Abstract
Crossing-over between the dispersed repeated sequences found in eukaryotic genomes would generate chromosomal rearrangements. The stability of the yeast genome suggests the existence of some constraint on the ability of these sequences to interact by recombination. We have constructed strains with two alleles of the his3 gene located on different chromosomes. Gene conversion accounts for the majority of the recombination events between these genes, but about 10% of the events are crossovers that result in a reciprocal translocation. When one of the alleles is on an autonomously replicating centromere plasmid, recombination is 5- to 10-fold more frequent than when both alleles are chromosomal, suggesting that higher-order chromosome structure may play a role in restricting interchromosomal recombination. We have also used the translocation to deduce the orientation of the his3 and rRNA genes relative to their centromeres.
Full text
PDF
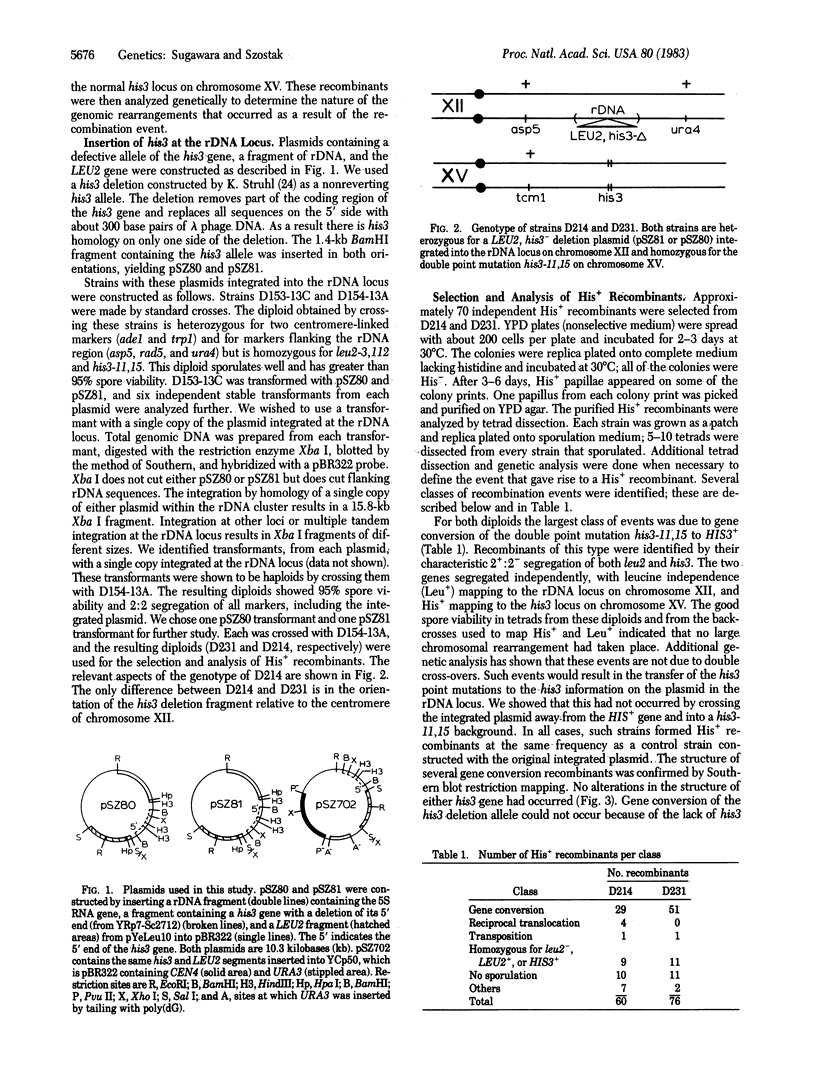


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Baltimore D. Gene conversion: some implications for immunoglobulin genes. Cell. 1981 Jun;24(3):592–594. doi: 10.1016/0092-8674(81)90082-9. [DOI] [PubMed] [Google Scholar]
- Bell G. I., DeGennaro L. J., Gelfand D. H., Bishop R. J., Valenzuela P., Rutter W. J. Ribosomal RNA genes of Saccharomyces cerevisiae. I. Physical map of the repeating unit and location of the regions coding for 5 S, 5.8 S, 18 S, and 25 S ribosomal RNAs. J Biol Chem. 1977 Nov 25;252(22):8118–8125. [PubMed] [Google Scholar]
- Cameron J. R., Loh E. Y., Davis R. W. Evidence for transposition of dispersed repetitive DNA families in yeast. Cell. 1979 Apr;16(4):739–751. doi: 10.1016/0092-8674(79)90090-4. [DOI] [PubMed] [Google Scholar]
- Chaleff D. T., Fink G. R. Genetic events associated with an insertion mutation in yeast. Cell. 1980 Aug;21(1):227–237. doi: 10.1016/0092-8674(80)90130-0. [DOI] [PubMed] [Google Scholar]
- Clark-Walker G. D., Azad A. A. Hybridizable sequences between cytoplasmic ribosomal RNAs and 3 micron circular DNAs of Saccharomyces cerevisiae and Torulopsis glabrata. Nucleic Acids Res. 1980 Mar 11;8(5):1009–1022. doi: 10.1093/nar/8.5.1009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Egel R. Intergenic conversion and reiterated genes. Nature. 1981 Mar 19;290(5803):191–192. doi: 10.1038/290191a0. [DOI] [PubMed] [Google Scholar]
- Ernst J. F., Stewart J. W., Sherman F. The cyc1-11 mutation in yeast reverts by recombination with a nonallelic gene: composite genes determining the iso-cytochromes c. Proc Natl Acad Sci U S A. 1981 Oct;78(10):6334–6338. doi: 10.1073/pnas.78.10.6334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Esposito M. S. Evidence that spontaneous mitotic recombination occurs at the two-strand stage. Proc Natl Acad Sci U S A. 1978 Sep;75(9):4436–4440. doi: 10.1073/pnas.75.9.4436. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fogel S., Mortimer R., Lusnak K., Tavares F. Meiotic gene conversion: a signal of the basic recombination event in yeast. Cold Spring Harb Symp Quant Biol. 1979;43(Pt 2):1325–1341. doi: 10.1101/sqb.1979.043.01.152. [DOI] [PubMed] [Google Scholar]
