moazed D. Role for perinuclear chromosome tethering in maintenance of genome stability. Nature. 2008;456(7222):667–670. doi: 10.1038/nature07460. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Peng JC, Karpen GH. H3K9 methylation and RNA interference regulate nucleolar organization and repeated DNA stability. Nat Cell Biol. 2007;9(1):25–35. doi: 10.1038/ncb1514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Stahl A, Hartung M, Vagner-Capodano AM, Fouet C. Chromosomal constitution of nucleolus-associated chromatin in man. Hum Genet. 1976;35(1):27–34. doi: 10.1007/BF00295616. [DOI] [PubMed] [Google Scholar]
- 49.Manuelidis L, Borden J. Reproducible compartmentalization of individual chromosome domains in human CNS cells revealed by in situ hybridization and three-dimensional reconstruction. Chromosoma. 1988;96(6):397–410. doi: 10.1007/BF00303033. [DOI] [PubMed] [Google Scholar]
- 50.Sullivan GJ, et al. Human acrocentric chromosomes with transcriptionally silent nucleolar organizer regions associate with nucleoli. EMBO J. 2001;20(11):2867–2874. doi: 10.1093/emboj/20.11.2867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Anastassova-Kristeva M. The nucleolar cycle in man. J Cell Sci. 1977;25:103–110. doi: 10.1242/jcs.25.1.103. [DOI] [PubMed] [Google Scholar]
- 52.Ding DQ, Yamamoto A, Haraguchi T, Hiraoka Y. Dynamics of homologous chromosome pairing during meiotic prophase in fission yeast. Dev Cell. 2004;6(3):329–341. doi: 10.1016/s1534-5807(04)00059-0. [DOI] [PubMed] [Google Scholar]
- 53.Zhao A, Guo A, Liu Z, Pape L. Molecular cloning and analysis of Schizosaccharomyces pombe Reb1p: Sequence-specific recognition of two sites in the far upstream rDNA intergenic spacer. Nucleic Acids Res. 1997;25(4):904–910. doi: 10.1093/nar/25.4.904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Melekhovets YF, Shwed PS, Nazar RN. In vivo analyses of RNA polymerase I termination in Schizosaccharomyces pombe. Nucleic Acids Res. 1997;25(24):5103–5109. doi: 10.1093/nar/25.24.5103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Rhind N, et al. Comparative functional genomics of the fission yeasts. Science. 2011;332(6032):930–936. doi: 10.1126/science.1203357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Win TZ, et al. Requirement of fission yeast Cid14 in polyadenylation of rRNAs. Mol Cell Biol. 2006;26(5):1710–1721. doi: 10.1128/MCB.26.5.1710-1721.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Buck SW, Sandmeier JJ, Smith JS. RNA polymerase I propagates unidirectional spreading of rDNA silent chromatin. Cell. 2002;111(7):1003–1014. doi: 10.1016/s0092-8674(02)01193-5. [DOI] [PubMed] [Google Scholar]
- 58.Cioci F, et al. Silencing in yeast rDNA chromatin: Reciprocal relationship in gene expression between RNA polymerase I and II. Mol Cell. 2003;12(1):135–145. doi: 10.1016/s1097-2765(03)00262-4. [DOI] [PubMed] [Google Scholar]
- 59.Schmitz KM, Mayer C, Postepska A, Grummt I. Interaction of noncoding RNA with the rDNA promoter mediates recruitment of DNMT3b and silencing of rRNA genes. Genes Dev. 2010;24(20):2264–2269. doi: 10.1101/gad.590910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Mohammad F, et al. Kcnq1ot1/Lit1 noncoding RNA mediates transcriptional silencing by targeting to the perinucleolar region. Mol Cell Biol. 2008;28(11):3713–3728. doi: 10.1128/MCB.02263-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Pandey RR, et al. Kcnq1ot1 antisense noncoding RNA mediates lineage-specific transcriptional silencing through chromatin-level regulation. Mol Cell. 2008;32(2):232–246. doi: 10.1016/j.molcel.2008.08.022. [DOI] [PubMed] [Google Scholar]
- 62.Fedoriw AM, Starmer J, Yee D, Magnuson T. Nucleolar association and transcriptional inhibition through 5S rDNA in mammals. PLoS Genet. 2012;8(1):e1002468. doi: 10.1371/journal.pgen.1002468. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Leupold U. Die Vererbung von Homothallie und Heterothallie bei Schizosaccharomyces pombe. C R Trav Lab Carlsberg SQr Physiol. 1950;24:381–480. [Google Scholar]
- 64.Lyne R, et al. Whole-genome microarrays of fission yeast: Characteristics, accuracy, reproducibility, and processing of array data. BMC Genomics. 2003;4(1):27. doi: 10.1186/1471-2164-4-27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Simon P. Q-Gene: Processing quantitative real-time RT-PCR data. Bioinformatics. 2003;19(11):1439–1440. doi: 10.1093/bioinformatics/btg157. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.