Abstract
Boundary or insulator elements set up independent territories of gene activity by establishing higher order domains of chromatin structure. The gypsy retrotransposon of Drosophila contains an insulator element that represses enhancer-promoter interactions and is responsible for the mutant phenotypes caused by insertion of this element. The gypsy insulator inhibits the interaction of promoter-distal enhancers with the transcription complex without affecting the functionality of promoter-proximal enhancers; in addition, these sequences can buffer a transgene from chromosomal position effects. Two proteins have been identified that bind gypsy insulator sequences and are responsible for their effects on transcription. The suppressor of Hairy-wing [su(Hw)] protein affects enhancer function both upstream and downstream of its binding site by causing a silencing effect similar to that of heterochromatin. The modifier of mdg4 [mod(mdg4)] protein interacts with su(Hw) to transform this bi-directional repression into the polar effect characteristic of insulators. These effects seem to be modulated by changes in chromatin structure.
Full text
PDF

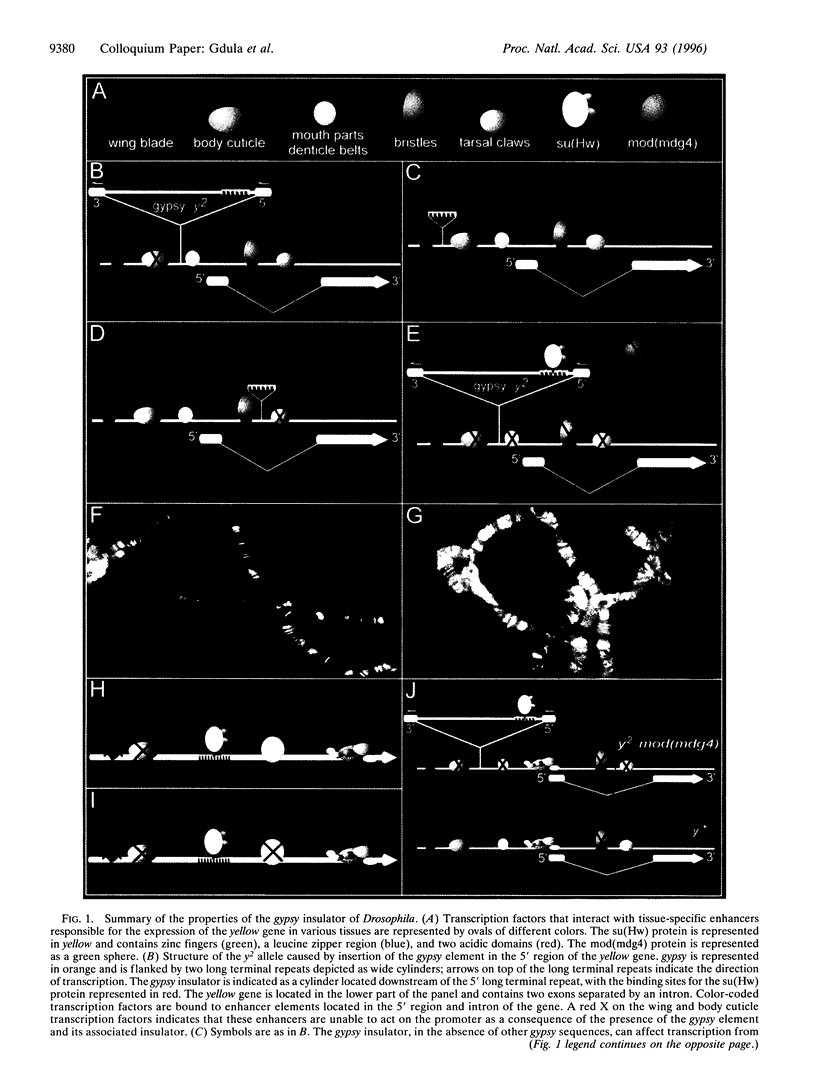



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bone J. R., Lavender J., Richman R., Palmer M. J., Turner B. M., Kuroda M. I. Acetylated histone H4 on the male X chromosome is associated with dosage compensation in Drosophila. Genes Dev. 1994 Jan;8(1):96–104. doi: 10.1101/gad.8.1.96. [DOI] [PubMed] [Google Scholar]
- Bonifer C., Vidal M., Grosveld F., Sippel A. E. Tissue specific and position independent expression of the complete gene domain for chicken lysozyme in transgenic mice. EMBO J. 1990 Sep;9(9):2843–2848. doi: 10.1002/j.1460-2075.1990.tb07473.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cai H., Levine M. Modulation of enhancer-promoter interactions by insulators in the Drosophila embryo. Nature. 1995 Aug 10;376(6540):533–536. doi: 10.1038/376533a0. [DOI] [PubMed] [Google Scholar]
- Chung J. H., Whiteley M., Felsenfeld G. A 5' element of the chicken beta-globin domain serves as an insulator in human erythroid cells and protects against position effect in Drosophila. Cell. 1993 Aug 13;74(3):505–514. doi: 10.1016/0092-8674(93)80052-g. [DOI] [PubMed] [Google Scholar]
- Corces V. G., Geyer P. K. Interactions of retrotransposons with the host genome: the case of the gypsy element of Drosophila. Trends Genet. 1991 Mar;7(3):86–90. doi: 10.1016/0168-9525(91)90277-W. [DOI] [PubMed] [Google Scholar]
- Dorn R., Krauss V., Reuter G., Saumweber H. The enhancer of position-effect variegation of Drosophila, E(var)3-93D, codes for a chromatin protein containing a conserved domain common to several transcriptional regulators. Proc Natl Acad Sci U S A. 1993 Dec 1;90(23):11376–11380. doi: 10.1073/pnas.90.23.11376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dorsett D. Potentiation of a polyadenylylation site by a downstream protein-DNA interaction. Proc Natl Acad Sci U S A. 1990 Jun;87(11):4373–4377. doi: 10.1073/pnas.87.11.4373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dreesen T. D., Henikoff S., Loughney K. A pairing-sensitive element that mediates trans-inactivation is associated with the Drosophila brown gene. Genes Dev. 1991 Mar;5(3):331–340. doi: 10.1101/gad.5.3.331. [DOI] [PubMed] [Google Scholar]