- Hawthorne D. C., Mortimer R. K. Genetic mapping of nonsense suppressors in yeast. Genetics. 1968 Dec;60(4):735–742. doi: 10.1093/genetics/60.4.735. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hereford L., Fahrner K., Woolford J., Jr, Rosbash M., Kaback D. B. Isolation of yeast histone genes H2A and H2B. Cell. 1979 Dec;18(4):1261–1271. doi: 10.1016/0092-8674(79)90237-x. [DOI] [PubMed] [Google Scholar]
- Holland J. P., Holland M. J. Structural comparison of two nontandemly repeated yeast glyceraldehyde-3-phosphate dehydrogenase genes. J Biol Chem. 1980 Mar 25;255(6):2596–2605. [PubMed] [Google Scholar]
- Holland M. J., Holland J. P., Thill G. P., Jackson K. A. The primary structures of two yeast enolase genes. Homology between the 5' noncoding flanking regions of yeast enolase and glyceraldehyde-3-phosphate dehydrogenase genes. J Biol Chem. 1981 Feb 10;256(3):1385–1395. [PubMed] [Google Scholar]
- Jackson J. A., Fink G. R. Gene conversion between duplicated genetic elements in yeast. Nature. 1981 Jul 23;292(5821):306–311. doi: 10.1038/292306a0. [DOI] [PubMed] [Google Scholar]
- Mikus M. D., Petes T. D. Recombination between genes located on nonhomologous chromosomes in Saccharomyces cerevisiae. Genetics. 1982 Jul-Aug;101(3-4):369–404. doi: 10.1093/genetics/101.3-4.369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olson M. V., Montgomery D. L., Hopper A. K., Page G. S., Horodyski F., Hall B. D. Molecular characterisation of the tyrosine tRNA genes of yeast. Nature. 1977 Jun 16;267(5612):639–641. doi: 10.1038/267639a0. [DOI] [PubMed] [Google Scholar]
- Orr-Weaver T. L., Szostak J. W., Rothstein R. J. Yeast transformation: a model system for the study of recombination. Proc Natl Acad Sci U S A. 1981 Oct;78(10):6354–6358. doi: 10.1073/pnas.78.10.6354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Potier S., Winsor B., Lacroute F. Genetic selection for reciprocal translocation at chosen chromosomal sites in Saccharomyces cerevisiae. Mol Cell Biol. 1982 Sep;2(9):1025–1032. doi: 10.1128/mcb.2.9.1025. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scherer S., Davis R. W. Recombination of dispersed repeated DNA sequences in yeast. Science. 1980 Sep 19;209(4463):1380–1384. doi: 10.1126/science.6251545. [DOI] [PubMed] [Google Scholar]
- Sherman F., Helms C. A chromosomal translocation causing overproduction of iso-2-cytochrome c in yeast. Genetics. 1978 Apr;88(4 Pt 1):689–707. [PMC free article] [PubMed] [Google Scholar]
- Struhl K., Davis R. W. Promotor mutants of the yeast his3 gene. J Mol Biol. 1981 Nov 5;152(3):553–568. doi: 10.1016/0022-2836(81)90268-0. [DOI] [PubMed] [Google Scholar]
- Struhl K., Davis R. W. Transcription of the his3 gene region in Saccharomyces cerevisiae. J Mol Biol. 1981 Nov 5;152(3):535–552. doi: 10.1016/0022-2836(81)90267-9. [DOI] [PubMed] [Google Scholar]
- Sugawara N., Szostak J. W. Construction of specific chromosomal rearrangements in yeast. Methods Enzymol. 1983;101:269–278. doi: 10.1016/0076-6879(83)01020-4. [DOI] [PubMed] [Google Scholar]
- Zamb T. J., Petes T. D. Analysis of the junction between ribosomal RNA genes and single-copy chromosomal sequences in the yeast Saccharomyces cerevisiae. Cell. 1982 Feb;28(2):355–364. doi: 10.1016/0092-8674(82)90353-1. [DOI] [PubMed] [Google Scholar]
- del Rey F. J., Donahue T. F., Fink G. R. sigma, a repetitive element found adjacent to tRNA genes of yeast. Proc Natl Acad Sci U S A. 1982 Jul;79(13):4138–4142. doi: 10.1073/pnas.79.13.4138. [DOI] [PMC free article] [PubMed] [Google Scholar]