- Farkas G., Gausz J., Galloni M., Reuter G., Gyurkovics H., Karch F. The Trithorax-like gene encodes the Drosophila GAGA factor. Nature. 1994 Oct 27;371(6500):806–808. doi: 10.1038/371806a0. [DOI] [PubMed] [Google Scholar]
- Georgiev P. G., Corces V. G. The su(Hw) protein bound to gypsy sequences in one chromosome can repress enhancer-promoter interactions in the paired gene located in the other homolog. Proc Natl Acad Sci U S A. 1995 May 23;92(11):5184–5188. doi: 10.1073/pnas.92.11.5184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Georgiev P. G., Gerasimova T. I. Novel genes influencing the expression of the yellow locus and mdg4 (gypsy) in Drosophila melanogaster. Mol Gen Genet. 1989 Dec;220(1):121–126. doi: 10.1007/BF00260865. [DOI] [PubMed] [Google Scholar]
- Georgiev P., Kozycina M. Interaction between mutations in the suppressor of Hairy wing and modifier of mdg4 genes of Drosophila melanogaster affecting the phenotype of gypsy-induced mutations. Genetics. 1996 Feb;142(2):425–436. doi: 10.1093/genetics/142.2.425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gerasimova T. I., Corces V. G. Boundary and insulator elements in chromosomes. Curr Opin Genet Dev. 1996 Apr;6(2):185–192. doi: 10.1016/s0959-437x(96)80049-9. [DOI] [PubMed] [Google Scholar]
- Gerasimova T. I., Gdula D. A., Gerasimov D. V., Simonova O., Corces V. G. A Drosophila protein that imparts directionality on a chromatin insulator is an enhancer of position-effect variegation. Cell. 1995 Aug 25;82(4):587–597. doi: 10.1016/0092-8674(95)90031-4. [DOI] [PubMed] [Google Scholar]
- Geyer P. K., Corces V. G. DNA position-specific repression of transcription by a Drosophila zinc finger protein. Genes Dev. 1992 Oct;6(10):1865–1873. doi: 10.1101/gad.6.10.1865. [DOI] [PubMed] [Google Scholar]
- Geyer P. K., Corces V. G. DNA position-specific repression of transcription by a Drosophila zinc finger protein. Genes Dev. 1992 Oct;6(10):1865–1873. doi: 10.1101/gad.6.10.1865. [DOI] [PubMed] [Google Scholar]
- Geyer P. K., Green M. M., Corces V. G. Mutant gene phenotypes mediated by a Drosophila melanogaster retrotransposon require sequences homologous to mammalian enhancers. Proc Natl Acad Sci U S A. 1988 Nov;85(22):8593–8597. doi: 10.1073/pnas.85.22.8593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geyer P. K., Spana C., Corces V. G. On the molecular mechanism of gypsy-induced mutations at the yellow locus of Drosophila melanogaster. EMBO J. 1986 Oct;5(10):2657–2662. doi: 10.1002/j.1460-2075.1986.tb04548.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grosveld F., van Assendelft G. B., Greaves D. R., Kollias G. Position-independent, high-level expression of the human beta-globin gene in transgenic mice. Cell. 1987 Dec 24;51(6):975–985. doi: 10.1016/0092-8674(87)90584-8. [DOI] [PubMed] [Google Scholar]
- Harrison D. A., Gdula D. A., Coyne R. S., Corces V. G. A leucine zipper domain of the suppressor of Hairy-wing protein mediates its repressive effect on enhancer function. Genes Dev. 1993 Oct;7(10):1966–1978. doi: 10.1101/gad.7.10.1966. [DOI] [PubMed] [Google Scholar]
- Holdridge C., Dorsett D. Repression of hsp70 heat shock gene transcription by the suppressor of hairy-wing protein of Drosophila melanogaster. Mol Cell Biol. 1991 Apr;11(4):1894–1900. doi: 10.1128/mcb.11.4.1894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jack J., Dorsett D., Delotto Y., Liu S. Expression of the cut locus in the Drosophila wing margin is required for cell type specification and is regulated by a distant enhancer. Development. 1991 Nov;113(3):735–747. doi: 10.1242/dev.113.3.735. [DOI] [PubMed] [Google Scholar]
- Kalos M., Fournier R. E. Position-independent transgene expression mediated by boundary elements from the apolipoprotein B chromatin domain. Mol Cell Biol. 1995 Jan;15(1):198–207. doi: 10.1128/mcb.15.1.198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kellum R., Schedl P. A group of scs elements function as domain boundaries in an enhancer-blocking assay. Mol Cell Biol. 1992 May;12(5):2424–2431. doi: 10.1128/mcb.12.5.2424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kellum R., Schedl P. A position-effect assay for boundaries of higher order chromosomal domains. Cell. 1991 Mar 8;64(5):941–950. doi: 10.1016/0092-8674(91)90318-s. [DOI] [PubMed] [Google Scholar]
- Levis R., Hazelrigg T., Rubin G. M. Effects of genomic position on the expression of transduced copies of the white gene of Drosophila. Science. 1985 Aug 9;229(4713):558–561. doi: 10.1126/science.2992080. [DOI] [PubMed] [Google Scholar]
- Mazo A. M., Mizrokhi L. J., Karavanov A. A., Sedkov Y. A., Krichevskaja A. A., Ilyin Y. V. Suppression in Drosophila: su(Hw) and su(f) gene products interact with a region of gypsy (mdg4) regulating its transcriptional activity. EMBO J. 1989 Mar;8(3):903–911. doi: 10.1002/j.1460-2075.1989.tb03451.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Modolell J., Bender W., Meselson M. Drosophila melanogaster mutations suppressible by the suppressor of Hairy-wing are insertions of a 7.3-kilobase mobile element. Proc Natl Acad Sci U S A. 1983 Mar;80(6):1678–1682. doi: 10.1073/pnas.80.6.1678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pazin M. J., Kamakaka R. T., Kadonaga J. T. ATP-dependent nucleosome reconfiguration and transcriptional activation from preassembled chromatin templates. Science. 1994 Dec 23;266(5193):2007–2011. doi: 10.1126/science.7801129. [DOI] [PubMed] [Google Scholar]
- Peifer M., Bender W. Sequences of the gypsy transposon of Drosophila necessary for its effects on adjacent genes. Proc Natl Acad Sci U S A. 1988 Dec;85(24):9650–9654. doi: 10.1073/pnas.85.24.9650. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roseman R. R., Pirrotta V., Geyer P. K. The su(Hw) protein insulates expression of the Drosophila melanogaster white gene from chromosomal position-effects. EMBO J. 1993 Feb;12(2):435–442. doi: 10.1002/j.1460-2075.1993.tb05675.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roseman R. R., Swan J. M., Geyer P. K. A Drosophila insulator protein facilitates dosage compensation of the X chromosome min-white gene located at autosomal insertion sites. Development. 1995 Nov;121(11):3573–3582. doi: 10.1242/dev.121.11.3573. [DOI] [PubMed] [Google Scholar]
- Scott K. S., Geyer P. K. Effects of the su(Hw) insulator protein on the expression of the divergently transcribed Drosophila yolk protein genes. EMBO J. 1995 Dec 15;14(24):6258–6267. doi: 10.1002/j.1460-2075.1995.tb00316.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith P. A., Corces V. G. The suppressor of Hairy-wing protein regulates the tissue-specific expression of the Drosophila gypsy retrotransposon. Genetics. 1995 Jan;139(1):215–228. doi: 10.1093/genetics/139.1.215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spana C., Corces V. G. DNA bending is a determinant of binding specificity for a Drosophila zinc finger protein. Genes Dev. 1990 Sep;4(9):1505–1515. doi: 10.1101/gad.4.9.1505. [DOI] [PubMed] [Google Scholar]
- Spana C., Harrison D. A., Corces V. G. The Drosophila melanogaster suppressor of Hairy-wing protein binds to specific sequences of the gypsy retrotransposon. Genes Dev. 1988 Nov;2(11):1414–1423. doi: 10.1101/gad.2.11.1414. [DOI] [PubMed] [Google Scholar]
- Tsukiyama T., Becker P. B., Wu C. ATP-dependent nucleosome disruption at a heat-shock promoter mediated by binding of GAGA transcription factor. Nature. 1994 Feb 10;367(6463):525–532. doi: 10.1038/367525a0. [DOI] [PubMed] [Google Scholar]
- Udvardy A., Maine E., Schedl P. The 87A7 chromomere. Identification of novel chromatin structures flanking the heat shock locus that may define the boundaries of higher order domains. J Mol Biol. 1985 Sep 20;185(2):341–358. doi: 10.1016/0022-2836(85)90408-5. [DOI] [PubMed] [Google Scholar]
- Vazquez J., Schedl P. Sequences required for enhancer blocking activity of scs are located within two nuclease-hypersensitive regions. EMBO J. 1994 Dec 15;13(24):5984–5993. doi: 10.1002/j.1460-2075.1994.tb06944.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wallrath L. L., Elgin S. C. Position effect variegation in Drosophila is associated with an altered chromatin structure. Genes Dev. 1995 May 15;9(10):1263–1277. doi: 10.1101/gad.9.10.1263. [DOI] [PubMed] [Google Scholar]
- Zollman S., Godt D., Privé G. G., Couderc J. L., Laski F. A. The BTB domain, found primarily in zinc finger proteins, defines an evolutionarily conserved family that includes several developmentally regulated genes in Drosophila. Proc Natl Acad Sci U S A. 1994 Oct 25;91(22):10717–10721. doi: 10.1073/pnas.91.22.10717. [DOI] [PMC free article] [PubMed] [Google Scholar]